1 Department of General and Industrial Microbiology, Faculty of Biology, Sofia University “St. Kliment Ohridski”, 1164 Sofia, Bulgaria
Abstract
Background: Traditional Bulgarian fermented foods are prominent for
their uniqueness of local ingredients, production methods, and endemic microbial
species. The present research investigated the diversity and beneficial
biological potential of lactic acid bacteria (LAB) isolated from various types of
unique Bulgarian fermented foods. Methods: Species identification was
performed via 16S rDNA sequencing. Biological activity was evaluated by
determining antibacterial activity (via agar well diffusion assay),
H
Keywords
- lactic acid bacteria
- functional foods
- bio-protection
- beneficial bacteria
- antibacterial potential
- acquired-antibiotic resistance
For centuries, fermented products derived from plant or animal materials have played an essential role in human nutrition worldwide. Food fermentation is considered one of the most ancient ways of processing and preserving food. Fermentation enhances the flavor and nutritional quality of food and increases its shelf life. Fermented foods are considered a good source of natural probiotics because their intake has been reported to improve intestinal function, help boost immunity, and prevent allergies [1]. These factors contribute to the growing interest in researching the health benefits of consuming fermented foods [2]. Among the most widespread and traditionally prepared fermented foods in Bulgaria are sauerkraut (whole sour cabbage), fermented green tomatoes, fermented cucumbers, kefir, boza, yogurt, etc. Their distinctive feature is their diverse and unique taste qualities. Moreover, their microbiota can vary depending on the raw substrate used in fermentation. Thus, exploring the beneficial microbial diversity in naturally fermented foods could be considered valuable with a strict focus on isolation, selection, and biotechnological application of bacterial strains, with the potential to act as probiotics [3].The term probiotics, however, refers to live microbes that, administered in sufficient amounts, lead to the improved health of the host [1].
Natural fermentation is a process that is inherent and carried out by a diverse microbiota, including bacteria, yeast, and molds [4, 5]. The final products of the different fermentation processes depend on the microbes, starting substrates, and fermentation conditions [6]. Microorganisms involved in food fermentation should not be pathogenic, and the enzymes produced during these processes (proteases, amylases, and lipases) help break down the initial substrates into final products with sought-after qualities [7].
Lactic acid bacteria (LAB) are the most prevalent microorganisms found in vegetable- or milk-fermented foods, and lactic acid fermentation is regarded as the primary contributor to the beneficial properties these foods possess [4]. The reported dominant microbiota in vegetable-fermented products (sauerkraut) consists of Lactiplantibacillus plantarum, Levilactobacillus brevis, and Leuconostoc mesenteroides [8]. Conversely, the prevalence of LAB microorganisms in traditional fermented milk products (kumis, jiaoke, cheese, cottage cheese, shubat, and yogurt) analyzed in Mongolia, Kazakhstan, and Russia showed significant diversity, including Lactobacillus helveticus, Lactococcus lactis, Streptococcus parauberis, Lactobacillus kefiranofaciens, Lactobacillus delbrueckii, Streptococcus salivarius, Lactococcus raffinolactis, Lactococcus piscium, and Streptococcus parasuis [9]. The significance of LAB is mostly related to their safe metabolic activity, which provides numerous functional and health benefits to fermented foods [10]. The LAB strains, which have the so-called Generally Recognized as Safe (GRAS) status and fulfill the requirements of qualified presumption of safety (QPS) in Europe, are considered suitable to be used in various products in food industries (dairy products, meat products, fermented cereals, etc.) [11]. Moreover, part of the LAB in fermented foods meets the criteria for probiotics [12]. Such valuable criteria could overcome the enzymatic activity in the oral cavity and survival through the harsh gastrointestinal tract (GIT) environment. Upon reaching the lower part of the small intestine and colon (their site of action), the LAB colonizes, multiplies, and adheres to the host cells, where they exert their beneficial effects [12].
Among the reported health benefits from the consumption of probiotic LAB are balancing the intestinal microbial community, stimulating the immune system, reducing the risk of bacterial and viral-associated diarrhea, and lowering serum cholesterol [13]. It has also been proposed that the mechanisms of probiotic activity include the production of antimicrobial substances, thereby preventing the pathogen from adhesion to the epithelial layer of the intestine and competing for nutrients [14]. The ability of probiotic bacteria to colonize in the host GIT epithelial cells is considered to be determined by their cell surface hydrophobicity [15]. This property prevents the adhesion of pathogens to mucosal layers and increases the persistence time of LAB in the host [12].
The synthesis of organic acids [16, 17], reuterin [18], proteinaceous substances [19], and cyclic dipeptides [20] has been linked to the antibacterial action of LAB. The bacteriocins synthesized by LAB have also been reported to be involved in two fundamental processes: (1) target cell membrane depolarization and/or suppression of the formation of the bacterial cell wall and (2) degradation of the murein layer [21]. The broad-spectrum antimicrobial substance reuterin has been thought to inhibit the ribonucleotide reductase enzyme competitively. It is also believed to degrade proteins and smaller molecules, which inhibits microorganisms from proliferating [22, 23]. Undissociated and hydrophobic organic acid molecules permeate the pathogens’ cell membranes, neutralizing their electrochemical properties and increasing permeation, ultimately leading to bacteriostasis and death [24, 25].
The food in each geographical region is distinguished by the specificity of local ingredients and production methods. This uniqueness is deeply anchored in the Bulgarian folk tradition and is related to the production of unique foods obtained with the participation of endemic microbial species. In light of this, the present research aimed to study the diversity and beneficial biological activity of the lactic acid microflora isolated from various types of unique Bulgarian fermented foods. The objects of the investigation are products that have not been subjected to such extensive examination previously (sauerkraut, fermented green tomatoes, fermented cucumbers). The isolated strains were identified and screened for their bioprotective attributes, alongside exploration of their beneficial potential in biotechnological applications.
Specimens of different types of fermented food products (sauerkraut, fermented green tomatoes, fermented cucumbers, kefir, white cheesе, and Izvara (curdled milk)) were prepared according to traditional Bulgarian recipes and used as sources of LAB isolation. Among these products were homemade and artisanal markets. The samples were collected in sterile containers from different regions in Bulgaria. The potential LAB in the samples were enriched via the cultivation of 1 g (mL) of each product in de Man, Rogosa, and Sharpe (MRS) broth (Merck, Darmstadt, Germany), separately. The resulting enriched cultures were diluted ten-fold, and aliquots of 100 µL were spread onto the MRS agar surface. The Petri dishes were incubated at 30–37 °C in an anaerobic jar (utilizing the BBL GasPack anaerobic system) for 48 hours. Colonies with different morphological characteristics were selected and re-streaked to obtain pure bacterial cultures. Preliminary selection of potential LAB isolates was performed according to results obtained for Gram-staining (Gram-staining Kit, Sigma Aldrich, St. Louis, MO, USA) to determine oxidase (Oxidase strips, Sigma-Aldrich) and catalase activities [26].
