- Academic Editor
Background: Myasthenia gravis (MG) is an autoantibodies-mediated
autoimmune disease with the complications of neuromuscular junction transmission.
In this study, we aimed to investigate the molecular regulatory roles of pentaxin
3 (PTX3) in patients and in animal model with MG and to explore its underlying
mechanism. Methods: Patients with MG were identified and enrolled at our
designated hospital and animal model was utilized for the proposed study. Enzyme-linked immunosorbent assay (ELISA)
kit were used to quantify the IL-1
Myasthenia gravis (MG) is an autoimmune disease caused by autoantibodies with neuromuscular junction transmission complications [1]. The global prevalence of MG is (150–250)/million, and the estimated annual incidence rate is (4–10)/million [2]. The incidence rate of MG in China is about 0.68/100,000, and interestingly, the female incidence rate is slightly higher [3]. The major reasons for death with MG included respiratory complications, infections in the lung, etc., with a 14.69% mortality rate in the hospital. It can occur at all ages, with double peaks at the age of 30 and 50 [4, 5]. In China, myasthenia gravis (MG) is diagnosed in up to 50% of children and teenagers [6].
MG, as an autoimmune disease, can lead to deregulated levels of immune factors
associated with anti-inflammation and pro-inflammation [7]. MG patients’ symptoms
have been shown to improve with some therapies that control the ratio of
anti-inflammatory to pro-inflammatory factors [8]. Some of the previous studies
indicated that the occurrence of MG is linked with the aberrant regulation of
immune T cells and inflammatory factors in patients, in which the level of CD
The NLRP3 inflammasome is present primarily in immune and inflammatory cells
following activation by inflammatory stimuli [11]; these cells include macrophages,
monocytes, DCs, and splenic neutrophils [12]. It was found that NLRP3
inflammatory bodies activate caspase-1 and release the downstream cytokine
IL-1
Pentaxin 3 (PTX3) is associated with the initiation and growth of cancer and has been
regarded as a sign of cancer progression [16]. PTX3 regulates the
TLR4/NF-
30 patients with MG were enrolled at designated hospital from May 2019 to July 2020. The blood samples from the patients were collected and immediately stored at 4 °C for further study. Informed consent was obtained from all the participants. The Medical Ethics committee of Beijing Friendship Hospital, Capital Medical University approved (Approve No. KR3234208) the study. All experiments were carried out following standard guidelines and regulations. The diagnosis of MG was made by an experienced neurologist based on clinical presentation, positive response to anticholinesterase medications, presence of autoantibodies, and characteristic electrophysiological findings. Patients with any history of inflammatory or autoimmune diseases such as autoimmune thyroid disease, inflammatory bowel disease, RA, and SLE were excluded from this study.
We built the EAMG model by using the C57BL/6 female mice [20]. Briefly, on day 1, mice were immunized with a synthetic immunogen containing complete Freund’s adjuvant, and on days 30 and 50, mice were immunized with a synthetic immunogen containing incomplete Freund’s adjuvant (SIMGA). After the first injection, we measured the EAMG scores every day; collected the serum; and sacrificed the model mice on the 21st day to conduct other essential experiments. All experiments were performed during the same period of the day (8:00 am to 4:00 pm) to exclude diurnal variations in pharmacological effects. The animals were handled in compliance with the procedures approved by the Animal Resources Centre of Capital Medical University.
We used Light Cycler® 480 SYBR Mix (Roche, Mannheim, Germany) in the LightCycler® 480 real-time PCR system to quantify the genes expression in the PCR system. We utilized the Invitrogen SuperScript double-stranded cDNA synthesis kit for the reverse transcription. NimbleGen’s one-color DNA labeling kit was used to synthesize double-stranded cDNA and then array hybridization was executed by using the NimbleGen hybridization system. In addition, the NimbleGen wash buffer kit was used for washing step. We employed GenePix4000B (Axon, Scottsdale, AZ, USA) and GenePix Pro 6.0 software (Axon, USA) microarray scanner (Molecular Devices) technology to scan the experimental outcomes. The sequences of primers for PTX3 gene used is Forward 5′-CCTGCGATCCTGCTTTGTG-3′, Reverse 5′-GGTGGGATGAAGTCCATTGTC-3′.
