1 Science Division, Pennsylvania State University, Berks Campus, Reading, PA 19610, USA
Abstract
Metabolic homeostasis is regulated by numerous genes, whose dysregulation leads to metabolic diseases such as obesity and diabetes. Several genes important for lipid storage were identified in a buoyancy-based screen in Drosophila larvae, including Glucose transporter 1 (Glut1), which encodes a glucose uniporter. Previous studies have identified metabolic functions of Glut1 in the whole fly brain; however, the specific neurons in which Glut1 acts to regulate nutrient storage remain unknown.
To determine the neuronal populations in which Glut1 regulates lipid and carbohydrate storage, Glut1 levels were decreased in specific neurons, and triglycerides (TAGs) and glycogen levels were measured. We specifically decreased Glut1 expression in corazonin (Crz)-expressing neurons, a neuronal population that expresses the corazonin gene (Crz), which encodes a neuropeptide involved in carbohydrate metabolism.
Targeting RNAi against Glut1 in Crz neurons reduced glycogen levels in males but did not alter TAG levels. To further characterize this nutrient storage phenotype, we measured the expression of two genes involved in glycogen storage, glycogen phosphorylase (Glyp) and glycogen synthase (Glys) as well as the Crz transcript. Notably, knocking down Glut1 in Crz-expressing neurons increased Glys and Crz transcript levels.
These data suggest that Glut1 acts in the Crz-expressing neurons to regulate Crz levels and organismal glycogen metabolism.
Keywords
- Drosophila
- corazonin (Crz)
- glycogen
- neuropeptides
- glycogen synthase
- glycogen phosphorylase
- carbohydrate metabolism
Over the years, various metabolic disorders such as type 2 diabetes and obesity have been linked to one or more genetic defects within the human genome [1]. Understanding how these genes function to regulate metabolic homeostasis has provided insight into potential drug development and treatments against these metabolic diseases. However, the full complement of genes that regulate organismal metabolism are not fully understood. The fruit fly, Drosophila melanogaster, has recently emerged as an excellent model for studying metabolism [2]. For example, the Drosophila genome contains homologs of approximately 75% of known human disease genes, many of which are involved in metabolism-related diseases [3]. Additionally, Drosophila has many conserved metabolic pathways and analogous metabolic organs compared with mammals allowing for information gained in Drosophila to be applied to mammalian systems [2]. Taking advantage of the rich genetic tools available in Drosophila, a number of genetic screens were performed to identify genes important for fat storage [4, 5, 6, 7]. One of the genes that was identified in one of these screens and we decided to study further was Glucose transporter 1 (Glut1) [7].
Glut1 encodes a glucose uniporter ubiquitously expressed in adult Drosophila; however, how Glut1 functions in different tissues to regulate the storage of triglycerides (TAGs) and glycogen is still unknown. Previous studies in our lab focused on the role of Glut1 in different metabolic tissues in Drosophila, such as its involvement within the fly’s adipocytes and neurons to regulate nutrient storage. For example, when RNAi was targeted against Glut1 in all adipocytes, there was a decrease in organismal TAG and glycogen levels [8]. Similarly, when RNAi was targeted against Glut1 in all neurons, there was a decrease in organismal TAG and glycogen storage [9]. To determine whether this metabolite storage defect is potentially due to Glut1 acting in specific neuronal populations to regulate nutrient storage, we aimed to characterize Glut1 function within neuronal populations that have been shown to be involved in modulating metabolic homeostasis and/or were glucose responsive in Drosophila. So far, decreasing Glut1 levels in the insulin-producing cells (IPCs) has been shown to recapitulate the phenotype seen when Glut1 levels were decreased in all neurons [9].
