1 Department of Gynecology, Clinical Oncology School of Fujian Medical University, Fujian Cancer Hospital, 350014 Fuzhou, Fujian, China
2 Department of Radiation Oncology, Fujian Medical University Union Hospital, 350001 Fuzhou, Fujian, China
3 Fujian Key Laboratory of Intelligent Imaging and Precision Radiotherapy for Tumors (Fujian Medical University), 350001 Fuzhou, Fujian, China
4 Clinical Research Center for Radiology and Radiotherapy of Fujian Province (Digestive, Hematological and Breast Malignancies), 350001 Fuzhou, Fujian, China
Abstract
Background: Ovarian cancer is the second leading cause of gynecologic
cancer-associated deaths. Cancer stemness and chemoresistance are responsible for
ovarian cancer metastasis and the poor prognosis of patients. In this study, we
determined the function of N
Keywords
- N6-methyladenosine
- PTGER2
- ovarian cancer
- stemness
- chemoresistance
Ovarian cancer is characterized by a high prevalence and high mortality rate and ranks as the second leading factor causing gynecologic cancer-related deaths in women worldwide [1]. Platinum-based chemotherapy is still the main treatment for advanced ovarian cancer [2]. Although ovarian cancer mortality has declined in recent years [3], ovarian cancer stemness, chemoresistance, and metastasis are common reasons for treatment failure and poor patient prognosis [4]. Therefore, identifying novel targets to overcome these tumorigenic properties is now an urgent priority for this disease.
N
Prostaglandin E2 receptor (subtype EP2), namely PTGER2, belongs to the G protein-coupled receptor superfamily. The activation of PTGER2 results in the increased activity of cAMP-dependent signaling [13]. Previous studies have revealed that PTGER2 acts as an oncogene in colorectal cancer [14] and prostate cancer [15] and serves as a tumor suppressor in neuroblastoma [16]. However, the biological role and clinical significance of PTGER2 and the mechanism of PTGER2 dysregulation in ovarian cancer remain unclear.
Here, we elucidated the role of m
Bioinformatics analysis was performed according to our previous study [17]. The genomic data, mRNA expression profiles, and clinical data of The Cancer Genome Atlas (TCGA) ovarian serous cystadenocarcinoma (OV) database were downloaded for conducting differential analyses, survival analyses, correlation analyses, unsupervised clustering analyses, gene set variation analysis (GSVA), and gene set enrichment analysis (GSEA). The gene set representing platinum drug resistance was constructed using data obtained from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database as previously reported [18]. Based on the best cut-off value obtained from survival analysis, samples were assigned as low and high expression groups.
Ovarian cancer cell lines OV90 and SKOV3 with short tandem repeat verification
were purchased from American Type Culture Collection (Manassas, VA, USA) and
cultured in Dulbecco’s Modified Eagle Medium (DMEM) harboring 10% fetal bovine
serum (FBS) at 37 °C with 5% CO
Single hairpin RNA (shRNA) negative control and shRNA targeting PTGER2 (sh-PTGER2) were constructed and inserted into lentiviral particles by GeneChem Corporation (Shanghai, China). Small interfering RNA (siRNA) against METTL3 (si-METTL3) was synthesized by RiboBio Corporation (Guangzhou, China). Polybrene and Lipofectamine 3000 were used for lentivirus particle transfection and siRNA transduction, respectively. After transfection or transduction for 48–72 h, ovarian cancer cells were subjected to further investigation.
