1 Department of Rehabilitation Therapy, The Second Rehabilitation Hospital of Shanghai, 200441 Shanghai, China
2 Department of Rehabilitation Medicine, Shanghai Pudong New Area People’s Hospital, 201299 Shanghai, China
3 Institute of Rehabilitation Medicine, Shanghai Academy of Traditional Chinese Medicine, 201203 Shanghai, China
4 School of Rehabilitation Science, Shanghai University of Traditional Chinese Medicine, 201203 Shanghai, China
†These authors contributed equally.
Abstract
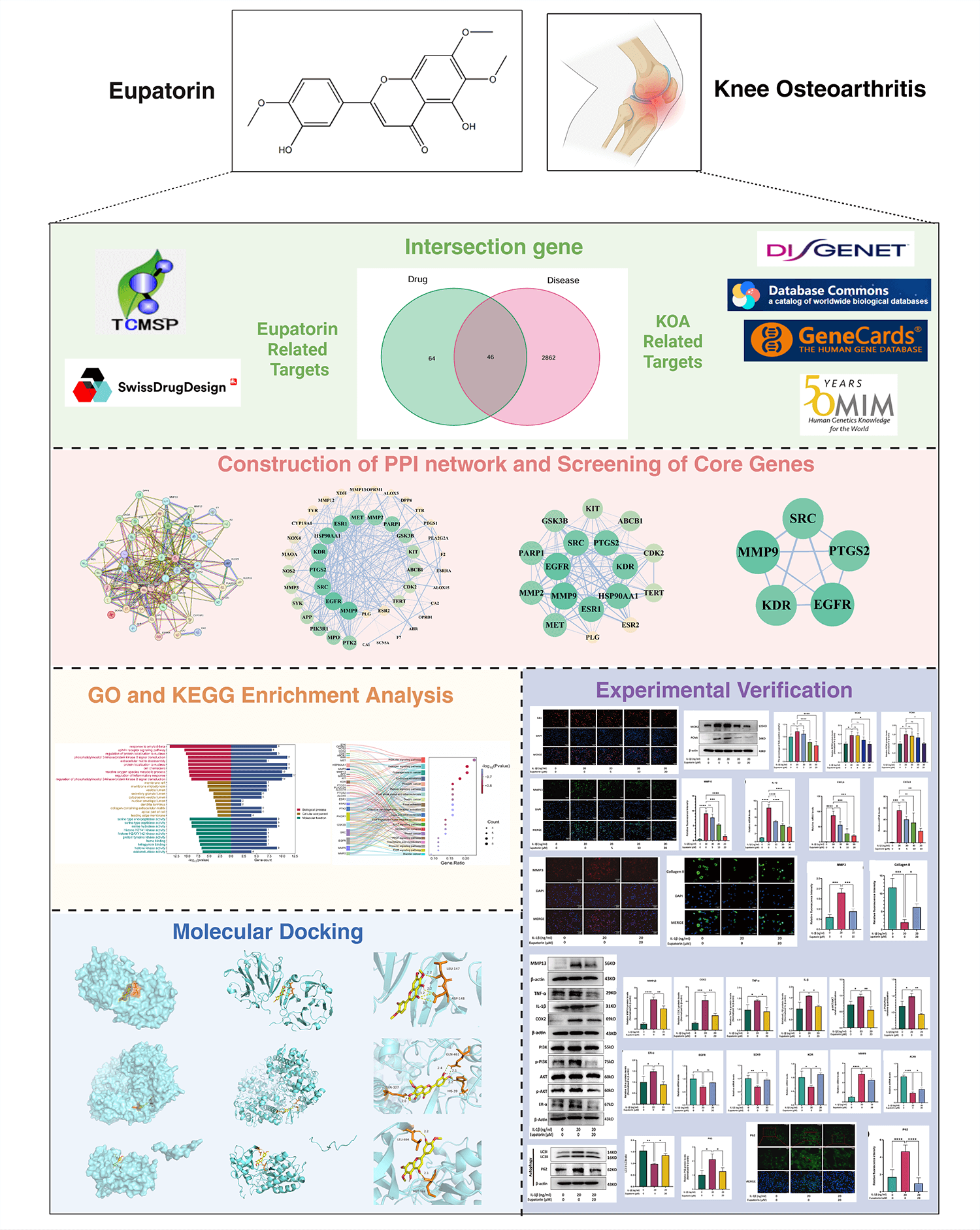
Knee osteoarthritis (KOA), a chronic degenerative joint disease, is primarily driven by inflammation-induced cartilage degradation, which represents its core pathological feature. Eupatorin, with its distinct anti-inflammatory properties, has emerged as a promising candidate for KOA research. This study aimed to explore the therapeutic potential of Eupatorin and elucidate its underlying mechanisms in KOA through an integration of network pharmacology analysis and experimental validation.
Potential targets of Eupatorin and KOA-related genes were retrieved from multiple databases, and the overlapping targets were utilized to build a protein‒protein interaction (PPI) network to identify core targets. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed to characterize the associated biological processes (BP), molecular functions (MF), and cellular components (CC). Additionally, molecular docking was performed to assess the binding affinities of Eupatorin with the core targets. Direct target engagement was confirmed using a cellular thermal shift assay (CETSA). Finally, biological experiments using interleukin-1β (IL-1β)-stimulated primary rat chondrocytes were carried out to validate the protective effects of Eupatorin through its anti-inflammatory activity.
Network pharmacology analysis revealed 46 overlapping targets, with Matrix Metallopeptidase 9 (MMP9), Epidermal Growth Factor Receptor (EGFR), and Prostaglandin G/H synthase 2 (PTGS2) as key nodes within the PPI network. GO and KEGG enrichment analyses revealed significant associations with inflammatory responses and extracellular matrix (ECM) metabolism, particularly the phosphatidylinositol 3-kinase (PI3K)/protein kinase B (AKT) and estrogen signalling pathways. Molecular docking further confirmed strong binding affinities between Eupatorin and key targets, including MMP9, EGFR, and PTGS2. CETSA validated the direct binding of Eupatorin to PTGS2. Eupatorin significantly inhibited IL-1β-induced cytokine expression and ECM degradation while promoting ECM synthesis and restoring impaired autophagy in inflamed chondrocytes, as indicated; however, no significant effect on cellular senescence was observed. Mechanistically, Eupatorin exerted its protective effects on chondrocytes by attenuating the upregulation of the PI3K/AKT and estrogen signalling pathways.
Eupatorin has demonstrated potential for use in KOA therapy by targeting inflammation and ECM, and by regulating the PI3K/AKT and estrogen-associated signaling pathways.
Keywords
- Eupatorin
- knee osteoarthritis
- network pharmacology
- chondrocytes
- inflammation
- extracellular matrix
Osteoarthritis (OA) represents a widespread degenerative disorder of the entire joint that impairs all articular tissues [1]. Pathologically, OA involves not only articular cartilage degeneration but also changes such as meniscal tears, inflammation/fibrosis of the infrapatellar fat pad, and synovial hyperplasia with fibrosis [2, 3]. Knee osteoarthritis (KOA) is the most common subtype of OA, with its primary features being progressive articular cartilage degeneration and secondary subchondral bone remodelling [4]. The pathogenesis of KOA is multifactorial and involves complex interactions among mechanical, biochemical, and inflammatory factors that collectively drive a series of pathological alterations in the articular cartilage, including fibrillation, fissures, ulceration, and wear [5]. In the treatment of KOA, anti-inflammatory therapy has long been considered a cornerstone strategy for alleviating pain and improving joint mobility [6]. Although emerging approaches such as stem cell therapy and ADAMTS-5 inhibition have shown considerable promise, significant challenges remain, including the risk of infection, limited capacity to restore cartilage thickness, and the potential for aberrant tissue overgrowth [7, 8, 9, 10]. These limitations underscore the need to further explore and validate novel therapeutic targets for treating KOA [11].
