1 Department of Anorectal Surgery, Changhai Hospital, Naval Medical University (Second Military Medical University), 200433 Shanghai, China
2 Department of Anorectal Surgery, Changzheng Hospital, Naval Medical University (Second Military Medical University), 200003 Shanghai, China
3 Department of Radiation Medicine, Faculty of Naval Medicine, Naval Medical University (Second Military Medical University), 200433 Shanghai, China
4 Department of General Surgery, Tongji Hospital, School of Medicine, Tongji University, 200065 Shanghai, China
†These authors contributed equally.
Abstract
Background: The cause of ulcerative colitis (UC) is not yet fully
understood. Previous research has pointed towards a potential role for mutations
in nucleotide-binding oligomerization domain-containing protein 2 (NOD2) in promoting the onset and progression of inflammatory bowel disease
(IBD) by altering the microbiota of the gut. However, the relationship between
toll-like receptor 4 (TLR4) and gut microbiota in IBD is not well understood. To
shed light on this, the interaction between TLR4 and gut microbiota was studied
using a mouse model of IBD. Methods: To examine the function of TLR4
signaling in intestinal injury repair, researchers developed Dextran Sulfate
Sodium Salt (DSS)-induced colitis and injury models in both wild-type (WT) mice
and TLR4 knockout (TLR4-KO) mice. To assess changes in the gut microbiota, 16S
rRNA sequencing was conducted on fecal samples from both the TLR4-KO and WT
enteritis mouse models. Results: The data obtained depicted a protective
function of TLR4 against DSS-induced colitis. The gut microbiota composition was
found to vary considerably between the WT and TLR4-KO mice groups as indicated by
Keywords
- 16S rRNA sequencing
- toll-like receptor 4
- gut microbiota
- DSS-induced colitis
A chronic inflammatory disorder of the colon, ulcerative colitis (UC)
[1], together with Crohn’s disease (CD), constitutes two primary types of
inflammatory bowel disease (IBD) [2]. The etiology of IBD is still elusive and is
supposed to be associated with environmental stimuli, immunological factors, and
genetic susceptibilities [3]. Studies have demonstrated that immunodeficiency is
a major cause of inflammation and tissue damage [4]. Excessive activation of the
toll-like receptor (TLR)/nuclear factor
As a vital part of the body, gut microbiota in the digestive tract contains large numbers of microorganisms [8]. Development in the body of the mother and the breastfeeding pathways after birth shape the gut microbiota of the infant, which gradually matures with environmental exposure during childhood. The gut microbiota from late childhood to adulthood remains quite stable, but as the immune status declines, the diversity of gut microbiota will decrease in the elderly [9]. The core functions of mature healthy gut microbiota include the short-chain fatty acids (SCFA) production through complex polysaccharides fermentation and the formation of certain lipopolysaccharides (LPS), as well as the biosynthesis of some essential amino acids and vitamins [10]. Gut microbiota dysfunction may cause damage to the intestinal mucosal immune barrier, which is believed to be strongly associated with the onset and progression of UC [11]. Microbiota of the gut has been reported as vital concerning inflammation in IBD in various experiments [12], which may help better understand the interaction between the microbiota in IBD and the immune system. TLRs and the cytoplasmic receptor, known as the nucleotide-binding oligomerization domain-containing protein 2 (NOD2) [13], are sensors for pathogen-associated molecular patterns (PAMPs) that include peptidoglycan, lipoteichoic acid, endotoxin, flagellin, lipopolysaccharide (LPS) and muramyl-peptide (MDP) [14, 15, 16]. NOD2 mutations have been reported to have a significant impact on the gut microbiota composition by increasing the number of mucosa-adherent bacteria [17] and decreasing the anti-inflammatory cytokine interleukin 10 (IL-10) transcription [18], thereby promoting the development of IBD. However, the relationship between TLR4 and gut microbiota in IBD remains poorly understood.
A variety of chemically-induced colitis models have been most utilized on the
laboratory scale [19], among which the DSS-induced mouse colitis model is the
predominantly utilized IBD animal model. DSS is formed by the esterification of
dextran and chlorosulfonic acid, with the molecular formula of (C
Animals used in this study included C57BL/6 WT adult male mice weighing 25–30
g, and C57BL/6 TLR4 gene-deleted type TLR4
The DSS-induced colitis model in all WT and TLR4
On day five after DSS treatment, about 2 mm of the mouse tail was cut off, from which about 40 µL blood was sampled, and then transferred to a K2-EDTA containing Eppendorf (EP) tube, followed by repeated inversions of the tube. Blood cells were counted utilizing a small animal blood cell counter.
The mice were euthanized via cervical dislocation on the third or fifth day following DSS treatment. and the intestinal tissues were isolated for colon bleeding observation. The dehydration of the gut tissues was conducted in an ascending series of ethanol, cleared, paraffin-embedded, and serially sliced into 2–7 micron-thick sections utilizing a rotatory microtome (Aihua, Tianjin, China). Hematoxylin-eosin (H & E) staining was commonly used to stain the deparaffinized sections [24].
Six WT mice and six TLR4-KO mice were classified into two groups and orally administered with 1.5% DSS. Following DSS treatment, the mice in both groups were separately transferred to 12 clean cages lined with sterile filter paper on the seventh day. Immediately following defecation, fecal samples were placed in sterile centrifuge tubes and kept at –80 °C for later examination and transplantation [25].
Extraction of total genomic DNA through the DNA Extraction Kit (Tiangen Biotechnology Company, Beijing, China) was conducted following the instructions of the kit. DNA was quantified and its quality was examined through NanoDrop and agarose gel, respectively. Dilution of the extracted DNA to 1 ng/µL concentration was done and was subsequently kept at –20 °C until further use. Barcoded primers and Takara Ex Taq (Takara, Tokyo, Japan) were used in PCR to amplify the bacterial 16S rRNA genes using the diluted DNA as a template. Concerning the amplification of the 16S rRNA genes’ variable regions V3–V4, the universal primers 343 F and 798 R were employed. The primers were connected with an Illumina sequencing adapter with the reverse primer containing a sample barcode. PCR products were purified and the concentrations were adjusted for sequencing on an Illumina Miseq PE300 system (Oebiotech, Shanghai, China). Utilizing their distinctive barcodes, the samples’ raw sequencing reads were sorted. After removing the barcode and linker, they were subjected to primer sequencing and clustering to generate operational taxonomic units (OTUs) using the software V search with a 97% similarity threshold [26]. All sequences were classified through the NCBI BLAST and SILVA databases [27]. R software (R-4.2.2, Auckland, New Zealand) was employed to carry out the 16S rRNA analysis of the fecal samples. The raw data of 16S rRNA analysis obtained in this study are available in the Gene Expression Omnibus (GEO) database with accession number GSE227565.
