1. Introduction
Vaccines have proven an effective strategy to protect members of the human
population from infectious agents. Significant advances have been made in vaccine
development, lately spurred by the Coronavirus disease 2019 (COVID-19) pandemic
that led to and was ameliorated by rapid development and deployment of mRNA
vaccines. There are several strategies for making vaccines, including mRNA,
expression of immunodominant epitopes/proteins, live attenuated viruses, and
inactivated (dead) whole viruses, each with strengths and weaknesses. For
example, mRNA vaccines can be rapidly synthesized but usually target portions of
a protein, whereas vaccine formulations composed of whole pathogens, whether
attenuated or inactivated, usually provide multiple immunologic targets. Vaccines
using inactivated viruses are made by exposing harmful viruses to certain
chemicals like formalin or ß-propiolactone. This alters the virus and stops
it from infecting cells or multiplying while preserving its ability to stimulate
an immune response within the body [1]. Alternatively, physical methods such as
Ultraviolet (UV) radiation can damage the viruses’ genetic material making them
non-infectious [1]. Although live viral vectors expressing foreign antigens show
promise as viral disease vaccines, safety remains a substantial challenge, even
with extensive attenuation of the vectors in immunocompromised patients, children
and elderly people [2, 3]. Furthermore, replicating the live attenuated viral
vector such as recombinant vesicular stomatitis (rVSV) within the host organism
could potentially result in a robust immune response against the rVSV itself,
which can overshadow the immune response of the heterologous antigen [3]. In
addition, like live viruses, live attenuated viral vector requires heavy demand
on storage and transportation in low and middle-income countries [4].
Consequently, the utilization of the vaccine in an inactivated state presents
distinct benefits [4, 5]. However, inactivated vaccines require a substantial
amount of antigen to elicit a satisfactory immune response. Furthermore, multiple
doses are needed to initiate and maintain immunity, as these vaccines cannot
self-replicate in vivo [1, 6].
The use of UV radiation to inactivate viruses has been extensively studied and
shown to be highly effective while minimizing the need for direct contact with
the inactivating agent [7]. UV light is a form of electromagnetic radiation
characterized by short wavelengths, which can cause damage to microorganisms,
including viruses [8]. Different types of UV radiation can be employed for this
purpose, including UV-A (315 to 400 nanometers wavelength), UV-B (280 to 315
nanometers wavelength), and UV-C, which has proven particularly effective
(wavelengths spanning 100 to 280 nanometers) in combating pathogens, such as
SARS-CoV-2 responsible for COVID-19, as well as influenza viruses, among others
[9, 10, 11, 12].
UV radiation offers numerous benefits compared to chemical, thermal, or
non-thermal approaches for viral inactivation. Noteworthy advantages include its
affordability, simplicity in handling and application, absence of toxicity and
irritability, and the absence of direct physical interaction with proteins or
epitopes [13]. In influenza vaccine development, influenza virus inactivation has
been proven effective [7, 14, 15, 16] with the use of different methods, including
formaldehyde, -ray or UV-ray inactivated whole influenza [7]. The
inactivation of live viral vectors such as the RABV and rNDV expressing EboGP and
SARS-CoV-2 has been proven safe and effective in generating a strong immune
response [4, 5, 17, 18, 19]. Likewise, inactivated or killed RABV has been described
to be safe and induced robust humoral immune responses against incorporated
antigens such as Hepatitis C virus and Bacillus anthrax [2, 20].
We have previously generated a recombinant vesicular stomatitis (rVSV)-based
vaccine that expressed a fusion protein consisting of the Dendritic Cell
(DC)-targeting domain of the Ebola Glycoprotein and four copies of the influenza
matrix protein ectodomain (EM-tM2e) and demonstrated this vaccine can
successfully induce robust immune responses against influenza M2e and protect
against lethal challenges from H1N1 and H3N2 strains [21, 22]. The present study
further indicates that UV-inactivated rVSV-EM-tM2e can induce
protective immune responses against influenza H1N1.
2. Materials and Methods
2.1 Ethical Statement
The BABL/c animals used in this study were maintained in a
specific-pathogen-free animal facility in the Basic Medical Science Building
(BSMB). They were all used according to the protocols approved by the Central
Animal Care Facility (CACF), University of Manitoba (Protocol Approval Number
20-017).
2.2 Construction of Plasmid
The construction of pCAGGS-EM-tM2e has been previously described
[21, 22, 23, 24, 25]. The tM2e consists of two copies of influenza M2e from the human strain
(SLLTEVETPIRNEWGCRCNDSSD), one from avian (SLLTEVETPTRNGWECKCSDSSD), and one from
swine (SLLTEVETPIRNGWECRCNDSSD). The pCAGGS-HuM2 was described previously [26].
For constructing the plasmids expressing the chimeric protein AvM2c or SwM2c that
contain the ectodomain of AvM2 or SwM2 with the HuM2 transmembrane and
cytoplasmic domains, a polymerase chain reaction (PCR)-based technique was used to amplify cDNA encoding
AvM2c or SwM2c. The 5-primers encoding for AvM2c (SLLTEVETPTRTGWECNCSGSSD)
and SwM2c (SLLTEVETPTRNEWECRCSDSSD) are 5-TCCGAATTCGTGCCGCCATGTCCCTGCTGACTGAAGTGGAAACACCAACACGGACTGGATGGGAATGCAACTGCAGCGGTAGCAGCGAC
and 5-TCCGAATTCGTGCCGCCATGTCCCTGCTGACTGAAGTGGAAACACCAACACGGAGTGAATGGGAA,
respectively. The 3-primer is 5-AAAAGATCTGCTAGCTCATTC. Then, each
PCR-amplified cDNA encoding for AvM2c or SwM2c was cloned into pCAGGS plasmid
with EcoRI and BglII sites. The sequence of AvM2c and SwM2c was
confirmed by sequencing analysis.
2.3 Antibodies and Chemicals
The influenza M2e monoclonal antibody (Santa Cruz Biotechnology, Dallas, TX,
USA; 14C2: sc-32238) was used. The peptides used are; the human M2e peptide
(SLLTEVETPIRNEWGCRCNDSSD; GenScript; RP20206), the swine influenza H1N1 M2e
peptide (SLLTEVETPTRSEWECRCSDSSD) and the avian influenza H7N9 M2e peptide
(SLLTEVETPTRTGWECNCSGSSD) and were synthesized by GenScript (Piscataway, NJ, USA)
using the methods described previously [27]. The Western Blot (WB) detection
enhanced chemiluminescence (ECL) kit was obtained from PerkinElmer Life Science
(Waltham, MA, USA) and the QUANTI-Luc™ 4 Lucia/Gaussia was purchased from
Invivogen (San Diego, CA, USA).