The total DNA of all isolates was extracted from overnight MRS broth cultures
using a Tissue and Bacterial DNA Purification kit (EURx Ltd., Gdansk, Poland),
following the manufacturer’s instructions. The DNA Purification kit purified the
isolated DNA (PCR/DNA Clean-up, GeneMatrix, EURx Ltd., Gdansk, Poland). The
quantity of purified DNA was measured by BioDrop µLITE+ (100
ng/µL
The antimicrobial activity of the isolated LAB was tested against the following
pathogenic test microorganisms: Listeria innocua F 4078,
Staphylococcus aureus ATCC 6538, Escherichia coli NBIMCC 3548,
Salmonella enterica subsp. enterica serotype
Enteritidis NBIMCC 8691, and Candida albicans ATCC 10231 using
the agar–well diffusion method [28, 29]. The pathogenic bacteria and yeast were
cultivated overnight on Brain Heart Infusion (BHI) broth (Merck KGaA, Darmstadt,
Germany) and Yeast Peptone Dextrose (YPD) broth (Merck KGaA, Darmstadt, Germany),
respectively, to obtain log cultures. The LAB isolates were cultivated in MRS
broth at 37 °C for 24 hours, and after that, cell-free supernatants
(CFS) were obtained by centrifugation (at 6000
The ability of the LAB isolates to produce H
Autoaggregation (AA) and co-aggregation (CA) assays were carried out according
to Tuo et al. [33]. The LAB isolates were grown on MRS broth at 37 °C
overnight. For AA, the bacterial cells were harvested by centrifugation at 10,000
The degree of hydrophobicity on the surface of bacteria was evaluated by
measuring their adherence to hexane using the modified protocol of Collado
et al. [34]. LAB isolates were cultivated in MRS at 37 °C for 24 hours,
washed twice in PBS, and resuspended in 3 mL of PBS buffer to achieve
approximately 1
With a few minor adjustments, an evaluation of the capacity of each LAB isolate
to produce a biofilm was conducted in accordance with earlier protocols by
Bujnakova et al. [35] and Gómez et al. [36]. The isolated
LAB strains were cultivated in 5 mL MRS broth and incubated at 37 °C for
24 hours. The resulting cultures were centrifuged (10,000
The hemolytic activity was evaluated following the methodology reported by
Carrillo et al. [38]. Pure LAB cultures were surface spot inoculated on
blood agar plates supplemented with 5% (v/v) horse blood and incubated at a
temperature of 37 °C for a duration of 24 to 48 hours. The hemolytic
activity of the isolated LAB strains was evaluated after the incubation period.
The strains were classified according to the changes in blood agar medium around
the spot culture as follows: (1) green zones around spot culture
(
The susceptibility of all isolates to different antibiotics was tested according
to the agar disc diffusion procedure described before by Bauer et al.
[40] and Yasmin et al. [1], with modifications. Various antibiotics
(AB), belonging to different groups were used (ampicillin/sulbactam (A/S)—10
µg/disc, vancomycin (VA)—5 µg/disc, chloramphenicol (C)—30
µg/disc, erythromycin (E)—15 µg/disc, tetracycline
(T)—10 µg/disc, ciprofloxacin (CP)—10 µg/disc,
cephalothin (CF)—10 µg/disc, gentamicin (G)—10
µg/disc, streptomycin (S) — 10 µg/disc, and neomycin
(N)—5 µg/disc). A total of 1 mL of each strain (approximately 1
The genes related to antibiotic resistance to gentamicin
(aac(6′)-aph(2′′)), chloramphenicol (cat), tetracycline
(tet(M)),
A total of 45 potential LAB strains (Table 1) were obtained from various traditional Bulgarian fermented foods (sauerkraut, fermented green tomatoes, fermented cucumbers, kefir, white cheese, and Izvara (curdled milk)). Of these, 25 strains were from fermented vegetable products, and 20 were from fermented milk products. Phenotypic characterization of the isolates showed that all strains formed white to creamy-colored colonies with smooth or jagged edges, predominantly with a convex profile and a uniform to slightly granular structure. Micromorphologically, the cells were rod-shaped with different lengths. All isolates were Gram-positive bacteria, catalase, and oxidase-negative. These phenotypic data are typical characteristics of the LAB group.
| Number | Strains | Origin of isolation | Year of isolation | Media | Cell morphology | Gram stain | Oxidase | Catalase | Production of H |
Species identification by 16S rDNA gene sequence |
| 1 | S1 | Sauerkraut* | 2019 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | |
| 2 | S2 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus brevis | |
| 3 | S3 | 2019 | MRS | Rod-shaped | + | – | – | + | Lactiplantibacillus plantarum | |
| 4 | S4 | 2019 | MRS | Rod-shaped | + | – | – | + | Lactiplantibacillus plantarum | |
| 5 | S5 | 2019 | MRS | Rod-shaped | + | – | – | + | Lactiplantibacillus plantarum | |
| 6 | S6 | Sauerkraut** | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | |
| 7 | S7 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 8 | S7’ | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 9 | S8 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 10 | S9 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 11 | S10 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 12 | S11 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 13 | S12 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 14 | S13 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 15 | S1 | 2018 | MRS | Rod-shaped | + | – | – | + | Lactiplantibacillus plantarum | |
| 16 | S15 | 2018 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 17 | S16 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus yonginensis | |
| 18 | FGT1 | Fermented green tomatoes** | 2019 | MRS | Rod-shaped | + | – | – | – | Levilactobacillus koreensis |
| 19 | FGT2 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus koreensis | |
| 20 | FGT3 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus koreensis | |
| 21 | FGT4 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus koreensis | |