Statistical treatment.Comparisons between TKR4
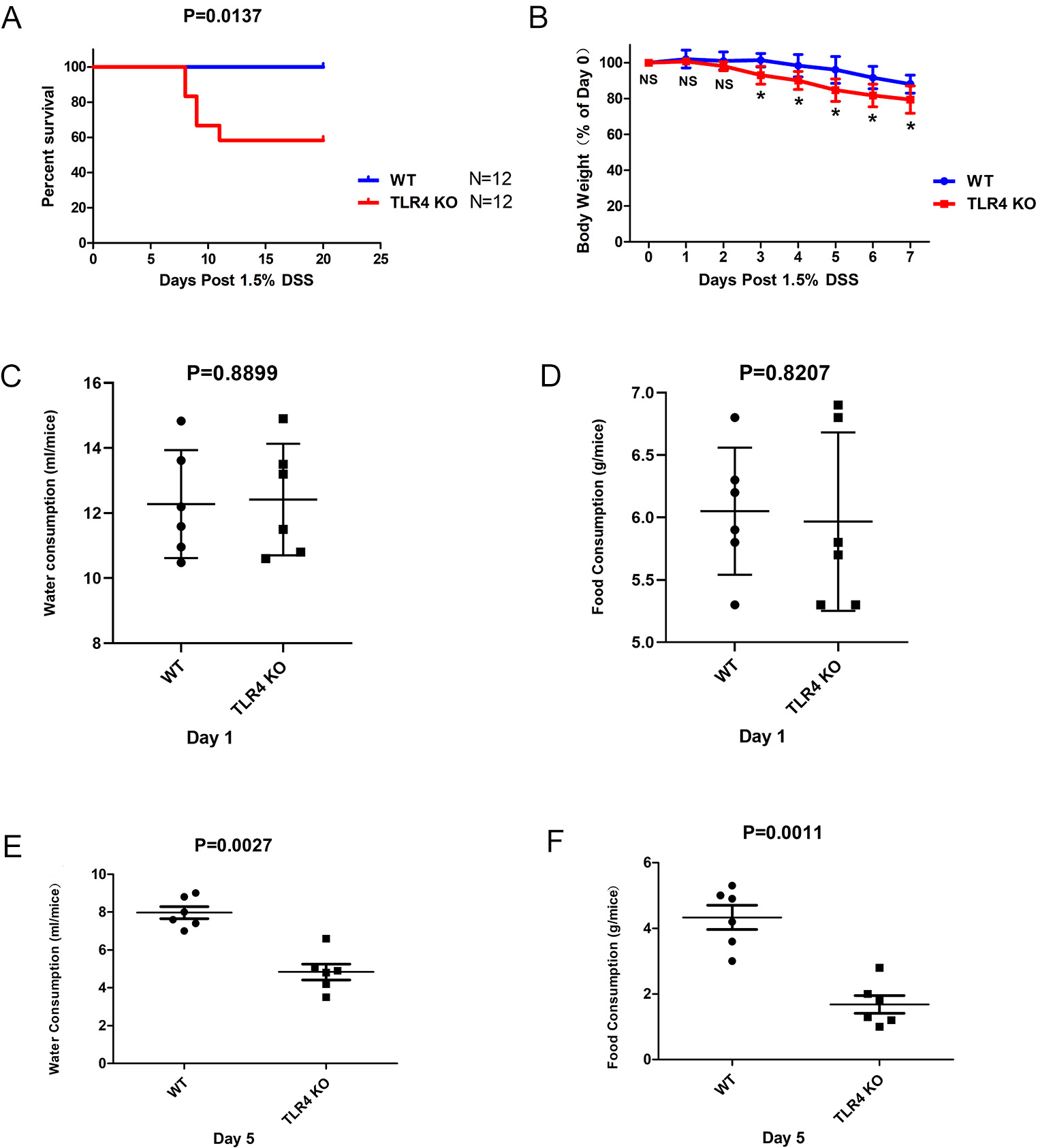
First, the function of TLR4 in DSS-induced intestinal injury was explored. It was found that the mortality in TLR4-KO mice was notably elevated in contrast to WT mice (p = 0.0137) (Fig. 1A). There were remarkable variations in the dynamic change of body weight between the two groups on day seven. TLR4-KO mice showed considerably more body weight loss than WT controls subsequent to orally administered 1.5% DSS (Fig. 1B). Simultaneously, the protective function of TLR4 against DSS-induced acute intestinal damage was analyzed using some basic daily behavioral parameters. The water and food consumption of WT and TLR4-KO mice treated with 1.5% DSS-supplemented water was recorded for seven consecutive days per mouse. The result showed that no significant difference in the water-drinking (p = 0.8899) and food-ingestion (p = 0.8207) abilities between TLR4-KO and WT mice at day 1 (Fig. 1C,D). However, the food-ingestion (p = 0.0011) and water-drinking (p = 0.0027) abilities of TLR4-KO mice began decreasing significantly from day five in comparison with those of the WT mice (Fig. 1E,F).
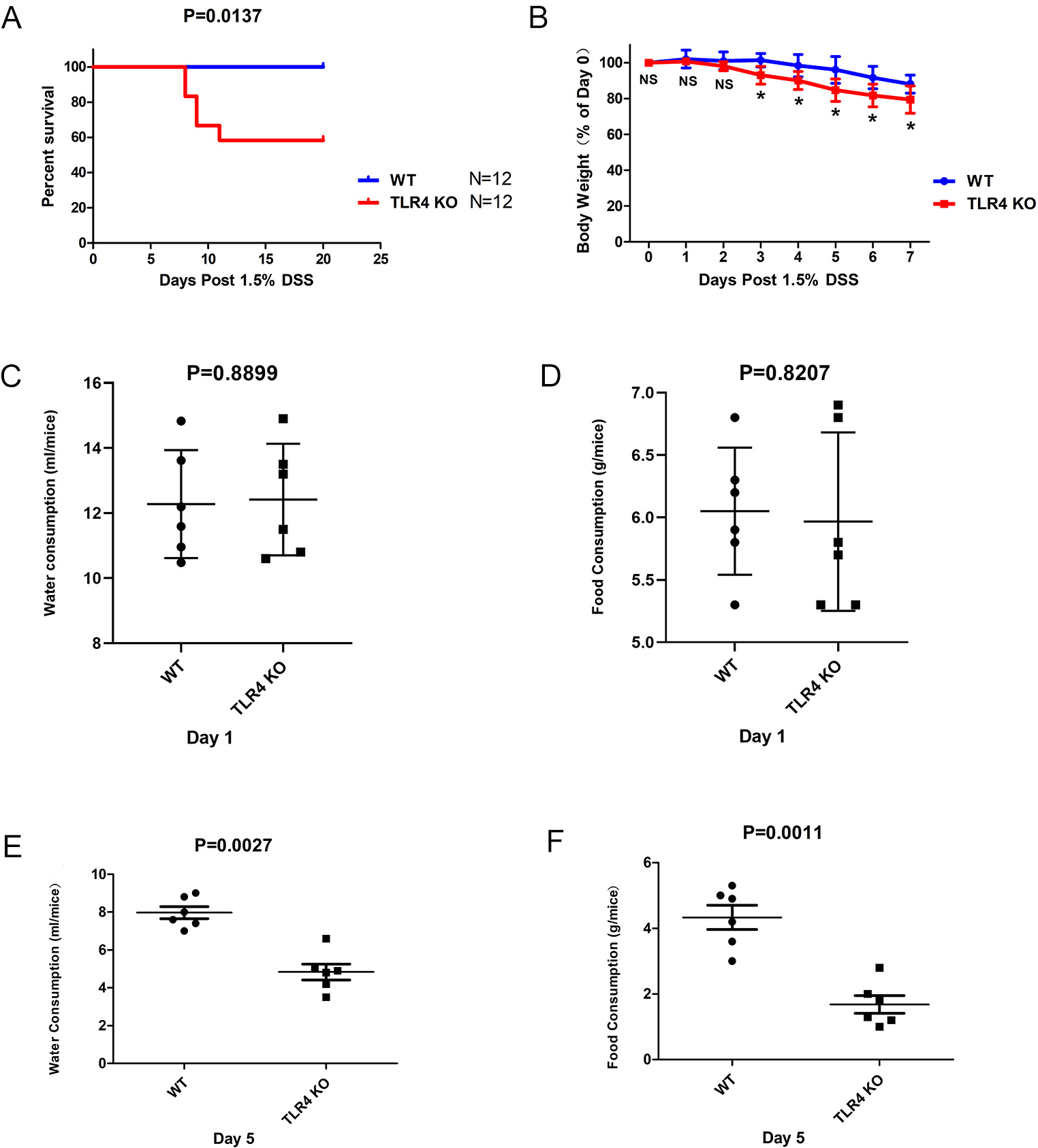 Fig. 1.