Cartilage degradation is a hallmark of KOA, during which chondrocytes attempt to
homeostasis by maintaining a balance of anabolic and catabolic processes [12, 13]. As key local sources of inflammatory factors, synovial tissue and the
infrapatellar fat pad (IFP) can synergistically secrete proinflammatory cytokines
including interleukin-1
Eupatorin (3′,5-dihydroxy-4′,6,7-trimethoxyflavone) is a natural
flavonoid compound predominantly isolated from diverse medicinal plants,
including Isatis indigotica fortune, Pratia nummularia, and members of the
Eupatorium genus [18]. This compound was shown to display
anti-inflammatory properties, primarily by suppressing the secretion of
inflammatory mediators through the inhibition of key signalling pathways such as
STAT1 and NF-
Given that inflammation is a critical driver of the onset and progression of
KOA, and that Eupatorin has been demonstrated to exert robust anti-inflammatory
properties in diverse disease models, we aimed to investigate its potential
protective role in cartilage under inflammatory conditions. Herein, we initially
utilized network pharmacology and molecular docking to systematically screen the
potential molecular targets and key signalling pathways through which Eupatorin
may exert its therapeutic effects in KOA. We then established an
IL-1
The core targets of Eupatorin were initially screened using the Traditional Chinese Medicine Systems Pharmacology Database (TCMSP 2.0; https://www.tcmsp-e.com/load_intro.php?id=43) [23]. The corresponding SMILES identifier for Eupatorin was retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih.gov) [24] and subsequently entered into the Swiss Target Prediction platform (http://swisstargetprediction.ch/) to predict potential targets, and only targets with a probability greater than zero were retained [25]. All of the predicted targets were normalized by converting their protein names to official gene symbols via the UniProt database (https://www.uniprot.org/) [26]. After removing duplicate entries, a list of potential target genes for Eupatorin was constructed.
The Medical Subject Headings (MeSH) terms “Knee Osteoarthritis” and “Osteoarthritis, Knee” were used to search for disease-associated target genes across multiple databases, including DisGeNET (https://www.disgenet.org/), GeneCards (https://www.genecards.org/), Online Mendelian Inheritance in Man (OMIM; https://omim.org/), and the Therapeutic Target Database (TTD; http://db.idrblab.net/ttd/). The obtained gene targets were subsequently normalized and converted into matching UniProt IDs through the UniProt database (https://www.uniprot.org/). Finally, R software (Version 4.4.2, R Core Team 2024, R Foundation for Statistical Computing, Vienna, Austria. Available from: https://www.R-project.org/) and the “Venn diagram” package were applied to identify the overlap of genes between Eupatorin-targeted genes and KOA-related genes, and a Venn diagram was generated.
The intersecting targets of Eupatorin and KOA were submitted to the STRING database (https://string-db.org), where the organism was specified as Homo sapiens and a medium confidence score was set as the minimum interaction threshold. Upon completion of the analysis, the resulting interaction data were exported in TSV format for downstream processing. A PPI network was subsequently constructed with Cytoscape (Version 3.9.1, Institute for Systems Biology, Seattle, WA, USA, Available from: https://www.cytoscape.org/). To enhance the comprehensiveness and accuracy of the network analysis, we employed two Cytoscape plugins, CytoNCA and CytoHubba [27, 28]. Using CytoNCA, centrality metrics for each node (including degree, betweenness centrality, and closeness centrality) were calculated to evaluate node importance within the network. Nodes with degree, betweenness centrality, and closeness centrality values exceeding the respective median thresholds were retained through two successive rounds of filtering to identify core targets. In parallel, CytoHubba was used to predict key genes and subnetworks on the basis of the maximal clique centrality (MCC) algorithm. During visualization, node size was scaled proportionally to its degree value (larger nodes indicating higher degrees), and node colour intensity reflected target influence (darker colours indicating greater influence).
For the purpose of further investigating the potential biological roles of the identified targets, using the R package “ClusterProfiler”, we conducted Gene Ontology (GO) enrichment analysis, including the three subcategories of biological process (BP), molecular function (MF), and cellular component (CC) subcategories, along with Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis [29]. Specifically, enrichment terms were filtered on the basis of p values, and the top 10 significant GO terms and the top 20 significant KEGG pathways were selected for in‑depth analysis. Finally, the results were visualized as bar charts and Sankey diagrams using an online bioinformatics platform to intuitively demonstrate the distribution and proportional flow of intersecting genes across different pathways (http://www.bioinformatics.com.cn/).
The 2D chemical structure file (SDF format) of Eupatorin was downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov). The minimum binding energy conformation of Eupatorin was calculated using ChemBio3D version 22.0.0.22 and saved in MOL2 format. Three-dimensional structures of the core target proteins Matrix Metallopeptidase 9 (MMP9) (PBD code: 1ITV), EGFR (PBD code: 2XKN), and PTGS2 (PBD code: 3OLT) were downloaded from the Protein Data Bank (PDB; https://www.rcsb.org). All protein structures underwent preparation through the removal of solvents and organics using PyMOL (Version 3.1, Schrödinger., New York, NY, USA, Available from: https://pymol.org/). The protein receptor grid box files were generated with AutoDockTools (Version 1.5.7, Scripps Research, La Jolla, CA, USA. Available from: https://ccsb.scripps.edu/), and molecular docking between Eupatorin and individual target proteins was carried out with AutoDock Vina. All the docking run options were default parameters (exhaustiveness = 10, num_modes = 20, and energy range = 5.0 kcal/mol). Finally, the binding patterns and key intermolecular interactions of candidate active compounds with core target proteins were evaluated and visualized in PyMOL version 4.6.0.
One-day-old Sprague–Dawley (SD) rats (Shanghai Sipur-Bikai Laboratory
Animal Co., Ltd. [Certificate No. SCXK (Shanghai) 2018-0006] licence no. SCXK
(Shanghai) 20180006) were euthanized by inhalation of an overdose of isoflurane
(at a concentration of 5%) and immediately immersed in 75% ethanol for 10
minutes. Subsequent to three consecutive washes using phosphate-buffered saline
(PBS), all the procedures were performed under sterile conditions. Articular
cartilage was carefully harvested from the tibial and femoral condyles and rinsed
extensively with PBS. Cartilage tissues were minced into approximately 1 mm3
fragments and transferred into 0.5% type II collagenase solution (SigmaAldrich,
C6885, St. Louis, MO, USA) for enzymatic digestion at 37 °C with gentle
agitation (150 rpm) for 60 minutes. The digestion mixture was subsequently
harvested and subjected to centrifugation at 1500
Primary chondrocytes were first plated into 96-well plates at a density
5
Thereafter, the chondrocytes were treated with increasing concentrations of
Eupatorin (TargetMol, T7032, Boston, MA, USA) (2, 5, 10, 20, and 50
µM). Simultaneously, blank wells (culture medium only) and vehicle
control wells (equivalent volume of DMSO) were included. Following 24 and 48 h of
culture, 10 µL of CCK-8 solution was added to every well prior to
incubation for 1–4 h at 37 °C. The absorbance at 450 nm (OD450)
was detected with a microplate reader, after which the cell viability was
computed using the following equation: cell viability (%) =
(ODsample-ODblank)/(ODcontrol-ODblank)
In accordance with the manufacturer’s instructions, chondrocyte proliferation
was assessed via a 5-ethynyl-2′-deoxyuridine (EdU) incorporation assay
(BeyoClick™ EdU Cell Proliferation Kit with AF555, Beyotime,
C0075). Briefly, cells were seeded in 24-well plates at 1
Primary articular chondrocytes were assigned to three groups: the non-intervened
group (untreated), the model group (20 ng/mL IL-1
Total RNA was isolated from chondrocytes of each experimental group using the RNAeasy™ Animal RNA Isolation Kit (column-based) (Beyotime, R0024) in accordance with the manufacturer’s instructions. Complementary DNA (cDNA) was synthesized via the Evo MMLV Reverse Transcription Kit (Accurate Biotechnology, AG11706, Hunan, China). qPCR was performed with the SYBR® Green Pro Taq HS Premix qPCR Kit (Accurate Biotechnology, AG11746) in a 20 µL reaction system comprising10 µL of SYBR Green Pro Taq HS Premix IV, 1 µL of forward and reverse primers (each), 5 µL of cDNA template, and 4 µL of RNase-free water. PCRs were performed on a QuantStudio 5 real-time PCR system (Applied Biosystems, Foster City, CA, USA). Relative mRNA expression levels were calculated using the 2-ΔΔCt method. qPCR primer sequences are provided in detail in Table 1.
