1 Russian Research Institute of Farm Animal Genetics and Breeding – Branch of the L. K. Ernst Federal Research Centre for Animal Husbandry, Pushkin, 196601 St. Petersburg, Russia
2 North-Western Center for Interdisciplinary Research of Food Security Problems, St. Petersburg Federal Research Center of the Russian Academy of Sciences, Pushkin, 196608 St. Petersburg, Russia
3 School of Natural Sciences, University of Kent, CT2 7NJ Canterbury, UK
4 Animal Genomics and Bioresource Research Unit (AGB Research Unit), Faculty of Science, Kasetsart University, Chatuchak, 10900 Bangkok, Thailand
5 L. K. Ernst Federal Research Center for Animal Husbandry, Dubrovitsy, Podolsk, 142132 Moscow Oblast, Russia
Abstract
Cryopreservation cannot be widely used for rooster sperm due to high incidences of cryoinjury, including damage to sperm membranes. Thus, cryopreserved rooster sperm has limited use due to low sperm motility and reduced fertilizing ability, which disrupts the mechanisms involved in sperm–egg interactions. Previously, we used an Illumina 60K single-nucleotide polymorphism (SNP) array to search for genes associated with rooster sperm quality, before and after freeze–thawing. As a continuation of these genome-wide association studies (GWAS), the present investigation used a denser 600K SNP chip. Consequently, the screen depth was expanded by many markers for cryo-resistance in rooster sperm while more candidate genes were identified. Thus, our study aimed to identify genome-wide associations with ejaculate quality indicators, including those concerning sperm membrane damage.
We selected sperm quality indicators after freezing–thawing using samples from a proprietary cryobank collection created to preserve generative and germ cells of rare and endangered breeds of chickens and other animal species. A total of 258 ejaculates from 96 roosters of 16 different breeds were analyzed. Moreover, 96 respective DNA samples were isolated for genotyping using a 600K Affymetrix® Axiom® high-density genotyping array.
In total, 31 SNPs and 26 candidate genes were associated with characteristics of sperm membrane damage, progressive motility, and sperm cell respiration induction using 2,4-dinitrophenol. In particular, we identified the ENSGALG00000029931 gene as a candidate for progressive motility, PHF14 and ARID1B for damaged sperm membranes, and KDELR3, DDX17, DMD, CDKL5, DGAT2, ST18, FAM150A, DIAPH2, MTMR7, NAV2, RAG2, PDE11A, IFT70A, AGPS, WDFY1, DEPDC5, TSC1, CASZ1, and PLEKHM2 for sperm cell respiration induction.
Our findings provide important information for understanding the genetic basis of sperm membrane integrity and other traits that can potentially compromise the mechanisms involved in sperm–egg interactions. These findings are relevant to the persistence of fertility after thawing previously frozen rooster semen.
Keywords
- frozen–thawed semen
- chicken (Gallus gallus)
- rooster sperm
- genome-wide association studies
- genetic predisposition
- damaged membranes
- impaired fertility
- compromised sperm–egg interaction
- sperm cryostability
Sperm–egg fusion is the initial stage in the formation of a new individual [1, 2, 3]. It is an intricate process that merges the gametes, unites the cytoplasm and fuses the two parental genomes. A crucial step in this process is membrane fusion; however, little is understood about the molecules and mechanisms involved [4]. Previous work suggested that membrane fusion requires a more conserved mechanism and set of molecules than the preceding processes in fertilization such as zona pellucida–sperm contact [5, 6, 7], since fusion exhibits less unique species-specificity. In recent times, numerous individual molecules on the sperm and/or egg have been identified as essential (or near-essential) to the process; however, more information is needed with regard to how they interact [4, 8].
The last several decades have seen a major development in our knowledge of the molecular mechanisms underlying sperm–egg contact in mammalian species [9, 10]. Nevertheless, due to the size of the egg and the challenge of simulating the physiological polyspermy that occurs during normal fertilization, the same degree of knowledge has not been gathered for birds. Because avian oocytes are so much larger than those of mammals, we are unable to observe the sperm–egg contact directly in vitro [11]. Nevertheless, male fertility issues in birds remain a significant factor that has a major impact on reproductive success related to sperm–egg interaction [12, 13].
The use of assisted reproductive technologies is thought to be crucial if the productivity of the poultry industry is to be improved [14, 15, 16]. One of the most important technologies is artificial insemination using frozen–thawed sperm and this has wide applicability to both “backyard farm” and large scale industrial animal and poultry (re)production [17, 18, 19]. The fertilizing potential of rooster sperm depends on the quality and quantity of sperm produced by the testes (e.g., [20, 21, 22, 23]). Furthermore, since each rooster is engaged in mating with several hens, sperm characteristics, and the genetic factors responsible for their formation, can have a large impact on the fertility of the breeding flock [24]. Currently, cryopreservation of reproductive cells is an effective means of preserving male genetic material in a cryobank [25]. On the other hand, technologies for bird sperm cryopreservation require further improvement. This includes work on optimizing cooling and freezing protocols, as well as optimizing the composition of cryoprotectants and semen extenders [26]. Traditional assessment of poultry semen quality and sperm function is mainly based on monitoring motility, viability, sperm concentration, sperm morphology and acrosome integrity [27, 28, 29, 30]. It is well known that cryopreserved rooster sperm has some limitations associated with low sperm motility and reduced fertilizing ability [31]. These include high frequency of cryotrauma, such as sperm membrane damage [32, 33, 34, 35].
Many factors affect sperm during the freezing process. One cause of impairment is oxidative stress, which seems to play a particularly important role [36]. Sperm have limited antioxidant defenses, high levels of polyunsaturated fatty acids, and an inability to synthesize proteins, so they are susceptible to oxidative stress. High levels of reactive oxygen species harm sperm by promoting oxidative damage to sperm lipids, proteins, and DNA, which then leads to infertility [37]. Reactive oxygen species can be considered a trigger for mitochondrial apoptosis. Low levels of reactive oxygen are produced by spermatozoa and are involved in physiological processes such as phosphorylation and hyperactivation of spermatozoa during fertilization. Avian sperm membranes contain high levels of polyunsaturated fatty acids (as compared to mammals), making them vulnerable to lipid peroxidation and reactive oxygen species that are produced by cellular components during freezing [31]. Therefore, an important rationale for further study of genetic features affecting resistance to oxidative stress is the test for sperm response to 2,4-dinitrophenol (2,4-DNP or simply DNP) [38, 39]. DNP is the best-known agent for uncoupling oxidative phosphorylation and turns off adenosine triphosphate (ATP) production through oxidation, causing the cell respiration (CR) rate to increase. If the respiratory chain is damaged, there is little or no response to DNP. This method has been tested in mammals and revealed a significant correlation of this parameter with fertilizing ability of the sperm [17]. On the other hand, the cryoprotective effect of DNP treatment, in small quantities, is such that can minimize the harmful effects of reactive oxygen species and is aimed at improving the oxidative status of sperm during cryopreservation [40].