2.4 Cells
Human Embryonic Kidney 293T and Vero E6 purchased from ATCC were cultured in
Dulbecco’s Modified Eagle Medium (DMEM), supplemented with 10% heat-inactivated
fetal bovine serum (FBS) in a humidified incubator at 37 °C with 5%
CO. The Jurkat-Lucia NFAT-CD16 Human T Lymphocytes (NFAT-Jurkat) purchased
from Invivogen (jktl-nfat-cd16) were grown and maintained in Iscove’s Modified
Dulbecco’s Media (IMDM) containing 2 mM L-glutamine, 25 mM HEPES and 10% FBS in
a humified incubator at 37 °C with 5% CO. The 293T, Vero-E6, and NFAT-Jurkat cells were validated by STR profiling and underwent
meticulous mycoplasma testing to ensure that they were free from contamination
using Myco-Sniff-Rapid™ Mycoplasma Luciferase Detection Kit
(ThermoFisher, Waltham, MA, USA).
2.5 Production of Recombinant Vesicular Stomatitis Virus Containing
EM-tM2e and UV-Inactivation of the rVSV EM-tM2e
The production of rVSV-EM-tM2e which expresses an Ebola Glycoprotein
(EM) fused with four copies of conserved M2e consisting of 2 copies
from influenza human strain, one from influenza avian and swine strains has been
described previously [22]. The mouse-adapted strains of influenza viruses used in
the animal challenge studies were A/Puerto Rico/8/1934 (H1N1) that were generated
using reverse genetics which allows for manipulating and generating specific
viral strains [28]. The titration of the rVSV-EM-tM2e virus was
quantified in Vero E6 cells using the TCID method [29]. The
UV-inactivation of rVSV-EM-tM2e was performed using a self-contained
UV-C (CleanSlateUV, Limestone Labs, Toronto, Canada) consisting of six GLP/4P 18W
germicidal lamps, three on top and three on the bottom, that emit UV-C light at a
wavelength of 254 nm (Fig. 1A). A 6-well plate of 35 mm in diameter with 1 mL of
rVSV-EM-tM2e viral material (titer, 10 TCID/100 µL)
was placed under the upper lamp at 128 mm, with plastic lid removed, and
irradiated for 30 seconds eight times with triple shaking of the liquid at
3-minute intervals. The efficiency of virus inactivation was monitored by viral
titration and the observation of the cytopathic effect on Vero cells.
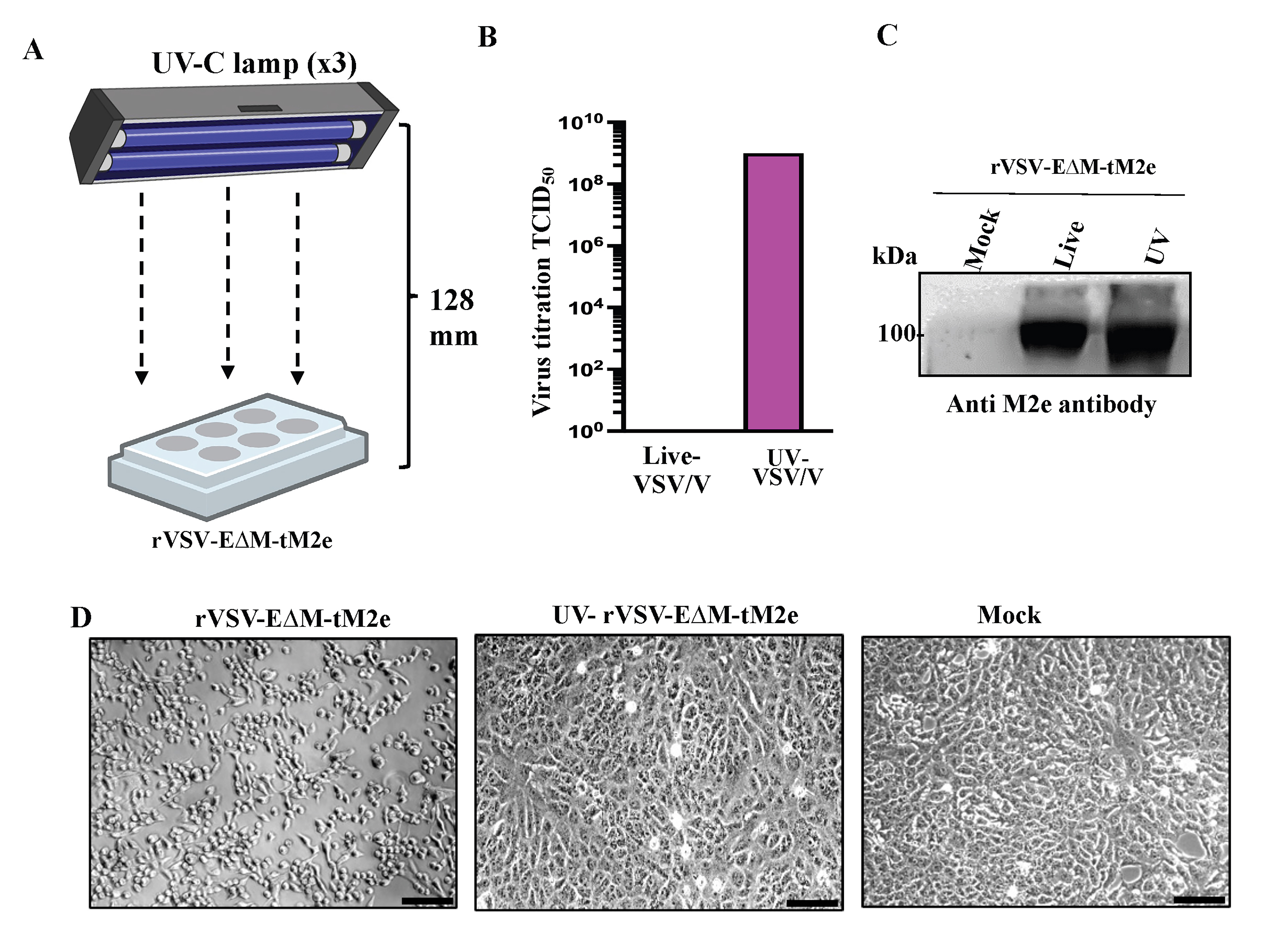 Fig. 1.
Fig. 1.