| 22 | FC1 | Fermented cucumber** | 2019 | MRS | Rod-shaped | + | – | – | – | Lactiplantibacillus plantarum |
| 23 | FC2 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus koreensis | |
| 24 | FC3 | 2019 | MRS | Rod-shaped | + | – | – | + | Levilactobacillus koreensis | |
| 25 | FC4 | 2019 | MRS | Rod-shaped | + | – | – | – | Levilactobacillus koreensis | |
| 26 | K1 | Kefir* | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | |
| 27 | K2 | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 28 | K3 | 2019 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 29 | K4 | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 30 | K5 | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 31 | K6 | 2019 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 32 | K7 | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 33 | K8 | 2019 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 34 | WC1 | White cheese* | 2018 | MRS | Rod-shaped | + | – | – | Lactobacillus helveticus | |
| 35 | WC2 | 2018 | MRS | Rod-shaped | + | – | – | – | Lactobacillus helveticus | |
| 36 | WC3 | 2018 | MRS | Rod-shaped | + | – | – | Lactobacillus helveticus | ||
| 37 | WC4 | 2018 | MRS | Rod-shaped | + | – | – | Lactobacillus helveticus | ||
| 38 | WC5 | 2018 | MRS | Rod-shaped | + | – | – | Lactobacillus helveticus | ||
| 39 | WC6 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 40 | I1 | Izvara* (curdled milk) | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | |
| 41 | I2 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 42 | I3 | 2018 | MRS | Rod-shaped | + | – | – | Levilactobacillus brevis | ||
| 43 | I4 | 2018 | MRS | Rod-shaped | + | – | – | Lactobacillus helveticus | ||
| 44 | I5 | 2018 | MRS | Rod-shaped | + | – | – | Lactiplantibacillus plantarum | ||
| 45 | I6 | 2018 | MRS | Rod-shaped | + | – | – | – | Levilactobacillus brevis |
*homemade products; **artisanal (market) products. LAB, Lactic acid bacteria; FGT, Fermented green
tomatoes; FC, Fermented cucumber; K, Kefir; WC, White cheese; I, Izvara; MRS, de
Man, Rogosa, and Sharpe; +, moderately positive reaction;
All 45 isolated strains from traditional homemade and artisanal Bulgarian fermented products were subjected to molecular identification. The genetic material of all potential LAB isolates was extracted, and the 16S rDNA gene was amplified using universal primers 27F/1492R [27]. The obtained PCR products (approximately 1500 bp) were purified and sequenced by Macrogen in The Netherlands. The results were processed and compared with the NCBI database. All LAB isolates with varying similarity percentages were identified at genus and species levels. Among them, representatives of three genera, according to the new reclassification within the family Lactobacillaceae proposed by Zheng et al. [43], were found (Fig. 1A). The most prevalent genera were found to be Lactiplantibacillus (47%) and Levilactobacillus (40%). Less represented was the genus Lactobacillus (13%). The LAB diversity established after species identification revealed the predominance of five bacterial species: L. plantarum, Levilactobacillus koreensis, L. brevis, L. helveticus, and Levilactobacillus yonginensis (Fig. 1B). L. plantarum was the most prominent (47%) species, as it was found in most tested products. It is interesting to note that this species represents 83% and 80% of all isolated LAB from the sauerkraut products: artisanal market and homemade, respectively. Similar results for the distribution of L. plantarum in traditional fermented foods have also been reported globally [3, 44]. Other authors have reported the presence of Leuconostoc, Weissela, and lactobacilli in sauerkraut fermentation, expressing the opinion that the composition of LAB microflora depends on the cultivar of the cabbage used in the fermentation process [45]. The only product where L. plantarum was not found was fermented green tomato (FGT), where the dominant LAB species was L. koreensis. Interestingly, L. koreensis was found only in two products (fermented green tomatoes, FGT, and fermented cucumbers, FC). Surprisingly, only one strain of L. yonginensis was isolated solely from artisanal sauerkraut. Moreover, both species (L. koreensis and L. yonginensis) are more commonly found in Asian fermented foods such as kimchi [46]. To our knowledge, this study is the first report of L. koreensis and L. yonginensis isolated from fermented vegetable foods in Bulgaria. L. helveticus was the predominant species in the white cheese and Izvara products. It seems that L. brevis is an irrevocable part of fermentative microflora, as it was found in 57% of the tested products in this study (Table 1).
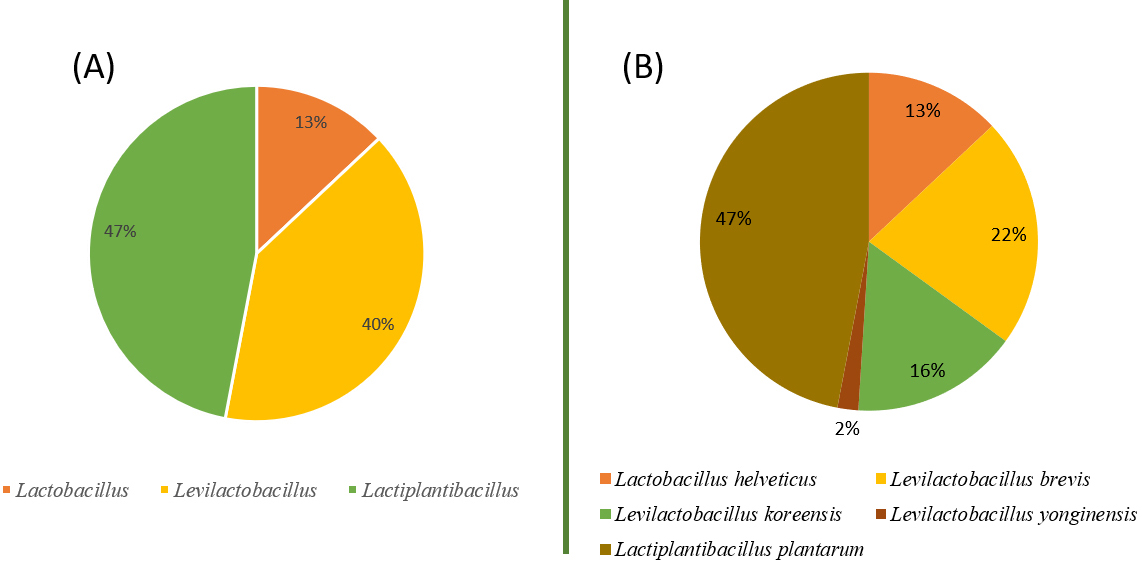 Fig. 1.
Fig. 1.Diversity of LAB in the studied foods. (A) Genus diversity of LAB strains isolated from traditional homemade and artisanal Bulgarian fermented products. (B) Species diversity of LAB isolated from traditional homemade and artisanal Bulgarian fermented products. LAB, lactic acid bacteria.
The fermentation process, which turns sugars into organic acids (lactic and acetic acids), resulting in the formation of acidic environments, is mostly responsible for the preservation behavior of LAB. However, these bacteria can generate and release inhibitory compounds in addition to lactic and acetic acids. Such substances have been reported to have antagonistic effects on various microorganisms. They include a variety of less well-defined or entirely unknown inhibitory substances, such as hydrogen peroxide, diacetyl, ammonia, ethanol, bacteriocins, antibiotics, etc. They are produced in much smaller amounts than lactic and acetic acids. Several of these compounds have demonstrated antagonistic effects against a variety of harmful bacteria found in food, such as Bacillus cereus, Clostridium botulinum, Clostridium perfringens, Listeria monocytogenes, and S. aureus [47, 48].