Corazonin (Crz) is a well-characterized neuropeptide that acts on the corazonin receptor (CrzR) to regulate various homeostatic processes, including ethanol metabolism, stress responses, male reproductive behavior, and systemic growth [10]. Crz is produced in a specific group of glucose-sensing neurons that act on the IPCs [11, 12]. In addition, targeting RNAi against CrzR on the IPCs causes an increase in starvation resistance and circulating glucose levels in well-fed male flies [11, 12]; however, targeting RNAi against Crz in cells producing Crz (using Crz-Gal4) results in no changes in ilp2, ilp3, and ilp5 transcript levels [11]. Furthermore, targeting RNAi against Crz in Crz-producing neurons causes an increase in glycogen, circulating glucose, and trehalose under well-fed conditions, with an increase in TAG levels after a 24-hour starvation period [11]. These results suggest that Crz-expressing neurons may function independently of the IPCs to regulate metabolic homeostasis in response to glucose availability; however, whether Glut1 acts in these neurons to regulate their function is unknown. This study will focus on identifying a potential role of Glut1 within the Crz-expressing neurons by decreasing Glut1 expression in these cells using RNAi and measuring TAGs and glycogen storage levels in well-fed male flies. The results of these experiments will increase our understanding of the role of the Crz-producing neurons in maintaining metabolic homeostasis.
The following flies were used in this study: w1118; P{w[+mC]=Crz-GAL4.391}4M (BL#51977, referred to as Crz-Gal4 here); y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF01355}attP2 (BL#31603, referred to as UAS-LucRNAi here), y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF03060}attP2 (BL#28645, referred to as UAS-Glut1RNAi-TRiP here), w1118; UAS-GFP RNAi (BL#9330, referred to as UAS-GFPRNAi here), w1118; UAS-Glut1 RNAi (VDRC#13326, referred to as UAS-Glut1RNAi-VDRC here). Flies were fed on cornmeal-sugar-yeast food (9 g Drosophila agar (66-103 Genesee Scientific, San Diego, CA, USA), 100 mL Karo Lite Corn Syrup, 65 g cornmeal, 40 g sucrose, and 25 g whole yeast in 1.25 L water) and grown at 25 °C in a 12 h:12 h light:dark cycle. Crz-Gal4 virgins were crossed with males from each UAS line and approximately one-week old male progeny were used. One-week old flies were chosen as the remnants of the larval fat body have been completely broken down and the adult fat body is fully formed by this time.
Two approximately one-week old male adult flies were homogenized in lysis buffer (140 mM NaCl; 50 mM Tris-HCl, pH 7.4; 0.1% Triton-X; 1X Pierce Protease Inhibitor Cocktail (A32965, Thermo Fisher Scientific, Waltham, MA, USA)). Homogenates were centrifuged at 13,000 rpm for 15 minutes at 4 °C. Protein levels were measured using the Pierce BCA Protein Assay Kit (PI23225, Thermo Fisher Scientific, Waltham, MA, USA), according to manufacturer instructions. TAG levels were measured using Infinity Triglyceride Reagent (TR22421, Thermo Fisher Scientific, Waltham, MA, USA), according to manufacturer instructions. Free glucose was measured using the Pointe Scientific Liquid Glucose Oxidase Reagent (23-666-288, Thermo Fisher Scientific, Waltham, MA, USA), according to manufacturer instructions. Total glucose was measured by combining each sample in a 1:1 ratio with 8 mg/mL amyloglucosidase (A1602-100mg, Sigma, St. Louis, MO, USA) in 0.2 M citrate buffer, pH. 5.0, and incubating at 37 °C for 2 hours. After incubation, total glucose was measured using the Liquid Glucose Oxidase Reagent (23-666-288, Thermo Fisher Scientific, Waltham, MA, USA). Glycogen concentrations were calculated by subtracting the free glucose from the total glucose. TAG and glycogen levels were normalized by their respective protein concentrations.