The production and characterization of the
UV-inactivated-rVSV-EM-tM2e. (A) The representative diagram of the
UV-C self-contained chamber light irradiation system of wavelength of 254 nm.
rVSV-EM-tM2e with a titer of 10 TCID/mL was placed in a 35
mm well in a 6-well plate dish, 128 mm below the UV-C light tube. (B) Viral titer
of the UV- rVSV-EM-tM2e and Live- rVSV-EM-tM2e in a Vero E6
cells. (C) The expressions of the EM-tM2e proteins in the
UV-inactivated rVSV-EM-tM2e and Live- rVSV-EM-tM2e by WB. (D)
Observation of cytopathic effect by UV-rVSV-EM-tM2e or
live-rVSV-EM-tM2e, with MOI 0.01 at 48 h after infection, Scale bars: 100 µm. UV,
ultraviolet light; rVSV, recombinant vesicular stomatitis virus; EM,
DC-targeting domain of Ebola Glycoprotein (with deleted mucin-like domain); tM2e,
four copies of influenza matrix 2 protein ectodomain (two from human strains, one
from avian and swine strains); WB, Western Blot.
2.6 Western Blot
To detect protein expression in cells, the HuM2, AvM2c or SwM2c transfected cell
lysates or lysed rVSV EM-tM2e samples were loaded onto a 10%
or 12% SDS-PAGE and analyzed by using corresponding antibodies, as described
previously [22].
2.7 Mice Immunization Experiments
A 6–8 weeks old of female BALB/c mice were immunized intranasally (IN) with live
rVSV EM-tM2e vaccine (1 10 TCID) or 1
10 TCID UV-inactivated rVSV EM-tM2e vaccine candidate and boosted with the same dosage on day 21 in a group of four.
The immunized and the PBS-treated mice were sacrificed on day 35, and blood
samples were collected for analyses on days 20 and 35. The influenza viral
challenge in mice was conducted two weeks after the final immunization by
infecting the mice intranasally with 2.1 10 PFU of H1N1, while
mice injected with PBS were challenged as a negative control. Weight loss or gain
of the mice was monitored daily for two weeks after the challenge.
2.8 Enzyme-Linked Immunosorbent Assay (ELISA)
To detect M2-specific antibodies in mouse sera, ELISA plates (NUNC Maxisorp,
Thermo Scientific) were prepared by coating them with 100 µL of human,
avian influenza H7N9, or swine influenza H1N1 M2e antigens. Subsequently, 100
µL of diluted mouse serum samples were added to each well and incubated for
two hours at 37 °C. Following washing steps, 100 µL of
peroxidase-conjugated goat anti-mouse immunoglobulin G (IgG), IgG1, or IgA
secondary antibodies were introduced and incubated for one hour at 37
°C. Finally, TMB (3,3,5,5-tetramethylbenzidine;
Mandel Scientific, Guelph, Ontario, Canada) was added, and the absorbance at 450
nm (OD450) was measured.
2.9 Antibody-Dependent Cellular Cytotoxicity (ADCC) Assay
The target cells were prepared to study influenza M2-induced ADCC by
transfecting HEK 293 T cells with HuM2-, AvM2c- or SwM2c-expressing plasmids.
After 24 hours of transfection, the cells were detached and washed once with DMEM
and then adjusted to 5 10 cells/well and added to the 96 well
plates (50 µL/well). Meanwhile, 1.5 10 cells/well
(50 µL/well) of effector cells (NFAT-Jurkat cells) were mixed with
serially diluted sera (50 µL/well) from the immunized mice and mixed
with the target cell, resulting in a final volume of 150 µL. The
cells were then incubated overnight. 30 µL supernatant from each
well was applied to 25 µL QUANTI-Luc™ 4 Lucia/Gaussia in a
96-well flexible plate for evaluation in a POLARstar OPTIMA microplate reader
(BMG Labtech, Ortenberg, Germany) [30, 31].
2.10 Flow Cytometry
The M2e expression on the surfaces of transfected 293T cells was evaluated using
anti-M2 (from the serum of rVSV EM-tM2e immunized mice) and analyzed by
fluorescence-activated cell sorting (FACS) [32]. Briefly, 293T cells were
transfected with HuM2-, AvM2c- or SwM2c-expressing plasmids in a 6-well plate.
After 24 hours of transfection, the cells were detached from the plate, incubated
with mouse serum at 4 °C for 2 hours, washed and stained with anti-mouse
IgG-FITC antibody in the dark for 30 min at 4 °C. After three washes
with PBS, cells were acquired immediately on BD LSR-Fortessa (Becton, Dickinson,
and Company (BD) Bioscience, Franklin Lakes, NJ, USA) and subsequently analyzed
on FlowJo version 10.6.1 (FlowJo, LLC, Ashland, OR, USA).
2.11 Statistics
Statistical analysis of antibodies was evaluated using the unpaired
t-test. Significance denoted by p 0.05, was determined
using GraphPad Prism 9.3.1 software (GraphPad Software, San Diego, CA, USA). With
the use of a one-way ANOVA, multiple comparison test and Tukey’s test with Prism,
the statistical analysis for the antibodies absorbance, endpoint titers, and
mouse weight loss was performed. The two-way ANOVA multiple comparison test was
used for the ADCC activity assessment. And the Log-rank (Mantel-Cox) test was
used for the survival rate.
3. Results
3.1 Characterization of the UV-Inactivated Vaccine
The rVSV-EM-tM2e vaccine candidate has been previously described and
characterized [21, 22]. Briefly, the vaccine candidate was constructed by fusing
the conserved four copies of influenza M2e from human strains (2 copies), bird (1
copy) and swine (1 copy) (tM2e) to the EM to form EM-tM2e and
expressed in the vesicular stomatitis virus (VSV) vector to generate the
rVSV-EM-tM2e as described [22]. We included M2e from different host
strains because the M2e of avian or swine-origin influenza viruses have some
amino acid substitutions. The rVSV-EM-tM2e was inactivated in this
study using UV-C light at the wavelength of 253.7 nm. Briefly, the
rVSV-EM-tM2e with a titer of 10 TCID/mL was placed in a 35
mm well in a 6-well plate dish, 12.8 cm below the UV-C light tube, and irradiated
for 30 seconds, 8 times at 3-minute intervals (Fig. 1A).
To ensure the complete deactivation of the virus, both live and UV-inactivated
rVSV-EM-tM2e (UV-rVSV-EM-tM2e) were subjected to titration on
Vero E6 cells. It was observed that the live rVSV-EM-tM2e virus
efficiently infected Vero E6 cells while the UV- rVSV-EM-tM2e had no
measurable titre detected after 48 hours of infection (Fig. 1B). The expression
of influenza M2e was detected in the UV-rVSV-EM-tM2e at an equivalent
level to the rVSV-EM-tM2e (Fig. 1C). The cytopathic effect was observed
on Vero E6 cells infected with the live rVSV-EM-tM2e virus, but not the
UV- rVSV-EM-tM2e (Fig. 1D). The collective data indicates that the
UV-inactivation of the rVSV-EM-tM2e was effectively achieved and the
inactivated rVSV-EM-tM2e lost its infection ability.
