1 Department of Gastroenterology and Hepatology, The Affiliated Hospital of Southwest Jiaotong University, The Third People's Hospital of Chengdu, Chongqing Medical University, 610031 Chengdu, Sichuan, China
2 Department of Critical Care Medicine, The Affiliated Hospital of Southwest Jiaotong University, The Third People's Hospital of Chengdu, Chongqing Medical University, 610031 Chengdu, Sichuan, China
3 Department of Infectious diseases, The Third Affiliated Hospital of Sun Yat-sen university, 510630 Guangzhou, Guangdong, China
†These authors contributed equally.
Abstract
Background: Hepatitis C virus (HCV) infection is a global health threat
to the public, and vaccines against it are not yet available. The HCV envelope
glycoprotein E2 is a key target for anti-HCV vaccines. The majority of previous
studies have focused on the hypervariable region and the glycosylation sites of
the_HCV structural protein. This study aims to investigate a conserved
domain of HCV E2 glycoprotein and explore its potential to induce an immune
response against HCV. Methods: HCV E2 conserved domain (encompassing
amino acids 505–702) was prepared in Escherichia coli (E. coli).
Peripheral blood mononuclear cells (PBMCs) were isolated from patients with HCV
or healthy controls. Interferon-gamma (IFN-
Keywords
- hepatitis C virus
- E2 glycoprotein
- conserved domain
- hCD81
- protein–protein interaction
- interferon-gamma
- immune response
Hepatitis C virus (HCV) infection remains a worldwide health threat to the public. Despite progress in therapeutic options, there is yet to be an effective and safe vaccine to prevent this virus. The HCV envelope E2 glycoprotein is one of three structural proteins (core, E1, E2) encompassing amino acids (aa) 384–746 of the viral polyprotein and carrying two hypervariable regions (HVR1 and HVR2) [1, 2, 3]. The high variation of HVR1 and HVR2 of HCV E2 glycoprotein and the use of error-prone RNA-dependent RNA polymerase for viral RNA replication generates a variety of HCV quasispecies, and they are considered a major viral escape mechanism. Several previous studies suggested that the epitopes aa 481–500 and aa 551–570 of HCV E2 glycoprotein may be critical for immunoreactivity. Furthermore, they added that most of the sites for binding to the human cell membrane protein CD81 (hCD81), including aa 420, 527, 529, 530, and 535, were found to be highly conserved across various HCV genotypes [4, 5, 6, 7, 8, 9]. Based on these previous findings, this study hypothesized that an HCV E2 fragment containing the highly conserved binding sites to hCD81 may circumvent this obstacle. To test this hypothesis, this study investigated the conserved domain and explored its potential as a target in developing a vaccine for HCV.
Extensive studies with soluble, truncated HCV E2 glycoprotein or virus-like particles containing E2 have shown its interaction with hCD81 and suggested that hCD81 was a receptor for HCV [10, 11, 12, 13, 14]. Moreover, a deglycosylated HCV E2 glycoprotein has been shown to bind hCD81 with similar binding efficiency compared with that of the core-glycosylated form. However, a highly glycosylated form of HCV E2 glycoprotein has shown a substantial reduction in binding affinity to CD81. Yurkova et al. [15] have reported the expression of glycosylated, soluble HCV E2 with a lack of its transmembrane domains in mammalian cells, and the expression level was low. In addition, the purified product was found to be highly heterogeneous, even when a single cellular compartment was used for low protein isolation [15]. To increase the yield sufficient for a further binding assay and a functional study, it was decided to express recombinant HCV E2 glycoprotein using a prokaryotic expression system.
Hepatitis C virus E2-induced specific immune responses were demonstrated in patients with HCV as supported by cytotoxic T lymphocyte, lymphocyte proliferation, and IFN-alpha production. More importantly, a stronger HCV E2-specific immune response was associated with a better clinical outcome of interferon therapy and viral clearance [16, 17, 18, 19]. However, little is known about whether a conserved fragment of HCV E2 could mediate specific T-cell responses. This study hypothesized that highly conserved and immuno-dominant domains within HCV E2 glycoprotein could elicit high cross-reactive immunity to different serotypes of HCV.