Ten approximately one-week old male flies were homogenized in Trizol (15-596-026, Thermo Fisher Scientific, Waltham, MA, USA), and RNA was extracted according to manufacturer instructions. 5 µg of RNA was DNase treated with a DNA-free kit (AM1906, Thermo Fisher Scientific, Waltham, MA, USA), according to manufacturer instructions, and then 0.25 µg of DNase-treated RNA was reverse-transcribed using qScript Ultra cDNA Supermix (76533-176, Quanta Biosciences, Beverly, MA, USA), according to manufacturer instructions. The relative expression of glycogen phosphorylase (Glyp), glycogen synthase (Glys) and corazonin (Crz) was quantitated using qPCR and the following reaction mixture: 1 µL cDNA, 200 nM primers, and 1X Perfecta SYBR Green Mix Master Mix (101414-160, Quanta Biosciences, Beverly, MA, USA) in a 25 µL reaction. The qPCR cycling conditions were 3 minutes at 95 °C and 40 cycles of the following conditions: 30 seconds at 95 °C, 1 minute at 60 °C, and 30 seconds at 72 °C followed by a melt curve. Glyp, Glys, Crz and rp49 expression was determined using a relative standard curve. Glyp, Glys, and Crz levels were then divided by rp49 levels for normalization. Relative expression of each normalized transcript in each genetic background control strain was set to 1 and expression in the genetic background matched Glut1RNAi line was calculated relative to the control. The following forward (F) and reverse (R) primer sequences were used: rp49-F 5′GACGCTTCAAGGGACAGTATCTG3′; rp49-R 5′AAACGCGGTTCTGCATGAG3′; Glyp-F 5′AACTTCCAGCGCAATGTAGC3′; Glyp-R 5′TGGGATCCTTCTTGATCCTG3′; Glys-F 5′CGCGAGGCTATAAAATCCAC3′; Glys-R 5′GGCAATCATAAAGCCAAGGA3′; Crz-F 5′GGCTCGAGCGCTGTCTATC3′; Crz-R 5′ACTCGGTTGGCATTGAAGTC3′.
Normality tests were performed using the Shapiro-Wilk test on the Statistics
Kingdom website (https://www.statskingdom.com/shapiro-wilk-test-calculator.html). The TAG and glycogen
data were not normally distributed, so a Mann-Whitney U test was performed to
compare the averages between the Glut1RNAi and appropriate control flies
using the Statistics Kingdom website
(https://www.statskingdom.com/170median_mann_whitney.html). The qPCR data were
normally distrubuted, so a two-tailed students t-test was performed to
compare the averages between the Glut1-RNAi and the appropriate control
flies using Excel software. The p-value
To understand the role of Glut1 in Crz-expressing neurons, Glut1 was decreased in all Crz-expressing neurons using the Gal4/UAS system, and the storage of triglycerides (TAGs) and glycogen was measured in male flies. Glut1 levels were decreased in these cells by combining the Crz-specific Crz-Gal4 line with two distinct Glut1-RNAi lines (indicated as VDRC & TRiP). TAG and glycogen levels were measured in whole animal extracts and normalized by their respective protein content. The glycogen/protein concentrations were decreased in these Crz-specific Glut1-RNAi flies compared to their respective genetic background controls (Fig. 1B), while there was no change in TAG/protein concentrations (Fig. 1A). This suggests that Glut1 functions specifically in the neurons that produce Crz to regulate organismal glycogen storage.
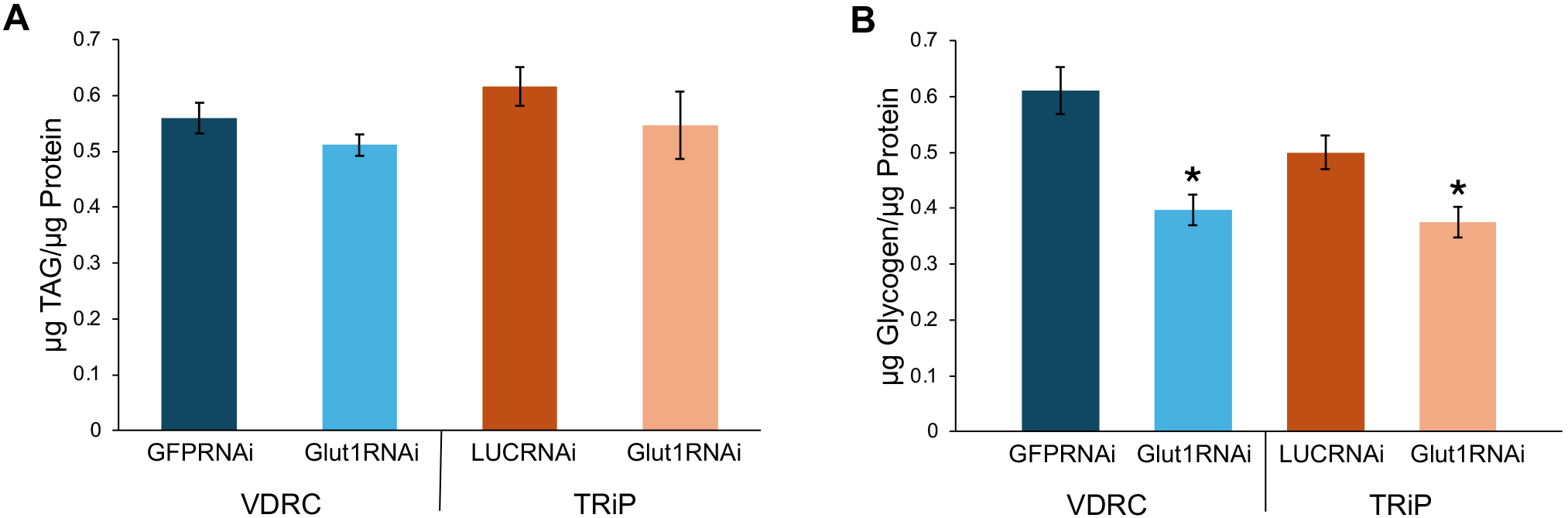 Fig. 1.