Fig. 1.Severe fatality and morbidity in TLR4-KO mice subsequent to
orally administered 1.5% DSS. (A) WT and TLR4-KO mice were orally administered
with 1.5% DSS for seven days. The survival duration was documented (p = 0.0137). (B) Body weight change (%) = (weight at day X / weight at day 0)
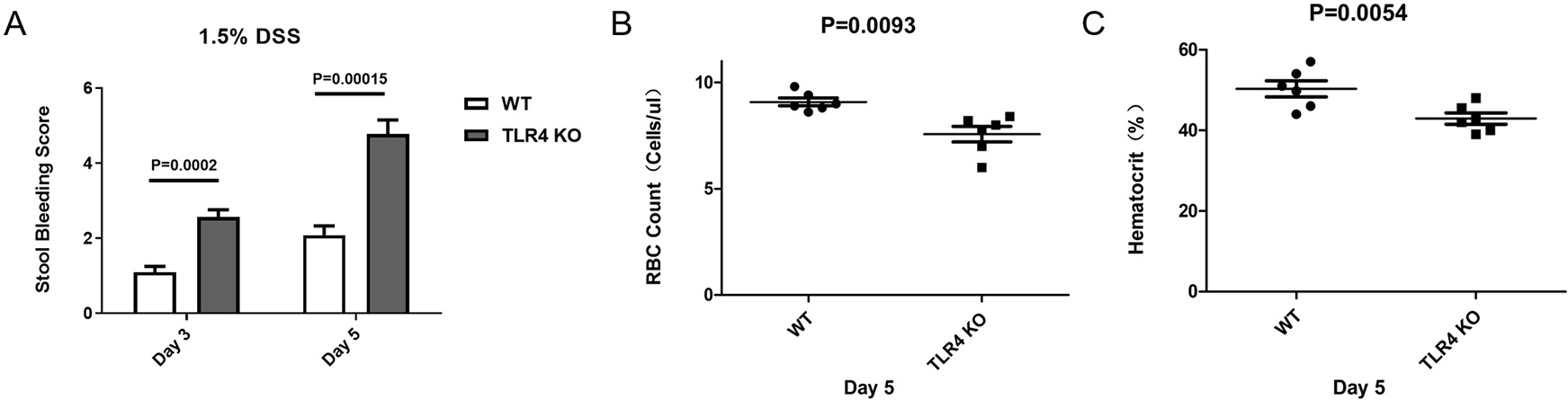
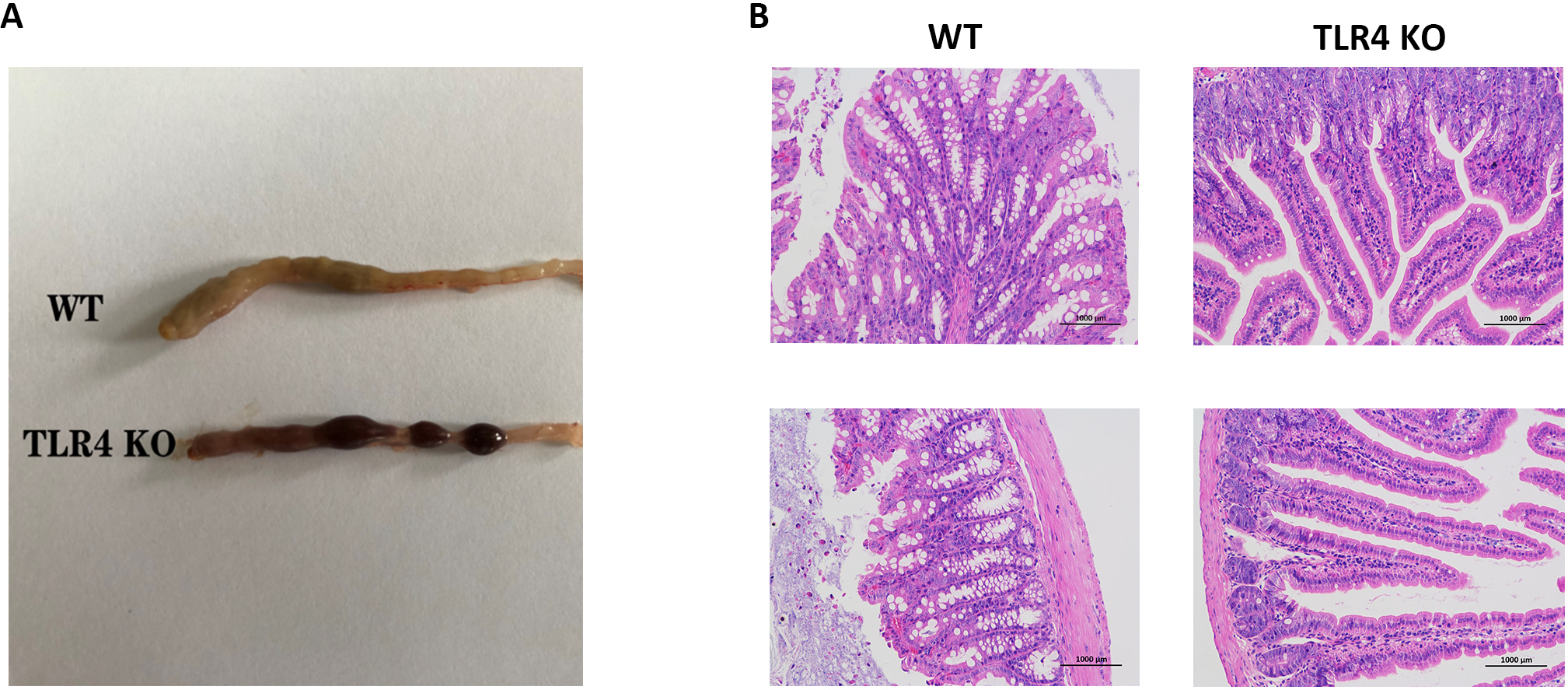
Morphological changes of the mice in the two groups were determined after oral administration of 1.5% DSS. Intestinal bleeding was detected in TLR4-KO and WT mice on days three (p = 0.0002) and five (p = 0.00015) respectively, but the severity of symptoms was increased in the TLR4-KO group in comparison to the WT group (Figs. 2A,3A). Consistent with colon bleeding, TLR4-KO mice showed more severe anemia by quantifying the peripheral red blood cell count on day five (RBC Count: p = 0.0093; Hematocrit: p = 0.0054) in both groups (Fig. 2B,C). Simultaneously, intestinal pathologic evaluation was performed on day five after oral administration of 1.5% DSS. It was found that intestinal inflammation and tissue necrosis in TLR4-KO mice were more severe than those in WT mice (Fig. 3B). All the above results demonstrated that TLR4-KO elevated the severity of intestinal epithelial injury in DSS-induced enteritis in mice.
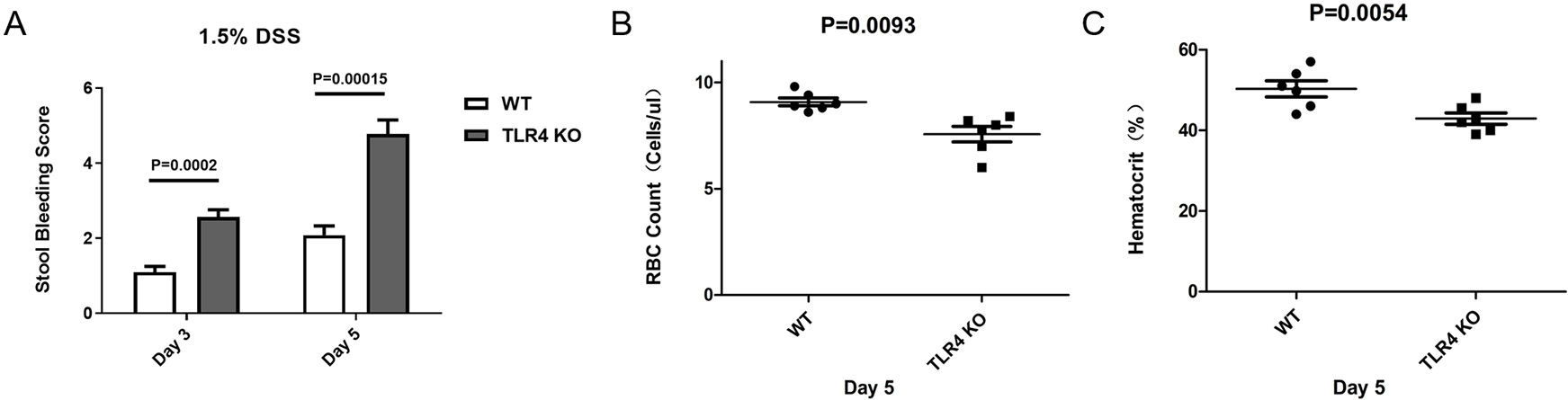 Fig. 2.
