1 Laboratory of Genomic Research, Research Institute for Genetic and Molecular Epidemiology, Kursk State Medical University, 305041 Kursk, Russia
2 Department of Obstetrics and Gynecology, Institute of Continuing Education, Kursk State Medical University, 305041 Kursk, Russia
3 Department of Biology, Medical Genetics and Ecology, Kursk State Medical University, 305041 Kursk, Russia
Abstract
Uterine fibroids (UF) is the most common benign tumour of the female reproductive system. We investigated the joint contribution of genome-wide association studies (GWAS)-significant loci and environment-associated risk factors to the UF risk, along with epistatic interactions between single nucleotide polymorphisms (SNPs).
DNA samples from 737 hospitalised patients with UF and 451 controls were genotyped using probe-based PCR for seven common GWAS SNPs: rs117245733 LINC00598, rs547025 SIRT3, rs2456181 ZNF346, rs7907606 STN1, SLK, rs58415480 SYNE1, rs7986407 FOXO1, and rs72709458 TERT.
We observed an association between rs547025 SIRT3 and the decreased risk of UF in overall group (effect allele C, odds ratio (OR) = 0.61, 95% confidence interval (CI) = 0.43–0.866, p = 0.005). SNP rs547025 exhibits protective effects against UF exclusively in patients with normal fruit and vegetable intake (OR = 0.39, 95% CI = 0.21–0.75, p = 0.002), no history of spontaneous abortions (OR = 0.48, 95% CI = 0.33–0.70, p = 0.0001), no pelvic inflammatory diseases (PID) in anamnesis (OR = 0.55, 95% CI = 0.38–0.80, p = 0.0016), and in smokers (OR = 0.20, 95% CI = 0.06–0.65, p = 0.006). In addition, rs7907606 STN1, SLK was associated with the risk of UF in patients without a history of pelvic inflammatory diseases (PID) (OR = 1.34, 95% CI = 1.03–1.74, p = 0.028). SNPs rs547025 SIRT3 and rs7907606 STN1, SLK, displayed the strongest mono-effects (0.71% and 0.52% contribution to UF entropy) and were characterized by the most pronounced gene-gene (G×G) effects when interacting with each other (0.60% contribution to entropy). The interaction Medical abortion×rs547025 SIRT3 served as the base for all the best gene-environment (G×E) models. Medical abortions have the most pronounced mono-effect (1.15% contribution to the entropy of UF), exceeding the mono-effects of SNPs involved in the most significant G×E-models (0.01%–0.49% contribution to entropy) and spontaneous abortions (0.48% of UF entropy) and exceeding the effects of G×E interactions (0.05–0.46% of UF entropy).
Bioinformatics analysis showed that GWAS SNPs are involved in the molecular mechanisms of UF mainly through the regulation of vasculogenesis, cell proliferation, apoptosis, DNA damage, inflammation, hypoxia, steroid hormone metabolism, cell signaling, organ formation.
Keywords
- uterine fibroids
- GWAS
- rs547025 SIRT3
- rs7907606 STN1, SLK
- gene-gene and gene-environmental interactions
- pelvic inflammatory diseases
- spontaneous abortions
- medical abortions
- fruit and vegetable intake
- smoking
Uterine fibroids (UF), also known as leiomyomas or fibromyomas, are the most common benign tumors of the female reproductive system, affecting up to 68.6% of women of reproductive age [1, 2]. Possible symptoms include prolonged or heavy menstrual bleeding, pelvic pressure or pain, impaired fertility, and consequent disruption of daily life and psychological well-being [3]. Currently, the leading treatment for fibroids remains surgical intervention [4]. The age of patients at first diagnosis of uterine fibroids is steadily decreasing [1, 5], which impacts their ability to conceive and successfully carry a pregnancy to term [6]. Due to this adverse and irreversible impact on the female reproductive system, risk factors for the development of uterine fibroids must be given particular attention.
Known risk factors for UF include premenopausal age, ethnicity (especially African descent), family history of UF, nulliparity, and increased body mass index (BMI) [7]. The molecular pathogenesis of uterine fibroids remains incompletely understood. Nonetheless, UF are recognised to be largely caused by oxidative stress [8], inflammation [9], hypoxia [10], steroid hormone imbalance [11], and extracellular matrix remodeling [12].
Genome-wide association studies (GWAS) have successfully identified genetic variants associated with multifactorial diseases, including UF [13, 14, 15, 16]. To date, the GWAS catalog contains data on more than 300 single nucleotide polymorphisms (SNPs) associated with UF (https://www.ebi.ac.uk/gwas/search?query=uterine%20fibroid). These data help to understand the molecular pathogenesis of the disease and stimulate the development of new drugs and preventive measures [17].
However, the role of GWAS-significant genetic variants in interaction with UF environment-associated risk factors, as well as the role of epistatic interactions between genetic variants, generally remains outside the focus of researchers, creating a significant gap in the functional analysis of GWAS-significant loci.
Our study aimed to replicate the associations of GWAS loci with UF risk in the Central Russian population; identify the most significant intergenic interactions associated with UF; conduct a comprehensive analysis of the combined influence of GWAS-significant loci and risk factors, such as smoking, intake of fresh vegetables and fruits, pelvic inflammatory diseases and medical/spontaneous abortions in history, on the risk of UF; and perform thorough functional annotation of SNPs.
The study included 1188 unrelated individuals from Central Russia, comprising 737 hospitalized patients with the diagnosis of uterine fibroid and 451 patients in the control group with no UF. The Ethical Review Committee of Kursk State Medical University approved the study protocol (protocol No. 5 from May 11, 2021), and all patients or their families/legal guardians provided written informed consent. The inclusion criteria for the study required participants to have self-declared Russian ancestry and to have been born in Central Russia. Table 1 provides the baseline and clinical characteristics of the study cohort.