Mouse myoblasts (C2C12) was cultured in RPMI 1640 medium (Gibco, Carlsbad, CA,
USA) with 10% fetal calf serum supplementation (FCS, Gibco, Carlsbad, CA, USA)
in a humid environment of 5% CO
We quantify the level of IL-1
Proteins were extracted from the cultured cells using RIPA and PMSF reagents (Beyotime, Beijing, China).We separated the protein lysates according to the molecular weight of respected protein on SDS/PAGE gels and then we transferred them to a Polyvinylidene Fluoride (PVDF, Millipore, Burlington, MA, USA) membrane system. The membranes containing protein were then blocked by using 5% non-fat-milk for 2 hours at room temperature and further, the membranes were placed on shaking incubator at 4 °C and incubated with anti-antibody of PTX3 (Abcam, ab90806), actin (Sigma, St. Louis, MO, USA, SAB4502543), Fl-GSDMD (Abcam, ab210070), N-GSDMD (Abcam, ab215203), NLRP3 (Abcam, ab263899), p-STAT3 (Abcam, ab30647), STAT3 (Abcam, ab68153). After incubation, the membranes were washed with 10% phosphate buffer saline (PBS) for ten minutes and repeated the washing steps three times. After that, membranes were incubated with a secondary antibody for 1 hour at room temperature and again repeated the washing step. The membranes were then visualized by machine.
Microarray data were obtained from NCBI’s Gene Expression Omnibus (GEO)
database. All available studies were considered, but only investigations that
included a DA/Ao comparison were selected for analysis. One-way ANOVA test was
used to analyze contrasts of interest, namely vessel type, to generate lists of
DEGs between DA and Ao. Permissive DEG lists (fold change
We fixed the cells by using 4% paraformaldehyde, permeabilized with 0.5% Triton X-100 in phosphate-buffered saline (PBS) for 15 minutes, and then the cells slides were blocked using the 5% BSA for 30 minutes at 37 °C. We then treated the cells with primary antibodies at 4 °C overnight. Then, incubated the cells by using the Cy3-conjugated goat anti-rabbit or goat anti-mouse IgG DyLight 488-conjugated secondary antibodies for 2 hours at room temperature. Furthermore, we stained the nuclei of the cells using the DAPI and we utilized a fluorescent illumination microscope (Olympus IX71, Tokyo, Japan) for observing the cells.
DNA was amplified using a methylation-specific primer set, PTX3-MF: 5′-CGTTTGCGGTTAGGAGTATTC-3′ and PTX3-MR: 5′-CAAAACGTCGTCCGTAACTTA-3′, or a non-methylation-specific primer set, PTX3-UF: 5′-TGTGT TTGTGGTTAGGAGTATTTG-3′ and PTX3-UR: 5′-CAA AACATCATCCATAACTTA-3′ [18], in a total volume of 20 µL, using 0.5 units of hot-start Taq-polymerase (Takara, Japan) per reaction. The size of the non-methylated amplicon was 105 bp, and the methylated amplicon was 103 bp.
We employed GraphPad 8.0 Software for analyzing the data and presented the data
as mean
To investigate the expression level of the PTX3 gene in MG, we used the PCR assay to identify the PTX3 expression level in patients with MG and mice. Our results revealed that the level of the PTX3 mRNA expression is significantly elevated (Fig. 1A). Serum mRNA expression of PTX3 showed a significant positive correlation with the serum IL-17 levels in patients with MG and the AUC = 0.9236 (Fig. 1B,C). In mice models of MG, we found an elevated level of PTX3 mRNA and protein quantity (Fig. 1D,E).
