1 Department of Thoracic Surgery, Affiliated Hospital of Southwest Medical University, 646000 Luzhou, Sichuan, China
Abstract
Background: Immune escape is a key factor influencing survival rate of
lung adenocarcinoma (LUAD) patients, but molecular mechanism of ubiquitin binding
enzyme E2T (UBE2T) affecting immune escape of LUAD remains unclear. The objective
was to probe role of UBE2T in LUAD. Methods: Bioinformatics means were
adopted for analyzing UBE2T and forkhead box A1 (FOXA1) expression in LUAD
tissues, the gene binding sites, the pathway UBE2T regulates, and the correlation
between UBE2T and glycolysis genes. Dual luciferase and chromatin
immunoprecipitation (ChIP) assays were conducted for validating the binding
relationship between the two genes. Quantitative reverse transcription polymerase
chain reaction (qRT-PCR) and western blot were employed to evaluate UBE2T, FOXA1,
and programmed death ligand 1 (PD-L1) levels in cancer cells. MTT assay was conducted for detecting cell
viability. Cytotoxicity assay detected CD8
Keywords
- FOXA1
- UBE2T
- glycolysis
- lung adenocarcinoma
- CD8+T cells
Currently, lung cancer is implicated in an unsatisfactory prognosis [1]. Among lung cancers, proportion of non-small cell lung cancer (NSCLC) is about 80%, and most of the NSCLCs are lung adenocarcinomas (LUADs) [2, 3]. In recent years, reversing effector T cell failure by immune checkpoint blocking with reagents such as programmed cell death protein 1 (PD-1) and programmed death ligand 1 (PD-L1) antibodies is a regular method for lung cancer [4]. However, a sizable percentage of patients did not respond to these manipulations, partly due to immune escape of tumor cells during contact with immune cells in microenvironment. Hence, exploring the mechanism of immune evasion is a necessary way to improve patient’s survival rate and can bring a new choice for the precise treatment of LUAD.
The process by which tumor cells avoid immune system monitoring and use various
strategies to facilitate their migration and invasion is known as tumor immune
escape [5]. PD-L1 shows a critical role in keeping immune homeostasis, in which
PD-L1 on tumor cell surface interplays with PD-1 in T cells to inhibit T
lymphocyte activation, proliferation, and cytokine secretion [6]. In the tumor
microenvironment (TME), this approach is adopted by cancer cells to avoid
T-cell-mediated tumor-specific immunity [6]. Numerous studies have investigated
the mechanism by which PD-L1 influences activity of CD8
Metabolic reprogramming is a signature of cancer [9]. Aerobic glycolysis, namely “Warburg effect”, is the most commonly seen mode of metabolic reprogramming and also the common way for cancer cells to metabolize glucose [10]. Somatic cells synthesize adenosine triphosphate (ATP) by oxidative phosphorylation, while tumor cells generate lactic acid by glycolysis [11]. Numerous reports supported the view that glycolysis regulates tumor development. As a study shows, succinate dehydrogenase B (SDHB) hinders occurrence and development of renal clear cell carcinoma by inhibiting glycolysis [12]. According to Zhao et al. [13], lncRNA MIR17HG facilitates colorectal cancer liver metastasis by mediating a positive feedback loop related to glycolysis. Furthermore, glycolysis metabolism was found to be associated with PD-L1-mediated immune evasion in TME of malignant tumors [14]. Glucose metabolism reprogramming drives immune evasion in hepatocellular carcinoma and pancreatic cancer [15, 16]. For example, high glucose stimulates pancreatic cancer cells to evade immune surveillance through the AMPK-Bmi1-GATA2-MICA/B pathway [17]. However, it is not clear how glycolytic metabolism in LUAD regulates tumor immune escape.
Ubiquitin binding enzyme E2T (UBE2T) is a representative E2 containing UBC
domain that can link to ring finger or HECT domain of E3 and promotes
mono-ubiquitination or poly-ubiquitination of substrates [18]. Accumulating
evidence shows that UBE2T exerts a carcinogenic effect in varying cancers and is
closely linked to cancer occurrence and progression, including hepatocellular
carcinoma [19], gastric cancer [20], and multiple myeloma [21]. The function of
UBE2T in development of lung cancer has also been investigated. For example,
UBE2T up-regulates the autophagy of NSCLC cells through activating the
p53/AMPK/mTOR pathway [18]. UBE2T promotes radiation resistance of NSCLC through
driving epithelial-mesenchymal transition (EMT) and FOXO1 degradation mediated by
ubiquitination [22]. In addition, UBE2T level is linked to infiltration of immune
cells. For example, Wang et al. [23] disclosed that UBE2T level is
substantially increased in retinoblastoma and is positively correlated with Th2
cells, acting as a prognostic biomarker for retinoblastoma. Similarly, survival
analysis presented that UBE2T is increased in breast cancer, and its
up-regulation is linked with unfavorable prognosis. UBE2T up-regulation is linked
to Th1 and Th2 cell balance, and Th1/Th2 balance shifts to Th2 in basal and
Luminal-B breast cancer (p
One pioneer factor that can bind to concentrated chromatin and start chromatin remodeling is the forkhead box A1 (FOXA1) protein [25]. Target genes are transcriptionally activated when FOXA1 interacts to its co-factors. FOXA1 mutations are a defining characteristic of estrogen receptor-positive (ER) breast cancer and contribute to the disease advancement through estrogen receptor-binding events [26]. When combined with HOXB13, FOXA1 can cause normal prostate epithelial cells to undergo a transformation [27]. SOX9 up-regulates FOXA1 and drives tumorigenic potential of lung cancer cells [28]. Thus, targeted interference with FOXA1 may serve as a substitute approach for LUAD therapy.