3.2 The UV- rVSV-EM-tM2e Induces Robust Influenza Humoral
Immune Responses
Given the established evidence that the UV-rVSV-EM-tM2e retains the
presence of influenza M2e fused with EM (EM-tM2e) while
lacking replication capabilities, we proceeded to examine the capacity of the
UV-rVSV-EM-tM2e in eliciting specific immune responses against
influenza. Briefly, via IN immunization, the BALB/c mice received 1.0
10 TCID of Live-rVSV-EM-tM2e or UV-rVSV-EM-tM2e
(1.0 10 TCID) on day 0 and day 21. On days 20 and 35,
mice sera were collected and measured for anti-M2e immune responses (Fig. 2A).
Results showed that both the Live or UV-rVSV-EM-tM2e immunization in
mice yielded a notable robust anti-M2e IgG against human, swine and avian
influenza M2e (Fig. 2B,C). Subsequent booster administration significantly
heightened the immune response titre. Notably, no statistically significant
difference was observed between the induced humoral immune responses by Live and
UV-rVSV-EM-tM2e. However, it is crucial to highlight that the
UV-rVSV-EM-tM2e dosage is 10 higher than the
Live-rVSV-EM-tM2e. We further examine their ability to induce M2e IgG,
IgG1 and IgA immune responses against human, swine, and avian influenza strains
(Fig. 2D–F). While the rVSV-EM-M2e vaccine demonstrated the ability to
generate substantial immune responses targeting various influenza strains, our
findings indicate notable differences in the immune responses elicited against
different strains (Fig. 2B–F). Notably, the immune responses triggered against
human strains were the most robust, whereas the responses against avian strains
were comparatively higher but not significantly higher than those against swine
strains (Fig. 2D,E). Interestingly, the IgA antibodies produced by either
Live-rVSV-EM-tM2e or UV-rVSV-EM-tM2e were not significantly
different (Fig. 2F). Overall, our data revealed that the inactivation of the
viral vector vaccine candidate rVSV-EM-tM2e with UV could still induce
robust, broad immune responses against influenza M2e from different strains.
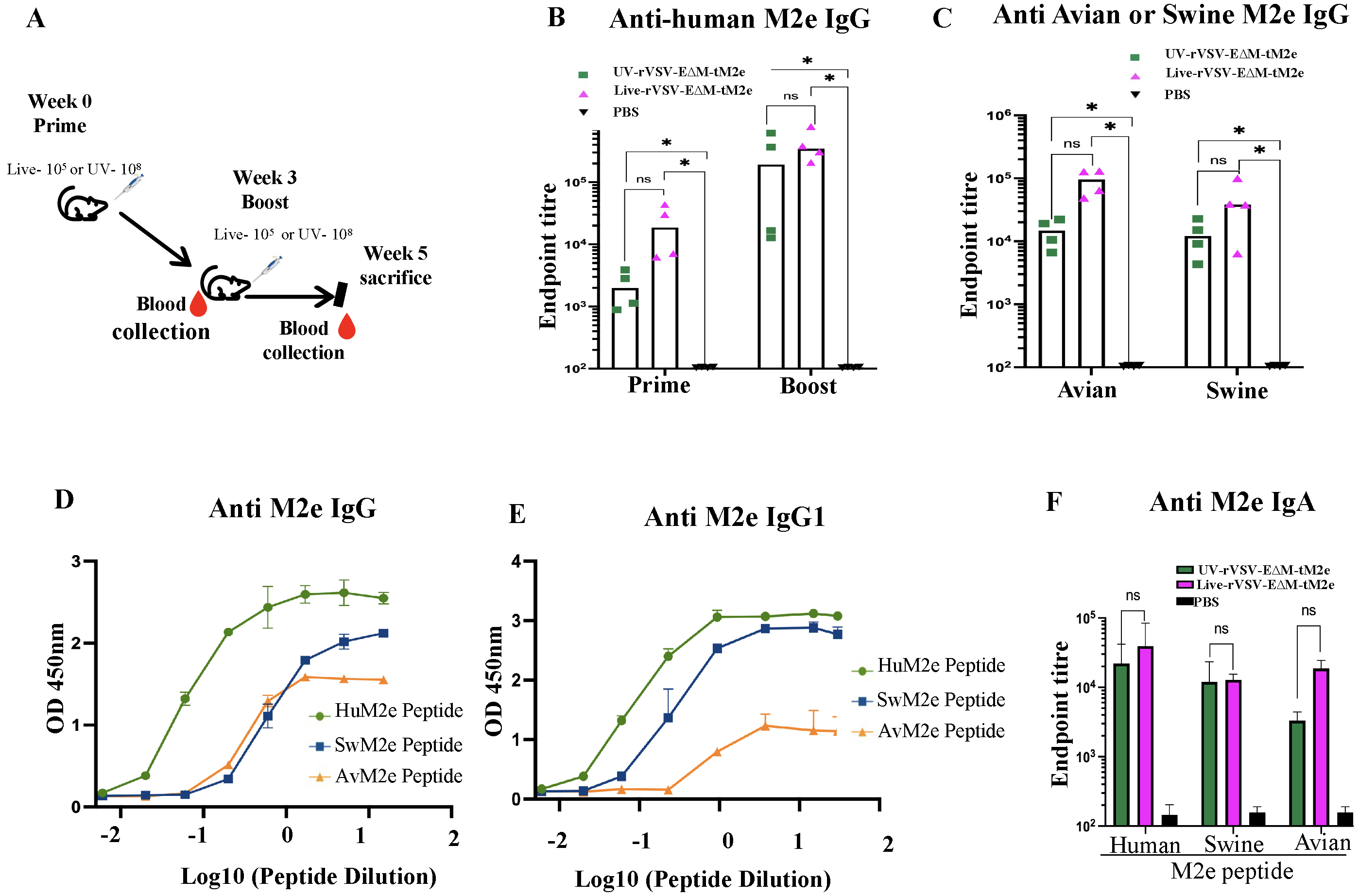 Fig. 2.
Fig. 2.
The specific anti-M2e elicited by UV-rVSV-EM-tM2e or
Live-rVSV-EM-HM2e in sera from immunized mice. (A) Following IN
immunization of the BALB/c mice with 1.0 10 TCID of
UV-rVSV-EM-tM2e or 1.0 10 TCID of
Live-rVSV-EM-tM2e in a prime-boost regimen (Day 0 and Day 21), the mice
were bled on days 20 and 35 (n = 4/5 mice per group). The enzyme-linked immunosorbent assay (ELISA) assay detected (B)
Anti-M2e IgG against M2e from human influenza strains after prime and boost
immunizations respectively. (C) Anti-M2e IgG levels against M2e from swine and
avian influenza strains after boost immunization. (D) Anti-M2e IgG levels present
in the serum of the Live-rVSV-EM-tM2e immunized mice against human
(HuM2e), swine (SwM2e) or avian (AvM2e) influenza M2e strains. (E) Anti-M2e IgG1
levels in the serum of the mice immunized with Live-rVSV-EM-tM2e
against human (HuM2e), swine (SwM2e) or avian (AvM2e) influenza M2e strains. (F)
Anti-M2e IgA levels against M2e from human, swine and avian influenza strains.
