Frontiers in Bioscience-Landmark (FBL) is published by IMR Press from Volume 26 Issue 5 (2021). Previous articles were published by another publisher on a subscription basis, and they are hosted by IMR Press on imrpress.com as a courtesy and upon agreement with Frontiers in Bioscience.
1 Department of Molecular Reproduction, Development and Genetics, Indian Institute of Science, Bangalore 560012, India
Abstract
Proteoglycans are essential constituents of tissue- and organ microenvironments, modulating both the structural scaffolds that surround cells as well as signaling cues that determine cellular phenotype. An important modification of proteoglycans is the sulfation of the monosaccharides that comprise them. Sulfates are added by sulfotransferases and desulfation occurs through the action of sulfatases. In this essay, we examine the biochemistry of a conserved family of desulfating enzymes known as arylsulfatases. A subset of these enzymes mediates the desulfation of proteoglycans. We review the consequences of their aberrant expression in the light of carcinogenesis and carcinomatosis: the dissemination of cancer cells. A closer understanding of their cellular-molecular roles reveals their promise for future strategies for cancer therapy.
Keywords
- Proteoglycans
- Arylsulfatase
- Cancer
- Epithelial To Mesenchymal Transition
- Review
The progression of cancer within solid tissues involves recurrent interactions between the proliferating and migrating cells, and the extracellular matrix (ECM) that surrounds them (1). The ECM consists of a spatiotemporally heterogeneous mixture of secreted glycoproteins and proteoglycans that acts both as a scaffold for, and a source of signaling cues to, transformed cells (2). Proteoglycans (PG) comprise a core protein with one or more variable linear chains of repeating oligosaccharide units known as glycosaminoglycans (or GAGs; each oligosaccharide unit consisting usually of an aminoglycan and an acidic glycan or a sugar; (Table 1) conjugated to the hydroxyl group of a serine through a tetrasaccharide bridge (3). The protein core determines localization of the proteoglycan: a transmembrane domain leads to its insertion in the membrane, otherwise proteoglycans often reside in the extracellular space (there are reports of localization of proteoglycans in the nucleus as well (4)).
| GAG | Aminoglycan | Acidic glycan/sugar | Sulfation | Degrees of sulfation/ disaccharide unit | Reference |
|---|---|---|---|---|---|
| Heparin | GlcN | IdoA | 2-O- of IdoA, 6-O- and 3-O- of GlcN | 2.7 | 5, 95 |
| Heparan Sulfates | GlcN | GlcA | 2-O- of GlcA, N-, 6-O and 3-O- of GlcN | 1 | 96 |
| Chondroitin | GalN | GlcA | 4-O- and/or 6-O- of GalNAc, 2-O- of GlcA | 0.1–1.3 | 6 |
| Dermatan | GalN | IdoA | 2-O-of IdoA | 1-3 | 6 |
| Keratan | GlcN | Gal | 6-O- of GlcN, 6-O- of Gal | - | 95 |
| Hyaluronic acid | GlcN | GlcA | None | - | 5 |
Based on the backbone of repeating disaccharide units comprising GAGs, they can be categorized into four main groups: heparin/heparan sulfate (HS), chondroitin sulfate (CS)/dermatan sulfate (DS), keratan sulfate (KS), and hyaluronan (HA, also known as hyaluronic acid) (Table 1). HS GAGs and CS GAGs are synthesized in the Golgi, where the individual GAG chains are O-linked to a core protein of PG (5). Dermatan sulfates (DSs) are stereoisomers of CSs (6). KS, on the other hand, can be either N-linked or O-linked to the core protein and are synthesized in the Golgi body. HA is unique GAG in that it is not synthesized by Golgi but by integral membrane protein synthases and also is not covalently attached to proteins to form a PG. It may however form complexes with proteins proteoglycans present on the cell surface and in the ECM (5).
The GAG attachment sites on the core protein of the PGs in most cases have a consensus Ser-Gly/Ala-X-Gly motif, where chain initiation begins from the Oγ atom of the serine (5). In most cases, GAGs (except for keratin sulfate) are linked to the protein core via a specific tetrasaccharide linker made up of a GlcA residue, two Gal residues, and a xylose (Xyl) residue making a structure such as: GAG(n)–GlcA–Gal–Gal–Xyl–Ser–protein (7). The process of linking takes place in endoplasmic reticulum and Golgi compartments (8). The tetrasaccharide linker is attached to the Ser residue in protein core through an O-glycosidic bond and is commonly found in PGs that contain GAGs like heparins, heparan sulfates, dermatan sulfates, and chondroitin sulfates. However, a trisaccharide linkage lacking GlcA residue has also been reported. Keratan sulfate is attached to Asn in core proteins via a complex-type N-linked branched oligosaccharide, or O-linked via GalN to Ser or Thr residues (9, 10).