Total RNAs of ovarian cancer cells were used for reverse transcription with a
kit purchased from TaKaRa Corporation (Dalian, China). RT-PCR and quantitative
PCR (qPCR) were conducted with the generated cDNAs and specific primers. Bio-Rad
GelDoc XR+ (Hercules, CA, USA) was applied to capture images for RT-PCR, and the
2
Total proteins of ovarian cancer cells in lysis buffer were quantified by the
BCA method as previously described in our investigation [19]. Then the separated
and transferred proteins were incubated with antibodies against the following
proteins: METTL3, PTGER2, Myc, cyclin D1 (CCND1), vimentin, and
The Methylated RNA Immunoprecipitation (MeRIP) m
PmirGLO luciferase reporter vectors (Promega Co., Madison, WI, USA) containing wild-type or mutant 3’-untranslated region (UTR) of PTGER2 were transfected into ovarian cancer cells. After transfection for 48 h, the luciferase activity in cells was measured to evaluate the effects of METTL3 on PTGER2 transcriptional levels on the BioTek Luminometer Synergy H1 (Winooski, VT, USA) using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Ovarian cancer cells seeded on 6-well ultra-low-attachment plates were incubated in serum-free DMEM/F12 (20 ng/mL epidermal growth factor, 20 ng/mL fibroblast growth factor, and 2% B27). The images of tumorspheres were captured, and the number of tumorspheres was counted after a 14-day culture. Three continuous generations of tumorspheres number were calculated for analysis.
Ovarian cancer cells seeded on coverslips were treated with paraformaldehyde
(4%) for fixation, incubated with Triton X-100 (0.2%) for permeabilization, and
subjected to incubation with antibodies against the following proteins: cluster
of differentiation 44 (CD44), CD133, and gamma-H2A histone family member X
(
