1 Department of Cardiology, The First Affiliated Hospital of Nanjing Medical University, 210029 Nanjing, Jiangsu, China
†These authors contributed equally.
Academic Editor: Rajesh Katare
Abstract
Background: Dilated cardiomyopathy (DCM) is one of the main causes of systolic heart failure and frequently has a genetic component. The molecular mechanisms underlying the onset and progression of DCM remain unclear. This study aimed to identify novel diagnostic biomarkers to aid in the treatment and diagnosis of DCM. Method: The Gene Expression Omnibus (GEO) database was explored to extract two microarray datasets, GSE120895 and GSE17800, which were subsequently merged into a single cohort. Differentially expressed genes were analyzed in the DCM and control groups, followed by weighted gene coexpression network analysis to determine the core modules. Core nodes were identified by gene significance (GS) and module membership (MM) values, and four hub genes were predicted by the Lasso regression model. The expression levels and diagnostic values of the four hub genes were further validated in the datasets GSE19303. Finally, potential therapeutic drugs and upstream molecules regulating genes were identified. Results: The turquoise module is the core module of DCM. Four hub genes were identified: GYPC (glycophorin C), MLF2 (myeloid leukemia factor 2), COPS7A (COP9 signalosome subunit 7A) and ARL2 (ADP ribosylation factor like GTPase 2). Subsequently, Hub genes showed significant differences in expression in both the dataset and the validation model by real-time quantitative PCR (qPCR). Four potential modulators and seven chemicals were also identified. Finally, molecular docking simulations of the gene-encoded proteins with small-molecule drugs were successfully performed. Conclusions: The results suggested that ARL2, MLF2, GYPC and COPS7A could be potential gene biomarkers for DCM.
Keywords
- weighted gene coexpression network analysis
- dilated cardiomyopathy
Dilated cardiomyopathy (DCM) is a clinical phenotype that manifests as heart failure due to a combination of genomic [1, 2], epigenetic [3] and external factors. The prevalence of DCM varies from 1/250 to 1/2500 [4], occurring in greater proportions than ischemic cardiomyopathy [5], and is the leading cause of heart failure. Many new drugs and devices are being used to improve the long-term prognosis of patients, such as angiotensin receptor neprilysin inhibitor (ARNI) [6], sodium-glucose cotransporter 2 (SLGT2) inhibitors [7], and left heart assist devices. However, clinical decision-making [8] in DCM is mainly based on heart failure without taking into account the heterogeneity [9] of DCM.
Previous studies have shown that mutations in genes encoding cytoskeletal, myosin, mitochondrial, bridging granule, nuclear membrane, and RNA-binding proteins are associated with DCM [5]. However, there is considerable heterogeneity in the genetic testing panel, especially with the development of whole-genome sequencing, which has seen many variations, making it difficult to distinguish pathogenic and nonpathogenic variants [10]. Abnormal genetic variants defined by individual studies may be normal in other populations due to sample size, ethnicity, and other factors. Currently, genetic diagnosis is used to rule out genes carried by other family members of the proband, but some carriers pass the genes on without any cardiac events. The fundamental mechanisms of DCM remain poorly understood. Finding efficient and low-cost diagnostic methods to identify hub genes has been a major challenge [11].
Thanks to the development of high-throughput technologies, transcriptional analysis based on multiple datasets has been used to determine the pathological mechanisms of diseases. Disease pathogenesis and progression are not caused by a single gene but by synergistic effects in a complex network [12]. Complementary to this is weighted gene coexpression network analysis (WGCNA), a widely used systems bioinformatics technique to assess associations between genomic and external sample features by constructing scale-free gene coexpression networks.
Molecular targeted therapies have been widely used in oncological diseases to assist physicians’ decisions based on the expression of star molecules, but have not yet been introduced to the clinic for use in cardiovascular diseases. This paper is dedicated to investigate the possibility of molecular targets for the diagnosis and treatment of clinical diseases, to find a new marker that can be used for general screening of DCM or to improve the prognosis of patients after intervention. Therefore, this study collected DCM-related genes and used multiple databases to further search for hub genes and targets for potential accurate treatment or diagnosis of DCM.
The raw data of two eligible microarray datasets (GSE120895 and GSE17800) based
on platform GPL570 were downloaded from the GEO database (87
patients with DCM and 16 controls after data merging). Moreover, GSE19303 (73
patients with DCM and 8 controls) was used as the validation set. The Limma
package was used to screen DEGs between DCM and controls, followed by data
normalization [13]. Bias and variability of the datasets were removed using the
Combat function in the sva package. p
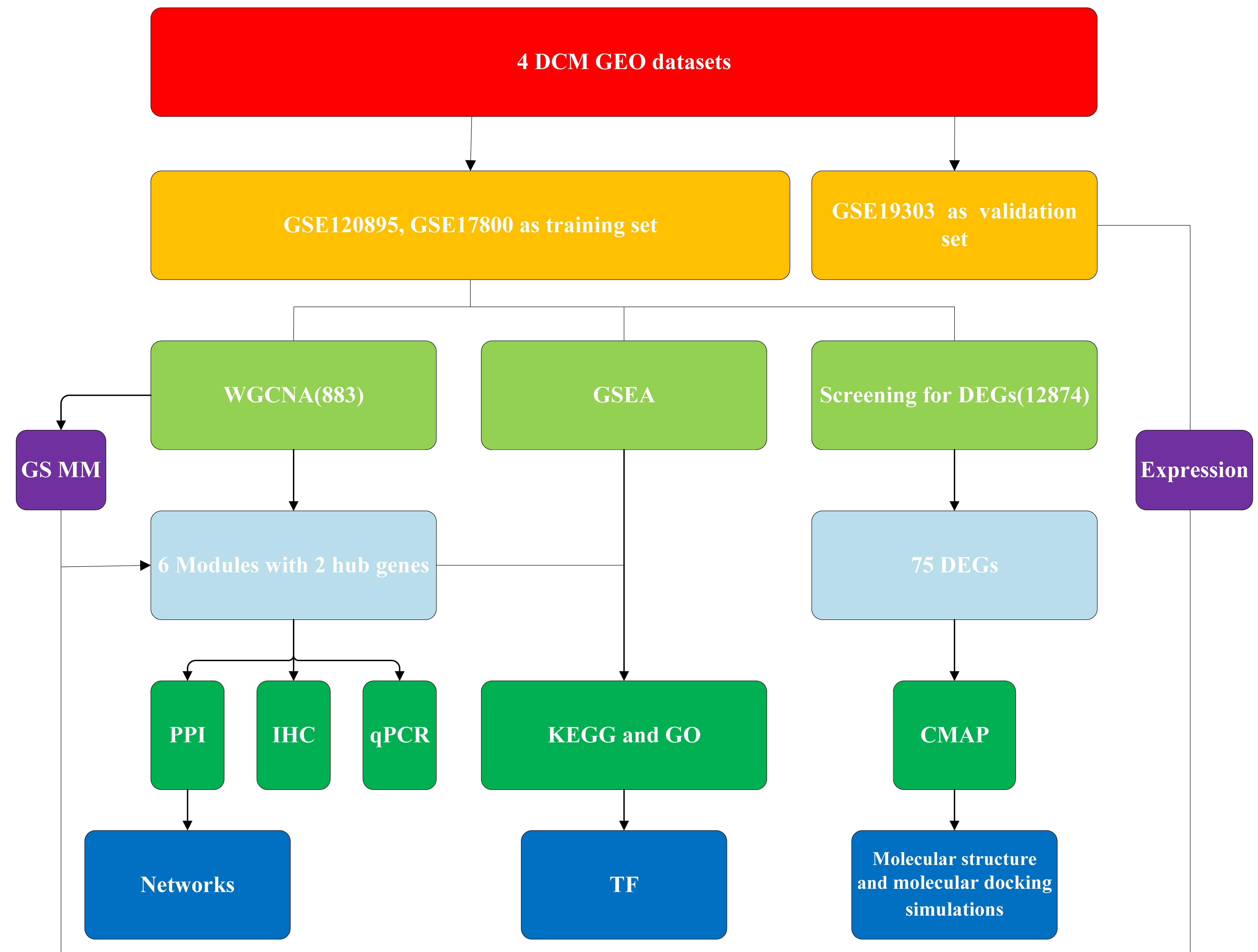
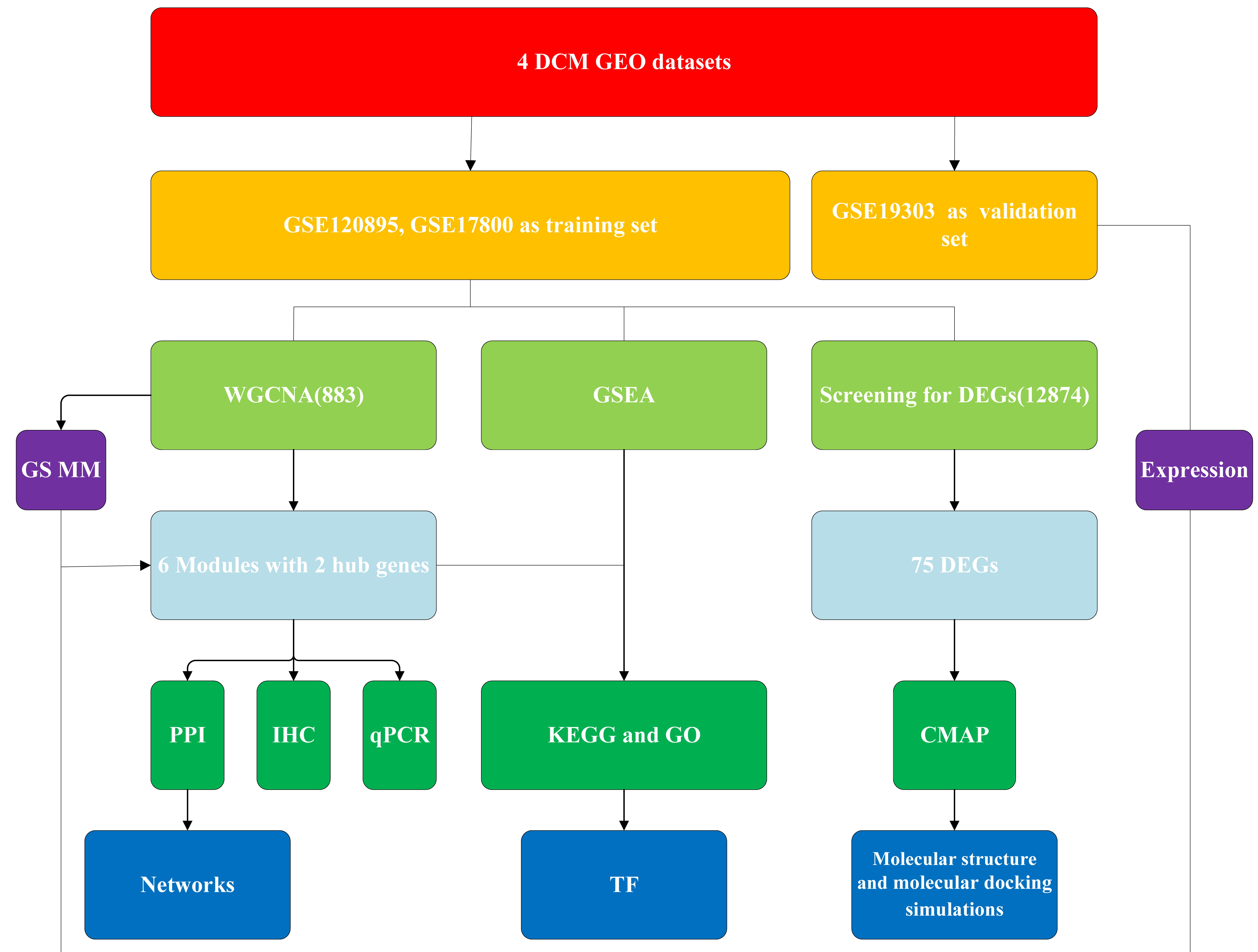 Fig. 1.