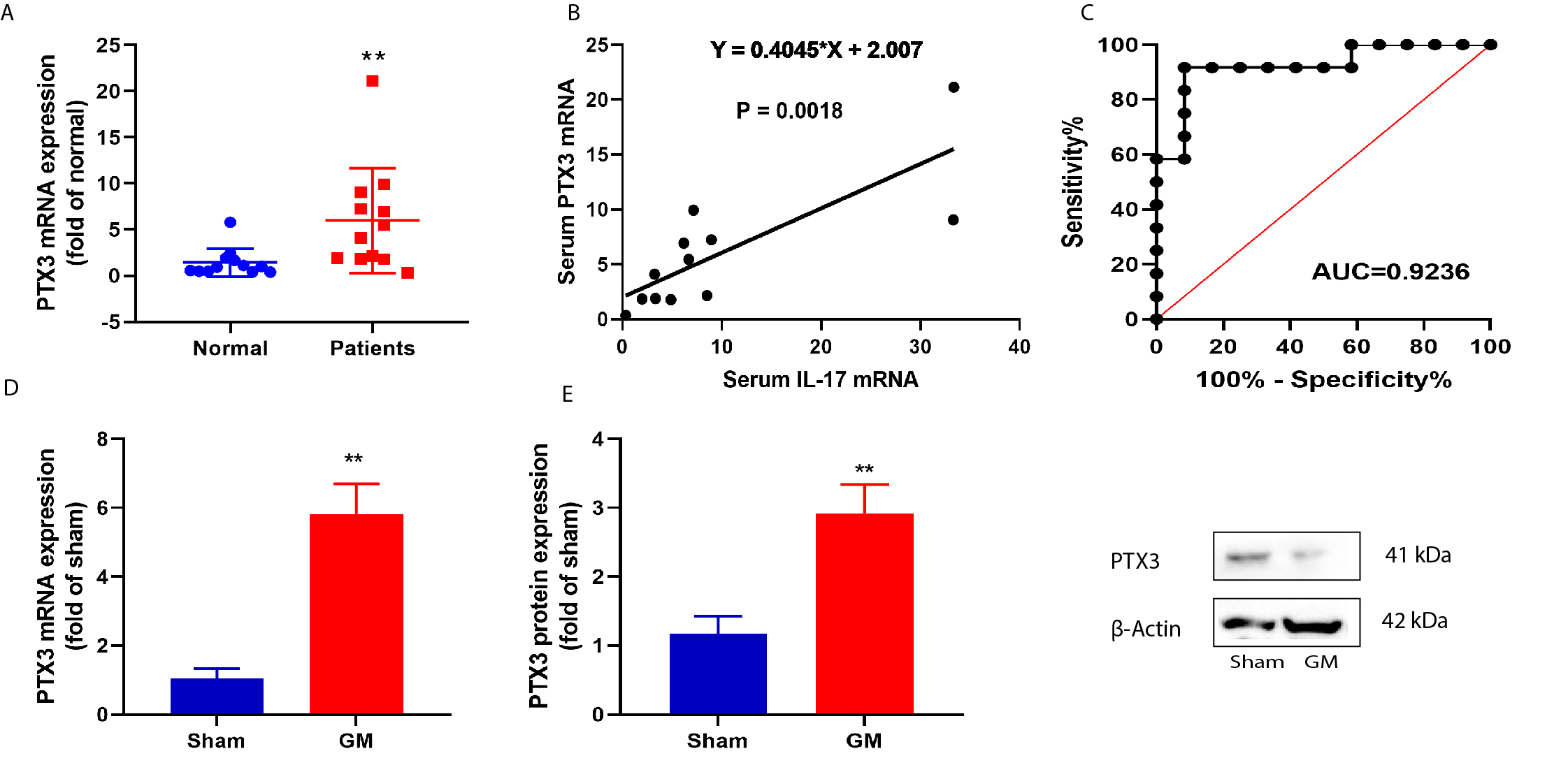 Fig. 1.
Fig. 1.Serum pentaxin 3 (PTX3) expression level in patients with Myasthenia gravis (MG) up-regulated.
PTX3 mRNA expression (A), Serum mRNA expression of PTX3 was in negative
correlation with serum IL-17 levels (B), AUC analysis results (C), Sensitivity
analysis in patients with MG (D), PTX3 mRNA expression (E), while the western
blot figure indicate protein expression in mice model. The experiments were
performed in triplicate and data is presented an average of these experiments.
Normal, normal volunteers’ group; Patients, Patients with MG; Sham, sham control
group; MG, mice with MG. **p
Next, we analyzed the correlation between PTX3 expression level and inflammation
level in MG models. PTX3 plasmid increased PTX3 mRNA expression and enhanced the
inflammatory factors, including IL-1
 Fig. 2.
Fig. 2.PTX3 promoted inflammation level using in vitro model.
