1 School of Life Sciences, Anhui Agricultural University, 230036 Hefei, Anhui, China
2 Faculty of Life Sciences, University of Strasbourg, 67000 Strasbourg, France
3 Zhangzhou Affiliated Hospital of Fujian Medical University, 363000 Zhangzhou, Fujian, China
†These authors contributed equally.
Abstract
Background: Drosophila ubiquitin carboxy-terminal hydrolase L5 (Uch-L5) functions as a critical component of the 26S proteasome to mediate degradation of polyubiquitinated proteins. It was recently shown to modulate tissue/organ development by targeting the Smoothened protein in the hedgehog pathway. However, whether it plays a role in controlling organismal immune response remains largely unknown. Methods: Reverse transcription plus quantitative polymerase chain reaction (RT-qPCR), dual-luciferase, and Western blot assays were used to explore the potential function of Uch-L5 in the innate immune regulation in cultured Drosophila S2 cells. Further genetic manipulations and bacterial infections were conducted to confirm the findings in vivo. Results: Silencing of Uch-L5 antagonizes the immune deficiency (IMD) but not the Toll innate immune signaling both in vitro and in vivo. Moreover, Uch-L5 positively contributes to the Drosophila innate immune response via its N-terminal Uch domain, which is the catalytical triad executing its deubiquitinase activity. Conclusions: Our studies shed light on a novel function of the deubiquitinase Uch-L5 in governing the anti-microbial defense in Drosophila.
Keywords
- Uch-L5
- IMD signaling
- innate immune response
- Drosophila melanogaster
Innate immunity reacts quickly and efficiently upon recognition of extracellular pathogenic invasion. This constitutes the first line of defense of most organisms [1]. Pattern recognition receptors (PRRs) have been suggested to be mainly responsible for recognizing pathogenic stimuli, in order to trigger downstream signaling transductions, thereby inducing expressions of various immune effectors [2, 3]. Several pioneering studies have demonstrated the pivotal roles of adaptor proteins and correlative modulators in the signal transductions from transmembrane and intracellular receptors to secreted effectors [reviewed in [1, 2, 3, 4, 5, 6]]. Identification and characterization of novel adaptors/modulators has always been one of the hotspots in the basic research of innate immunity.
Drosophila melanogaster (fruit fly) is an excellent animal model for uncovering novel modulators in the innate immune signaling pathways, due to its powerful genetic approaches and worldwide resources of Drosophila mutants and transgenes. The fly innate immune response is mainly governed by two signaling pathways, namely the Toll and the immune deficiency (IMD) pathways [7, 8, 9], which are similar to the mammalian Myd88 (myeloid differentiation factor 88)-dependent TLR (Toll-like receptor) and TNFR (tumor necrosis factor receptor) signaling pathways, respectively. Upon infection by most fungi or Gram-positive bacteria, the Toll receptor-involved signaling is activated in order to trigger the induction of downstream anti-microbial peptides (AMPs), for instance drosomycin (Drs) and metchnikowin (Mtk) [10]. On the other hand, the IMD signaling pathway is mostly activated by Gram-negative bacteria and some types of Gram-positive bacteria, resulting in secretion of another set of AMPs like attacins (Att), cecropin (Cec), and diptericin (Dpt) for the host immune defense [11].
The IMD signaling pathway is normally activated by the binding of
bacteria-derived DAP (meso-diaminopimelic acid)-type PGN (peptidoglycan) to the
PGN recognition proteins PGRP-LC (located on the cell membrane) and/or PGRP-LE
(intracellular) [12]. This binding further leads to the recruitment of the
adaptor protein, Imd, to the receptor, together with Fadd (Fas-associated death
domain) and the caspase Dredd (death related ced3/nedd2-like caspase) [13]. Dredd
participates in the cleavage of Imd and the activation of the downstream kinase
complex Tak1 (TGF-
Uncontrolled IMD signaling is detrimental to flies [21]. The maintenance of IMD homeostasis requires a series of regulators, among which are the ubiquitin-related enzymes including E3 ligases and Dubs (deubiquitinases). For instance, E3 ligases including Dnr1 (defense repressor 1) [22], Faf (fat facets) [23], Posh (plenty of SH3s) [24], and SkpA (Skp1-related A) [25] negatively contribute to IMD signaling through mediating the ubiquitin assembly of targeted substrates. Additionally, a Dub Usp36 (ubiquitin-specific protease 36) associates with Imd to antagonize its K63-linked ubiquitination, thereby down-regulating the IMD signals [26]. Some other Dubs such as Cyld (cylindromatosis) [27], Trabid (Traf-binding domain) [28, 29], and Otu (ovarian tumor) [30] have been found to limit excessive activation of the IMD pathway under different physiological conditions. Recently, we noted that the ubiquitin carboxy-terminal hydrolase L5 (Uch-L5, also known as Uch37) functions as a typical Dub (belonging to the Uch sub-family of Dubs) to positively regulate Hh (hedgehog) signaling by restricting ubiquitination-mediated degradation of Smo (smoothened) [31]. However, little is known about the immunological role of Uch-L5 in the fly defense against microbial challenges.
In the present study, we focused mainly on exploring the potential role of Uch-L5 in regulating Drosophila innate immunity. We show that silencing of Uch-L5 results in markedly decreased IMD signaling, whereas ectopic expression of Uch-L5 behaves oppositely. The N-terminal Uch domain is both required and sufficient for Uch-L5 positively impacting on IMD signaling, suggesting that Uch-L5 executes its immune function relying on the Dub enzymatical activity. We further provide evidence displaying that Uch-L5 is essential for the immune defense of adult flies upon bacterial infections. Collectively, our results uncover a novel biological role of Uch-L5 in mediating the IMD innate immune reaction in Drosophila.
Flies (Drosophila melanogaster) were raised at 25 °C with a 12
h of light/dark cycle and 65% humidity. Standard Drosophila medium
(6.65% cornmeal, 7.15% dextrose, 5% yeast, 0.66% agar, 2.2% nipagin, and 3.4
mL/L propionic acid) was used to feed all flies. The w
All dsRNAs were synthesized according to methods described previously [33]. In brief, DNA templates were amplified using specific primers (Supplementary Table 1). PCR products were then combined with 1/10 volume of 5 M NaCl and 2.5-fold volume of EtOH, followed by centrifugation for 10 min at high speed. The pellet was washed with 75% EtOH and dissolved in RNase/DNase-free water. The T7 in vitro transcription kit (Promega, Cat#P1300, Madison, Wisconson, USA) was used for dsRNA synthesis and purification according to the manufacture’s protocol. Purified dsRNAs were diluted in RNase/DNase-free water to a final concentration of 1 µg/µL. The quality of dsRNAs was confirmed by agarose gel electrophoresis (Supplementary Fig. 1A). For gene silencing in S2 cells, dsRNA at the amount of 3 µg was added directly into S2 cells for 48 h.