Fig. 1.The workflow of our research. DCM, dilated cardiomyopathy; GEO, Gene Expression Omnibus; GO, Gene Ontology; PPI, protein-protein interaction; CMAP, Connectivity Map; KEGG, Kyoto Encyclopedia of Genes and Genomes; GSEA, gene set enrichment analysis; TF, transcription factor; DEG, differentially expressed gene; WGCNA, weighted gene coexpression network analysis; IHC, immunohistochemical; qPCR, quantitative polymerase chain reaction.
A coexpression network of 884 genes with commonly upregulated or downregulated
expression in DCM/control was constructed using WGCNA, which is a widely used
systems biology approach that helps identify the relationship between genes and
disease phenotypes. The main processes were as follows: (1) the hclust function
was used for hierarchical clustering analysis; (2) a power of
In WGCNA, module membership (MM) is defined as the correlation of the module
eigengene and the gene expression profile, while gene significance (GS) is
defined as the correlation between the gene and the trait. Among the modules of
interest in our study, GS
The least absolute contraction and selection operator (LASSO) method [14], which is suitable for high-dimensional data restoration, was used to select the optimal risk factor prediction characteristics from gene datasets. The A radiomics score (Rad-score) of each gene was calculated by selecting a linear combination of features, which are weighted by their respective coefficients. Then, the variables selected by the LASSO method were used to obtain the final factors for establishing the model. Harrell’s C index and the area under the curve (AUC) were measured to quantify the discrimination performance of the model based on the experimental set and validation set.
GSEA is a computational method used to assess whether a predefined set of genes
displays statistical significance and consistency differences between two
biological states [15]. The c5.all.v7.4.entrez and c2.cp.kegg.v7.4.entrez
datasets in the MsigDB database were used as reference gene sets, and the
clusterprofile package was utilized to perform GSEA with integrated gene
expression data. p
The promoter sequences of target genes were searched using the University of California Santa Cruz (UCSC) and National Center for Biotechnology Information databases (NCBI). Potential binding transcription factors were searched using the JASPAR database and further screened according to transcription direction and related levels. Gene expression profiling interactive analysis (GEPIA) was used to verify the correlations and identify potential transcription factors. Finally, the structural information combined with the interval key structural domain was presented using UniProt.
The CMap database establishes links between genes, compounds and diseases based
on similar and opposite gene expression profiles. In this study, the DEGs of the
DCM and control groups were grouped according to expression differences. Then,
the DEGs were loaded to the “Query” page. In this study, drugs with
connectivity scores
The study was approved by the Medical Ethics Committee of
Nanjing Medical University, and a total of 16 patients were recruited to provide
blood samples, and informed consent was obtained from patients before
participation. Expression levels of 4 hub genes were verified in 8 DCM blood
samples and 8 normal blood samples (Supplementary Table 1). Total RNA
was extracted using TRIzol reagent according to the manufacturer’s instructions.
Total RNA was reverse transcribed into cDNA by PrimeScript RT Master Mix (TaKaRa,
Japan) after measuring the corresponding concentration. Then qRT-PCR was
performed using Power SYBR Green PCR Master Mix (No. A25742; Thermo Fisher
Scientific, Waltham, MA, USA). Finally, the relative expression levels of miRNAs
and target genes were calculated according to the 2 -
The Human Protein Atlas was used to validate the immunohistochemistry of potential target genes. This database facilitates the systematic study of transcriptome and gene pathology expression of coding genes in different tissue types. Staining of core gene proteins in human myocardial tissue based on immunohistochemical techniques. In addition, tissue types and staining levels were retrieved from the database to analyze the quality of the data to interpret the results [16, 17].
We retrieved the 3D structure of the receptor from the Uniprot and RCSB protein databases, and the corresponding simplified molecular-input line-entry system (SMILES) for the receptor ligand was obtained from the PubChem database. After the ligand energy was adjusted to a minimum by using Chem 3D software 18.0 (PerkinElmer, Waltham, MA, USA), the crystals were imported into Pymol 2.4.0 software (Schrödinger, L. & DeLano, W., CA, USA) for dehydration, hydrogenation and ligand separation, followed by AutoDockTools-1.5.7 [18] to construct docking grid boxes for each target. Docking is done with Autodock Vina 1.1.2 (The Scripps Research Institute, San Diego, CA, USA) [19].
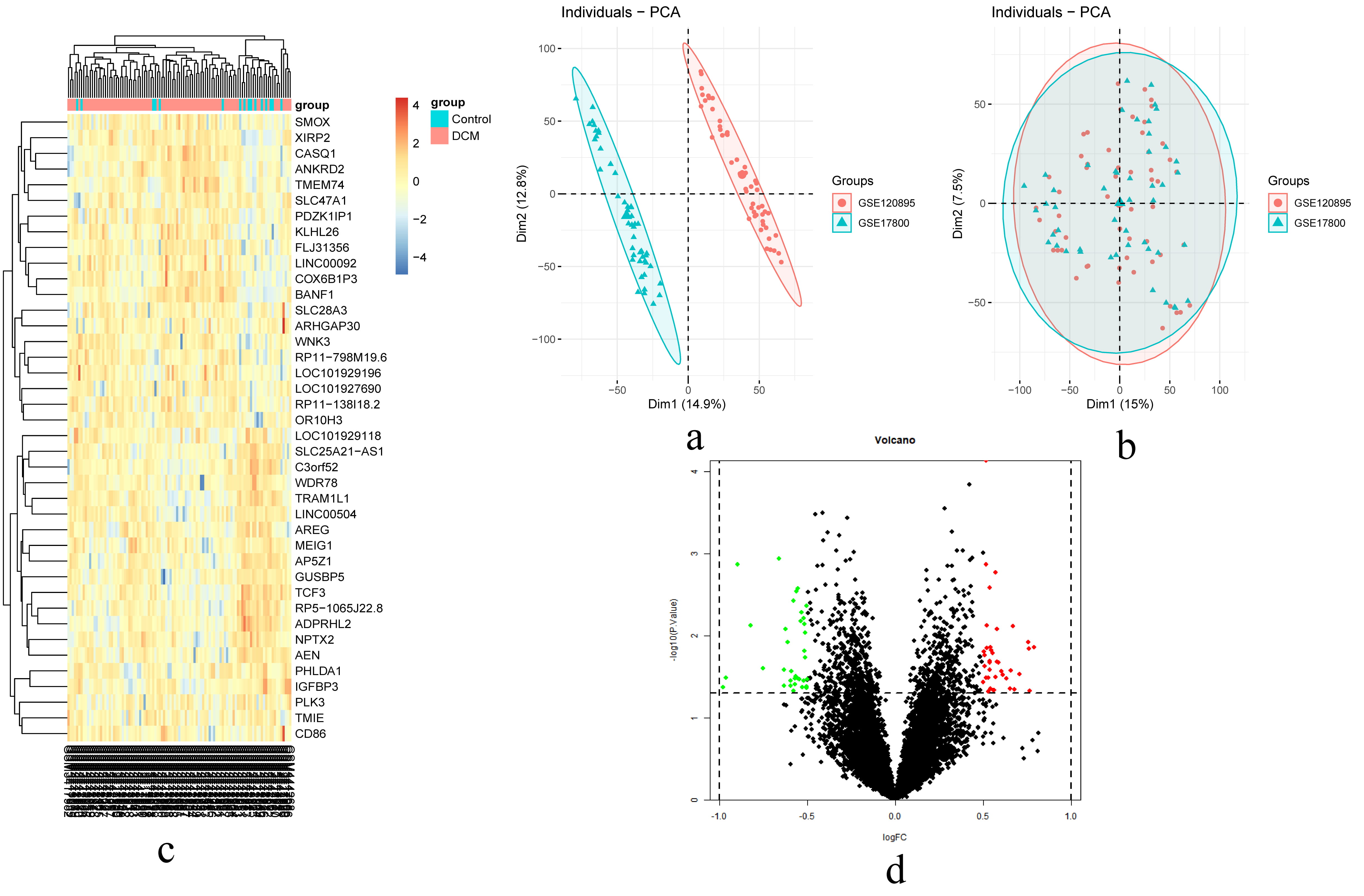
To identify DEGs associated with DCM, we screened the integrated normalized data and obtained 12,748 genes after adjusting for batch effects. As shown in the volcano map (Fig. 2), compared to controls, there were 75 DEGs in the DCM sample, of which 39 were upregulated and 36 were downregulated. The 20 most upregulated and 20 downregulated genes in DCM are visualized in the heatmap.
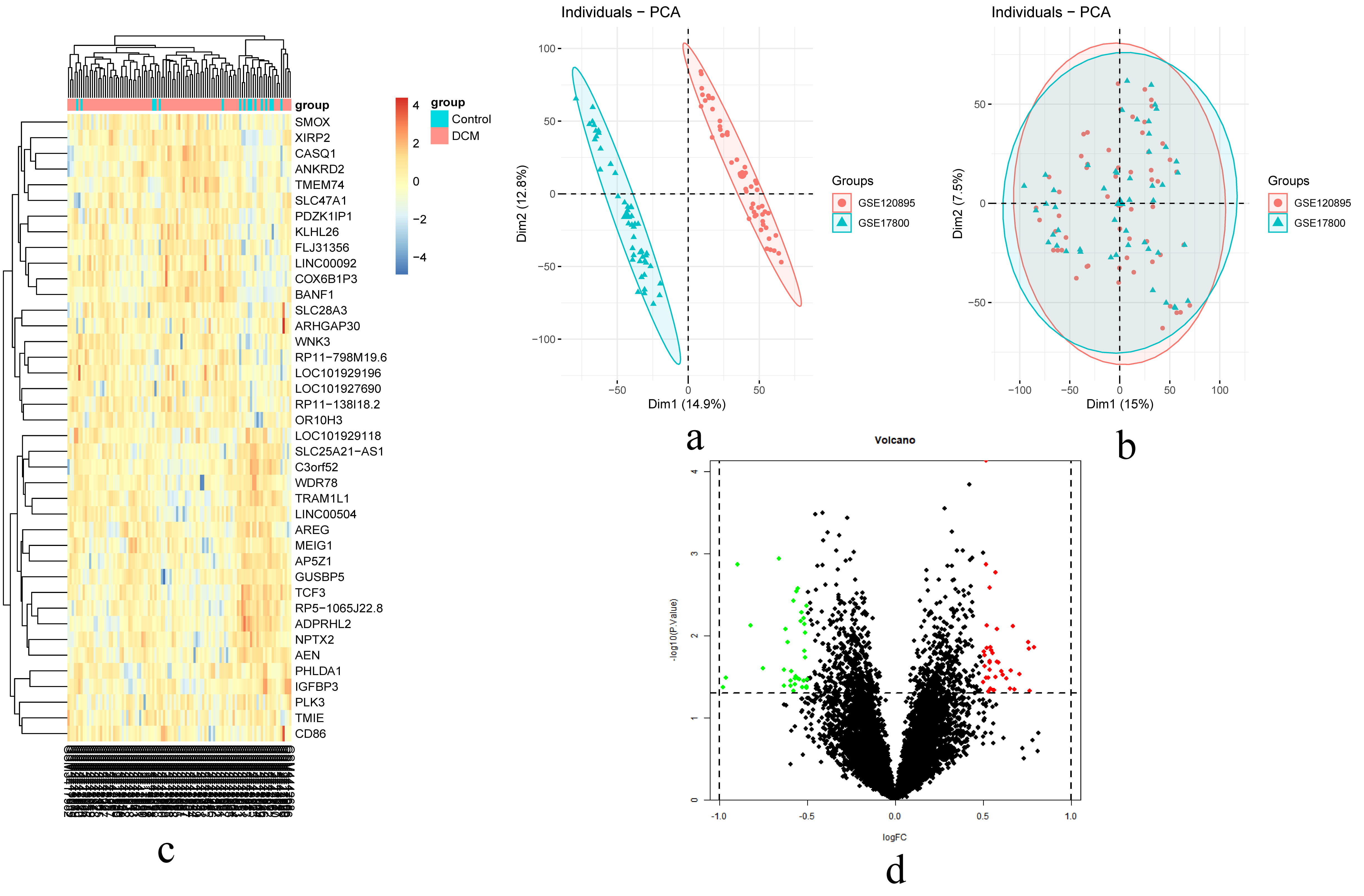 Fig. 2.