In this work, we focused on clarifying function of UBE2T in LUAD. As our results
displayed, UBE2T was increased in LUAD tissues and cells and repressed activity
of CD8
mRNA expression data (59 normal cases, 539 tumor cases) of LUAD were downloaded
from The Cancer Genome Atlas (TCGA) and subjected to differential analysis (
Human lung bronchial epithelial cells (BEAS-2B), LUAD cells (A549, H460, and
H1650), 293T cells, and peripheral blood mononuclear cells (PBMCs) were purchased
from ATCC (Manassas, VA, USA), and mouse LUAD cells (LA795) were purchased from
Pricella (Wuhan, China). BEAS-2B cells were maintained in MEM with 10% fetal
bovine serum (FBS; PAN, Aidenbach, Germany), A549 cells in F12K medium
(Invitrogen, Waltham, MA, USA) containing 10% FBS (PAN, Germany), H460 cells and
H1650 cells in RPMI-1640 medium (Invitrogen, Waltham, MA, USA) with 10% FBS
(PAN, Germany), 293T cells in DMEM (Invitrogen, Waltham, MA, USA) with 10% FBS
(PAN, Germany) and 2 mM L-glutamine (Invitrogen, Waltham, MA, USA), and PBMC
cells in HBSS medium (Invitrogen, Waltham, MA, USA) containing 10% FBS (PAN,
Germany). LA795 cells were placed in RPMI-1640 culture-medium (Invitrogen,
Waltham, MA, USA) mixed with 10% FBS (PAN, Germany), streptomycin (100
µg/mL, Corning, Shanghai, China), penicillin G (100 U/mL, Beyotime,
Shanghai, China). The cultured cells were incubated in humidified atmosphere with
5% CO
Short hairpin RNAs (shRNAs) targeting UBE2T and FOXA1 (sh-UBE2T, sh-FOXA1) and their negative controls (sh-NC) were produced by GenePharma (Shanghai, China), and the carrier pcDNA-UBE2T and its negative control pcDNA-NC were accessed from RiboBio (Guangzhou, China). As per the instructions, sh-UBE2T, sh-FOXA1, sh-NC, pcDNA-UBE2T, and pcDNA-NC were used to transfect LUAD cells using Lipofectamine 2000 (Invitrogen, Waltham, MA, USA).
Total RNA extraction from tissues and cells was conducted by using TRIzol
reagents (Invitrogen, Waltham, MA, USA), and RNA concentration and purity were
quantitatively determined with a NanoDrop ND-1000 spectrophotometer (NanoDrop,
Waltham, MA, USA). Revert Aid™ First Strand cDNA Synthesis Kit
(Thermo Fisher, Waltham, MA, USA) was applied to synthesize cDNA from RNA. ChamQ
SYBR Color qPCR Master Mix (High ROX Premixed; Vazyme, Nanjing, China) was
utilized to perform qPCR on ABI 7500 PCR instrument (Applied Biosystems, Waltham,
MA, USA). Total reaction volume was 20 µL. Relative expression of genes was
computed by applying 2
| Primer | Forward (5 |
Reverse (5 |
| UBE2T | ATCCCTCAACATCGCAACTGT | CAGCCTCTGGTAGATTATCAAGC |
| LDHA | TGGAGATTCCAGTGTGCCTGTATGG | CACCTCATAAGCACTCTCAACCACC |
| GLUT1 | CTTTGTGGCCTTCTTTGAAGT | CCACACAGTTGCTCCACAT |
| HK2 | GAGCCACCACTCACCCTACT | CCAGGCATTCGGCAATGTG |
| FOXA1 | ACAGCTACTACGCAGACACG | ACAGCTACTACGCAGACACG |
| CTACGTCGCCCTGGACTTCGAGC | GATGGAGCCGCCGATCCACACGG |
UBE2T, ubiquitin binding enzyme E2T; GLUT1, glucose transporter type 1; LDHA, lactate dehydrogenase A; HK2, hexokinase 2; FOXA1, forkhead box A1.
LUAD cells were inoculated into 96-well plates (1
Total protein extraction from tissues and cells was performed with RIPA buffer
(Sigma, St.Louis, MO, USA), followed by protein concentration determination by a
BCA protein detection kit (Thermo Fisher, Waltham, MA, USA). SDS-PAGE was run for
protein isolation with 10% polyacrylamide gel (Beyotime, Shanghai, China) and
transferred to a polyvinylidene fluoride membrane (Beyotime, Shanghai, China).
Thereafter, membrane was incubated with primary antibodies (Abcam, Cambridge, UK)
of the proteins PD-L1 (1:1000), UBE2T (1:1000), GLUT1 (1:100,000), LDHA (1:5000),
HK2 (1:1000) and FOXA1(1:5000) overnight. The next step was incubation for 1 h
with second antibody IgG (Abcam, Cambridge, UK) at room temperature.
T cells were purified from PBMCs using magnetic beads from Miltenyi Biotech
(Bergisch Gladbach, North Rhine-Westphalia, Germany). Then, CD8
CD8
Interferon
We measured cellular glycolysis capacity and mitochondrial function by extracellular flow analyzer (Seahorse Bioscience, North Billerica, MA, USA) as per instructions. The day before assay, cells were placed in cell culture microporous boards (Seahorse Bioscience, North Billerica, MA, USA). Seahorse buffer was a mixture of DMEM, 2 mM sodium pyruvate, phenol red, 25 mM glucose, and 2 mM glutamine. For ECAR value test, 10 mM glucose, 1 µM oligomycin, and 100 mm 2-DG were automatically added. Following baseline respiration monitoring, 1 µM oligomycin, 1 µM carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone (FCCP), and 1 rotenone were automatically injected into the microporous boards to assess OCR [1].