The sulfation of linker saccharide has also been reported but only in case of DS/CS and not in heparin/HS. Both CS and DS may have C4 sulfation of the second Gal in the linker region. It has been proposed that the modification by sulfate might influence the degree of synthesis of CS/DS versus HS/heparin (7). According to another report, sulfation of the linker region is restricted to the CS/DS pathway, wherein 4-O-sulfation of the second galactose and 6-O-sulfation of both the first and the second galactose units is found. Here again of the galactose units of the linker region has been suggested to promote CS/DS synthesis while the 3-O sulfation of the fourth sugar unit of the linker (GlcA) produces opposite effect, by preventing CS GAG synthesis acting as potential GAG chain termination signal. Same cell type can produce different GAG chains depending on the extent of sulfation of linker region (11).
Apart from linker tetrasaccharide, GAGs also show variable length, composition and degree of sulfation (12, 13). Sulfation alters the chemical properties of the milieu in which the proteoglycan resides: the negative charge along with the hydrophilicity of the glycan chains mediates binding of diffusible proteins such as growth factors (14, 15). Differential sulfation of PGs for example regulates the balance between self-propagation and differentiation within stem cell niches (16) (6) and can determine the switch between homoeostatic and transformative phenotypes of tissues (17). Sulfation, the final step of proteoglycan synthesis is mediated by sulfotransferases (18-20). Therefore, cell- and tissue-specific regulation of sulfotransferases results in differential sulfation of proteoglycans within, and between, ECMs of different organs. The role of sulfotransferases has been extensively studied in the context of development, homeostasis and diseases, especially cancer (21, 22).
A second set of enzymes are involved in desulfating the proteoglycans as part of their degradation. These enzymes, known as sulfatases belong to a larger family of enzymes known as aryl-sulfate sulfohydrases or arylsulfatases that catalyze the hydrolysis of sulfate ester bonds from a wide range of substrates including, but not limited to, GAGs (23-26). Our knowledge of tissue-specific functions of arylsulfatases, or of the consequence of their perturbation on organ and organismal phenotype, trails behind the extensive research built around sulfotransferases. However, in the last decade or so, a formidable body of investigations on arylsulfatases has begun to shed light on their involvement in cancer.
Here, we review the emerging literature on arylsulfatases in the context of insights into the role of the microenvironment in the invasion and dissemination of cancer, with an emphasis on the subset of this family that is involved in proteoglycan metabolism. The first section introduces the molecular and cellular characteristics of arylsulfatases. The second section surveys the contribution of arylsulfatases to the modification of the tumor microenvironment during cancer progression. In the third and concluding section, we discuss open questions relating to the tumor microenvironment and how investigations on oncogenic roles of arylsulfatases may advance translational research in cancer therapy.
Arylsulfatases (ARS) are a class of phylogenetically-related enzymes that hydrolyze sulfate esters of a variety of substrates such as sulfated GAGs, sulfo-lipids and -proteins, and steroids. The class derives its name from the fact that its enzymes are able to hydrolyze small synthetic sulfated aromatic substrates such as p-nitrocatechol sulfate, p-nitrophenyl sulfate, and 4-methylumbelliferyl sulfate in vitro (27-30). The human genome is known to encode seventeen arylsulfatases (31). Since the substrates on which they exert their action are diverse in chemical nature, arylsulfatases contribute to cellular structure and function in manifold ways. Similarly, the dysregulation of their expression and/or activities is associated with disorders of development (such as lysosomal storage diseases) and pathologies such as cancer.
Based on their canonical subcellular localization, arylsulfatases have broadly been divided into lysosomal and non-lysosomal subgroups. The former subgroup comprises six sulfatases: arylsulfatases A (ARSA), and -B (ARSB), iduronate-2-sulfatase (IDS), N-sulfoglucosamine sulfohydrolase (SGSH), glucosamine-6-sulfatase (GNS), and galactosamine-6-sulfatase (GALNS). These enzymes typically localize to the lysosomes and exhibit maximal activity in an acidic milieu (26).
