1 School of Mental Health, Jining Medical University, 272000 Jining, Shandong, China
2 Department of Psychiatry, Shandong Daizhuang Hospital, 272000 Jining, Shandong, China
3 Cheeloo College of Medicine, Shandong University, 250000 Jinan, Shandong, China
†These authors contributed equally.
§DIRECT: Depression Imaging Research Consortium.
Abstract
Major depressive disorder (MDD) is associated with altered organization of functional brain networks. This study aims to evaluate the classification efficacy of three brain networks constructed by Pearson correlation (PC), sparse representation (SR), and group sparse representation (GSR) in distinguishing patients with MDD from healthy controls (HCs).
The present study involved the recruitment of 117 Chinese participants, comprising 61 individuals diagnosed with MDD and 56 HCs, all of whom underwent functional magnetic resonance imaging (fMRI). Brain time-series signals were extracted from 116 regions to construct whole-brain networks utilizing PC, SR, and GSR. A linear support vector machine (SVM) classifier with least absolute shrinkage and selection operator (LASSO) feature selection was trained using leave-one-out cross-validation (LOOCV) to optimize generalizability. An independent dataset of Chinese (124 first-episode drug-naïve MDD and 105 HCs) was utilized for additional validation.
Compared to the PC and SR, the GSR network yielded superior classification results, with an area under the receiver operating characteristic curve of 0.85, an accuracy of 0.81, and a sensitivity of 0.95. Similar results were observed in the independent MDD dataset. We identified 17 brain connections and 27 brain regions within the GSR network.
Our findings support the adoption of GSR-based brain networks as a robust tool for MDD diagnosis, challenging the conventional reliance on PC in neuroimaging research.
Keywords
- major depressive disorder
- sparse representation
- support vector machine
- classification
- brain networks
1. The Group Sparse Representation (GSR) method built a brain network that classified Major Depressive Disorder patients significantly better than standard methods, achieving high accuracy and sensitivity.
2. This superior performance was validated in an independent patient dataset, confirming its robustness.
3. The findings establish GSR as a powerful and preferable alternative to the commonly used Pearson Correlation for building diagnostic brain networks in psychiatry research.
Major depressive disorder (MDD) represents a significant contributor to the global psychiatric burden, impacting approximately 300 million individuals worldwide [1]. Over the past decades, substantial progress has been achieved in elucidating the pathophysiological mechanisms underlying MDD [2, 3, 4]. Notably, the brain network has emerged as a prevalent tool for assessing brain functions in individuals with MDD [5, 6, 7].
Brain functional connectivity (FC) between distinct brain regions serves as a primary method for analyzing resting-state functional magnetic resonance imaging (rs-fMRI) data [8, 9, 10]. The construction of a functional brain network typically involves the use of Pearson correlation (PC) [11, 12], which assesses the pairwise linear interactions between different brain regions, with the PC coefficient representing the connectivity weight. Previous studies [13, 14, 15] have extensively documented the application of sparse representation (SR) in various domains such as decomposition, dimensionality reduction, reconstruction [16], face recognition [17], and brain tissue segmentation [18]. In the context of functional brain network research, SR facilitates the construction of sparse networks by evaluating the interrelationships among multiple brain regions at an individual level [14]. The signals from a specific brain region can be represented as a linear combination of signals from other regions, with the corresponding combination weights interpreted as the connections among these regions [14]. The SR method provides enhanced options for constructing brain networks in both healthy individuals [19] and patients with mild cognitive impairment (MCI) [20, 21] and Alzheimer’s disease (AD) [22].
Group sparse representation (GSR) is predominantly employed in the classification of multi-feature, multimodal biometrics [23] and hyperspectral images [24]. In contrast to SR, the GSR network incorporates group constraints and maintains uniformity across all subjects while preserving individual-specific information [25]. The GSR method generates a sparse network through the joint selection or elimination of specific connectivity links applicable to all subjects. Research has demonstrated that the GSR network can be used to effectively classify patients with MCI [25].