S2 cells were cultured in a 27 °C incubator using insect medium (Gibco)
with 10% FBS (Hyclone). The MycoAlert
S2 cells or fly samples were homogenized in Trizol Reagent (Invitrogen, Cat#15596026, Waltham, Massachusetts, USA). Chloroform solution (1/5 volume) was added into the sample, followed by intense votexing for 15 sec, and centrifugation at high speed for 15 min. The upper layer of the supernatant was transferred into a fresh tube and an equal volume of isopropanol was added. After centrifugation again at high speed for 10 min, the pellet was washed with 75% EtOH and diluted in RNase/DNase-free water. Samples were further incubated with DNase for 30 min, and the quality of RNA was assessed by examining the 260:280 ratio, and the pattern in the agarose gel electrophoresis assay. The first-strand cDNA synthesis kit (Transgen, Cat#AT341-01, Beijing, China) was used to reverse-transcribe RNA (1 µg) into cDNA. Quantitative PCR assays (in 3 technical repetitions) were performed using a SYBR Green Master Mix (Thermo Fisher, Cat#A46012, Waltham, Massachusetts, USA) in the Lightcycler 480 PCR platform (Roche, Cat#05015278001, Basel, Halbkanton, Switzerland). Rp49 was used as the internal control. Results were analyzed based on data from 7 independent biological replications. The primers used in RT-qPCR experiments are shown in Supplementary Table 1.
Infection experiments were performed as previously described [36]. In brief,
overnight bacterial cultures were harvested and diluted in sterile PBS at a
concentration of OD
All statistical analyses were performed by using GraphPad Prism 9 (v. 9.4.0,
Dotmatics, Boston, MA, USA). Data were shown as means plus standard errors.
Statistical significance was determined by using the ANOVA or Mann-Whitney tests
except for survival assays, in which the Log-Rank test (Kaplan-Meier method) was
used. The p value of less than 0.05 was considered statistically
significant. * p
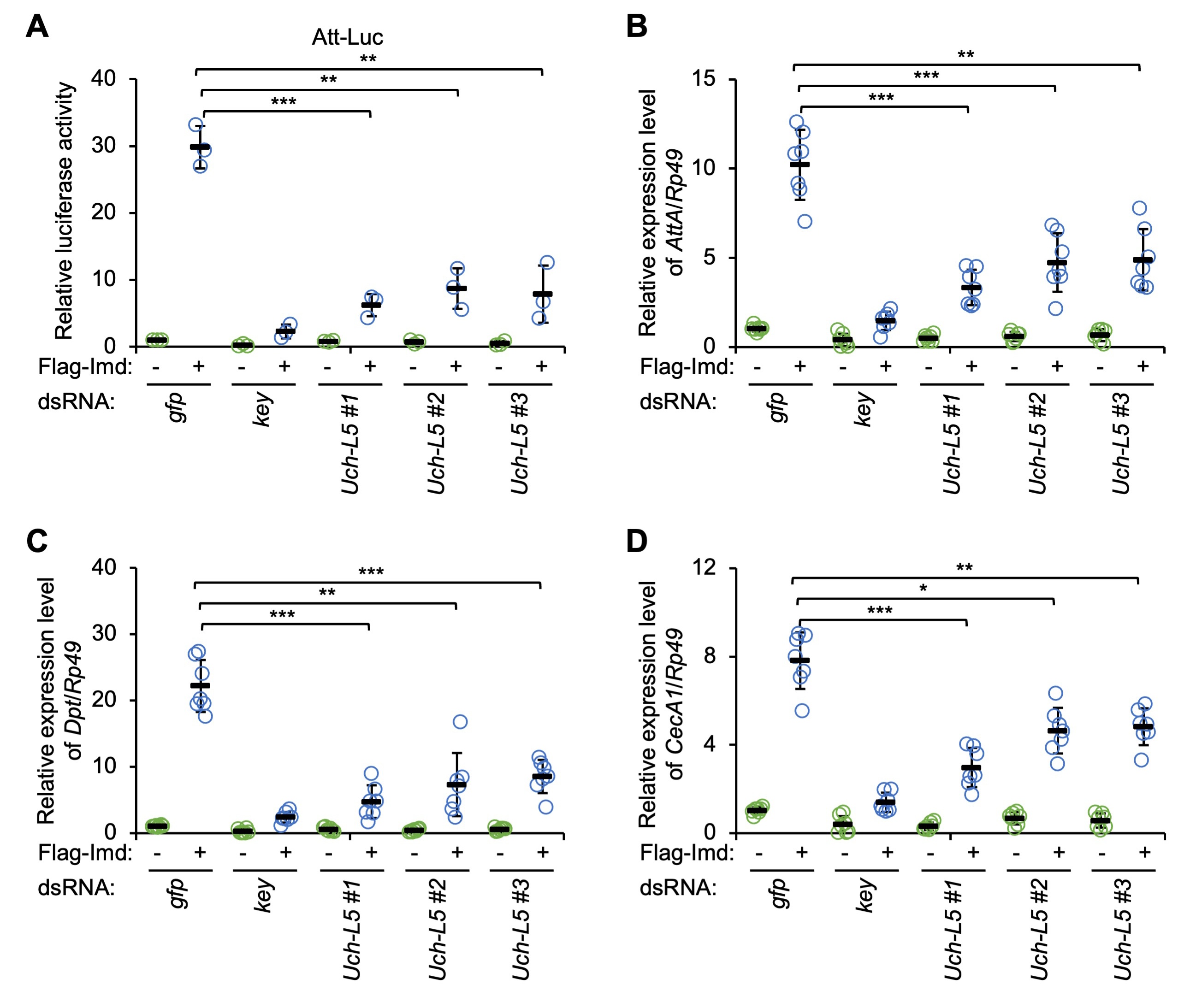
We examined the potential role of Uch-L5, a previously described Dub modulating Hh signaling [31], in affecting the innate immune reaction in Drosophila. To do this, we first designed 3 types of dsRNAs that targeted different regions of the coding sequence of Uch-L5 (referred to as Uch-L5 dsRNAs #1, #2, and #3, respectively, Supplementary Fig. 1A). Cultured Drosophila S2 cells were treated with these Uch-L5 dsRNAs (dsRNA that targeted GFP was used as the control), and the knockdown efficiency was monitored by RT-qPCR assays (Supplementary Fig. 1B). As illustrated in Fig. 1A, silencing of Uch-L5 resulted in drastic decreases (by ~75% to ~80%) of the Att-Luc activities upon Imd over-expression, indicating that Uch-L5 is a potentially positive regulator in the IMD signaling pathway in cultured S2 cells. Further, we performed RT-qPCR assays to detect the endogenous inductions of the AMPs that are downstream of the IMD pathway, including AttA, Dpt, and CecA1. We obtained consistent results: down-regulation of Uch-L5 significantly prevented the Imd-driven transcription of several AMP genes (Fig. 1B–D).