More recently, two new enzymes, arylsulfatases G (ARSG) and K (ARSK) have also been shown to localize to lysosomes (29, 32). However, this canonical localization may change depending on tissue-specific, or pathological contexts. For example, both ARSA and ARSB are also found on the cell surface of sinusoidal endothelia (where they are localized with heparan sulfate proteoglycans) and macrophages (Kupffer cells) of liver (30). ARSB may also be localized to the nuclear and cell membranes of colonic epithelial cells, cerebrovascular cells, hepatocytes and in adenocarcinomas of colon (33). The levels of ARSB has also been reported to be several-fold higher in the membrane fractions of cultured bronchial epithelial cells (34).
The non-lysosomal arylsulfatases show diverse localizations: arylsulfatases C, D and F, which are localized in the endoplasmic reticulum (ER), arylsulfatase E, which resides in the Golgi organelles and Sulfatase 1 and Sulfatase 2, which reside on the cell surface. These enzymes exhibit optimal activity at neutral pH. In addition to ER, arylsulfatase C (ARSC), may localize to almost all membrane compartments—the rough and smooth endoplasmic reticulum, the Golgi cisternae, the endosomes, the dense lysosomes, and the plasma membrane (35-37).
Of the four new arylsulfatases termed as arylsulfatase H, I, J, and K, identified in 2005 by a genome-wide screen for a conserved sulfatase-specific signature sequence (C/S-X-P-X-R) (31), Arylsulfatase I (ARSI) displays arylsulfatase activity at neutral pH (38) and is thought to be secreted (39). Less is known about the localization of arylsulfatases H (ARSH) and J (ARSJ) although prediction programs suggest an extracellular and ER-based localization for the latter and an ER-based localization for the former.
In Table 2, we have, for better visual representation, tabulated the known substrates, tissue of expression, and genetic diseases associated with each arylsulfatase (Table 2)
| Gene | Enzyme |
Associated genetic disease | Physiological substrate | Organ with Highest expression (Transcripts) | Reference |
|---|---|---|---|---|---|
| ARSA | Arylsulfatase A |
Metachromatic dysplasia (accumulation of sulfatides to toxic levels in the nervous system). | Sphingolipid sulfate ester (cerebroside 3-sulfate, sulfatides) | Testes, spleen | 97 |
| ARSB | Arylsulfatase B |
Mucopolysaccharidosis type VI syndrome (MPS VI) (accumulation of sulfated GAGs. like dermatan sulfate and chondroitin sulfate in vital organs). | N-Acetyl-D-galactosamine, chondroitin-4-sulfate (C4S) and dermatan sulfate (DS) | Kidney, placenta | 98 |
| ARSC/STS | Steroid sulfatase (Arylsulfatase C, estrone sulfatas, steryl-sulfate sulfohydrolase) | X-linked ichthyosis characterized by excessive skin scaling or hyperkeratosis, due to lack of breakdown of cholesterol sulfate. | 3-beta-hydroxysteroid sulfates | Placenta, fat | 99 |
| IDS | Iduronate-2-sulfatase |
Hunter syndrome, also known as Mucopolysaccharidosis type II, an X-linked lysosomal storage disease characterized by coarse facial features, stiff joints, skeletal abnormalities, hepatosplenomegaly, cardiovascular and respiratory disorders, developmental delay and deteriorating intellectual function. | Heparan sulfate and dermatan sulfate | Brain | 100 |
| SGSH | N-sulfoglucosamine sulfohydrolase |
Sanfilippo syndrome (mucopolysaccharidosis (MPS1) type III) causing progressive neurological deterioration. | Heparan sulfate | Spleen, adrenal, prostate | 101 |
| GNS | N-acetylglucosamine-6-sulfatase |