Fig. 2.The result of DEGs identification. (a) Principal component analysis (PCA) before batch effects removement. (b) PCA after batch effects removement. (c) A heatmap of 20 most up-regulated and 20 most down-regulated genes. (d) Volcano plot of the DCM-Control. Abbreviations: DCM, dilated cardiomyopathy; logFC, log2 fold-change.
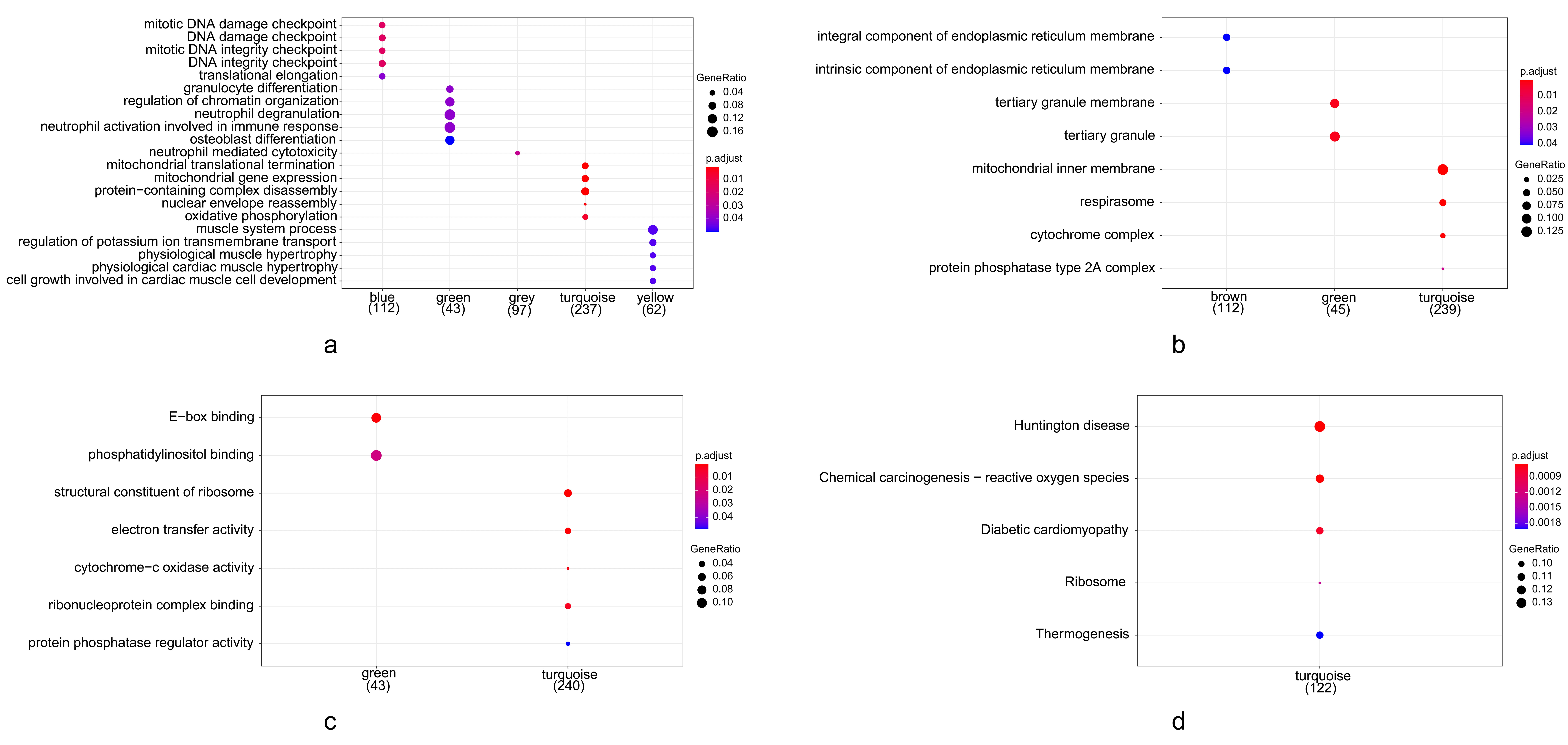
It is reasonable to consider that certain genes with similar expression patterns may perform their functions as a global network and participate in similar biological processes. The 883 DEGs with absolute deviation were used for WGCNA. A total of six gene modules were obtained, which are represented by branches of the clustering tree, and different colors are shown in Fig. 3. In addition, modular enrichment analysis showed that primary enrichment in biological process (BP) corresponding to each module included mitotic DNA damage checkpoint, neutrophil degranulation, protein-containing complex disassembly and muscle system process. Primary enrichments in cell component (CC) involved integral components of endoplasmic reticulum membrane, tertiary granule and mitochondrial inner membrane. Primary enrichments in molecule function (MF) consisted of E-box binding and structural constituent of ribosome. Primary enrichments in the KEGG pathway were chemical carcinogenesis - reactive oxygen species and diabetic cardiomyopathy (Fig. 4). The PPI network of DEGs was constructed with STRING and visualized with Cytoscape (3.8.2). As shown in the Supplementary Fig. 1, the PPI network contained nodes and edges in a circular distribution. The nine most important modules were identified by the Cytoscape plugin MCODE.
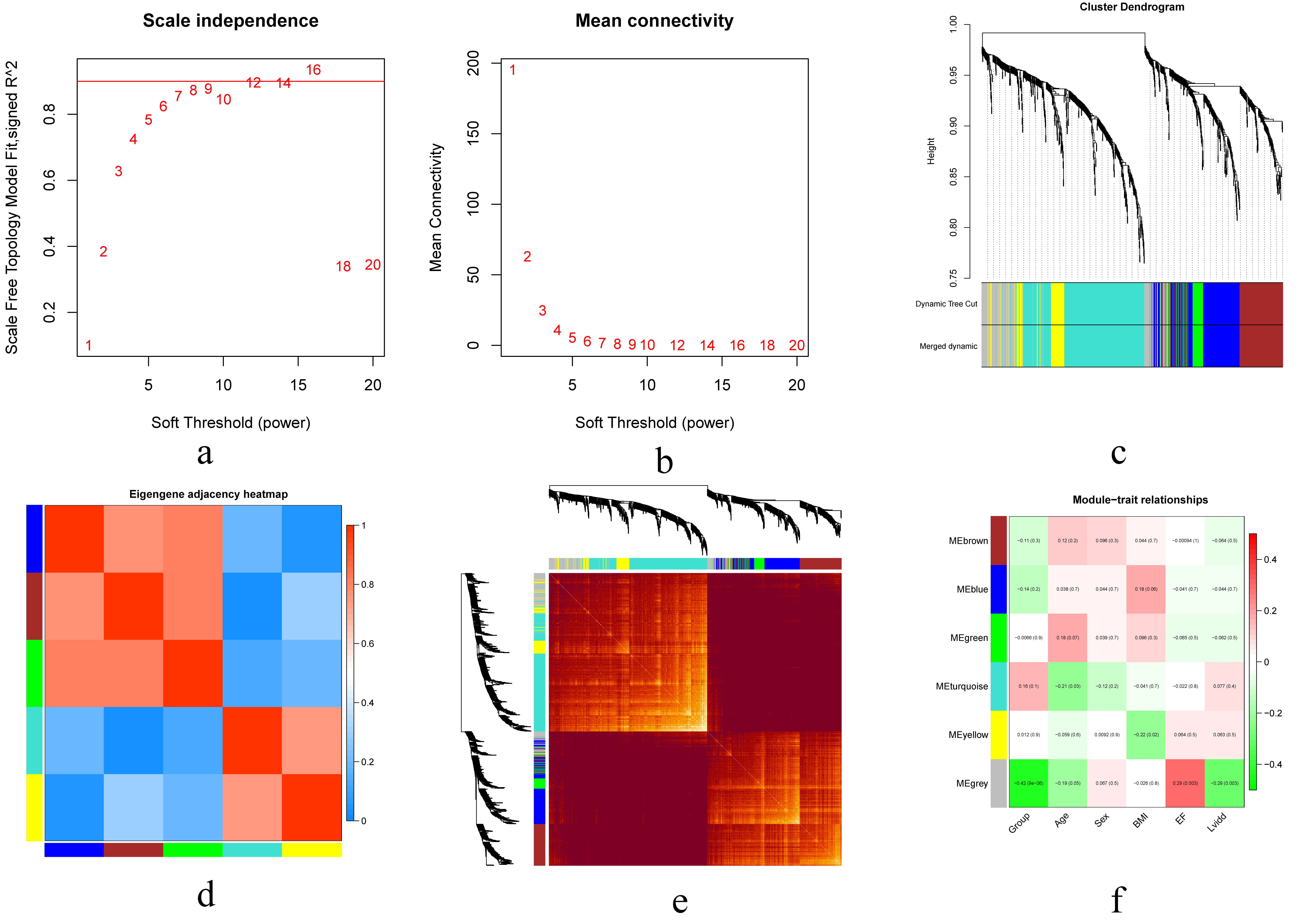
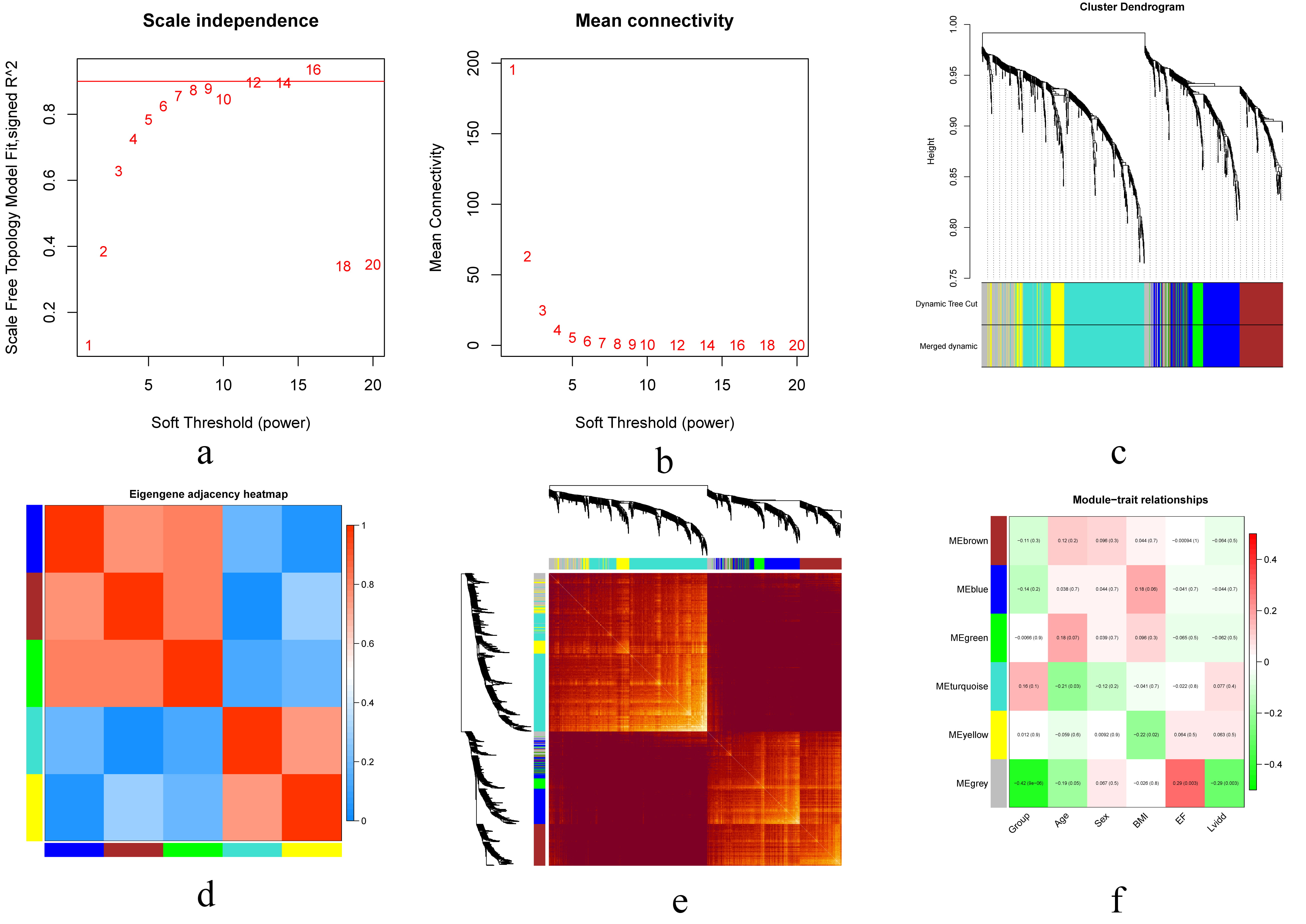 Fig. 3.