| Gene | Primer Sequence (5′-3′) | Product Length (bp) |
| Forward: GCTACAGCTTCACCACCACA | 94 | |
| Reverse: GCCATCTCTTGCTCGAAGTC | ||
| IL-1 |
Forward: GACTTCACCATGGAACCCGT | 104 |
| Reverse: GGAGACTGCCCATTCTCGAC | ||
| CXCL3 | Forward: ACCAGCCTTCAGGGACTGT | 140 |
| Reverse: GGCTATGACTTCTGTCTGGGT | ||
| CXCL6 | Forward: TTCTGCTGCTGTTCACACTG | 233 |
| Reverse: TATCAACGGAGCTTGTGGGT | ||
| MMP9 | Forward: GATCCCCAGAGCGTTACTCG | 132 |
| Reverse: GTTGTGGAAACTCACACGCC | ||
| SOX9 | Forward: CAGACCAGTACCCGCATCTG | 77 |
| Reverse: GCTCTCGTTCAGCAGTCTCC | ||
| ACAN | Forward: GCCTCTCAAGCCCTTGTCTG | 101 |
| Reverse: GATCTCACACAGGTCCCCTC | ||
| EGFR | Forward: CCTATGGGCCAAAGATCCCA | 324 |
| Reverse: CCTTGATGGCCACAGGATTT | ||
| KDR | Forward: CAGCTCGTCATCCTAGAGCG | 197 |
| Reverse: GTTAGGTTCCGGTTCCCGTC |
IL-1
Total protein was extracted from chondrocytes using RIPA lysis buffer
(Servicebio, G2002, Wuhan, China) supplemented with a phosphorylated protease
inhibitor (Servicebio, G2007) and a serine protease inhibitor (PMSF, Servicebio,
G2008). After determining protein concentration via a BCA protein assay kit
(Thermo Fisher Scientific, 23225, Waltham, MA, USA), 10 µg of protein
sample was subjected to electrophoresis separation using the
Omni-Easy™ One-step Colour PAGE Gel Rapid Preparation Kit (Epizyme
Biotech, PG212, Cambridge, MA, USA). The separated proteins were subsequently
transferred from the gel to a PVDF membrane (MerckMillipore, IPVH00010,
Darmstadt, Hesse, Germany) and blocked with a rapid blocking solution (Epizyme
Biotech, PS108P) to prevent nonspecific binding sites. Next, the membrane was
incubated overnight with primary antibodies at 4 °C including:
Chondrocytes were fixed with Immunol Staining Fix Solution (Beyotime, P0098) for 10 minutes. Thereafter, the cells were subjected to permeabilization with 0.5% Triton X-100 and rinsed three times with PBS. The samples were then blocked with immunostaining blocking buffer (Beyotime, P0102) for 1 hour at room temperature to reduce nonspecific binding. Afterwards, the cells were incubated with primary antibodies targeting collagen II (Santa Cruz, sc52658), MMP13 (Santa Cruz, sc-30073) and MMP3 (Santa Cruz, sc21732) overnight at 4 °C, and both were diluted 1:500. The following day, the unbound primary antibodies were removed by washing the cells three times with PBST. The cells were then incubated with the appropriate fluorescently labelled secondary antibodies, namely, goat anti-rabbit IgG (H+L) cross-adsorbed secondary antibody, Alexa Fluor™ 488 (Thermo Fisher Technology, A11008, Waltham, MA, USA), Alexa Fluor® 488-conjugated goat anti-mouse IgG (H+L) (Servicebio, GB25301), goat anti-rabbit IgG (H+L) cross-adsorbed secondary antibody, and Alexa Fluor™ 555 (Thermo Fisher Technology, A21428) (diluted 1:1000), for 1 hour at room temperature. Afterwards, the nuclei were counterstained with DAPI (Proteintech, CM07245) for 10 minutes at room temperature to facilitate nuclear localization. Finally, the samples were washed three times with PBST in the dark to remove any unbound secondary antibody and excess DAPI. Fluorescence images were acquired with a digital inverted fluorescence microscope (Olympus, IX73). The mean fluorescence intensities were quantified across six randomly selected fields per sample using ImageJ software to assess target protein expression levels.
Primary rate chondrocytes were cultured to 80–90% confluence, harvested, and resuspended in ice-cold PBS supplemented with 1% PMSF. Cells were treated with Eupatorin (20 µM, dissolved in DMSO) or vehicle (DMSO) at 37 °C for 1 h. Samples were heated in a metal heat block (45 °C, 50 °C, 55 °C, 60 °C, 65 °C, 70 °C, 3 min) and then cooled (liquid nitrogen, 3 min). The lysates were prepared via three freeze‒thaw cycles, centrifuged (14,000 rpm, 15 min, 4 °C) to collect soluble fractions, and then subjected to Western blotting.
All statistical analyses and graphical visualizations were conducted with
GraphPad Prism v10.0.0 for Windows (GraphPad Software, Boston, MA, USA;
https://www.graphpad.com). One-way analysis of variance (ANOVA) was used to
compare multiple groups, with subsequent Tukey’s post hoc test for pairwise
comparisons. Statistical significance was considered present when the p
value was
In total, 110 putative targets of Eupatorin were obtained from the TCMSP and Swiss Target Prediction databases. Specifically, the TCMSP database provided target information for 14 active Eupatorin-related compounds, whereas Swiss Target Prediction yielded 100 predicted targets. After removing duplicate entries, 110 unique targets remained (Supplementary Table 1A). Concurrently, 2908 KOA-associated targets were retrieved from four disease databases (Fig. 2a and Supplementary Table 1B). To elucidate the potential mechanisms by which Eupatorin may exert therapeutic effects in KOA, overlapping targets between Eupatorin and KOA were identified via R software, and a Venn diagram was generated (Fig. 2b). The analysis revealed 46 common targets that may play critical roles in the pharmacological action of Eupatorin against KOA.
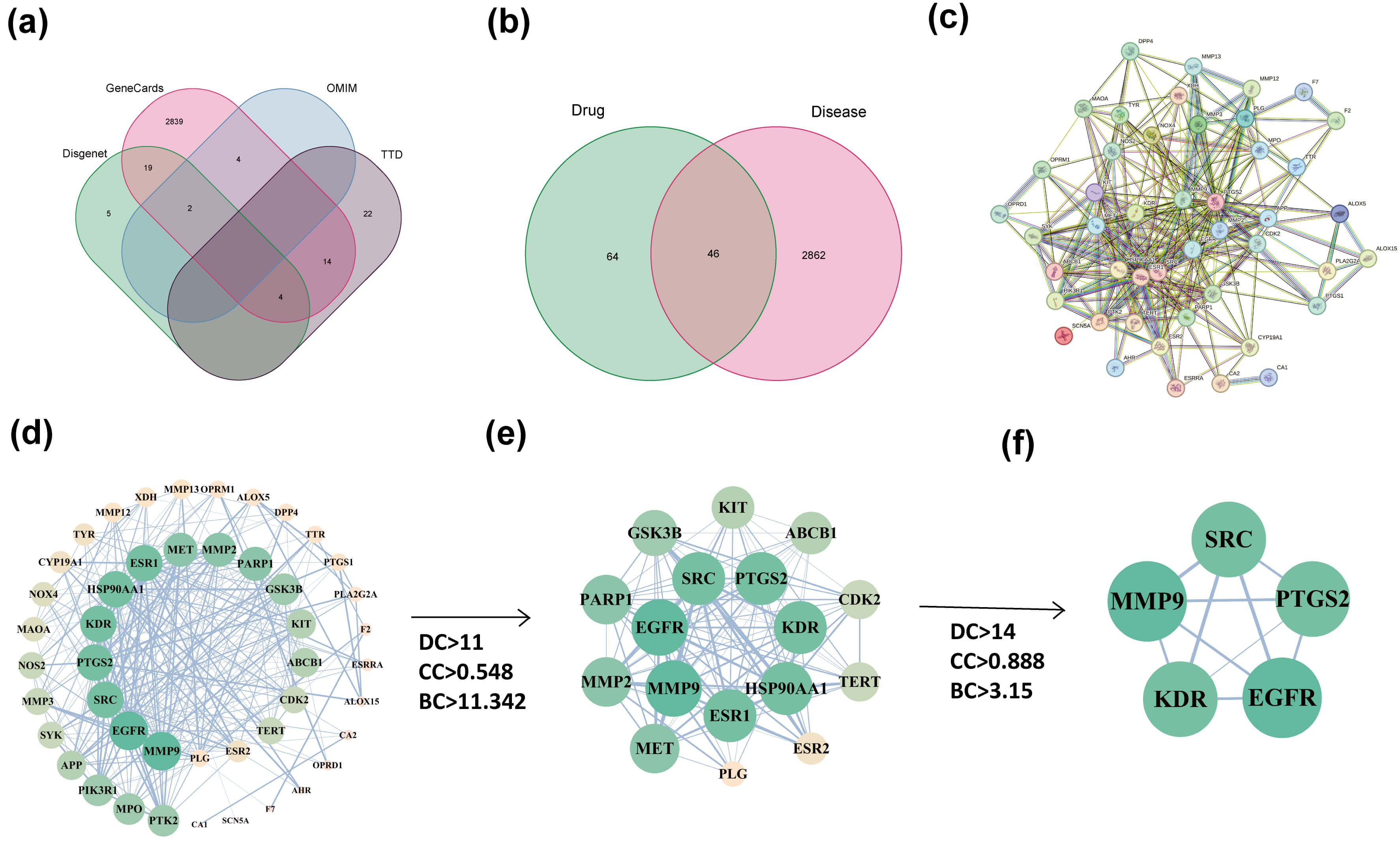 Fig. 2.