Our LAB isolates were evaluated for their antimicrobial potential and ability to
produce hydrogen peroxide. We found that only 11 strains showed moderate ability
to produce H
The obtained results showed that antibacterial activity was observed only for
CFS (Table 2). No inhibitory activity was detected for NtCFS and htNtCFS. The
lack of such activity means the isolated LAB does not produce bacteriocins or
bacteriocin-like substances. The observed inhibition activity of CFS against some
of the test microorganisms is probably due to the production of lactic acid,
hydrogen peroxide, or other substances with an inhibitory effect. Different
percentages of the strains of L. plantarum showed apparent antagonistic
effects against E. coli and Sal. Enteritidis (57% and 19%,
respectively). All these strains were isolated from artisanal markets and
homemade sauerkraut. Very weak inhibitory activity (inhibition zones 1–2 mm)
against Listeria innocua F and S. aureus was detected in 24%
and 29% CFS of all L. plantarum isolates, respectively. Similar weak,
barely detectable activity against Sal. Enteritidis, E. coli,
and S. aureus were also established for two strains of L.
koreensis (FGT2 and FC2). Very weak antimicrobial activity by CFS, obtained from
lactobacilli isolated from sauerkraut, was previously reported against
Listeria monocytogenes [45]. Only one of our strains L. brevis
(S15—isolated from artisanal sauerkraut) showed activity against
E. coli (Table 2). None of our isolates showed inhibitory properties
against C. albicans. According to our results and those reported by
other authors, we can conclude that no correlation between antibacterial activity
and H
| Strains | Test microorganisms (inhibition zones, mm) | ||||
| Sal. Enteritidis | L. innocua | E. coli | S. aureus | C. albicans | |
| L. plantarum S1 | – | – | – | – | – |
| L. brevis S2 | – | – | – | – | – |
| L. plantarum S3 | – | – | – | – | |
| L. plantarum S4 | – | – | 14.0 |
– | |
| L. plantarum S5 | – | – | 16.0 |
– | |
| L. plantarum S6 | – | – | 14.0 |
– | |
| L. plantarum S7 | 12.0 |
– | 12.0 |
– | |
| L. plantarum S7’ | – | – | 11.0 |
– | – |
| L. plantarum S8 | – | 11.0 |
– | – | |
| L. plantarum S9 | 11.0 |
– | 13.0 |
– | – |
| L. plantarum S10 | 11.0 |
12.0 |
– | – | |
| L. plantarum S11 | 11.0 |
13.0 |
– | – | |
| L. plantarum S12 | 11.0 |
– | – | ||
| L. plantarum S13 | – | – | 16.0 |
– | – |
| L. plantarum S14 | – | – | 12.0 |
– | – |
| L. brevis S15 | – | – | 11.0 |
– | – |
| L. yonginensis S16 | – | – | – | – | – |
| L.koreensis FGT1 | – | – | – | – | – |
| L. koreensis FGT2 | – | – | – | ||
| L.koreensis FGT3 | – | – | – | – | – |
| L. koreensis FGT4 | – | – | – | – | – |
| L. plantarum FC1 | – | – | – | – | – |
| L. koreensis FC2 | – | – | |||
| L. koreensis FC3 | – | – | – | – | – |
| L. koreensis FC4 | – | – | – | – | – |
| L. brevis K1 | – | – | – | – | – |
| L. brevis K2 | – | – | – | – | – |
| L. plantarum K3 | – | – | – | ||
| L. brevis K4 | – | – | – | – | – |
| L.brevis K5 | – | – | – | – | – |
| L. plantarum K6 | – | – | – | – | – |
| L. brevis K7 | – | – | – | – | – |
| L. brevis K8 | – | – | – | – | – |
| L. helveticus WC1 | – | – | – | – | |
| L. helveticus WC2 | – | – | – | – | – |
| L. helveticus WC3 | – | – | – | – | – |
| L. helveticus WC4 | – | – | – | – | – |
| L. helveticus WC5 | – | – | – | – | – |
| L. plantarum WC6 | – | – | – | – | – |
| L. plantarum I1 | – | – | – | – | – |
| L. plantarum I2 | – | – | – | – | – |
| L. brevis I3 | – | – | – | – | – |
| L. helveticus I4 | – | – | – | – | – |
| L. plantarum I5 | – | – | |||
| L. brevis I6 | – | – | – | – | – |
*
The ability to autoaggregate is an essential property of LAB to form a barrier on the GIT mucosa, which prevents pathogenic bacteria from attaching to it. Some authors claim that lactobacilli have a modest to moderate capability for autoaggregation [50]. To verify the autoaggregation potential of our strains, all isolates were tested for their ability to autoaggregate, and the results were analyzed at 4 and 24 hours. From the group of isolates identified as L. plantarum, four strains (S8, WC6, I1, and I2) showed very good autoaggregation properties after 4 hours—more than 40% of the cells autoaggregated (Fig. 2). However, the best autoaggregation ability after 4 hours was observed for L. koreensis FC4 (48%) and L. brevis S2 (44%). The lowest AA activity was observed for two strains of L. helveticus isolated from white cheese (WC4, 8%, and WC5, 11%). The percent of AA in the rest of the LAB isolates varied from approximately 15% to 25%.
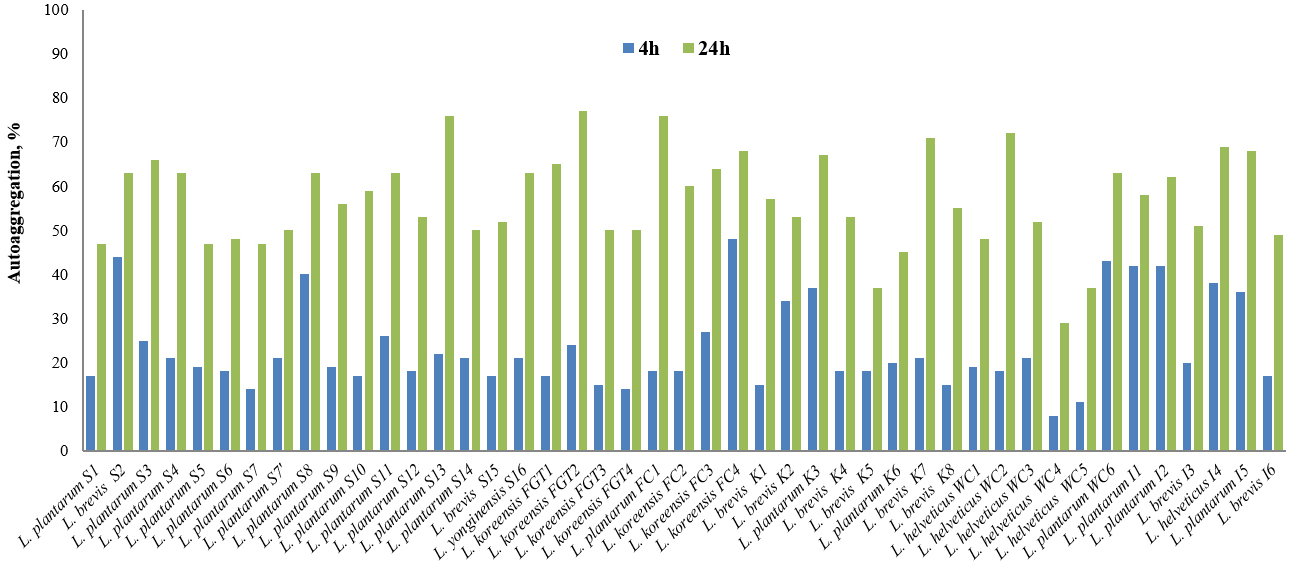 Fig. 2.