Fig. 3.Construction of gene co-expression modules. (a,b) Analysis of network topology for various soft-thresholding powers. (c) Hierarchical cluster dendrogram of DCM-relate genes based on one dissimilarity measure. (d) Hierarchical cluster heatmap of the adjacencies in the eigengene network. (e) Topology overlay heat map: Both rows and columns indicate individual genes, and dark yellow and red indicate a high degree of topological overlap. (f) Module-phenotype associations. Each row corresponds to a module eigengene, and each column corresponds to a clinical feature. Abbreviations: DCM, dilated cardiomyopathy; BMI, body mass index; EF, ejection fraction; Lvidd, left ventricular internal diameter at end-diastole.
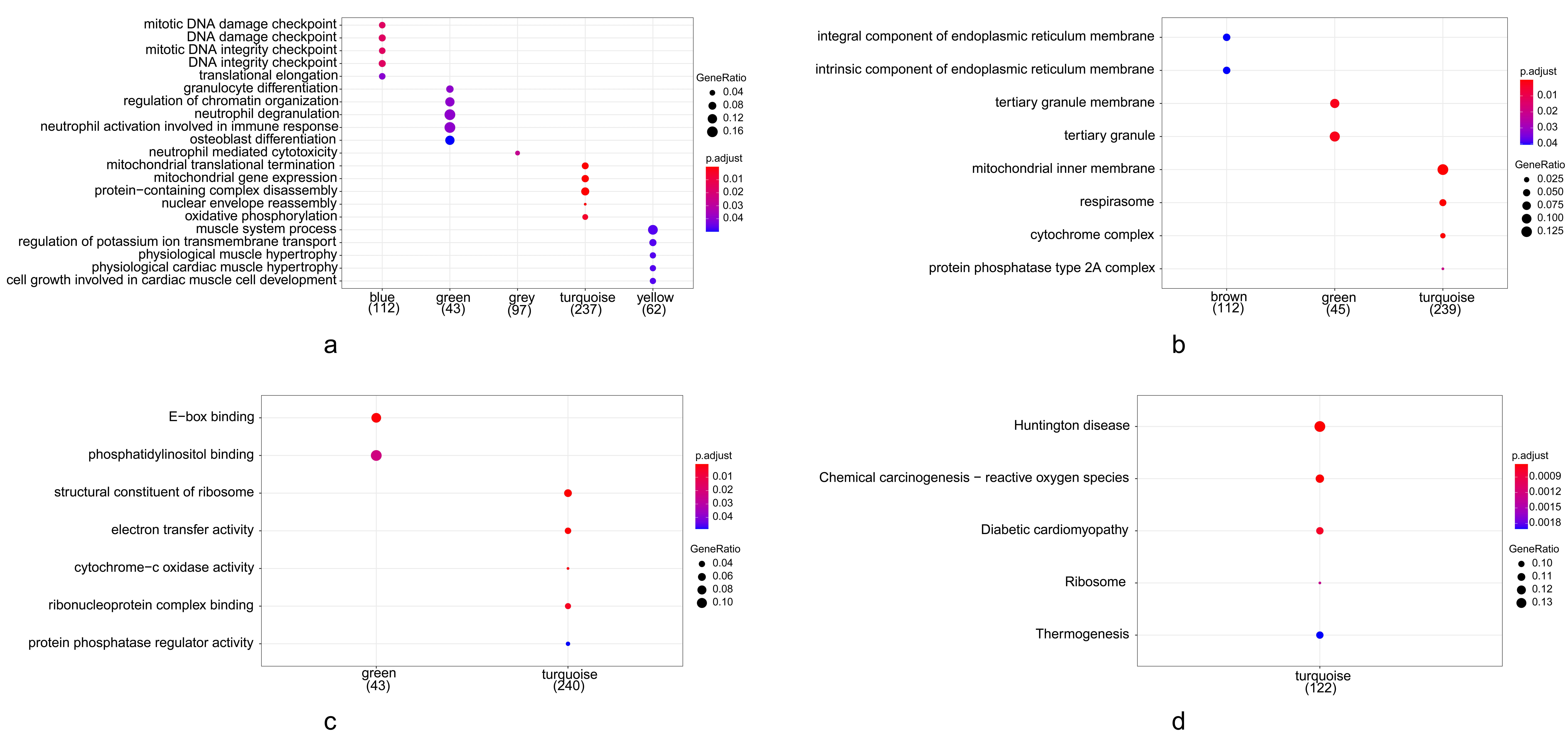 Fig. 4.
Fig. 4.GO annotation and KEGG pathway enrichment analysis. (a) Biological process of module. (b) Cell component of module. (c) Molecule function of module. (d) KEGG pathway of module. Abbreviations: GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Gene and Genomes.
Correlations between the modules of the DCM and control groups and the phenotypes of the six clinics (group, age, sex, body mass index, ejection fraction, left ventricular internal diameter at end-diastole) were calculated, and their corresponding p values were analyzed for magnitude. Except for the gray module, the blue module (r = –0.14) had the strongest negative correlation, while the turquoise module (r = 0.16) had the strongest positive correlation (Fig. 3).
Based on GS and MM values (Supplementary Fig. 2), 12 genes in the
yellow module were identified as hub genes RPS15, OXA1L, MLF2, GYPC, GNB2L1,
EMC4, COPS7A, BANF1, ATP5G2, ARL2, AP2S1 and AK2. Through LASSO regression, four
variables, GYPC (glycophorin C), MLF2 (myeloid leukemia factor 2), COPS7A (COP9
signalosome subunit 7A) and ARL2 (ADP ribosylation factor like GTPase 2), were
ultimately identified and used to construct the model. ROC curve analysis was
performed on the regression model to predict DCM in the training set with an area
under the curve (AUC) of 0.853. To further test the diagnostic efficacy, we used
the GSE19303 datasets as the validation set, and the AUC was 0.89, indicating
that the model gene had a high diagnostic value. The C-index for the prediction
model of the cohort was 0.90, and it was 0.85 through bootstrapping validation,
demonstrating the model’s good discrimination. In addition, we found that there
were significantly upregulated in the expression levels of GYPC (p
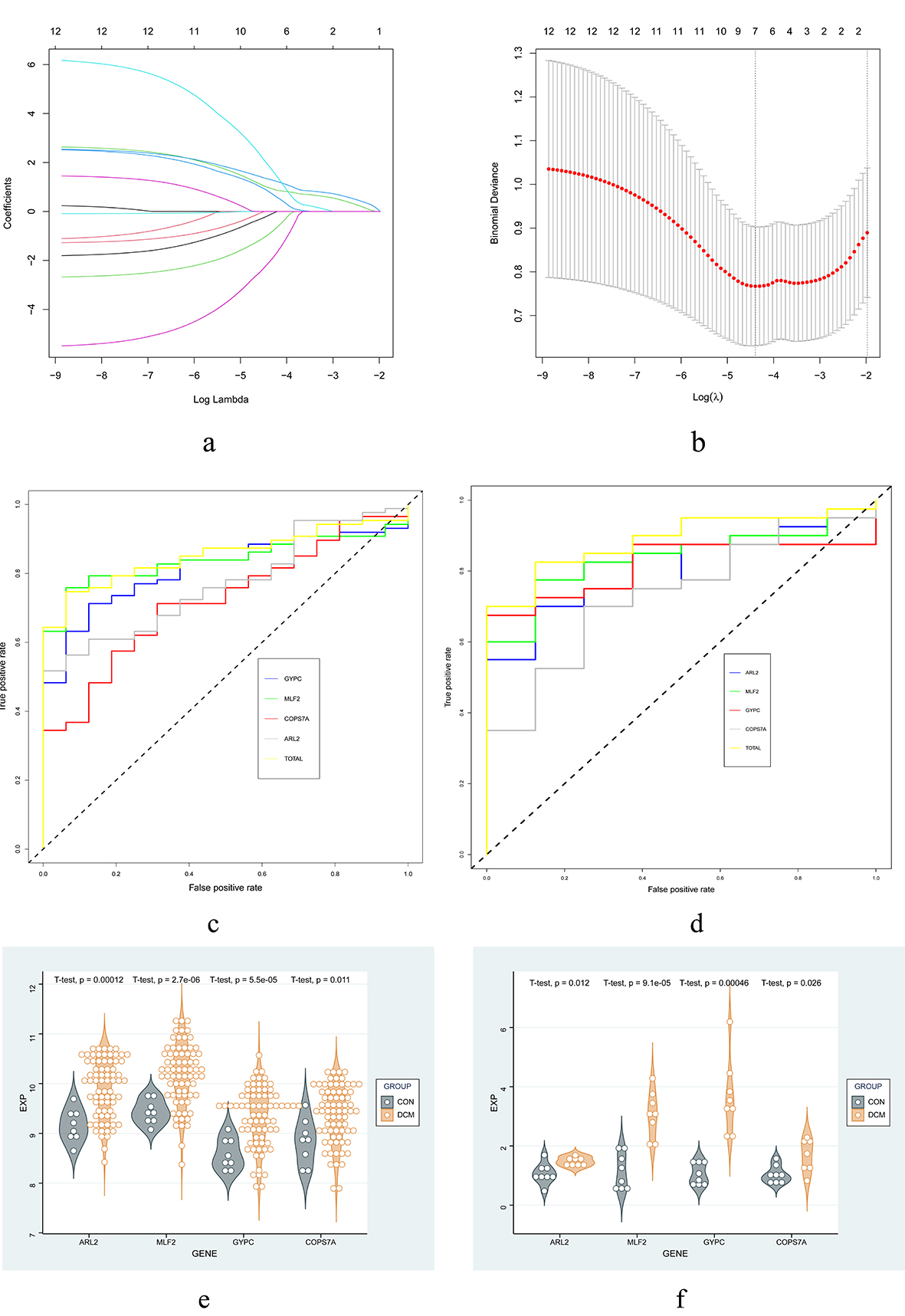 Fig. 5.