Sanfilippo syndrome type D (Mucopolysaccharidosis type IIID), the least common form of the four subtypes of Sanfilippo syndrome. | Heparin, heparan sulfate, and keratan sulfate | Adrenal, kidney | 102, 103 |
| GALNS | N-acetylgalactosamine-6-sulfatase |
Mucopolysaccharidosis IV A (also known as MPS IV A and Morquio A), characterized by symptoms like severe skeletal abnormalities, hearing loss, corneal clouding, and heart valve disease. | Keratan sulfate, and chondroitin 6-sulfate | Bone marrow | 103, 104 |
| ARSD | Arylsulfatase D |
Chondrodysplasia Punctata, Tibia-Metacarpal Type, characterized by flat midface and nose, short limbs, and otherwise normal development. | Sulfate esters | Kidney | 105, 106 |
| ARSE | Arylsulfatase E | X-linked Recessive Chondrodysplasia Punctata (CDPX1) |
Sulfate esters | Kidney, liver | 105, 106 |
| ARSF | Arylsulfatase F | Chondrodysplasia Punctata | Sulfate esters | Kidney skin | 105-107 |
| ARSG | Arylsulfatase G | Mucopolysaccharidosis IIIE, an autosomal recessive, neurodegenerative disorders with primary characteristic like degeneration of the central nervous system, mental retardation and hyperactivity | 3-sulfoglucosamine of heparan sulfatase | Thyroid, placenta, heart | 106, 108 |
| ARSH | Arylsulfatase family member H | - | Sulfate esters from sulfated steroids, carbohydrates, proteoglycans, and glycolipids | Esophagus, gall bladder, stomach | 31, 106 |
| ARSI | Arylsulfatase family member I | - | Sulfate esters and sulfamates | Skin, placenta, lung | 31, 106 |
| ARSJ | Arylsulfatase family member J | - | Sulfate esters from sulfated steroids, carbohydrates, proteoglycans, and glycolipids | Gall bladder, prostate | 38, 106 |
| ARSK | Arylsulfatase family member K | - | 2-sulfoglucuronate | Thyroid, ovary | 106 |
| SULF1 | Sulfatase 1 |
Mesomelia-Synostoses Syndrome, a rare autosomal-dominant disorder characterized by mesomelic limb shortening, acral synostoses, and multiple congenital malformations. | Heparan sulfate | Urinary bladder, prostate | 106, 109 |
| SULF2 | Sulfatase 2 | - | Heparan sulfate | Ovary, endometrium | 106, 110 |
Given that we can classify arylsulfatases broadly based on their localization and substrate specificity, it is pertinent to ask if enzymes with similar characteristics are related more closely in their phylogeny. A careful look at a maximum-likelihood phylogenetic tree, constructed with the primary structures of 17 human arylsulfatases shows that this is not the case. The proteins segregate into two major subclusters with the smaller one comprising of SULF1, SULF2, GNS and ARSK. As described in the previous section, the first two is localized extracellularly and the GNS and ARSK are lysosomal in their cellular location. In the larger subcluster, although GALNS constitutes a subgroup with ARSA and ARSG, the substrates on which they act are chemically distinct.
ARSI and ARSJ cluster closely with ARSB in consonance with their mutual paralogy (39). That ARSC, -D, -E, -F and -H form a separate subgroup can also be explained by the fact that they reside within a common locus on the short arm of chromosome X and are presumably products of gene duplications (31). Therefore, while their molecular origin through duplication has likely determined their phylogenetic relationship, their primary and especially tertiary structure likely diverged to distinguish their location and function (Figure 1).
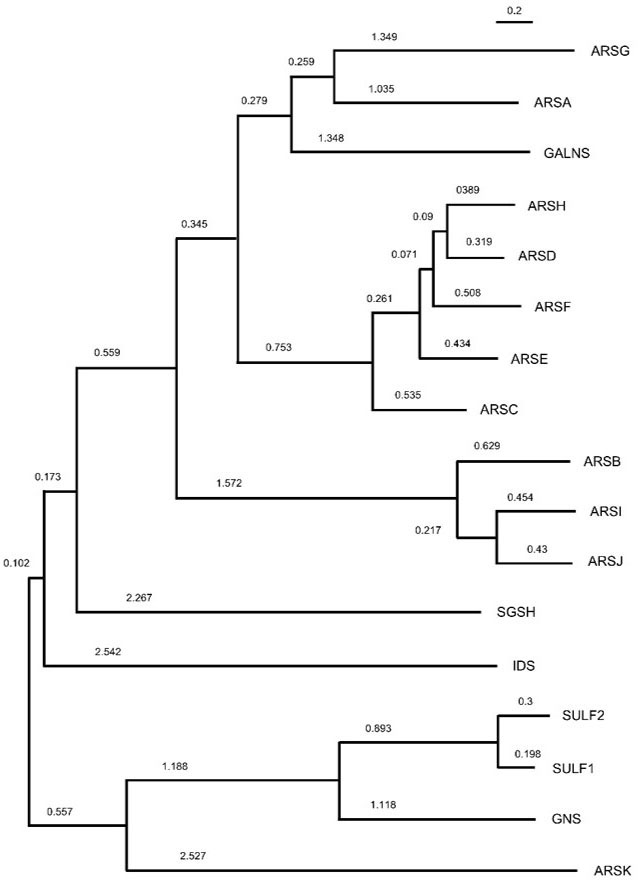 Figure 1
Figure 1Maximum-likelihood phylogenetic tree constructed using amino acid sequences of human arylsulfatase proteins. The numbers represent the length of branch.