Fig. 2.Elevated susceptibility of TLR4-KO mice to contract anemia. (A)
Stool bleeding Score on days three and five. The following is the stool bleeding
score: 0 = absence of any gross blood visible in the entirety of the colon; 1 =
presence of gross blood in
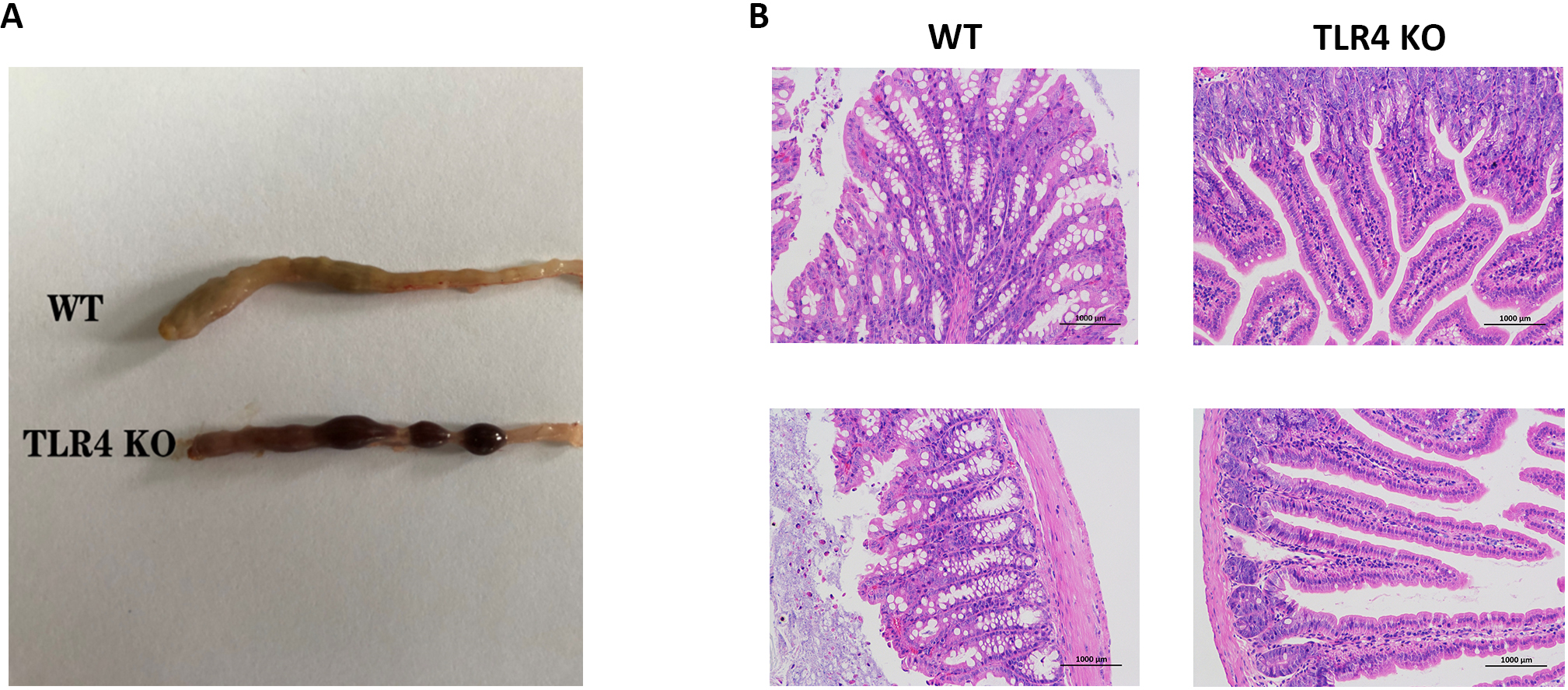 Fig. 3.
Fig. 3.Colon histopathology of the two mouse groups. (A) Photographs of bleeding colon from WT and TLR4-KO mice on day seven of orally administered 1.5% DSS. (B) H&E staining photographs of the colon in the two mice groups seven days following 1.5% DSS-infused drinking water treatment.
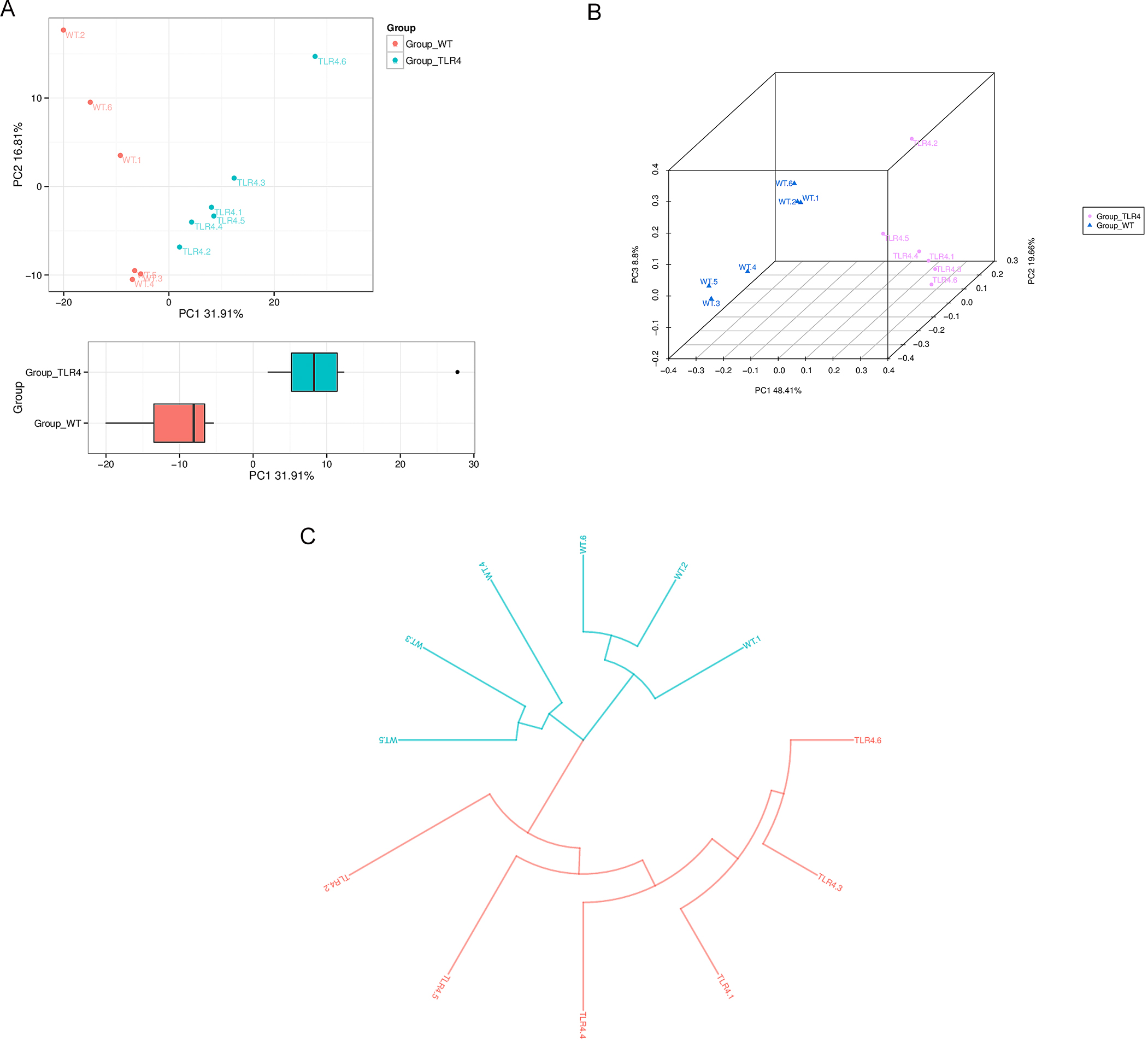
Differences in the composition of the microbiota of the gut in fecal samples
were compared through sequencing 16S rRNA gene between TLR4-KO and WT mice at day
seven after orally administered 1.5% DSS using Operational Taxonomic Unit (OTU)
[29] partitioning with 97% sequence similarity in gut microbiota. From the
flower plot, shared intestinal microbiota OTUs of all fecal samples were found to
be 42, the number on the petal indicated the total OTUs of each sample minus the
number of common OTUs (Supplementary Fig. 1). The abundance of each OTU
was analyzed and the first 50 with the most abundant OTU were selected, an
evolutionary tree was established, and the abundance of OTU in different samples
was displayed with a heatmap graph (Supplementary Fig. 2).