| Baseline and clinical characteristics | Uterine fibroid patients (n = 737) | Controls (n = 451) | p-value | |
| Age, Ме [Q1; Q3] | 48 [43; 52] | 51 [43; 59] | ||
| Smoking | Yes, N (%) | 108 (14.7%) | 44 (9.7%) | |
| No, N (%) | 629 (85.3%) | 407 (90.3%) | ||
| Low fruit/vegetable consumption | Yes, N (%) | 603 (81.8%) | ND | - |
| No, N (%) | 134 (18.2%) | ND | ||
| History of infertility | Yes, N (%) | 8 (1.1%) | 2 (0.4%) | |
| No, N (%) | 566 (76.8%) | 376 (83.4%) | ||
| ND, N (%) | 163 (22.1%) | 73 (16.2%) | ||
| Pelvic inflammatory diseases (PID) | Yes, N (%) | 108 (14.6%) | 61 (13.5%) | |
| No, N (%) | 464 (63.0%) | 318 (70.5%) | ||
| ND, N (%) | 165 (22.4%) | 72 (16%) | ||
| Family history of UF | Yes, N (%) | 205 (27.8%) | 36 (8%) | |
| No, N (%) | 532 (72.2%) | 415 (92%) | ||
| Age of menarche | Ме [Q1; Q3] | 12 [12; 14] | 13 [12; 14] | |
| Pregnancy | Yes, N (%) | 622 (84.4%) | 401 (88.9%) | |
| No, N (%) | 29 (3.9%) | 13 (2.9%) | ||
| ND, N (%) | 86 (11.7%) | 37 (8.2%) | ||
| Number of pregnancies | Ме [Q1; Q3] | 3 [2; 5] | 3 [2; 4] | |
| Parity | Yes, N (%) | 604 (81.9%) | 394 (87.4%) | |
| No, N (%) | 44 (6%) | 19 (4.2%) | ||
| ND, N (%) | 89 (12.1%) | 38 (8.4%) | ||
| Number of labors | Ме [Q1; Q3] | 2 [1; 2] | 2 [1; 2] | |
| Medical abortion in anamnesis | Yes, N (%) | 438 (59.4%) | 225 (49.9%) | |
| No, N (%) | 201 (27.3%) | 178 (39.5%) | ||
| ND, N (%) | 98 (13.3%) | 48 (10.6%) | ||
| Number of medical abortions | Ме [Q1; Q3] | 1 [0; 2] | 1 [0; 2] | |
| Spontaneous abortion in anamnesis | Yes, N (%) | 133 (18%) | 82 (18.2%) | |
| No, N (%) | 478 (64.9%) | 318 (70.5%) | ||
| ND, N (%) | 126 (17.1%) | 51 (11.3%) | ||
| Number of spontaneous abortions | Ме [Q1; Q3] | 0 [0; 0] | 0 [0; 0] | |
| Prolonged periods ( |
Yes, N (%) | 180 (24.4%) | 83 (18.4%) | |
| No, N (%) | 442 (60%) | 305 (67.6%) | ||
| ND, N (%) | 115 (15.6%) | 63 (14%) | ||
| Regular periods | Yes, N (%) | 357 (48.4%) | 200 (44.3%) | |
| No, N (%) | 159 (21.6%) | 2 (0.5%) | ||
| ND, N (%) | 221 (30%) | 249 (55.2%) | ||
| Dysmenorrhea | Yes, N (%) | 227 (30.8%) | 23 (5.1%) | |
| No, N (%) | 236 (32%) | 179 (39.7%) | ||
| ND, N (%) | 274 (37.2%) | 249 (55.2%) | ||
| Menorrhagia (heavy menstrual bleeding) | Yes, N (%) | 370 (50.2%) | 36 (8%) | |
| No, N (%) | 144 (19.5%) | 166 (36.8%) | ||
| ND, N (%) | 223 (30.3%) | 249 (55.2%) | ||
| Multiple form of UF | Yes, N (%) | 411 (55.8%) | - | - |
| No, N (%) | 217 (29.4%) | - | ||
| ND, N (%) | 109 (14.8%) | - | ||
| Age of UF diagnosis | Ме [Q1; Q3] | 40 [36; 45] | - | - |
| Uterine size at age of diagnosis (weeks of pregnancy) | Ме [Q1; Q3] | 7 [6; 9] | - | - |
| Current uterine size (weeks of pregnancy) | Ме [Q1; Q3] | 9 [7; 12] | - | - |
Note: Me, median; Q1, the first quartile; Q3, the third quartile; ND, no data; UF, uterine fibroids; differences that are statistically significant are indicated in bold.
The patients were enrolled in the study in 2021–2023 at the Perinatal Center and Kursk City Maternity Hospital from gynecological departments. All patients had an ultrasound-confirmed diagnosis of UF.
The control group consisted of relatively healthy patients without clinical or ultrasound signs of uterine fibroids. Controls from the Kursk region were selected from hospitals during routine medical examinations conducted in public institutions and industrial establishments [18, 19, 20, 21]. This group was recruited from the same population and during the same time period.
As risk factors for the development of uterine fibroids, we considered the following:
• Smoking: Some studies suggest that smoking affects the risk of developing uterine fibroids [7, 22] by lowering estrogen levels [23] and altering regulatory cytokines and receptivity markers, such as C-X-C motif chemokine ligand 12 (CXCL12) and fibroblast growth factor 2 (FGF2) [24].
• Intake of fresh vegetables and fruits: This factor has been identified by researchers as a significant risk factor for uterine fibroids [25, 26]. Phytochemicals in these foods, including flavonoids, carotenoids, and polyphenols, are known for their ability to regulate cell proliferation, inflammation, fibrosis, apoptosis, and angiogenesis [27] through managing the response to oxidative stress [28]. In accordance with World Health Organization (WHO) guidelines, low fruit and vegetable consumption was defined as consuming less than 400 g per day. Adequate consumption of fresh vegetables and fruits was defined as consuming 400 g or more, equivalent to 3–4 servings per day, excluding starchy tubers like potatoes [29].
• Spontaneous abortions are considered by Song et al. [30] and Parazzini et al. [31] as a risk factor for uterine fibroids. Early and especially late spontaneous abortions lead to a sharp decrease in steroid sex hormone levels after miscarriage relative to baseline. Abortions influence the expression of estrogen and progesterone receptors, subsequently affecting tumor formation [32].
• Medical (induced) abortions influence the risk of developing uterine fibroids through local inflammation following invasive methods like vacuum extraction or D&C (dilation and curettage) [33].
• Pelvic inflammatory diseases: Trauma, infection, and subsequent inflammation cause an imbalance in the immune system by increasing T-helper cytokines and decreasing the function of regulatory T cells (Treg). This immune response leads to the formation and proliferation of fibrous tissue [34].
For this study, we selected SNPs using the GWAS catalog data
(https://www.ebi.ac.uk/gwas/, assessed on February 15, 2023), which contains data
on 238 SNPs located in 169 loci that have been found to be associated with UF
risk by 22 GWAS studies. Our study included genetic variants that were associated
with the risk of UF in European populations at least in two studies. SNPs with a
minor allele frequency
The Laboratory of Genomic Research at the Research Institute for Genetic and Molecular Epidemiology of Kursk State Medical University (Kursk, Russia) performed genotyping. Each participant provided up to 5 mL of venous blood from a cubital vein, which was stored in EDTA-coated tubes at –20 °C until processing. Defrosted blood samples were utilised to extract genomic DNA using typical procedures such as phenol/chloroform extraction and ethanol precipitation. The extracted DNA samples’ purity, quality, and concentration were evaluated using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
Genotyping of SNPs was carried out employing allele-specific probe-based polymerase chain reaction (PCR) techniques designed in the Laboratory of Genomic Research. The Primer3 software (https://primer3.ut.ee/) was used for primer design [35]. A real-time PCR procedure was performed in a 25 µL reaction solution containing 1.5 units of Hot Start Taq DNA polymerase (Biolabmix, Novosibirsk, Russia), approximately 10 ng of DNA, and the following concentrations of reagents: 0.25 µM of each primer; 0.1 µM of each probe; 250 µM of each dNTP; 3 mM MgCl2 for rs117245733 and rs547025; 3.5 mM MgCl2 for rs2456181 and rs7907606; and 2.5 mM MgCl2 for the rs58415480, rs7986407, and rs72709458; 1xPCR buffer (67 mM Tris-HCl, pH 8.8, 16.6 mM (NH4)2SO4, 0.01% Tween-20). The PCR procedure comprised an initial denaturation for 10 minutes at 95 °C, followed by 39 cycles of 92 °C for 30 s and 64 °C, 65 °C, 63 °C, 62 °C, 60 °C, 59 °C, 59 °C for 1 min (for rs117245733 LINC00598, rs547025 SIRT3, rs2456181 ZNF346, rs7907606 STN1, SLK, rs58415480 SYNE1, rs7986407 FOXO1, rs72709458 TERT, respectively). 10% of the DNA samples were genotyped twice, blinded to the case-control status, in order to assure quality control. Over 99% of the data were consistant. Due to the Hardy-Weinberg equilibrium deviation in the control group for SNP rs7907606 STN1, SLK, all locus samples underwent re-genotyping. The results were entirely consistent (100%) with the initial genotypes.
The STATISTICA software (version 13.3, Santa Clara, CA, USA) was utilized for statistical processing. The normality of the distribution for quantitative data was assessed using the Shapiro-Wilk’s test. Given that the majority of quantitative values had deviations from normal distribution, they were given as the median (Me), along with the first and third quartiles [Q1 and Q3]. To compare the quantitative variables between two independent groups, the Mann-Whitney test was performed. Differences in statistical significance between categorical variables were assessed using Pearson’s chi-squared test with Yates’ correction for continuity. Fisher’s exact test was used to determine if genotype distributions were consistent with Hardy-Weinberg equilibrium. The study groups’ genotype frequencies and their associations with disease risk were analyzed using regression analysis using the SNPStats software resource (https://www.snpstats.net/start.htm (accessed on 6 June, 2024)). The additive model was considered for the genotype association analysis. Associations within the entire group of UF patients/controls were adjusted for age and UF family history. Environmental risk factors can significantly influence the connection of genetic markers with disease [36, 37]. Therefore, associations were analyzed depending on the presence or absence of the risk factor. Since data on fresh vegetable and fruit consumption was unavailable for the control group, SNP associations were analyzed based solely on the presence or absence of this risk factor within the patient group, using the overall control group as a baseline. To adjust for multiple comparisons, a Bonferroni correction was applied, accounting for the two comparison groups.