Crystal structures of several human arylsulfatases have been established till date. These enzymes possess a signal peptide and a variable number of glycosylations (ARSA has three mannose-type N-linked glycan chains (40); ARSB, ARSC and IDS have 6, 4 and 8 potential N-glycosites respectively (41)). The active site of the enzymes consists of a cysteine, whose (-CH2-SH) moiety undergoes post-translational oxidation to (-CH=S). The resultant production of formylglycine (fGly) is facilitated by the fGly-generating enzyme (FGE) (42), which is encoded by the sulfatase-modifying factor 1 (SUMF1) gene (43). Upon impairment of this enzyme, all arylsulfatases are argued to be catalytically inactive, as evidenced in the lysosomal storage disorder known as multiple sulfatase deficiency (44).
The fGly residue is located in a positively-charged pocket of enzyme active site and acts as the ligand of an octahedrally-coordinated metal ion which could be Mg2+ or Ca2+, depending on the type of arylsulfatase. The metal ion may stabilize the formation of the sulfate ester (45). The catalysis for sulfate ester hydrolysis, mediated by fGly is unique, highly efficient and includes hydroxylation of the fGly residue by a water molecule, forming the activated hydroxyl-formylglycine (46). The active site of sulfatases consists to variable extents of positively charged residues for efficient binding of negatively charged substrates. Sulfatases such as ARSB and GALNS have ‘open flat positively charged active sites’ for polyanionic proteoglycan substrates (27, 47). In contrast the sphingolipid desulfating ARSA forms a suitably accommodative small active site pocket (45). On the other hand, the steroid ARSC possesses an active site consisting of hydrophilic and hydrophobic residues (48). The configuration of ARSA depends on the pH of the environment in its vicinity. At neutral pH, it is predominantly dimeric, while at lysosomal acidic pH it exists as a homo-octamer, composed of 4 dimers arranged in a ring-like structure (45). Unlike ARSA, ARSB and IDS occur as monomers (41, 49) whereas SGSH and GALNS present as homodimers (47, 50).
Arylsulfatases have variable domain architectures. For example, the first 33 residues of IDS comprise an N-terminal ER signal peptide and a pro-peptide and are cleaved before secretion. The residues from 34-550 adopt a compact, globular α/β sandwich fold and consists of two subdomains, the N-terminal SD1 (residues 34–443) and the C-terminal SD2 (residues 455–550). SD1 includes the ‘heavy' 42 kDa chain and contains the catalytic core while SD2 corresponds to the ‘light' 14 kDa chain (49). Each SGSH monomer also comprise two domains: a large N-terminal domain (domain 1) and a smaller C-terminal domain (domain 2). Domain 1 possesses the consensus active site along with metal ion Ca2+ (50).
The two extracellular sulfatases, Sulf1 and Sulf2 (the Sulfs), are endosulfatases with specific substrate specificity toward 6O-sulfate groups of heparan sulfate (51, 52). Sulf1 and Sulf2 have a large hydrophilic domain (HD), which is located between the N-terminal catalytic domain and the C-terminal domain. The HD is a unique feature of the extracellular sulfatases and is not found in other sulfatases. The HD of human Sulf1 has a size of ∼320 amino acid residues, 27% of which are basic and 14% acidic, resulting in a strong positive charge at neutral pH. On the other hand, the C-terminal end of the HD is composed of 12 basic amino acid residues (53). HSulf-1 and HSulf-2 protein consist of four domains. The order from the N to C terminus includes a signal peptide, a catalytic domain of 374 amino acids, a basic hydrophilic domain of 346/366 amino acids, and a C-terminal domain of 109/127 amino acids (51).