The fragment of UBE2T was inserted into pGL3 promoter vector to construct pGL3-UBE2T-promoter-WT (wild type) or pGL3-UBE2T-promoter-MUT (mutation type) vector. The luciferase reporter vector and sh-FOXA1/sh-NC were used for co-transfection of 293T cells by Lipofectamine 2000 reagent. The 293T cells were incubated for 48 h, followed by luciferase activity measurement in dual luciferase assay system.
After 37% formaldehyde (Beyotime, Shanghai, China) treatment, the cells were
collected for sonication with VCX750 (SONICS, Newtown, CT, USA) at 25% power.
Sonication was performed for 4.5 s each time, with 14 times in total and 9 s
intervals. Next, centrifugation at 10,000
The paraffin-embedded sample was sliced into 4 µm-thick slices using a paraffin microtome (Leica, Wetzlar, Germany). IHC staining was completed according to the standard protocol described previously [18]. After initial dewaxing, antigen repair, and sealing, the slices were incubated overnight with anti-UBE2T antibody (Abcam, Cambridge, UK) in a humid container at 4 °C. The next day, slices were treated with horseradish peroxidase-labeled secondary antibody IgG (Abcam, Cambridge, UK). DAB (Novus Biologicals, Littleton, CO, USA) was adopted to develop UBE2T staining, and hematoxylin counterstaining was conducted.
Thirty male mice (BALB/c, 4 weeks old, 18–22 g) were from SLAC Laboratory
Animal Co., Ltd. (Shanghai, China). Animal experiments were conducted with
approval from the Ethics Committee of Affiliated Hospital of Southwest Medical
University. Mice were kept in controlled conditions with 25
Data were processed using GraphPad Prism 8.0 (GraphPad Software, Inc., San
Diego, CA, USA). All measured data were presented in the form of mean
Previous studies have found that UBE2T is up-regulated and promotes cancers of various types, such as hepatocellular carcinoma [34] and esophageal squamous cell carcinoma [35]. Hence, we explored UBE2T expression in LUAD. Bioinformatics results showed a significant up-regulation of UBE2T in LUAD tissues (Fig. 1A). qRT-PCR and western blot detection also showed an evident higher UBE2T expression in LUAD cells than in normal lung epithelial cells (Fig. 1B,C). These results suggested the aberrantly high expression of UBE2T in LUAD. Since UBE2T had higher relative expression in A549 cells and lower expression in H460 cells among the tested cell lines, A549 was selected for the subsequent knockdown experiment and H460 for the overexpression experiment.
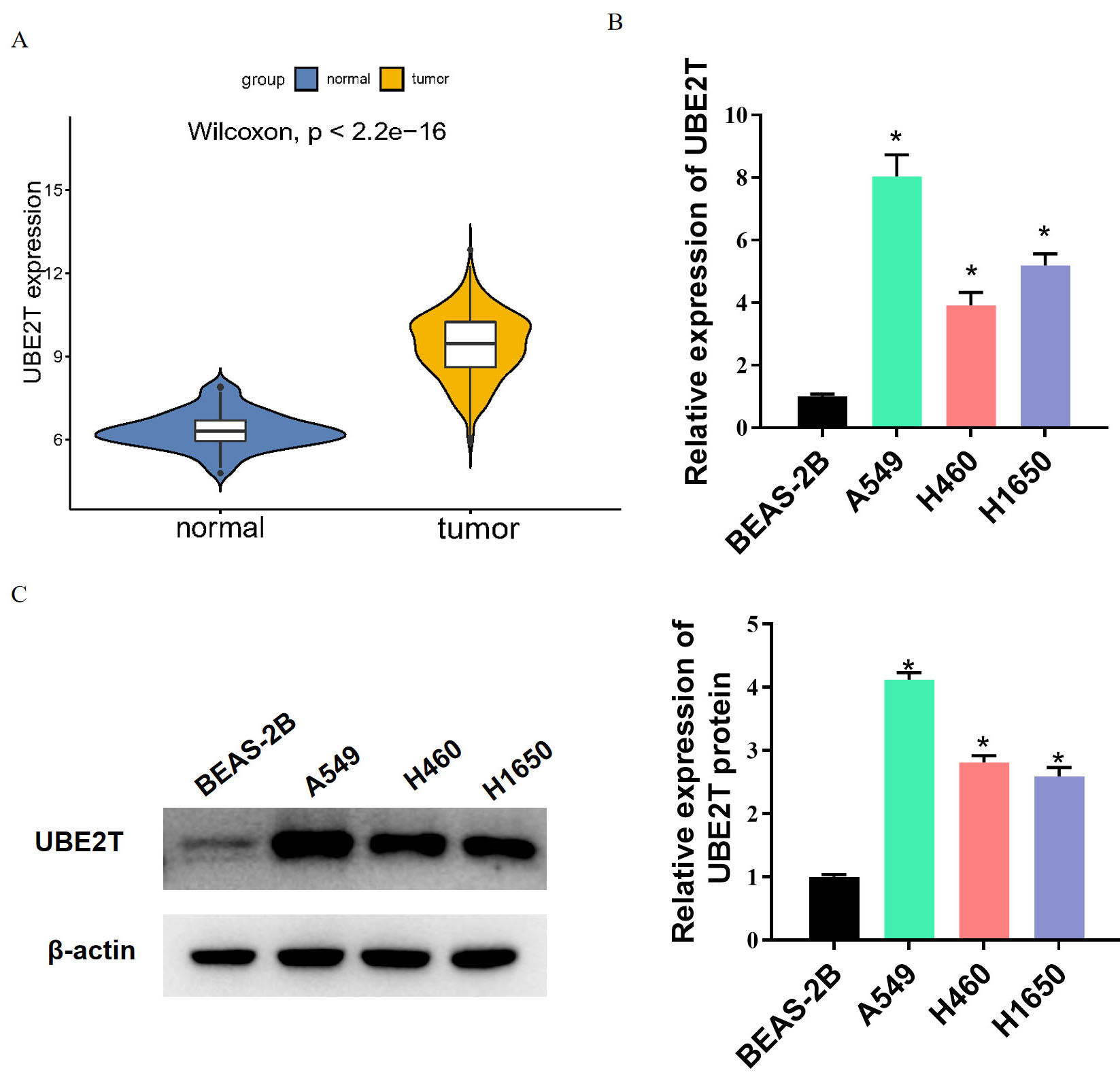 Fig. 1.