Fig. 1.
Targeting RNAi against Glucose transporter 1
(Glut1) in Crz-expressing neurons decreases glycogen storage.
(A) TAGs normalized by protein content were measured in
Crz-Gal4
To further characterize the glycogen storage defect in Crz-specific Glut1-RNAi flies, we measured the expression of two regulators of glycogen metabolism, glycogen phosphorylase (Glyp) and glycogen synthase (Glys), using quantitative PCR. Glyp cleaves the alpha glycosidic linkages between glucose molecules stored in glycogen promoting glycogen breakdown while Glys makes the alpha glycosidic linkages between glucose molecules to synthesize glycogen [13]. While there was no consistent change in Glyp expression in Glut1-RNAi flies (Fig. 2A), Glys levels were increased when Glut1 was decreased in Crz neurons (Fig. 2B). Since glycogen levels were blunted in Crz-specific Glut1-RNAi flies (Fig. 1B), this elevation of Glys expression is likely compensating for the low glycogen storage.
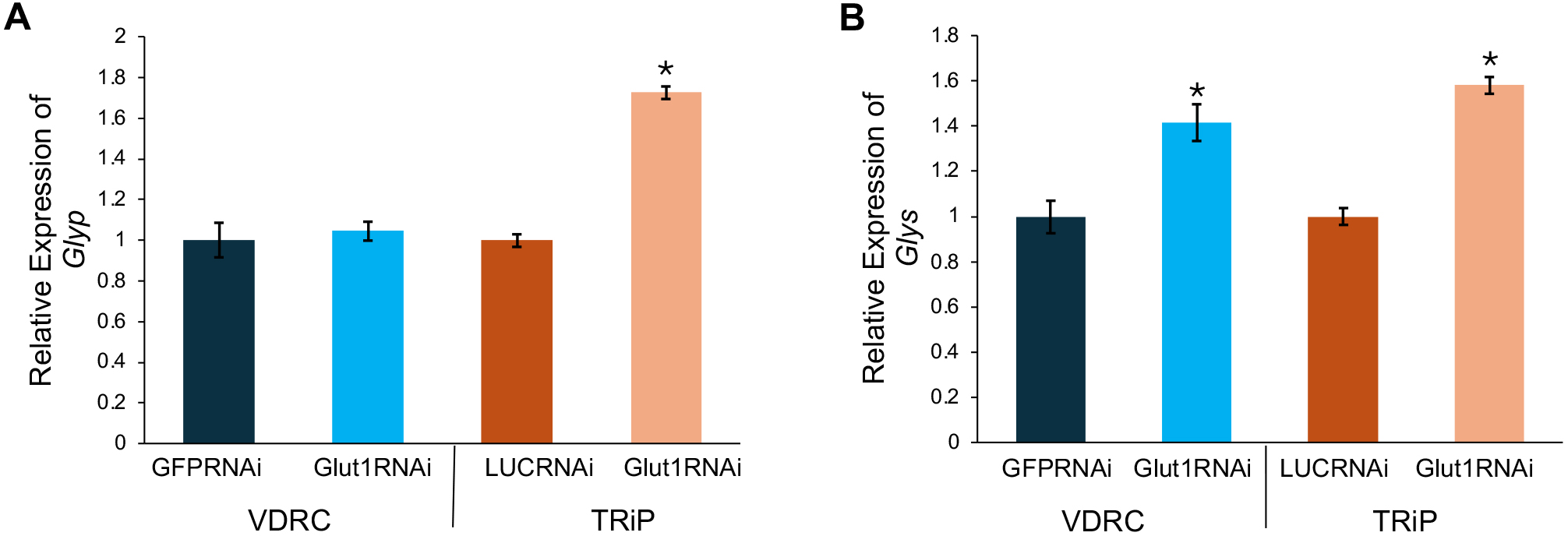 Fig. 2.
