1 Department of Gynecology, The Obstetrics and Gynecology Hospital of Fudan University, 200090 Shanghai, China
2 Shanghai Key Laboratory of Female Reproductive Endocrine-Related Diseases, The Obstetrics and Gynecology Hospital, Fudan University, 200011 Shanghai, China
Abstract
Background: Cervical cancer has high morbidity and intratumor heterogeneity. Anoikis, a form of programmed cell death preventing detached cancer cells from readhering, may serve as a potential prognostic signature for cervical cancer. This study aimed to assess the predictive performance of anoikis patterns in cervical cancer prognosis. Methods: Differentially expressed anoikis-related genes (DEARGs) were identified between normal and cancer samples using data from the Gene Expression Omnibus database with the elucidation of mutation status and bio-function. Novel anoikis molecular subtypes were defined in The Cancer Genome Atlas (TCGA) cohort with consensus clustering analysis. A multigene prognostic signature was constructed through least absolute shrinkage and selection operator (LASSO) Cox analysis with internal and external validation. The nomogram-based survival probability of cervical cancer over 3 and 5 years was predicted and assessed with calibration, receiver operating characteristic, decision curve analysis, and Kaplan-Meier curves. Additionally, mutation, function, and immune analysis were conducted among different risk groups. Results: We identified 77 DEARGs between normal and cervical cancer tissues and explored their mutation status and functions. The TCGA cohort could be categorized into two subtypes based on these genes. Furthermore, seven prognostic signature genes were constructed, and the nomogram involving DEARGs and clinicopathological characteristics showed satisfactory predictive performance. Functional analysis indicated that immune-related genes were enriched, and immune status, as well as sensitivity of chemotherapies and targeting drugs, were correlated with the risk model. Conclusions: Anoikis patterns play important roles in tumor immunity and can be used to predict the prognosis of cervical cancers.
Keywords
- cervical cancer
- anoikis
- molecular subtype
- prognosis
- immune response
- drug sensitivity
- somatic mutation
Cervical cancer (CC) is a common gynecological malignant tumor that poses a threat to women’s health globally [1, 2]. The widespread use of cervical screening programs and human papillomavirus (HPV) vaccines in recent years [3], has resulted in a reduction in the incidence of CC in developed countries [4]. Despite being one of the most preventable cancers, CC remains the second leading cause of premature cancer death among women aged 20–39 years in the United States [5], and a high burden in many low- and middle-income countries [6, 7, 8, 9, 10]. While HPV infections has been known to promote CC [11], its impact on CC prognosis is currently inconclusive and the FIGO (International Federation of Gynaecology and Obstetrics) staging system is still the main prognostic indicator [12]. Nevertheless, FIGO stage is inapplicable to the heterogeneity analysis of CC patients [13]. Therefore, the identification of subgroups of CC with accurate prognostic biomarkers and the development of predictive models are urgently needed for better treatment guidance.
Anoikis or matrix detachment-induced apoptosis, first introduced in 1993 by Meredith et al. [14], is a specific form of programmed apoptosis triggered by losing contact with their extracellular matrix (ECM) or neighboring cells. It plays a pivotal role in maintaining tissue homeostasis by preventing dislodged cells from readhering to other substrates for abnormal proliferation [15]. Cancer cells must develop anoikis resistance in order to survive under anchorage-independent conditions, such as in the blood or lymphatics circulation, before forming metastatic foci in distant organs [16, 17]. Metastasis is responsible for up to 90% of cancer-related deaths [18], and reducing metastasis-associated mortality remains a major clinical unmet need [19]. Therefore, understanding the mechanisms driving anoikis resistance may help to counteract tumor progression and prevent metastasis.
An increasing number of anoikis-related genes (ARGs) have been identified in various tumors [20, 21, 22, 23, 24]. Fonseca et al. [20] found that high expression levels of Plk4 induce anoikis resistance of breast cancer cells partially through P-cadherin upregulation. A previous study revealed that nuclear MYH9 promotes CTNNB1 transcription, which conferred resistance to anoikis in gastric cancer [21]. In a study by Jin et al. [22], GDH1-mediated metabolic reprogramming of glutaminolysis has been verified to promote anoikis resistance and tumor metastasis in LKB1-deficient lung cancer. In addition, ARGs have been proven to be associated with prognosis in several tumors through bioinformatic analyses [25, 26, 27, 28, 29, 30]. While several studies have identified that anoikis resistance exerts important functions in the oncogenesis of CC [31, 32, 33, 34], the comprehensive analyses of the implications of anoikis in CC are still limited.
In this study, differentially expressed ARGs (DEARGs) were screened between normal and CC tissues based on the datasets from the Gene Expression Omnibus (GEO) database with mutation and function analysis. Then, the subgroups of CC based on these DEARGs and a gene signature was identified in The Cancer Genome Atlas (TCGA) cohort. The prognostic value of the gene signature was validated in external cohort and its expression was validated through the Gene Expression Profiling Interactive Analysis (GEPIA). Nomogram analysis in combination with the DEARGs and other clinical characteristics for predicting the survival probability of CC patients in 3 and 5 years was conducted and assessed. We then performed mutation and function analysis among differentially expressed genes (DEGs) between low- and high-risk groups. The correlation between anoikis molecular subtypes and the immune microenvironment landscape of CC was also explored.
We retrieved expression profile microarray datasets from the GEO database (https://www.ncbi.nlm.nih.gov/geo/) with similar platforms, prioritizing tissue-based ones for normal cervix and CC analysis while excluding cell line based and other datasets. The final cohort included data from GSE6791 (the code number of cohort in GEO database, including 20 CC samples and 8 normal cervix samples with platform coded GPL570, similarly hereinafter), GSE7803 (21 CC samples and 10 normal cervix samples with platform GPL96), GSE9750 (33 CC samples and 24 normal cervix samples with platform GPL96), GSE39001 (19 CC samples and 5 normal cervix samples with platform GPL6244), GSE52903 (55 CC samples and 17 normal cervix samples with platform GPL6244), GSE63514 (28 CC samples and 24 normal cervix samples with platform GPL570), with a total of 264 samples (176 CC samples and 88 normal samples). We integrated expression profiles using the Combat function of the ‘sva’ R package (version 4.2.1, R: A language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria, similarly hereinafter) to remove batch effects [35, 36] and downloaded 801 ARGs from the GeneCards database (https://www.genecards.org/). DEARGs were analyzed when a given gene expression was detected across all six datasets.