Fig. 1.UBE2T expression is up-regulated in LUAD. (A) UBE2T expression
in LUAD tissues analyzed by bioinformatics approaches. (B,C) Expression of UBE2T
in LUAD cells analyzed by western blot and qRT-PCR. * indicates p
UBE2T is associated with immune cell infiltration [24]. We found by
bioinformatics analysis that UBE2T had a negative correlation with CD8
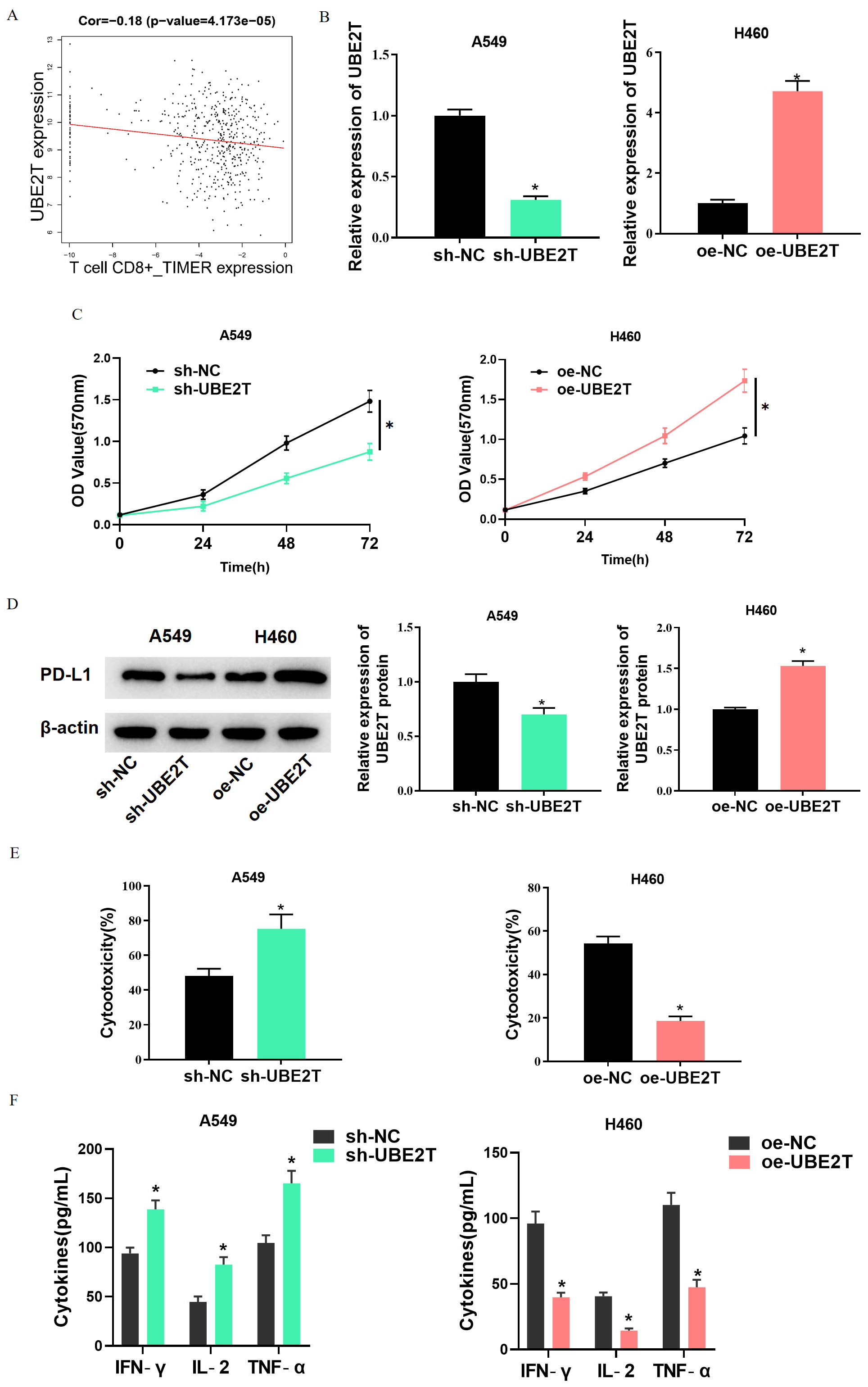 Fig. 2.