Based on these previous findings and to test this hypothesis, this study
constructed a prokaryotic expression plasmid containing the coding region of the
conserved domain of E2 protein (aa 505–702, including the positions at aa 527,
529, 530, and 535 for binding to hCD81). It was then examined whether the
bacterially expressed conserved domain of HCV E2 glycoprotein could be functional
by an assay binding to hCD81 and interferon-gamma (IFN-
Blood samples were collected from 10 patients [6 men and 4 women, with a mean
age of 42 years (range 22
Peripheral blood mononuclear cells (PBMCs) were isolated from the whole blood samples using standard Ficoll density gradient centrifugation. Blood samples were mixed with Hank’s balanced salt solution at an equal volume, followed by Ficoll-Hypaque density gradient centrifugation (GE HealthCare, Little Chalfont, Buckinghamshire, UK). After removing the supernatant, the resulting cells were washed twice and re-suspended in a complete cell culture medium [mg/mL streptomycin, and 50 mM 2-mercaptoethanol].
Total RNA was extracted from the serum of patients with HCV or the PBMCs of the healthy control using the RNeasy Midi-Prep Kit (QIAGEN GmbH, hilden, Germany) following the manufacturer’s manual. DNase I (Roche Diagnostics GmbH, Mannheim, Germany) was applied to remove potential genomic DNA contamination. Reverse transcription-polymerase chain reaction (RT-PCR) (MBI Fermentas, Vilnius, Lithuania) was conducted using the purified total RNA as a template, oligo(dT) as the RT primer, followed by PCR amplification using specific primers as listed in Table 1. The PCR conditions used in the RT-PCR were as follows: 94 °C for 30 s, 58 °C for 40 s, and 72 °C for 1 min, for 30 cycles. Finally, the reaction was maintained at 72 °C for 10 min. The PCR amplified fragment encoding the conserved domain of HCV E2 [encompassing amino acids (aa) 505–702] and the large extracellular loop of human CD81 (hCD81) were used for the construction of their expression plasmids in Escherichia coli (E. coli).
| Primer | Polarity | Primer sequence (5′–3′) | Product | Position |
| P1 | Forward | TCAGGATCCTCCAGTGTATTGCTTCAC | E2 | aa505–702 |
| P2 | Reverse | TGCAAGCTTCAGGTATTGCACGTCCA | ||
| P3 | Forward | ATAGGATCCGATGGGAGTGGAGGGCTGCAC | hCD81 | aa41–751 |
| P4 | Reverse | TCGAAGCTTTTAGTACACGGAGCTGTTCCGGATG |
Note: The forward primers (P1, P3) contain a start codon ATG
incorporated into a BamH1 site (shown in italics) followed by
nucleotides representing the 5
The Novagen pET Expression System 22b, pET22b (+) vector, was used to construct prokaryotic expression plasmids for the preparation of the conserved domain of HCV E2 glycoprotein [encompassing amino acids (aa) 505–702, GenBank accession number QQ354893] and human CD81 (hCD81), respectively. The HCV E2 fragment encoding the conserved domain (encompassing aa 505–702) was amplified using the primers listed in Table 1 and ligated into the expression plasmid pET22b (+). The recombinant plasmids were purified, and the fragment was subjected to DNA sequencing.
The constructed plasmids were transformed into E. coli BL21 (DE3), BL21
(DE3) pLysS, Rosetta-gami DE3, or Rosetta-gami pLysS cells (Novagen, San Diego,
CA,USA) to express the conserved domain of HCV E2 or hCD81. The E. coli cells were grown in
a lysogeny broth medium to reach an A
Upon the completion of the protein expression, the E. coli cells were harvested, washed, and then resuspended in BugBuster (Novagen, San Diego, CA, USA). Cell suspensions were sonicated on ice for 4 min (cycles of 10 s on followed by 30 s off), and this step was repeated twice, followed by centrifugation at 10,000 g for 20 min at 4 °C. The resulting supernatant and pellet were collected.