PTX3 mRNA expression (A). IL-1
We next further examined the function of PTX3 in MG using PTX3 recombinant
protein. PTX3 recombinant protein increased global clinical score and AChR level
and expanded anti-TAChR IgG/IgG1/IgG2b/IgG2c levels, induced Ccl12, Ccl19, and
Ccl13 mRNA expressions, while increased the levels of inflammatory regulators,
including IL-1
 Fig. 3.
Fig. 3.The inhibition of PTX3 presented MG and reduced inflammation
levels in the mice model. Global clinical score (A). AChR level (B). anti-TAChR
IgG (C)/IgG1 (D)/IgG2b (E)/IgG2c (F) levels. Ccl12 (G), Ccl19 (H) and Ccl13
(I) mRNA expressions. IL-1
Furthermore, we analyzed the regulatory roles of PTX3 on pyroptosis in the in-vitro model of MG. PTX3 up-regulation decreased cell proliferation and increased the activity of LDH in the in-vitro model (Fig. 4A,B). Down-regulation of PTX3 enhanced the cellular proliferative charactertristics and decreased the activities of LDH in the in-vitro model (Fig. 4C,D). Additionally, we discovered in the in-vitro model that downregulating PTX3 decreased the protein level of GSDMD while upregulating PTX3 increased the expression of GSDMD (Fig. 4E,F). Furthermore, in the mice model, PTX3 recombinant protein promoted GSDMD protein expression (Fig. 4G).
 Fig. 4.
Fig. 4.PTX3 promoted pyroptosis in mice model or in vitro model. Cell
growth and LDH activity levels (A and B) in vitro model by
over-expression of PTX3. Cell growth and LDH activity levels (C and D) in
vitro model by down-regulation of PTX3. GSDMD protein expression in
vitro model by over-expression of PTX3 (E). Full-length GSDMD (Fl-GSDMD) and
N-terminal domain of GSDMD (N-GSDMD) protein expression in vitro model
by down-regulation of PTX3 (F). GSDMD protein expression in mice with MG by sh-
PTX3 (G). The experiments were performed in triplicate and data is presented an
average of these experiments. **p
To understand the molecular regulatory role of PTX3 in the MG model, we used microarray technology to perform gene expression profiling (Fig. 5A,B). Our results indicated that PTX3 might regulate STAT3 and NLRP3 levels (Fig. 5B). The recombinant protein product of PTX3 stimulated the expression of p-STAT3/NLRP3 protein in the mice model (Fig. 5C). In the mice model of MG, PTX3 up-regulation induced PTX3 and p-STAT3/NLRP3 protein expressions (Fig. 6A) while, PTX3 down-regulation suppressed PTX3 and p-STAT3/NLRP3 protein expressions in vitro model (Fig. 6B). Confocal showed that PTX3 up-regulation increased PTX3 and p-STAT3 expression levels in vitro model (Fig. 6C). We searched that the gene promoter of PTX3 and STAT3, showing the molecular interaction between PTX3 and STAT3 (Fig. 7A,B). Based on the luciferase reporter assay, the PTX3 WT substantially correlated with the expression of STAT3 (Fig. 7C). Then, the STAT3 inhibitor (2 µM of STAT3-IN-10) downregulated the protein level of p-STAT3 and reduced the protein expression level of GSDMD/NLRP3 in the in-vitro model (Fig. 7D). Moreover, the STAT3 agonist (5 µM of Oleoylcarnitine) induced the protein level of p-STAT3 and stimulated the protein level of GSDMD/NLRP3 in the in-vitro model (Fig. 7E).
 Fig. 5.
Fig. 5.PTX3 induced STAT3/NLRP3 inflammasome. Heat map (A). Result
workflow and analysis (B). Presentation of p-STAT3/NLRP3 protein expression in
mice with MG (C). **p
 Fig. 6.
Fig. 6.PTX3 promoted gene synthesis of STAT3. PTX3/p-STAT3/NLRP3
protein expression in the in-vitro model by over-expression of PTX3 (A).
PTX3/p-STAT3/NLRP3 protein expression in vitro model by
down-regulation of PTX3 (B). PTX3/p-STAT3 expression by confocal microscope with the scale bar of 10 μM (C).