Both Sulfs are produced as preproenzymes and are later cleaved by a furin-type proteinase to form heterodimers of 75- and 50-kDa subunits. Even though the catalytic center resides in the N-terminal 75-kDa subunit, the C-terminal 50-kDa subunit is essential for both arylsulfatase and glucosamine 6-O-sulfate-endosulfatase activity of the Sulf enzymes. Hydrophilic regions of the Sulfs is critical for endosulfatase activity but not essential for arylsulfatase activity (52, 53).
The altered degradation of sulfated conjugates as a result of perturbation in expression and activity of arylsulfatases may lead to the disruption of cellular homoeostasis. The effect may be felt at the scale of an organelle, such as the lysosomes (54), or organisms such as developmental abnormalities and cancer (6). In this section, we review the literature on the effects of arylsulfatases that act on proteoglycans.
Of the lysosomal sulfatases, the most well-studied one shown to act on GAG desulfation is ARSB. ARSB acts on, and participates in, the degradation of chondroitin and dermatan sulfates that contain N-acetylgalactosamine-4 sulfates (55). In general, investigators have noted a decrease in the activity or expression of ARSB in malignantly transformed cells in comparison with their untransformed counterparts (56-58). In fact, investigations by Tobacman’s group shows that the depletion of ARSB mimics hypoxia and leads to increase in nuclear levels of hypoxia inducible factor (HIF1α). Hypoxia and ARSB depletion individually lead to increase in sulfated GAG (sGAG) levels as well as accumulation of Galectin-3 (GAL-3) in the nucleus. GAL-3, which can otherwise bind to sGAGs, upon hypersulfation of the latter (as a result of ARSB knockdown), changes preference and instead binds to Sp1 (specificity factor-1) which in turn binds to the promoter of Wnt-9a and increases its expression in colon cancer (59). The Gal-3 unbound hypersulfated sGAGs, especially chondroitin-4 sulfate (C4S) is free to bind to the non-receptor tyrosine phosphatase SHP2, which results in upregulation of both MMP-2 and Erk signaling activity in malignant melanoma cells (58).
The expression and activity of ARSB has also been shown to be lower in prostate cancer in comparison with normal tissues, leading in turn to an increase of total sulfated glycosaminoglycans and chondroitin-4-sulfate content within the prostatic tumor microenvironment. According to an interesting report, ARSB can be a useful prognostic biomarker of prostate cancer than prostate-specific antigen (PSA), the gold standard for diagnosis of prostate carcinoma (57). Increased chondroitin 4- sulfate levels as a result of low ARSB activity leads to up-regulation of EGFR and EGFR mediated tyrosine kinase signaling pathways and thus escalate the proliferation of prostate cells (60). Since EGFR plays a key role in cellular proliferation and cancer, the EGFR-mediated malignancies can be managed by restoration of ARSB activity with the associated reduction in chondroitin-4 sulfate levels.
There is a very little information on the functions of lysosomal sulfatase sulfamidase (also known as N-sulfoglucosamine sulfohydrolase (SGSH)) and Iduronate 2-sulfatase (IDS) in cancer. SGSH is ubiquitously expressed throughout the body with the highest protein levels seen in kidney, lung prostate and gall bladder, all organs with an epithelial parenchyma (61). Prostate cancer sections may show moderate immunoreactivity against SGSH antibodies although why it is specifically upregulated in this cancer remains unknown. For IDS, a survey of its gene expression in various cancers shows a progressive stage-specific decrease in carcinoma of breast, bladder and lung (62). In addition, its expression is significantly lower in tumors of prostate and ovaries and in glioblastoma compared with their non-malignant tissue counterparts (61).
The enzyme N-acetylglucosamine-6-sulfatase (GNS) mediates the hydrolysis of 6-sulfate groups of the N-acetyl-D-glucosamine 6-sulfate units of heparan- and keratan- sulfates and has been not been subjected to any oncological investigations, although transcript levels show lower levels in primary breast and lung tumors compared with normal tissues whereas levels in glioblastoma are higher than in controls (62). The expression of the Morquio syndrome-causing N-acetylgalactosamine-6-sulfatase (GALNS) is elevated in primary tumors of the breast and lung (and depleted in primary tumors of the prostate) (62), although whether it is a cause of oncological progression of the disease or a consequence is yet to be investigated.