The model-based multivariate dimensionality reduction (MB-MDR) method analysis
tested two-, three-, and four-level genotype combinations (gene-gene,
G
Additionally, the most significant G
The functional effects of SNPs were examined using bioinformatics resources, the methodologies and functionalities of which were comprehensively described in our prior research [39, 40, 41]:
• The bioinformatic tool GTExportal (http://www.gtexportal.org/ (accessed on June 10, 2024)) was used to analyze the link of SNPs with expression quantitative trait loci (eQTLs) in reproductive organs, adipose tissue, vessels, whole blood, thyroid, adrenal, and pituitary glands [42].
• For additional examination of binding SNPs to expression quantitative trait loci (eQTL) in peripheral blood, the eQTLGen resource available at https://www.eqtlgen.org/ (accessed on June 10, 2024) was employed [43].
• HaploReg (v4.2), a bioinformatics tool available at
https://pubs.broadinstitute.org/mammals/haploreg/haploreg.php (accessed on June
11, 2024), was utilized to assess the associations between GWAS SNPs and specific
histone modifications marking promoters and enhancers. These modifications
included acetylation of lysine residues at positions 27 and 9 of the histone H3
protein, as well as mono-methylation at position 4 (H3K4me1) and tri-methylation
at position 4 (H3K4me3) of the histone H3 protein. Additionally, the tool was
applied to investigate the positioning of SNPs in DNase hypersensitive regions
and to search for SNPs that are strongly linked (r2
• The atSNP Function Prediction online tool (http://atsnp.biostat.wisc.edu/search (accessed on June 11, 2024)) was used to evaluate the impact of SNPs on the gene affinity to transcription factors (TFs) depending on the carriage of the reference/alternative alleles [45]. TFs were included based on the degree of influence of SNPs on the interaction of TFs with DNA, calculated on the basis of a positional weight matrix.
• Using the Gene Ontology online tool (http://geneontology.org/ (accessed on June 11, 2024)), it was feasible to analyze the joint involvement of TFs linked to the reference/SNP alleles in overrepresented biological processes directly related to the pathogenesis of UF [46]. Biological functions controlled by transcription factors associated with SNPs were used as functional groups.
• The Reproductive System Knowledge Portal (RSKP) (https://cd.hugeamp.org/ (accessed on June 15, 2024)), which combines and analyzes the results of genetic associations of the largest consortiums for the study of reproductive system diseases, was used for bioinformatics analysis of associations of SNPs with UF and intermediate phenotypes (such as body mass index, heavy menstrual bleeding, etc.).
The genotype frequencies of SNPs within the study cohorts are detailed in
Supplementary Table 2. Because connections between genetic markers and
disease can produce deviations from equilibrium, we relied on the control group’s
Hardy-Weinberg equilibrium results. Within the control group, all studied SNPs
exhibited genotype frequencies consistent with Hardy-Weinberg equilibrium
(p
The analysis of the entire group (Table 2) revealed an association between rs547025 SIRT3 and the decreased risk of UF, regardless of age and family history of UF: effect allele C, OR = 0.61, 95% CI = 0.43–0.866, p = 0.005.
| Genetic variant | Effect allele (minor) | Other allele (major) | N | OR [95% CI]1 | p2 |
| rs117245733 LINC00598 | A | G | 952 | 0.70 [0.38–1.29] | 0.25 |
| rs547025 SIRT3 | C | T | 953 | 0.61 [0.43–0.86] | 0.005 |
| rs2456181 ZNF346 | C | G | 953 | 0.90 [0.75–1.09] | 0.29 |
| rs7907606 STN1, SLK | G | T | 953 | 1.28 [1.00–1.63] | 0.044 |
| rs58415480 SYNE1 | C | G | 954 | 1.10 [0.86–1.41] | 0.44 |
| rs7986407 FOXO1 | A | G | 952 | 1.07 [0.87–1.31] | 0.54 |
| rs72709458 TERT | C | T | 952 | 1.06 [0.84–1.34] | 0.61 |
All calculations were performed relative to the minor alleles (Effect allele) with adjustment for age. SNPs, single nucleotide polymorphisms; UF family history; GWAS, genome-wide association studies; 1OR, odds ratio; 95% CI, 95% confidence interval; 2p-value; statistically significant differences are marked in bold.
Using the MB-MDR approach, four most significant models of gene-gene
interactions of polymorphic loci of GWAS-significant genes associated with
uterine fibroids were identified: one two-locus, two three-locus, and one
four-locus (pperm
| Gene-gene interaction models | NH | beta H | WH | NL | beta L | WL | Wmax | pperm |
| The best two-locus models of intergenic interactions (for G×G models with pmin. | ||||||||
| rs547025 SIRT3 × rs117245733 LINC00598 | 1 | 0.11912 | 10.376 | 1 | –0.11358 | 6.991 | 10.376 | 0.01 |
| The best three-locus models of intergenic interactions (for G×G models with pmin. | ||||||||
| rs72709458 TERT × rs7907606 STN1, SLK × rs547025 SIRT3 | 1 | 0.0989 | 5.449 | 4 | –0.1775 | 14.01 | 14.01 | 0.029 |
| rs72709458 TERT × rs58415480 SYNE1 × rs547025 SIRT3 | 0 | NA | NA | 4 | –0.2061 | 13.46 | 13.46 | 0.033 |
| The best four-locus models of gene-gene interactions (for G×G models with pmin. | ||||||||
| rs72709458 TERT × rs7907606 STN1, SLK × rs547025 SIRT3 × rs117245733 LINC00598 | 1 | 0.1011 | 5.426 | 5 | –0.1976 | 17.32 | 17.32 | 0.01 |
Note: MB-MDR, model-based multivariate dimensionality reduction; NH is the
number of interacting high-risk genotypes; beta H, regression coefficient for
high-risk interactions identified at the 2nd stage of analysis; WH, Wald
statistics for high-risk interactions; NL, number of interacting low-risk
genotypes; beta L, regression coefficient for low-risk interactions identified at
the 2nd stage of analysis; WL, Wald statistics for low-risk interactions;
pperm, permutational significance levels for models (all models are
adjusted for age; UF family history); G
In the next step, interactions between genetic variants characterizing at least two of the most significant models were analyzed using the multivariate dimensionality reduction (MDR) method (Fig. 1).
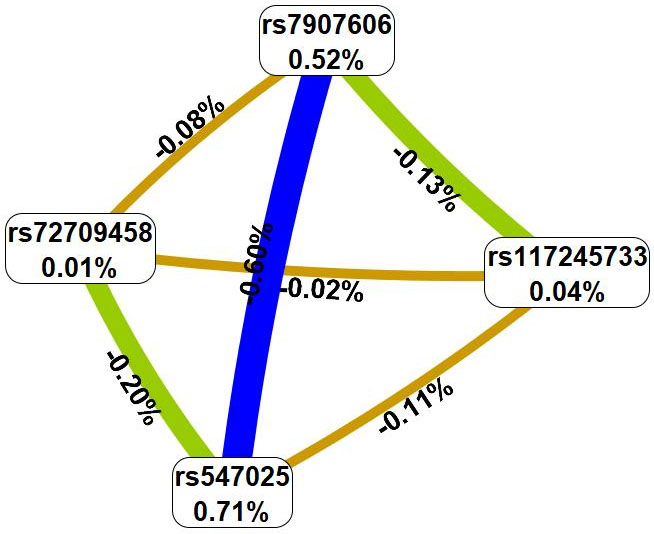 Fig. 1.
Fig. 1.