The genetic basis of sperm cryodamage and, alternatively, preservation after freezing–thawing in male chickens (Gallus gallus; GGA) is, however, still poorly understood. Genome-wide association studies (GWAS), nonetheless, represent a powerful instrument for identifying loci (genes) associated with traits of interest (e.g., [41, 42, 43]). GWAS can be carried out using both single-nucleotide polymorphism (SNP) chip technology and whole-genome resequencing, which takes into account hundreds of thousands or even millions of SNPs and other polymorphisms in the genome [44, 45].
Previously, using an Illumina 60K chip, we identified some genes associated with sperm motility before and after freezing–thawing [46, 47]. The present study is a more detailed analysis of genomic associations using a higher density chip of 600K SNPs. The aim of the current study was to identify a larger number of genetic markers for cryoresistance in rooster sperm through a more detailed search for related candidate genes. In doing so, the ultimate objective was to glean a greater insight into impaired sperm–egg interaction mechanisms caused by cryopreservation. Specifically, the present investigation aimed to seek genome-wide associations with ejaculate quality and sperm membrane integrity using samples from the inhouse collection of the institutional Cryobank of Generative and Germ Cells from Rare and Endangered Breeds of Chickens and Other Animal Species created at the Russian Research Institute of Farm Animal Genetics and Breeding (RRIFAGB; Pushkin, Russia) [48, 49]. To search for these associations, we selected an appropriate suite of sperm quality and membrane integrity indicators after freezing–thawing. The results of this study will, we expect, provide important information for understanding the genetic factors of fertility preservation/impairment traits after thawing of frozen semen in roosters.
The study was conducted using a total of 96 males from 16 chicken breeds (see the breed list in Table 1) maintained in the RRIFAGB bioresource collection entitled Genetic Collection of Rare and Endangered Breeds of Chickens (GCREBC, Pushkin, Russia) [47, 50, 51, 52], which is a part of the national Network Bioresource Collection of Farm Animals, Birds, Fish and Insects [53].
| Breed | N | FPM, % | FDM, % | CR |
| Pushkin | 6 | 18.83 |
33.92 |
1.72 |
| Leningrad Golden Gray | 10 | 39.90 |
73.77 |
1.37 |
| Orloff Mille Fleur | 13 | 42.38 |
70.96 |
1.84 |
| Poltava Clay | 4 | 38.75 |
52.13 |
1.60 |
| Pantsirevka Black | 3 | 66.67 |
91.33 |
1.00 |
| Tsarskoye Selo | 14 | 38.50 |
68.82 |
2.29 |
| Australorp Black/Black Speckled | 5 | 32.60 |
15.70 |
1.08 |
| Leghorn Light Brown (or Italian Partridge) | 1 | 55.0 | 91.00 | 1.00 |
| Frizzle | 3 | 35.00 |
87.08 |
1.13 |
| Kuchino Jubilee | 2 | 65.00 |
40.19 |
3.21 |
| Pervomai | 3 | 50.00 |
43.00 |
1.15 |
| Uzbek Game | 5 | 43.00 |
50.74 |
1.26 |
| Russian Crested | 3 | 41.33 |
59.13 |
3.53 |
| Czech Golden | 3 | 20.67 |
87.50 |
2.11 |
| Yurlov Crower Golden/Silver | 5 | 36.60 |
63.80 |
1.16 |
| Minorca Black | 8 | 50.38 |
75.19 |
1.28 |
* N, number of roosters per breed; FPM, forward progressive motility; FDM, frequency of damaged membranes; CR, cell respiration rate determined using dinitrophenol (DNP) induction.
To assess the quality of sperm, roosters of the same age were selected. The keeping, feeding, lighting, temperature conditions and other environmental factors conformed to the regulations for the maintenance of breeder roosters and were not expected to have an effect, since all the birds were housed in the same premises under the same conditions. The roosters chosen for the experimental group were characterized by good health and sperm production quality corresponding to the GCREBC requirements [52, 54], so that no differences would be observed between breeds, although correction for breed affiliation was carried out (see section 2.4). Nevertheless, we assumed that the genetic loci identified in this study did not include markers that depend on breed affiliation and that there was no need to include environmental factors in the analysis, as well. The selected animals were kept in individual cages, and the roosters’ peritoneum was regularly massaged to stimulate sperm production. To obtain sperm samples suitable for evaluation and examination, a preliminary control of the sperm quality and the health condition of the birds was carried out to meet the data homogeneity assumption. This was possible because the semen quality before freezing was adequate (as roosters were selected that produced sperm that met the standards for freezing) and there were no differences between breeds. In particular, sperm quality requirements in the preliminary control analysis were as follows: minimum volume of 0.2 mL, concentration not less than 2 billion/mL, total motility not less than 70%, and progressive motility not less than 50%. The acceptable health conditions were: well-developed red comb, good plumage, legs corresponded to the breed standard, general health factors in accordance with standards for keeping roosters, and assessment of the general health condition that was determined by a veterinarian. The appropriate biological material (i.e., semen and blood) collected from experimental groups of roosters was used for the subsequent analyses.
To evaluate potentially impaired fertility traits, a total of 258 ejaculates were obtained from 96 roosters that had been also subject to genotyping. Diluted semen samples were equilibrated from 18 °C to 5 °C for 40 min. After cooling, dimethylacetamide (DMA; Sigma Aldrich, St. Louis, MO, USA) was added to each sample at a final concentration of 6%. After adding DMA, the samples were incubated at 5 °C for 1 min. The ejaculates were frozen into pellets by directly dripping the semen into, and stored in, liquid nitrogen at –196 ℃. The Leningrad Cryoprotective Medium [55] was used for semen freezing that had the following components supplied by Dia-m (Moscow, Russia) (per 100 mL of distilled water): monosodium glutamate, 1.92 g (114 mM); fructose, 0.8 g (44 mM); potassium acetate, 0.5 g (51 mM); polyvinylpyrrolidone, 0.3 g (8.3 µM); protamine sulfate, 0.032 g (3.27 µM) [56]. Because not all roosters produced sufficient semen volume to be examined, suitable samples from 88 roosters (Table 1) remained in the analysis after freezing, storage and thawing. These sperm samples were assessed for forward progressive motility (FPM), frequency of damaged membranes (FDM) index, and sperm CR rate after post-frozen DNP (Dia-m) treatment as described elsewhere [17, 57].