Fig. 2.High expression of UBE2T inhibits the activity of CD8
Next, we investigated mechanism that UBE2T affects activity of CD8
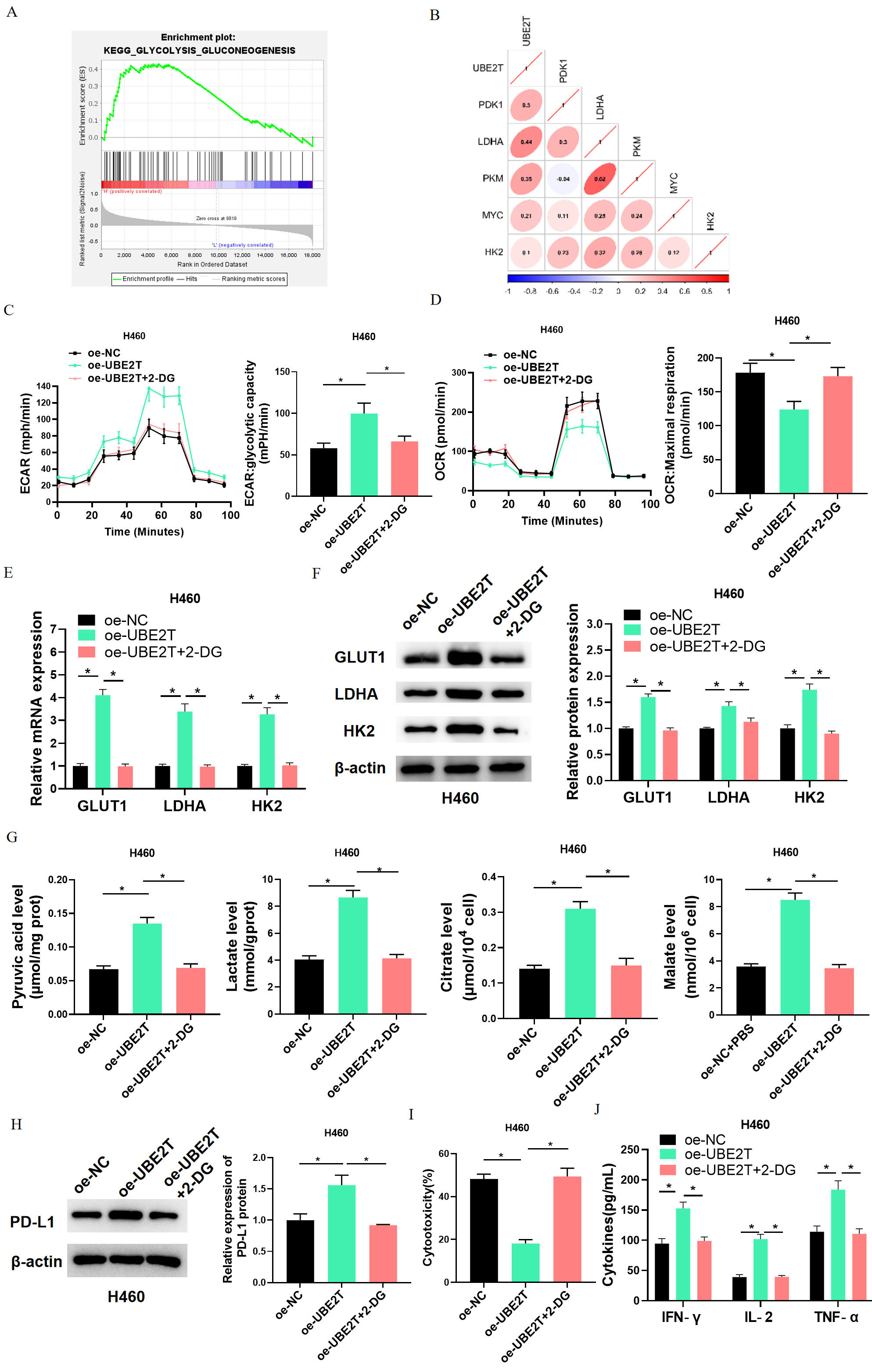 Fig. 3.
Fig. 3.High expression of UBE2T inhibits CD8
Next, H460 cells with oe-NC, oe-UBE2T, or oe-UBE2T+2-DG were subjected to
co-culture with activated CD8
To explore the transcription factors upstream of UBE2T, hTFtarget was utilized to predict UBE2T upstream potential transcription factors, which were then intersected with up-regulated DEmRNAs. Finally, a total of 17 potential transcription factors were obtained (Fig. 4A). FOXA1 is associated with the growth and cell characteristics of LUAD [36]. Further, Pearson correlation analysis demonstrated a positive correlation between UBE2T and FOXA1 (Fig. 4B). JASPAR prediction showed a binding site of UBE2T to FOXA1 in the promoter region of UBE2T (Fig. 4C). Bioinformatics analysis told high FOXA1 expression in LUAD tissues (Fig. 4D), while qRT-PCR and western blot results presented up-regulated FOXA1 level in LUAD cells (Fig. 4E,F). Thereafter, we continued to probe mechanism of influence of FOXA1 and UBE2T on LUAD. As the FOXA1 level in A549 cells was the highest among the tested cell lines, A549 was selected for subsequent experiments. Subsequent ChIP assay showed that FOXA1 antibody could significantly enrich UBE2T compared with IgG (Fig. 4G). The dual luciferase assay found that sh-FOXA1 led to a substantially reduced luciferase activity in UBE2T-WT but caused no great change in the UBE2T-MUT group (Fig. 4H). Finally, the expression of UBE2T in sh-FOXA1-treated A549 cells was assayed by qRT-PCR. UBE2T was significantly down-regulated (Fig. 4I). The above experiments showed that FOXA1 activated UBE2T transcription.
