1 Department of Animal Sciences, University of Illinois at Urbana-Champaign, Urbana, 61801 IL, USA
2 Illinois Informatics Institute, University of Illinois at Urbana-Champaign, Urbana, 61801 IL, USA
3 Neuroscience Program, University of Illinois at Urbana-Champaign, Urbana, 61801 IL, USA
4 Department of Chemistry, University of Illinois at Urbana-Champaign, Urbana, 61801 IL, USA
5 Department of Statistics, University of Illinois at Urbana-Champaign, Urbana, 61801 IL, USA
Abstract
The prolonged effects of maternal immune activation in response stressors
during gestation on the offspring’s molecular pathways after birth are beginning
to be understood. An association between maternal immune activation and neurodevelopmental and behavior
disorders such as autism and schizophrenia spectrum disorders has been detected
in long-term gene dysregulation. The incidence of alternative splicing among
neuropeptides and neuropeptide receptor genes, critical cell-cell signaling
molecules, associated with behavior may compromise the replicability of reported
maternal immune activation effects at the gene level. This study aims to advance the understanding of
the effect of maternal immune activation on transcript isoforms of the neuropeptide system (including
neuropeptide, receptor and connecting pathway genes) underlying behavior
disorders later in life. Recognizing the wide range of bioactive peptides and
functional receptors stemming from alternative splicing, we studied the effects
of maternal immune activation at the transcript isoform level on the hippocampus and amygdala of
three-week-old pigs exposed to maternal immune activation due to viral infection during gestation. In
the hippocampus and amygdala, 29 and 9 transcript isoforms, respectively, had maternal immune activation effects
(P-value
Keywords
- Maternal immune activation
- Neuropeptide
- Autism
- Alternative splicing
The activation of the immune system in the gestating female triggered by infection or other stressors elicit signals that can reach the developing fetus and can cause prolonged disruption of molecular pathways in the offspring’s brain [1, 2, 3, 4]. The relationship between maternal immune activation (MIA) and neurodevelopmental disorders, such as autism and schizophrenia spectrum disorders, and other neurodegenerative disorders such as Alzheimer’s disease, has been well documented [5, 6, 7].
Neuropeptides, hormones and neuropeptide receptors participate in biological processes underlying behavior and memory [8, 9, 10, 11, 12], and some neuropeptides such as insulin-like growth factor 1 (IGF1) have been associated with MIA-related disorders such as autism spectrum disorders [13, 14]. Our analysis of gene-level expression profiles in the amygdala of pigs exposed to viral infection during gestation exposed the effect of MIA on some members of the neuropeptide system, including the prohormone genes proenkephalin (PENK) and proopiomelanocortin (POMC), and receptor genes corticotropin-releasing hormone receptor 2 (CRHR2), parathyroid hormone 1 receptor (PTH1R), and vasoactive intestinal peptide receptor 2 (VIPR2) [15]. Despite the established role of neuropeptide and associated receptors on cell signaling and participation on pathways associated with social, feeding and aggression behavior, evidence gathered at the gene-level of expression has been limited. Bioactive neuropeptides result from complex transcription and translation processes [16, 17]. We hypothesize that MIA may have a varying effect on the products alternatively transcribed from neuropeptide genes. Therefore, analysis at the gene-level may hinder the exposition of MIA effects and may hinder the detection of antagonistic effect at the isoform level. Furthermore, the effect of MIA on the transcript isoform profiles may vary across brain regions involved in behaviors.
The amygdala and hippocampus brain regions are part of the limbic system and modulate behavior, emotion and memory. Amygdala and hippocampus processes have been associated with social behavior [18, 19], and disruption of the amygdala has been proposed to underlie autism spectrum disorders [20, 21]. Despite the established association between behavior disorders and MIA and between behavior disorders and neuropeptide systems, disruptions of the neuropeptide systems associated by MIA in the hippocampus and amygdala are only partially understood.