Fig. 2.
Targeting RNAi against Glut1 in Crz-expressing
neurons increases glycogen synthase (Glys) expression.
(A) Glycogen phosphorylase (Glyp) transcript levels
normalized by rp49 levels were measured in
Crz-Gal4
A previous study has shown that targeting Crz with RNAi increases glycogen levels [11]; thus, the decrease in glycogen storage shown here could be due to excess Crz upregulation in these neurons in response to low glucose levels when Glut1 was knocked down in these cells. To test this hypothesis, Crz transcript levels were measured in Glut1-RNAi flies using quantitative PCR. There was a nearly 2-fold increase in Crz transcript levels in these Crz-specific Glut1-RNAi flies compared to their respective controls (Fig. 3). This suggests that in response to low glucose levels, Crz-expressing neurons upregulate Crz, leading to a decrease in glycogen storage presumably caused by activated downstream effectors of Crz signaling in target tissues.
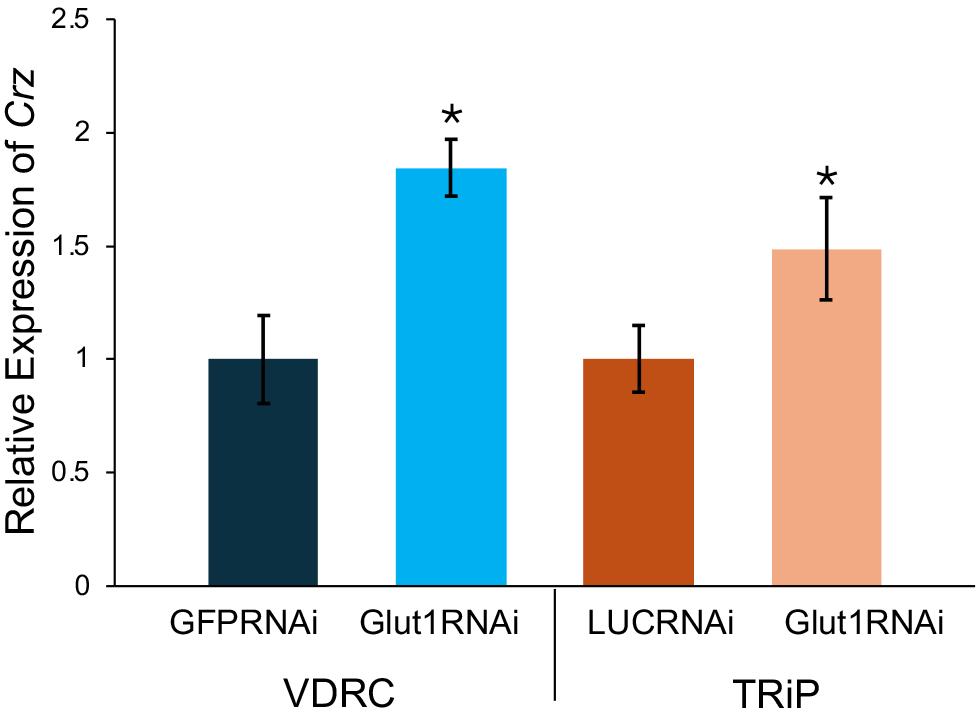 Fig. 3.
Fig. 3.