Graph reflecting the structure and power of the most significant
G
The MDR method, firstly, showed that the genetic variants involved in the best
G
Fifth, the strongest associations with UF have the following combinations of
genotypes: rs547025 SIRT3 T/T
Risk factor-stratified analysis is detailed in Supplementary Table 4. We discovered that rs547025 SIRT3 was related to UF regardless of a history of medical abortions in the anamnesis; nonetheless, it was modified by all other risk factors that were evaluated (Table 4). Thus, rs547025 SIRT3 exhibited protective effects against UF exclusively in patients with normal fruit and vegetable intake (OR = 0.39, 95% CI = 0.21–0.75, pbonf = 0.002), no history of spontaneous abortions (OR = 0.48, 95% CI = 0.33–0.70, p = 0.0001), no pelvic inflammatory diseases (PID) in anamnesis (OR = 0.55, 95% CI = 0.38–0.80, p = 0.0016), and in smokers (OR = 0.20, 95% CI = 0.06–0.65, p = 0.006). In addition, rs7907606 STN1, SLK was associated with the risk of UF in patients without a history of PID (OR = 1.34, 95% CI = 1.03–1.74, p = 0.028) (Table 4).
| Genetic variant | Effect allele | Other allele | N | OR [95% CI]1 | p2 (pbonf) | N | OR [95% CI]1 | p2 (pbonf) |
| Nonsmokers | Smokers | |||||||
| rs547025 | C | T | 621 | 0.73 [0.49–1.08] | 0.11 | 130 | 0.20 [0.06–0.65] | 0.006 |
| SIRT3 | ||||||||
| Normal fruit and vegetable intake | Low fruit and vegetable intake | |||||||
| rs547025 | C | T | 529 | 0.39 [0.21–0.75] | 0.001 (0.002) | 927 | 0.71 [0.52–0.97] | 0.034 (0.068) |
| SIRT3 | ||||||||
| No spontaneous abortion | Spontaneous abortion | |||||||
| rs547025 | C | T | 699 | 0.48 [0.33–0.70] | 0.0001 | 189 | 1.53 [0.67–3.50] | 0.3 |
| SIRT3 | ||||||||
| No medical abortion | Medical abortion | |||||||
| rs547025 | C | T | 334 | 0.56 [0.32–0.97] | 0.037 | 585 | 0.62 [0.41–0.93] | 0.021 |
| SIRT3 | ||||||||
| No PID in anamnesis | PID in anamnesis | |||||||
| rs547025 | C | T | 696 | 0.55 [0.38–0.80] | 0.0016 | 135 | 0.68 [0.30–1.56] | 0.36 |
| SIRT3 | ||||||||
| rs7907606 | G | T | 696 | 1.34 [1.03–1.74] | 0.028 | 135 | 1.25 [0.64–2.45] | 0.51 |
| STN1, SLK | ||||||||
All calculations were performed relative to the minor alleles (Effect allele); SNP, single nucleotide polymorphisms; 1OR, odds ratio; 95% CI, 95% confidence interval; 2p-value; statistically significant differences are marked in bold.
Using the MB-MDR approach, nine most significant gene-environment interactions
associated with UF were identified: two two-level, three three-level, and four
four-level (Table 5). Notably, SNPs rs547025 SIRT3, rs2456181
ZNF346, and rs72709458 TERT were involved in two or more of the
best G
| Gene-gene interaction models | NH | beta H | WH | NL | beta L | WL | Wmax | pperm |
| The best two-order models of gene-smoking interactions (for G×E models with pmin. | ||||||||
| Med_abort × rs547025 SIRT3 | 1 | 0.14406 | 22.620 | 3 | –0.1278 | 16.43 | 22.62 | |
| Med_abort × rs2456181 ZNF346 | 1 | 0.08028 | 4.678 | 1 | –0.1825 | 19.97 | 19.97 | 0.001 |
| The best three-order models of gene-interactions (for G×E models with pmin. | ||||||||
| Spont_abort × Med_abort × rs547025 SIRT3 | 3 | 0.15962 | 26.705 | 4 | –0.2160 | 30.42 | 30.42 | |
| Med_abort × rs547025 SIRT3 × rs117245733 LINC00598 | 1 | 0.15619 | 26.791 | 3 | –0.1226 | 14.62 | 26.79 | |
| Med_abort × rs72709458 TERT × rs547025 SIRT3 | 2 | 0.14167 | 21.931 | 5 | –0.1551 | 24.32 | 24.32 | |
| The best four-order models of gene-interactions (for G×E models with pmin. | ||||||||
| Spont_abort × Med_abort × rs7986407 FOXO1 × rs547025 SIRT3 | 2 | 0.1271 | 12.15 | 6 | –0.3142 | 41.33 | 41.33 | |
| Spont_abort × Med_abort × rs72709458 TERT × rs547025 SIRT3 | 3 | 0.1463 | 15.26 | 5 | –0.2802 | 35.26 | 35.26 | |
| Spont_abort × Med_abort × rs2456181 ZNF346 × rs547025 SIRT3 | 3 | 0.1273 | 11.05 | 5 | –0.2191 | 34.27 | 34.27 | |
| Spont_abort × Med_abort × rs58415480 SYNE1 × rs547025 SIRT3 | 3 | 0.1209 | 11.80 | 7 | –0.2461 | 33.72 | 33.72 | |
Note: NH is the number of high-risk interactions; beta H, regression coefficient
for high-risk interactions identified at the 2nd stage of analysis; WH, Wald
statistics for high-risk interactions; NL, number of interacting low-risk
interactions; beta L, regression coefficient for low-risk interactions identified
at the 2nd stage of analysis; WL, Wald statistics for low-risk interactions;
pperm, permutational significance levels for models (all models are
adjusted for age, UF family history); G
In the next step, interactions between these risk factors and genetic variants characterizing at least two of the most significant models were analyzed using the multivariate dimensionality reduction (MDR) method (Fig. 2).
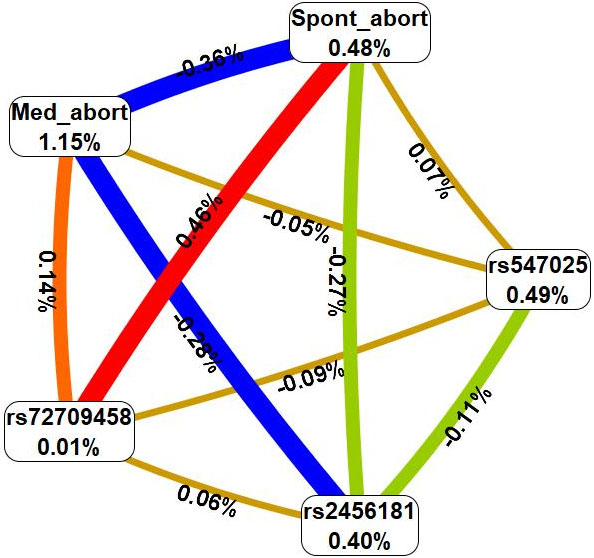 Fig. 2.
Fig. 2.
Graph reflecting the structure and power of the most significant
G
First, MDR revealed that such an environmental risk factor as medical abortions
has the most pronounced mono-effect (1.15% contribution to the entropy of UF),
exceeding the mono-effects of SNPs involved in the most significant models of
gene-environment interactions (0.01%–0.49% contribution to entropy) or
spontaneous abortions (0.48% of UF entropy), and exceeding the effects of
gene-environment interactions (0.05%–0.46% of UF entropy) (Fig. 2). Secondly,
in interaction with spontaneous/medical abortions, rs72709458 TERT is
characterized by synergism, rs2456181 ZNF346 by antagonism, and rs547025
SIRT3 by independent (additive) effects. Thirdly, spontaneous and
medical abortions interact antagonistically. Fourthly, SNPs characterizing the
best G
Fifth, the strongest associations with UF have the following gene-environmental
interactions: medical abortion
The summary of the effects of GWAS SNPs on the contribution to UF is provided in Table 6.
| SNP | G×G interactions | G×E interactions | ||
| Mono-effect | GG-effect | Mono-effect | GE-effect | |
| rs547025 SIRT3 | 0.71% | 0.91% | 0.49% | 0.32% |
| rs117245733 LINC00598 | 0.04% | 0.26% | - | |
| rs72709458 TERT | 0.01% | 0.30% | 0.01% | 0.66% |
| rs7907606 STN1, SLK | 0.52% | 0.81% | - | |
| rs2456181 ZNF346 | - | 0.40% | 0.72% | |
In our analysis of genetic variants, gene-gene, and gene-environment interactions, we identified significant associations between UF and the following GWAS-identified loci: rs547025 SIRT3, rs117245733 LINC00598, rs72709458 TERT, rs7907606 STN1, SLK, and rs2456181 ZNF346. A comprehensive functional annotation of these SNPs was also performed.