To the best of our knowledge, these brain network construction methods have predominantly been employed for the classification of patients with MCI [20, 25, 26] and AD [22]. However, their application in the context of MDD remains inadequately explored. MDD is characterized by significant heterogeneity, state-dependent manifestations, and complex symptomatology, yet it also exhibits certain intrinsic pathological markers whose stability is underrecognized. While neurodegenerative research frequently concentrates on fixed patterns of degeneration, MDD-optimized GSR is capable of capturing both transient state effects and stable trait markers through the application of group constraints. This capability is essential for the study of mood disorders.
The present study sought to evaluate and compare the efficacy of prevalent
methodologies in the automatic classification of patients with MDD. A cohort
comprising 61 individuals diagnosed with MDD and 56 healthy controls (HCs) was
recruited. Brain networks were constructed utilizing PC, SR, and GSR. A linear
kernel support vector machine (SVM) classifier was developed to distinguish
between MDD patients and HCs. The least absolute shrinkage and selection operator
(LASSO) algorithm was employed to select brain connections within each network as
features for SVM training. The diagnostic performance was assessed using the
leave-one-out cross-validation (LOOCV) approach. The classification performance
was quantified through metrics including the area under the receiver operating
characteristic (ROC) curve (AUC), accuracy (ACC), sensitivity (SEN), and
specificity (SPE). To enhance the validity of our findings, we employed an
independent dataset of MDD, sourced from the R-fMRI Maps Project (https://rfmri.org/maps)
[27]. We hypothesized that the GSR would surpass alternative methodologies by
simultaneously modeling consistent MDD network pathology through the application
of
The study enrolled 61 patients diagnosed with MDD and 56 HCs, all of whom were Chinese, between 18 and 50 years, and who possessed a minimum of seven years of education, and were right-handed (refer to Table 1). The recruitment of MDD patients was conducted through outpatient services from August 2023 to December 2024, while HCs were recruited concurrently via online advertisements. All participants underwent interviews conducted by two experienced psychiatrists utilizing the Structured Clinical Interview for DSM-IV (SCID) [28] and the Mini International Neuropsychiatric Interview (MINI). The severity of depression was evaluated using the 17-item Hamilton Rating Scale for Depression (HAMD-17) [29].
| MDD (n = 61) | HCs (n = 56) | t/z/ |
p (two-tailed) | |
| Age | 35.07 |
31.58 |
t = 1.49 | 0.14 |
| Gender (male/female) | 26/35 | 31/25 | 0.17 | |
| HAMD | 13.98 (7.00, 35.00) | 0.69 (0.00, 4.00) | z = 8.41 |
Note: The HAMD scores in the HC group were non-normally distributed, whereas
those in the MDD group followed a normal distribution (mean
The inclusion criteria for participants with MDD were as follows: (1) age between 18 and 50 years; (2) minimum primary education completion; (3) provision of written informed consent; (4) diagnosis of unipolar MDD episodes following the SCID and MINI; (5) a score of 8 or higher on the HAMD-17; (6) avoidance of all psychotropic medications for a minimum of two weeks prior to study enrollment (four weeks for fluoxetine due to its prolonged half-life). The exclusion criteria for all participants included: (1) a history of epilepsy or brain trauma; (2) severe physical diseases; (3) high risk of suicide; (4) electroconvulsive therapy or transcranial magnetic stimulation within 6 months; (5) pregnancy. In this study, all participants provided written informed consent, and the study protocol received approval from our institutional review board (NO. 202311-HY-1).