Sperm FPM was explored using a computer-assisted sperm analysis (CASA, ArgusSoft
Ltd., St. Petersburg, Russia; [58]). CASA settings were properly configured to
identify rooster sperm; the respective identification mode was preliminarily
calibrated using several relevant videos and fixed in the program settings. The
membranes were assessed for integrity vs. damage using the Viability module of
the CASA software. Damage to the plasma membranes of spermatozoa was detected
using the Sperm VitalStain dye (Nidacon International AB, Mölndal, Sweden)
[57]. Staining was carried out in Eppendorf tubes (50 µL sperm mixed with
50 µL dye) and smears were then made on glass slides. The preparations were
viewed at
Respiratory activity of the sperm cells was evaluated using the DNP test substance [38], with the final concentration of 0.025 mM/L DNP [17]. The appropriate procedure included use of the Expert-001MTX ion meter and a Clarke electrode (Econix-Expert Research and Production Company, Moscow, Russia) to measure the CR rate [17, 59]. After adding 1 mL of 11% lactose and 100 µL semen to the chamber, the rate at which the oxygen content decreased was recorded; 10 µL DNP was then added. The CR rate in response to DNP divided by the CR rate without DNP was calculated as a measure of CR induction and its disruption in the frozen–thawed sperm samples. The resultant ratio was equal to one (1) or more and was considered a respiratory response to the addition of DNP [17]. Herewith, the value of this indicator equal to 1 characterized the absence of CR induction due to the destructive (suppressing) effect of freezing–thawing on the respiratory chain of the sperm cells. The higher this indicator was, the less the process of oxidative phosphorylation in sperm experienced a destructive (suppressing) effect due to freezing–thawing.
Sperm quality parameters were estimated within breeds and SNP genotypes as means
(
Blood samples from experimental roosters were taken from a wing vein. For
genome-wide genotyping, a total of 96 DNA samples were isolated. A high density
SNP chip for chicken, i.e., the 600K Affymetrix®
Axiom® (Thermo Fisher Scientific Inc., Waltham, MA, USA)
genotyping array [61], was employed for genotyping. All samples passed quality
control and were suitable for further research. In particular, DNA samples with
the genotyping quality conforming to more than 95% SNP loci were selected. As a
result, a total of 86 samples fit the required genotyping quality and were thus
included in the examined dataset subject to the GWAS analysis. The GWAS was
carried out on all chromosomes using the PLINK 1.9 program (Purcell Lab, Brigham
& Women’s Hospital, Boston, MA, USA) [62] and Ensembl chicken (red jungle fowl)
genome assembly GRCg6a [63, 64]. The following parameter settings for filtering
SNPs suitable for the further analysis were used: minor allele frequency (MAF)
At this stage, the respective calculations were carried out using the EMMAX software [41]. The GWAS procedure in EMMAX was performed using identity-by-state (IBS) kinship matrices to search for the influence of genetic factors on cryostability of rooster semen using the appropriate EMMAX in-built statistical tests to confirm the respective association p-values. The standard models for GWAS implemented in the EMMAX program, including usage of relative trait values (percentages), were used that are widely employed in similar EMMAX-assisted GWAS experiments (e.g., [66, 67]). Because roosters of various breeds were included in the analysis (Table 1), we implemented a breed-adjusted GWAS in the EMMAX model. Accordingly, the following mathematical model was used to calculate the effect of SNPs on sperm motility:
in which Y is a vector of phenotypes, b is the effect of
n SNPs, X is a matrix of SNP genotypes, u is a vector
of additive genetic effects considered to follow a normal distribution with a
mean equal to zero and (co)variance
Next, Manhattan plots were generated using the GWAS results and the qqman
package in R software (version 4.3.2; The R Project for Statistical Computing, R
Foundation for Statistical Computing, Institute for Statistics and Mathematics,
Vienna University of Economics and Business, Vienna, Austria) [68]. The resultant
significant and suggestive SNPs were localized, with significant and suggestive
levels for an SNP effect being set as Bonferroni correction to exclude false
positive results at p = 1.031694
We examined a few characteristics of sperm quality (Table 1) that represent the most important reproductive traits when breeding poultry in small populations and/or preserving the gene pool [70, 71, 72, 73]. A total of 31 quantitative trait nucleotides (QTNs) with significant effects on frozen–thawed sperm quality traits were detected on chicken chromosomes GGA1, GGA2, GGA3, GGA4, GGA5, GGA7, GGA9, GGA15, GGA17, and GGA21 as will be detailed below.
In Table 1, the measured breed-specific values are shown for the indices of
sperm FPM, FDM, and CR rate after DNP treatment post freezing–thawing.
Evaluation of the rooster sperm quality showed high individual variability for
all three main studied indicators. In addition, the volume of an individual
ejaculate varied from 0.4 to 1.4 mL, cell motility before freezing from 20 to
95%, and after freezing from 5 to 80%. The amount of membrane damages depended
significantly on a particular animal and ranged from 3 to 96%. The magnitude of
sperm CR induction in roosters by DNP essentially ranged from 1.00 to 8.09, with
an average of 1.66
One potential SNP, AX-75329814, associated with FPM was identified on GGA1 and
is presented in Table 2 and Fig. 1A and Supplementary Fig. 1A. This
suggestive SNP (p = 1.54
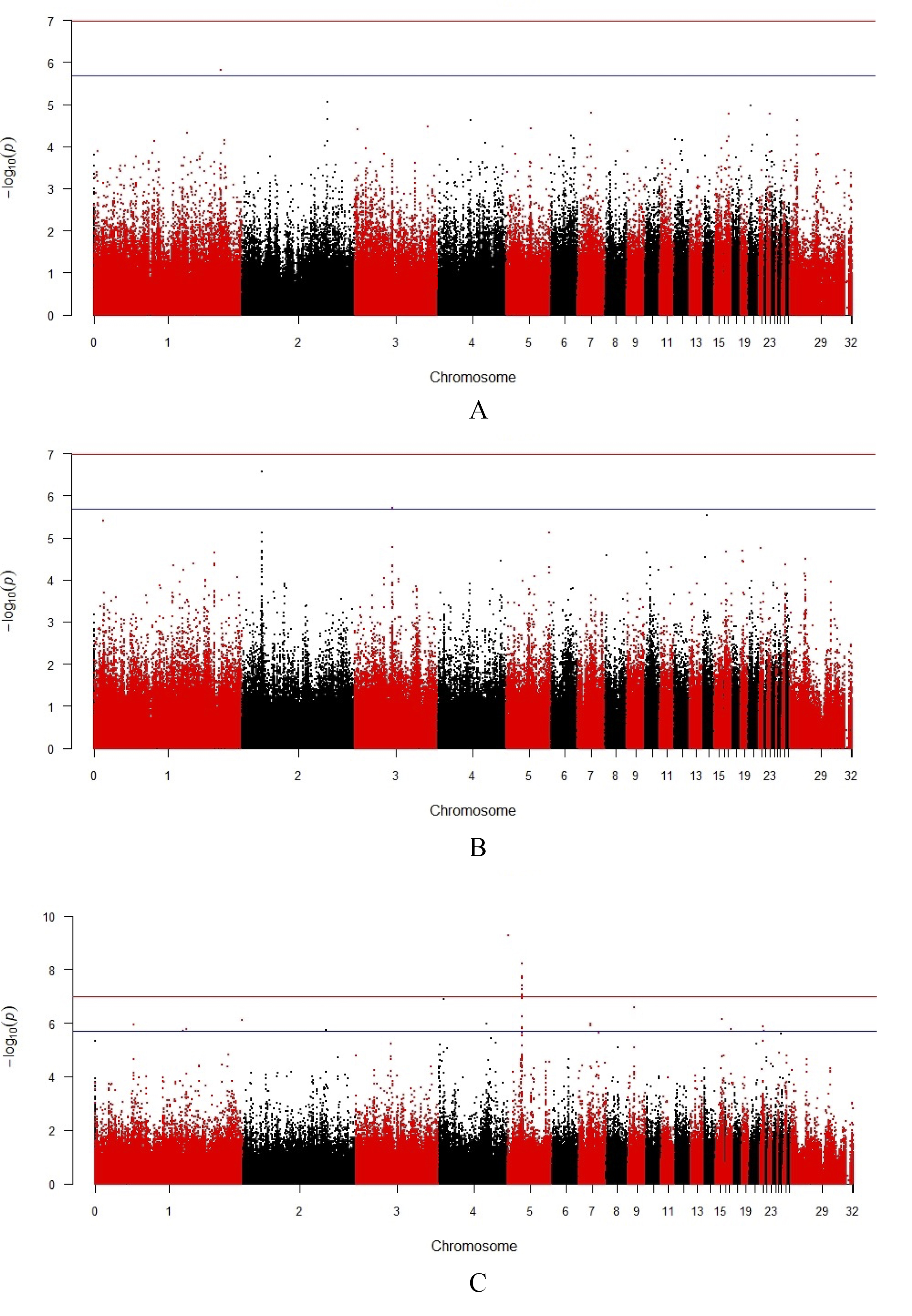 Fig. 1.