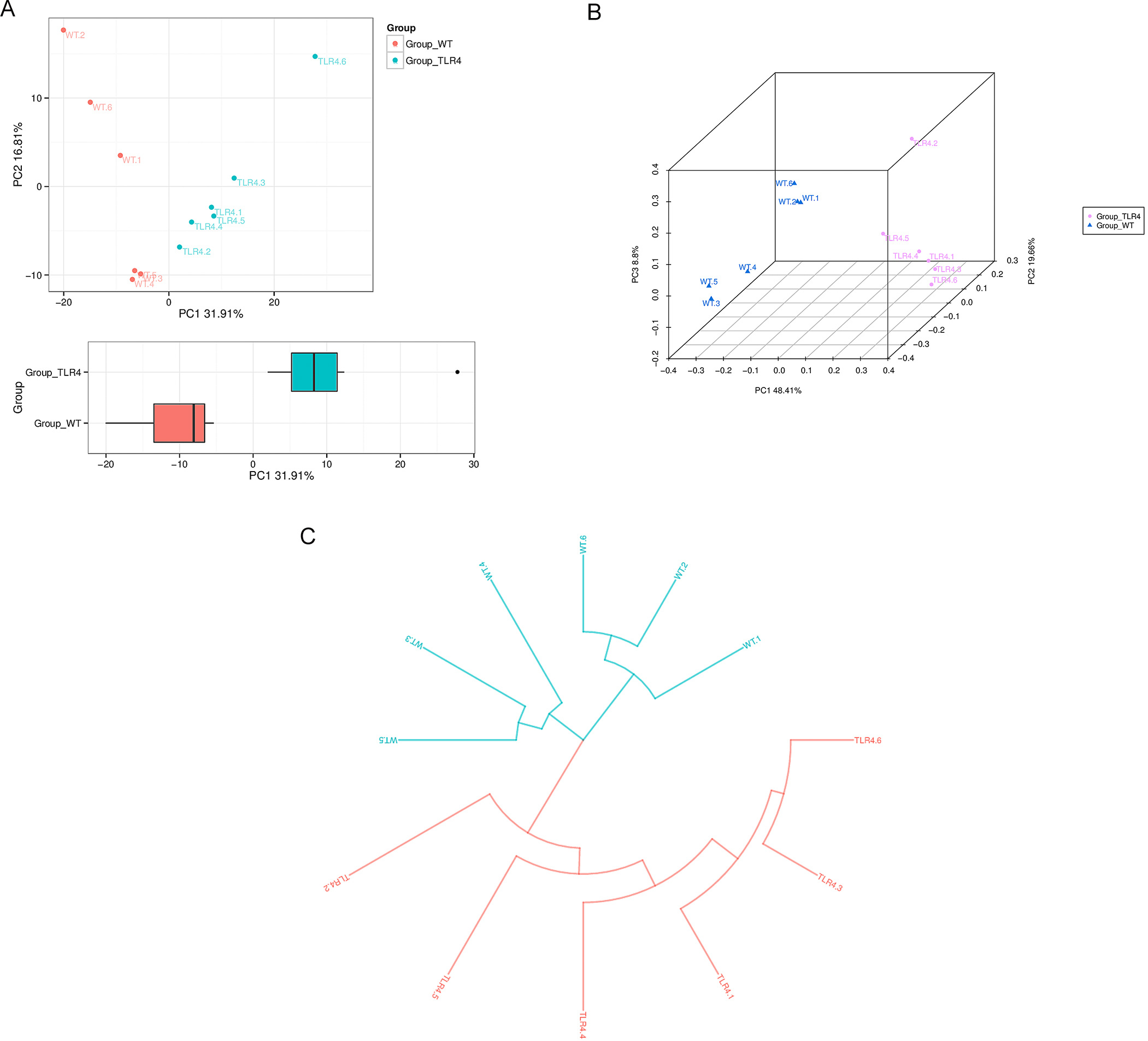 Fig. 4.
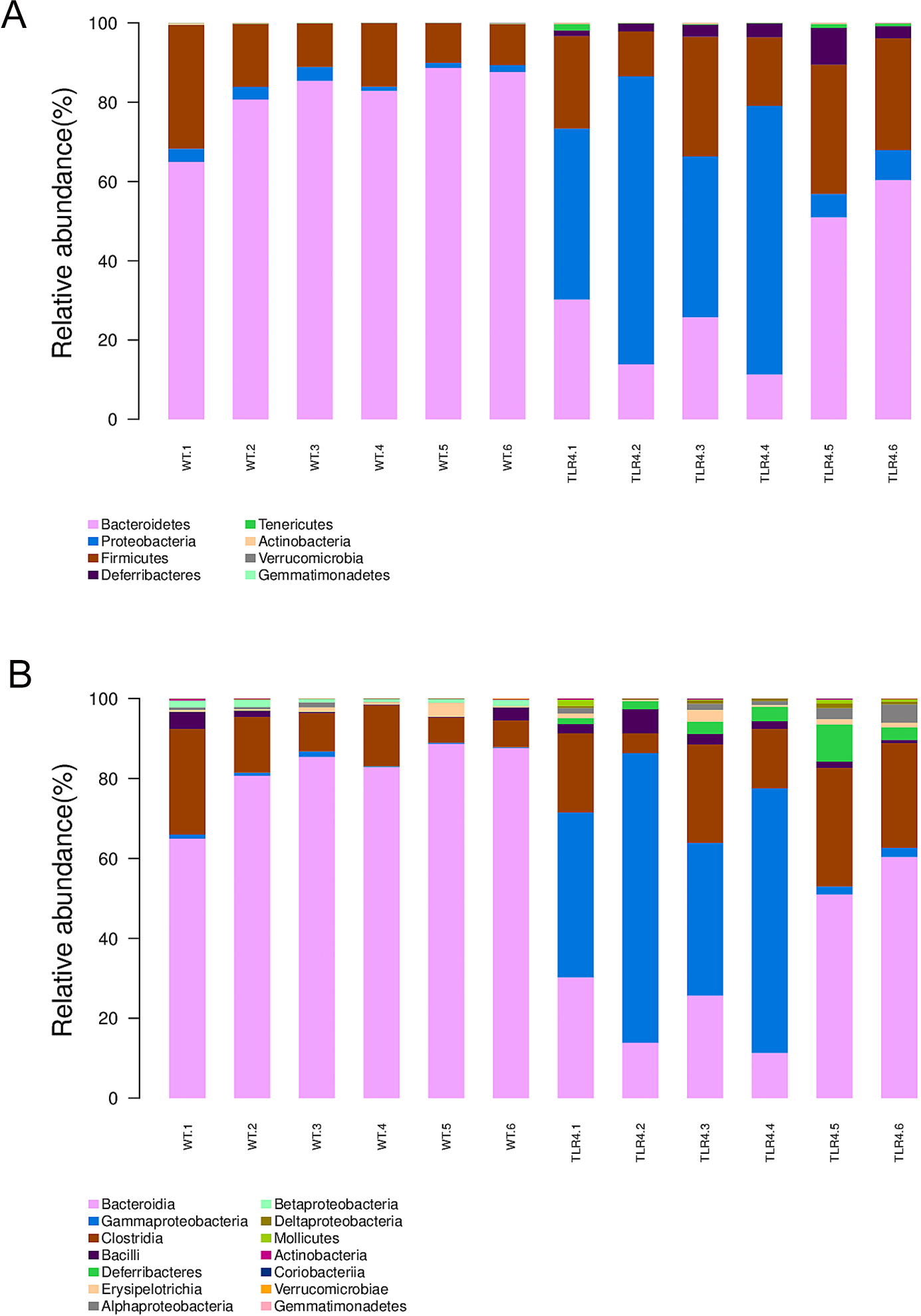
Fig. 4.According to the microbiota taxonomic levels of phylum, class, order, family, and genus, the relative abundance of each sample at various levels was recorded. The average number of reads per sample was listed in Supplementary Table 1. After a seven-day treatment with 1.5% DSS, the dominant phylum, class, order, family, and genus in WT mice were Bacteroidetes, however, they were considerably reduced in TLR4-KO mice. Simultaneously, the phylum Proteobacteria, the class Gammaproteobacteria (Fig. 5A,B), the order Enterobacteriales, the family Enterobacteriaceae, and the genus Escherichia/Shigella and Rikenellaceae were increased in TLR4-KO mice in contrast to WT controls (Supplementary Fig. 4A–C). Furthermore. ANOVA statistical analysis was carried out on OTUs and the five aforementioned phylogenetic levels, and the first ten different OUTs (species abundances) were selected for relative abundance boxplot analysis for the sake of quickly obtaining intra- and inter-group differences between the dominant and differential species. The results of ANOVA difference statistics analysis showed at the family level abundant Enterobacteriaceae, Moraxellaceae, Deferribacteraceae, Desulfovibrionaceae, Rikenellaceae, Xanthomonadaceae, and Pseudomonadaceae in gut microbiota in TLR4-KO mice, and abundant Bacteroidales, Pepococcaceae, Alcaligenaceae and Porphyromonadaceae in gut microbiota in WT mice following DSS treatment (Fig. 6A,B). The linear discriminant analysis (LDA) coupled with effect size measurements (LEfSe) analysis indicated that the gut microbiota genera were differentially numerous between WT and TLR4-KO mice after orally administered 1.5% DSS. Gladography revealed a considerably increased abundance of Bacteroidetes at the genus level in WT mice, whereas an increased abundance of Deferribacteres, Tenerlcutes, and Proteobacteria was determined in TLR4-KO mice (Fig. 7).