All participants underwent scanning using a 3.0 Tesla Siemens Trio MRI scanner
(Erlangen, Germany) following specific imaging protocols. Participants were
instructed to keep their eyes closed, remain awake and relaxed, and minimize
movement during the scan. The resting-state functional MRI (rs-fMRI) parameters
were as follows: repetition time (TR)/echo time (TE) = 2000/30 ms, flip angle =
90°, matrix size = 64
We preprocessed the fMRI data utilizing the Data Processing Assistant for
Resting-State fMRI (DPARSF, V4.2, https://rfmri.org/DPARSF) toolbox [30], which operates within the MATLAB (R2022b, https://www.mathworks.com/products/new_products/release2022b.html)
environment. The preprocessing steps were as follows: (1) the first ten time
points of the rs-fMRI images were discarded due to instability of the initial MRI
signal, leaving 230-time points; (2) correction for the acquisition time delay
between slices and further realigned to the first volume to correct for head
motion incorporating nuisance covariate regression using the Friston 24-parameter
model; we reduced respiratory and cardiac effects by using signals from
segmentation of the white matter (WM) and cerebrospinal fluid (CSF) compartments
in the 3D T1-weighted image as regressors; (3) co-registered T1 structural images
to functional images via a nonlinear image registration approach, segmented using
a new segment algorithm with diffeomorphic anatomical registration through
exponentiated lie algebra (DARTEL); (4) movement parameters for each participant
were assessed and participants were excluded if movement exceeded 2 mm or
2° of translation or rotation in any direction; (5) A band-pass
frequency filter method (0.01 to 0.08 Hz) was applied to reduce physiological
high-frequency noise. The rs-fMRI images were spatially normalized into the
Montreal Neurological Institute (MNI) template, and resampled into a spatial
resolution of 3
In this study, blood oxygen level-dependent (BOLD) signals were extracted from
the entire brain using the Automated Anatomical Labeling (AAL) template, which
comprises 116 regions of interest (ROI) [31]. It was assumed that the time series
signal
For any two ROI with time series vectors x and y of length T (time points), the Pearson correlation coefficient r was calculated as:
where
In contrast to the PC method, the SR brain network was constructed utilizing
linear regression. Brain signals from the rth ROI
where
Based on the SR method, the GSR network was constructed by using l2,1-norm regularization across all subjects within-group [15] as Eqn. (3):
where
The connection coefficients calculated using the PC, SR, and GSR methods were extracted as features to represent network properties. To minimize the feature set, we employed the widely adopted LASSO technique for feature selection in the training of the SVM classifier.
In this study, a linear kernel SVM was employed to evaluate the discriminative
capability of features extracted from three distinct network methodologies.
Feature selection was conducted using the LASSO. The regularization parameter
(
We used an independent dataset of Chinese individuals diagnosed with MDD from the R-fMRI Maps Project (rfmri.org/maps) [27] to further substantiate our findings. In this study, we included a total of 229 participants, comprising 124 unmedicated patients experiencing a drug-naïve first episode of MDD and 105 HCs. This independent validation cohort with both similarities and distinctions versus our primary cohort, and matched Chinese ethnicity, and age range. Both used SCID/MINI diagnosis, and HAMD. The demographic details of the participants in this publicly available dataset are presented in Supplementary Table 2. The preprocessing of fMRI data and the analysis of classification performance were conducted using the same methodologies as applied to our own MDD sample set.
All continuous demographic variables (age, HAMD-17 scores) were assessed for
normality using the Shapiro-Wilk test (
Functional brain networks constructed via PC, SR, and GSR exhibited distinct topological patterns between MDD patients and HCs (Fig. 1A–F). The detailed numerical values corresponding to these six connectivity patterns displayed in Fig. 1A–F are shown in the Supplementary Excel Files.
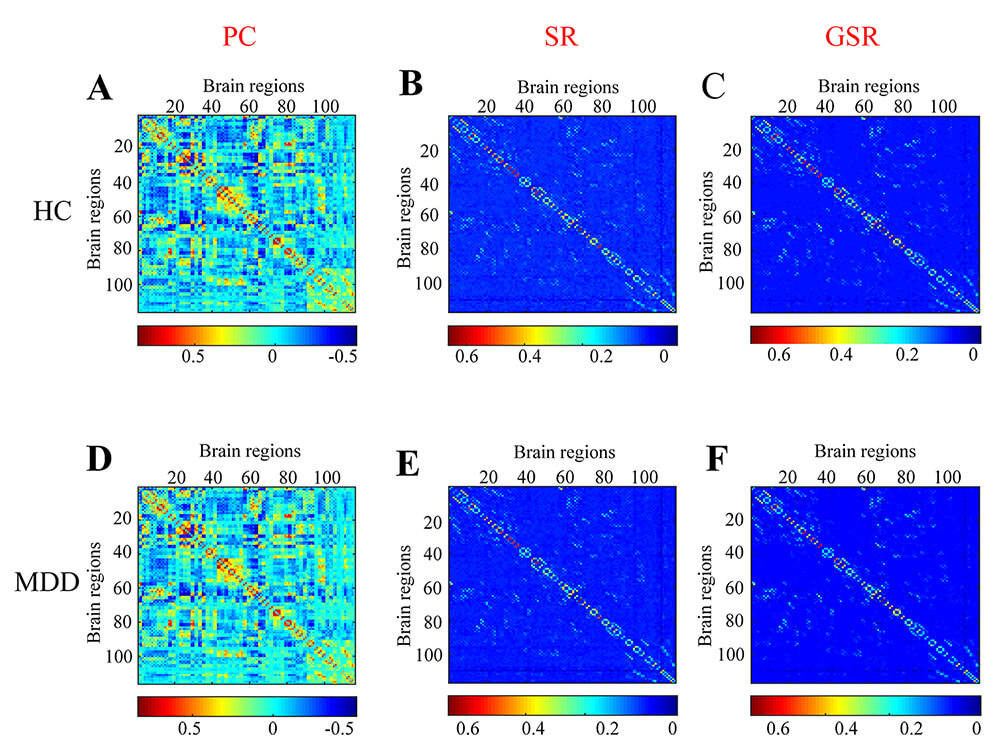 Fig. 1.