Targeting RNAi against Glut1 in corazonin
(Crz)-expressing neurons increases Crz transcript
levels. Crz transcript levels normalized by rp49 levels were
measured in Crz-Gal4
Here we show that decreasing Glut1 levels in Crz-expressing neurons in male flies led to a decrease in glycogen storage, but no changes in TAG levels (Fig. 1). This is consistent with a previous study where glycogen levels were increased when RNAi was targeted towards Crz in well-fed male flies [11]. However, in the same study when Crz levels were decreased with RNAi, there was only an increase in TAG levels after a 24-hour starvation period [11], suggesting that Crz has a starvation-specific role in regulating TAG levels; therefore, it is possible that TAG levels could be altered in Crz-specific Glut1-RNAi flies after a 24-hour starvation period. Additionally, the decrease in glycogen storage phenotype shown here matches the phenotype observed when RNAi was targeted against Glut1 in all neurons [9], suggesting that Glut1 may act in Crz neurons to regulate nutrient homeostasis by modulating glycogen metabolism.
Glycogen metabolism is regulated by two major enzymes, Glyp and Glys [13], and we tested whether their expression was altered in Glut1-RNAi flies. Interestingly, Glys transcript levels were elevated when Glut1 was decreased in Crz-producing neurons (Fig. 2B), despite the low glycogen in these flies suggesting a compensatory transcriptional mechanism to regulate glycogen storage. However, we did not measure Glyp or Glys enzyme activity in this study and it is possible that the activity of the glycogen breakdown enzyme Glyp could be increased or the activity of the glycogen synthesis enzyme Glys could be decreased resulting in the low glycogen storage phenotype observed here. Future experiments designed to measure Glyp and Glys enzymatic activity in Glut1-RNAi flies could help clarify the function of the Crz-producing neurons in regulating glycogen metabolism.
We have also shown that decreasing Glut1 levels in Crz-expressing neurons led to an increase in Crz transcript levels (Fig. 3). This suggests that when glucose levels are low, Crz levels increase so that Crz can be secreted to act on Crz receptors (CrzR) expressed on target tissues. However, while we show here that Crz transcript levels are increased in Glut1-RNAi flies, we did not measure Crz secretion directly and future experiments aimed at measuring Crz secretion in these flies and perhaps other Crz-regulated phenotypes would help support this claim. Interestingly, when CrzR levels are decreased in fat body cells, there is an increase in glycogen storage, but no change in circulating glucose levels in well-fed male flies [14]. Therefore, it is possible that when Glut1 levels were decreased in Crz-expressing neurons, the excess Crz likely acted directly on the fat body to promote glycogen breakdown. While little is known about CrzR and its downstream targets in the fat body, multiple studies have characterized the receptor in Drosophila and other insects as a GPCR. Interestingly, CrzR shares a high percentage of amino-acid identity with the adipokinetic hormone receptor (AKHR) [15], which promotes glycogen breakdown in fly fat tissue [16]. Therefore, it is possible that the downstream target effectors of AKHR involved in promoting glycogen breakdown can also be targeted by CrzR to perform that same function. Further investigation is required to identify the downstream targets of CrzR in the fat body to understand the mechanism of how Crz modulates glycogen storage.
Overall, this study has characterized the role of Glut1 in Crz-expressing neurons to regulate Crz expression and overall glucose homeostasis. Crz-producing neurons are part of a growing group of neuronal populations in Drosophila such as the IPCs, adipokinetic hormone-producing neurons and neuropeptide F-producing neurons that respond to nutrients and regulate metabolism [9, 17, 18, 19]. The results described here will help to expand our understanding of the mechanisms whereby the nervous system detects changes in nutrient availability and alters organismal metabolism accordingly.
One of the limitations of this study is that the increases in Glys and Crz reported here are at the RNA level; the production of these proteins was not measured. While in many instances the levels of a gene’s transcript correlate with the protein produced, this is not always the case, so future analysis of Glys protein production and enzyme activity and Crz protein production and secretion will increase our understanding of how Glut1 acts in Crz-expressing neurons to regulate glycogen metabolism.
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Conceptualization, JRD; methodology, JRD; validation, PP; formal analysis, PP; investigation, JRD and PP; resources, JRD; writing—original draft preparation, PP; writing—review and editing, JRD and PP; visualization, PP; supervision, JRD; project administration, JRD; funding acquisition, JRD. Both authors have read and agreed to the final version of the manuscript. Both authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
The authors would like to thank Matthew Kauffman for technical assistance and helpful discussions. Stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) and the Vienna Drosophila Resource Center (VDRC, https://www.vdrc.at) were used in this study.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.