To detect cell proliferation and chemoresistance, ovarian cancer cells were subjected to treatment with carboplatin for 6 h at the indicated concentrations and then seeded in wells for 10 days. The generated colonies were subjected to fixation, staining, and recording for analysis.
The migration and invasion ability of ovarian cancer cells were detected by transwell assays. For the migration assay, serum-free cells were seeded in the upper chamber of a transwell without Matrigel coating. For the invasion assay, serum-free cells were plated in the upper chamber of a transwell with Matrigel coating. DMEM containing 10% FBS was added to the lower chambers of a transwell were for both the migration and invasion assays. The migrated and invaded cells were subjected to fixation, staining, and recording for analysis.
One hundred and fifty-eight ovarian cancer specimens described in our previous study [19] were collected for analysis. All specimens had a pathological diagnosis. Patient written consent and ethics approval from the Ethics Committee of the hospital were obtained (K2020-036-01). All procedures strictly adhered to the policies approved by the Institutional Ethics Committee and Declaration of Helsinki.
Immunohistochemistry (IHC) and staining evaluation were carried out as previously described [19]. Paraffin-fixed sections from ovarian cancer specimens were deparaffinized and rehydrated before antigen retrieval in citrate buffer. Then the sections were subjected to the eradication of endogenous peroxidase activity and blocking of non-specific antigens. After incubation with anti-PTGER2 antibody, the sections were treated with Diaminobenzidine substrate and subjected to the evaluation of staining intensities. For statistical analysis, a score of 7–12 was assigned as high expression, and a score of 0–6 was considered low expression.
SPSS 22.0 (IBM SPSS Inc., Chicago, IL, USA) or RStudio 9.0 was applied for analysis