TCGA database (https://portal.gdc.cancer.gov/) was used to obtain the RNA
sequencing, mutation, and clinical data of 304 cervical squamous cell carcinoma
and endocervical adenocarcinoma (CESC) samples for constructing prognostic
features. These data were pre-processed through the R packages and algorithms to
extract the expression profile matrix and the clinical and pathological
characteristics, including their survival status, survival times, CC stage and
grade, and the acceptance of radiotherapy. The samples with a follow-up time of
The DEARGs between CC and normal cervix tissues were identified by the ‘limma’
package, during which an empirical bayes technique was performed with six crucial
steps: building a gene expression matrix, building an experimental design matrix,
building a contrast matrix, fitting a linear model, Bayes test, and generating a
result report. To prevent the possibility of false-positive effects,
The consensus clustering analysis of the DEARGs in TCGA-CESC cohort was
performed with ‘ConsensusClusterPlus’ R package [37]. The Univariate Cox
regression analyses were performed on DEARGs for association with overall
survival (OS) [38]. The candidate prognostic DEARGs that might predict OS were
presented using a forest plot with a criterion of p
The R package ‘caret’ was used to randomly divide the 244 TCGA-CESC samples into
the training and testing sets at a 6:4 ratio. In the training cohort, a least
absolute shrinkage and selection operator (LASSO) Cox regression algorithm was
performed to determine the optimal prognostic DEARGs (penalty parameter, the
Subsequently, the best threshold point of ROC in training cohort was used to
stratify each patient in entire set into the high-risk group (HRG) and the
low-risk group (LRG). Therefore, the signature was validated with the testing set
and the entire TCGA dataset. Kaplan-Meier (KM) curve was generated to evaluate
the difference in OS between the HRG and LRG by using the ‘survival’ and
‘survminer’ R packages (p
ROC curves were drawn with the ‘pROC’ and ‘ggplot2’ R package to compare the discrimination performance of age, stage, and risk score calculated from the signature in predicting survival outcome. To predict the survival probability of CC patients in 3 and 5 years, a nomogram was generated according to the risk score and clinicopathologic factors (age and stage) by using the ‘rms’ and ‘survival’ packages. The prognostic accuracy of the nomogram was assessed by the Hosmer-Lemeshow calibration curves, the time-dependent ROC curves for 1-, 3-, and 5-years, and the decision curve analysis (DCA) curves in the entire TCGA dataset, which was achieved by the ‘rms’, ‘timeROC’, and ‘ggDCA’ R package. Finally, based on the best threshold point of ROC, the entire TCGA cohort was divided into HRG and LRG and the survival differences between two groups were illustrated using KM survival analysis.
The expression level of the prognostic DEARGs in prediction model were visualized as boxplot with the ‘ggplot2’ R package. Then the Spearman correlation coefficients among these DEARGs were calculated using the ‘corrplot’ R package. With the GEPIA public database (http://gepia.cancer-pku.cn/index.html) as external validation cohort (including 306 CESC samples and 13 normal cervix samples from TCGA database and The Genotype-Tissue Expression (GTEx) database), we reanalyzed the expression level of these DEARGs included in prognostic signature.
The somatic mutation data were obtained from TCGA and the mutation information was extracted in R software. The positions of the DEARGs between normal cervix and CC samples, and the DEGs between different risk groups were profiled through ‘RCircos’ R package [39]. Somatic variation of the DEARGs in the TCGA-CESC dataset was profiled using ‘maftools’ package. Simultaneously, the tumor mutation burden (TMB) of each sample was calculated with the ‘tmb’ function in ‘maftools’ package and demonstrated with waterfall plot and scatter diagram. Then, the ‘maftools’ package was also used for creating waterfall diagrams to show the distribution of top 15 genes with highest somatic mutation frequency in HRG and LRG categorized by final prognostic model [40].
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment
analysis were performed through the ‘clusterProfiler’, ‘GOplot’, and ‘ggthemes’ R
packages to elucidate the potential biological mechanism and functions of the
DEGs, which included the DEARGs between CC and normal cervix samples in the
cohort merged by 6 datasets from GEO database, and the DEGs between low- and
high-risk groups in TCGA-CESC dataset. The biological process, cellular
processes, and molecular functions were assessed respectively in GO analysis. The
functions or pathways with p
To investigate the relationship between the immune infiltration status and our prognostic model, single-sample gene set enrichment analysis (ssGSEA) algorithms was used to calculate the immune cell infiltration profile of the TCGA-CESC dataset with ‘limma’ and ‘GSVA’ packages. The results were presented as a boxplot to compare the different immune cell abundance between HRG and LRG. Furthermore, the Spearman correlation coefficients of risk scores calculated from the final prognostic model and the immune cells abundance/immune checkpoints were visualized using ‘corrplot’ R package.
Statistical analyses of this study were performed using the R program (version 4.2.1, R Foundation for Statistical Computing, Vienna, Austria). Single-factor analysis of variance was applied to compare the gene expression levels between the normal cervix and CC tissues, while the Pearson chi-square test was used to compare the categorical variables. The prognoses between different risk groups were compared using the KM analysis with a two-sided log-rank test. To assess the independent prognostic value of the risk model, we used univariate and multivariate Cox regression models. When comparing the immune cell infiltration between the two groups, the Mann–Whitney test was used.
The flowchart of this study is illustrated in Fig. 1A. We obtained RNA transcriptome data and corresponding clinical data of 264 cervix samples (including 176 CC and 88 normal cervix) from the GEO database (merged by GSE6791, GSE7803, GSE9750, GSE39001, GSE52903, GSE63514), and 11,965 genes were detected across all 6 datasets (Fig. 1B,C). Among 801 ARGs identified from the GeneCards dataset, 673 ARGs were sequenced in our cohort and their expression data were used to screen DEARGs by comparing them between CC and normal tissues.
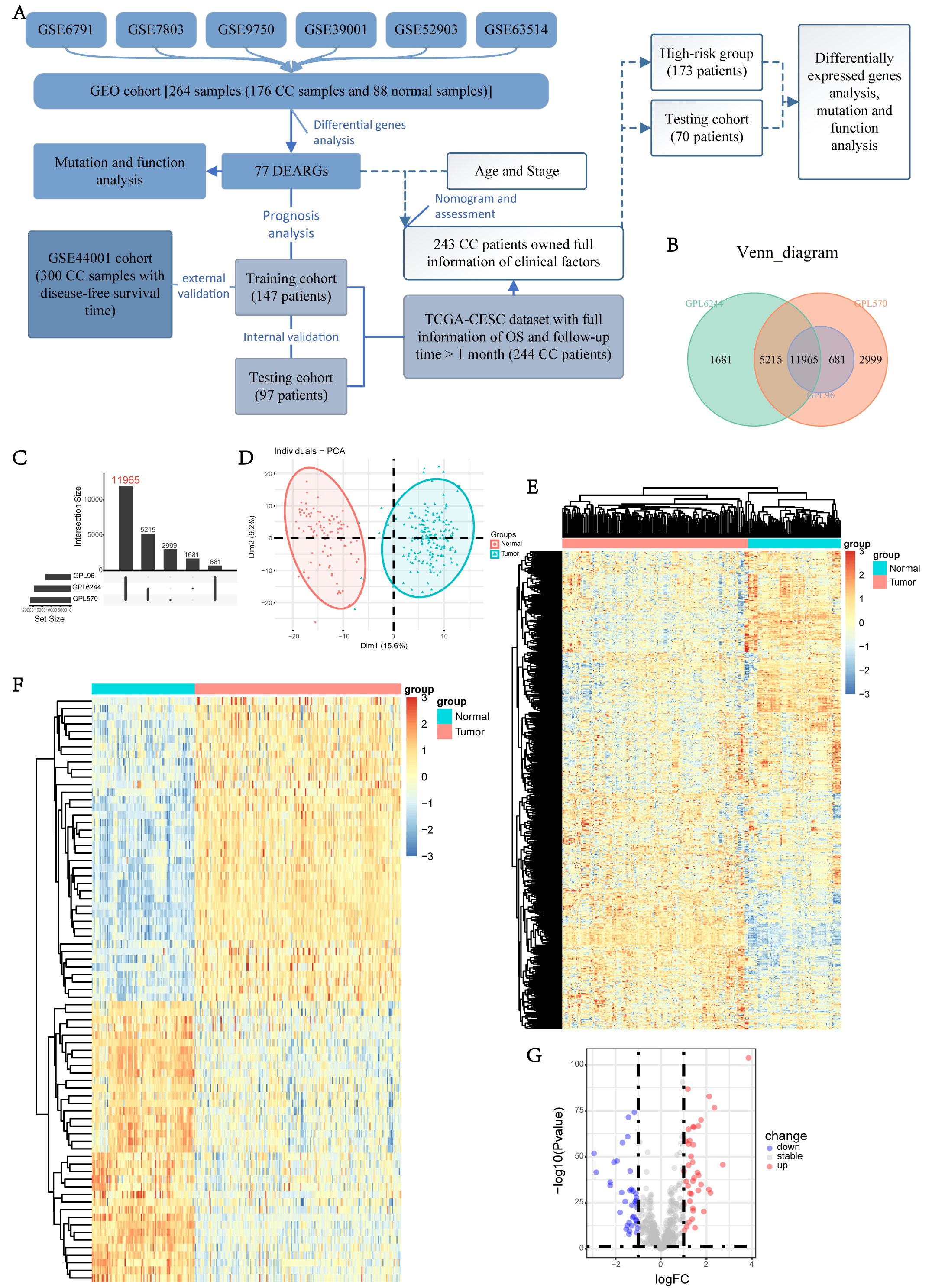 Fig. 1.