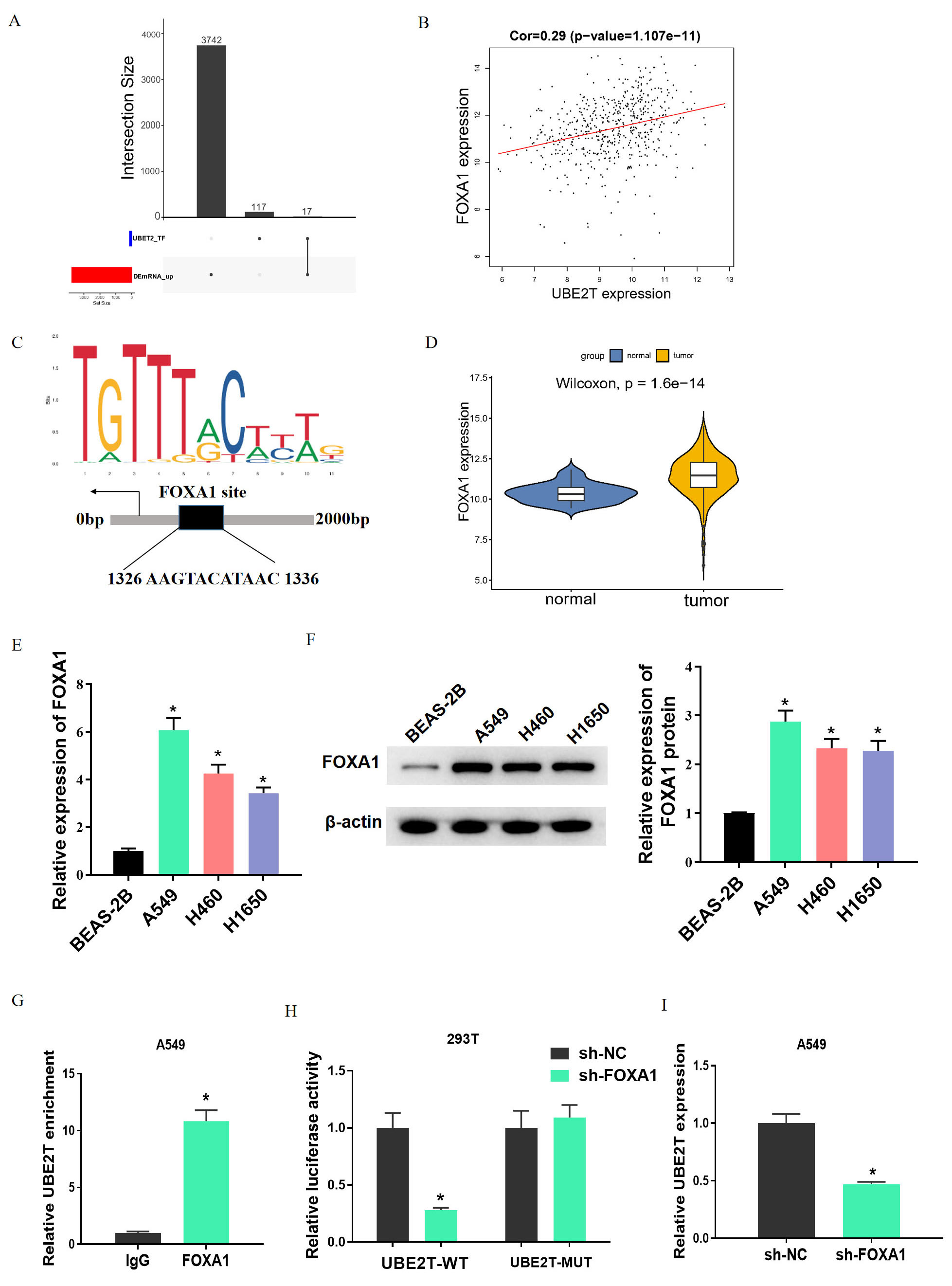 Fig. 4.
Fig. 4.FOXA1 activates UBE2T transcription. (A) Upset plot of the
intersection between hTFtarget predicted transcription factors and up-regulated
DEmRNAs. (B) Pearson correlation analysis of UBE2T and FOXA1. (C) JASPAR
predicted binding site between UBE2T and FOXA1. (D) FOXA1 expression in LUAD
analyzed by bioinformatics means. (E,F) qRT-PCR and western blot results on FOXA1
expression in LUAD cells. (G,H) ChIP and dual luciferase assay results on the
binding relationship between UBE2T and FOXA1. (I) qPCR results on UBE2T
expression after sh-FOXA1 treatment. * indicates p