of the data expressed as the mean
To explore the role of m
To further examine how the 23 m
To identify specific genes that could be affected by m
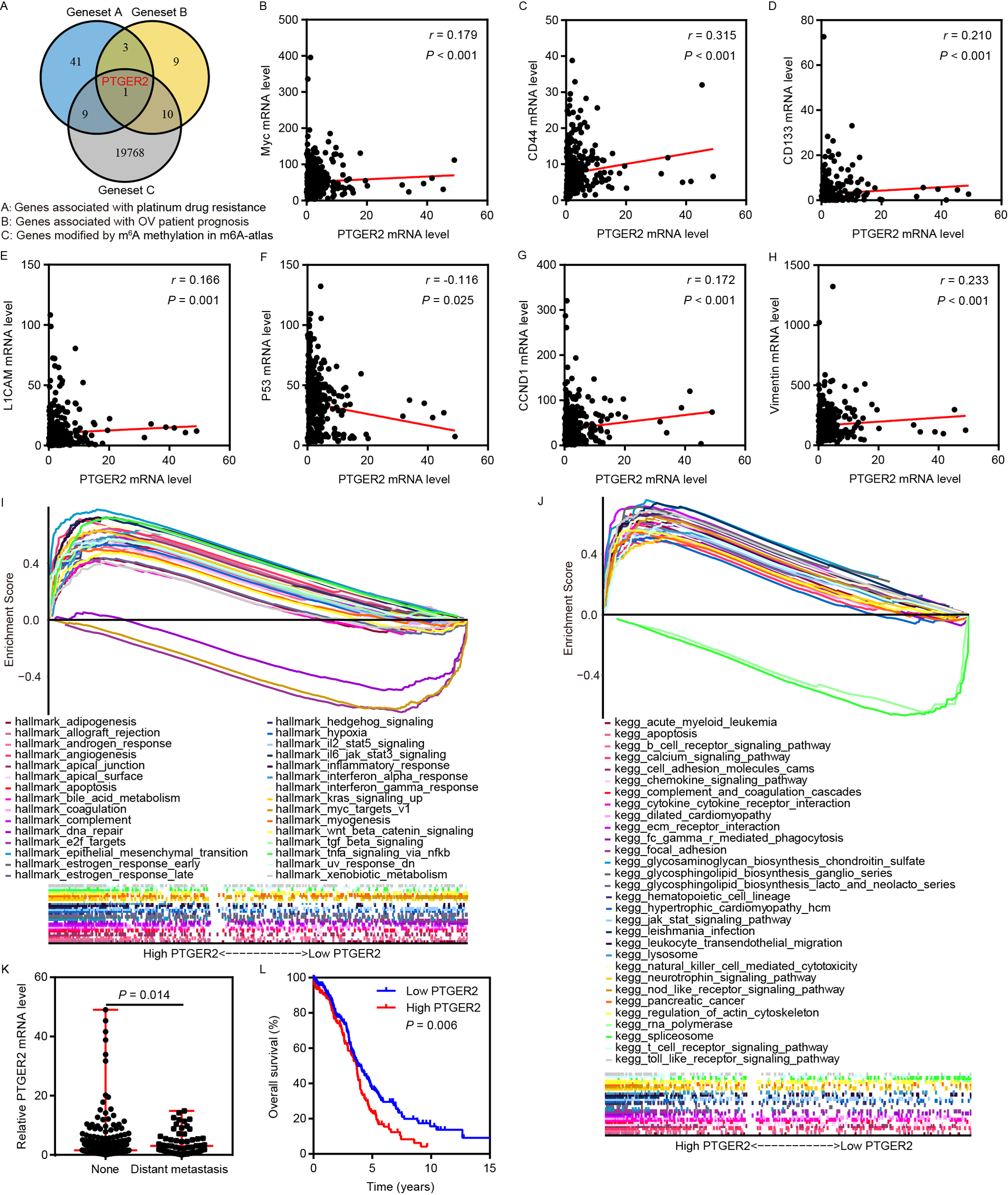 Fig. 1.
Fig. 1.PTGER2 expression serves as an m
Subsequent correlation analyses revealed a positive correlation between PTGER2
expression and the expression of Myc, CD44, CD133, L1 cell adhesion molecule
(CAM), CCND1, and vimentin, which serve as stemness, proliferation, and
epithelial-mesenchymal transition (EMT)-associated markers, and showed a negative
association between PTGER2 expression and expression of the DNA damage
repair-associated marker p53 (Fig. 1B–H). Moreover, GSEA confirmed that PTGER2
positively participated in oncogenic processes and pathways such as DNA repair,
E2F targets, EMT, Myc targets, Wnt/
Based on the findings of the above-mentioned bioinformatics analyses, we further
investigated whether PTGER2 expression is modulated by m
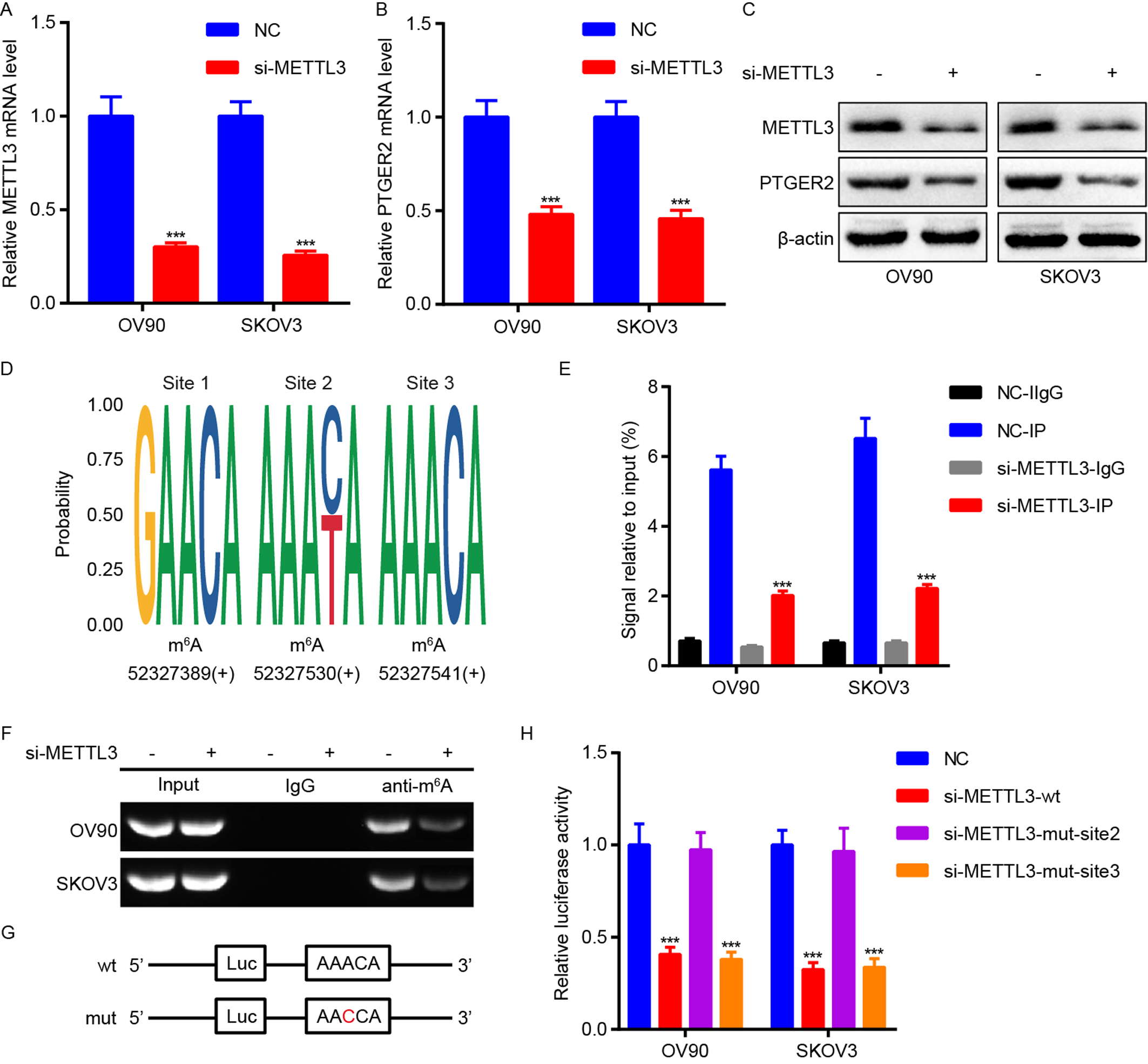 Fig. 2.
Fig. 2.METTL3 is responsible for m
According to the results obtained from the above bioinformatics analyses, we
determined the impact of PTGER2 on ovarian cancer cell stemness, chemoresistance,
proliferation, and metastasis. First, shRNA targeting PTGER2 (sh-PTGER2) was used
for PTGER2 silencing in ovarian cells. Then tumorsphere formation and
immunofluorescence assays showed that PTGER2 knockdown suppressed the stemness of
ovarian cancer cells and impaired the expression of stem cell markers CD44 and
CD133 in ovarian cancer cells (Fig. 3A–C). Colony formation assays indicated
that PTGER2 depletion inhibited the carboplatin resistance and proliferation of
ovarian cancer cells (Fig. 3D,E), and transwell assays measured the inhibition of
migration and invasion by PTGER2 knockdown in ovarian cancer cells (Fig. 3F,G).