The objective of the present study is to advance the understanding of transcript isoform dysregulation in the neuropeptide system caused by MIA on two brain regions that regulate behavior. The characterization of the MIA effect at the transcript isoform level will complement reports from MIA studies of neurodevelopmental disorders at the gene level, and assist in the detection of potentially antagonistic effects of MIA on isoforms within a gene. A pig model of viral-elicited MIA was used to assess the impact of the immune response during gestation on the hippocampus and amygdala neuropeptide transcriptome. The supporting objective included the identification of MIA effects on the neuropeptide transcriptome that were brain region-dependent. Neuropeptide network reconstruction and visualization enabled us to uncover interrelationships between the neuropeptide genes impacted by MIA.
A study of the impact of MIA on the transcript isoforms was undertaken in recognition that neuropeptides and their receptors play an essential role in various physiological functions. Yet, the activity of these molecules depends on the transcribed isoform. Transcript isoform profiles from neuropeptide, corresponding receptors and genes connected through protein-protein interactions were studied in previously unpublished hippocampus samples and complemented with published amygdala samples [15] from pigs exposed to MIA relative to controls. This porcine model is suitable for MIA due to the many similarities to human physiology and the immune system [22]. The Porcine Reproductive and Respiratory Syndrome virus used to expose pigs to MIA belongs to the Arteriviridae family within the Nidovirales order, and this order includes the Coronaviridae family [23]. Viruses from these families and order have high incidence, causing colds and severe respiratory tract infections in human populations. These viruses can elicit immune activation in gestating females, and can cause significant financial cost to the swine industry due to reproductive failure and respiratory disease [24].
The experimental design and animal care protocols were approved by the
Institutional Animal Care and Use Committee (IACUC) at the University of Illinois
and were previously described [15, 25]. Briefly, pregnant gilts were intranasally
inoculated with the Porcine Reproductive and Respiratory Syndrome virus (5 mL of
1
For each brain region, RNA was isolated using EZNA isolation kit following the manufacturer’s instructions (Omega Biotek, Norcross, GA, United States). The hippocampus and amygdala RNA integrity was at least 7.5 and the RNA-Seq libraries were prepared with TruSeq Stranded mRNAseq Sample Prep kit’ (Illumina Inc, San Diego, CA). The individual pig libraries of the hippocampus (n = 48) and amygdala (n = 24) were sequenced in two lanes on a NovaSeq 6000 resulting in 150 nt long paired-end reads. Prior analysis of the amygdala expression patterns at the gene level identified differential abundance on a handful of neuropeptide and receptor genes, yet insufficient to support the enrichment of the Gene Ontology neuropeptide signaling pathways [15]. The present study broadens our previous gene-level analysis into the transcript isoform profiles in the amygdala and the hippocampus, another brain region that modulates behavior. We found that transcript isoforms provided more accurate gene networks than gene-level abundance [27].
The hippocampus and amygdala RNA-seq sequence reads were mapped to the Sus scofa genome v11.1 genome using Kallisto v0.43.0 with default settings [28]. Previously unpublished transcript isoform from neuropeptide and neuropeptide receptor genes from the hippocampus and amygdala were identified and analyzed. For each region, the count of each transcript was analyzed using a negative binomial generalized linear model of MIA, sex and the interaction between MIA and sex. The statistical analyses were conducted using SAS v9.4 (SAS Institute Inc., Cary, NC, United States). A preliminary analysis of the transcript isoforms corresponding to 185 neuropeptide and receptor genes demonstrated that less than 5% presented significant interaction effects, and therefore interaction effects are not reported. Of interest was the effect of MIA on the transcript isoform profiles, irrespectively of the sex; therefore, only the estimates of the effect of MIA, adjusted for sex effects, are presented.
Protein-protein and other molecular interaction databases were searched for genes known to connect neuropeptides and receptors that had transcript isoforms impacted by MIA in each brain region using the BisoGenet plugin [29] in the Cytoscape environment [30]. The visualization of the molecular relationships employed a maximum of two connections between neuropeptide-related genes based on the human gene and protein interactions identified by BisoGenet. Transcript isoforms of connecting pathway genes were analyzed for MIA effects using the same neuropeptide and receptor genes.