Fig. 1.Profile of the study and the screening of DEARGs. (A) The workflow of data analysis. (B,C) Venn diagram of 6 datasets from GEO database. (D) Principal component analysis of the ARGs between normal and tumor samples in GEO cohort. (E) Expression heatmap of the ARGs in GEO cohort. (F,G) Heatmap and volcano plot of the DEARGs in GEO cohort. GEO, Gene Expression Omnibus; DEARGs, differentially expressed anoikis-related genes; ARGs, anoikis-related genes; TCGA, The Cancer Genome Atlas; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma.
Principal component analysis (PCA) showed that the expression of these 673 ARGs
varied between normal cervix and CC samples, suggesting a potential role for
anoikis in CC development (Fig. 1D). We screened 77 DEARGs (40 upregulated and 37
downregulated in tumors) using a threshold of
The position of DEARGs were plotted on the diagram of the chromosome, their
expression heatmap and the regulated status were also shown using the “RCircos”
package (Fig. 2A). We investigated somatic mutations of the 77 DEARGs in 289
TCGA-CESC patients and found the top 5 most frequently mutated DEARGs were BRCA1,
PIK3R1, BRCA2, AR, and DNMT1. The top 30 were analyzed and displayed through
waterfall plots (Fig. 2B). It was shown that missense mutations constituted the
vast majority of mutations after further categorization, the single nucleotide
polymorphism was the most frequent mutation type and the most frequent single
nucleotide variant was the C
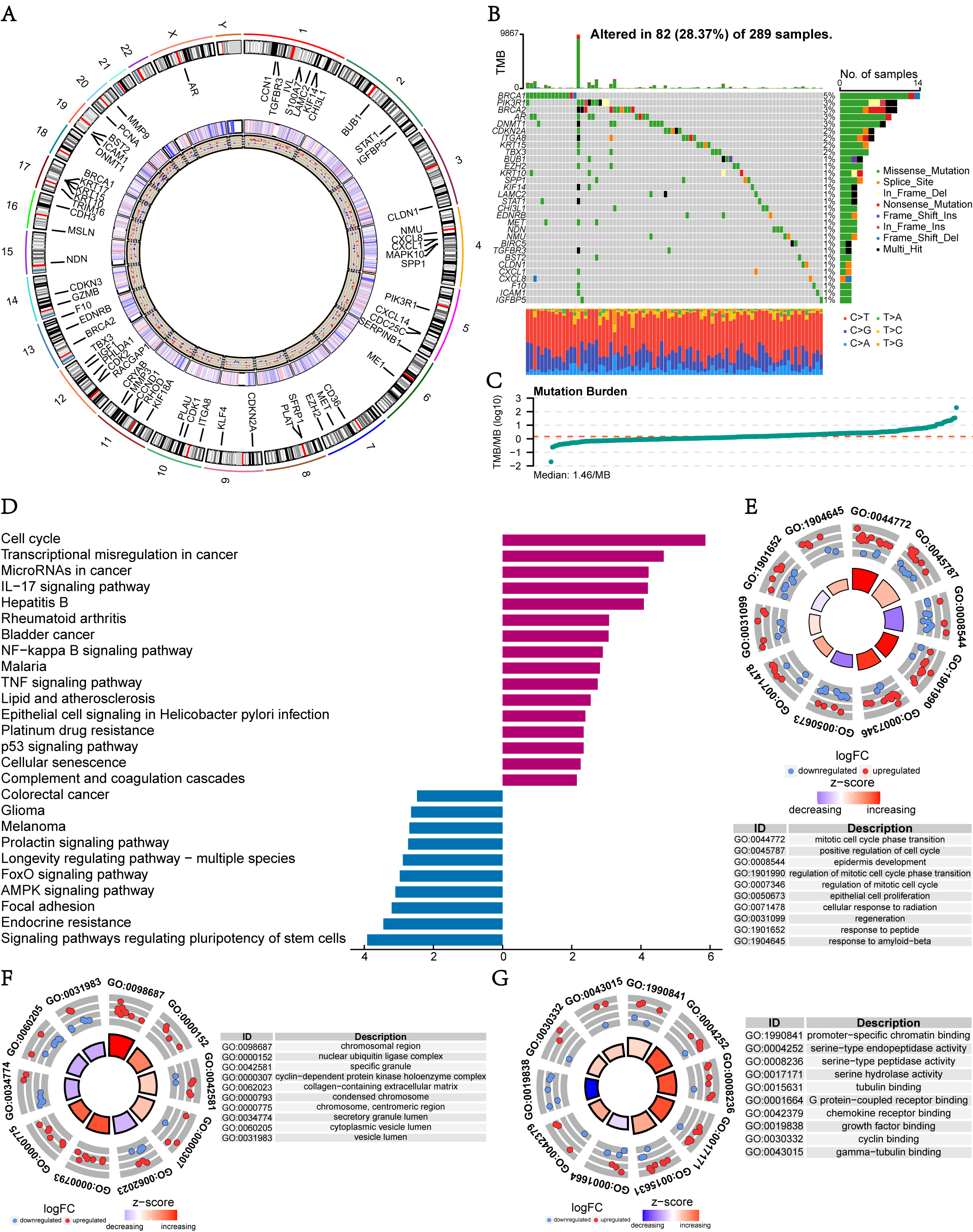 Fig. 2.
Fig. 2.Mutation and function analyses of DEARGs. (A) The diagram of chromosome labeled the DEARGs, their expression heatmap and the regulated status. (B,C) Waterfall plot and TMB distribution of DEARGs in TCGA dataset. (D) Bar diagram showed KEGG analysis of DEARGs, in which two bars in different direction represent upregulated (red bar) or downregulated (blue bar) signaling pathways respectively. (E–G) GO analysis of DEARGs in biological process, cell component, and molecular function respectively. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; TMB, tumor mutation burden.