Fig. 2.
Construction of the target and PPI network of Eupatorin against KOA. (a) Targets associated with KOA. (b) Venn diagram of overlapping genes between Eupatorin and KOA. (c) PPI network and identification of hub genes. (d) The initial and hub networks were constructed on the basis of the values of DC, CC and BC. (e) Top 17 hub genes. (f) Top 5 hub genes. The arrows indicate the process of screening for hub genes using the Cytohubba plugin. Disgenet, Disease Gene Network; OMIM, Online Mendelian Inheritance in Man; TTD, Therapeutic Target Database; DC, degree centrality; CC, closeness centrality; BC, betweenness centrality.
To further identify potential targets of Eupatorin in KOA, the 46 overlapping
targets were imported into the STRING database (https://string-db.org) with the
organism restricted to Homo sapiens and a confidence score threshold of
| No. | UniProt ID | Gene Symbol | Protein name | BC | CC | DC |
| 1 | P14780 | MMP9 | Matrix metalloproteinase 9 | 230.799 | 0.762 | 31 |
| 2 | P00533 | EGFR | Epidermal growth factor receptor erbB1 | 160.349 | 0.737 | 30 |
| 3 | Q05769 | PTGS2 | Prostaglandin G/H synthase 2 | 263.268 | 0.762 | 32 |
| Cyclooxygenase-2 |
DC, degree centrality (reflecting the node’s direct connection ability); BC, betweenness centrality (measuring the node’s mediating role in network paths); CC, closeness centrality (characterizing the node’s proximity to other nodes).
To further elucidate the biological functions of Eupatorin in the treatment of KOA, functional enrichment analyses were performed. In total, 1466 GO terms were enriched, including 1285 BP, 33 CC, and 148 MF terms (Supplementary Table 4). The top-ranked enriched terms for each category are visualized in a histogram; Fig. 3a displays the top 10 enriched terms in each GO category. These results demonstrate that Eupatorin may be implicated in biological processes, including the inflammatory response, ECM catabolism, and reactive oxygen species metabolism.
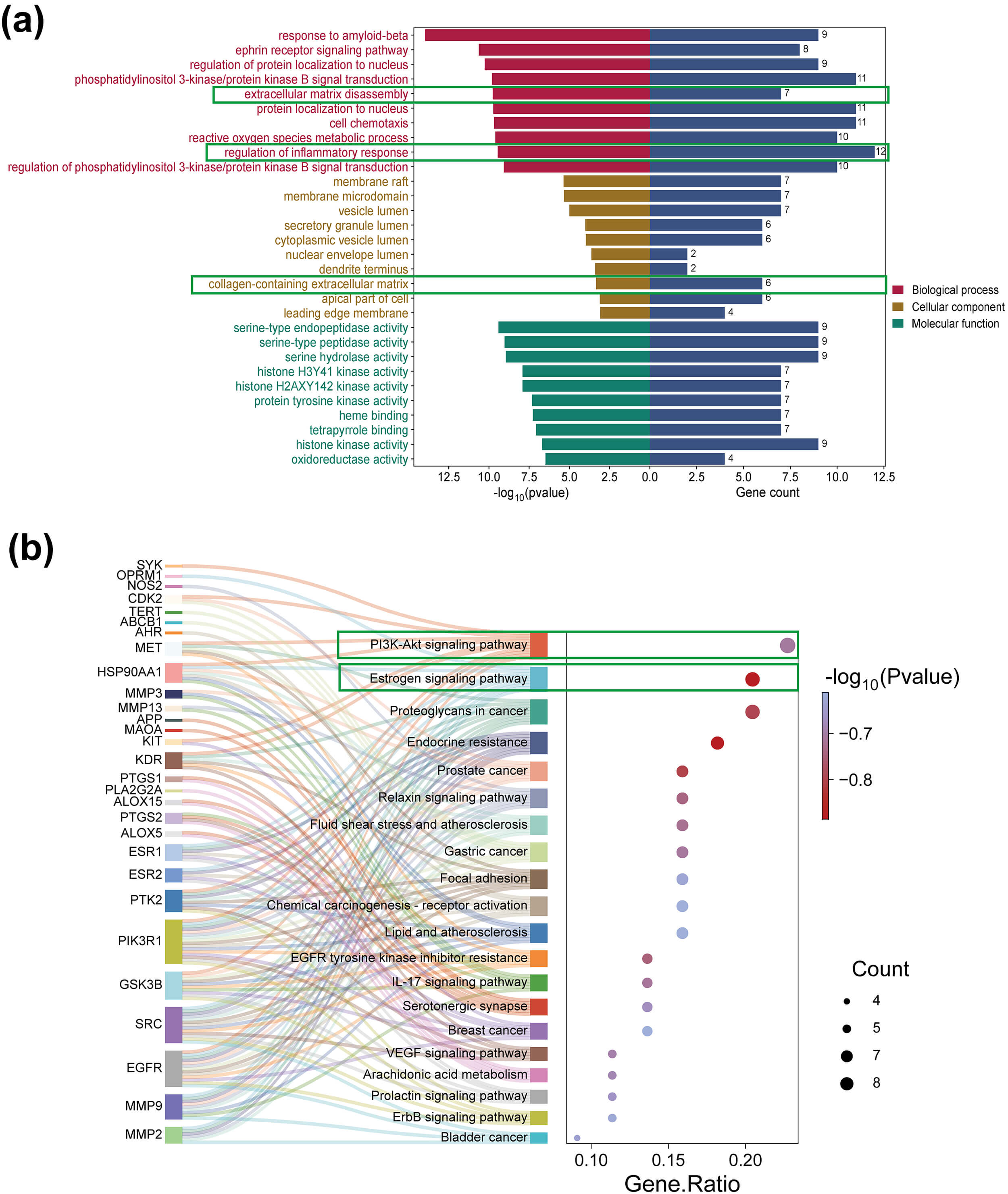 Fig. 3.
Fig. 3.
GO and KEGG analysis. (a) Bar chart of GO analysis for the enrichment of 46 targets, including BP, CC, and MF. (b) Bubble plot and Sankey diagram of the results of the KEGG enrichment analysis for the 46 targets. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes. Green boxes highlight the GO terms and KEGG pathways that were the focus of this study, which are hypothesized to be key mediators of Eupatorin’s therapeutic effects on KOA.