Fig. 1.
Manhattan plots of the GWAS results for the studied sperm
quality traits. (A) forward progressive motility, (B) frequency of damaged
membranes, and (C) sperm cell respiration rate. The analysis displays the
distribution of single trait SNPs in rooster chromosomes in relation to the
thresholds for the genome-wide nominal significance level (–log10
(p)), based on estimated probability values of p
| SNP position (bp) | p-value | Alleles | SNP location | Candidate genes | Genotype | n* | FPM |
| 168,850,183 | 1.54 × 10–6 | C/T | ENSGALG00000029931 exons | lncRNA** ENSGALG00000029931 | CC | 50 | 33.70 |
| CT | 30 | 47.00 | |||||
| TT | 6 | 65.50 |
* n, number of roosters per genotype. ** lncRNA, long non-coding RNA.
a,b,c Pairwise significance of difference (a–b, b–c, and a–c) for FPM
genotypes at p
Noteworthy SNPs associated with FDM are presented in Table 3 and Fig. 1B and
Supplementary Fig. 1B. Two suggestive SNPs were found on GGA2 and GGA3.
One of these SNPs, AX-76063628 (p = 2.61
| SNP | Chromosome | SNP position (bp) | p-value | Alleles | SNP location | Candidate genes | Genotype | n* | FDM |
| AX-76 063628 | GGA2 | 26,182,792 | 2.61 × 10–7 | T/C | intron | PHF14 | CC | 23 | 26.74 |
| CT | 39 | 74.07 | |||||||
| TT | 23 | 78.36 | |||||||
| AX-76 495998 | GGA3 | 51,262,693 | 1.93 × 10–6 | A/G | intron | ARID1B | AA | 42 | 78.40 |
| AG | 31 | 54.91 | |||||||
| GG | 12 | 36.83 |
* n, number of roosters per genotype. a,b,c Pairwise significance
of difference (a–b, b–c, and a–c) for FDM genotypes at p
The results of the performed GWAS for the index of sperm CR induction in
roosters using DNP is shown in Table 4 and Fig. 1C and Supplementary Fig.
1C. For example, one suggestive SNP AX-80992139 (p = 1.89
| SNP | Chromosome | SNP position (bp) | p-value | Alleles | SNP location | Candidate genes | Genotype | n* | CR |
| AX-75466971 | GGA1 | 50,898,160 | 1.14 × 10–6 | C/T | KDELR3 intron, downstream DDX17 | KDELR3, DDX17 | GG | 4 | 4.85 |
| GT | 14 | 2.38 | |||||||
| TT | 67 | 1.38 | |||||||
| AX-75221789 | GGA1 | 116,157,001 | 1.94 × 10–6 | G/A | ENSGALG00000056838 intron, downstream DMD | lncRNA** ENSGALG00000056838, DMD | AA | 73 | 1.49 |
| AG | 11 | 2.28 | |||||||
| GG | 2 | 6.44 | |||||||
| AX-75231769 | GGA1 | 122,024,645 | 1.77 × 10–6 | C/T | intron | CDKL5 | CC | 1 | 6.75 |
| CT | 15 | 2.71 | |||||||
| TT | 69 | 1.42 | |||||||
| AX-75397985 | GGA1 | 196,966,714 | 7.73 × 10–7 | C/T | downstream ENSGALG00000064293, upstream DGAT2 | lncRNA ENSGALG00000064293, DGAT2 | AA | 65 | 1.36 |
| AC | 16 | 2.29 | |||||||
| CC | 4 | 4.85 | |||||||
| AX-80992139 | GGA2 | 109,830,505 | 1.89 × 10–6 | A/T | intergenic (between ST18 and FAM150A) | ST18, FAM150A (ALKAL1) | CC | 74 | 1.47 |
| CT | 8 | 2.87 | |||||||
| TT | 3 | 4.47 | |||||||
| AX-80778510 | GGA4 | 5,664,389 | 1.31 × 10–7 | T/A | intergenic (between ENSGALG00000068051 and DIAPH2) | lncRNA ENSGALG00000068051, DIAPH2 | CC | 57 | 1.31 |
| CT | 21 | 2.06 | |||||||
| TT | 8 | 3.57 | |||||||
| AX-76705102 | GGA4 | 63,101,468 | 1.05 × 10–6 | C/T | intron | MTMR7 | CC | 4 | 4.85 |
| CT | 13 | 2.49 | |||||||
| TT | 68 | 1.38 | |||||||
| AX-76788932 | GGA5 | 1,970,758 | 5.26 × 10–10 | C/T | intron | NAV2 | CC | 78 | 1.49 |
| CT | 6 | 3.55 | |||||||
| TT | 2 | 4.55 | |||||||
| AX-76791439 | GGA5 | 19,979,152 | 1.41 × 10–6 | G/A | ENSGALG00000066890 intron, upstream RAG2 | lncRNA ENSGALG00000066890, RAG2 | AA | 5 | 5.17 |
| AG | 26 | 1.71 | |||||||
| GG | 55 | 1.38 | |||||||
| AX-76791443 | GGA5 | 19,981,011 | 5.49 × 10–7 | G/A | same as above | same as above | AA | 4 | 4.85 |
| AC | 14 | 2.38 | |||||||
| CC | 68 | 1.38 | |||||||
| AX-76791454 | GGA5 | 19,985,728 | 1.87 × 10–8 | G/A | same as above | same as above | AA | 77 | 1.47 |
| AT | 9 | 3.66 | |||||||
| TT | 0 | – | |||||||
| AX-76791501 | GGA5 | 20,013,308 | 1.63 × 10–6 | C/T | same as above | same as above | AA | 76 | 1.43 |
| AG | 9 | 3.65 | |||||||
| GG | 0 | – | |||||||
| AX-76791503 | GGA5 | 20,014,711 | 1.18 × 10–7 | C/T | same as above | same as above | AA | 4 | 4.85 |
| AG | 15 | 2.37 | |||||||
| GG | 66 | 1.37 | |||||||
| AX-76791535 | GGA5 | 20,030,837 | 1.83 × 10–8 | C/A | same as above | same as above | CC | 76 | 1.47 |
| CT | 8 | 3.22 | |||||||
| TT | 2 | 4.55 | |||||||