Fig. 2.Autoaggregation ability (in percentage, %) of the isolated LAB strains, measured at 4 and 24 hours.
The autoaggregation activity increased alongside the incubation time, and after 24 hours, it reached between 29 and 77%. The highest AA ability was detected for L. koreensis FGT2 (77%), L. plantarum S13 and FC1 (both 76%), L. helveticus WC2 (72%), and L. brevis K7 (71%). However, for FC1 and WC2, the AA ability detected after 4 hours was relatively low, at 18%. Interestingly, the two strains with the highest autoaggregation ability after 4 hours did not possess the maximum AA activity after 24 hours (S2: 63%; FC4: 68%). Fifteen of the isolates (S2, S3, S4, S8, S11, S16, FGT1, FC2, FC3, FC4, K3, WC6, I2, I4, and I5) showed over 60% autoaggregation after 24 hours. Recently, numerous authors have investigated the autoaggregation abilities of lactobacilli as part of their probiotic potential. There is still no single scale for determining the degree of autoaggregation. However, most authors accept 35–40% as sufficient for lactobacilli to be defined as strains with a high aggregation potential. Notably, the strains of the L. plantarum species show different degrees of autoaggregation. Ramos et al. [51] reported that one L. plantarum and one L. fermentum strain reached autoaggregation values of 61.83% and 55.61%, respectively. L. plantarum strain 557, isolated from vegetables, exhibited 34% autoaggregation [49]. Comparing these reported results with those obtained by our study, we can conclude that our L. plantarum isolates showed similar AA abilities. Seven strains of the L. fermentum species isolated from fermented Chinese products presented highly varying autoaggregation limits ranging from 0.86% to 65.15% [52]. In a study of nine strains isolated from milk and milk products (cheese, yogurt), vegetables, and the intestinal tract, Ren et al. [49] described a high rate of autoaggregation only in Lactobacillus salivarius subsp. salicinius CICC 23174 (46%), and Lactobacillus acidophilus CGMCC 1.1854 (45%), compared to the reference strain L. rhamnosus LGG (33%). Our results are similar to those for the strains isolated from milk and milk products, as three L. plantarum strains isolated from white cheese and Izvara (WC6, I1, and I2) showed high autoaggregation potential.
The co-aggregation between probiotic microorganisms and harmful microorganisms creates an unsuitable environment for developing pathogens. This reduces their growth, eliminates the pathogen, and restores the normal gastric microbiota [53]. In the present study, the pathogenic yeast species C. albicans and the studied lactic acid isolates were co-inoculated to monitor their co-aggregation.
In the studied population of LAB isolates, four strains showed
the highest percentage of CA with the pathogen after 4 hours: FGT1 (70%), S2
(54%), S13 (51%), and S6 (50%). The FGT1, K7, S2, and FC2 strains reached the
greatest CA after 24 hours: 95%, 89%, 72%, and 70%, respectively (Fig. 3).
Thus, based on our results, we can conclude that L. koreensis FGT1 could
be considered a candidate for potential probiotic application, according to this
criterion. Interestingly, another group of strains, including FGT2, I6, FC4, S9,
and S6, showed great CA activity at the moment of contact between the LAB and the
pathogen cells (t
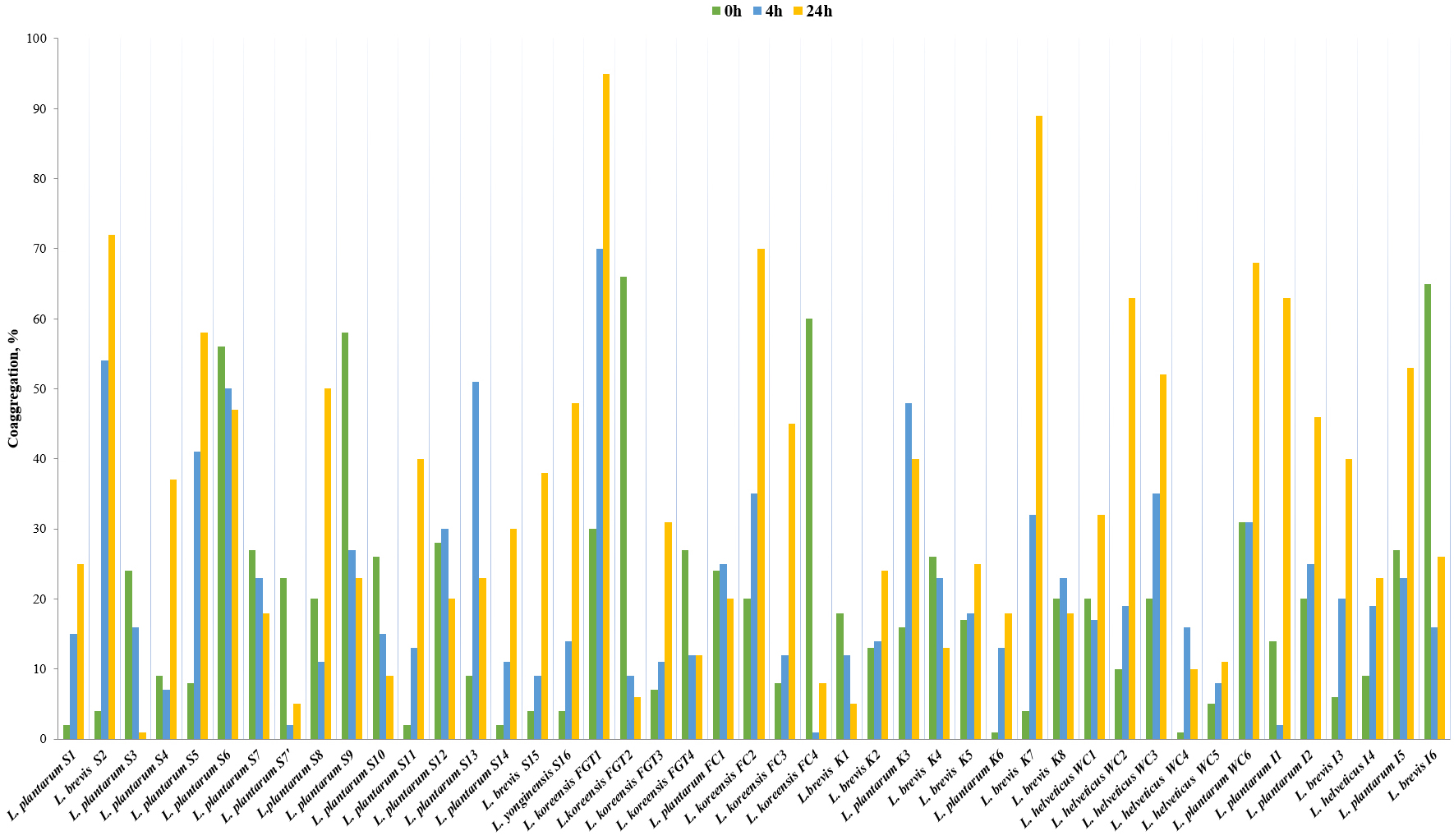 Fig. 3.