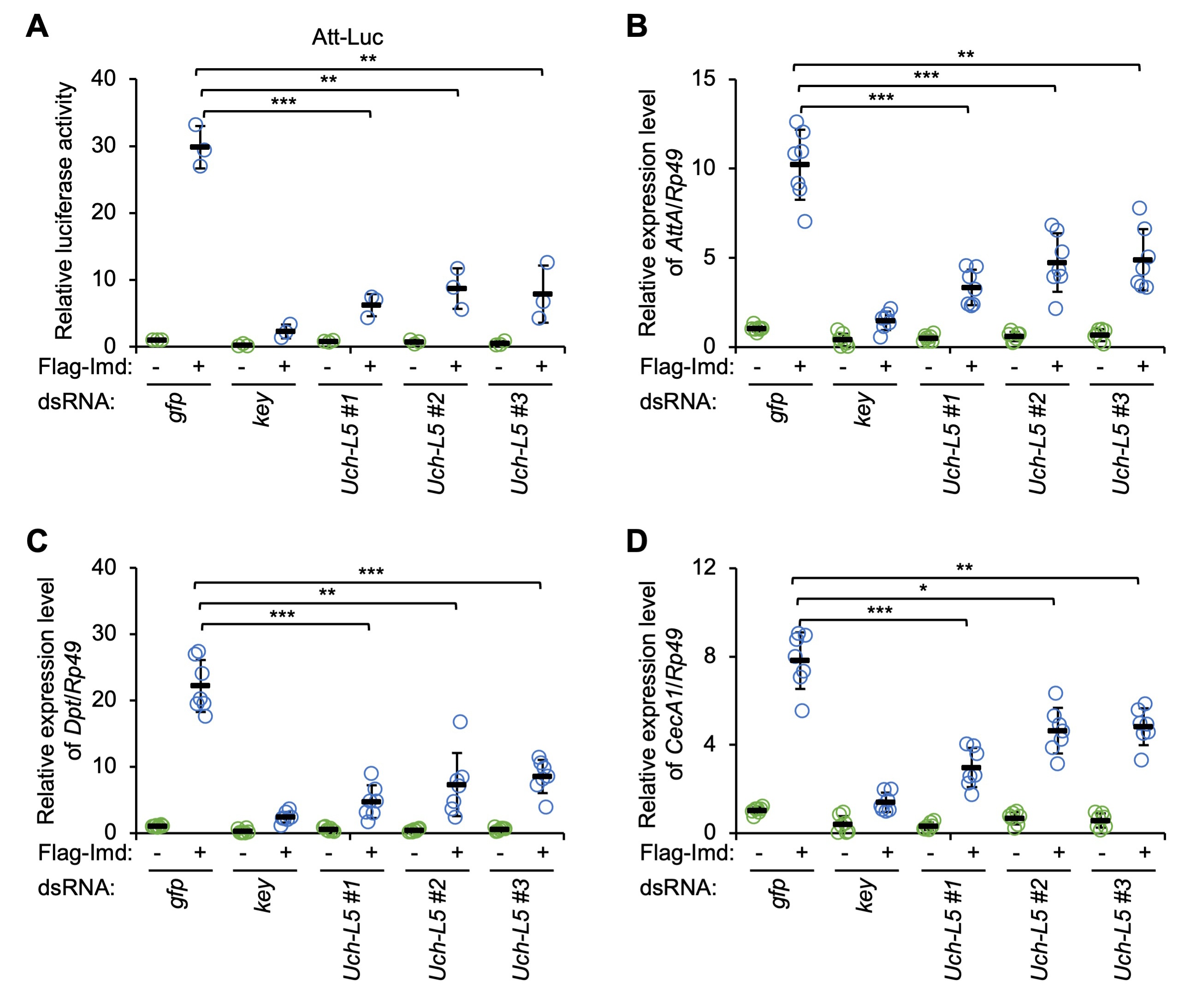 Fig. 1.
Fig. 1.Silencing of ubiquitin carboxy-terminal hydrolase L5 (Uch-L5) reduces immune deficiency (IMD) signaling in Drosophila
S2 cells. (A) S2 cells were pretreated with indicated dsRNAs. 48 h later, cells
were transfected with various combinations of expressing plasmids, followed by
dual-luciferase assays. (B–D) S2 cells were pretreated with dsRNAs as in (A) for
48 h. Cells were then transfected with indicated expressing plasmids for 36 h and
harvested for RT-qPCR assays to detect the mRNA levels of AttA (B),
Dpt (C), and CecA1 (D). In (A–D), each dot represents one
independent replicate and data are shown as means plus standard errors. *
p
As mentioned in the Introduction, the Drosophila innate immune response is mostly governed by two signaling cascades, the Toll and the IMD pathways. We then conducted another luciferase reporter assay (Drs-Luc) in S2 cells, as previously reported, aiming to test whether Uch-L5 is also involved in affecting Toll signaling. However, we did not observe apparent alterations in the Myd88-driven Drs-Luc activities when Uch-L5 was