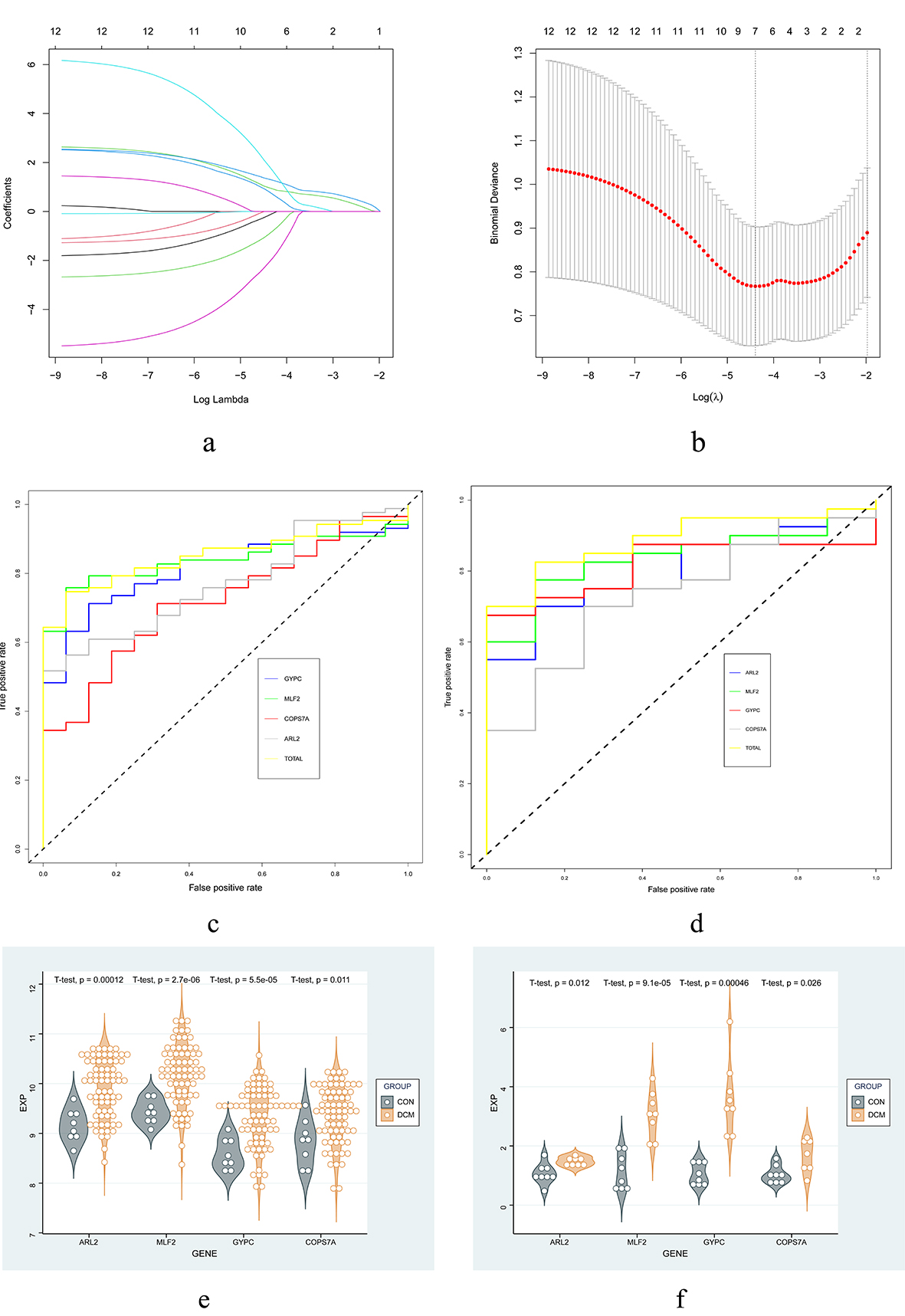
Fig. 5.Screening and verification of key genes. (a,b) LASSO model. (c,d) ROC curves for training set and validation set. (e,f) Expression of hub genes in the validation set and qPCR. Abbreviations: LASSO, least absolute shrinkage and selection operator; ROC, receiver operator characteristic; qPCR, quantitative polymerase chain reaction.
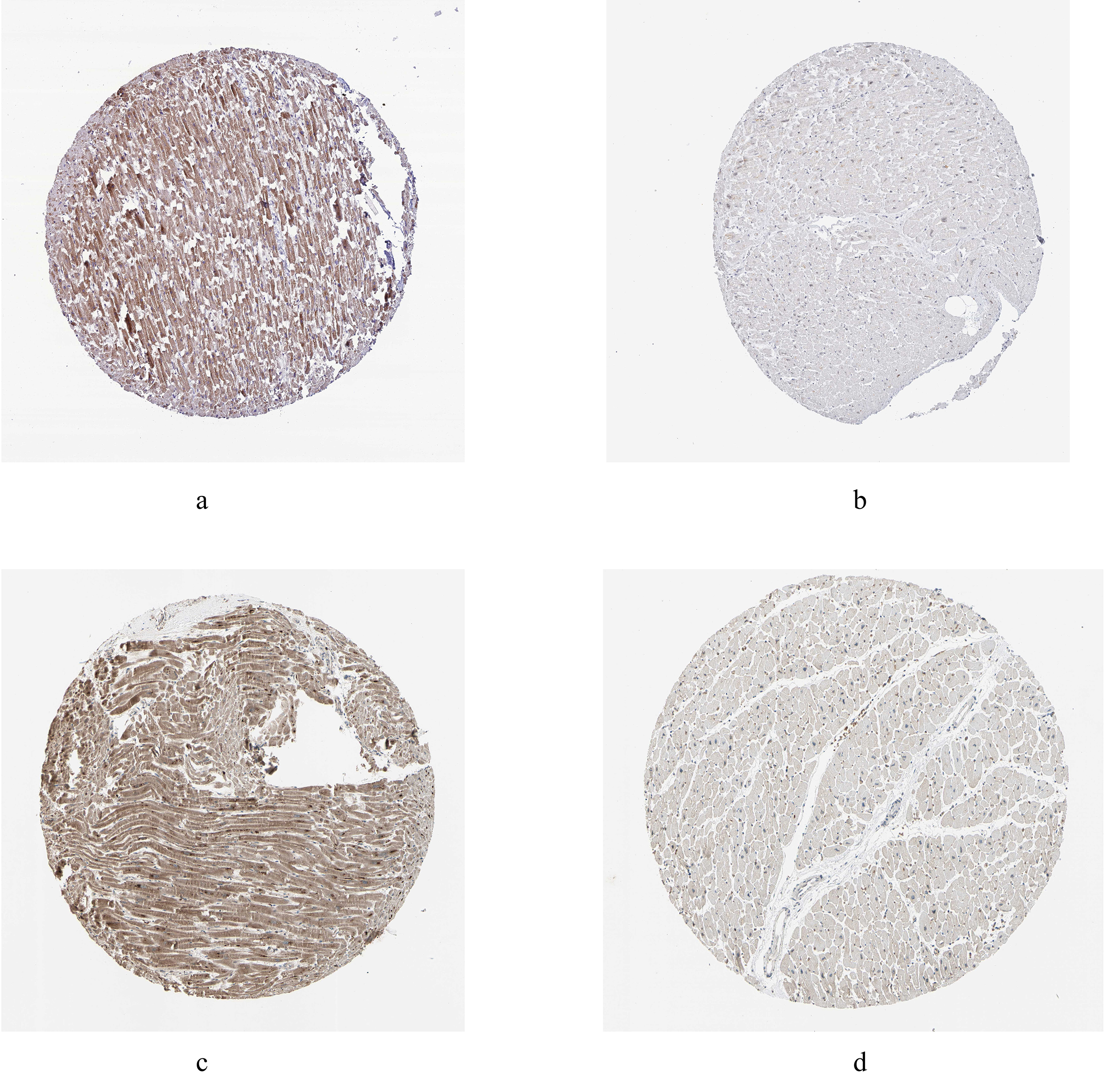 Fig. 6.
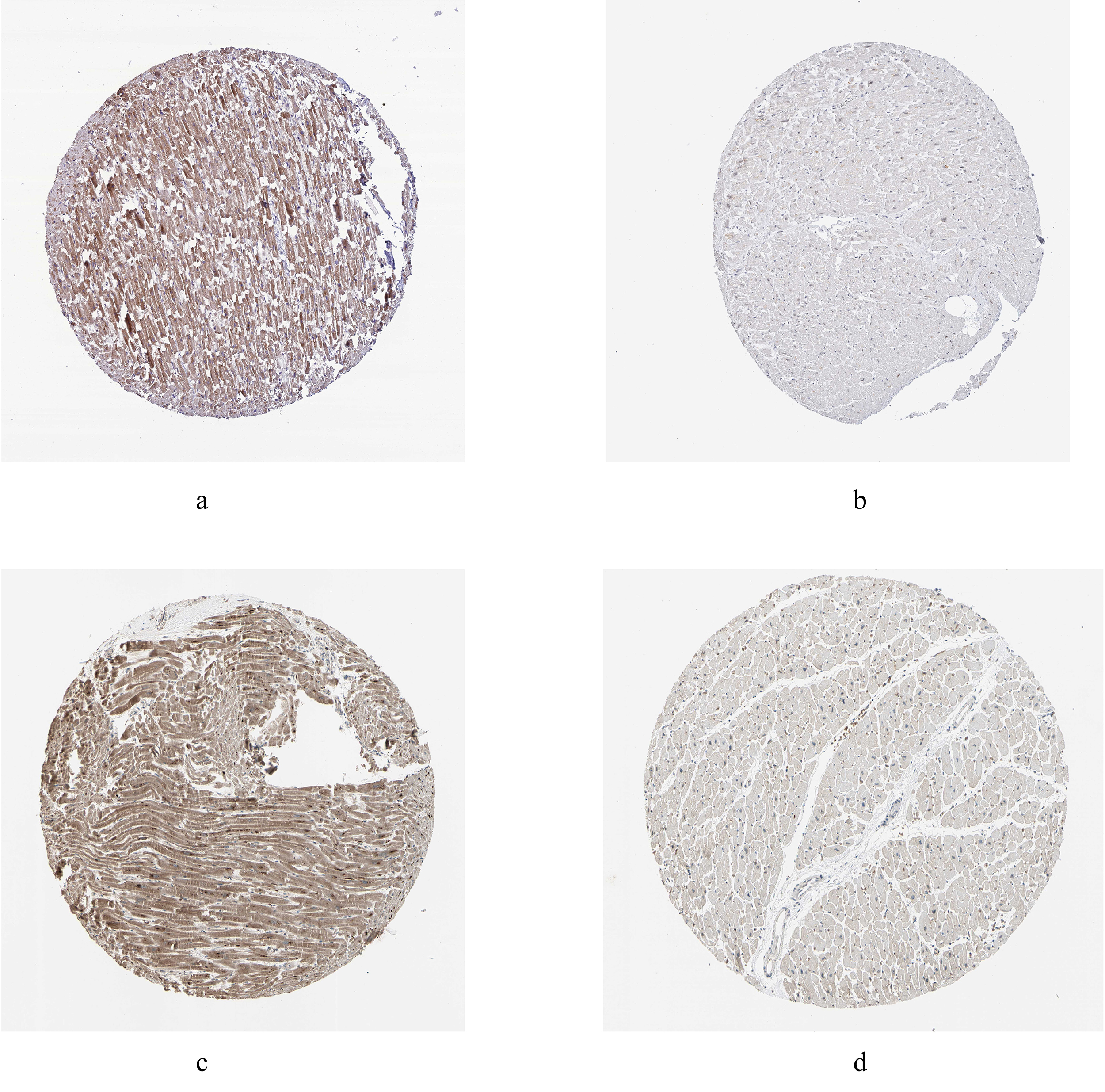
Fig. 6.Immunohistochemistry of the hub genes based on the Human Protein Atlas database (HPAD). (a) Protein levels of ARL2 in myocardial tissue. (b) Protein levels of COPS7A in myocardial tissue. (c) Protein levels of MLF2 in myocardial tissue. (d) Protein levels of GYPC in myocardial tissue.