Fig. 3.Co-aggregation ability (in percentage, %) of the isolated LAB strains, measured at three time points: 0, 4, and 24 hours.
Bacterial attachment and colonization to the inner side of the GIT depend on cell surface hydrophobicity and AA. The LAB strain must possess a minimum value of 40% hydrophobicity to be called a probiotic strain [56]. Based on this criterion, 84% of our isolates could not be considered presumptive probiotic strains (Fig. 4). However, seven of the isolated LAB strains showed hydrophobicity above 40%. The great ability to adhere to hexane was detected in three LAB strains isolated from milk—fermented products (kefir and white cheese): L. helveticus WC5, 86%; L. brevis K2, 72%, and L. plantarum K6, 52%. The other four LAB strains were isolated from fermented vegetable products (sauerkraut and fermented cucumbers): L. yonginensis S16, 50%; L. plantarum S11, 46%; L. plantarum S12, 44%, and L. koreensis FC4, 43%. The last four stains, except strain S12, also showed good AA ability and were detected after 24 hours. Microbial cell adhesion is affected by a combination of electrostatic and van der Waals forces, as well as the hydrophobicity of the surfaces [57]. Hydrophobicity is important in the initial interaction between bacterial cells and mucosal or epithelial cells [58]. Lactobacilli adhesion is also connected with particular interactions involving the receptor on the bacterial surface recognizing a particular area or ligand [59]. It turns out that the ability of LAB to attach to hydrocarbons is encountered relatively rarely among LAB. For example, Ren et al. [49] investigated nine strains, of which only two strains (22%) showed relatively high levels of hydrophobicity (59% and 43%, respectively). Hoxha et al. [3] reported that none of the twelve LAB strains tested had hydrophobicity above 40%.
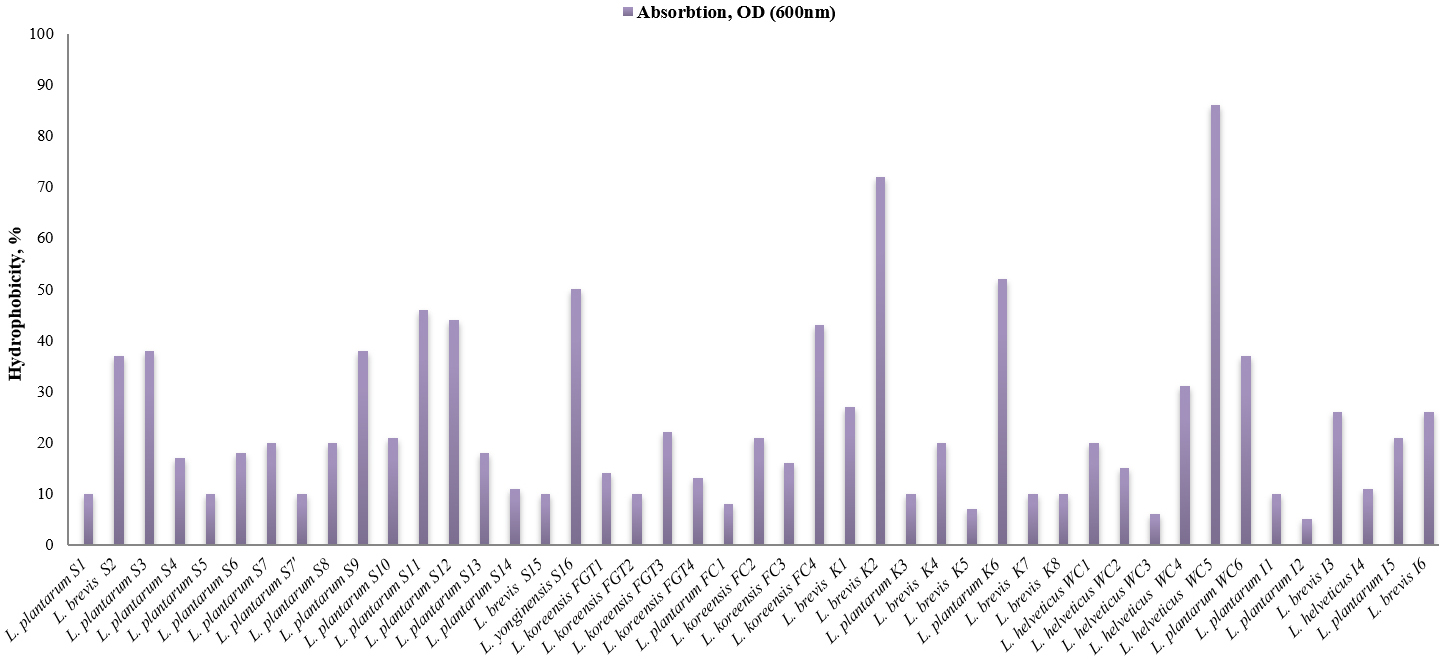 Fig. 4.
Fig. 4.Cell surface hydrophobicity (in percentage, %) of the isolated LAB strains.
Bacterial biofilms are crucial in understanding how bacteria adapt to environmental stress and colonize diverse habitats. To investigate the ability of the LAB strains isolated from traditional Bulgarian fermented food to form biofilms, they were cultivated in a 96-well plate for 24 hours. Based on the obtained results, the strains were grouped as follows: Non-biofilm producers, 18% of the strains (S2, S3, S4, S6, S7, S7’, S8, S14); weak biofilm producers, 13% of the strains (S11, S12, S15, K2, K4, K8); moderate biofilm producers, 53% (S5, S9, S10, S13, K1, K6, WC1, WC5, I1, I2, I4, I5, I6, WC3, WC4, FC3, FGT1, FC1, FC4, FGT2, FGT3, FGT4, S16, FC2); strong biofilm producers, 16% of the strains (S1, K3, K5, K7, WC6, I3, WC2). Based on these results, we can conclude that the ability to form a biofilm is rather strain-dependent.