For pathway analysis, the enrichment analysis of the 46 core genes revealed 80
significantly enriched pathways (p
On the basis of the identified compound–target relationships, molecular docking
was performed between Eupatorin and the core targets MMP9, PTGS2 and EGFR, and in
consideration of both the binding energy values and the centrality of these
targets within the PPI network, MMP9, PTGS2, and EGFR were selected for detailed
visualization (Fig. 4). Binding energy scores
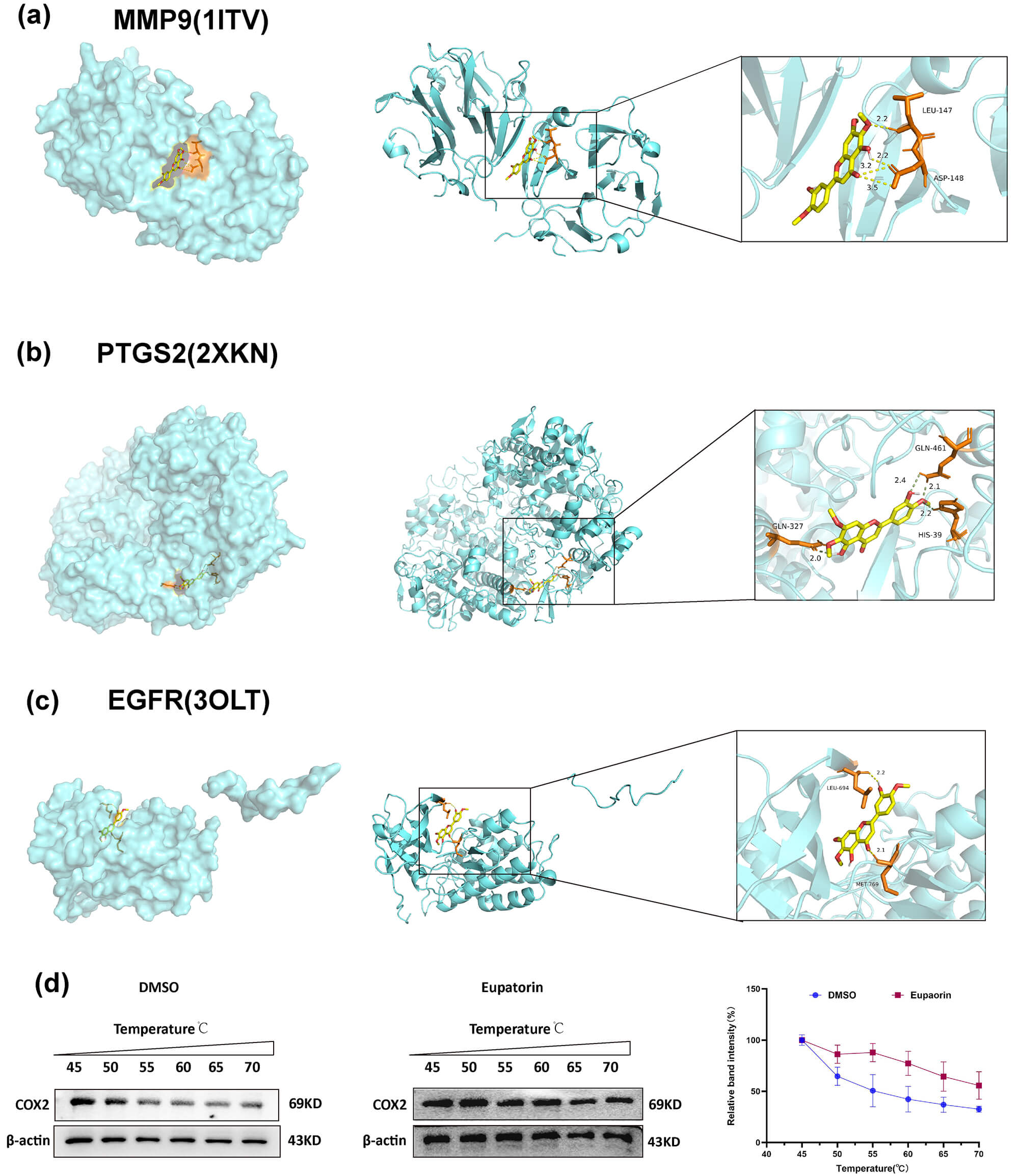 Fig. 4.
Fig. 4.
Interaction of Eupatorin with three core targets. (a)
MMP9–Eupatorin. (b) PTGS2–Eupatorin. (c) EGFR–Eupatorin. The left figure
represents the atom binding diagram of a molecule and protein. The middle figure
represents the overall binding pattern between the two, and the right figure
shows an enlarged view of the molecule and protein binding, where the text
indicates the residue name. The yellow dashed lines indicate hydrogen bonds.
(d) Eupatorin protected the COX2 protein against temperature-dependent
degeneration in chondrocytes. The values are presented as the means
| NO. | Receptors | PDB ID | Binding Energy (kcal/mol) | Centre | Size | ||||
| X | Y | Z | X | Y | Z | ||||
| 1 | MMP9 | 1ITV | –7.5 | –27.68 | –40.35 | –15.84 | 68 | 60 | 46 |
| 2 | EGFR | 2XKN | –7.8 | 6.17 | 7.48 | 57.74 | 90 | 62 | 46 |
| 3 | PTGS2 | 3OLT | –9.4 | 23.52 | 41.69 | 39.61 | 70 | 80 | 92 |
To further validate the binding interaction between Eupatorin and its highest-affinity target PTGS2 (also known as COX2; binding energy: –9.4 kcal/mol), we performed a CETSA. Compared with the DMSO control, Eupatorin treatment (20 µM) increased the thermal stability of COX2 across the temperature range (Fig. 4d). These findings provide compelling evidence that Eupatorin directly targets COX2, a core target identified through integrated network pharmacology and molecular docking approaches.
The results of the CCK-8 assay revealed that Eupatorin (
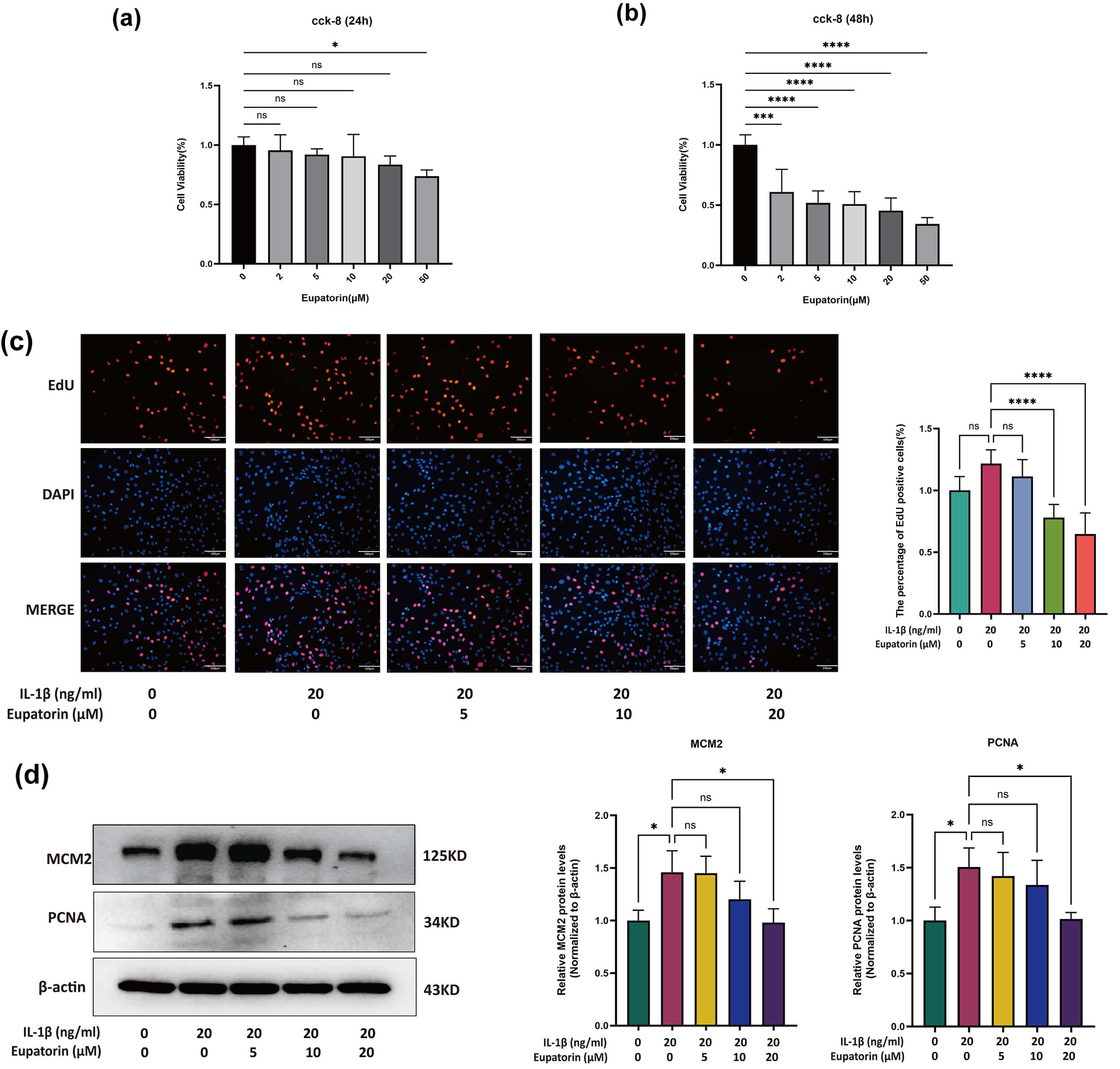 Fig. 5.