The hippocampus study yielded 207 transcript isoforms identified from 97
neuropeptide genes, 196 transcript isoforms from 87 neuropeptide receptor genes,
and 211 transcript isoforms from 41 connecting pathway genes. Significant MIA
effects (P-value
The amygdala study yielded 207 transcript isoforms identified from 97
neuropeptide genes, 196 transcript isoforms from 87 neuropeptide receptor genes,
and 178 transcript isoforms from 49 connecting pathway genes. Significant MIA
effect (P-value
Table 1 summarizes the number of transcript isoforms and genes differentially
expressed (P-value
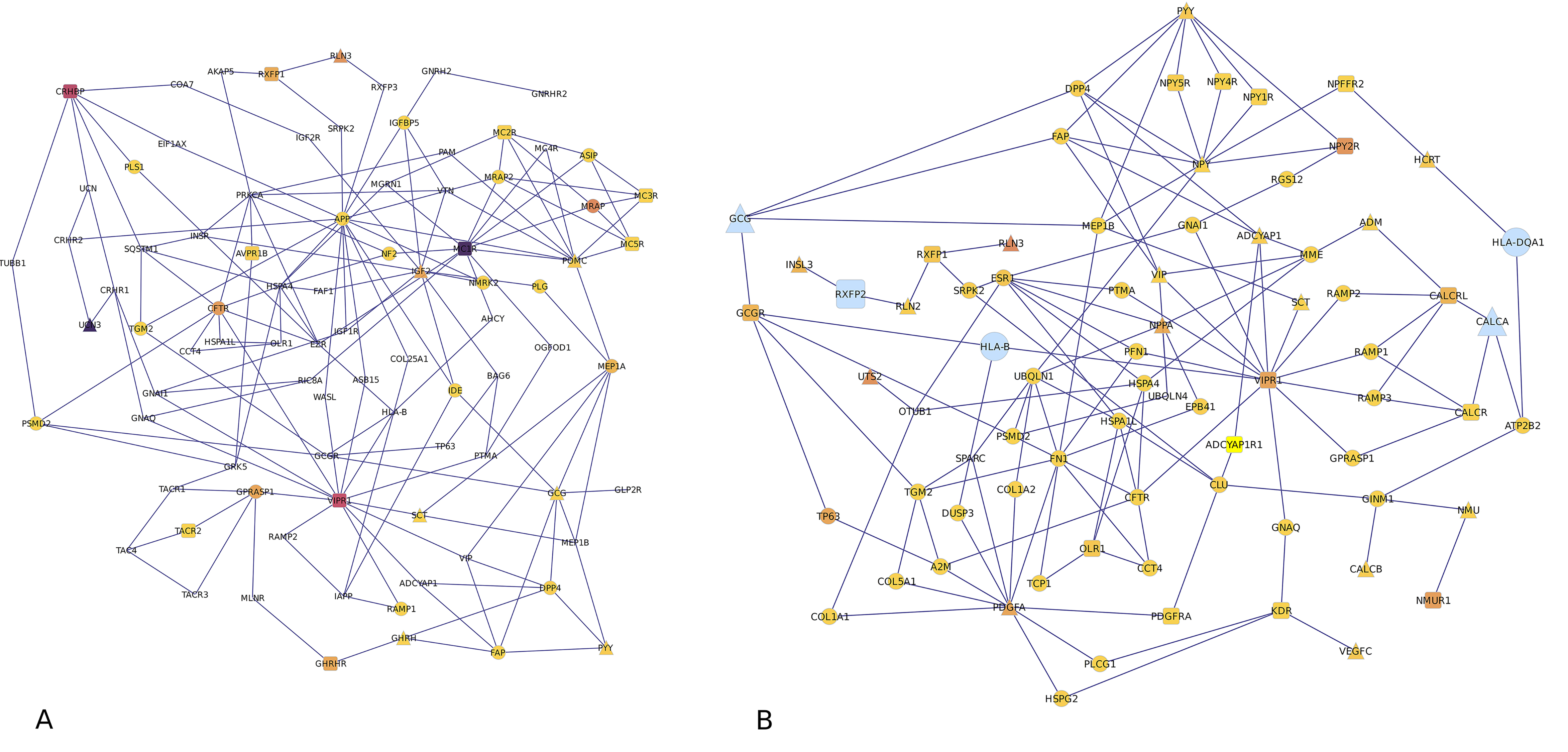 Fig. 1.