An important requirement for probiotic strains is their safety. An important initial step in developing or selecting new beneficial microorganisms is to evaluate the lack of hemolytic activity or cytotoxicity [60]. Although hemolytic activity is not inherent to lactobacilli and is relatively rarely found in enterococci, evaluating each case is necessary. Even though many LAB strains have been used for a long time due to their technological properties for food preparation or as probiotics in humans and animals, they are not directly granted GRAS status [61, 62]. This makes verifying that the isolated LAB meets this criterion mandatory. For this reason, the hemolytic activity of all isolated 45 strains was evaluated on blood agar. Hemolytic activity was not detected in any of the tested strains, as evidenced by the absence of hydrolysis zones around the colonies of the isolates. Thus, the tested strains cover one of the safety criteria, such as the absence of hemolytic activity.
Natural resistance to ciprofloxacin, bacitracin, streptomycin, cefoxitin,
fusidic acid, kanamycin, nitrofurantoin, gentamicin, metronidazole, norfloxacin,
sulphadiazine, teicoplanin, trimethoprim/sulphamethoxazole, and vancomycin has
been widely reported for some LAB. [63]. A significantly elevated rate of
spontaneous mutation to nitrofurazone (10
The antibiotic susceptibility of our LAB isolates to 10 antibiotics (ampicillin, tetracycline, erythromycin, gentamicin, streptomycin, vancomycin, neomycin, cephalothin, ciprofloxacin, and chloramphenicol) was evaluated using the agar disc diffusion method. The obtained results are summarized in Table 3, where the isolates were categorized according to Yasmin et al. [67]. Variable susceptibility to the tested antibiotics was observed even at the intraspecies level. All tested strains showed resistance to vancomycin and streptomycin (except L. plantarum S13, which showed strong sensitivity). Our results differ from those reported by Ren et al. [49], showing that all LAB isolated from fermented food were susceptible to streptomycin. A large proportion of strains showed resistance or moderate susceptibility to cephalothin (71%), neomycin (58%), and ciprofloxacin (80%). Three strains (I2, I3, and I4) showed resistance to chloramphenicol; the remaining strains were strongly sensitive. The group of strains (47%) isolated from all fermented products showed resistance or moderate susceptibility to gentamicin. The strains were strongly susceptible to three of the tested antibiotics: Ampicillin, tetracycline, and erythromycin. L. yonginensis S16 was the exception since it showed resistance to tetracycline.
| Strains | Test antibiotics (inhibition zone, mm) | |||||||||
| A/S | T | VA | CP | CF | C | G | E | S | N | |
| L. plantarum S1 | 35/SS | 21/SS | 0/R | 0/R | 0/R | 32/SS | 10/MS | 26/SS | 0/R | 11/SS |
| L. brevis S2 | 30/SS | 16/SS | 0/R | 0/R | 0/R | 31/SS | 13/SS | 25/SS | 0/R | 15/SS |
| L. plantarum S3 | 31/SS | 21/SS | 0/R | 0/R | 0/R | 33/SS | 10/MS | 26/SS | 0/R | 12/SS |
| L. plantarum S4 | 34/SS | 21/SS | 0/R | 0/R | 0/R | 34/SS | 11/SS | 25/SS | 0/R | 11/SS |
| L. plantarum S5 | 24/SS | 22/SS | 0/R | 0/R | 0/R | 32/SS | 10/MS | 26/SS | 0/R | 8/MS |
| L. plantarum S6 | 40/SS | 22/SS | 0/R | 0/R | 17/SS | 35/SS | 13/SS | 27/SS | 0/R | 15/SS |
| L. plantarum S7 | 30/SS | 22/SS | 0/R | 0/R | 17/SS | 33/SS | 12/SS | 27/SS | 0/R | 13/SS |
| L. plantarum S7’ | 36/SS | 21/SS | 0/R | 0/R | 13/SS | 32/SS | 15/SS | 29/SS | 0/R | 9/MS |
| L. plantarum S8 | 27/SS | 25/SS | 0/R | 0/R | 14/SS | 36/SS | 9/MS | 27/SS | 0/R | 0/R |
| L. plantarum S9 | 40/SS | 25/SS | 0/R | 0/R | 16/SS | 35/SS | 11/SS | 27/SS | 0/R | 9/MS |
| L. plantarum S10 | 40/SS | 24/SS | 0/R | 0/R | 0/R | 35/SS | 14/SS | 29/SS | 0/R | 0/R |
| L. plantarum S11 | 35/SS | 23/SS | 0/R | 0/R | 19/SS | 34/SS | 10/MS | 30/SS | 0/R | 15/SS |
| L. plantarum S12 | 32/SS | 25/SS | 0/R | 0/R | 10/MS | 36/SS | 10/MS | 30/SS | 0/R | 10/MS |
| L. plantarum S13 | 35/SS | 20/SS | 0/R | 0/R | 0/R | 33/SS | 18/SS | 27/SS | 17/SS | 15/SS |
| L. plantarum S14 | 35/SS | 25/SS | 0/R | 0/R | 0/R | 35/SS | 8/MS | 27/SS | 0/R | 20/SS |
| L. brevis S15 | 26/SS | 15/SS | 0/R | 0/R | 0/R | 31/SS | 12/SS | 29/SS | 0/R | 0/R |
| L. yonginensis S16 | 30/SS | 0/R | 0/R | 15/SS | 0/R | 32/SS | 15/SS | 35/SS | 0/R | 18/SS |
| L. koreensis FGT1 | 30/SS | 16/SS | 0/R | 0/R | 0/R | 30/SS | 0/R | 26/SS | 0/R | 0/R |
| L. koreensis FGT2 | 37/SS | 15/SS | 0/R | 0/R | 0/R | 30/SS | 0/R | 25/SS | 0/R | 0/R |
| L. koreensis FGT3 | 26/SS | 15/SS | 0/R | 0/R | 0/R | 32/SS | 0/R | 30/SS | 0/R | 0/R |
| L. koreensis FGT4 | 25/SS | 18/SS | 0/R | 0/R | 0/R | 30/SS | 0/R | 26/SS | 0/R | 0/R |
| L. plantarum FC1 | 25/SS | 18/SS | 0/R | 0/R | 0/R | 30/SS | 0/R | 25/SS | 0/R | 10/MS |
| L. koreensis FC2 | 31/SS | 21/SS | 0/R | 0/R | 0/R | 36/SS | 0/R | 30/SS | 0/R | 10/MS |
| L. koreensis FC3 | 30/SS | 16/SS | 0/R | 0/R | 0/R | 36/SS | 0/R | 30/SS | 0/R | 0/R |
| L. koreensis FC4 | 25/SS | 17/SS | 0/R | 0/R | 0/R | 33/SS | 0/R | 25/SS | 0/R | 0/R |
| L.brevis K1 | 29/SS | 20/SS | 0/R | 10/MS | 0/R | 32/SS | 22/SS | 30/SS | 0/R | 13/SS |
| L. brevis K2 | 39/SS | 30/SS | 0/R | 12/SS | 0/R | 39/SS | 9/MS | 31/SS | 0/R | 13/SS |
| L. plantarum K3 | 26/SS | 15/SS | 0/R | 0/R | 0/R | 27/SS | 10/MS | 29/SS | 0/R | 12/SS |
| L. brevis K4 | 25/SS | 12/SS | 0/R | 0/R | 0/R | 25/SS | 10/MS | 15/SS | 0/R | 11/SS |