Furthermore, immunofluorescence assays detected the upregulation of
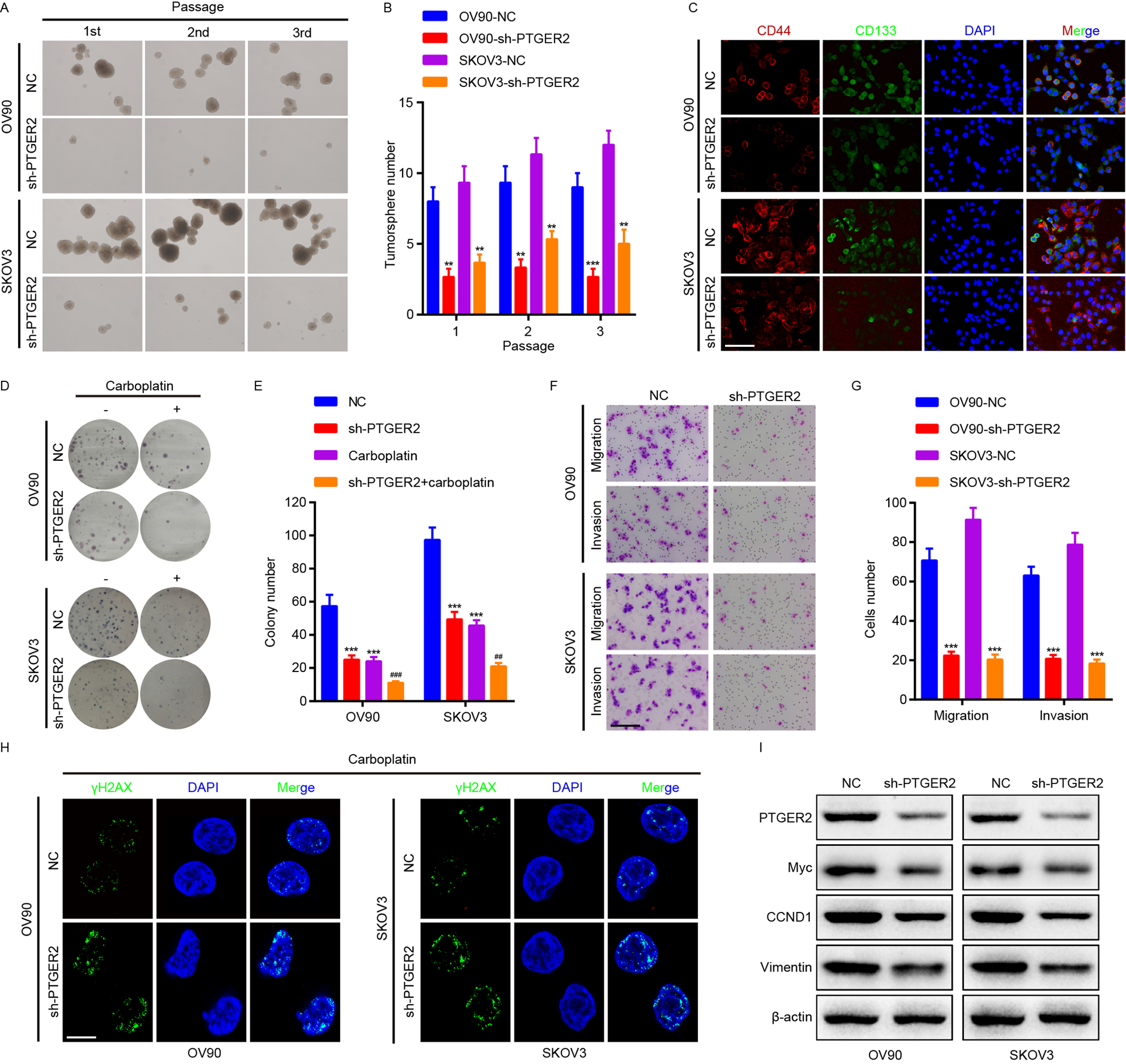 Fig. 3.