knocked down, compared to that in the control (Supplementary Fig. 2A). Similar results were obtained when we looked at the transcript levels of Drs and Mtk, two well-known downstream AMP genes in the Toll pathway (Supplementary Fig. 2B,C). These data indicated that Drosophila Dub Uch-L5 is specifically involved in controlling the IMD other than the Toll innate immune signaling pathway in S2 cell cultures.
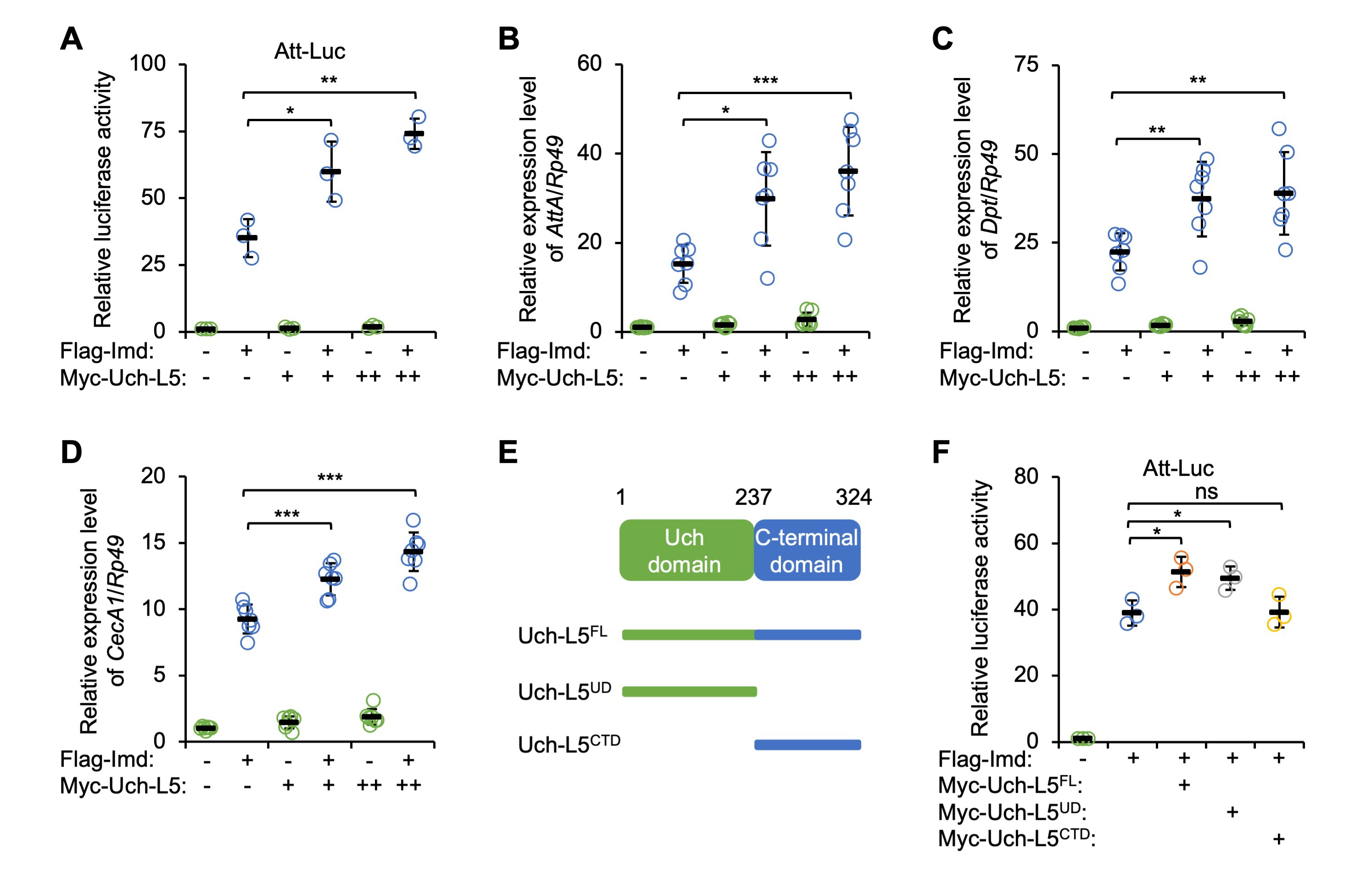
We collected more evidence on the functional role of Uch-L5 in controlling IMD innate immunity by constructing a plasmid expressing Myc-tagged Uch-L5 in S2 cells. By again using the Att-Luc reporter system, we observed that ectopic expression of Myc-Uch-L5 enhanced IMD signaling in a dose-dependent manner (Fig. 2A). Accordingly, co-transfection of Uch-L5 expressing plasmid resulted in increased inductions (by ~32% to ~136%) of the IMD-downstream AMPs upon Imd over-expression (Fig. 2B–D), which indicated a positive role of Uch-L5 in modulating IMD signaling.
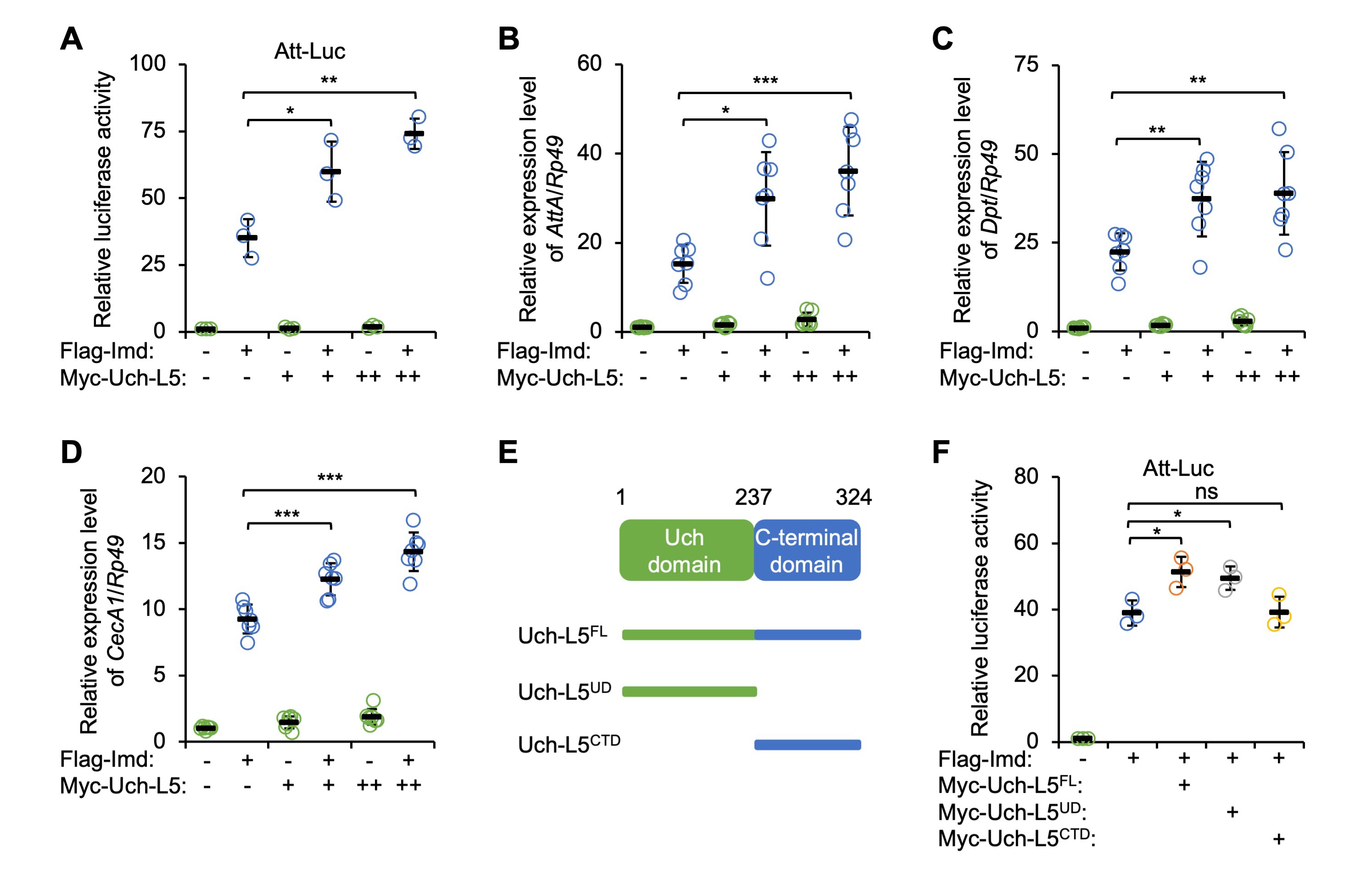 Fig. 2.
Fig. 2.Uch-L5 affects IMD signaling in a Uch domain-dependent manner.