After validating the JASPAR preselected transcription factors by gene expression
correlation (p
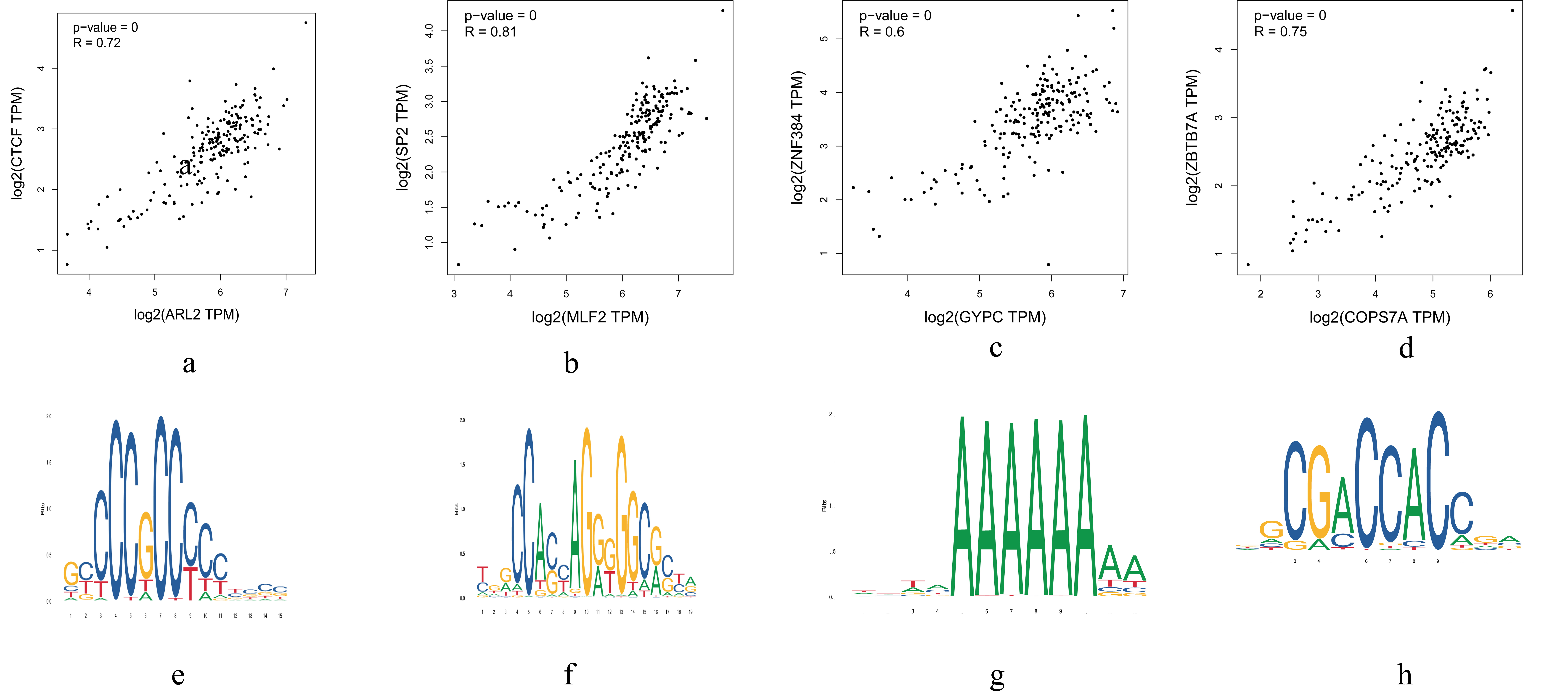 Fig. 7.
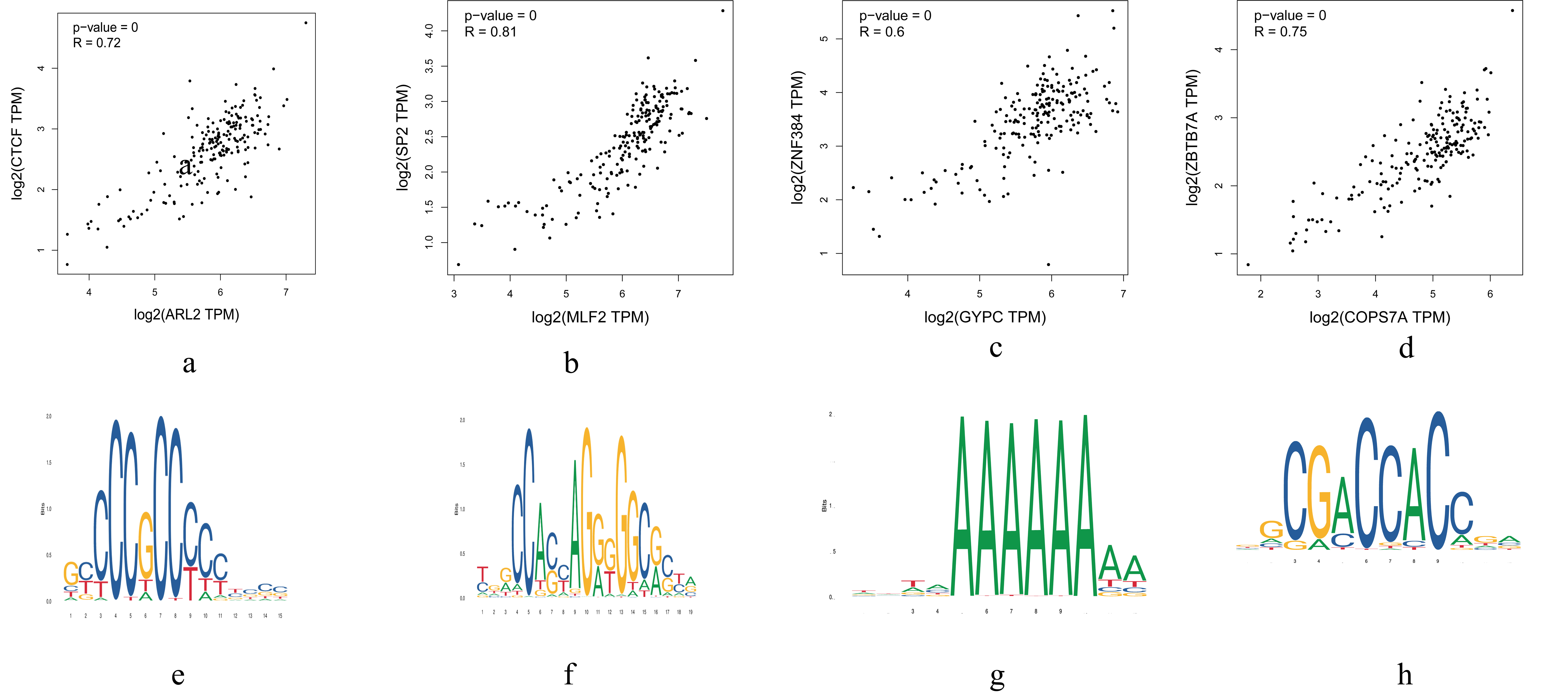
Fig. 7.Transcription factor validation and binding threshold. (a–d) Scatter plots of gene expression correlations corresponding to the most likely transcription factors bound by the hub genes, respectively. (e–h) Binding sites for transcription factors.
To find drugs for DCM, we searched the Cmap database and subsequently obtained seven drug candidates, namely, trihexyphenidyl, meclofenamic acid, daunorubicin, simvastatin, doxorubicin, dirithromycin and spiperone (Table 1). The 2D (Supplementary Fig. 3) and 3D structures of these drugs were provided by PubChem, and their corresponding active human validated targets were identified. The spatial structure of the gene and the spatial structure of the drug can be used for further molecular docking simulations to seek possible mechanisms of occurrence.
| Cmap name | Mean | Enrichment | Specificity | Percent non-null | Canonical SMILES | Targetname |
| Trihexyphenidyl | 0.666 | 0.946 | 0.0056 | 100 | C1CCC(CC1)C(CCN2CCCCC2) | NULL |
| (C3=CC=CC=C3)O | ||||||
| Meclofenamic acid | 0.543 | 0.796 | 0 | 100 | CC1=C(C(=C(C=C1)Cl) | PTGS1, TTR, AKR1C3, PTGS2, AKR1C1, AKR1C2, ABCB11 |
| NC2=CC=CC=C2C(=O)O)Cl | ||||||
| Daunorubicin | –0.623 | –0.846 | 0.0175 | 100 | CC1C(C(CC(O1)OC2CC | BLM, CBFB, GNAI1, HDAC1, HDAC10, HDAC11, HDAC2, HDAC3, HDAC4, HDAC5, HDAC6, HDAC7, HDAC8, HDAC9, HTT, MAPT, POLK, RGS12, RUNX1, TOP1, TOP2A, USP1, YAP1 |
| (CC3=C2C(=C4C(=C3O)C(=O)C5=C | ||||||
| (C4=O)C(=CC=C5)OC)O)(C(=O)C)O)N)O | ||||||
| Simvastatin | –0.581 | –0.822 | 0 | 100 | CCC(C)(C)C(=O)OC1CC(C=C2C1C(C(C=C2)C) | BSLCO1B1, NR2E3, MDM4, MDM2, IDH1, ICMT, HMGCR, CYP3A4, CYP2D6, CYP2C9, CYP2C8, CYP2C19, CYP1A2, ABCB11 |
| CCC3CC(CC(=O)O3)O)C | ||||||
| Doxorubicin | –0.65 | –0.845 | 0.1353 | 100 | CC1C(C(CC(O1)OC2CC(CC3=C2C(=C4C | ABCC3, ABCC4, BCL2, BCL2L1, C1SD1, CYP1A2, CYP2C19, CYP2C9, CYP2D6, DHCR7, EBP, EPAS1, HIF1A, POLK, PPM1D, SLCO1B3, TOP1, TOP2A, TOP2B, TP53 |
| (=C3O)C(=O)C5=C(C4=O)C(=CC=C5)OC)O) | ||||||
| (C(=O)CO)O)N)O | ||||||
| Dirithromycin | –0.559 | –0.826 | 0 | 100 | CCC1C(C2C(C(C(CC(C(C(C(C(C(=O)O1)C) | PTGER2, NLRP1 |
| OC3CC(C(C(O3)C)O)(C)OC)C)OC4C(C(CC(O4) | ||||||
| C)N(C)C)O)(C)O)C)NC(O2)COCCOC)C)(C)O | ||||||
| Spiperone | –0.529 | –0.873 | 0.0614 | 100 | C1CN(CCC12C(=O)NCN2C3=CC=CC=C3) | ACBC11, ADRA1A, AVPR1A, CHRM1, CHRM4, CHRM5, CRHBP, CRHR2, DRD1, DRD2, DRD3, DRD4, GNA15, HCRTR1, HTR1A, HTR1D, HTR2A, HTR2B, HTR6, HTR7, MDM2, MDM4, OXTR, TAAR1, TRHR |
| CCCC(=O)C4=CC=C(C=C4)F | ||||||
| Canonical SMILE, Internationally recognized atomic structure; Targetname, Validated targets with activity. | ||||||
Molecular docking simulations were used to delve into the possible therapeutic mechanisms of these drugs. The binding energy between two counterparts was calculated to predict their affinity. Binding energies below 0 indicated that the two molecules bind spontaneously, with smaller binding energies leading to a more stable conformation. The 3D structure of MLF2 was not available in the Protein Data Bank and ligands corresponding to GYPC and COPS7A were not suitable for molecular docking simulation. The binding energies of ARL2 with Trihexyphenidyl, Meclofenamic acid, Daunorubicin, Simvastatin, Doxorubicin, Dirithromycin and Spiperone were –6.0, –6.7, –7.2, –7.6, –7.4, –6.7 and –5.0 kcal/mol, respectively. Fig. 8 detailed the local structure of molecular docking. It has been illustrated that Doxorubicin acts as an induction target for DCM as positive controls [20]. From the point of view of binding energy this suggested that Simvastatin may be a target for the treatment of DCM (the binding energy of the protein is lower than that of the positive control).