| AX-76791553 | GGA5 | 20,037,753 | 9.94 × 10–8 | G/T | same as above | same as above | AA | 3 | 4.88 |
| AG | 13 | 2.48 | |||||||
| GG | 70 | 1.42 | |||||||
| AX-76791570 | GGA5 | 20,046,742 | 4.04 × 10–8 | A/C | same as above | same as above | CC | 76 | 1.50 |
| CT | 9 | 2.67 | |||||||
| TT | 1 | 8.09 | |||||||
| AX-76791596 | GGA5 | 20,057,754 | 6.20 × 10–9 | T/C | same as above | same as above | AA | 75 | 1.50 |
| AG | 10 | 2.57 | |||||||
| GG | 1 | 8.09 | |||||||
| AX-76791611 | GGA5 | 20,066,805 | 1.83 × 10–8 | G/T | same as above | same as above | CC | 68 | 1.44 |
| CT | 15 | 2.14 | |||||||
| TT | 3 | 5.59 | |||||||
| AX-76791629 | GGA5 | 20,079,418 | 2.11 × 10–8 | T/G | same as above | same as above | AA | 63 | 1.36 |
| AT | 18 | 2.10 | |||||||
| TT | 5 | 4.59 | |||||||
| AX-76791640 | GGA5 | 20,085,610 | 8.84 × 10–8 | C/A | ENSGALG00000066890 exon, upstream RAG2 | lncRNA ENSGALG00000066890, RAG2 | AA | 5 | 3.99 |
| AG | 14 | 2.38 | |||||||
| GG | 67 | 1.39 | |||||||
| AX-76791651 | GGA5 | 20,089,359 | 5.43 × 10–8 | C/T | downstream ENSGALG00000066890, upstream RAG2 | lncRNA ENSGALG00000066890, RAG2 | CC | 70 | 1.36 |
| CT | 15 | 2.88 | |||||||
| TT | 1 | 8.09 | |||||||
| AX-76986124 | GGA7 | 15,619,919 | 1.25 × 10–6 | G/A | intron | PDE11A | AA | 4 | 4.85 |
| AC | 18 | 2.22 | |||||||
| CC | 64 | 1.36 | |||||||
| AX-76986304 | GGA7 | 15,687,930 | 1.05 × 10–6 | A/G | downstream IFT70A, intergenic (between IFT70A and AGPS) | IFT70A (TTC30B), AGPS | GG | 4 | 4.85 |
| GT | 14 | 2.38 | |||||||
| TT | 68 | 1.38 | |||||||
| AX-77181439 | GGA9 | 8,463,525 | 2.68 × 10–7 | T/C | intron | WDFY1 | CC | 58 | 1.30 |
| CT | 20 | 2.12 | |||||||
| TT | 7 | 3.93 | |||||||
| AX-75848147 | GGA15 | 9,143,016 | 7.44 × 10–7 | T/C | intron | DEPDC5 | GG | 8 | 4.45 |
| GT | 25 | 1.60 | |||||||
| TT | 53 | 1.34 | |||||||
| AX-75873724 | GGA17 | 7,048,201 | 1.67 × 10–6 | C/T | intron | TSC1 | CC | 0 | – |
| CT | 12 | 3.47 | |||||||
| TT | 74 | 1.42 | |||||||
| AX-76244713 | GGA21 | 3,983,187 | 1.30 × 10–6 | A/G | upstream CASZ1 | CASZ1 | CC | 67 | 1.38 |
| CT | 15 | 2.29 | |||||||
| TT | 4 | 4.85 | |||||||
| AX-76245698 | GGA21 | 4,201,372 | 1.97 × 10–6 | C/T | intron | PLEKHM2 | CC | 1 | 8.09 |
| CT | 20 | 2.39 | |||||||
| TT | 65 | 1.39 |
* n, number of roosters per genotype. ** lncRNA, long non-coding RNA. a,b,c Pairwise significance
of difference (a–b, a–c and b–c) for FDM genotypes at p
One significant SNP, AX-76788932 (p = 5.26
In addition, a number of suggestive SNPs were found on GGA1, GGA7, GGA9, GGA15, GGA17, and GGA21, for which the following candidate genes were proposed: KDELR3 (KDEL endoplasmic reticulum protein retention receptor 3), DDX17 (DEAD-box helicase 17), DMD (dystrophin), CDKL5 (cyclin dependent kinase like 5), DGAT2 (diacylglycerol O-acyltransferase 2), PDE11A (phosphodiesterase 11A), IFT70A (intraflagellar transport 70A), AGPS (alkylglycerone phosphate synthase), WDFY1 (WD repeat and FYVE domain containing 1), DEPDC5 (DEP domain containing 5), TSC1 (TSC complex subunit 1), CASZ1 (Castor zinc finger 1), and PLEKHM2 (pleckstrin homology and RUN domain containing M2).
Genetic and genomic associations relative to reproductive traits and issues are pivotal for progress in animal and poultry production and reproduction [77, 78, 79, 80, 81, 82, 83]. Semen cryobanks play an important role in conserving the genetic resources of breeds and species of animals, including birds. Sperm properties are indicators of the reproductive qualities of roosters, and the appropriate freezing technology is the basis for preserving sperm suitable for further effective fertilization [84, 85, 86, 87]. It is well known that use of cryopreserved rooster sperm has some limitations, significantly associated with low sperm motility and reduced fertilizing ability, due to disruptions in the mechanisms of sperm–egg interaction (e.g., [88, 89, 90]). Cryopreservation techniques in birds have not yet found widespread use due to the high incidence of cryotrauma, including damage to sperm membranes (e.g., [91, 92, 93, 94]). In the present GWAS study, novel phenotypic traits to search for genetic factors influencing the fertilization process in birds after semen freezing–thawing have been used, as discussed below.