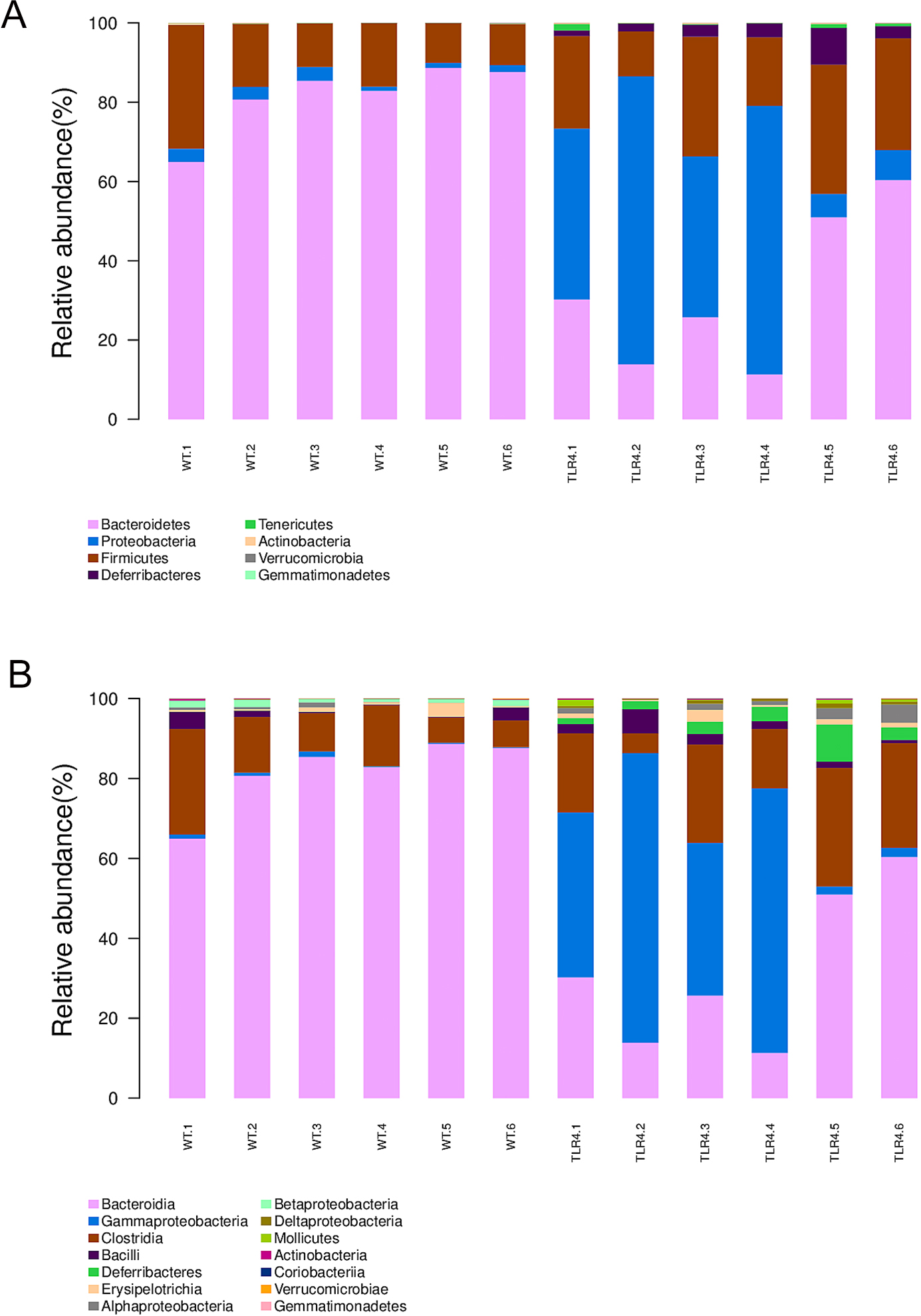 Fig. 5.
Fig. 5.Gut microbiota variation in TLR4-KO and WT mice after DSS treatment at levels of phylum and class. (A) The relative abundance of the main phylum. (B) The relative abundance of the main class.