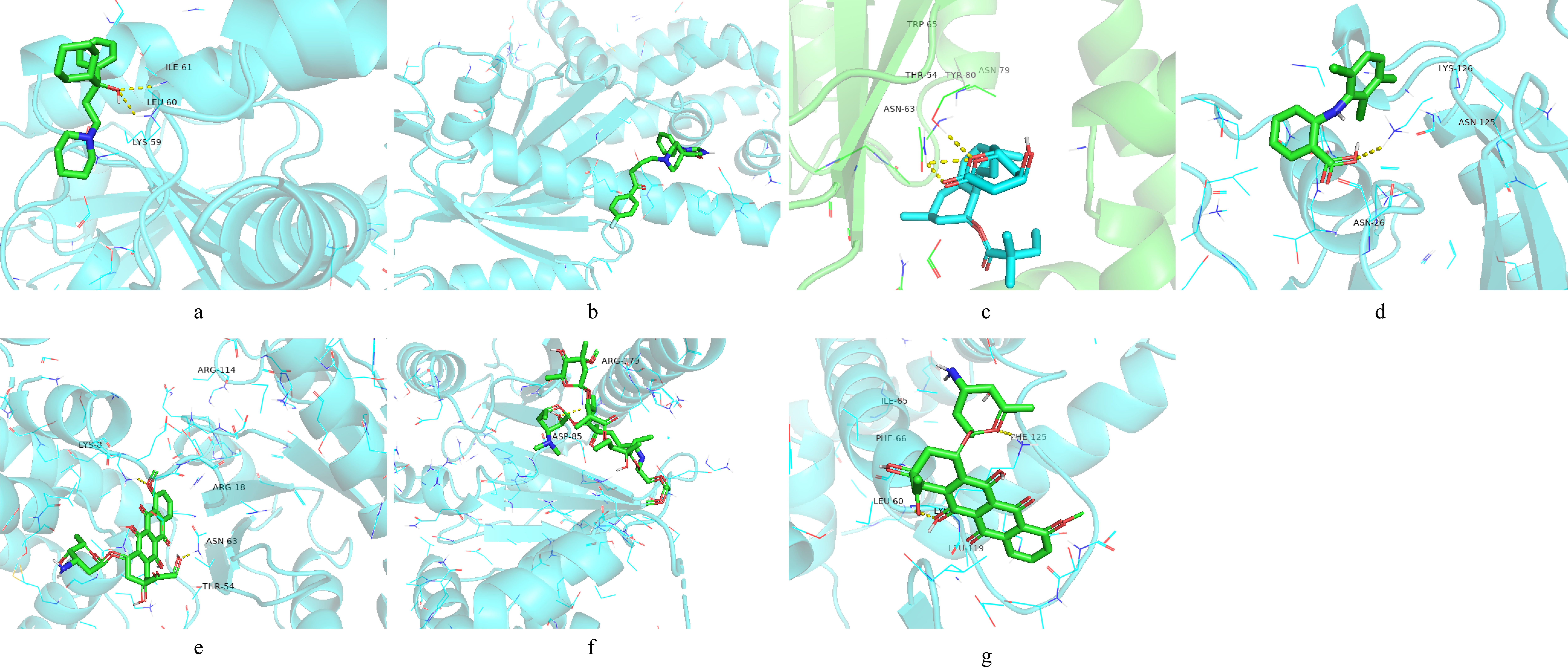 Fig. 8.
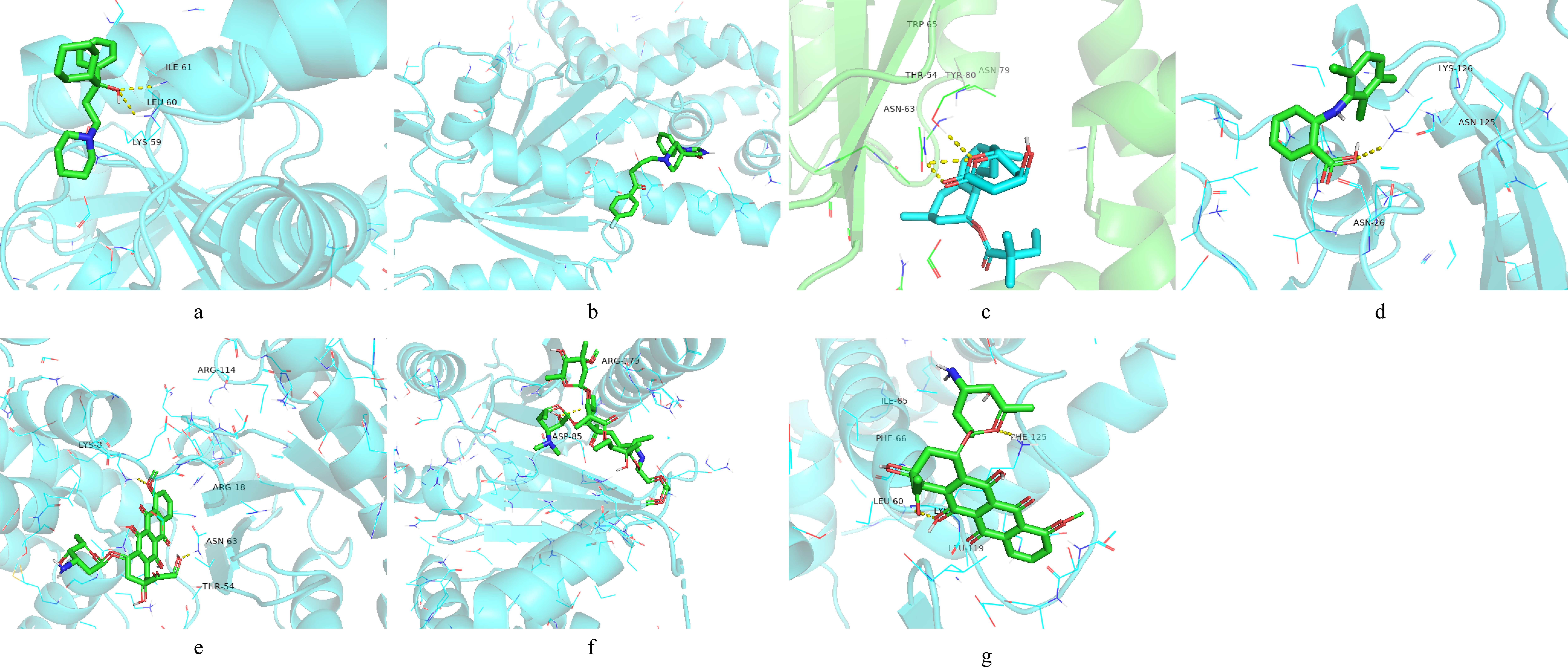
Fig. 8.The results of the molecular docking simulations. (a) ARL2 with Trihexyphenidyl. (b) ARL2 with Meclofenamic acid. (c) ARL2 with Daunorubicin. (d) ARL2 with Simvastatin. (e) ARL2 with Doxorubicin. (f) ARL2 with Dirithromycin. (g) ARL2 with Spiperone.
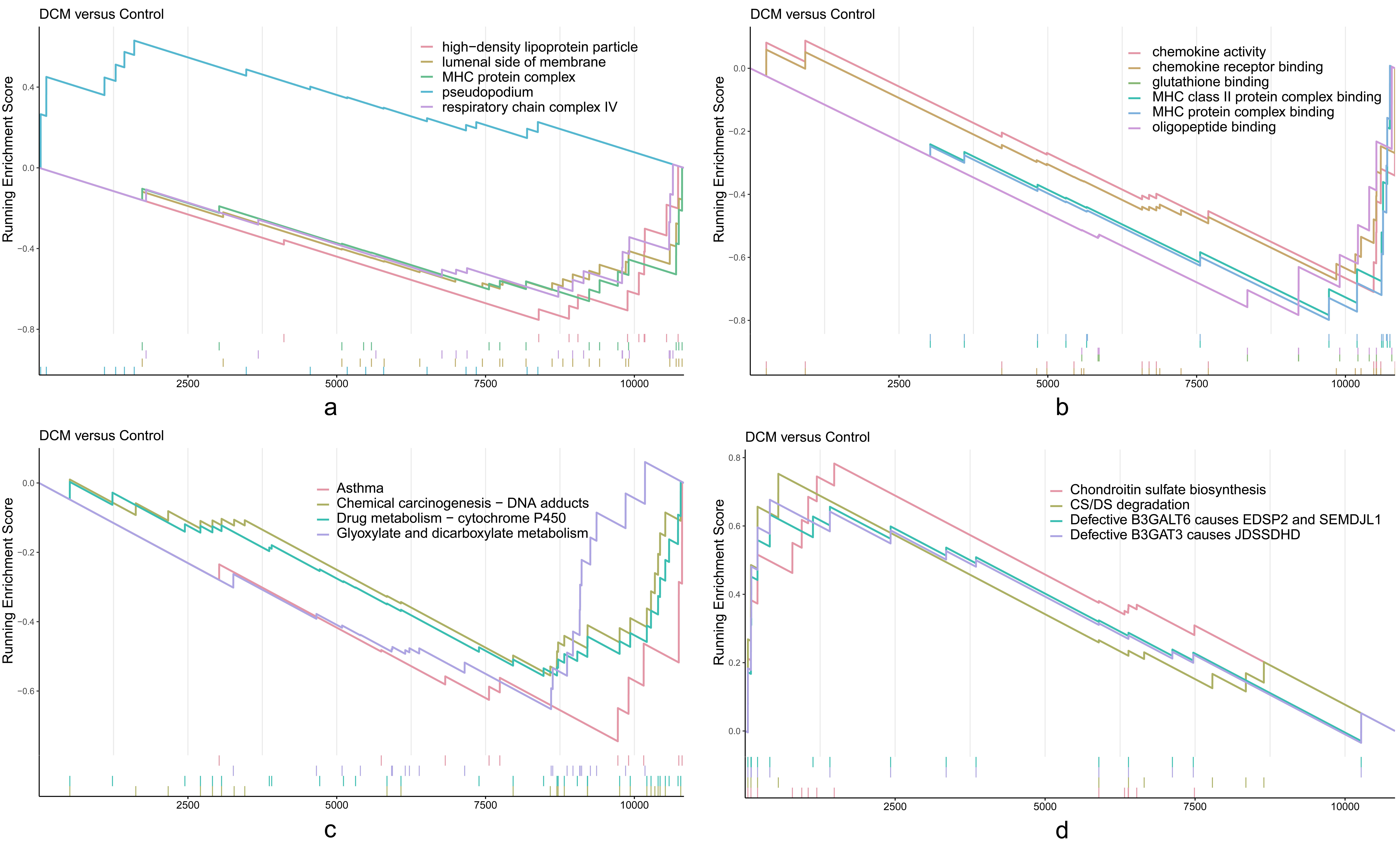
Gene set enrichment analysis showed that, compared to control samples, the DCM group was significantly enriched in cell component (CC) such as pseudopodium, lumenal side of membrane, respiratory chain complex IV, MHC protein complex and high-density lipoprotein particle; in molecule function (MF) such as chemokine receptor binding, chemokine activity, glutathione binding, oligopeptide binding, MHC class II protein complex binding and MHC protein complex binding; in signaling pathways such as Chemical carcinogenesis - DNA adducts, Drug metabolism - cytochrome P450, Glyoxylate and dicarboxylate metabolism and Asthma; and in reactome such as Chondroitin sulfate biosynthesis, CS/DS degradation, Defective B3GAT3 causes JDSSDHD and Defective B3GALT6 causes EDSP2 and SEMDJL1 (Fig. 9 and Table 2).