FPM estimate is the most widely used indicator for sperm quality assessment
[56, 95, 96]. Here, rooster sperm quality evaluation resulted in high individual
variability in FPM: from 20 to 95% before freezing and from 5 to 80% after
freezing (data not shown). Previously, we [46] also noted significant variation
in this indicator before freezing (40–95%) and after thawing (0–75%), with
its mean values being 87.9
One suggestive SNP, AX-75329814, associated with the FPM index was found on GGA1
(Table 2 and Fig. 1A and Supplementary Fig. 1A). This was an exonic
polymorphic variant of the ENSGALG00000029931 lncRNA gene. It is known
that lncRNAs have lengths more than 200 nucleotides and are incapable of encoding
protein. They are found in the cytoplasm and nucleus and are transcribed by RNA
polymerase II [98, 99, 100]. There is a connection between male fertility and
noncoding RNA markers. As reviewed by Hitit et al. [100], lncRNAs play
critical functions in controlling a wide range of biological processes that are
strongly expressed in mature sperm and the testes of mice and bulls. They are
expressed differently in human and bovine motility phenotypes, suggesting that
sperm lncRNAs may have functional involvement in fertility. Sperm bearing RNA was
discovered to be conversed among various species, including stallions, goats,
boars, and rams [100]. Involvement of lncRNAs in the formation of male germ cells
and sperm motility has been demonstrated in chickens and turkeys [101, 102, 103, 104]. Our
analysis of the corresponding FPM phenotypes revealed significant differences
between the ENSGALG00000029931 lncRNA genotypes (p
An important indicator of semen viability after freezing–thawing is the degree of damage to the sperm plasma membranes [105, 106]. Membrane rafts are present on the surface of sperm and play an important role in the mechanisms of sperm–egg interaction in the fertilization process [35]. The freezing process significantly deteriorates the condition of the sperm plasma membranes. In the work by Olexikova et al. [107], only about a quarter of the spermatozoa assessed belonged to quality class I. In our investigation, the individual FDM values in the studied group of roosters ranged from 3 to 96% (data not shown).
Suggestive SNPs associated with the FDM index were found on GGA2 and GGA3 (Table 3 and Fig. 1B and Supplementary Fig. 1B). The candidate PHF14 gene located on GGA2 affects cell proliferation. In our study, genotype analysis for FDM revealed a significant negative effect of the T allele in the PHF14 gene on the integrity of sperm membranes during cryopreservation. As reported by Huang et al. [108], knockout of the PHF14 gene in mice led to neonatal lethality due to respiratory failure. Pathological analysis revealed severe disturbances in tissue and cellular structures. PHF14 may be an epigenetic regulator and play an important role in the development of many organs. PHF14 also reduced the expression of the Cdkn1a gene, which in turn controlled the expression level of the H3K4me3 gene. PHF14 plays an important role in the formation of the germinal center by regulating the proliferation of germinal center B cells in the spleen [108, 109]. In a human study for the MeCP2-TCF20 complex that plays a critical role in brain function, a missense mutation in the PHF14 subunit of the TCF20 complex causes neurological symptoms due to the abolition of the MeCP2-PHF14-TCF20 interaction [110]. The putative candidate gene ARID1B located on GGA3 is a chromatin regulator, and mutations in this gene cause neurodevelopmental disorders in humans [111, 112].
A significant number of factors influence cryodamage of spermatozoa, with an important role among these factors being played by oxidative stress. Biochemical processes that use ATP to generate energy, such as glycolysis and oxidative phosphorylation, play a major role in the essential activity of cells. The intensity of energy metabolism is a very important criterion to assess the sperm quality. A crucial justification for studying this sperm parameter is its connection with fertilizing ability. Oxidative phosphorylation prevails over glycolysis and is the main source of ATP synthesis. Therefore, disruption of the oxidative phosphorylation process during freezing–thawing of spermatozoa can be the main reason for the decrease in their fertilizing ability.
Previously, we showed the identity of responses to DNP in terms of changes in sperm CR in boars, bulls, stallions, reindeer and chickens [17]. The only difference across different animal species is the strength of the response to the test substance. After freezing and thawing, sperm CR induction using DNP declined. It was found that the sperm CR rate correlates with the pregnancy rate after fertilization [17]. Thus, the test for an increased CR rate after the DNP addition can be a good test for the sperm fertilizing ability. That is why we also chose this indicator of sperm quality after freezing–thawing to search for genome-wide associations.
The sperm CR induction variability in roosters was high in the present experiment (Table 1). The coefficient of variation for FPM ranged from 0.5 to 19%, for FDM from 0.1 to 40%, and for CR from 0.02 to 4.4%. High reproducibility of DNP sperm CR induction in roosters across samples suggests the presence of certain genetic factors. Their identification can serve as a biomarker for the selection of animals whose sperm retains high fertilizing ability after freezing–thawing. As a result of GWAS for such factors, five suggestive SNPs were found on GGA1 (Table 4) and the following candidate genes were proposed: KDELR3, DDX17, DMD, CDKL5, and DGAT2. KDELR3 is responsible for the regulation of cellular processes; in studies on cows, the KDELR3 gene expression was reduced in normally fat cows compared to excessively fat ones [113]. The adjacent DDX17 gene encoding the RNA helicase binds and hydrolyzes ATP, altering RNA structures that affect steroid hormone signaling, including phosphorylation of estrogen receptor 1 (ESR1). Gene expression dynamics during and after meiosis demonstrated that DDX17 transcript levels grow, while most transcripts decrease or remain stable, suggesting a role for DDX17 in subsequent sperm functions, i.e., egg fertilization [114].
The candidate DMD gene is expressed in mitochondria. Its encoded protein forms a component of the dystrophin-glycoprotein (DGC) complex that connects the internal cytoskeleton and extracellular matrix. Although mdx/utrn-/- mice lacking both dystrophin and its autosomal homolog murine utrophin are infertile, the mechanisms underlying this phenomenon remain unclear [115]. One more discovered candidate gene, CDKL5, encodes a phosphorylated protein with protein kinase activity and binds ATP and nucleotides. CDKL5 kinase activity is important for the suppression of gene transcription induced by double-strand DNA breaks [116]. The DGAT2 gene encodes one of two enzymes that catalyze the final reaction of triglyceride synthesis. Accordingly, it can affect glycerol metabolism, lipid biosynthesis, and lipid metabolism. The methylation level of the DGAT2 gene in sperm DNA is critical for fertility and viability of offspring [42].
On GGA4, one significant (AX-80778510, p = 1.31
One of the two candidate genes on GGA5 that we identified was the NAV2 gene. This gene encodes a member of the neuronal navigator gene family, which may play a role in cell growth and migration [119], although its role remains largely unknown. Next, a large region containing a cluster of significant and putatively significant SNPs was found on GGA5. They were located in the vicinity of the RAG2 gene. A protein that is produced by this gene plays a role in the development of B- and T-cells [120]. Mutations in this gene lead to immunodeficiency diseases, and RAG2 knockout is used in genome editing during transplantation [121, 122]. Previously, other researchers discovered in human sperm cells that RAG1 and RAG2 are required for immunoglobulin G (IgG) synthesis. Anti-IgG antibodies were found to be able to prevent sperm penetration into zona pellucida-negative hamster eggs. It is suggested that IgG can be produced by human sperm and may play a role during fertilization [123]. RAG2 has also been suggested to be involved in pre-B cell decisions between apoptosis vs. DNA repair and cell cycle progression after DNA damage [124]. We speculate that this gene’s polymorphisms are linked to fertilization selectivity and that the gene’s efficiency is related to ATP-dependent remodeling of ATPase subunits. The significant SNPs found (Table 4) confirm the association of RAG2 polymorphic variants with the sperm response to DNP-mediated blockade of ATP production.