Functional enrichment analyses were performed to reveal the potential biological functions associated with the DEARGs. As illustrated in Fig. 2D, KEGG enrichment analysis demonstrated that the correlated genes were mainly clustered in several terms, such as cell cycle, transcriptional misregulation in cancer, microRNAs in cancer, IL-17 signaling pathway, signaling pathways regulating pluripotency of stem cells, endocrine resistance, focal adhesion, and the AMPK signaling pathway. At the same the time, GO functional terms of DEARGs in biological process (BP) were significantly enriched in mitotic cell cycle phase transition, positive regulation of cell cycle, and epidermis development categories (Fig. 2E). Cell component related functions of these DEARGs were mainly enriched in the chromosomal region, nuclear ubiquitin ligase complex, and specific granules, among others (Fig. 2F). Molecular function (MF) mainly focuses on promoter-specific chromatin binding, serine-type endopeptidase activity, and serine-type peptidase activity categories (Fig. 2G). Taken together, it was suggested that the DEARGs may be mainly associated with CC proliferation and migration, such as the regulation of cell cycle and adhesion ability. Consistent with known dysfunctions in CC, these results suggest the reliability of our findings.
To explore the connections between the expression of the 77 DEARGs and CC subtypes, we performed a consensus clustering analysis on all 244 CC patients who met the inclusion and exclusion criteria in the TCGA cohort. By increasing the clustering variable (k) from 2 to 10, we found that when k = 2, the intragroup correlations were the highest and the intergroup correlations were low, indicating that the 244 CC patients could be divided into two clusters based on the 77 DEARGs (Fig. 3A). The different expressions of DEARGs between the two clusters were further profiled through PCA (Fig. 3B). The OS time was compared between the two clusters, but no obvious differences were found (p = 0.37, Fig. 3C), suggesting that further screening of DEARGs for establishing the prognostic predicting signature in CC patients was needed.
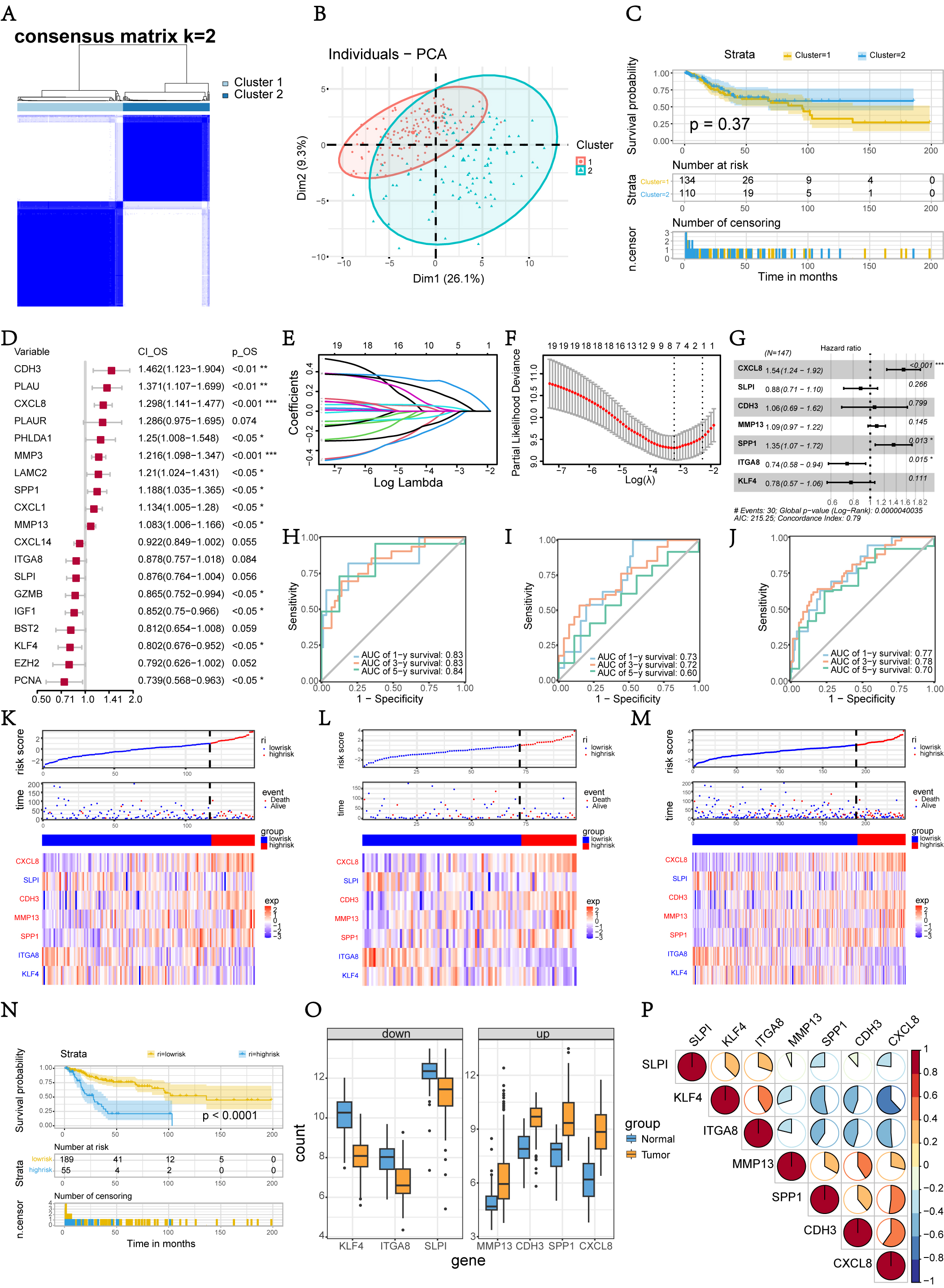 Fig. 3.
Fig. 3.Tumor classification based on the DEARGs and constructions of 7-genes risk signature. (A) 244 CC patients were grouped into two clusters according to the consensus clustering matrix (k = 2). (B) Principal component analysis of the DEARGs between two clusters in TCGA cohort. (C) KM curves of OS between two clusters. (D) Forest plot of 19 DEARGs screened out by Univariate Cox regression analysis. (E,F) LASSO regression analysis determined 7 genes for the construction of prognostic signature. (G) Forest plot of 7-genes prognostic scoring model. (H–J) Time-dependent ROC curves of the signature predicting 1, 3, and 5-years survival in training, internal testing and entire TCGA cohorts. (K–M) Distribution of risk scores, survival state, and the expression heat map of selected DEARGs in training, internal testing and entire TCGA cohorts. (N) KM curves of OS between two risk groups. (O) Boxplot of 7 DEARGs between two risk group. (P) Correlation coefficients heatmap of 7 DEARGs. KM, Kaplan-Meier; OS, overall survival; ROC, receiver operating characteristic; LASSO, least absolute shrinkage and selection operator.