(A–D) S2 cells were transfected with expressing plasmids as indicated, followed
by dual-luciferase assays (A) or RT-qPCR assays to examine the expression levels
of AttA (B), Dpt (C), and CecA1 (D). (E) Domain
analysis of Uch-L5. (F) S2 cells were transfected with indicated combinations of
expressing plasmids, followed by dual-luciferase assays. In (A–D,F), each dot
represents one independent replicate and data are shown as means plus standard
errors. * p
A recent study that explored the biochemical characteristic of Uch-L5
demonstrated that Uch-L5 harbors a typical Uch domain at its N-terminus, which is
the catalytical region for its Dub enzymatical activity [31]. We therefore
constructed two kinds of plasmids that expressed truncated forms of Uch-L5,
including Uch-L5
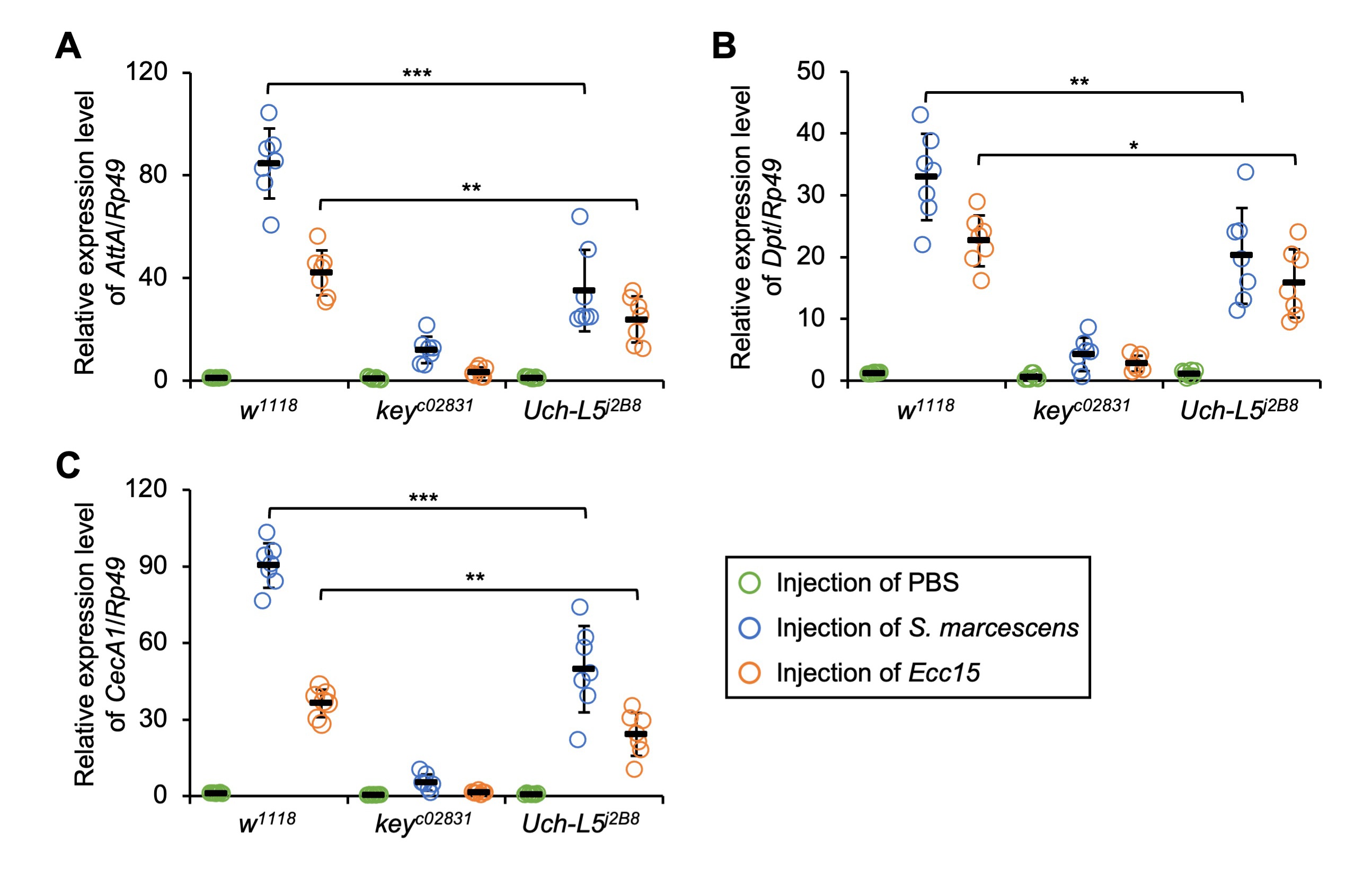
We next sought to decipher the immune function of Uch-L5 in vivo. To
this end, we collected w
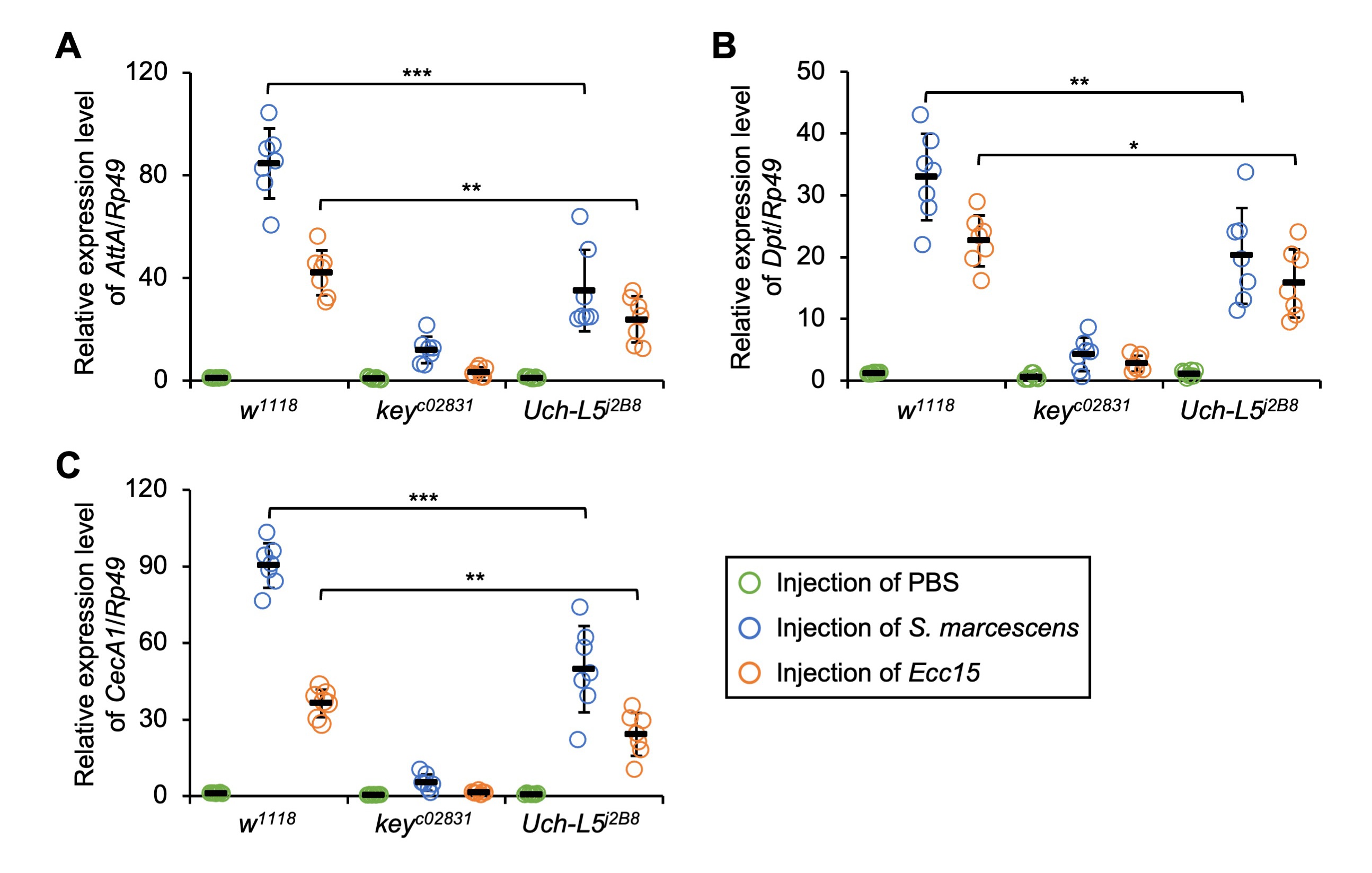 Fig. 3.