The prokaryotic expression of the recombinant HCV E2 and hCD81 was examined using 10% sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis on a Mini Protean III apparatus (Bio-Rad, Hercules, CA, USA). Protein gels were stained with Coomassie brilliant blue R-250 and destained with 5% acetic acid and 25% methanol.
Western blot (WB) analysis was conducted to examine the expression and purification of recombinant HCV E2 and hCD81. Total proteins were separated using 12% SDS-PAGE and electro-transferred onto the nitrocellulose membrane (Millipore, Billerica, MA, USA). The resulting membranes were blocked with 3% non-fat milk (KPL) in a phosphate-buffered saline (PBS) containing 0.05% Tween-20 (PBS-T) for 2 h at room temperature (RT, 20 °C). The membranes were then incubated with a histidine (HIS)-tag monoclonal antibody (Novagen, San Diego, CA, USA) (dilution, 1:2000) or HCV E2-specific monoclonal antibody (Biodesign, Denver, CO, USA) (dilution, 1:100) for 2 h. For the detection of hCD81, the primary monoclonal antibody against CD81 (Santa Cruz Biotechnology, Santa Cruz, CA, USA) was used, and the dilution was 1:1000 in 1% non-fat milk in PBS-T. Subsequently, the membranes were incubated for 1 h with peroxidase-conjugated anti-mouse or anti-rabbit IgG antibody (Sigma Chemical Co., St. Louis, Mo, USA) (dilution, 1:2000 in 1% non-fat milk in PBS-T) at RT. The bound antibodies were detected using tetramethylbenzidine antibodies for 10 min at RT.
The HIS-tagged HCV E2 fusion protein was purified by affinity chromatography
using the nickel-nitrilotriacetic acid (Ni-NTA) superflow resin (Novagen, San Diego,
CA,USA) following the manufacturer’s instructions. The E. coli cells with the
expression of HIS-tagged HCV E2 fusion protein were suspended with a lysis buffer
(50 mM NaH
Freshly prepared soluble fractions of E. coli cell lysate containing
the recombinant conserved domain of HIS-HCV E2 (5 mL) or hCD81 (5 mL) were mixed
and incubated at 30 °C for 90 min. The mixture with potential
HIS-E2-hCD81 complex was loaded onto the Ni-NTA affinity column pre-equilibrated
with a lysis buffer (QIAexpressionist, Qiagen, hilden, Germany) and was allowed to bind to the
Ni-NTA resin for at least 1 h. After washing three times with a 5
The Human IFN-
Statistical analysis was conducted using the SPSS software (version 23.0, IBM,
Armonk, New York, USA), and differences in mean values were determined
using the Student’s t-test. Furthermore, p
This study generated an expression vector harboring the conserved domain of HCV
E2 glycoprotein using the pET22b prokaryotic expression system. A 6
To optimize conditions for the target protein expression, this study tested different strains of host cells [BL21 (DE3), BL21(DE3) pLysS, and Rosetta-gami strains] and a wide temperature range (+20, +25, +30, and +37 °C) (the data is not shown). By comparison, it was decided to use BL21(DE3) pLysS as host cells to express the target protein at 37 °C. As shown in Fig. 1, the conserved domain of HCV E2 glycoprotein was highly expressed in the E. coli strain BL21(DE3) pLysS (Fig. 1). The results suggested that the prokaryotic expression system gave the best yield of the soluble recombinant HCV E2 protein (Fig. 1A,B1,C1,D1,E1).