Using an unpaired t-test, and significant p-values are
represented with asterisks, *p 0.01, the statistical significance between
the groups was determined. ns, no significance.
In addition, we tested whether the influenza M2e antibodies in the immunized
mice sera could recognize and react with the influenza M2e present on the cell
surface. To achieve this, we constructed plasmids expressing human M2 (HuM2) and
chimeric proteins (AvM2c and SwM2c) in which the HuM2e was replaced by the AvM2e
or SwM2e (Fig. 3A), and transfected each of them into 293T cells. The expression
of the HuM2, AvM2c, and SwM2c was detected by WB using
Live-rVSV-EM-tM2e immunized mice sera (Fig. 3B). Similarly,
fluorescence-activated cell sorting (FACS) analysis was conducted on HuM2, AvM2c,
or SwM2c transfected 293T cells that were probed with the mice sera. Meanwhile,
EM-tM2e or influenza HA transfected cells, were used as control.
Interestingly, our results showed that the sera effectively recognized and bound
with huM2, SwM2c, AvM2c and EM-tM2e, but not HA (Fig. 3C).
These findings strongly indicate that the vaccine candidates possess the
potential for cross-protection against influenza strains affecting humans, swine,
and birds.
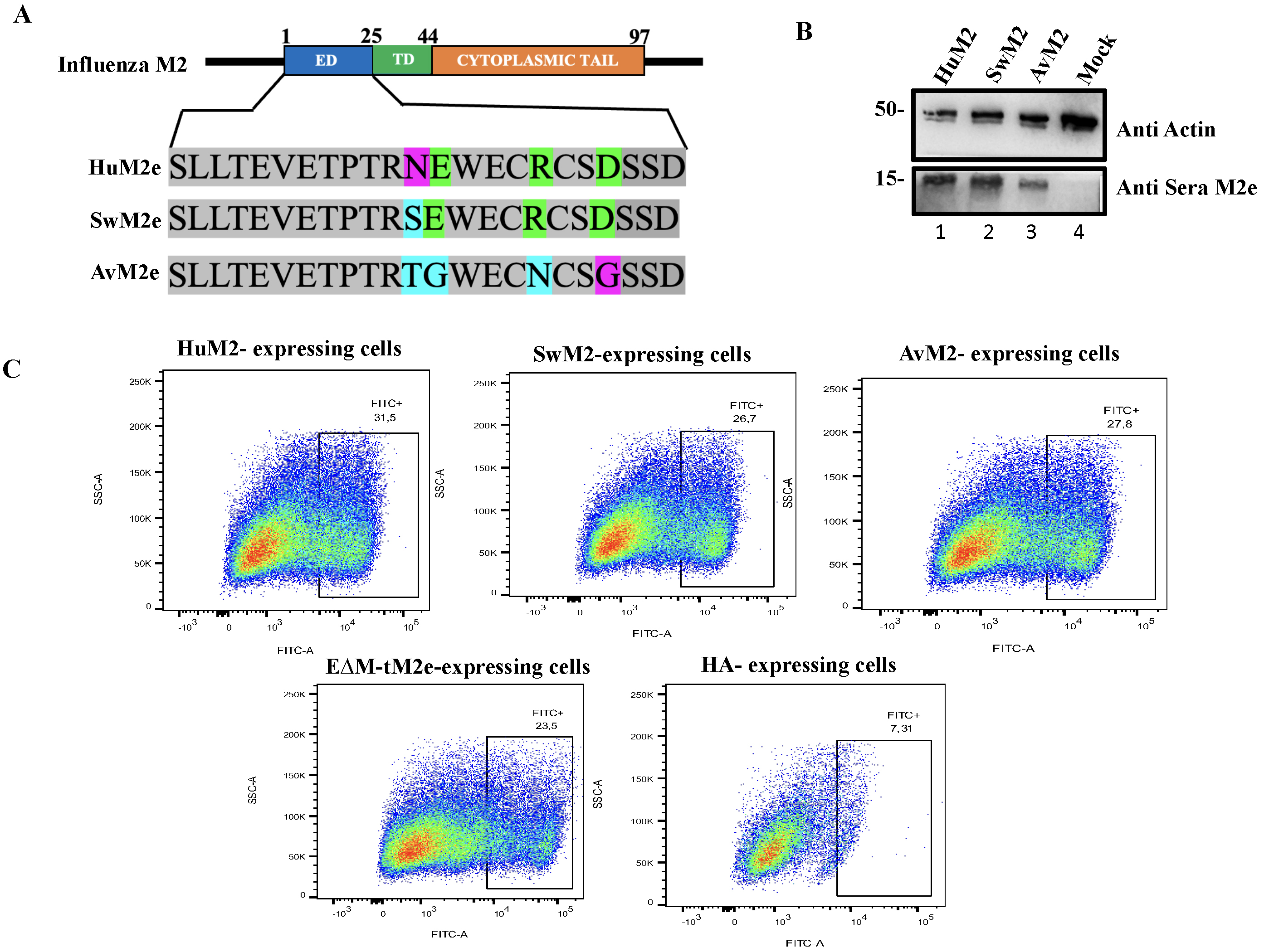 Fig. 3.
Fig. 3.
Characterization of anti-M2e induced immune responses by
Live-rVSV-EM-tM2e. (A) Schematic representation of the influenza HuM2
expressing M2e from human (HuM2e), avian (AvM2e) or swine (SwM2e) and are named
as HuM2, AvM2c or SwM2c. The ectodomain of the pCAGGS M2H5N1 was deleted and
replaced with the ectodomain of avian (SLLTEVETPTRTGWECNCSGSSD) or swine
(SLLTEVETPTRSEWECRCSDSSD). The M2e expression on the surfaces of transfected 293T
cells was evaluated by anti-M2 staining (from the serum of immunized mice) and
analyzed by WB (B) and fluorescence-activated cell sorter (FACS) and (C) analyzed
on FlowJo version 10.6.1.