The data is presented an average of triplicated experiments. **p
 Fig. 7.
Fig. 7.Gene promoter of PTX3 and STAT3. The gene promoter of PTX3 and
STAT3 (A), molecular interaction within PTX3 and PTX3 (B), luciferase reporter
(C), p-STAT3/GSDMD/NLRP3 protein expression in vitro model by PTX3+STAT3
inhibitor (D), p-STAT3/GSDMD/NLRP3 protein expression in vitro model by
sh-PTX3+STAT3 agonist (E). All experiments were performed in triplicates and the
data is presented an average of these experiments. **p
We further examined the mechanism of PTX3 to check the progression of MG. We found that serum mRNA level of PTX3 was positively correlated with the level of METTL3 in MG patients (Fig. 8A). However, PTX3 level (serum mRNA) is not correlated with METTL14 levels in MG patients (Fig. 8B). In the immunoprecipitated fraction, an elevated level of PTX3 was found by using the m6A antibody (Fig. 8C). METTL3 reduced the m6A methylation level of PTX3 (Fig. 8D). Si-METTL3 decreased PTX3 expression levels (Fig. 8E), indicating the stable stimulation of m6A methylation of PTX3. The suspicious methylation modification sites of PTX3 are present at the stop codon region (Fig. 8F). METTL3 reduced PTX3 mRNA levels expression levels (Fig. 8G). In conclusion, these results suggest that METTL3-mediated m6A modification decreases PTX3 stability.
 Fig. 8.
Fig. 8.METTL3-mediated m6A modification PTX3 stability. Serum mRNA of
PTX3 was in negative correlation with METTL3 or METTL14 levels
in patients with MG (A and B), fraction immunoprecipitated by m6A antibody (C),
m6A methylation level of PTX3 (D), presentation of PTX3 expression levels (E),
multiple suspicious methylation modification sites (F), PTX3 mRNA levels
expression levels (G). The experiments were performed in triplicate and the data
is an average of these expeirments. **p
MG, an autoimmune disorder, is crucially regulated by immune mediators, including antibodies of acetylcholine receptor, complement, and cytokines. In neuromuscular junction, the postsynaptic membrane acetylcholine receptors are substantially associated with the development of MG [21]. It is often manifested as pathological fatigue of partial or systemic skeletal muscle [22]. The development of MG is related to the change in the expression level of inflammatory factors, which participate in the pathogenesis of MG through immune regulation [23, 24]. Here in, we quantified the elevated level of serum PTX3 in MG patients. A study concluded that dysregulated PTX3 contributes to ischemia and reperfusion injury [25]. Thus, it indicates that the PCBP2 regulates the disease progression of MG.
IL-1
The development of MG and the expression level of inflammatory factors are
activated, and participate in the response of cell apoptosis, while the STAT3
regulatory signal is associated with the inflammatory and cell death process
mediated by MG [32, 33, 34]. The development of myasthenia gravis has been shown to
regulate inflammatory response through the STAT3 signaling pathway. STAT3 can
activate the IL-1
In the end, the first time, we demonstrated that the expression level of PTX3 is elevated in patients with MG. PTX3 promoted pyroptosis and inflammation in the progression of MG through STAT3/NLRP3 signaling pathway. Moreover, PTX3 represents a potential therapeutic target/spot for the treatment of MG. Finally, this research discovered the mechanisms at molecular levels for suppressing pyroptosis and inflammation in patients with MG.
Our study suggests that the PTX3 is associated with the enhancement of inflammation and pyroptosis through regulating the STAT3/NLRP3 inflammasome signaling pathway at the early stage of the disease. The pro-inflammatory PTX3 facilitates the development of MG and it can be used as a potantial MG-associated diagnostic biomarker for MG.
The data used to support this study is available from the corresponding author upon request.
Conceptualization, YZ; Data curation, YPL; Formal analysis, YL, SW and YPL; Investigation, YL and SW; Methodology, YL and YZ; Resources, YZ; Software, YL and SW; Validation, YZ; Writing – original draft, YL, SW, YPL and YZ; Writing – review & editing, YZ. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
This study was approved by The Medical Ethics committee of Beijing Friendship Hospital, Capital Medical University (Approval No. KR3234208). Informed consent was obtained from all the participants.
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