To verify the influence of FOXA1/UBE2T on CD8
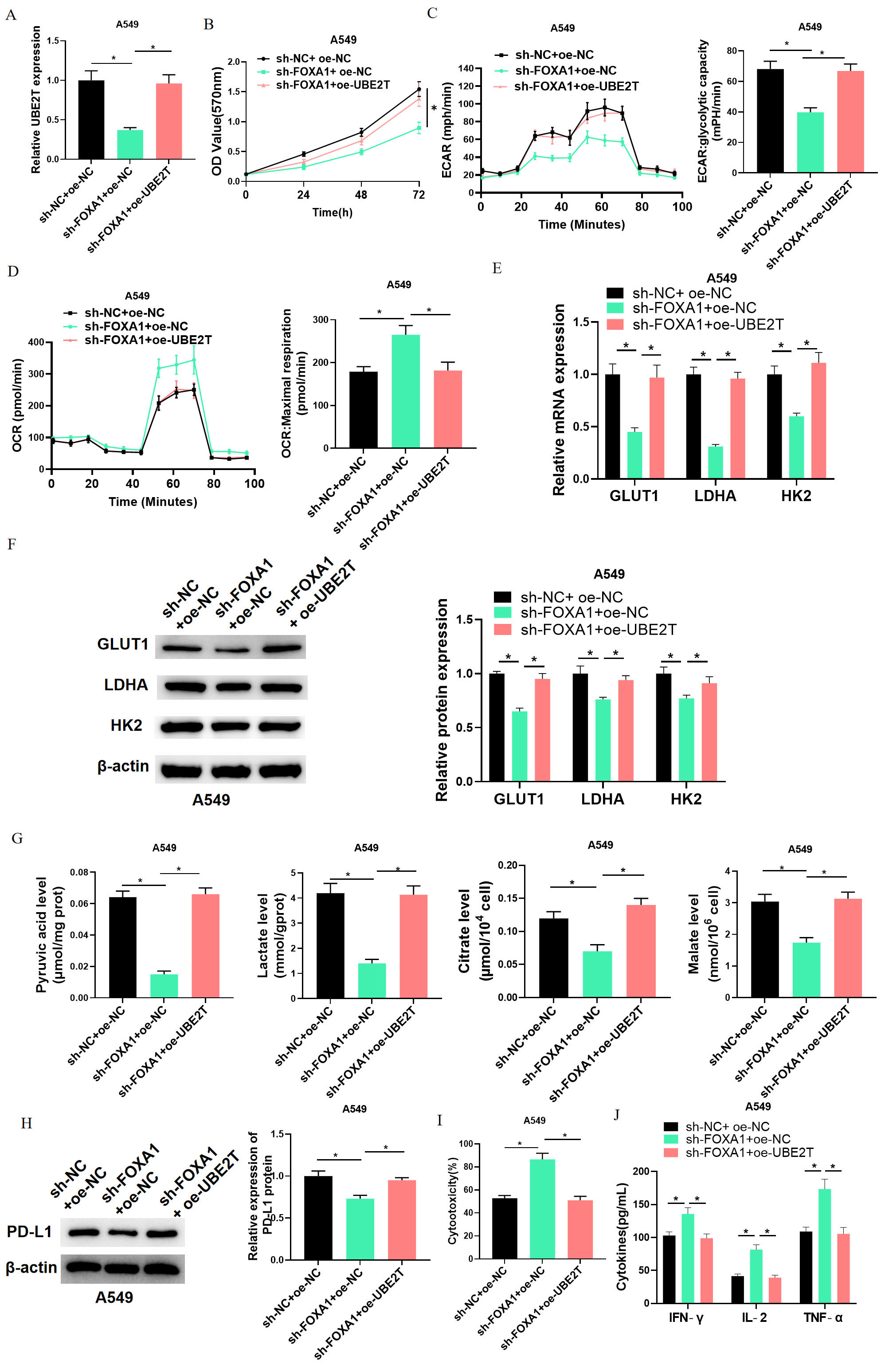 Fig. 5.
Fig. 5.FOXA1/UBE2T inhibits CD8
These cells were subjected to co-culture with activated CD8
We constructed an allograft LUAD model to investigate the tumorigenicity of
UBE2T in vivo. Compared with sh-NC, sh-UBE2T had a significant reduction
effect on tumor volume and weight (Fig. 6A–C). qRT-PCR, western blot, and IHC
analyses showed decreased UBE2T levels and PD-L1 expression after UBE2T knockdown
(Fig. 6D,E). According to IHC results, mice treated with sh-UBE2T cells had
higher infiltration of CD8
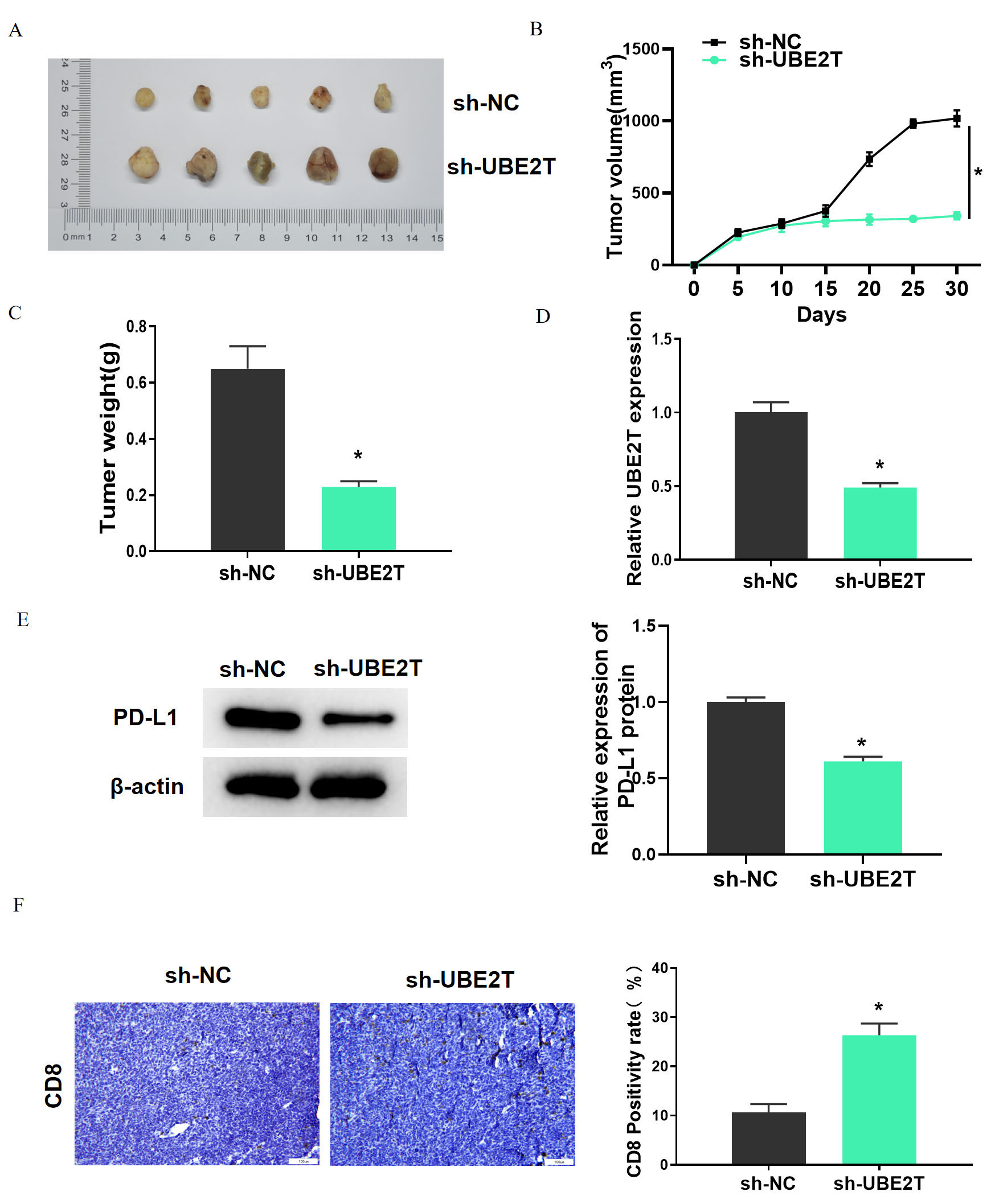 Fig. 6.