Fig. 3.PTGER2 increases ovarian cancer stemness,
chemoresistance, proliferation, and metastasis. (A–G) Tumorsphere formation
(A,B), immunofluorescence (scale bar: 12.5 µm) (C), Clonogenic (D,E),
transwell (Scale bar: 50 µm) (F,G) assays were adopted for detecting the
stemness, chemoresistance, proliferation, and metastasis of PTGER2-depleted OV90
and SKOV3 cells and the controls. (H) Immunofluorescence analysis (scale bar: 8
µm) was applied to measure
To gain further knowledge of the vital role of PTGER2 in the progression of ovarian cancer, we performed IHC in 158 ovarian cancer tissues. IHC showed the different levels of PTGER2 protein expression in the detected ovarian cancer samples (Fig. 4A), and PTGER2 protein expression was upregulated in ovarian cancer samples with metastasis compared to those without metastasis (p = 0.021) (Fig. 4B, Supplementary Table 1). However, PTGER2 protein expression had no significant correlation with other clinical and pathological characteristics. Survival analysis was performed and revealed that high PTGER2 protein expression conferred poor overall survival for patients with ovarian cancer (p = 0.009, log-rank test) (Fig. 4C). Univariate analyses were subsequently conducted and elucidated the positive associations between high PTGER2 protein expression, old age, advanced T classification, advanced N classification, distant metastasis, and poor overall survival of patients with ovarian cancer (Fig. 4D). Multivariate analyses were performed and revealed that PTGER2 protein expression (HR = 1.095, 95% confidence interval [CI] 1.061–1.180, p = 0.027), T classification, and lymph node metastasis served as independent and unfavorable prognostic factors for patients with ovarian cancer (Fig. 4E).
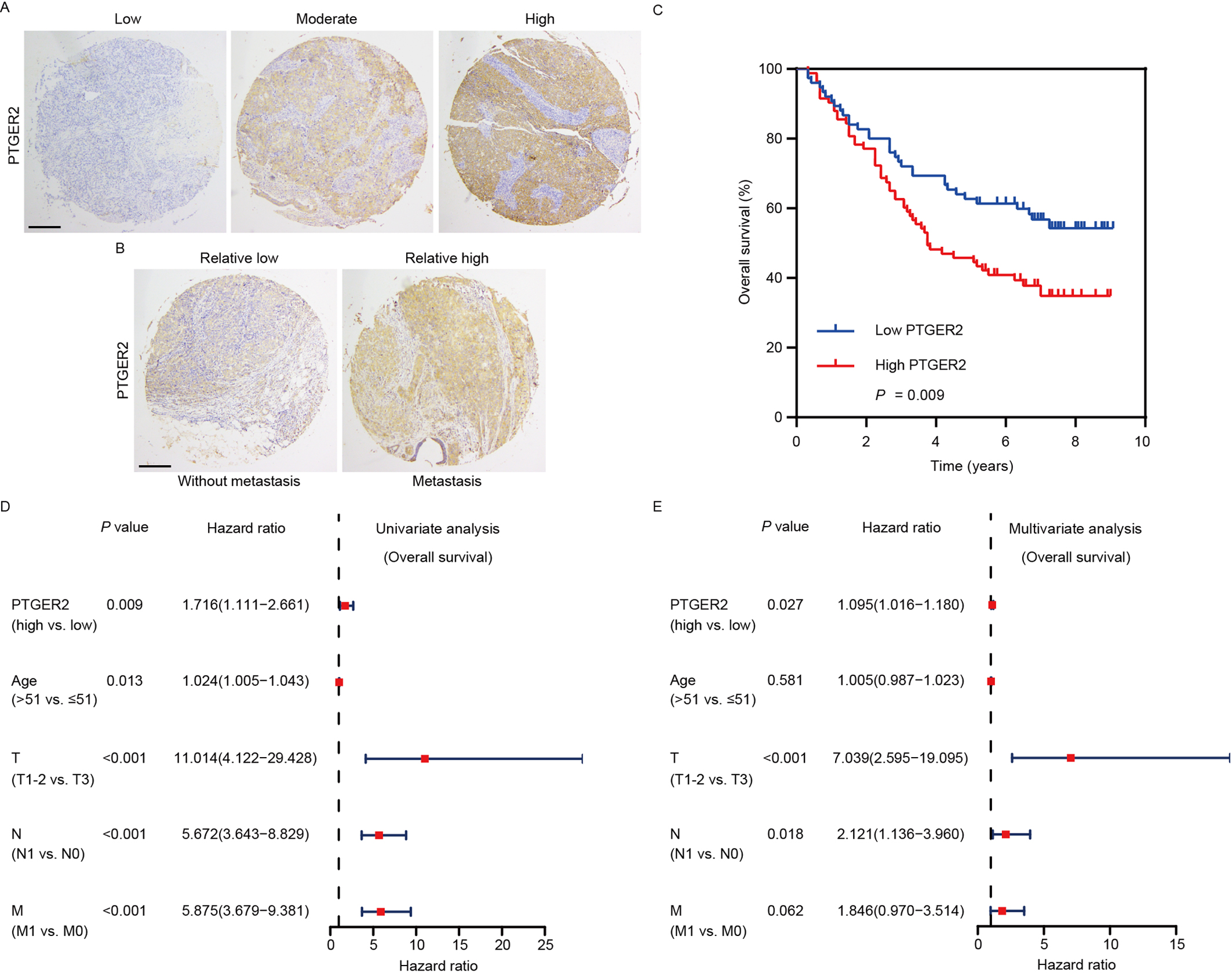 Fig. 4.