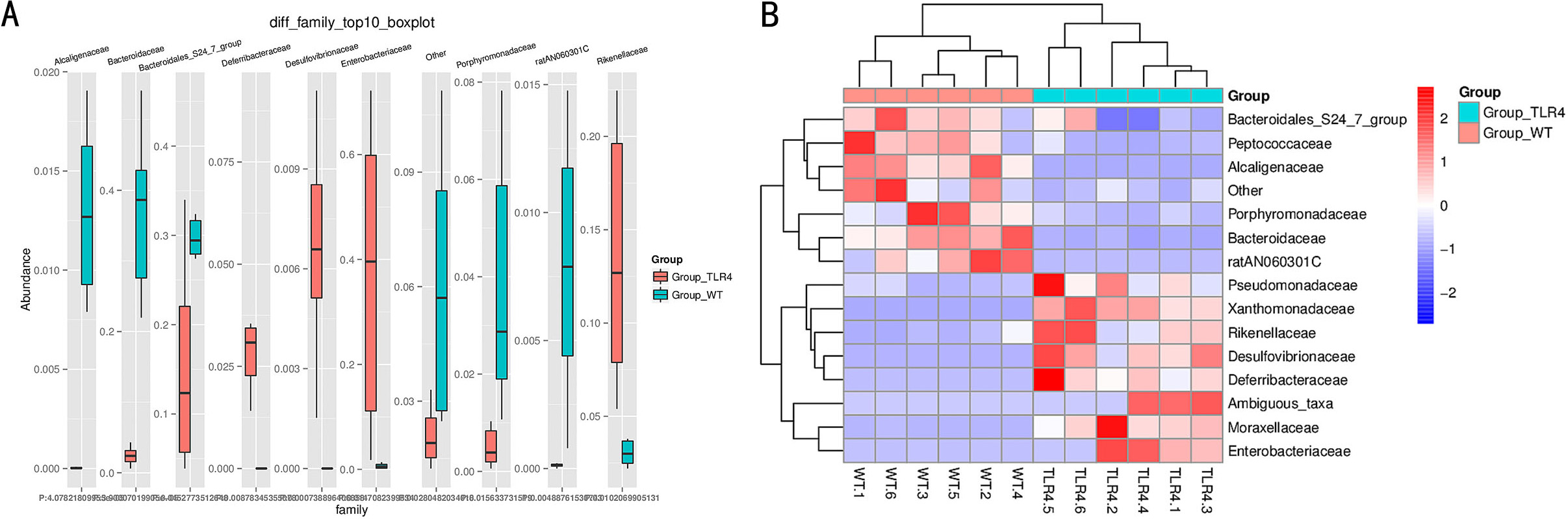
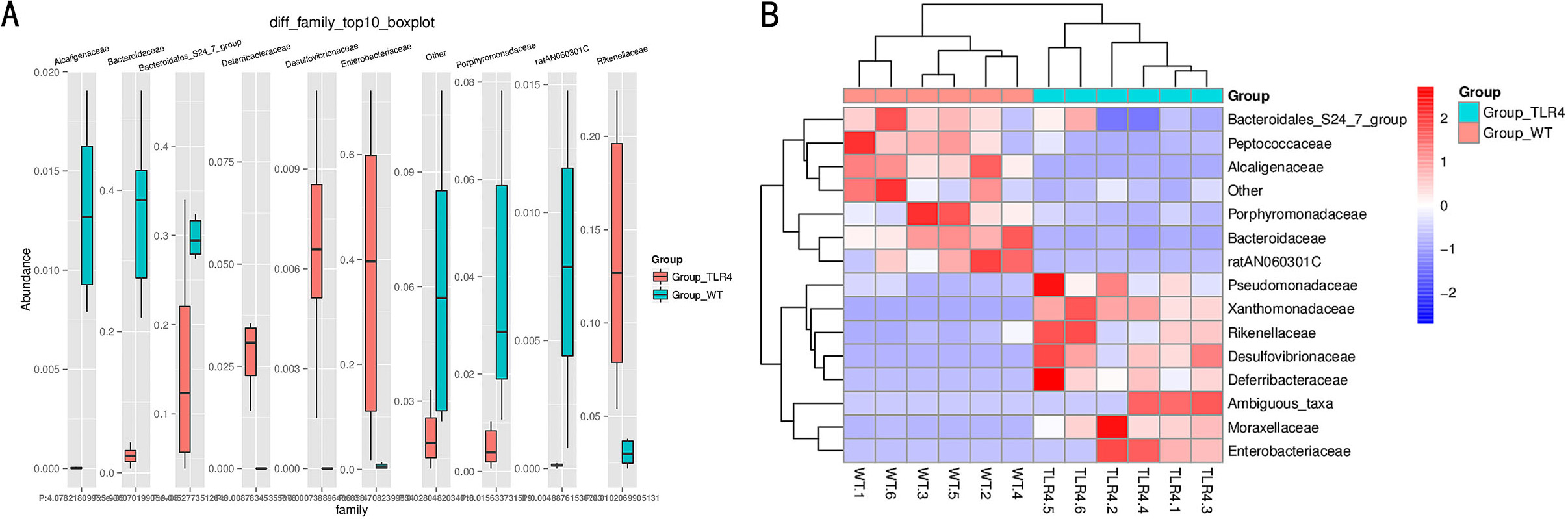 Fig. 6.
Fig. 6.Statistical analysis of microbial multivariate variables between TLR4-KO and WT mice following orally administered 1.5% DSS. (A,B) ANOVA statistical analysis of differences between TLR4-KO and WT mice following oral administration of 1.5% DSS at the family level.
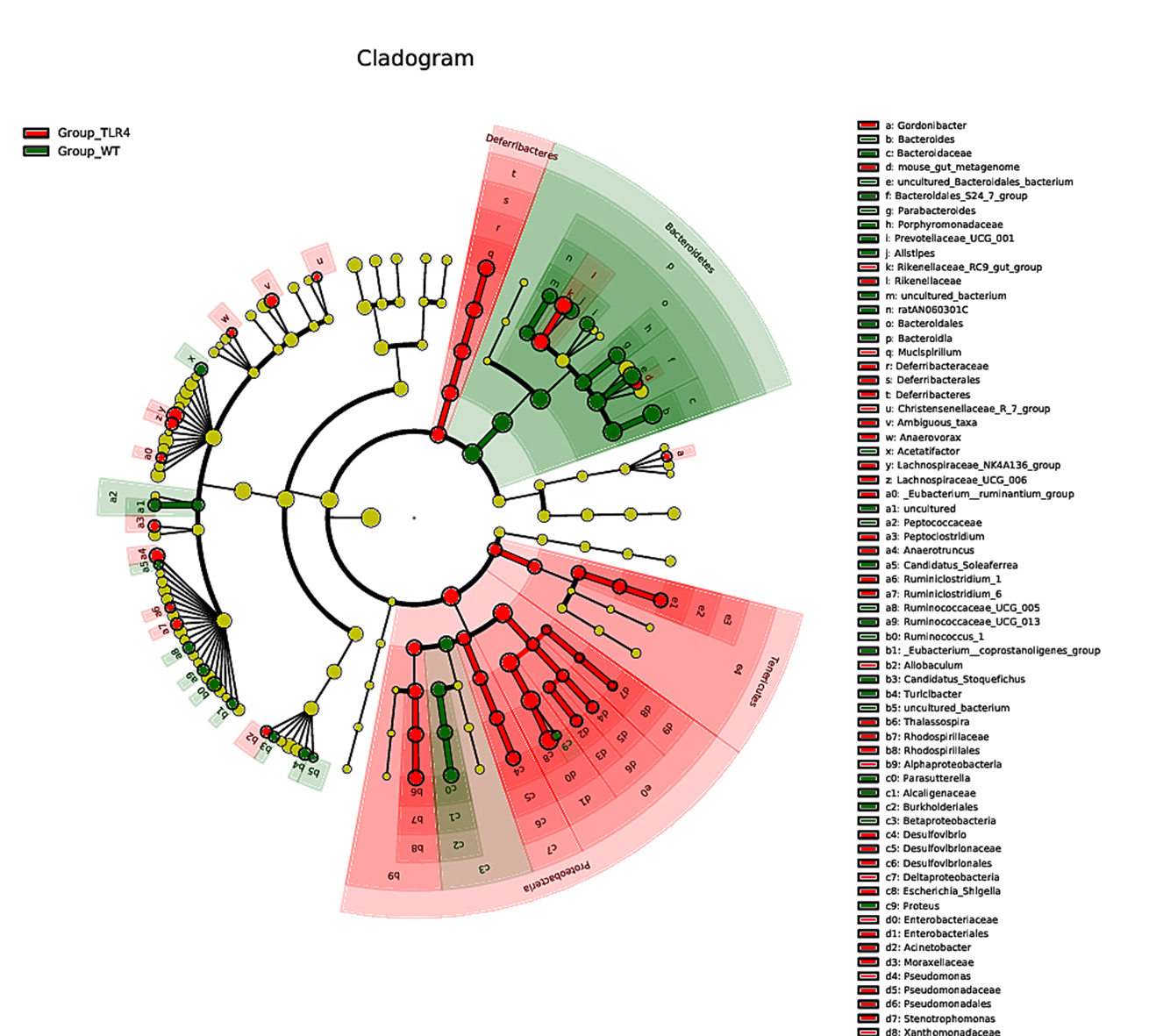
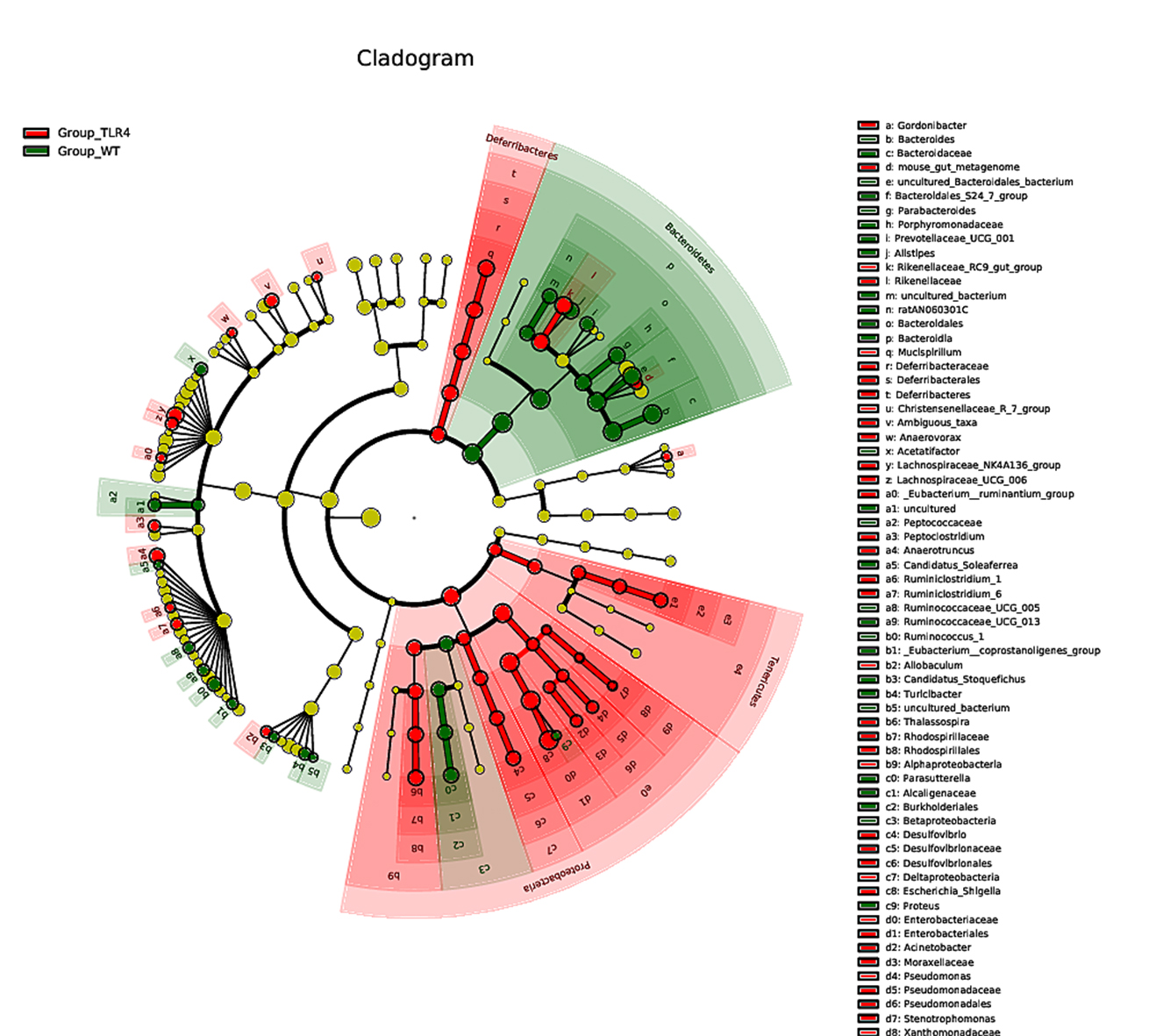 Fig. 7.
Fig. 7.Differential enrichment of multiple taxa in the TLR4-KO and WT control mice feces as per coupled with effect size measurements (LEfSe) analysis.
Due to the limitations of human differences, as well as ethical and moral factors, the pathogenesis of IBD has not been clearly defined, and there is no effective clinical treatment [32]. Therefore, it is very important to create a simple and easy animal model mimicking the clinical symptoms. The mature IBD models include spontaneous animal colitis models, cell transplantation models, and chemical-induced models [33]. In this study, the spontaneous mouse colitis model induced by DSS was employed. DSS is a synthetic heparin polysaccharide and the DSS modeling method is simple, with good repeatability and maintenance. As the signs and pathological features of the model mimic human IBD, it is currently regarded as the gold standard in the field of colitis modeling research [34]. In this experiment, the mouse colitis model was induced by drinking DSS. After modeling, the weight of the mice decreased, and the intestinal histopathology showed inflammatory changes, confirming that the DSS-induced colitis mouse model was successfully established. In this study, survival, body weight change, basic daily behavioral parameters, and intestinal bleeding between TLR4 KO and WT mouse groups were compared. The data implied that TLR4 had a protective function against DSS-induced intestinal damage. Some studies have shown that TLR4 is the primary medium of LPS response both in vitro and in vivo. It has been established that TLR4 signaling exacerbates intestinal injury in mice undergoing DSS-induced colitis [34, 35, 36]. Contrary to prior research, our previous study found that moderately activating TLR4 signaling elevated inflammation and facilitated repair of the intestinal epithelium in DSS-induced colitis, both in vitro and in vivo [37]. Moderate activation of TLR4 signaling upregulated the levels of expression of repair factors PGE2 and GM-CSF [38, 39] in the later stage of DSS-induced colitis. Meanwhile, Cario E et al. [40] reported that intestinal stem cells that could be found at the intestinal crypt base could promote the repopulation of the depleted crypt after intestinal epithelial injury, and this process is termed compensatory proliferation. This process may be involved in the moderate activation of the TLR4 signaling pathway.
Gut microbiota diversity is strongly linked to human health. The number of
microbiota in the gut of healthy adults far exceeds the number of cells in the
