†These authors contributed equally.
Academic Editor: Giovanni Monni
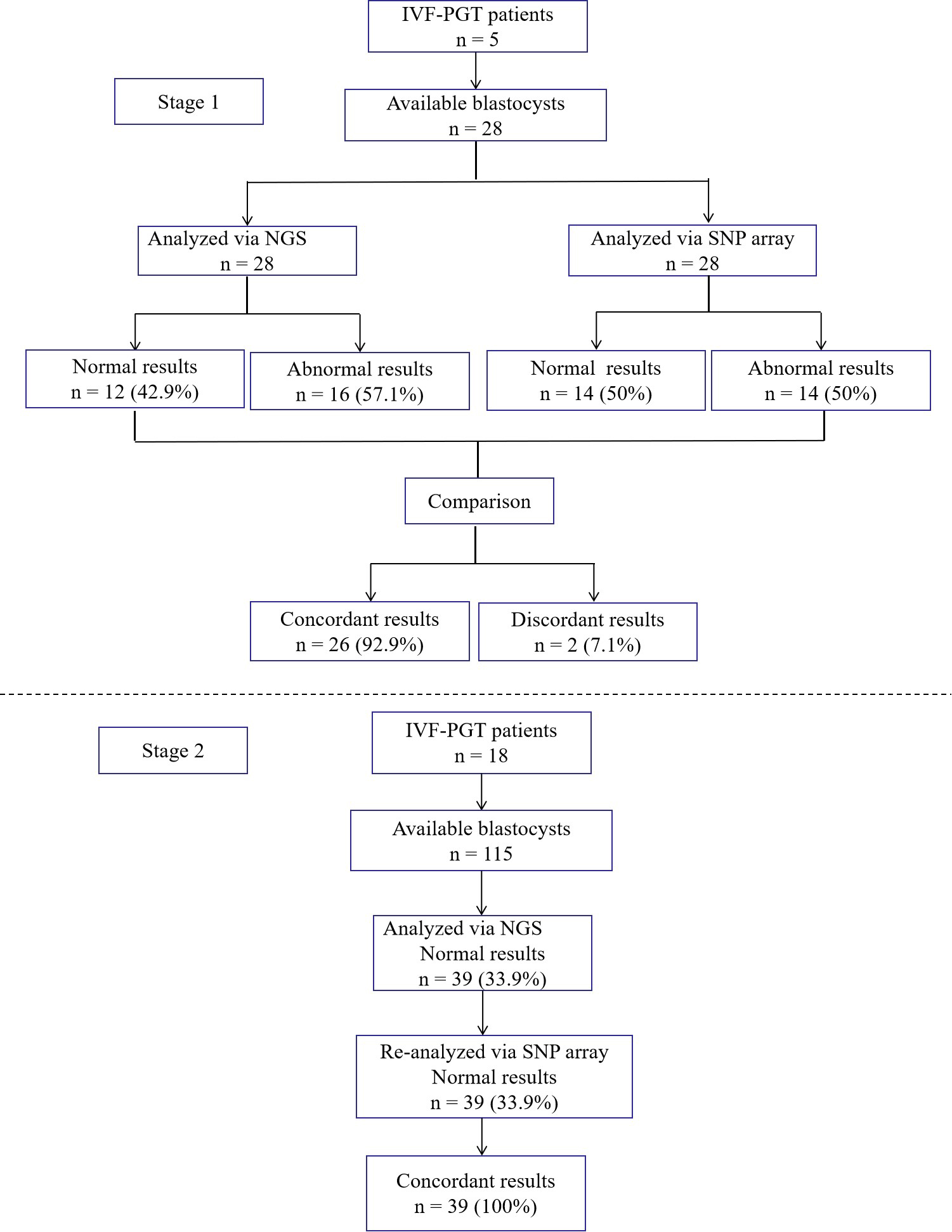
Background: This study aimed to compare the use of next-generation sequencing (NGS) and single-nucleotide polymorphism (SNP) array in preimplantation genetic testing for aneuploidy (PGT-A) in the same blastocyst. Methods: We performed a retrospective study on 67 embryos (from 23 couples), where PGT-A was carried out. A trophectoderm (TE) biopsy was performed on the blastocyst, and the 24-chromosomal ploidy status was analyzed. Initially, 28 blastocysts with unknown ploidy were analyzed using both NGS and SNP array. Thereafter, 39 blastocysts with euploidy detected via NGS were re-analyzed using SNP array. Results: In the first stage, the concordance rate was 92.9% (26/28). Among the 28 blastocysts, 16 were abnormal, and 12 were euploid when analyzed using NGS. Among the 16 abnormal blastocysts, two showed mosaicisms when analyzed using NGS but were found to be euploid using the SNP array. In the second stage, the concordance rate was 100% (39/39) when analyzing the normal blastocysts. After single blastocyst transfer in 29 frozen embryo transfer cycles, the clinical pregnancy rate was 75.9% (22/29), the ongoing pregnancy rate was 69.0% (20/29), and the live birth rate was 69.0% (20/29). Nineteen couples (20 babies) had healthy babies. Their prenatal diagnosis results and karyotype analysis after delivery were concordant with the PGT results. Two cycles miscarried, and the abortion villus exhibited euploidy. Conclusions: There was a high concordance rate between NGS and SNP array. TE biopsy combined with NGS for PGT was an efficient strategy to identify the suitability of embryos for transfer.