3.3 The UV-rVSV-EM-tM2e Protected against H1N1 Influenza A
Challenge
Considering that the UV-rVSV-EM-tM2e exhibited the capacity to elicit
an immune response like that of the Live-rVSV-EM-tM2e, we proceeded to
subject the mice that had been immunized with the UV-rVSV-EM-tM2e to
influenza H1N1 challenge. Groups of 4 mice were immunized with 10
TCID of rVSV-EM-tM2e, 10 TCID of
UV-rVSV-EM-tM2e or PBS through the IN route on Day 0 and boosted with
the same dosage on Day 21. After two weeks following the booster dose
administration, the mice that had received immunization were challenged with the
influenza virus, H1N1, with a viral titer of 2.1 10 TCID,
employing the same methodology described earlier [22] (Fig. 4A). The mice were
observed for 14 days for weight loss and survival. Both the Live
rVSV-EM-tM2e and the UV-rVSV-EM-tM2e displayed a gradual
decrease in weight of approximately 10% until Day 4, followed by a progressive
weight gain without any instances of mortality. In contrast, mice in the PBS
group deteriorated and lost weight sporadically within the four days of the viral
challenge without survival (Fig. 4B,C). Interestingly, the anti-M2e immune
responses increased after the challenge in both live and UV-
rVSV-EM-tM2e immunized mice, but not in the group of mice that received
PBS (Fig. 4D). These results revealed the potency of the immune response induced
by the UV-rVSV-EM-tM2e to protect against influenza challenge.
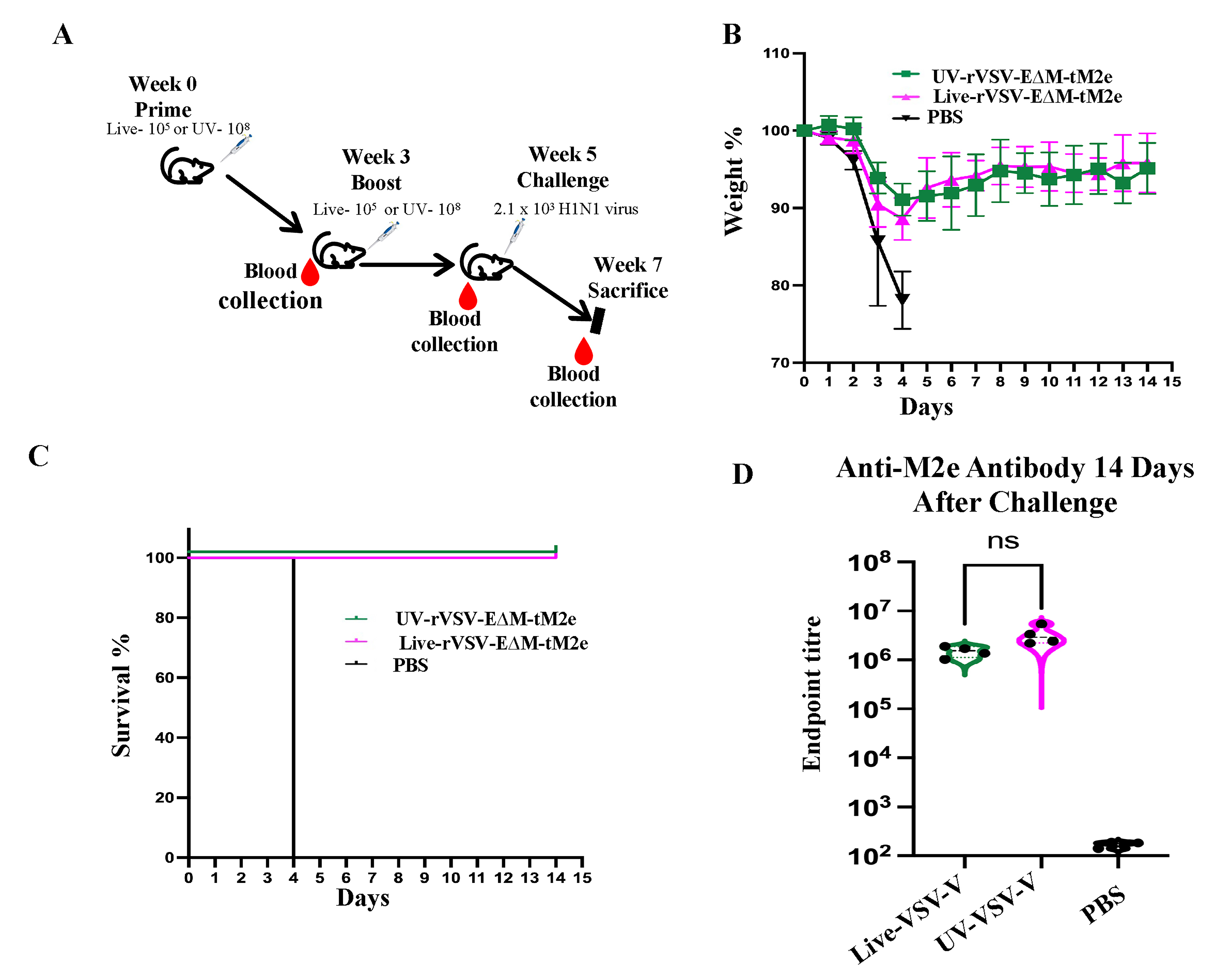 Fig. 4.
Fig. 4.
The UV-rVSV-EM-tM2e vaccination protected against the
influenza H1N1. (A) The female BALB/c mice immunized IN with 1.0
10 TCID of UV-rVSV-EM-tM2e or 1.0 10
TCID Live-rVSV-EM-tM2e on days 0 and 21 were challenged with 2.1
10 PFU of H1N1 on Day 14 post-immunization. (B) The percentage
increase or decrease in weight loss or gain of the mice. (C) The survival rates
of mice after infection with H1N1. (D) Levels of anti-M2e IgG in the serum after
the challenge detected by ELISA. Using an unpaired t-test, the
statistical significance between the groups was determined. ns, no significance.
3.4 The rVSV-EM-tM2e Vaccination Induces ADCC against
Influenza M2e from Different Host Strains
Studies have suggested that the anti-influenza M2e that confers protection
against influenza challenge could be through the ADCC [33, 34, 35, 36]. We, therefore,
expanded our investigation to determine whether the Live rVSV-EM-tM2e
or UV-rVSV-EM-tM2e vaccine can also elicit ADCC activity against
influenza M2e proteins from avian and swine influenza strains. This is
particularly significant because the vaccine formulation includes M2e regions
from human, swine, and avian sources, making it crucial to evaluate its potential
effectiveness against these diverse strains. ADCC is a key immune mechanism where
effector cells with Fc receptors eliminate target cells coated with antibodies
expressing pathogen-derived antigens [37]. Associations between ADCC activity and
Fc receptor polymorphisms are observed in vaccination [37, 38]. The NFAT
signalling pathway is crucial for Fc receptor (FcR)
activation, playing a critical role in ADCC [39]. In this study, we employed
NFAT-reporter systems, utilizing human FcRIIa (CD16a) expressed in
Jurkat cells as effector cells, and 293T cells transfected with influenza human
M2 as target cells. We used Influenza huM2, SwM2c, or AvM2c transfected 293 T
cells as targets, mixed with effector cells (NFAT-Jurkat cells) and serum from
immunized mice. After overnight incubation, we evaluated the supernatant using
QUANTI-LucTM 4 Lucia/Gaussia and a microplate reader (BMG Labtech, Germany) [30, 31]. The Live rVSV-EM-tM2e vaccine could induce immune responses that
elicited ADCC activity against the HuM2e, SwM2c and AvM2c. Interestingly, the
UV-rVSV-EM-tM2e vaccine also demonstrated ADCC activity against the
HuM2, SwM2c and AvM2c, even though the level of ADCC activity was notably lower
compared to that induced by the Live rVSV-EM-tM2e vaccine (Fig. 5A–C).