The results of the cis-eQTL analysis (Table 7), shed light on the impact of specific genetic variants on gene expression. According to the GTEx Portal, rs547025 SIRT3 increases the expression of RP11-326C3.12 in the adrenal gland and decreasea the expression of RIC8A in adipose tissue (subcutaneous) and PSMD13 in cultured fibroblasts. SNP rs2456181 ZNF346 increases the expression of FGFR4 in subcutaneous adipose tissue, arteries, pituitary, cultured fibroblasts, and whole blood and increases UIMC1 expression in subcutaneous adipose tissue, ovaries, and whole blood. Additionally, rs2456181 ZNF346 decreases the expression of ZNF346 and ZNF346-IT1 in the thyroid gland.
| SNP | Gene Expressed | p-value | Effect (NES) | Tissue |
| rs547025 SIRT3 (T/C) | RP11–326C3.12 | 3.1 × 10–6 | 0.82 | Adrenal Gland |
| PSMD13 | 1.0 × 10–7 | –0.15 | Cells - Cultured fibroblasts | |
| RIC8A | 1.6 × 10–5 | –0.23 | Adipose - Subcutaneous | |
| rs2456181 ZNF346 (G/C) | FGFR4 | 1.6 × 10–7 | 0.24 | Adipose - Subcutaneous |
| UIMC1 | 5.5 × 10–7 | 0.099 | Adipose - Subcutaneous | |
| FGFR4 | 4.2 × 10–5 | 0.22 | Artery - Aorta | |
| FGFR4 | 2.3 × 10–9 | 0.26 | Artery - Tibial | |
| FGFR4 | 1.4 × 10–9 | 0.30 | Cells - Cultured fibroblasts | |
| UIMC1 | 1.8 × 10–5 | 0.26 | Ovary | |
| FGFR4 | 2.7 × 10–5 | 0.27 | Pituitary | |
| ZNF346 | 3.8 × 10–22 | –0.30 | Thyroid | |
| ZNF346–IT1 | 3.9 × 10–5 | –0.16 | Thyroid | |
| FGFR4 | 4.5 × 10–12 | 0.28 | Whole Blood | |
| UIMC1 | 4.1 × 10–5 | 0.067 | Whole Blood |
Note: Effect alleles are marked in bold. eQTL, expression quantitative trait loci; NES, normalized effect size.
Moreover, data from the eQTLGen Browser indicated (Table 8) that rs547025 SIRT3 is associated with reduced expression levels of PSMD13, BET1L, SCGB1C1, RIC8A, ODF3, IFITM3, PTDSS2, and an elevation in the expression of IFITM2 and IFITM1 in blood. SNP rs7907606 STN1, SLK is correlated with decreased expression of RP11-541N10.3, OBFC, and increased expression levels of SLK, SH3PXD2A, GSTO1, RP11-127L20.6, ITPRIP, and CALHM2 in the blood. And finally, SNP rs2456181 ZNF346 was correlated with increased UIMC1, FGFR4, and HK3 and decreased ZNF346-IT1 in blood.
| SNP | Symbol | Z-score | Assessed | Other | FDR |
| rs547025 SIRT3 (T/C) | PSMD13 | –16.7775 | C | T | 0 |
| BET1L | –16.5046 | C | T | 0 | |
| SCGB1C1 | –14.2627 | C | T | 0 | |
| RIC8A | –7.7756 | C | T | 0 | |
| ODF3 | –7.0824 | C | T | 0 | |
| IFITM2 | 6.4087 | C | T | 0 | |
| IFITM1 | 5.74 | C | T | 5.8 × 10–5 | |
| IFITM3 | –4.6478 | C | T | 0.01 | |
| PTDSS2 | –4.3283 | C | T | 0.04 | |
| rs7907606 STN1, SLK (T/G) | SLK | 13.0742 | G | T | 0 |
| SH3PXD2A | 9.3992 | G | T | 0 | |
| RP11-541N10.3 | –7.5035 | G | T | 0 | |
| GSTO1 | 6.8781 | G | T | 0 | |
| OBFC1 | –6.7461 | G | T | 0 | |
| RP11-127L20.6 | 6.2581 | G | T | 0 | |
| ITPRIP | 5.6728 | G | T | 7 × 10–5 | |
| CALHM2 | 4.5757 | G | T | 0.01 | |
| rs2456181 ZNF346 (G/C) | UIMC1 | 34.9529 | G | C | 0 |
| FGFR4 | 13.9163 | G | C | 0 | |
| ZNF346-IT1 | –4.477 | G | C | 0.02 | |
| HK3 | 4.2789 | G | C | 0.047 |
Note: FDR, false discovery rate.
The analysis of transcription factors of GWAS loci is presented in
Supplementary Tables 6–10. It was revealed that the risk
allele G rs7907606 STN1, SLK creates DNA binding sites for 37
TFs, co-controlling cell differentiation (GO:0030154; false discovery rate (FDR) = 1.24
SNP allele G rs2456181 ZNF346 creates DNA binding sites for 45 TFs, co-controlling positive regulation of angiogenesis (GO:0045766; FDR = 0.01), regulation of cell population proliferation (GO:0042127; FDR = 0.002), interleukin-9-mediated signaling pathway (GO:0038113; FDR = 0.007), growth hormone receptor signaling pathway via JAK-STAT (GO:0060397; FDR = 0.013), cellular response to interleukin-17 (GO:0097398; FDR = 0.025), positive regulation of vascular endothelial growth factor production (GO:0010575; FDR = 0.002), cell surface receptor signaling pathway via JAK-STAT (GO:0007259; FDR = 0.008) (Supplementary Table 7).
SNP allele A rs117245733 LINC00598 creates DNA binding sites for 37
TFs, jointly involved in cellular response to steroid hormone stimulus
(GO:0071383; FDR = 0.01) and negative regulation of Wnt signaling pathway
(GO:0030178; FDR = 0.014) (Supplementary Table 7). Reference allele G
rs117245733 LINC00598 creates DNA binding sites for 32 TFs, regulating
the following biological processes: hormone-mediated signaling pathway
(GO:0009755; FDR = 1.68
Carriage of the SNP T allele rs72709458 TERT results in loss of DNA
binding to TFs that are jointly involved in DNA damage response, signal
transduction by p53 class mediator resulting in transcription of p21 class
mediator (GO:0006978; FDR = 0.015), response to hypoxia (GO:0001666; FDR =
0.014), negative regulation of cell population proliferation (GO:0008285; FDR =
3.29
Using the bioinformatics tool HaploReg (v4.2), we analyzed histone modifications associated with SNPs identified in our study as linked to an increased risk of UF (Supplementary Table 11).
SNPs rs547025 SIRT3, rs7907606 STN1, SLK, rs117245733 LINC00598, and rs2456181 ZNF346 are characterized by histone H3 mono-methylation at the 4th lysine residue (H3K4me1), acetylation of the lysine residues at N-terminal position 27 of the histone H3 protein (H3K27ac), and acetylation at the 9th lysine residues of the histone H3 protein (H3K9ac) in the blood. Similarly, SNPs rs117245733 LINC00598 and rs2456181 ZNF346 are associated with histone tags in blood vessels, ovary, and adipose tissue (Supplementary Table 11).