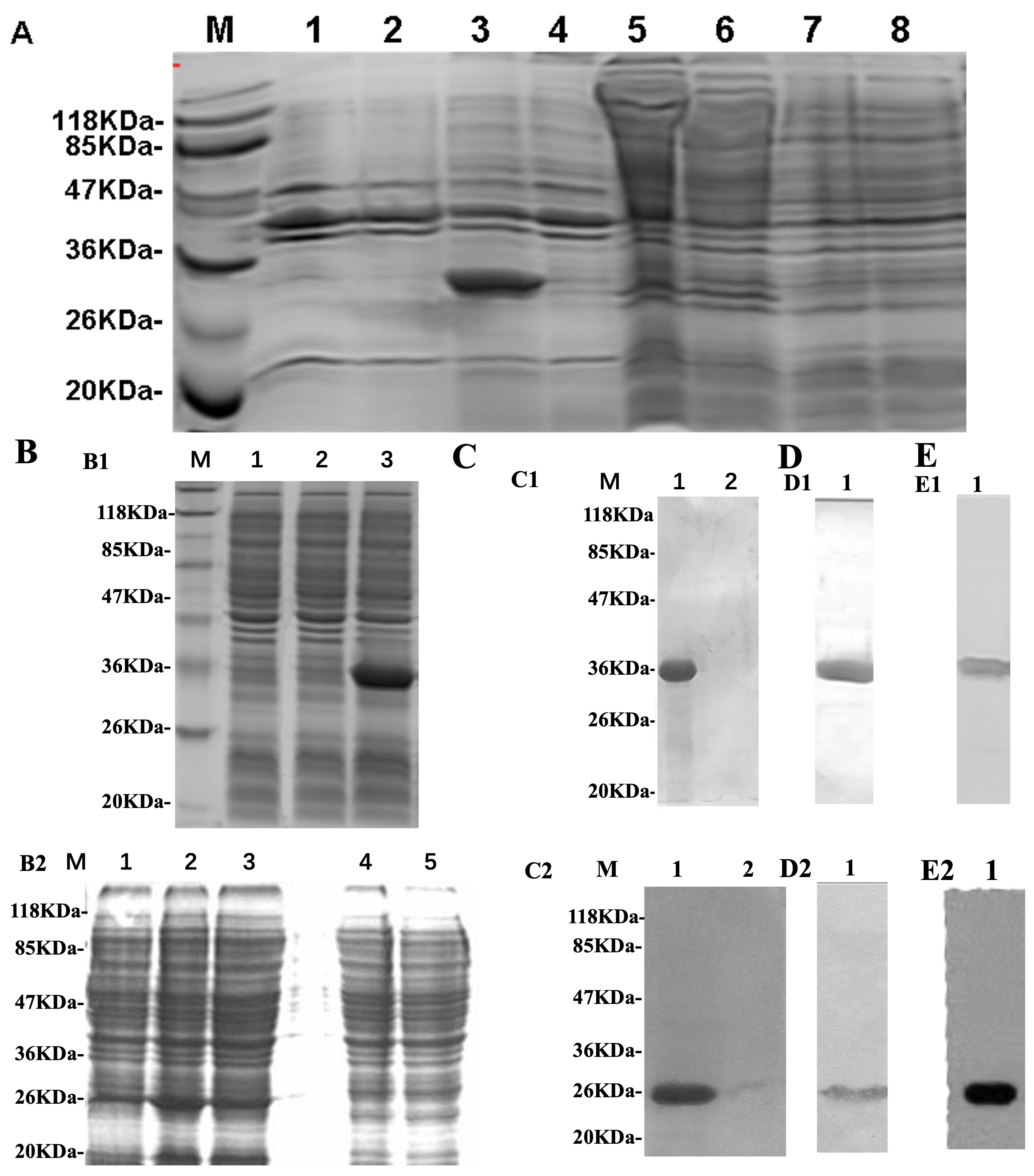 Fig. 1.
Fig. 1.Analysis of hepatitis C virus E2 and hCD81 protein expression
and purification. Prokaryotic expression plasmids were transformed into
Escherichia coli strain BL21, and hepatitis C virus E2 or hCD81 protein
(the large extracellular loop) expression was induced by
propylthio-
With the optimized prokaryotic expression system, the vector containing the hCD81 coding region was transformed into E. coli strain BL21(DE3) pLysS. As shown in Fig. 1B2,C2,D2,E2, the target protein hCD81 was highly expressed in BL21(DE3) pLysS.