Fig. 1.
The functional brain networks derived using the Pearson correlation (PC), sparse representation (SR), and group sparse representation (GSR) methods for HCs (A–C) and MDD (D–F).
Fig. 2 demonstrates that the GSR method attained the highest values for AUC, ACC, and SEN, with 0.85, 0.81, and 0.95, respectively. In contrast, the PC network exhibited the lowest ACC and SEN, with values of 0.54 and 0.55, and achieved the highest SPE with 0.50. The SR method recorded the lowest AUC and SPE, with values of 0.52 and 0.28, respectively.
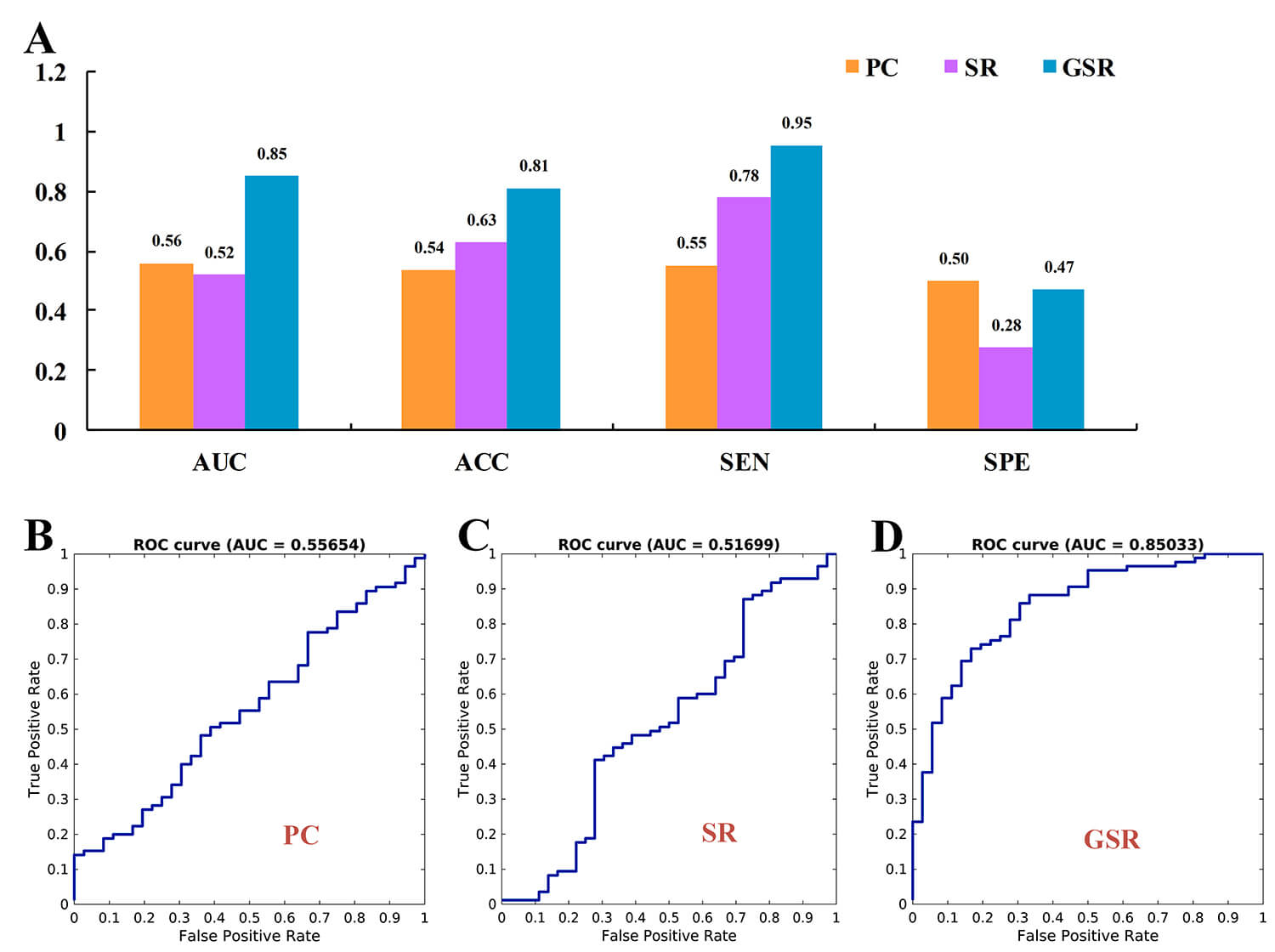 Fig. 2.