Fig. 5.
Effects of Eupatorin on chondrocyte proliferation. (a)
Chondrocyte viability was measured using the CCK-8 assay after 24 h of incubation
with Eupatorin at 2, 5, 10, 20, or 50 µM. (b) Chondrocyte viability was
measured using the CCK-8 assay after 48 h of incubation with Eupatorin at 2, 5,
10, 20, or 50 µM. (c) Proliferation of chondrocytes treated with
20 ng/mL IL-1
IF staining revealed a dose-dependent reduction in MMP13 expression in response
to 5–20 µM Eupatorin (Fig. 6a). To evaluate the regulatory effect of
IL-1
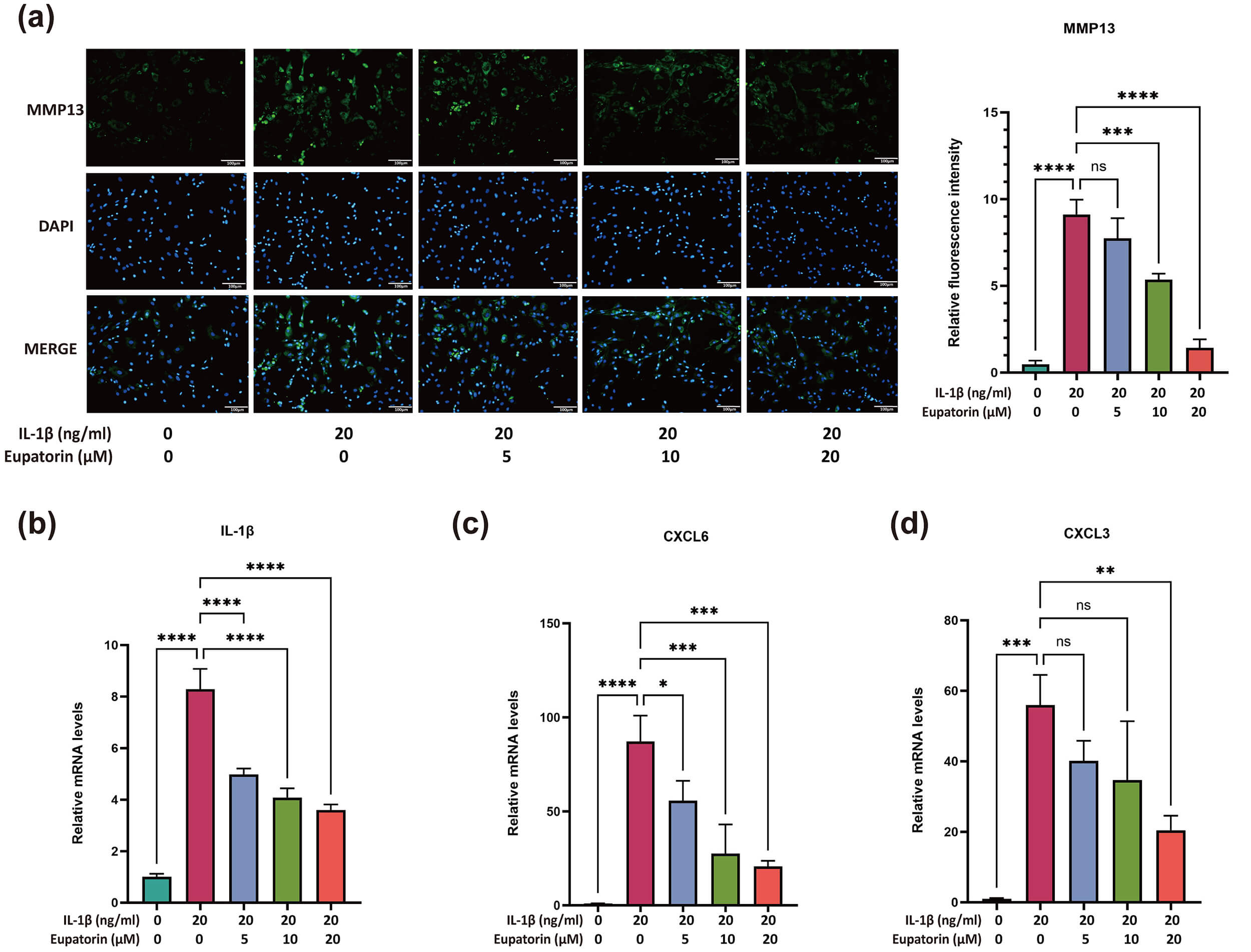 Fig. 6.
Fig. 6.
Effects of Eupatorin on chondrocyte catabolism and inflammatory
responses. (a) MMP13 expression in IL-1
Given that 20 µM Eupatorin had the strongest protective effects on chondrocyte inflammatory responses, extracellular matrix (ECM) damage, and abnormal chondrocyte proliferation without compromising cell viability, this concentration was therefore selected for subsequent investigations.
The expression of matrix degradation markers (MMP3, MMP13 and MMP9) and matrix
synthesis markers (collagen II and ACAN) was evaluated. Immunofluorescence
analysis revealed that Eupatorin significantly reduced the protein expression
level of MMP3 induced by IL-1
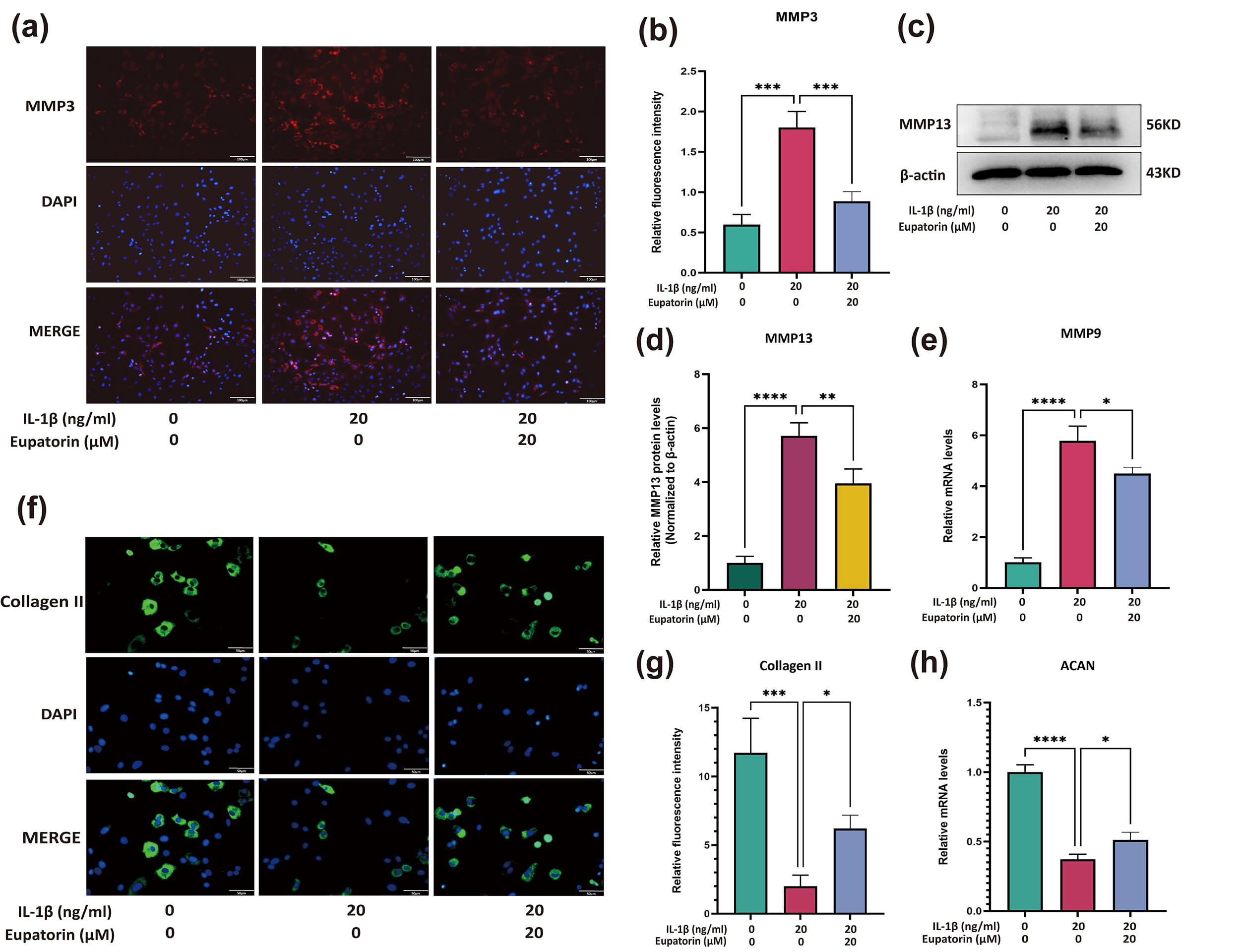 Fig. 7.