Fig. 3.Loss of Uch-L5 antagonizes IMD downstream AMP
expressions. (A–C) w
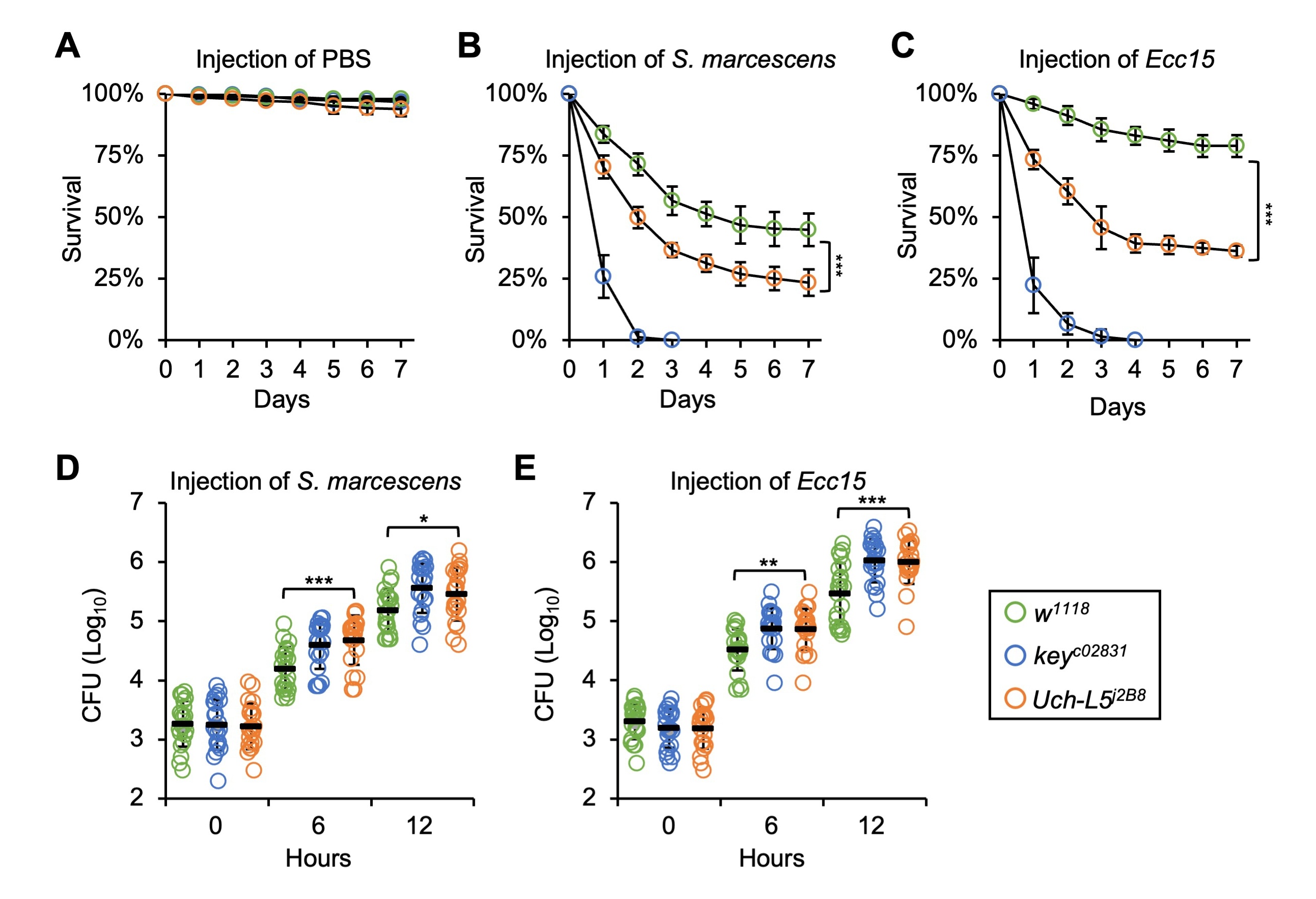 Fig. 4.
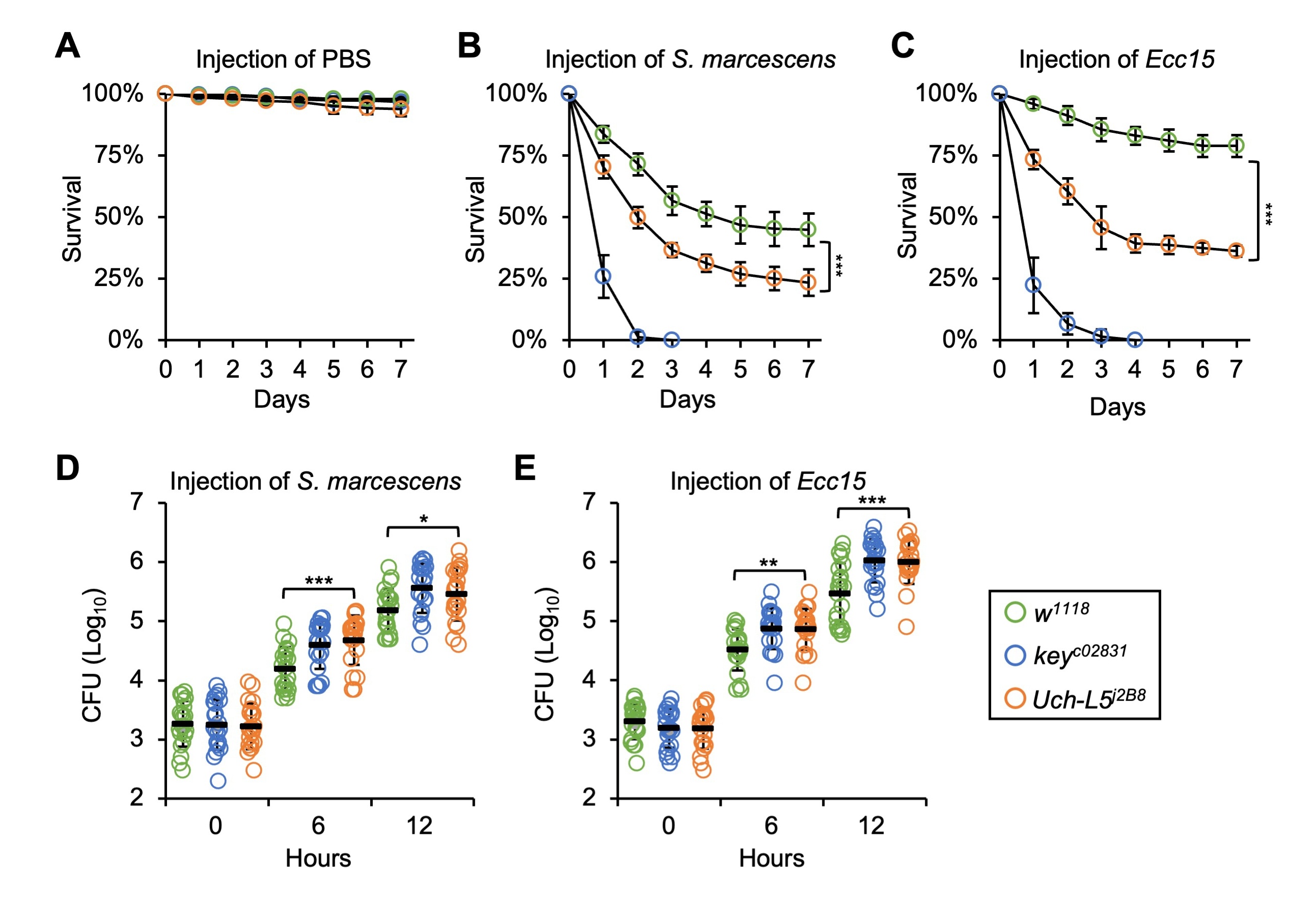
Fig. 4.Uch-L5 plays a critical role in the fly anti-microbial
defense. (A–C) w
To test the involvement of Uch-L5 in affecting the proliferation of
injected pathogens (S. marcescens or Ecc15), we performed CFU
(colony-forming-units) assays at different time points (0, 6, and 12 h) after
injection. As shown in Fig. 4D,E, the burdens of S. marcescens and
Ecc15 in Uch-L5
Fat body is one of the main responsible immune tissues/organs during systemic
infection in Drosophila. As the Uch-L5 transcript is relatively
abundant in the fat body cells, according to the high-throughput sequencing or
array data in the FlyBase website (https://flybase.org/reports/FBgn0011327), we
sought to investigate the functional role of Uch-L5 in
Drosophila fat body. Two different Uch-L5 RNAi (RNA
interference) flies (Uch-L5 RNAi #1 and #2, for detailed
information, see Materials and Methods) were crossed with the fat body-specific
driver c564-gal4. The Tub-gal80
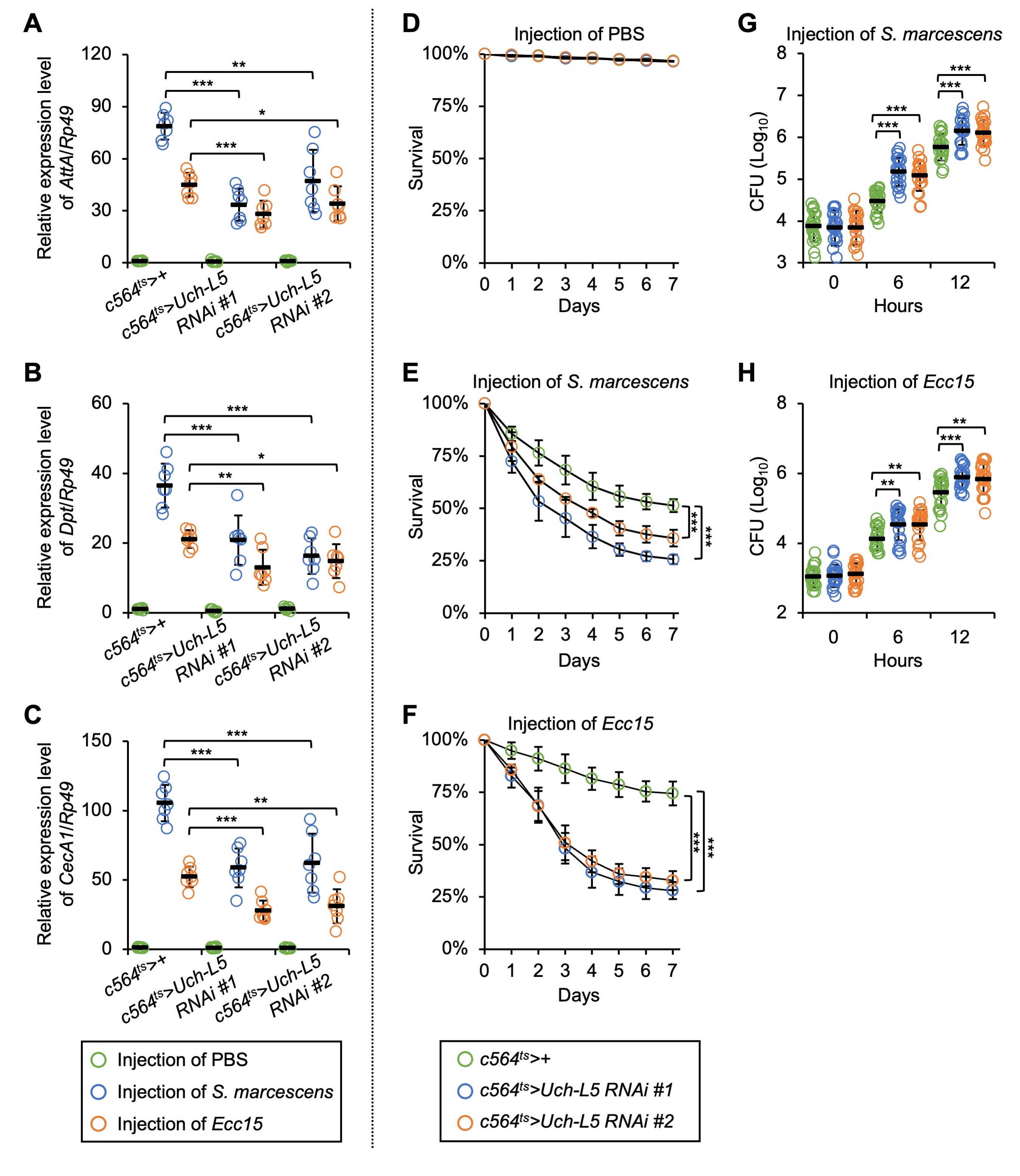
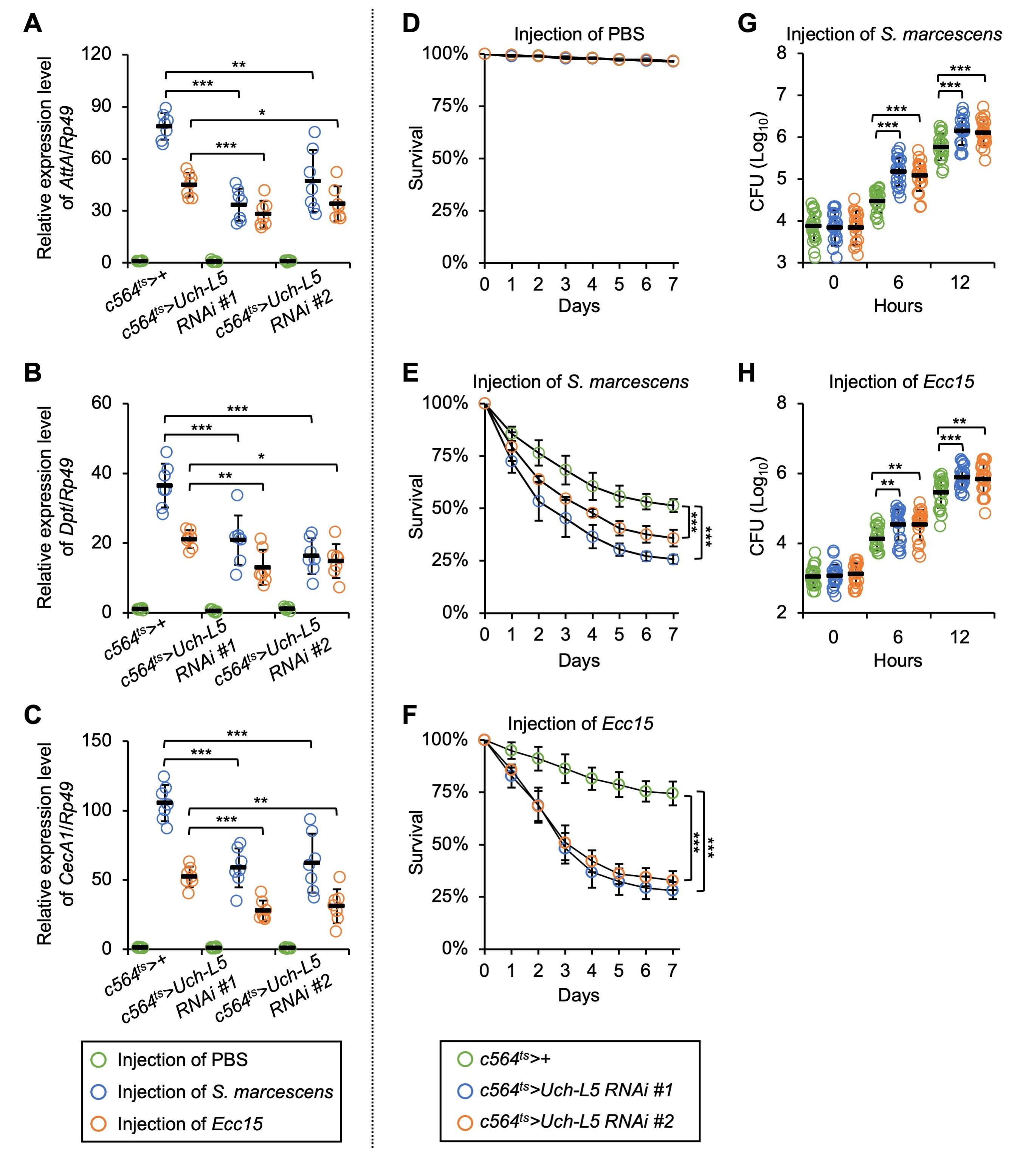 Fig. 5.
Fig. 5.Silencing of Uch-L5 in fat body results in immune
defects. (A–H) c564
Drosophila Uch-L5 belongs to the Uch sub-family of Dubs [31, 37]. Dub-mediated cleavage of ubiquitin/poly-ubiquitin from ubiquitinated substrates has been widely studied and demonstrated to be involved in a broad range of cellular processes [reviewed in [38, 39, 40]]. However, our knowledge regarding the biological function of the fly Uch-L5 is incomplete. In the present study, we carried out a series of investigations using both in vitro and in vivo models. We showed that Uch-L5 behaves as a positive modulator in the fly IMD innate immune defense against bacterial stimuli. Our data shed light on a previously undescribed role of Uch-L5 in controlling organismal innate immunity.