Fig. 2.
Classification performance metrics for the Pearson correlation (PC), sparse representation (SR), and group sparse representation (GSR) methods. (A) presents the area under the curve (AUC), accuracy (ACC), sensitivity (SEN), and specificity (SPE). (B–D) display the receiver operating characteristic (ROC) curves for PC, SR, and GSR, respectively.
In the independent validation cohort, the GSR method demonstrated superior performance, achieving the highest values for AUC, ACC, SEN, and SPE, with respective values of 0.65, 0.66, 0.72, and 0.59. In contrast, the PC network recorded the lowest performance metrics, with AUC, ACC, SEN, and SPE values of 0.50, 0.46, 0.20, and 0.50, respectively. The SR method exhibited moderate performance, with AUC, ACC, SEN, and SPE values of 0.60, 0.59, 0.64, and 0.54, respectively. See Supplementary Fig. 1.
Fig. 3A,B illustrates the most discriminative brain connections within the GSR
network as identified by the SVM classifier. The mean weight assigned to each
connection and the normalized frequency of each connection’s occurrence are shown
in Supplementary Excel Files. The GSR network achieved a peak accuracy
of 0.81 (Fig. 3C) and demonstrated the highest frequency of occurrence in the
model robustness evaluation (Fig. 3D) when the regularization parameter
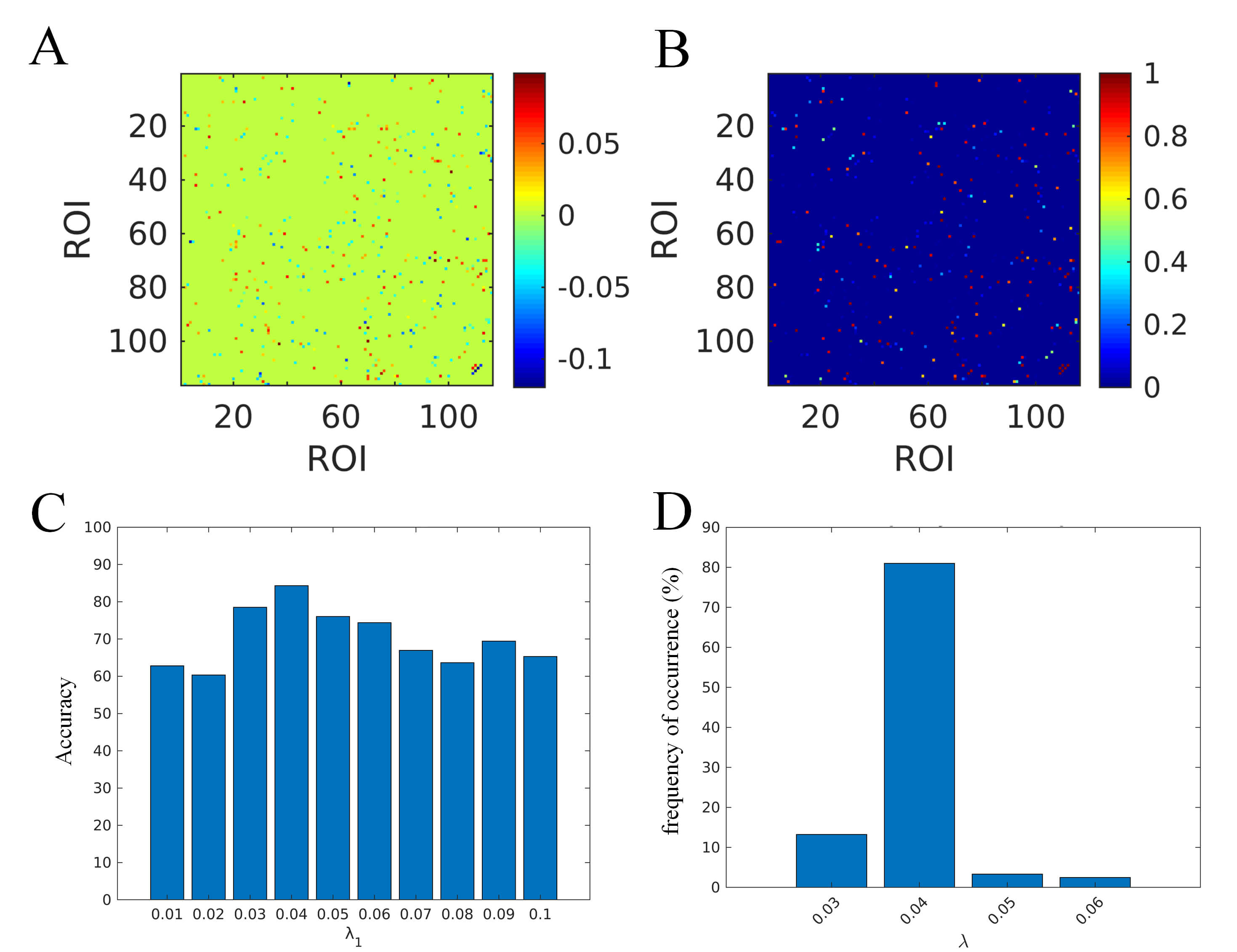 Fig. 3.
Fig. 3.
The following elements pertain to the optimal classification model for GSR. (A) the mean weight assigned to each connection; (B) the normalized frequency of each connection’s occurrence; (C) the classification accuracy associated with each lambda value; (D) the frequency of occurrence for each lambda value. ROI, regions of interest.
The performance comparison results and optimal models for these three methods, as applied to the public dataset, are presented in the Supplementary Figs. 1,4,5,6). The GSR method demonstrated superior performance within this dataset, as illustrated in Supplementary Fig. 1.