Fig. 7.
Eupatorin attenuates ECM breakdown in
IL-1
Compared with those in the control group, the protein expression levels of
TNF-
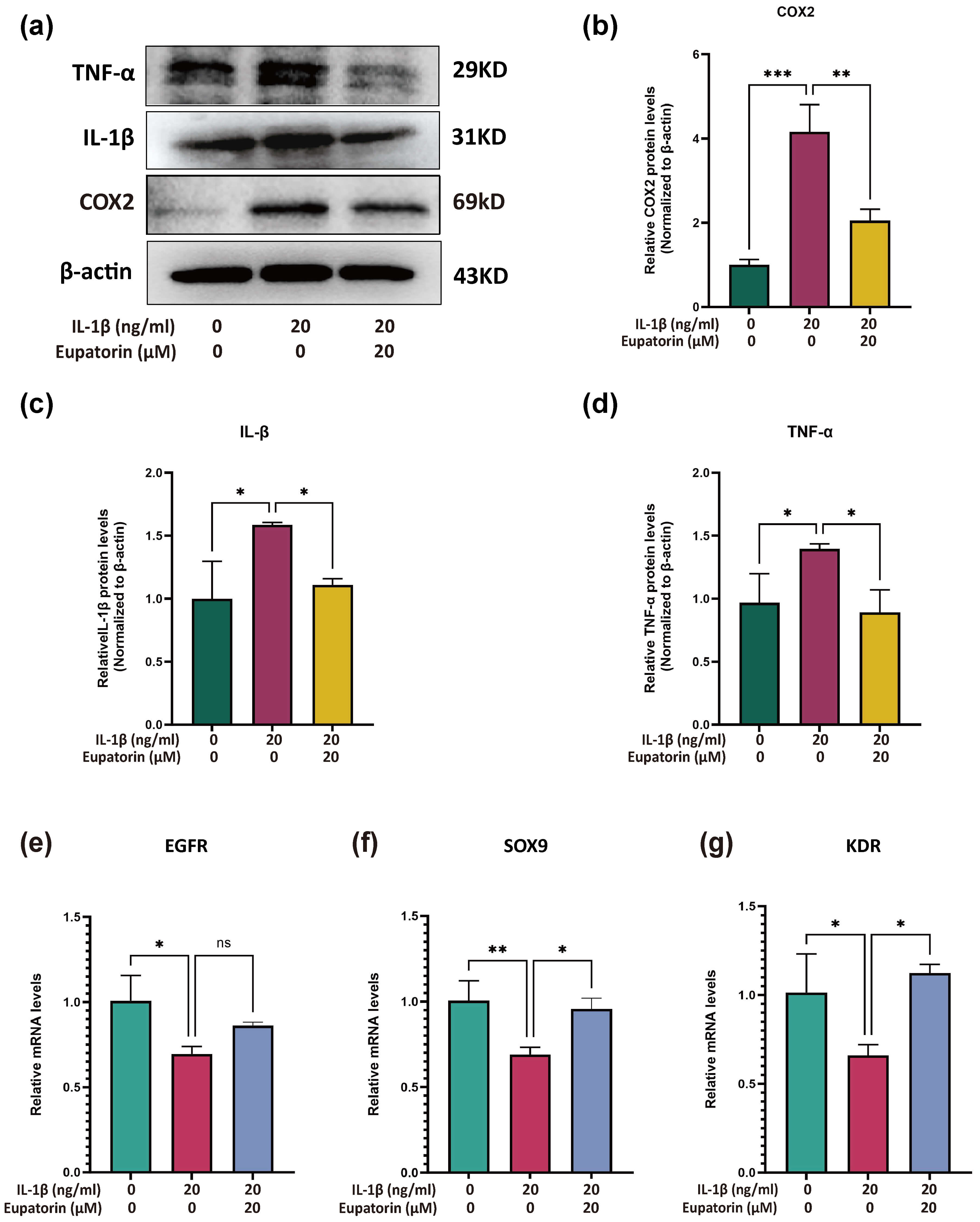 Fig. 8.
Fig. 8.
Effects of Eupatorin on IL-1
To determine whether Eupatorin also modulates key anabolic regulators in
inflamed chondrocytes, we quantified the mRNA levels of EGFR,
SOX9 and KDR by RT–qPCR. Compared with no treatment,
IL‑1
We investigated the regulatory effects of Eupatorin on the expression of PI3K,
AKT, and estrogen receptor
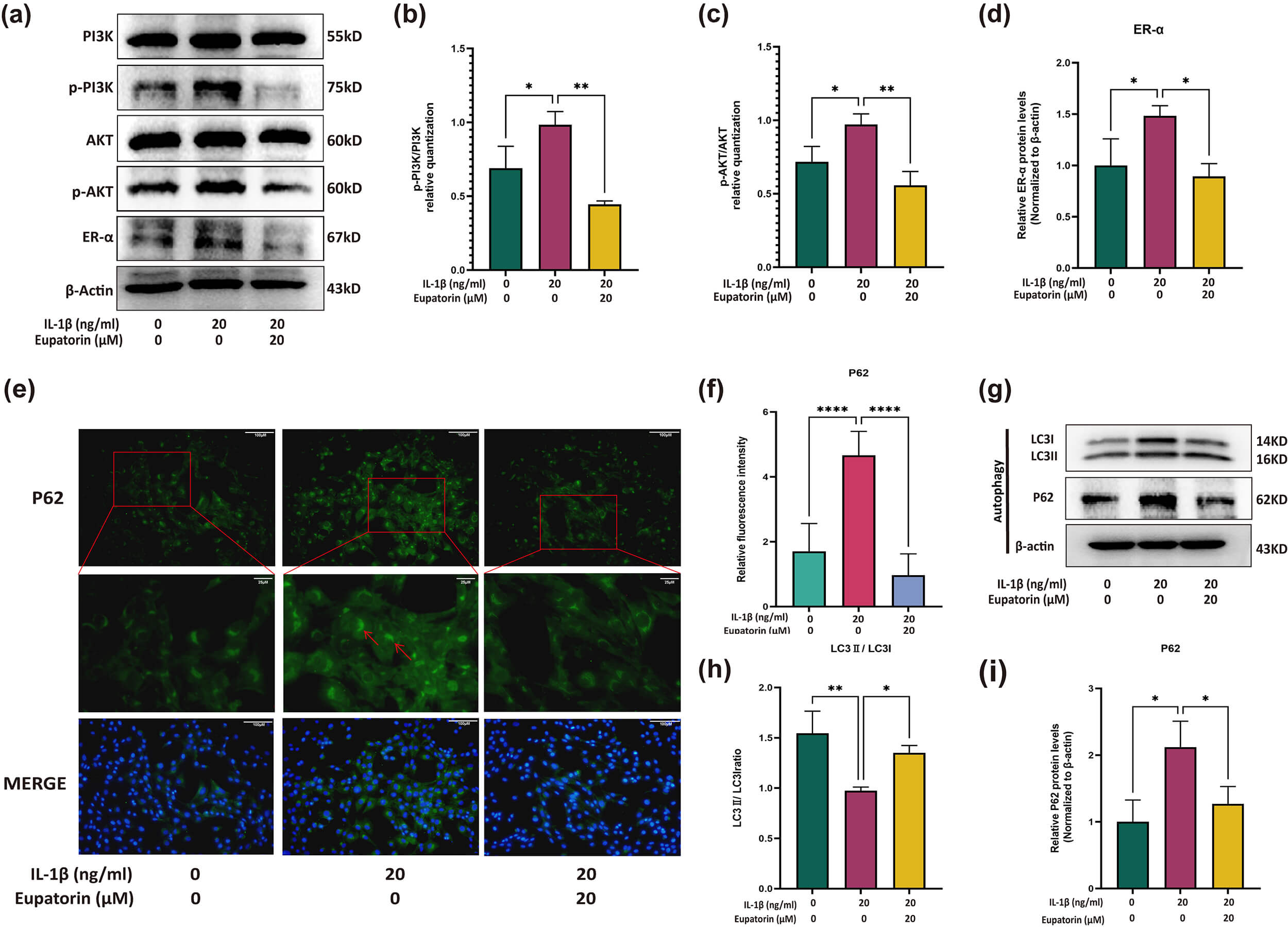 Fig. 9.