Fig. 6.UBE2T can promote lung adenocarcinoma tumor growth and inhibit
CD8
LUAD is a malignancy characterized by uncontrolled cell growth in the lungs and bronchus [37]. Currently, the treatment methods of LUAD are mainly surgical resection, chemotherapy, radiotherapy, and targeted therapy, but the clinical outcomes are unsatisfactory due to the lack of reliable biomarkers [38]. Hence, an effective molecular marker is in need to project early onset of LUAD and formulate therapies. Recently, immunotherapy targeting immune checkpoints has shown great promise in cancer treatment, and immune escape is the biggest obstacle to effective cancer treatment [39]. However, the molecular mechanism of immune escape of LUAD has not been perfected. Hence, this study aimed to search for new biomarkers as a theoretical basis for new therapies for LUAD.
UBE2T is an oncogene whose up-regulated expression has been noticed in many kinds of cancer. Moreover, the carcinogenic effect of UBE2T in LUAD has also been demonstrated. For example, UBE2T is up-regulated in LUAD, targeted by NEDD4, and inhibits PI3K-AKT signal through ubiquitination and degradation, which collectively lead to LUAD progression [40]. As Zhu et al. [18] found, UBE2T overexpression could promote the proliferation of LUAD cells and autophagy triggered by cisplatin, resulting in cisplatin resistance of A549 cells. Here we also found an aberrant high UBE2T expression in LUAD, which is in line with previous studies. Additionally, UBE2T participates in metabolic reprogramming during the development of cancer. For example, UBE2T features in promoting tumor cell proliferation, invasion, and glycolysis via PI3K/AKT pathway in breast cancer [29]. Through bioinformatics analysis, we know that a high UBE2T level in LUAD could activate glycolysis signaling pathways. This suggests that UBE2T affecting the progression of LUAD through glycolysis is one of the mechanisms. UBE2T has a potentially pivotal role in LUAD progression.
Immune escape is a critical pathway to avoid T cell-mediated specific immune
killing for tumor cells in the TME [32]. A diet low in glucose enhances the host
lung immune responses and hinders the formation of tumors in experimental lung
adenocarcinomas [41]. Cancer cells undergo a high glucose consumption and lactic
acid production during the reprogramming of glucose metabolism, which leaves the
TME deficient in oxygen and energy. This has a significant effect on immunity and
helps cancer cells evade immune surveillance [15]. Guo et al. [42]
discovered that in human glioblastoma cells, aerobic glycolysis mediated by
hexokinase 2 (HK2) upregulates PD-L1 via phosphorylation of
I
We also found that FOXA1 was the upstream transcription factor for UBE2T, and UBE2T expression was positively regulated by this factor. FOXA1 has been reported to regulate cancer development by activating target genes. For example, FOXA1 can act with the promoter of TGFB3 and inhibit its accumulation in the nuclear; miR-93-5p can reduce FOXA1 expression and elevate TGFB3 expression; thus, apoptosis of colorectal cancer cells is inhibited [43]. FOXA1 promotes cell cycle progression in chondrosarcoma cells by activating the cyclin B1 expression [44]. A clinical study showed that adjuvant endocrine therapy is implicated in FOXA1 downregulation in pleural metastases [45]. FOXA1 intimately participates in production and function of regulatory T cells (Tregs). Liang et al. [46] demonstrated that FOXA1 Tregs are enhanced in lung cancer and suppress anti-tumor immunity of T cells. In the current study, we found through bioinformatic analysis that FOXA1 was a transcription factor for UBE2T. By binding to UBE2T promoter, FOXA1 targeted UBE2T expression and promoted LUAD immune escape by activating the glycolysis pathway.
However, our experiment was conducted only at the cellular and animal levels,
and the repeated experimental results were not collected from clinical LUAD
patients, which is a limitation. In addition, specific molecular mechanism of
UBE2T activation of aerobic glycolysis affecting activity of CD8
Collectively, according to our results, UBE2T was significantly overexpressed in
LUAD; UBE2T expression was positively regulated by FOXA1, and UBE2T up-regulated
PD-L1 expression by activating glycolytic metabolism, thus inhibiting CD8
The data and materials in the current study are available from the corresponding author on reasonable request.
Conceptualization: JP. Visualization: KW and BW. Acquiring: ZY. Analyzing: QS and DZ. Interpreting data: PZ, WY and ZY. Resources: DZ. Writing - Original Draft: KW, ZY, JP and DZ. Writing - Review & Editing: QS, PZ, BW and WY. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The animal experiments in this study were performed with the approval from the Ethics Committee of Affiliated Hospital of Southwest Medical University (No. 20221206-007).
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