Table 2 provides a summary of the 17 discriminative brain connections and 27 brain regions identified within the GSR network, as determined by the weighting coefficients of the SVM analysis. These 27 brain regions comprise 14 cortical regions, 3 subcortical regions, and 10 cerebellar areas. These connections and regions demonstrate that both cortical and cerebellar regions participate in local connections (within cortical or cerebellar regions) as well as global connections (extending beyond their respective regions to interact with other regions). The subcortical regions exhibit two connections with cerebellar regions and one connection with the cortical regions.
| Connection number | ROI1 | ROI2 | ||
| Index | Brain region names | Index | Brain region names | |
| 1 | 24 | Right superior frontal gyrus, medial | 11 | Left inferior frontal gyrus, opercular part |
| 2 | 64 | Right supramarginal gyrus | 35 | Left posterior cingulate gyrus |
| 3 | 65 | Left angular gyrus | 38 | Right hippocampus |
| 4 | 66 | Right angular gyrus | 47 | Left lingual gyrus |
| 5 | 65 | Left angular gyrus | 55 | Left fusiform gyrus |
| 6 | 86 | Right middle temporal gyrus | 78 | Right thalamus |
| 7 | 95 | Left cerebellum.3 | 67 | Left precuneus |
| 8 | 93 | Left cerebellum.Crus2 | 69 | Left paracentral lobule |
| 9 | 95 | Left cerebellum.3 | 70 | Right paracentral lobule |
| 10 | 96 | Right cerebellum.3 | 11 | Left inferior frontal gyrus, opercular part |
| 11 | 100 | Right cerebellum.6 | 70 | Right paracentral lobule |
| 12 | 105 | Left cerebellum.9 | 71 | Left caudate nucleus |
| 13 | 112 | Vermis.6 | 75 | Left lenticular nucleus, pallidum |
| 14 | 112 | Vermis.6 | 30 | Right insula |
| 15 | 112 | Vermis.6 | 109 | Vermis.1.2 |
| 16 | 114 | Vermis.8 | 70 | Right paracentral lobule |
| 17 | 111 | Vermis.4.5 | 110 | Vermis.3 |
Note: In the same row, a connection exists between ROI1 and ROI2. The regions of interest (ROI) were labeled according to the AAL116 template, with both the indices and the names of the brain regions referenced from this template.