First, the 244 CC patients were randomly grouped into training (147 patients)
and testing sets (97 patients) at a 6:4 ratio. Using the univariate Cox
regression analysis, 19 DEARGs possibly associated with OS in the training set
were screened with a criterion of p
Subsequently, the area under the curve (AUC) value under the time-dependent ROC curve was calculated to assess the prognostic performance of the risk score, with respective AUC values pertaining to 1-, 3-, and 5-year survival outcomes of 0.83, 0.83, and 0.84 in the training cohort (Fig. 3H). To study the prognostic and risk verification abilities of the model, the risk score of each individual in the testing set was calculated using the same calculation as that used in the training set. According to the time ROC analysis, the 1-, 3-, and 5-year AUCs of this signature were 0.73, 0.72, and 0.60, respectively in the testing cohort (Fig. 3I). Based on the risk scores of all of the enrolled patients, the 1-, 3-, and 5-year AUCs under the time-dependent ROC were 0.77, 0.78, and 0.70 respectively in the entire cohort (Fig. 3J). In summary, the ROC curve analyses demonstrated that the DEARGs signature possessed a satisfactory predicting accuracy.
Patients in the training, testing, and the entire sets were stratified into two
groups: HRG and LRG, based on the threshold point of the ROC curve calculated
from the formula in the training set. Then, the distribution of risk score,
survival state, and the expression heat map of selected DEARGs signatures were
compared between the LRG and HRG (Fig. 3K–M). It was shown that established
anoikis-related signatures had a strong ability to predict OS, whereas increasing
risk score was positively correlated with the patients’ poor survival status. The
lower (SLPI, ITGA8, and KLF4) or higher
(CXCL8, CDH3, MMP13, and SPP1) the expression
of these selected DEARGs, the higher the risk rate, suggesting that these DEARGs
help to predict the risk among patients. This conclusion was consistently
validated in all three sets. The KM curve of the entire TCGA set demonstrated
that the survival probability of CC patients at the same time in the HRG was
significantly lower than the LRG; and that the OS in the HRG was much shorter
than that in the HRG (p
Finally, the expression level of the 7 selected DEARGs between normal and CC samples in the merged GEO cohort was presented as a boxplot (Fig. 3O). The expression of these DEARGs were validated using data from the GEPIA public database with 306 CC and 13 normal samples (Supplementary Fig. 2). The Spearman correlation coefficients among the 7 DEARGs were then analyzed (Fig. 3P). CXCL8, CDH3, MMP13, and SPP1, the upregulated DEARGs in CC, were positively correlated with each other, and negatively correlated with SLPI, ITGA8, and KLF4 (the downregulated DEARGs in CC).
The GSE44001 cohort of 300 CC patients with relapse information was used as a
validation set to further evaluate the prognostic performance of our scoring
model. The recurrence risk score was calculated for each individual in the
GSE44001 dataset based on the predicting model, which included 7 DEARGs. The AUC
of 1-, 3-, and 5-year no relapse rates under time-dependent ROC curves were 0.71,
0.72, and 0.77, confirming the prediction accuracy of this model (Fig. 4A).
Therefore, this DEARGs signature was validated to have an important application
in predicting the prognosis, DFS and OS, for patients with CC. These patients
from the GSE44001 cohort were then divided into HRG (n = 116) and LRG (n = 184)
groups using the threshold point of the ROC curve (Fig. 4B). The distribution of
risk scores showed that the patients with elevated risk scores had higher
recurrence rates. The expression levels of the 7 selected DEARGs involved in the
signature are also visualized through heatmap. Significant differences in DFS
were evident between the HRG and the LRG of the GSE44001 cohort (p
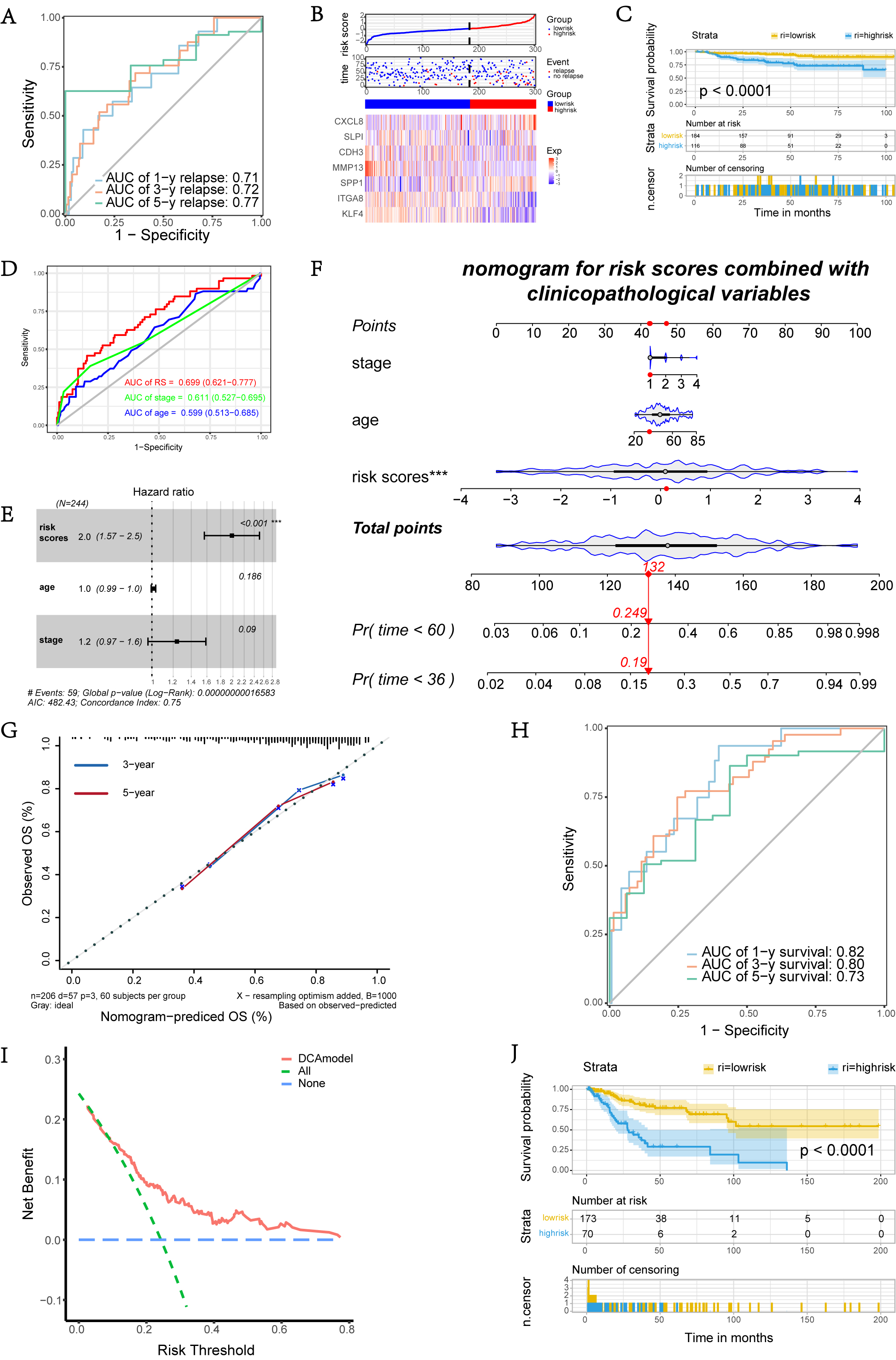 Fig. 4.