Two significant SNPs were found on GGA7. One of the respective candidate genes was PDE11A. By controlling the intracellular concentration of the cyclic nucleotides, i.e., cyclic adenosine monophosphate (cAMP) and cyclic guanosine monophosphate (cGMP), phosphodiesterase 11A, which is expressed by this gene, contributes to signal transduction. It channelizes the hydrolysis of cAMP and cGMP to 5′-AMP and 5′-GMP, respectively [125]. All these facts support the possible influence of polymorphic variants of the PDE11A gene on the stability of cellular processes in frozen-thawed semen. To increase sperm viability, various substances that interact with phosphodiesterase are used, e.g., thiophylline and IBMX (3-isobutyl-1-methylxanthine) [126, 127, 128]. The study of the effect of stimulants that improve semen quality on various PDE11A genotypes has the potential for practical use to increase the fertilizing ability of sperm after freezing–thawing and to form the composition of sperm extenders. Another gene, IFT70A (or TTC30B, i.e., tetratricopeptide repeat domain 30B), encodes a protein involved in the intraflagellar transport machinery and ciliogenesis [129]. Herewith, sperm tail production is impacted by genetic abnormalities in motile cilia that causes male infertility [130]. Another potential candidate on GGA7, AGPS, is known to cause male sterility when mutated in mice [131].
A candidate gene WDFY1 found on GGA9 encodes the phosphatidylinositol-3-phosphate-binding protein required for spermatogenesis and male fertility, particularly in mice [132]. The candidate gene DEPDC5 that is located on GGA15, with a molecular function to activate GTPase, was identified as influencing male infertility in yak-cow hybrids (cattleyaks) [133]. The TSC1 gene located on GGA17 controls cell growth, proliferation, and size [134, 135]. In a complex with TSC complex subunit 2 (TSC2) in mammals, it inhibits the phosphorylation of S6K1 and EIF4EBP1 by regulating signaling to the mechanistic target of rapamycin complex 1 (mTORC1) [136]. In particular, this candidate gene in mice positively plays an inhibitory role in the functioning of mTORC1, a central regulator of cell growth, proliferation, and differentiation, including in the process of the proliferation and differentiation of spermatogonia. In the TSC1 knockout mice, there were abnormalities in testicular development, partial spermatogenic arrest, reduced sperm count, excessive loss of germ cells, and subfertility [137]. Another important candidate gene on this chromosome is PLEKHM2 encoding a kinesin-related protein that is putatively associated with male factor infertility [138].
In addition, we identified several lncRNAs (ENSGALG00000056838, ENSGALG00000064293, ENSGALG00000068051, and ENSGALG00000066890), but their possible roles in spermatogenesis and egg fertilization activity require further study. The distinct ncRNAs found in male sperm that consistently exhibit fertility phenotypes have the potential to be effective fertility biomarkers [98, 100].
Thus, the present study delivers a theoretical basis for understanding the factors regulating the fertilizing ability of rooster sperm after freeze–thaw and provides further insights into the molecular mechanisms for semen selection during cryopreservation. In Fig. 2 (Ref. [37, 139]), we provided an overview of a number of positional candidate genes identified in our study, alongside their biological influence on functional molecular mechanisms that may contribute to the fertilization process (i.e., sperm–egg fusion [140]). Many of the discovered candidate genes are involved in the mechanisms of interaction of the Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathways in preparation for egg fertilization (Fig. 2).
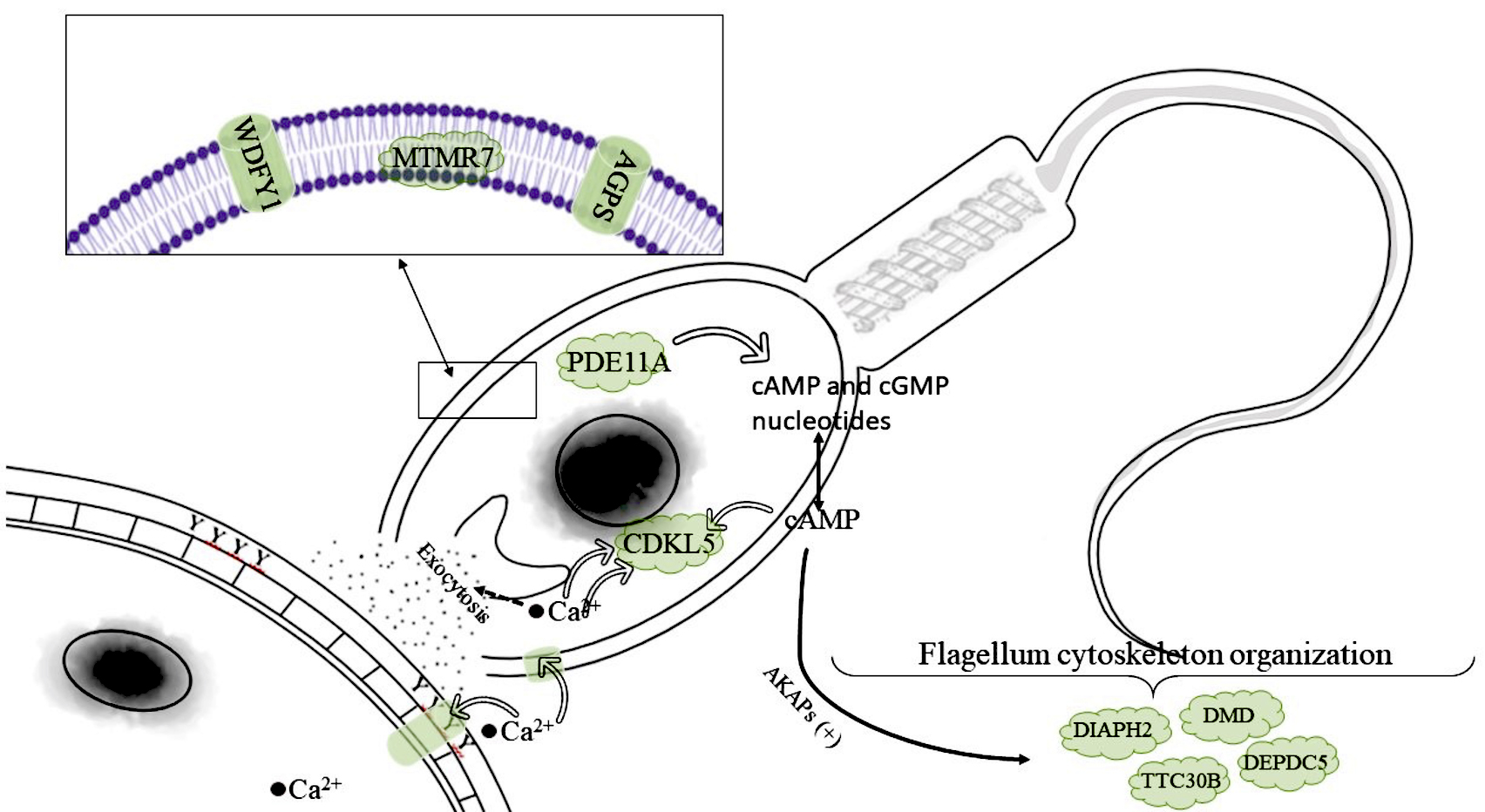 Fig. 2.