According to the bioinformatic resource Reproductive System Knowledge Portal, the GWAS-significant SNPs rs547025 SIRT3, rs117245733 LINC00598, rs72709458 TERT, rs7907606 STN1, SLK and rs2456181 ZNF346, associated with UF in our study, are linked to the increased risk of UF and heavy menstrual bleeding in other populations (all SNPs), to age at natural menopause (rs547025 SIRT3, rs2456181 ZNF346), and body mass index (rs7907606 STN1, SLK) (Table 9).
| No | SNP | Phenotype | p-value | Beta (OR) | Sample Size |
| 1. | rs547025 SIRT3 (C/T) | Uterine fibroids | 4.78 × 10–13 | OR▼0.8724 | 244,324 |
| 2. | Uterine fibroids and heavy menstrual bleeding | 1.70 × 10–6 | OR▼0.9967 | 13,406 | |
| 3. | Heavy menstrual bleeding | 0.01 | OR▼0.9965 | 37,507 | |
| 4. | Age at natural menopause (ANM) | 0.036 | Beta▼–0.0120 | 244,171 | |
| 5. | rs7907606 STN1, SLK (G/T) | Uterine fibroids | 6.14 × 10–9 | OR▲1.0827 | 244,324 |
| 6. | Body mass index (BMI) | 1.05 × 10–4 | Beta▼–0.0084 | 2,140,420 | |
| 7. | Uterine fibroids and heavy menstrual bleeding | 0.001 | OR▲1.0016 | 13,406 | |
| 8. | rs117245733 LINC00598 (G/A) | Uterine fibroids | 2.52 × 10–2 | OR▲1.3127 | 244,324 |
| 9. | Uterine fibroids and heavy menstrual bleeding | 0.0019 | OR▲1.0046 | 13,406 | |
| 10. | rs2456181 ZNF346 (C/G) | Age at natural menopause (ANM) | 3.12 × 10–134 | Beta▲0.0853 | 162,657 |
| 11. | Uterine fibroids and heavy menstrual bleeding | 4.00 × 10–10 | OR▲1.0023 | 13,406 | |
| 12. | Uterine fibroids | 5.62 × 10–9 | OR▲1.0677 | 239,139 | |
| 13. | Heavy menstrual bleeding | 0.000059 | OR▲1.0028 | 37,507 | |
| 14. | rs72709458 TERT (C/T) | Uterine fibroids | 6.07 × 10–15 | OR▲1.1049 | 244,324 |
| 15. | Heavy menstrual bleeding | 1.70 × 10–8 | OR▲1.0050 | 37,507 | |
| 16. | Uterine fibroids and heavy menstrual bleeding | 3.30 × 10–8 | OR▲1.0025 | 13,406 |
Data obtained using the bioinformatic resource Reproductive System Knowledge Portal (https://reproductive.hugeamp.org/). ▼, indicates a decrease in the effect; ▲, indicates a increase in the effect. Effect alleles are marked in bold.
For a more in-depth analysis of the phenotypic effects of UF-GWAS-significant
SNPs, we included assessement of polymorphic loci strongly linked (with r2
In the present study, we replicated the associations of the GWAS loci rs547025 SIRT3 and rs7907606 STN1, SLK with the occurrence of uterine myoma among the Caucasian population of Central Russia. We also discovered, for the first time, that environment-associated risk factors like as cigarette smoking, consuming fresh vegetables and fruits, and undergoing abortions have an important impact on these connections (Fig. 3).
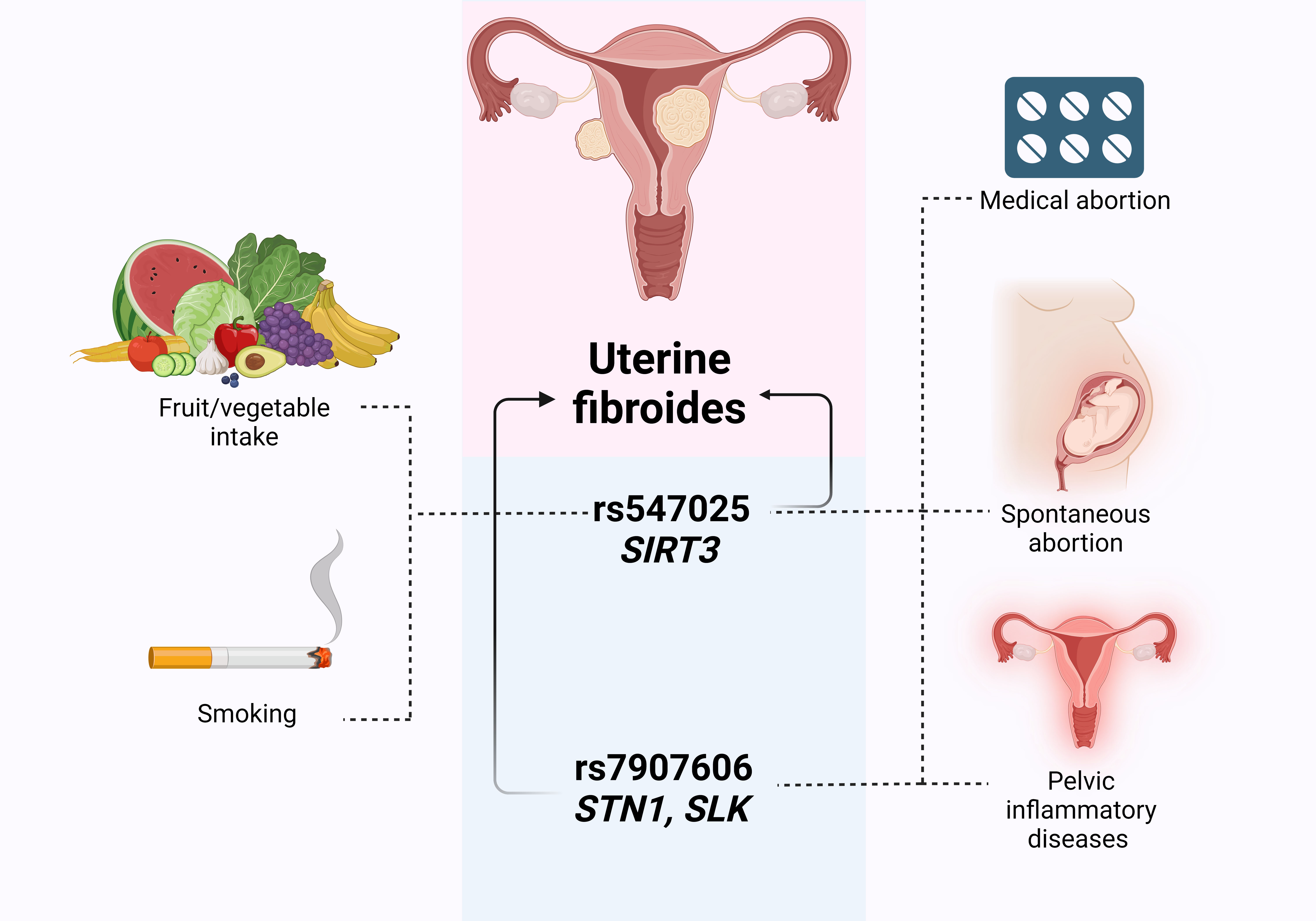 Fig. 3.
Fig. 3.
The outline of associations between environment-associated risk factors, GWAS SNPs and UF risk. Created with Biorender.
We identified GWAS SNPs with particularly prominent epistatic interactions and performed a comprehensive bioinformatics inquiry of UF-associated variants (Fig. 4).