How does Uch-L5 execute its essential role in regulating the IMD signaling
pathway? A recent study by Zhou and colleagues illustrated that Uch-L5 depends on
its N-terminal Uch domain to positively contribute to Hh signaling [31],
highlighting the critical requirement of the Dub catalytical triad for Uch-L5
functioning. Indeed, when we utilized several truncated forms of Uch-L5
expressing plasmids and conducted Att-Luc reporter assays in cultured S2 cells,
we observed that Uch-L5 without the Uch domain no longer promoted IMD signaling.
It is of interest to note that the Dub enzymatical activity of
Drosophila Uch-L5 is somehow autoinhibited by the CTD via
oligomerization, which can be alleviated by the association of a co-factor such
as Rpn13 [31, 37]. However, we would like to conclude that this is not the case
for the regulatory role of Uch-L5 in innate immune regulation, based on the
following reasons: (1) ectopic expression of Uch-L5 without CTD showed a similar
contribution to the IMD signaling as that of the Uch-L5
Recently, numerous efforts have been made to decipher the critical role of Dubs in regulating IMD signaling. To date, only Imd, Tak1, and Dredd in the canonical IMD pathway, have been clearly demonstrated to be modified by ubiquitination/deubiquitination. Whether the other key factors also involve a ubiquitin-mediated modulation remains a mystery. Uch-L5 might therefore target a factor, or some of these factors, to enhance IMD signaling. The 48th lysine (K48)-linked ubiquitination has been suggested to primarily mediate protein recognition and degradation by the 26S proteasome, whereas the 63rd lysine (K63)-linked ubiquitination commonly governs signal transduction [41]. According to our current knowledge and the results of the present study, we speculate that the Dub Uch-L5 may