Fig. 4.External validation of the 7-genes signature and construction of nomogram involved DEARGs and clinical factors. (A) Time-dependent ROC curves of 1, 3, and 5-years relapse in GSE44001 cohorts. (B) Distribution of risk scores, survival state, and the expression heatmap of selected DEARGs in GSE44001 cohorts. (C) KM curves of DFS between two risk groups. (D) ROC curves of DEARGs signature, age, and stage predicting survival respectively. (E) Forest plot of prognostic scoring model constructed by anoikis pattern combined with clinical factors. (F) Nomogram of the risk model involved anoikis pattern and clinical factors. (G) Calibration curves compare nomogram predicted survival probabilities with actual ones. (H) Time-dependent ROC curves of the risk model predicted 1, 3, and 5-years survival in TCGA dataset. (I) DCA analysis of the risk model. (J) KM curves of OS between two risk groups defined by risk model. DFS, disease-free survival; DCA, decision curve analysis.
The ROC curve indicated that the OS predictive power of this signature relative
to other clinical signatures (including age and FIGO stage) was high in the TCGA
dataset, of which the AUC was 0.699 and the 95% confident interval (95% CI) was
between 0.621 to 0.777 (Fig. 4D). To further verify the accuracy of the ARGs
signatures, an independent prognostic analysis was performed among the DEARGs
risk scores and other clinical signatures through multivariate Cox regression
(Fig. 4E). As depicted in the forest plot, the anoikis risk score was an
independent predictor as compared with the other clinical signatures (p
To provide a better quantitative method for clinicians to predict cancer prognosis, we constructed a mixed nomogram combining the prognostic scoring model and clinicopathological features to score patients and predict their OS at 3 and 5 years (Fig. 4F). We also corrected the predictive performance of this nomogram, and the calibration plots showed that predicting 3- or 5-year OS based on the nomogram exhibited a consistent agreement between the actual probability and the predicted probability (Fig. 4G). In addition, the time-dependent AUCs were 0.82, 0.80, and 0.73 for predicting OS at 1-, 3-, and 5-year time points, suggesting that the predictive performance of the nomogram was satisfactory (Fig. 4H). We also calculated other metrics of our nomogram and compared them with using only age or stage to predict the prognosis of the TCGA cohort. The specificity (SP), sensitivity (SN), accuracy (ACC) and Matthews correlation coefficient (MCC) of the model were 0.799 (0.741–0.857, 95% CI), 0.559 (0.433–0.686, 95% CI), 0.741 and 0.338. The MCC of the model based on age or stage were 0.178 and 0.235 respectively, which further verified that the prognostic ability in our model has been increased by including the DEARGs. Almost all of the red curve was in the area above the green and blue solid lines in the DCA (Fig. 4I), demonstrating a high net benefit could be obtained by using the nomogram to make clinical decisions. According to the risk scores generated from the nomogram, CC patients in the entire TCGA set were grouped into different risk groups by the threshold point of the ROC curve. It was demonstrated that patients in the HRG possessed significantly inferior OS compared to those in the LRG in the KM curves (Fig. 4J).
Through differential gene screening, we identified 118 DEGs associated with HRG
and LRG in the TCGA-CESC cohort, including 34 upregulated and 84 downregulated in
the HRG (Fig. 5A,B). Among the 787 ARGs sequenced in the TCGA dataset, 21 DEARGs,
including 13 upregulated and 8 downregulated, were identified between the HRG and
the LRG (Fig. 5C). The threshold of
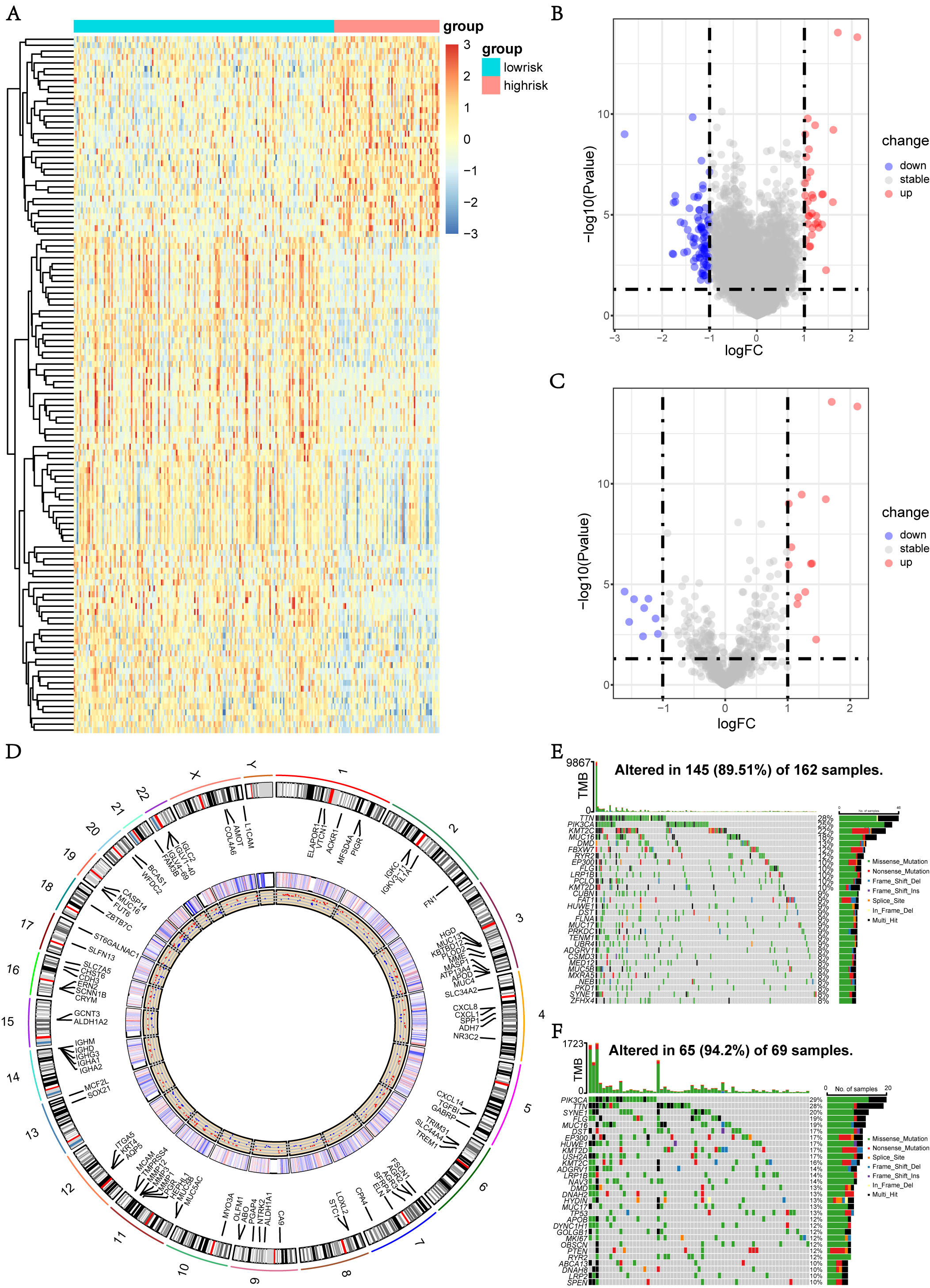 Fig. 5.