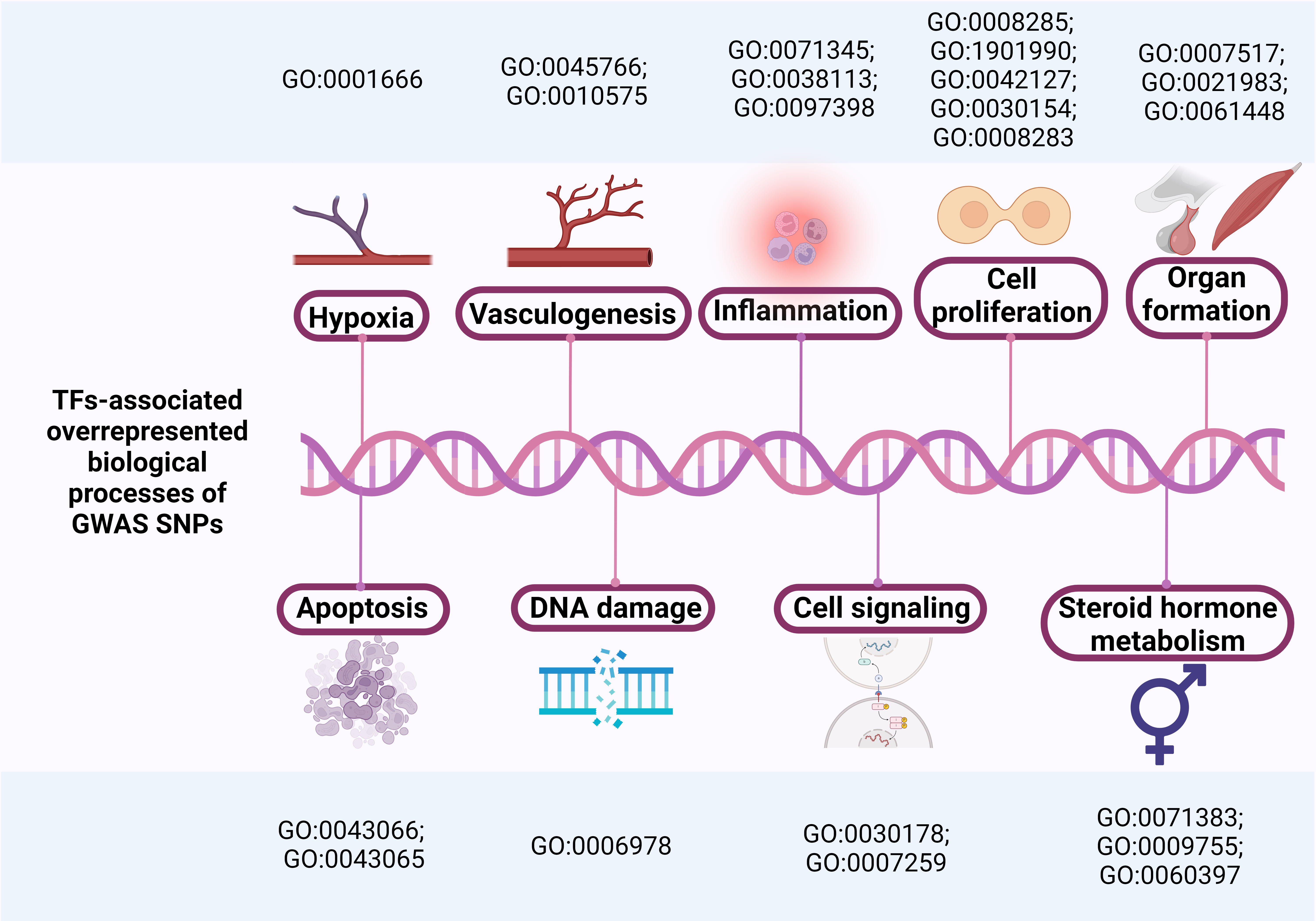 Fig. 4.
Fig. 4.
Overrepresented biological processes associated with TFs binding to GWAS SNPs.
According to our data, rs547025 SIRT3, previously described as reducing the likelihood of UF [47, 48], was also linked with a decreased risk of UF in the present study in the entire group, regardless of the history of medical abortions, but this was modified by environmental risk factors such as smoking and the level of consumption of fresh vegetables and fruits. It is interesting that, on the one hand, these environmental factors themselves act as significant regulators of cellular oxidative stress and modifiers of associations of genetic markers with the risk of developing multifactorial human diseases [49, 50, 51, 52]; on the other hand, it has been shown that rs547025 SIRT3 has the capacity to impact tumor cell proliferation and progesterone production by controlling the response to oxidative stress [28].
SIRT3 is known as a regulator of oxidative stress, and SIRT3 mRNA levels are upregulated by reactive oxygen species [28]. However, SIRT3 also positively regulates the expression of genes associated with folliculogenesis, luteinization, and progesterone secretion in human ovarian tissue [53]. Thus, high levels of oxidative stress in patients with low consumption of fresh vegetables and fruits might serve as a factor in increasing SIRT3 expression and driving enhanced progesterone release, a substantial risk factor for the UF [54, 55, 56]. In contrast, lower SIRT3 levels lead to decreased mRNA expression of many agents associated with steroidogenesis and, as a consequence, potentially lower levels of progesterone secretion in patients with normal levels of fresh vegetable and fruit consumption, which explains the protective effect of rs547025 SIRT3 in this group of patients.
Surprisingly, we observed an association of rs547025 SIRT3 specifically in smokers, despite the fact that smoking is known to increase reactive oxygen species (ROS) levels [57]. However, tobacco smoke pollution has been shown to decrease SIRT3 mRNA expression [58], and benzapyrene, a component of cigarette smoke, resulted in increased methylation of SIRT3 3′ UTR [59]. Given that increased methylation is a regulating cause of decreased gene expression, this finding provides support for the protective effects of smoking on UF. In addition, a fairly large number of studies indicate the protective effects of smoking on UF development [7, 22, 60], likely due to the influence of smoking on hormone levels. Women who smoke tend to exhibit lower urinary estrogen levels during the luteal phase compared to non-smokers [61], as nicotine can inhibit aromatase activity, reducing the conversion of androgens to estrone [62]. Thus, smoking is associated with impaired production and reduced levels of endogenous circulating estrogens [22], suggesting that these effects may be mediated through the influence of rs547025 SIRT3 in smokers. Thus, it is possible that the protective impact of SIRT3 is particularly pronounced in smokers, possibly through its role in regulating estrogen levels [63].
We also observed a protective effect of rs547025 SIRT3 against UF exclusively in patients without a history of spontaneous abortion. Spontaneous abortion is diagnosed up to 22 weeks of pregnancy, therefore, the level of estrogen and progesterone at these stages of pregnancy is many times higher in a pregnant woman until the moment of miscarriage [64]. Most likely, additional factors, such as a more dramatic decrease in steroid sex hormone levels relative to baseline after placental abruption [7], as well as the highly invasive nature of the method for removing fetal remnants, which entails severe local inflammation [33], are strong risk factors for the development of UF, exceeding the effect of rs547025 SIRT3.
The existence of pelvic inflammatory disorders also altered the relationship between rs547025 SIRT3 and the UF risk; rs547025’s protective effect was detected only in women who had no history of pelvic inflammatory diseases. Most likely, inflammatory diseases, which can be a risk factor for the development of uterine fibroids, according to many previous studies [9, 65], may exceed the protective effect of rs547025 SIRT3 due to a more significant contribution to the pathogenesis of the disease.
The bioinformatics analysis assisted in understanding the functional impact of rs547025 SIRT3. Thus, rs547025 SIRT3 is characterized by a cis-eQTL effect on reducing the expression of RIC8A in subcutaneous fat and blood. RIC8A has already been described as a risk factor for UF in Japanese [15] and European [14, 66] populations.
In blood, rs547025 SIRT3 affects expression levels of PSMD13, previously identified as a risk factor for UF in GWAS [13, 67] and transcriptome-wide association study (TWAS) [68] studies. PSMD13 is thought to be linked to platelet count, which acts as a mediator in immunological and inflammatory responses [69]. A cis-eQTL-mediated effect of rs547025 SIRT3 on the reduction of BET1L (Bet1 Golgi vesicular membrane trafficking protein like) gene expression was also found, implicated in UF risk and UF volume, as previously described in Chinese [70], European American [71], and Taiwanese [72] populations. The role of the rs547025-cis-eQTL-linked gene SCGB1C1 in UF risk was described by Sakai K. et al. [16] in 2020; altered ODF3 gene expression has been shown in various tumors of the reproductive system [73, 74]. IFITM2, IFITM1, and IFITM3, also regulated by rs547025 SIRT3 via cis-eQTL effects, are described by P. Cha as risk factors for UF [69]. Cis-eQTL-linked gene PTDSS2 was also implicated in tumorigenesis [75, 76].