| L. brevis K5 | 23/SS | 15/SS | 0/R | 0/R | 0/R | 27/SS | 11/MS | 25/SS | 0/R | 11/SS |
| L. plantarum K6 | 28/SS | 18/SS | 0/R | 0/R | 0/R | 26/SS | 0/R | 24/SS | 0/R | 8/MS |
| L. brevis K7 | 27/SS | 20/SS | 0/R | 10/MS | 0/R | 30/SS | 15/SS | 30/SS | 0/R | 16/SS |
| L. brevis K8 | 25/SS | 23/SS | 0/R | 0/R | 0/R | 33/SS | 20/SS | 30/SS | 0/R | 25/SS |
| L. helveticus WC1 | 37/SS | 20/SS | 0/R | 11/SS | 9/MS | 31/SS | 13/SS | 28/SS | 0/R | 0/R |
| L. helveticus WC2 | 28/SS | 18/SS | 0/R | 10/MS | 13/SS | 34/SS | 13/SS | 25/SS | 0/R | 0/R |
| L. helveticus WC3 | 45/SS | 19/SS | 0/R | 0/R | 0/R | 35/SS | 20/SS | 30/SS | 0/R | 14/SS |
| L. helveticus WC4 | 36/SS | 27/SS | 0/R | 10/MS | 0/R | 35/SS | 10/MS | 29/SS | 0/R | 13/SS |
| L. helveticus WC5 | 40/SS | 30/SS | 0/R | 12/SS | 17/SS | 37/SS | 14/SS | 31/SS | 0/R | 0/R |
| L. plantarum WC6 | 33/SS | 25/SS | 0/R | 13/SS | 15/SS | 34/SS | 18/SS | 29/SS | 0/R | 0/R |
| L. plantarum I1 | 35/SS | 27/SS | 0/R | 0/R | 15/SS | 36/SS | 14/SS | 30/SS | 0/R | 0/R |
| L. plantarum I2 | 35/SS | 20/SS | 0/R | 38/SS | 0/R | 0/R | 19/SS | 30/SS | 0/R | 0/R |
| L. brevis I3 | 35/SS | 26/SS | 0/R | 37/SS | 15/SS | 0/R | 14/SS | 29/SS | 0/R | 0/R |
| L. helveticus I4 | 30/SS | 26/SS | 0/R | 35/SS | 0/R | 0/R | 19/SS | 30/SS | 0/R | 9/MS |
| L. plantarum I5 | 31/SS | 26/SS | 0/R | 11/SS | 12/SS | 34/SS | 14/SS | 28/SS | 0/R | 0/R |
| L. brevis I 6 | 40/SS | 25/SS | 0/R | 8/MS | 17/SS | 36/SS | 11/SS | 31/SS | 0/R | 0/R |
A/S, ampicillin/sulbactam; T, tetracycline; VA, vancomycin; CP, ciprofloxacin; CF, cephalothin; C, chloramphenicol; G, gentamicin; E, erythromycin; S, streptomycin; N, neomycin.
Distinguishing between intrinsic (non-specific, non-transferable) and acquired resistance by the beneficial strains is also necessary [68]. Molecular techniques that directly screen bacterial strains for the presence of antibiotic resistance genes could be considered an important addition to phenotypic testing. Usually, these include PCR-based methods [69, 70] or DNA microarrays that combine various antibiotic resistance genes [71].
Phenotypic analyses showed that all LAB isolates in this study, belonging to the
family Lactobacillaceae, showed resistance to one or more antibiotics.
To clarify the nature of this antibiotic resistance, PCR amplification was
performed with specific primers for acquired resistance genes: Gentamicin,
aac(6′)-aph(2′′); chloramphenicol, cat; tetracycline,
tet(M);
This study analyzed 45 LAB strains originating from traditional Bulgarian fermented foods. Species identification revealed the presence of five species: L. plantarum, L. brevis, L. helveticus, L. koreensis, and L. yonginensis. The strains isolated from fermented milk products most commonly belonged to the species L. plantarum, L. brevis, and L. helveticus, as well as in fermented vegetable foods—the dominant species was L. plantarum. This study reports the isolation of two species, L. koreensis and L. yonginensis, from Bulgarian fermented vegetable foods for the first time. All strains were tested for a set of abilities regarding their beneficial potential and safety application. We can summarize some significant findings: Among the strains, there were several that showed an apparent antagonistic effect, mainly against Gram-negative pathogenic bacteria; the ability of autoaggregation, co-aggregation, adhesion to hydrocarbons (hexane), and biofilm production were differentially represented among strains; we established that the observed phenotypic antibiotic resistance to different antibiotics is not the result of the presence of resistance genes and none of the strains showed hemolytic activity.
In conclusion, traditional fermented Bulgarian foods can be considered sources of variable beneficial LAB. The main observation in our study was that none of the tested strains simultaneously met all probiotic criteria. However, it can be proposed that the complex action of the LAB members in the microflora of fermented foods prevents the development of pathogenic bacteria and creates safe food. Further, a significant number of strains possessing beneficial properties were isolated from the traditional Bulgarian food, sauerkraut; thus, it can be considered a functional food. Moreover, this type of cabbage preservation is strictly typical in our country; therefore, we can speculate that its natural microbial diversity is some kind of endemic. However, the obtained preliminary results for the beneficial potential of traditional Bulgarian fermented foods, reported in this paper, could serve as a solid basis for further elaboration of complex starter cultures with the potential for biotechnological applications.
All used materials and generated data are included in this manuscript.
Substantial contributions to conception and design: VYMY, PKH and YKK; performed the research: VYMY; analysis and interpretation of data: PKH, YKK and IKR; writing the original draft of the manuscript: VYMY; review the draft version of the manuscript: PKH and YKK; All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work. All authors contributed to editorial changes in the manuscript.
Not applicable.
Not applicable.
This work was supported by project No. 80-10-30/ 20.04.2023, Sofia University “St. Kliment Ohridski”.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.