Of the non-lysosomal sulfatases, ARSC is involved in desulfation of steroid substrates and the functions of ARSD, E and F are yet to be clearly established. Two other sulfatases, Heparan sulfate 6-O-endosulfatases, encoded by Sulf1 and 2 are secreted to the extracellular milieu. By acting on heparan sulfate proteoglycans (HSPGs) residing on the cell surface, Sulf1/2 modulate a variety of signal transduction pathways that are involved in regulation of cell division, migration and polarity. Sulf1 has been shown to be upregulated in cancers of liver, stomach, breast and pancreas (63-66).
Sulf1 has in general, been shown to suppress the induction of tumorigenic signal transduction mediated by heparin-binding growth factor (67). In ovarian cancer, its expression, otherwise observed in normal surface epithelia is diminished in cancer cell lines (68). Loss of Sulf1 is accompanied by an increased sulfation of HSPGs followed by increased Erk signaling resulting in loss of the proapoptotic protein Bim (69). Sulf1 levels also regulate the status of autophagy: loss of Hsulf1 decreases autophagic activity and increases lipid biogenesis and glycolysis (70, 71). Sulf1 expression also downregulates growth in hepatocellular carcinoma and induces apoptosis in pancreatic cancer (65, 67, 72, 73). The effect of Sulf1 levels on cancer progression was also observed in vivo: MDA-MB-468 derived xenografts showed diminished burden and reduced angiogenic vessel density in a VEGF-dependent manner when Sulf1 expression was knocked down (74).
Unlike Sulf1, the endosulfatase Sulf2 plays a largely inductive role in cancer progression. Higher levels of Sulf2 have been detected in hepatocellular cancer cell lines (75). Sulf2-deficient mice show relative resistance to diethylnitrosamine-induced carcinogenesis. Overexpression of Sulf2, on the other hand, brings about increased proliferation, migration and neoangiogenesis. The angiogenic action is mediated through a TGF-β-dependent upregulation of periostin (76). Sulf2 overexpression in prostate cancer cells leads to increased viability, migration and an enhanced ability to form colonies in noble agar. In addition, it led to increased Wnt signaling and spurred epithelial to mesenchymal transition of cancer epithelia (77). Jung and coworkers confirmed using culture and murine models that cancer cells that survive ionizing radiation show elevated levels of Sulf2 that stabilize β-catenin in order to upregulate IL6: elevated levels of both Sulf2 and IL-6 enhanced radiation-induced metastasis in murine models (78). Increased SUlf-2 levels were also seen in sections of lungs from patients afflicted with the non-small cell lung carcinoma (NSCLC). In NSCLC-derived cell lines, depletion of SUlf-2 leads to a phenotypic reversion of cancer epithelia, through the abolition of oncogenic autocrine Wnt signaling (79). This also results in decreased xenograft tumors in nude mice. Overexpressing SUlf-2 in untransformed bronchial epithelia leads to an increased ability of the cells to migrate, proliferate and give rise to colonies in noble agar. Using cell lines and immunocompromised mice, Solari and coworkers show that MYCN amplification manifests its effects on proliferation and growth through upregulation of Sulf-2. Higher levels of the latter strongly correlated with poorer outcome in neuroblastoma patients (80).
That the effect of Sulf-2 in onco-progression is dependent on tissue-specificity is borne out by its role in renal cell carcinoma (RCC): Patients with higher expression of Sulf-2 showed better survival than those with lower levels. In RCC cell lines, overexpression of Sulf-2 led to higher Wnt-specific proliferation whereas low levels led to greater proliferation as well as invasion in an FGF-VEGF-dependent manner (81).
Upon transformation of epithelia within an organ, the ECM starts undergoing important changes in not just its composition but also its mechanical, physical and material properties (82). This is because tumorigenesis results in the recruitment and activation of connective tissue- and immune cells, and the tumor ECM is remodeled through degradation and synthesis of molecules by these cells along with cancer cells (83). This process, known as desmoplasia, results in an increase in stiffness of the ECM which correlates with increased invasion and epithelial to mesenchymal transition of cancer cells as a result of the transduction of altered ECM to the cellular adhesome (84). Increased stiffness results in the nuclear translocation of the mechanosensing protein Twist1, that drives EMT, invasion and metastasis (85). Desmoplastic stiffness in pancreatic cancer cell lines results in the decrease in E-cadherin expression and increased nuclear translocation of β-catenin and expression of vimentin (86).