Bioinformatics analysis of data from the Reproductive System Knowledge Portal
showed associations of rs547025 and strongly linked (r2
Our study also found that rs7907606 STN1, SLK was associated with an increased risk of UF in patients without pelvic inflammatory disease. Inflammation is a significant risk factor for uterine fibroids. Trauma, infection, and subsequent inflammation cause an imbalance in the immune system by increasing T-helper cytokines and decreasing target cell function [78], and the inflammatory response of the immune system leads to the formation and proliferation of fibrous tissue [34, 79]. Most likely, the contribution of inflammation to the formation of this trait exceeds the contribution of rs7907606 STN1, SLK, explaining the lack of link in patients with a history of PID.
The association between rs7907606 STN1, SLK and the risk of UF has already been described by Cha P.C. et al. [67], Ishigaki K. et al. [80] and Masuda T. et al. [15] in the Japanese population, as well as by Edwards T.L. et al. [81] in the European and African populations. Rafnar T. et al. [48] and Välimäki N. et al. [82] also described the association between rs7907606 STN1, SLK and UF risk in the European population. This SNP affects the expression of various genes in the blood: increases the expression of SH3PXD2A and GSTO1, which, according to the Reproductive System Knowledge Portal, are associated with the risk of UF with a moderate and very strong evidence range (HuGE score levels 3.0 (https://reproductive.hugeamp.org/gene.html?gene=SH3PXD2A) and 45.0 (https://reproductive.hugeamp.org/gene.html?gene=GSTO1), respectively; TPRIP, which was also noted in the impact on UF risk [82]. Furthermore, modified OBFC1 expression as a risk factor for UF has been described in the studies of Edwards T. L. et al. [81], Rafnar T. et al. [48] and Välimäki N. et al. [82].
The analysis of transcription factors revealed that the risk allele G rs7907606
STN1, SLK creates DNA binding sites for TFs involved in cell
differentiation and proliferation, which provides further evidence for its
possible role in tumorigenesis and UF development [83, 84] (Supplementary
Table 5). According to the bioinformatics resource “Reproductive System
Knowledge Portal”, rs7907606 STN1, SLK, as well as six SNPs
strongly linked to it (r2
Analysis of gene-gene interactions through the MDR method revealed that SNPs
associated with UF at the stage of single genetic variant analysis (rs547025
SIRT3 and rs7907606 STN1, SLK) possessed the greatest
pronounced mono-effect (0.71% and 0.52% contribution to UF entropy,
respectively) and effect of gene-gene interactions with each other (0.60%
contribution to entropy). G
The association between SNP rs2456181 ZNF346 and the risk of UF was described by Gallagher C.S. et al. [14] and Sliz E. et al. [85]. Bioinformatics analysis revealed that rs2456181 ZNF34 is characterized by a cis-eQTL effect on the expression level of genes that have shown an association with UF in previous studies: UIMC1 [66], HK3 [48], and FGFR4 [14]. The role of FGFR4 in the regulation of cell apoptosis was noted in studies of uterine sarcoma [86]. Transcription factor analysis showed that the SNP G allele creates DNA binding sites for 45 TFs, co-controlling positive regulation of angiogenesis, regulation of cell population proliferation, the interleukin-9-mediated signaling pathway, the growth hormone receptor signaling pathway via JAK-STAT, the cellular response to interleukin-17, positive regulation of vascular endothelial growth factor production, and the cell surface receptor signaling pathway via JAK-STAT (Supplementary Table 6). Interestingly, one study found a significant decrease in circulating growth factors, including VEGF and IL-7, in women with UF [87]. In contrast, during UF development, there is an increase in growth factors associated with hematopoiesis and angiogenesis in the myometrial and leiomyoma tissues [88, 89]. Additionally, JAK-STAT pathway activation in leiomyoma cells may drive extracellular matrix production, a key feature of UF [90, 91].
Numerous studies have linked rs117245733 LINC00598 to the risk of UF [14, 48, 85, 92]. LINC00598 (Long Intergenic Non-Protein Coding RNA 598) is an RNA gene, and is affiliated with the lncRNA class (https://www.genecards.org/cgi-bin/carddisp.pl?gene=LINC00598&keywords=LINC00598). Our bioinformatics analysis showed that allele A rs117245733 LINC00598 creates DNA binding sites for 37 TFs, jointly involved in cellular response to steroid hormone stimulus and negative regulation of the Wnt signaling pathway. This suggests a significant role for this SNP in UF, as Wnt signaling is known to promote the formation and growth of fibroids [93, 94], while steroid hormones are fundamental in UF pathogenesis [56, 95]. Reference allele G rs117245733 creates DNA binding sites for 32 TFs, regulating the hormone-mediated signaling pathway, positive regulation of the apoptotic process, and muscle organ development. According to bioinformatics analysis of the Reproductive System Knowledge Portal, rs117245733 LINC00598 increases the risk of UF and heavy menstrual bleeding.
Finally, we found the association of rs72709458 TERT, which was previously described in the study of Gallagher C.S. et al. [14] as a risk factor for UF. According to the bioinformatic resource “Reproductive System Knowledge Portal”, rs72709458 TERT increases the risk of UF and heavy menstrual bleeding. Carriage of the SNP T allele rs72709458 TERT results in loss of DNA binding to TFs that are jointly involved in DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator, response to hypoxia, negative regulation of cell population proliferation, regulation of mitotic cell cycle phase transition, cellular response to cytokine stimulus, and negative regulation of the apoptotic process. These processes are particuraly interesting, as they are closely intertwined and significantly contribute to UF development [96]. The tumor suppressor protein p53, for instance, can be down-regulated by estradiol, illustrating a primary mechanism through which estrogen promotes UF growth [97]. p53 regulates the cell cycle by influencing factors related to cell proliferation and apoptosis, such as p21 and Bax [98, 99, 100, 101]. When estrogen reduces p53 levels, apoptosis is suppressed, thereby enabling tumor proliferation.
First, we were limited in the number of SNP markers studied. Second, we chose the genotyping technique using Taq-man probes, and therefore several SNPs were excluded from the analysis due to methodological problems. Third, we were limited in the availability of a sample of patients with UF from another Russian population, which prevented us from performing a replication study of our associations. Fourth, we did not have information on the level of consumption of fresh vegetables and fruits in the control group, so this environmental risk factor was not included in MB-MDR. Fifth, some clinical data were missing in some patients, and also belonged to the category of low-frequency factors, and therefore they could not be assessed in terms of gene-environment interactions and the influence of SNPs on clinical manifestations.
In conclusion, our study replicated the associations of GWAS-significant loci with UF in Russians. For the first time, we showed that GWAS loci interact substantially with medical and spontaneous abortion history in terms of associations with UF risk, that may additionally be significantly altered by risk factors such as pelvic inflammatory disorders, smoking, and fruit/vegetable intake. Comprehensive bioinformatics analysis revealed that the molecular mechanisms of involvement of GWAS SNP in UF risk are determined by their role in the regulation of cell proliferation, inflammation, apoptosis, hypoxia, vasculogenesis, DNA damage, and cell signaling, all of playing crucial parts in the pathogenesis of uterine fibroids. Unlike previous studies focused primarily on identifying GWAS-significant loci, our findings emphasize that environmental factors can modify associations with the risk of UF. This integrated approach to assessing genetic and environmental contributions may advance personalized medicine for UF and open new possibilities for applying GWAS findings to the prevention and management of uterine fibroids in particular and of multifactorial human pathology in general.
All data reported in this paper will also be shared by the corresponding author, upon reasonable request.
OB designed the research study. LP, KK, and OB performed the research. LP, KK, and OB analyzed the data. LP and OB wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The research protocol was approved by the Ethics Committee of Kursk State Medical University (protocol No. 5, from May 11, 2021), and all patients or their families/legal guardians provided signed informed consent. The study was carried out in accordance with the guidelines of the Declaration of Helsinki.
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/j.fbs1604024.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.