The present study investigated the classification performance of brain networks which were constructed using PC, SR, and GSR in Chinese patients diagnosed with MDD. The results indicated that GSR achieved the highest classification performance, suggesting that brain connections that incorporate both inter-subject variability and within-group similarity may be more effective in identifying patients with MDD. This conclusion was further confirmed by an independent dataset of Chinese MDD patients. The most discriminative brain connections were identified between the cerebral cortex and the cerebellum, suggesting their pivotal role in distinguishing MDD patients from HCs and challenging the traditional limbic-centric model of depression pathophysiology. The findings of this study make significant contributions to advancing the field of MDD identification and provide additional evidence regarding the classification efficacy of sparse brain networks.
The PC network has been extensively investigated in the context of various neurological disorders [33]. The PC method was the predominant technique employed in neuroimaging research [34, 35]. This correlation method cannot reveal the interaction effects among several brain regions [34]. In the present study, the SR network, which incorporated inter-regional brain effects into its network construction, demonstrated superior classification performance compared to the PC method. Moreover, the GSR network, which integrates both individual and group-level information into the SR network, achieved the highest classification performance. These findings are consistent with previous research on patients with MCI [20, 25, 26] and AD [22].
The SR network exhibited increased inter-subject variability, resulting in distinct network topological structures for individual subjects [36]. This variability could potentially impair generalization ability due to the heterogeneity or inconsistency across subjects [25, 36], despite achieving superior classification performance compared to PC. In contrast, the GSR method sparsely represented brain connections at the group level, simultaneously enforcing intrinsic local sparsity and nonlocal self-similarity within a unified framework. The brain network derived from GSR encompassed both inter-subject variability and within-group similarity [37], thereby contributing to optimal classification performance in MDD patients.
The GSR approach exhibits heightened sensitivity to network abnormalities specific to MDD in both datasets, potentially due to its distinctive capability to model concurrently. Specifically, the l2,1-norm regularization enforces group-wise sparsity by selectively preserving connections that consistently exhibit alterations across MDD patients, while simultaneously eliminating noisy individual variations. Furthermore, GSR maintains group consistency while allowing for subject-specific weight adjustments for preserved connections [15, 38], thereby accommodating the clinical heterogeneity inherent in MDD. This dual capacity of GSR to identify both shared biomarkers and personalized variants renders it particularly well-suited for addressing the complex pathophysiology of MDD, where population-level abnormalities coexist with clinically significant heterogeneity.
The relatively low SPE observed across methods (PC: 0.50, SR: 0.28, GSR: 0.47)
highlights two critical considerations. First, the extreme sparsity of SR may
lead to the exclusion of essential negative controls that are present in the
denser PC networks, which also led to the lowest AUC value of 0.52. In contrast,
GSR constraints more effectively preserve these discriminative null connections.