However, even with this lower ADCC activity, the antibody induced by
UV-rVSV-EM-tM2e was sufficient to protect mice against the highly
virulent H1N1 influenza strain. Moreover, to further prove the specificity of the
anti-M2e-mediated ADCC activity, the M2e-antibody-induced ADCC assay was
performed in the presence of different concentrations of HuM2e peptides, and
results showed that the anti-HuM2e ADCC were blocked using HuM2e peptides at
different levels with various concentrations. Our results revealed that at the
concentration of HuM2e peptide (10 µg/mL), the ADCC activity against
HuM2 was significantly reduced (Fig. 5D). These data support the specificity and
immunogenicity of the ADCC activity induced by the rVSV-EM-tM2e.
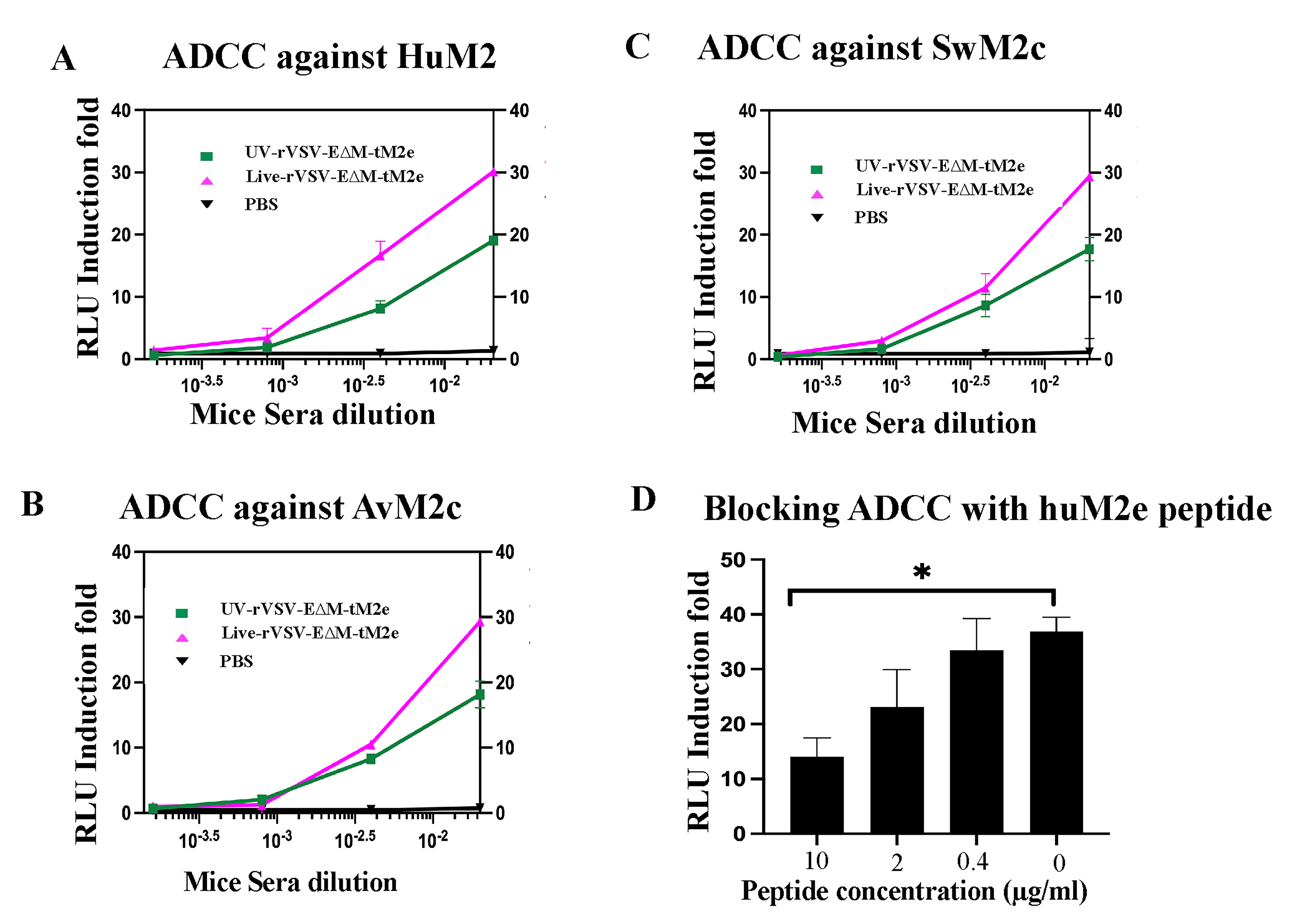 Fig. 5.
Fig. 5.
The Live or UV-rVSV-EM-tM2e immunized serum-mediated
anti-M2e ADCC activity against influenza M2e from different host strains. ADCC
activity induced by UV-rVSV-EM-tM2e or Live rVSV-EM-tM2e in
the serum against (A) HuM2 (B) SwM2c or (C) AvM2c. (D) The Antibody-Dependent
Cellular Cytotoxicity (ADCC) activity levels of the anti-M2e antibodies in the
presence of HuM2e peptide. Using an unpaired t-test, and significant
p-values are represented with asterisks, *p 0.01, the statistical
significance between the groups was determined.
4. Discussion
The primary study objective was to investigate concerns associated with the
utilization of replication-competent rVSV vaccine candidates. We explored the
effectiveness of a UV light-based inactivation technology to enhance the safety
of vaccine candidates [40]. The inactivation of rVSV using UV light was deemed
advantageous compared to attenuating the virus, as the growth of attenuated virus
strains often presents challenges during manufacturing [40]. In this study, we
successfully demonstrated the efficacy of a UV-inactivated rVSV-carrying
influenza antigen -M2e to elicit robust immune responses against influenza M2e
from human, swine, and avian hosts. These immune responses proved highly
effective in protecting against a lethal H1N1 challenge. Moreover, our research
results emphasize that the immune responses triggered by M2e can induce ADCC
activity against the HuM2, SwM2c, and AvM2c proteins present on the cell surface.