To evaluate the similarity of amino acid sequences of the acquired conserved domain of HCV E2 glycoprotein among the main HCV serotypes and strains across different geographic regions, amino acid sequences (aa 500–700) were aligned with those of 15 representative strains of different HCV subtypes using the DNAssist software (version 2.0, Softonic, Barcelona, Spain). As schematically presented in Fig. 2 (Ref. [21]), the positions of amino acid residues 527, 529, 530, 535, 544, and 551, known as critical sites for binding to hCD81, shared 100% similarity and were identical among all the strains from HCV serotypes as expected.
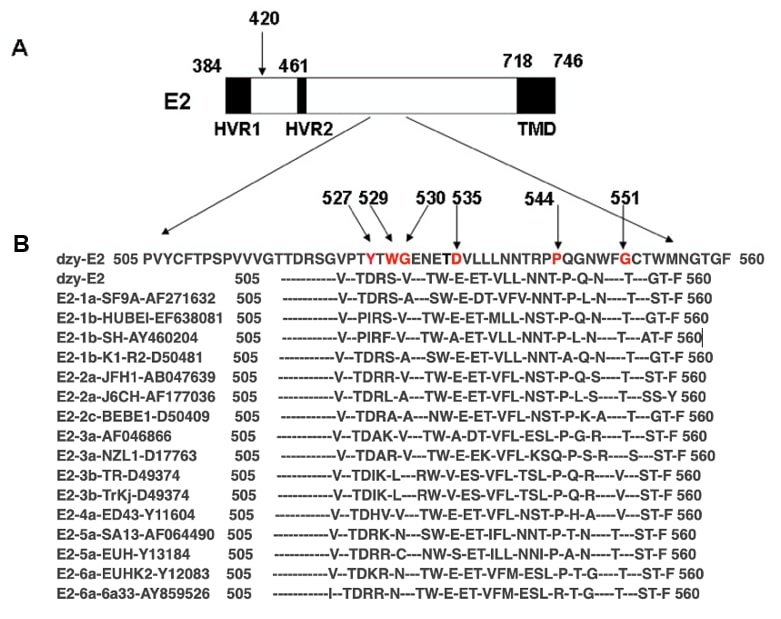 Fig. 2.
Fig. 2.Alignment of amino acid sequences (aa 505–560) in hepatitis C virus envelope glycoprotein E2 among different hepatitis C virus genotypes. (A) Schematic representation of hepatitis C virus envelope glycoprotein E2. The numbers corresponded to the amino acid positions in the hepatitis C virus polyprotein of the human prototype strain H of the hepatitis C virus as a reference strain (GenBank accession no. AY460204). Hypervariable regions 1 (HVR1) and 2 (HVR2) and the transmembrane domain (TMD) were denoted. (B) Alignment of amino acid sequences (aa 505–560) in dzy-E2 (GenBank accession no. QQ354893) and hepatitis C virus E2 among 15 representative hepatitis C virus strains, including hepatitis C virus genotype 1a (GenBank accession no. AF271632); hepatitis C virus genotype 1b (EF638081, AY460204, D50481); hepatitis C virus genotype 2a (AB047639, AF177036); 2c (D50409); hepatitis C virus genotype 3a (AF046866, D17763); hepatitis C virus genotype 3b (D49374); 4a (Y11604); hepatitis C virus genotype 5a (AF064490, Y13184); and hepatitis C virus genotype 6a (Y12083, AY859526). As indicated by prefixes, the 15 subtypes represented hepatitis C virus isolates of each hepatitis C virus genotype. Amino acids identical to those in the consensus sequence were indicated by a hyphen. The arrows indicated the positions of the amino acid at 420 (A), and at 527, 529, 530, 535, 544, and 551 in the conserved domain (B), which were previously reported to be potentially involved in binding to hCD81 [21].
This study assayed the binding of the recombinant conserved domain of HCV E2 glycoprotein to hCD81 as described in Materials and Methods. Freshly prepared HIS-E2 protein was incubated with non-HIS-hCD81, and a complex was expectedly formed. As illustrated in Fig. 3A, SDS-PAGE analysis of the final eluted protein samples showed two dominant bands of a bound complex on the Ni-NTA column. Western blot analysis was further carried out using anti-HIS tag monoclonal antibody to detect HIS-E2 (Fig. 3B) and anti-hCD81 monoclonal antibody (Fig. 3C). The Western blot images indicated that the bound complex resulted from the direct interaction between the conserved domain of HCV E2 glycoprotein and hCD81 (Fig. 3B,C).