Fig. 9.
Regulatory effects of Eupatorin on PI3K/AKT, the
estrogen signalling pathway, and autophagy-related proteins. (a) Expression of
the PI3K, AKT, p-PI3K, p-AKT and ER-
Given that autophagy is a downstream event of the PI3K/AKT pathway, we therefore
examined markers of chondrocyte autophagy. IL-1
These results indicate that Eupatorin modulates the activity of the PI3K/AKT and estrogen signalling pathways, which may contribute to the functional regulation of chondrocytes.
To investigate the potential mechanisms of Eupatorin in KOA, this study applied a network pharmacology approach, and validated these predictions through molecular docking and in vitro experiments. Inflammation and ECM degradation act synergistically to drive cartilage deterioration and pain in KOA [31]. In particular, inflammatory mediators upregulate the expression of matrix-degrading enzymes, such as matrix metalloproteinases (MMPs) and aggrecanase, which cleave collagen and proteoglycans, thus accelerating ECM breakdown [32, 33]. Given that articular cartilage lacks vasculature and innervation, its repair capacity is limited, leading to the rapid accumulation of damage [34, 35]. As inflammation serves as a key pathogenic driver in KOA, by accelerating the breakdown of the cartilage extracellular matrix and amplifying pain signalling, targeting inflammation is therefore crucial for preserving cartilage structural integrity, halting disease progression, and alleviating clinical symptoms.
In recent years, natural phytochemicals have been shown to have a protective effect on cartilage tissue [36, 37]. Previous studies have demonstrated the broad pharmacological properties of Eupatorin, especially its anti-inflammatory potential [38, 39]. For instance, it has been shown to suppress the lipopolysaccharide (LPS)-induced upregulation of iNOS and COX-2 expression and to exert significant anti-inflammatory effects in carrageenan-induced mouse paw inflammation models [20]. However, its anti-inflammatory roles in the setting of OA remain largely uncharacterized.
In the present study, network pharmacology analysis demonstrated that Eupatorin
shares multiple targets with KOA, highlighting several genes closely related to
inflammation and extracellular matrix remodelling. Among them, PTGS2, MMP9, and
EGFR emerged as pivotal nodes in the PPI network. Molecular docking further
supported their potential relevance by demonstrating favourable binding
affinities with Eupatorin. Our findings indicate that Eupatorin exhibit
protective effects against IL-1
For a deeper exploration of the underlying mechanisms of Eupatorin’s protective
role in KOA, we performed GO and KEGG enrichment analyses, and the findings
revealed its involvement in pathways associated with inflammatory regulation and
extracellular matrix degradation. These processes are central to KOA
pathogenesis, as proinflammatory cytokines such as IL-1
From another perspective, KEGG analysis revealed that the PI3K-AKT pathway was
the Greatest enrichment magnitude and that the estrogen signalling pathway was
the most significantly enriched. Estrogen and its receptors are widely expressed
in joint-resident cells, suggesting that joint tissues are responsive to
estrogenic regulation [53]. ER-
It is well known that estrogen can exert biological effects through both
classical genomic and nongenomic signalling pathways [57]. Among these, the
activation of nongenomic signalling pathways involves various protein and lipid
kinases, including PI3K/AKT [58, 59]. ER-
In chondrocytes, inflammation and oxidative stress often coexist and collectively disrupt the balance of chondrocyte autophagy [66]; notably, the PI3K/AKT pathway functions as a pivotal regulator of chondrocyte autophagy, a conserved cellular process essential for maintaining intracellular homeostasis [67]. In the context of KOA, excessive PI3K/AKT activation further impairs chondrocyte autophagy through the phosphorylation and activation of mammalian target of rapamycin (mTOR), thus inducing a “damage overload” state within chondrocytes [68, 69]. In the present study, Eupatorin modulated autophagy activation, as indicated by reduced P62 expression and an elevated LC3II/LC3I ratio (Fig. 9e–i); however, our results revealed no statistically significant effect of Eupatorin on chondrocyte senescence (Supplementary Fig. 2). This observation aligns with the notion that chondrocyte senescence is typically a late-stage consequence of prolonged autophagy impairment rather than an immediate response to autophagy-related changes [70]. Alternatively, the regulatory effect of Eupatorin may be restricted to the autophagic process itself, without extending to the subsequent senescence pathway. Thus, more systematic studies are needed to confirm this. Within this context, we found that Eupatorin may exert protective actions, at least in part, by modulating PI3K/AKT activity, thus attenuating inflammatory and catabolic signalling while promoting chondrocyte autophagy in KOA.
Taken together, these findings strengthen the validity of network pharmacology
predictions and highlight Eupatorin as a potential therapeutic candidate for KOA.
Notably, Eupatorin is a bioactive flavonoid predominantly isolated from plants of
the Eupatorium genus, which have been documented to exert potential therapeutic
effects on chronic inflammatory diseases [71, 72]. Furthermore, given that KOA is
recognized as a local inflammatory disease, leveraging well-established
nanocarrier-based drug delivery strategies for flavonoids may enable targeted
drug accumulation in joint tissues, enhance therapeutic efficacy, and reduce
systemic risks [73, 74], thereby underscoring considerable clinical value that
warrants further exploration. Nonetheless, a number of limitations are worth
noting. The current analysis relied primarily on existing databases and published
literature, which may introduce biases related to data quality and completeness.
More importantly, as a network pharmacology-based analysis, it is inherently
characterized by predictivity rather than practical verification. Moreover, it
overlooks the dynamic microenvironment of organisms, such as differences in cell
types and tissue specificity, which may lead to deviations between predictions
and actual biological effects. Mechanistic evidence relies primarily on changes
in key phosphorylation nodes; subsequent validation with activator intervention
experiments is needed. Furthermore, we recognize that Eupatorin, as a natural
flavonoid, exhibits polypharmacological characteristics and may engage with
multiple signalling pathways beyond our initial hypotheses. While our study
focused on the PI3K/AKT and estrogen signalling pathways, we cannot definitively
exclude contributions from other enriched pathways to the effects of Eupatorin.
Our experiments provide foundational insights into the regulation of chondrocytes
by Eupatorin, yet KOA involves chronic, multifactorial inflammation, cartilage
degradation, and systemic immune crosstalk [75], a complexity not fully captured
by the high-dose IL-1
This represents a key limitation, as our data do not rule out off-target interactions or unexplored pathway involvement. To address these limitations, future studies will incorporate expanded phosphoproteomic profiling to determine Eupatorin’s activation of distinct pathways in KOA. Additionally, investigations will prioritize KOA-relevant cell types, including synovial fibroblasts and macrophages, alongside multicellular co-culture models, to fully elucidate its integrated biological effects within the disease context. Additionally, more clinically relevant models, such as medial meniscus destabilization (DMM) in rodents, should be adopted to better mimic KOA progression.
In conclusion, network pharmacology analysis, molecular docking and experimental validation suggest that Eupatorin may exert chondroprotective effects by regulating the activity of the PI3K/AKT and estrogen signalling pathways to modulate downstream inflammation and autophagy, but this pathway dependency requires further confirmation. These findings offer preliminary insights for subsequent studies exploring its role in KOA.
The datasets generated and analyzed during the current study are available from the corresponding author upon reasonable request.
MJZ; JLY: study design, data acquisition and statistical analysis, cell-based experiments, drafting of the original manuscript. JHL; YSG; CMH: database searching, visualization of the main results. TTM; YQZ; XHH; LLC; JWD: supplementary data collection, figure preparation, supplementary data analysis, manuscript revision. XBW; DFD: conception, research framework design, critical review of key results, final manuscript review and approval. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The study was conducted in compliance with the laboratory animal guideline for ethical review of animal welfare and approved by Ethics Committee of Shanghai University of Traditional Chinese Medicine (PZSHUTCM2311290001).
Not applicable.
This study was supported by the National Natural Science Foundation of China (grant numbers 81902306); Shanghai Municipal Health Commission Traditional Chinese Medicine Research Project (grant numbers 2024QN012).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/FBL46747.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.