Notably, the Live rVSV-EM2e merely necessitated 10 TCID,
whereas the UV-rVSV-EM2e necessitated 10 TCID, suggesting
that accomplishing these outcomes entailed administering a relatively higher
amount of UV-rVSV-EM-tM2e.
A notable drawback associated with the utilization of inactivated viruses is the
requirement for elevated dosages and repeated administrations, owing to the
inability of the virus to replicate within the host [1, 6]. It is therefore worth
noting that our UV-inactivated vaccine candidate only required two
administrations, like the Live rVSV-EM-tM2e, to induce protective
immune responses. Similarly, Jones et al. [40] mentioned in their study
that a psoralen/UV light-inactivated vaccinia virus vector (VV) delivering HIV-1
multi-envelope produces the same outcome as Live VV in immune responses and
protection against heterologous SHIV 89.6P in the macaque. The immune responses
triggered by the rVSV-EM-tM2e vaccine against the M2e protein could
recognize and bind with the ectodomain of the M2 proteins from different
influenza viruses, including those found in humans, swine and avian. This is
expected since the vaccine contains two copies of the M2e protein from human
strains, one from the avian and the swine influenza strains. However, the immune
response showed a higher specificity towards M2e proteins from human strains,
likely due to the higher number of human M2e copies in the vaccine construct. The
broad immune responses exhibited by our vaccine against human, swine, and avian
strains indicate its significance for universal influenza vaccine development.
Studies have demonstrated the pivotal role of influenza M2e in developing a
universal influenza A vaccine because of its high conservation [34]. Despite
being less immunogenic, influenza M2e is highly conserved among influenza
strains, making it an ideal target for universal influenza vaccines [41]. Several
attempts have been made to enhance its immunogenicity [27, 42]. Our concept has
shown that the fusion of four copies of influenza with the DC-targeting domain of
Ebola GP (EM) can improve its immunogenicity [21, 43, 44, 45].
Likewise, our study demonstrated that intranasal immunization of either live or
UV-rVSV vaccine candidates is safe when administered in mice. Notable advantages
of rVSV in mucosal immunization include stimulating all immune system components,
including IgA, and IgG [46]. Intranasal administration is a painless and
non-invasive method that does not require sterilization. It can be
self-administered and induces protection against respiratory tract infections.
This method generates lymphoid tissue associated with the nasal cavity, providing
long-term immune responses [46]. Our previous studies have also shown that the
intramuscular, intranasal, or oral immunization of rVSV-M2e-based vaccines
triggered robust immune responses against influenza M2e [22, 47]. This suggests
that this vaccine can be administered intranasally.
The spread of avian influenza A (HPAI) H5N1, highly pathogenic in wild birds,
has caused occasional severe human infections and sometimes human-human
transmission, increasing the likelihood of the next influenza pandemic [48]. Our
vaccine candidate is, therefore, promising to be used against HPAI H5N1 thereby
reducing the chances of a pandemic and can even be used in pandemic preparedness.
Likewise, the 2009 pandemic H1N1 swine flu which is now circulating as seasonal
flu is of great health concern [49]. Many other strains of swine flu have also
emerged [49]. This also calls for the need to develop a vaccine that is not only
targeting human strains but also other strains of influenza. In line with the One
Health campaign, our vaccine design is potentially suitable for human and animal
health use.
It is noteworthy that the immune response against the M2e protein also activated
the activity of ADCC against HuM2, SwM2c, and AvM2c. This indicates that the
vaccine’s means of protection against influenza involves the ADCC mechanism.
Although there is some disagreement about the functional activity of the
M2e-induced antibody in the protective role, our previous and current data
suggest that ADCC may play a significant role [47]. The Fc receptors (FcR) have
been shown to play a critical role in cross-protective immunity through M2e [50, 51]. This suggested that the viral load can be cleared through various mechanisms
such as ADCC, antibody-dependent cellular phagocytosis, and complement-dependent
cytotoxicity (CDC). A study showed that H7N9+4M2e had higher CDC than H7N9 alone
in immunized mice serum [52]. Whereas another study suggests ADCC as a potential
protective mechanism [50]. The immune serum of the M2e-H3 stalk showed increased
binding of IgG antibodies to viral antigens of groups 1 and 2 on the cell
surface. This was associated with an increase in the activity of Jurkat cells,
acting as surrogate effector cells for ADCC, targeting Group 1 or Group 2
infected Madin-Darby Canine Kidney (MDCK) cells [50].
5. Conclusions
Conclusively, the UV-based inactivation of the rVSV vector vaccine against the
influenza virus has shown remarkable success in generating immune responses and
protecting against the H1N1 viral challenge. These findings suggest that
UV-inactivated rVSV-EM-tM2e could be used as an inactivated vaccine
against influenza viruses. It also demonstrated the proof-of-concept of its usage
as a vaccine against human, avian and swine influenza strains in animal health.
However, it is important to assess the effectiveness of the UV-inactivated
vaccine against other influenza virus strains, such as H3N2 and H5N1.
Availability of Data and Materials
The data sets generated and/or analyzed during the current study are available
from the corresponding author.
Author Contributions
XY, TAO, ZA, KC, LM, GK and DK designed the research study. TAO, ZA, HA, TO, and
MJO, performed the research. TAO, ZA and TO analyzed the data. TAO, ZA, XY and KC
wrote the manuscript. All authors contributed to the editorial changes, read, and
approved the final manuscript and participated substantially in this work. All
authors agreed to be accountable for all aspects of the work.
Ethics Approval and Consent to Participate
The study was conducted following the Central Animal Care Facility (CACF),
University of Manitoba (Protocol Approval Number 20-017) or the Animal Care Committee at the Canadian Center for Human and
Animal Health for studies involving animals.
Acknowledgment
We thank Ruth Mwatelah for proofreading and providing linguistic editing of the
manuscript.
Funding
This research was supported by the funding from Canadian Institutes of Health Research (OV5-170710),
Research Manitoba (https://researchmanitoba.ca/). This research was
supported by the Manitoba Research Chair Award from Research Manitoba (RM) to
X.Y. T.A.O., who is also a recipient of the Allan Rolland Studentship by the
Department of Medical Microbiology and Infectious Diseases, University of
Manitoba.
Conflict of Interest
The authors declare no conflict of interest.
 Fig. 1.
Fig. 1. Fig. 2.
Fig. 2. Fig. 3.
Fig. 3. Fig. 4.
Fig. 4. Fig. 5.
Fig. 5.