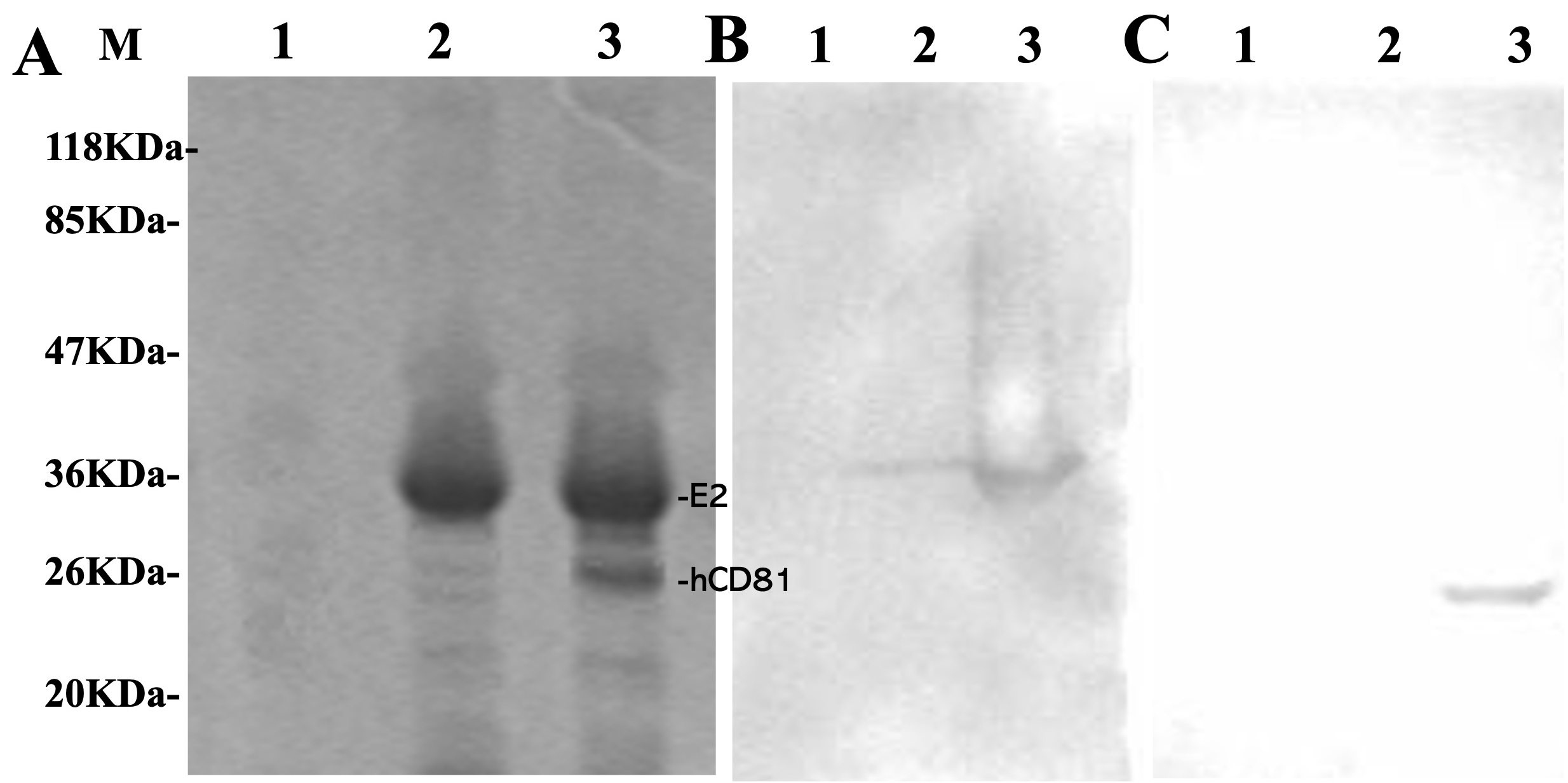 Fig. 3.