Fig. 4.PTGER2 expression predicts the prognosis of ovarian cancer patients. (A) Images representing the differential PTGER2 expression in ovarian cancer tissues (scale bar: 125 µm). (B) Images representing the differential PTGER2 expression between ovarian cancer patients with or without distant metastasis (scale bar: 125 µm). (C) Kaplan-Meier curve displaying the overall survival of patients with ovarian cancer based on PTGER2 protein levels. (D) Univariate analysis was used to correlate PTGER2 protein levels, clinicopathological parameters, and overall survival of patients with ovarian cancer. (E) Multivariate analysis was conducted to correlate PTGER2 protein levels, clinicopathological parameters, and overall survival of patients with ovarian cancer.
Developing new biomarkers for cancer diagnosis, treatment, and prognosis
evaluation are the key issues in cancer research [20, 21]. Stemness and
chemoresistance are responsible for proliferation and metastasis and are regarded
as contributors to poor prognosis of patients with ovarian cancer [19]. m
Distinct m
Previous studies have shown that METTL3 promotes ovarian cancer proliferation,
migration, and invasion [24, 25]. In addition, METTL3 is an inducer of cancer
stem cell self-renewal and chemoresistance in human cancers [26, 27]. In this
work, the regulatory impact of m
PTGER2 plays dual roles as an oncogene or a tumor suppressor in human cancers [13, 15, 16]. PTGER2 facilitates cancer stemness, chemoresistance, proliferation, and metastasis [14, 29]. In the present work, we also revealed the role of PTGER2 in promoting ovarian cancer stemness, chemoresistance, proliferation, and metastasis. Cancer stem cell self-renewal properties contribute to regulating the EMT process and DNA damage repair [30]. The facilitation of cancer chemoresistance can be regulated by several mechanisms, including modulation of cancer stemness, DNA damage repair, and the EMT [31]. In our investigation, we further discovered that PTGER2 increased cancer stem cell self-renewal properties, the EMT, and DNA damage repair to enhance cell stemness, resistance to carboplatin, proliferation, and metastasis, thus potentiating ovarian cancer progression. Moreover, bioinformatics analysis of TCGA ovarian cancer samples and immunohistochemical assays in our collected clinical samples confirmed that PTGER2 functioned as an oncogene and was correlated with ovarian cancer distant metastasis. Importantly, PTGER2 expression was identified as an independent and unfavorable factor in patient tissues, further supporting the conclusions drawn from ovarian cancer cells.
Overall, our study provides a new regulatory axis consisting of METTL3 and
PTGER2 in the modulation of ovarian cancer progression. METTL3-mediated m
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
YL and BX conceived and designed the present study. YL wrote the manuscript. BX checked and revised the manuscript. Both authors contributed to the article and approved the submitted version. Both authors read and approved the final manuscript. Both authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Patient written consent and ethics approval from the Ethics Committee of the hospital were obtained (K2020-036-01).
Not applicable.
This work was founded by Startup Fund for scientific research, Fujian Medical University (Grant number: 2021QH1141).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.