Fig. 5.DEGs and mutation analysis between different risk groups. (A,B) Heatmap and volcano plots of the DEGs between high-and low-risk groups in TCGA cohort. (C) Volcano plot of the DEARGs between high-and low-risk groups in TCGA cohort. (D) The diagram of chromosome labeled the DEGs between high-and low-risk groups, their expression heatmap and the regulated status. (E,F) Waterfall plots of top 30 mutated genes in high-and low-risk groups of TCGA dataset.
In addition, typical somatic mutations of individuals in the HRG and the LRG
were investigated using the Waterfall plots, in which the genes with the top 30
mutation frequency were visualized (Fig. 5E,F). Missense mutations still
constituted the vast majority of mutations in both groups after further
categorization based on numerous comprehensive classifications. A significant
difference was observed between the HRG (94.20%) and the LRG (89.51%) in terms
of the total mutation rate. The mutation rate of the 12 genes was higher than
10% in the LRG, while all of the top 30 mutated genes in the HRG showed a
mutation frequency higher than 10%, suggesting that not only the mutation rates
were elevated, but also the genes with a high mutated rate were increased in the
HRG. Furthermore, it was shown in our study that there was a
significant but not so strong relationship between TMB and risk scores
(correlation coefficient = 0.14, p
GO and KEGG pathway enrichment analyses of the 118 DEGs between the HRG and the LRG were conducted to investigate their potential biological functions. GO enrichment analysis of BP showed that they were mainly enriched in humoral immune response, complement activation, defense response to bacterium, recognition of phagocytosis, and tissue homeostasis (Fig. 6A). Several cell components related to these DEGs included immunoglobulin complex, blood microparticles, the external side of the plasma membrane, and the apical plasma membrane (Fig. 6B). The MF of these genes were mainly clustered in terms of immunoglobulin receptor binding, antigen binding, cytokine activity, receptor ligand activity and signaling receptor activator activity (Fig. 6C). KEGG pathway enrichment analysis revealed that most enriched pathways were mainly involved in the regulation of immune functions (Supplementary Fig. 4).
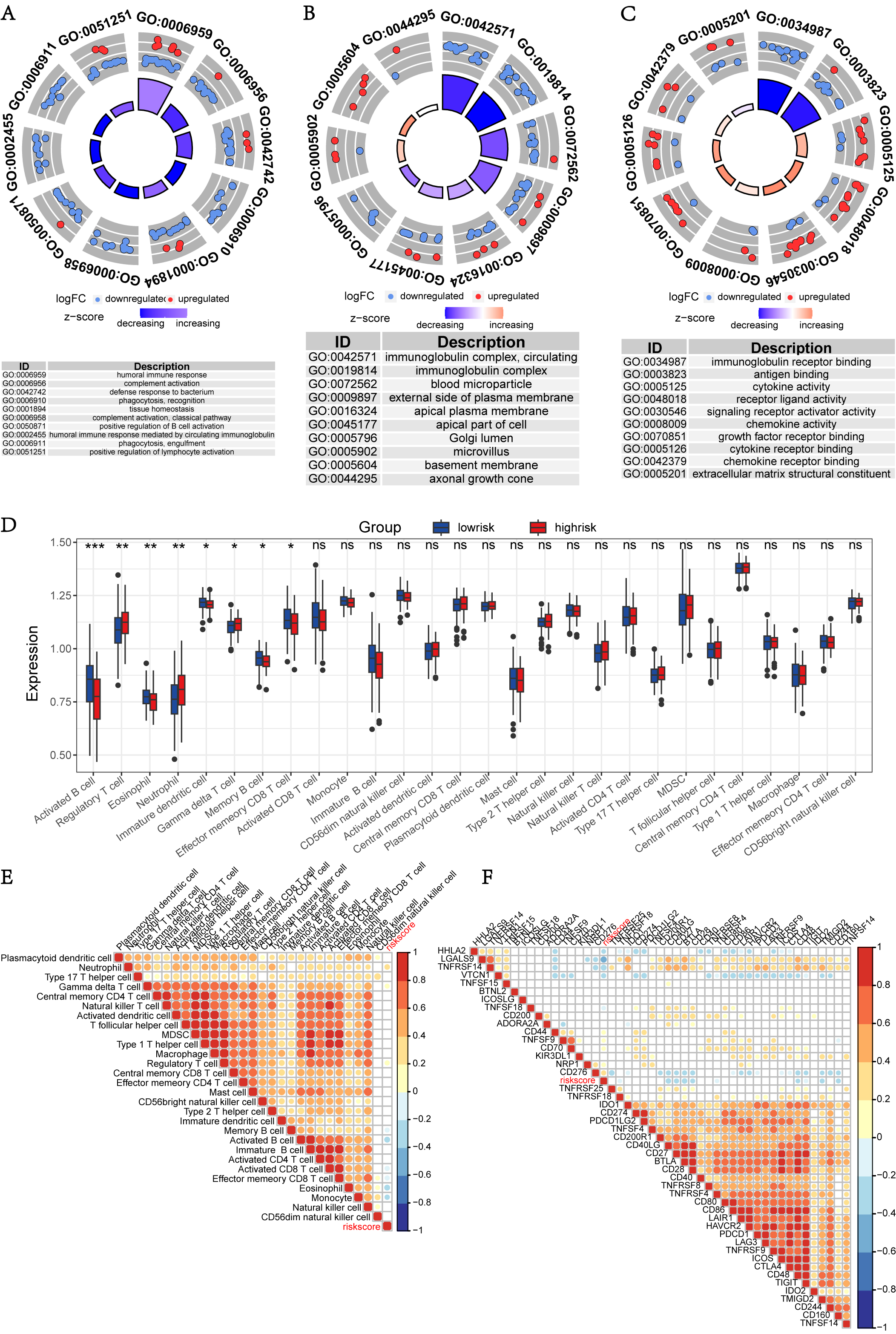 Fig. 6.
Fig. 6.Functional analysis of DEGs and immune analysis between low- and high-risk groups. (A–C) GO analysis of DEGs between low- and high-risk groups in biological process, cell component, and molecular function respectively. (D) Boxplot of immune cells infiltration between two risk group. (E) Heatmap of spearman correlation coefficients between risk scores and the abundances of immune cells. (F) Heatmap of spearman correlation coefficients between risk scores and the expressions of immune checkpoints.
Taken together, these results suggest that the genes involved in immune pathways
may have significant differences between the HRG and the LRG which are
discriminated by our risk model. Therefore, we continued to study the
relationship between the risk score and tumor immune microenvironment. First, in
terms of immune cells, immune penetration is represented by a boxplot constructed
using the ssGSEA (Fig. 6D). It was shown that the abundances of regulatory T
cells, neutrophils, and
Cervical cancer is one of the malignancies that imperils women health, which resulted in approximately more than 60,000 deaths of Chinese females due to CC in 2022 [41]. While the incidence has significantly decreased in developed countries due to screening programs and HPV vaccines, the survival of CC has still not improved significantly since the 1970s and treatment options are still restricted, especially for advanced CC [42, 43, 44]. Anoikis could be viewed as a physiological process which ensures tissue homeostasis. Resistance to anoikis might result in adherent cells proliferating at ectopic sites, which is emerging as a hallmark of metastatic cancers [20, 21, 22, 23, 25, 26, 27, 28, 29, 30, 45]. Therefore, ARGs have attracted much attention for their important functions in the development of CC [31, 32, 33, 34]. Several attempts have been made to explore the promising prognostic values of ARGs [46]. However, the anoikis-related molecular subtypes of CC and the performance of anoikis pattern in predicting mutational status and sensitivities to therapies has not been verified.