Fig. 3.Analysis of the interaction between histidine-hepatitis C virus E2 and non-histidine-hCD81. Freshly prepared soluble fractions of Escherichia coli cell lysate containing histidine-E2, non-histidine-hCD81, or a mixture of both were loaded onto the nickel-nitrilotriacetic acid affinity chromatography column. After unbound proteins were washed thoroughly, bound proteins were eluted and examined by (A) Sodium dodecyl-sulfate polyacrylamide gel electrophoresis and Western blot analysis using (B) an anti-histidine tag antibody or (C) an anti-hCD81 monoclonal antibody. Lane M: protein marker; Lane 1: non-histidine-hCD81; Lane 2: histidine-HCV E2; Lane 3: a mixture of histidine-hepatitis C virus E2 and non-histidine-hCD81.
Finally, this study evaluated whether the recombinant conserved domain of HCV E2
glycoprotein could induce specific T-cell responses as reflected by the secretion of IFN-
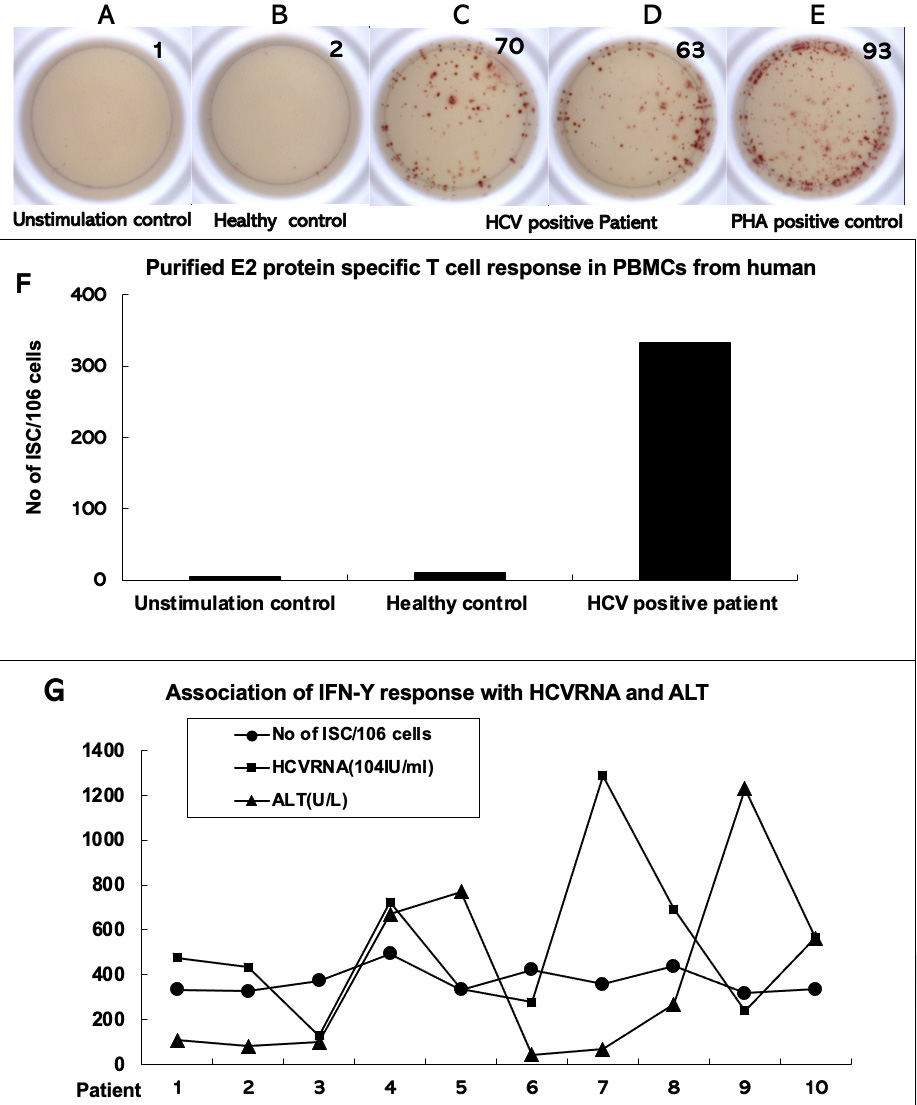 Fig. 4.