Fig. 2.
Mechanisms of interaction of KEGG signaling pathways in preparation for egg fertilization based on positional candidate genes identified by GWAS for sperm parameters post freeze–thaw. Such genes as MTMR7, WDFY1, and AGPS encode structural membrane components and are elements of intracellular signaling pathways, mediating membrane trafficking and autophagy. One of the main signaling pathways involved in sperm motility is the cAMP-dependent pathway. The PDE11A gene metabolites play a role in signal transduction by regulating the intracellular concentration of cyclic nucleotides cAMP and cGMP that act as second messengers, including indirect effects on tyrosine phosphorylation proteins (A-kinase anchor proteins, or AKAPs). AKAPs are a group of structurally diverse scaffolding proteins that bind to protein kinase A. Their interaction affects the genes DMD, DIAPH2, TTC30B (IFT70A), and DEPDC5, causing a reorganization of the sperm cytoskeleton during capacitation [37, 139]. This presumably leads to changes in flagellar curvature that modify the amplitude of the tail wave, which directly affects sperm fertility. The activity of the protein kinase encoded by the CDKL5 gene is regulated by several events (e.g., DNA damage), along with chemical signals in the cAMP-dependent pathway, including cAMP, cGMP and Ca2+ ions, which trigger exocytosis and serve as transmitters of various cellular signals, thereby being sensors of this process. WDFY1, WD repeat and FYVE domain containing 1; MTMR7, myotubularin related protein 7; AGPS, alkylglycerone phosphate synthase; PDE11A, phosphodiesterase 11A; CDKL5, cyclin dependent kinase like 5; DIAPH2, diaphanous related formin 2; DMD, dystrophin; DEPDC5, DEP domain containing 5; KEGG, Kyoto Encyclopedia of Genes and Genomes; cAMP, cyclic adenosine monophosphate; cGMP, cyclic guanosine monophosphate; TTC30B, tetratricopeptide repeat domain 30B.
For discussion purposes, we acknowledge that there may have been limitations in our study that were associated with the selection of roosters for obtaining sperm. In ideal world, we would have needed a larger flock of birds with high-quality semen. Roosters produce an average of 0.25 mL of semen per ejaculation that should be assessed before each ejaculate is frozen and after freezing. This procedure would require significant effort and semen volume given that low-quality samples should be rejected before freezing. In the main flocks used in this study, roosters are normally not tested and selected for semen quality, since they are used for natural mating. Artificial insemination, due to its labor intensity, can only be used on a limited scale within these flocks.
Our associative studies of the whole genome and traits associated with the processes of fertilizing ability of cryopreserved rooster sperm after freezing–thawing provide important information for understanding the genetic basis of sperm fertility. Further studies of polymorphic variants of candidate genes are necessary to develop a strategy for selecting high-quality biological material that retains fertility after freezing–thawing. This is especially important for preserving valuable genetic resources [141, 142, 143, 144, 145, 146, 147].
In the present study, we have focused on seeking the relevant candidate genes in the chicken genome that is identical in all cells. Our research objective did not include gene expression analysis in the sperm (in this case, there may be differences between the studied DNA sources, i.e., blood and semen). Because of that, it was not necessary to use spermatozoa for DNA extraction. Therefore, our association studies of the entire genome and traits associated with fertilizing ability of cryopreserved rooster sperm are pivotal for providing important information for understanding the genetic factors of, and potent markers for, sperm fertility and related impairments [19, 148, 149, 150, 151]. However, if one additionally aims to study expression of genes in spermatozoa, this approach could provide further information in this regard.
In addition, further research of polymorphic variants of candidate genes are necessary to develop a strategy for selecting high-quality biological material that retains fertility after freezing–thawing. This is especially important for depositing valuable genetic resources in semen cryobanks. Research on how different gene variants (e.g., those of PDE11A [126, 127, 128]) are affected by certain stimulating compounds that enhance semen quality may be useful in developing sperm extenders and improving sperm’s capacity to fertilize after freezing–thawing. The issue of cryoadditives requires more in-depth studies on the expression of candidate genes in the sperm of birds with different genotypes for candidate genes.
This study sought to identify genome-wide correlations between characteristics associated with spermatozoa’s capacity to fertilize following cryopreservation. We examined 96 roosters’ ejaculates, totaling 258 samples. Using a denser 600K SNP chip, this study was an extension of our earlier GWAS investigations [46, 47]. This led to the identification of more candidate genes and a greater number of rooster sperm cryoresistance markers, which increased the study’s depth. Following cryopreservation, FPM, FDM, and sperm CR induction by DNP were evaluated. We identified 26 potential genes after 31 statistically significant/suggestive SNPs were discovered for the three parameters under study. For the FPM trait cryopreservation, one candidate gene, ENSGALG00000029931 lncRNA, was found; for the FDM indicator, the PHF14 and ARID1B genes; and for the DNP sperm CR induction, KDELR3, DDX17, DMD, CDKL5, DGAT2, ST18, FAM150A (ALKAL1), DIAPH2, MTMR7, NAV2, RAG2, PDE11A, IFT70A (TTC30B), AGPS, WDFY1, DEPDC5, TSC1, CASZ1, and PLEKHM2 genes. The relationship between several detected lncRNAs and the studied structural and functional features of spermatozoa remains unclear.
The loci revealed in this GWAS can be used for breeding work aimed at increasing the cryotolerance of rooster sperm. The study of the candidate genes resulted from the sperm CR enhancer application, i.e., under the influence of DNP, is especially important since they are assumed to be associated with the fertilizing ability of semen. Our findings offer crucial data for further research and comprehension of the mechanisms underlying sperm–egg interactions, the genetic basis and characteristics of sperm membrane integrity, and the maintenance of fertility in roosters following the thawing of frozen sperm.
Data presented in this study are contained within this article and in the Supplementary Material, or are available upon request to the corresponding author.
Conceptualization, NVD, EVN; data curation, NVP, YLS, AAM, MNR; formal analysis, YSS; funding acquisition, NVD, EVN, MNR; investigation, NVD, AER, AIA; methodology, EVN, YSS, DKG; project administration, MNR; resources, NVP, YLS; software, YSS; supervision, DKG; validation, MNR; visualization, AER, MNR; writing – original draft, NVD, EVN, MNR, DKG; writing – review & editing, MNR, DKG. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The principles of laboratory animal care were followed, and all procedures were conducted, according to the ethical guidelines of the L. K. Ernst Federal Research Center for Animal Husbandry. The protocol was approved by the Commission on the Ethics of Animal Experiments at the L. K. Ernst Federal Research Center for Animal Husbandry (Protocol No. 2020/2) and the Law of the Russian Federation on Veterinary Medicine No. 4979-1 (dated 14 May 1993).
Not applicable.
This research was funded by the Ministry of Science and Higher Education of the Russian Federation (State Assignment Program No. 124020200114-7).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/FBS26022.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.