In this study, we first studied the expression profile of 673 ARGs in 176 CC and 88 normal cervix tissues from the GEO database. It was found that 77 were differentially expressed, including 40 upregulated (51.9%) and 37 downregulated in tumors (48.1%). Their mutation status as well as bio-functions were further explored. However, the KM analysis of the two clusters produced by the consensus clustering analysis based on the DEARGs did not show any significant differences for OS in the TCGA cohort. To further assess the prognostic value of these DEARGs, a 7-gene risk signature was identified using the univariate Cox model combined with LASSO regression analysis, which was then validated by internal and external datasets. Subsequently, a nomogram involving DEARGs risk scores and clinical factors was established to stratify patients into the LRG and the HRG of CC. More genes with higher mutation rate were detected in HRG. Furthermore, the DEGs between the LRG and HRG were related to immune-related pathways. The immune cell infiltration in the LRG and HRG were compared and it was found that the HRG had universally decreased levels of infiltrating immune cells and increased immune-suppressive cells. Finally, the correlation coefficient of the risk scores with expression of immune checkpoints and the DEARGs with the sensitivity to chemotherapeutic and targeting drugs were calculated, suggesting that the risk model might predict the therapeutic effect of CC.
It still remained unknown how the 7 DEARGs (CXCL8, CDH3, MMP13, SPP1, ITGA8, KLF4, and SLPI) affect the oncogenesis and development of CC. The potential mechanism of these DEARGs might be diverse according to previous studies. The elevated expressions of CXCL8, CDH3, MMP13, and SPP1 were identified as risk factors for the prognosis of CC in our study. Interleukin-8 (IL8), alternatively known as CXCL8, is a proinflammatory CXC chemokine that is associated with multiple components of tumor microenvironment to regulate the proliferation and migration of various cancer cells. Increasing evidence has shown that CXCL8/CXCR1/2 signaling plays a substantial regulatory role in tumor metastasis. For example, it was elevated to contribute toward melanoma progression upon anoikis resistance and reattachment from cell suspension [47]. It was also demonstrated that CXCL8 inhibits anoikis of colorectal carcinoma cells via increasing TLAK cell-originated protein kinase (TOPK, a MAPKK-like serine/threonine protein kinase) levels and activating AKT and ERK signaling [48]. A similar pathway was suggested in head and neck squamous cell carcinoma cells which showed that blockade of CXCL8 by gene silencing in endothelial cells inhibited phosphorylation of STAT3, AKT, and ERK of tumor cells, thus increased their anoikis [49]. In addition, the autocrine CXCL8 was identified to enhance the anoikis-resistance via the JNK/p38-ATF-2 axis as a pro-invasive pathway in lung cancer cells [50]. Cadherin 3 (CDH3), or P-cadherin, belongs to calcium-dependent cell adhesion proteins. It has been proposed that P-cadherin upregulates carbon flux through the pentose phosphate pathway and decreases oxidative stress in matrix-detached breast cancer cells, thus promoting anoikis in circulation and metastatic sites [51]. Its upstream regulation could be varied, for instance, the homeobox gene HOXA9 promotes aggregation and inhibits anoikis in floating epithelial ovarian cancer (EOC) cells through its induction of P-cadherin [52]. Matrix metallopeptidase 13 (MMP13), or collagenase 3, could function in the degradation of multiple extracellular matrix proteins. It has been shown to shed the Nerve/glial antigen 2 (NG2) on the cell surface, which is a transmembrane proteoglycan receptor that interacts with extracellular matrix to mediate cell adhesion and proliferation, to attenuate anoikis [53]. Interestingly, CXCL8/CXCR1/2 signaling might lead to p38 MAPK activation that triggered downstream MMP13 actions [54]. Secreted phosphoprotein 1 (SPP1), or osteopontin, could be viewed as a non-collagenous bone protein which is important to cell-matrix interaction or to cytokines involved in immune-related pathways. It has been found that invasive tumor cells generate three splice variants of the metastasis gene osteopontin (a, b, and c), a coalescence in cancer cells between osteopontin-a and osteopontin-c, which increases the cellular glucose levels and utilizes this glucose to generate energy respectively, which could offer energy for detached cancer cells in anoikis resistance [55]. In addition, osteopontin-c, as the soluble form of osteopontin, could protect cancer cells from anoikis during anchorage-independent growth by inducing the expression of oxidoreductases [56]. For the study of upstream, it was shown that breast cancer metastasis suppressor 1 (BRMS1) sensitizes HCC cells to anoikis by suppressing osteopontin expression [57].
In contrast, the activation of ITGA8, KLF4, and SLPI
were shown as protective factors for the prognosis of CC. Integrin
The functional enrichment analysis of DEARGs between normal cervix and CC samples revealed that the underlying biological mechanisms were mainly enriched in terms of cell cycle, transcriptional regulation and focal adhesion, which are correlated with the biological process of anoikis and consistent with previous studies [63]. While CESC patients with different TMB were previously shown to have different survival outcomes [64], the OS of DEARGs high- and low-TMB groups showed no significant difference in our study. It was also shown that TMB has significant but not such a strong positive relationship with risk scores in our model, which included DEARGs and clinicopathological factors. However, different risk groups defined by our model showed different mutation rates, suggesting that the mutation status may affect the up or down stream genes rather than ARGs directly. We analysed the 118 DEGs between different risk groups and found that they were mainly involved in immune responses in addition to tissue homeostasis. Based on the results of our GO and KEGG analyses, it is reasonable to speculate that anoikis can regulate the composition of the tumor immune microenvironment.
Related function analysis of immune cell subsets showed that HRG had universally
decreased levels of infiltrating immune cells, while regulatory T cell,
We explored the anoikis-related biomarkers that could be used for discriminating subgroups of CC and predicting prognosis and therapeutic responses. However, there are still a few limitations in our research. First, our research data comes from public databases and some specific clinical information may have been unavailable. Moreover, in order to validate our findings and to uncover the mechanism of anoikis-related genes in CC, further functional experiments are needed in our laboratory based on cell lines and tissue samples.
In summary, the finding of our study revealed potential biomarkers and therapeutic targets for anoikis-related signatures in cervical cancer. Novel subgroups of CC were defined through clustering based on the anoikis patterns. The prognostic model based on the ARGs and clinical factors could refine the predicted performance of CC survival outcomes, evaluate mutation and immune conditions, and predict the sensitivities of therapies for CC patients.
All data can be retrieved from the public database.
JD designed the study, analyzed the data, and revised the paper. XX collected the data and wrote the paper. Both authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work. Both authors read and approved the final manuscript.
Not applicable.
Not applicable.
This study was supported by Fudan University’s “Tomorrow Star” Famous Physicians Cultivation Project.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