Second, the notably low SEN and AUC of SR are consistent with evidence indicating
that subtypes of MDD exhibit divergent patterns [39, 40, 41], resulting in the
misclassification of true negatives when employing individual-level sparse
networks. Additionally, LASSO’s tendency to favor positive correlations may lead
to a disproportionate selection of connections that exhibit MDD
hyperconnectivity. However, this bias was mitigated in GSR through the use of the
The discriminative connections identified through GSR network analysis align with established neuropathological findings in MDD, elucidating a coherent framework of circuit-level dysfunction [42, 43]. The left inferior frontal gyrus (IFG), recognized as a hub for cognitive control and regulation of emotion [44, 45, 46], exhibits altered connectivity patterns that directly contribute to core depressive symptomatology [47, 48, 49]. Its impaired connections with both the right superior frontal gyrus and the cerebellum indicate: (1) disrupted top-down cognitive control, as evidenced by deficits in executive function; (2) abnormal emotional processing, manifesting as a negative bias; and (3) dysregulated monoaminergic modulation, supported by cerebellar-prefrontal neurotransmitter pathways observed in animal studies [50, 51, 52, 53]. This dysfunction within the prefrontal-cerebellar circuit operates in conjunction with abnormalities in the limbic system, where alterations in angular gyrus-hippocampus connectivity are associated with maladaptive memory processes, such as overgeneralized negative recall [54, 55]. Furthermore, insula-vermis dysregulation underlines characteristic somatic symptoms, including fatigue and altered pain perception. The involvement of bilateral paracentral lobules further indicates potential deficits in sensorimotor integration, which may contribute to psychomotor disturbances observed in MDD [56]. These network abnormalities collectively impair higher-order executive functioning through three primary mechanisms: (1) cognitive-emotional integration failure, where prefrontal-cerebellar dysconnectivity disrupts the capacity for regulation of emotion; (2) memory processing bias, in which limbic-cerebellar abnormalities promote a preferential recall of negative experiences; and (3) sensorimotor dysynchronization, where the involvement of the paracentral lobes alters bodily perception and motor response. Future research should explore whether these connectivity patterns exhibit differential sensitivity to specific treatment modalities, such as repetitive transcranial magnetic stimulation targeting prefrontal versus cerebellar nodes.
It should be noted that the limbic system is frequently identified as an
atypical brain region in individuals with MDD [57, 58, 59]. The absence of limbic
connections (e.g., amygdala-hippocampus-prefrontal pathways) in our GSR reflects
the critical GSR’s group-consistency requirement (
Several methodological limitations should be acknowledged in this study. First,
our analysis was restricted to undirected functional connectivity networks,
which, while computationally efficient, may not fully capture the directional
information flow between brain regions that could provide greater neurobiological
insight. Future research should more effectively incorporate connectivity methods
[60] and alternative network construction approaches (partial correlation,
graphical LASSO, mutual information) to better characterize the complex,
potentially directional interactions in MDD. Second, while this study employed
three methods for network construction, there are additional methodologies worth
considering, such as the strength and similarity-guided GSR approach [15]. The
primary objective of this study was to evaluate the classification performance of
methods commonly utilized in brain network research within a cohort of
individuals with MDD. Consequently, recently proposed methods that have not been
widely adopted were not included. Moreover, Sparse methods (SR/GSR) enhance
interpretability by emphasizing dominant neural connections. However, the
sparsity induced by the choice of the regularization parameter (
In this study, we evaluated the classification performance of functional brain networks constructed using PC, SR, and GSR in patients with MDD. Utilizing the SVM algorithm, our results demonstrated that the GSR approach yielded superior classification performance. This finding suggests that brain networks that integrate individual connectivity information into a group framework may be more effective for identifying MDD patients. Our results imply that while PC-based networks are prevalent in numerous studies, they may not represent the optimal method. This research contributes additional evidence regarding the efficacy of the classification of sparse brain networks.
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
DZ , CL, HY, and SJ conceived the study, designed the experiments, and drafted the manuscript. AZ performed data analysis and created visualizations. XW and WX provided key reagents and validated the findings. YW and YD analyzed and interpreted data for the work. HY and SJ designed the research study, jointly supervised the project, acquired funding, and reviewed and edited the manuscript. All authors contributedto critical revision of the manuscript for important intellectual content. All authors read and approved the finalmanuscript. All authors have participated sufficiently inthe work and agreed to be accountable for all aspects of thework. The DIRECT Consortium provided shared data.
This study was approved by the Institutional Review Board of Shandong Daizhuang Hospital (Approval No. 202311-HY-1). All participants provided written informed consent, and the study was conducted in accordance with the principles of the Declaration of Helsinki.
Not applicable.
This work was supported in part by the National Key Technology R&D Program of China (2023YFC2506204), Natural Science Foundation of Shandong Province (ZR2024QH652), and Jining Key Research and Development Program (2022YXNS098).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/AP40685.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.