human body, which is a necessary condition for the normal functioning of the
human gut. Research has increasingly demonstrated that gut microbiota dysfunction
is the main cause of IBD [11]. Previous 16S rRNA sequencing analyses on the gut
microbiota composition in IBD patients [41, 42, 43, 44] demonstrated that
These results demonstrated severe susceptibility of TLR4-KO mice to DSS-induced colitis. This is consistent with the results of the research we have published. The data depicted the repairing effect of TLR4 on DSS-induced intestinal damage and that it can up-regulate CSF3, IL6, and CCL2 [7]. On this basis, this research depicted remarkable variation in the number of gut microbiota between TLR4-KO and WT mouse groups treated with 1.5% DSS via 16S rRNA gene sequencing. There were abundant Enterobacter, and Escherichia coli/Shigella in the gut microbiota of TLR4-KO mice, whereas the abundance of Bacteroidetes was considerably reduced at five phylogenetic levels as compared with WT controls. Collectively, moderate activation of TLR4 may perform some function in repairing the intestinal epithelium by up-regulating Enterobacter, Escherichia coli/Shigella, and down-regulating Bacteroidetes. The limitation of this study was that the gut microbiota screened by 16S rRNA sequencing was not been verified and the microbiota profiles of WT and TLR4-KO mice before 1.5% DSS administration was not be compared. Future research is planned to verify the various core gut microbiota that has been screened out between TLR4-KO and WT mice and establish a network map of intestinal microbiota with TLR4 as the core to repair IBD-related intestinal mucosal damage. The analysis of the relationship between downstream genes and key gut microbiota of TLR4 signaling in the repair of intestinal mucosal damage caused by DSS is also a key part of it.
All data generated or analyzed during this study are included in this published article. The experimental data analyzed and displayed in the present manuscript are available from the corresponding author upon reasonable request.
(I) Conception and design—HW, CL, ZQH; (II) Administrative support—HW; (III) Provision of study materials—ZQH; (IV) Collection and assembly of data—YJS, CH; (V) Data analysis and interpretation—YJS, PQG; (VI) Manuscript writing—All authors; (VII) Final approval of manuscript—All authors.
All animal experiments conformed to the National Institute of Health Guide for the Care and Use of Laboratory Animals’ (NIH Publication No. 85-23, National Academy Press, Washington, DC, revised 1996), with the approval of the Laboratory Animal Center of the Naval Medical University, Shanghai (No. 96-01).
Not applicable.
This work was supported by the National Key Basic Research Development Program of China (Grant No. 2015CB554000); the National Natural Science Foundation of China (Grant No. 82173005, 81573092, 81872046 & 81872559); and Shanghai Municipal Science and Technology Commission Research Program (Subject No. 17411951100).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.