Fig. 4.Interferon gamma enzyme-linked immunosorbent spot assays to test
hepatitis C virus E2 protein-stimulated T-cell response in peripheral blood
mononuclear cells of patients with chronic hepatitis C. Peripheral blood
mononuclear cells were isolated from the whole blood samples of patients with
chronic hepatitis C (n = 10). They were stimulated in vitro with
purified hepatitis C virus E2 protein for IFN-
Previous studies on the expression of the recombinant HCV E2 glycoprotein in
bacteria resulted in an insoluble product at a low yield [15, 22, 23, 24]. This study
constructed a prokaryotic expression vector carrying the conserved domain of HCV
E2 glycoprotein, and the target protein was highly expressed in E. coli
in a soluble form. The major novel findings were summarized as follows: (1) the
conserved domain of HCV E2 glycoprotein prepared in the E. coli strain
BL21 was recognized by an anti-HCV E2 monoclonal antibody; (2) the bacterially
expressed conserved domain of HCV E2 glycoprotein showed a direct interaction
with hCD81 in the binding assay; (3) the recombinant conserved domain of HCV E2
glycoprotein markedly induced IFN-
One of the strengths of this study was the intention to investigate the function of cross-serotype conserved domain with HCV E2 glycoprotein. Evaluation of the similarity of amino acid sequences showed that the bacterially expressed conserved domain of HCV E2 glycoprotein contained amino acid residues 527, 529, 530, 535, 544, and 551, which were well documented as important sites for binding to hCD81. The conserved domain shared 100% similarity with that among all the strains from HCV serotypes, which was expected. It was found that the high variable E2 sequence also has several conservative regions that would help maintain the function stabilization of E2 and support vaccine design for effective cross-serotype protection against HCV.
To test whether the bacterially expressed conserved domain of HCV E2 glycoprotein as prepared in this study is functional, a modified method like the “pull-down” assay [25, 26] was applied to determine its interaction with hCD81. The results indicated that the bacterially expressed conserved domain of HCV E2 glycoprotein was functional. This study showed that the conserved domain of HCV E2 glycoprotein directly interacted with the HCV receptor, hCD81, in vitro. In the binding assay, it was found that the eluted samples were recognized by HIS-tag McAb or E2 McAb. The antigenic and hCD81-binding properties of the conserved domain of HCV E2 glycoprotein indicated similar functions compared with its native glycosylated counterpart. The results of the binding assay also demonstrated that the non-glycosylated recombinant conserved domain of HCV E2 glycoprotein may retain the functionality of the native glycosylated counterpart. Considering that the soluble conserved domain of HCV E2 glycoprotein can be prepared in adequate amounts, it can be used as a surrogate model of a native counterpart. It was also confirmed that the binding domains of HCV E2 to hCD81 existed in a respective conserved region. These findings may help design hCD81-based mimics that inhibit the binding of HCV-E2 to CD81.
The conserved domain of HCV E2 stimulated the production of IFN-
In conclusion, this study has shown that the conserved domain of HCV E2
glycoprotein directly interacts with hCD81, through which it activates
IFN-
HCV, Hepatitis C virus; HVR1 and HVR2, hypervariable regions; hCD81, human cell
membrane protein CD81; VLP, virus-like particles; IFN-
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
ZYD and GL designed the research study. ZYD and WZ performed the research. XS analyzed the data. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
This study had been approved by the Ethics Committee of the Sun Yat-sen University. All methods were carried out in accordance with relevant guidelines and regulations (SYSU, 2022017882). All the patients/participants provided their written informed consent to participate in this study
This work was supported by the Research Unit and Vaccine Research Institute of the Third Affiliated Hospital of Sun Yat-sen University, and Gene Therapy Program of the University of Pennsylvania.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.