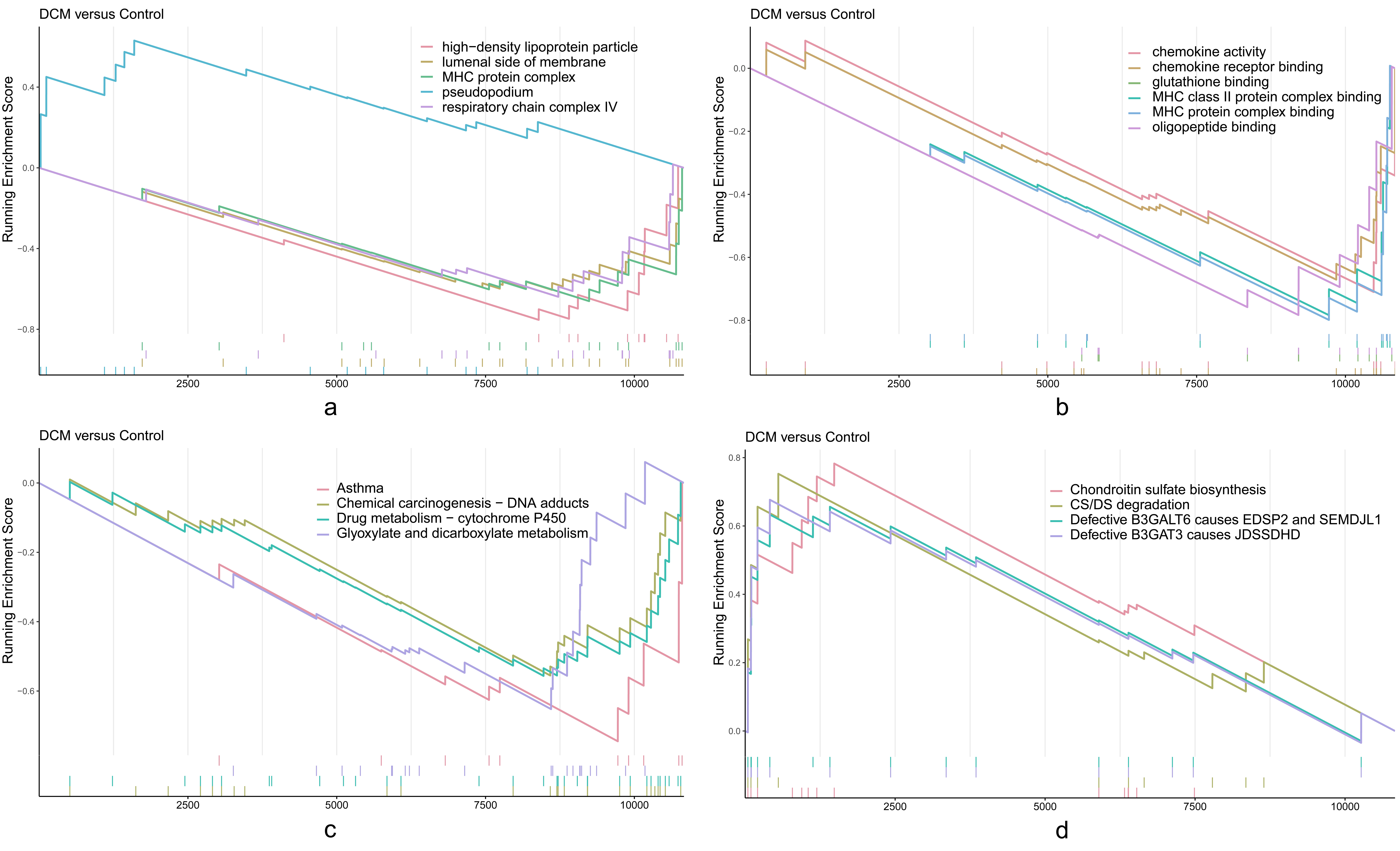 Fig. 9.
Fig. 9.GSEA. (a) Cell component of gene set.(b) Molecule function of gene set. (c) KEGG pathway of gene set. (d) Reactome of gene set. Abbreviations: GSEA, gene set enrichment analysis; KEGG, Kyoto Encyclopedia of Genes and Genomes.
| Term | Description | set size | enrichment score | NES | p value | q values |
| GO:0031143 | pseudopodium | 16 | 0.630406438 | 1.765318418 | 0.005649718 | 0.094103654 |
| GO:0098576 | lumenal side of membrane | 23 | –0.604675899 | –1.906359382 | 0.002123142 | 0.059312504 |
| GO:0045277 | respiratory chain complex IV | 16 | –0.639024812 | –1.813987412 | 0.006369427 | 0.097489377 |
| GO:0042611 | MHC protein complex | 15 | –0.660610096 | –1.860053648 | 0.004158004 | 0.078901538 |
| GO:0034364 | high-density lipoprotein particle | 10 | –0.753329992 | –1.876851266 | 0.004048583 | 0.078901538 |
| GO:0042379 | chemokine receptor binding | 22 | –0.670673966 | –2.092976246 | 0.00209205 | 0.144831662 |
| GO:0008009 | chemokine activity | 14 | –0.709970379 | –1.957303241 | 0.001976285 | 0.144831662 |
| GO:0043295 | glutathione binding | 11 | –0.782922238 | –1.997545034 | 0.002057613 | 0.144831662 |
| GO:1900750 | oligopeptide binding | 11 | –0.782922238 | –1.997545034 | 0.002057613 | 0.144831662 |
| GO:0023026 | MHC class II protein complex binding | 12 | –0.783391223 | –2.040328703 | 0.002053388 | 0.144831662 |
| GO:0023023 | MHC protein complex binding | 14 | –0.798539963 | –2.201478969 | 0.001976285 | 0.144831662 |
| hsa05204 | Chemical carcinogenesis - DNA adducts | 27 | –0.555294552 | –1.802179492 | 0.002053388 | 0.097124772 |
| hsa00982 | Drug metabolism - cytochrome P450 | 32 | –0.556436183 | –1.909942264 | 0.003960396 | 0.137320359 |
| hsa00630 | Glyoxylate and dicarboxylate metabolism | 21 | –0.652250567 | –2.059525422 | 0.001934236 | 0.097124772 |
| hsa05310 | Asthma | 10 | –0.745038804 | –1.854621025 | 0.006048387 | 0.153475695 |
| R-HSA-2022870 | Chondroitin sulfate biosynthesis | 14 | 0.782742829 | 2.134302347 | 0.001941748 | 0.216045732 |
| R-HSA-2024101 | CS/DS degradation | 10 | 0.752675863 | 1.845047523 | 0.004048583 | 0.222934147 |
| R-HSA-3560801 | Defective B3GAT3 causes JDSSDHD | 14 | 0.676721314 | 1.845213822 | 0.001941748 | 0.216045732 |
| R-HSA-4420332 | Defective B3GALT6 causes EDSP2 and SEMDJL1 | 15 | 0.65615414 | 1.832055703 | 0.003868472 | 0.222934147 |
| Abbreviations: GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Gene and Genomes; DCM, dilated cardiomyopathy; DEG, differentially expressed gene. | ||||||
DCM is one of the leading causes of cardiac insufficiency. Despite the increasing number of improved treatments, due to the aging population structure, the proportion of disease occurrence and the number of illnesses is increasing, especially in China. DCM has a serious impact on the quality of life of patients and causes a heavy social and economic burden. The comprehensive molecular mechanisms involved in DCM are still unclear, and advances in high-throughput technologies and bioinformatics distribution can provide a more comprehensive and in-depth understanding of the disease process.
In this study, a systematic collection of DCM-associated genes, followed by WGCNA using multiple clinical features, resulted in the identification of six modules, of which the turquoise coexpression module was the most significantly associated with the occurrence of DCM. We selected the turquoise module as the key module for DCM occurrence and used it for the subsequent hub gene search. In addition, these coexpression modules may interact with each other during DCM. The expression profiles of the hub genes were extracted to construct the LASSO model, and ROC curve analysis showed that the LASSO model had high AUC values for both the training and test sets and could be used as a biomarker for DCM.
TFs regulate gene expression by binding to the promoter regions of target genes; therefore, regulating the biology and binding properties of TFs can be used for targeted therapy [21]. SP2 acts as a cofactor to recruit and increase interactions between the SP2 – Pbx1:Prep1 – Nf-y complex, which in turn promotes genomic binding [22]. The complex activates downstream targeted proteins to control cell proliferation and apoptosis. CTCF is a transcriptional regulatory protein that encodes 11 highly conserved zinc finger structural domains that allow different combinations of structural domains to bind different DNA target sequences and proteins [23]. For example, these domains can bind to a complex containing histone acetyltransferase (HAT) and act as a transcriptional activator or to a complex containing histone deacetylase (HDAC) and act as a transcriptional repressor; if a domain binds to a transcriptional insulator element, it can block communication between the enhancer and the upstream promoter, thereby regulating blot expression [24, 25]. Subsequent in vitro and in vivo experiments are needed for verification of the modes of action of the TFs screened in this study.
Our results indicated that ARL2 and MLF2 were the central genes obtained after modular analysis and expression validation in DCM. ARL2 is a small G-protein that belongs to the Arf-like small G-protein subfamily, which promotes mitosis by acting on cytoskeletal tissue [26]. Previous studies [27] have shown that ARL2 overexpression increases polymerizable soluble heterodimers, while ARL2 depletion increases microtubule dynamic instability. Thus, ARL2 depletion significantly reduced the percentage of cells in G2/M phase and mitotic cells. In addition, ARL2 is involved in regulating mitochondrial functions [28], including the maintenance of mitochondrial morphology, motility and ATP levels. ARL2 consumption reduces mitochondrial membrane potential and regulates downstream protein factors to promote mitochondrial fusion and activity [29]. MLF2 and MLF1 are members of the myeloid leukemia factor (MLF) family and are important paraphyletic homologs, sharing nearly 40% identity [30]. The MLF family regulates apoptosis and transcription processes by blocking the association between HS1-associated protein X-1 and HtrA serine peptidase 2 to inhibit the maturation of HtrA2, thus maintaining normal mitochondrial function [31]. These regulatory patterns and action sites are similar to those of the previous DCM gene families. Considering the findings our study, ARL2 and MLF2 may serve as therapeutic targets.
We used enrichment analysis of modules and GSEA to explore the key mechanisms of DCM. Excitingly, in contrast to previous results [32], although different datasets were used, the key to the analysis was all about functional changes in the cellular matrix with abnormal mitochondrial function. We also found that the regulated factor MLF2 may act in both leukemia and DCM, similar to previous studies [33] in which the MLLT family was involved in the pathology of DCM, suggesting that modulation of this core target may benefit both difficult diseasesUnlike previous studies [33], we did not only perform integration of multiple datasets and validation at the dataset level, we also performed expression validation through molecular biology experiments, which made our experimental results more reliable. Moreover, we further explored subsequent therapeutic strategies from a pharmacological perspective.
We must acknowledge that there are still limitations to this research. First, the verification of hub genes and their functions has only been tested in human venous blood, but not in other animals model or even human clinical trials. Second, inhibition and overexpression experiments of hub genes have not been completed and need to be further supplemented.
Overall, our study demonstrated that ARL2, MLF2, GYPC and COPS7A could be potential gene biomarkers for DCM. However, the identification of the potential key pathways and genes was based on bioinformatic tools and will require further validation by molecular experiments. The extent to which the upregulation and downregulation of ARL2 and MLF2 contribute to DCM development and the specific modes of action of TFs in DCM patients remain to be tested.
The deidentified participant data will be shared on a request basis. Please directly contact the corresponding author to request data sharing.
QG—Conceptualization, Methodology, Software, Investigation, Formal Analysis, Writing - Original Draft; QQ—Data Curation, Visualization, Writing - Original Draft; LW—Visualization, Data Curation, Investigation; SL—Resources, Supervision; XZ—Software, Validation; AD—Visualization, Writing - Review & Editing; QZ—Validation, Investigation; IC—Data Curation, Writing - Review & Editing; RG—Software, Methodology; XL—Conceptualization, Resources, Supervision, Writing - Review & Editing.
The research protocol was approved by the Ethics Committee of Nanjing Medical University (license number: 2017-SR-086).
The authors thanked the patients and investigators who participated in GEO for providing the data and the authors appreciated Jiajin Chen, Department of Biostatistics, School of Public Health, Nanjing Medical Univeristy for the statistical guidance, appreciated Mengli Chen and Mengsha Shi for experimental instruction.
This research received no external funding.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/j.fbl2708246.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.