inhibit the K48-linked ubiquitination of its substrate(s), thus improving its (their) stability. Future biochemical approaches, for instance co-immunoprecipitation and ubiquitination assays, will greatly help us further understand the molecular mechanism by which Uch-L5 benefits the IMD innate immune defense in Drosophila.
On the other hand, it is somehow lagged off that no Dubs have been shown to play a role in the Toll innate immune defense of Drosophila. This may be due to the fact that ubiquitination modification in Toll signaling is not as important as that in IMD signaling. Even though we failed to observe in the present study any involvement of Uch-L5 in controlling the Toll pathway, we cannot exclude the possibility that the Toll pathway is regulated by other Dubs. One strategy would be to explore Dubs that can physically associate with MyD88, a pivotal Toll downstream adaptor protein that undergoes ubiquitination [33].
We have demonstrated a critical regulatory role of the Dub Uch-L5 in Drosophila IMD innate immunity. The present study has provided insight into the understanding of the precise dynamic modulation of IMD signaling in response to bacterial infection.
The datasets used and analysed during the current study are available from the corresponding author on reasonable request.
SJ and YH designed the research study. CZ, SZ, FK, YX, KS, YJ, JL, AQ, and XZ performed the research. CZ, SZ, FK, and YH analyzed the data. SJ and YH wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript.
Not applicable.
We are grateful for the Bloomington Drosophila Stock Center, the Vienna Drosophila Resource Center, and the Tsinghua Fly Center for fly resources.
This research was funded by the National Natural Science Foundation of China (32100702) and the Anhui Provincial Natural Science Foundation (2008085J14).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.