The altered rheology of the ECM can be understood by the concurrent occurrence of multiple processes: cancer cells and associated microenvironmental fibroblasts are known to secrete higher levels of fibrillar matrix proteins such as collagen (87). In addition, cells express high levels of the enzyme lysyl oxidase (LOX), which enhance the cross-linking of collagen fibers (88). The proteoglycan content of tumor matrix is also altered (89). Increased hyaluronan and dermatan sulfates (DS) have been shown to correlate with an increase in stiffness in ECM, as well as invasion and EMT in cancer cells (90-92). The presence of acidic sugars along with sulfates makes these GAGs highly negatively charged, which in turn makes the matrix more hydrated, gel-like and consequently resistant to compressive forces (93). Heightened expression of sulfotransferases or the downregulation of sulfatases will potentially enhance the sulfation of such GAGs accentuating in turn their induction of tumor stiffness.
In this review, we have endeavored to explore the role of desulfation of glycosaminoglycans (GAGs) in the context of cancer progression and metastasis. We review literature that argues that sulfated GAG levels may impact cellular processes involved in cancer invasion, such as their transition from epithelial to mesenchymal morphologies. This may be through their modulation of signaling in cancer epithelia or through altering the biophysical nature of the tumor matrix (Figure 2).
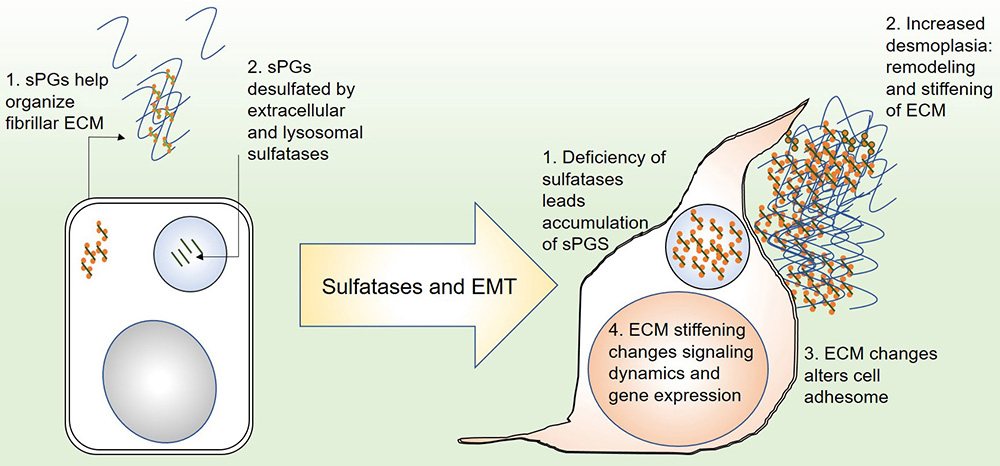 Figure 2
Figure 2Depiction of the role of proteoglycan sulfatases in epithelial to mesenchymal transition: (left) in an epithelial state, secreted sulfated proteoglycans organize local fibrillar matrix. Their degradation in the extracellular milieu and in lysosomes maintains reciprocal homoeostasis between cells and their matrix microenvironment. (right) Deficiency or downregulation of sulfatases leads to accumulation of sulfated proteoglycans in the matrix, leading to abnormal remodeling and increased stiffness (desmoplasia). This in turn alters the cortical cytoskeleton of the cells and the localization of matrix adhesion cues. At the same time, outside in signaling is altered leading to aberrant transcriptional response which changes cell behavior.
Several open questions remain: Given that the perturbations of some sulfatases may alter the levels of multiple GAGs, the cumulative effect of concurrent alteration of multiple GAGs on matrix stiffness and cell behavior is difficult to predict. Moreover, the effect of even discrete GAGs on tissue phenotype is reported to be complex: breast cancer epithelia exposed to different levels of DS show opposite phenotypes (94). Non-canonical localization in cancer cells suggests that the enzymes may also exert functions independent of their enzymatic activity and catalytic domain. Future cell biological investigations will allow us to gain better insight into the functions of such arylsulfatases, which may then be examined as druggable targets for future cancer therapy.
The authors declare no competing interests. RB would like to acknowledge funding support from SERB ECR fellowship (DSTO1586) and CSIR (1671). VS would like to acknowledge support from SERB NPDF fellowship (DSTO1751).