Fig. 1.Pathway of significantly differently expressed neuropeptide (triangle), receptor (square) and connecting (circle) genes in the A) amygdala and B) hippocampus. Gene symbol size represents the transcript isoform fold change between MIA and control pigs, and gene symbols with darker colors more significantly differential expression of transcript isoform than gene symbols with lighter colors.
| Brain Region | Transcript | Gene | ||||
| Neurop | Recep | Connect | Neurop | Recep | Connect | |
| Hippocampus | 11 | 8 | 8 | 10 | 8 | 7 |
| Amygdala | 2 | 3 | 2 | 2 | 3 | 2 |
| Overlap | 1 | 1 | 0 | 1 | 1 | 0 |
| Connect, number of transcript isoforms from connecting pathway transcripts or genes; Neurop, number of transcript isoforms from neuropeptide transcripts or genes; Recep, number of transcript isoforms from neuropeptide receptor transcripts or genes. | ||||||
The evaluation of the patterns of transcript isoform expression offered further
insights into the impact of MIA. Tables 2 and 3 summarize the transcript’s
hippocampus profiles isoforms over- and under-expressed in MIA relative to
control pigs (P-value
| Type | Symbol | Accession | Name | Diff | P-Value |
| Connect | EPB41 | XM_021095719.1 | erythrocyte membrane protein band 4.1, transcript variant X46 | 22.29 | 5.65E-03 |
| Connect | ESR1 | XM_021083031.1 | estrogen receptor 1, transcript variant X6 | 11.82 | 5.66E-06 |
| Connect | PLCG1 | XM_021078391.1 | phospholipase C gamma 1, transcript variant X4 | 1.69 | 5.53E-03 |
| Connect | PLCG1 | XM_021078394.1 | phospholipase C gamma 1, transcript variant X7 | 1.63 | 1.60E-03 |
| Connect | SLA-DQA1 | NM_001114062.2 | MHC class II histocompatibility antigen SLA-DQA | 6.39 | 7.97E-04 |
| Connect | SRPK2 | XM_021102581.1 | SRSF protein kinase 2, transcript variant X5 | 10.28 | 3.17E-03 |
| Connect | UBQLN1 | XM_021064659.1 | ubiquilin 1, transcript variant X2 | 1.28 | 3.40E-03 |
| Neurop | ADCYAP1 | XM_021093591.1 | adenylate cyclase activating polypeptide 1, transcript variant X2 | 3.17 | 2.33E-03 |
| Neurop | CALCB | NM_001102473.1 | calcitonin-related polypeptide beta | 11.60 | 7.23E-04 |
| Neurop | CRSP1 | XM_021080192.1 | calcitonin-related polypeptide beta, transcript variant X3 | 1.61 | 5.31E-03 |
| Neurop | IGF1 | XM_005664195.3 | insulin like growth factor 1, transcript variant X1 | 1.87 | 9.83E-03 |
| Neurop | IGF1 | XM_005664199.3 | insulin like growth factor 1, transcript variant X7 | 1.53 | 7.61E-03 |
| Neurop | RLN2 | XM_021082276.1 | relaxin 2, transcript variant X1 | 2.86 | 6.20E-03 |
| Neurop | SCG2 | NM_001012299.1 | secretogranin II | 1.62 | 6.19E-03 |
| Neurop | TRH | XM_005669803.3 | thyrotropin releasing hormone, transcript variant X2 | 3.05 | 6.72E-03 |
| Recep | GLP1R | NM_001256594.1 | glucagon like peptide 1 receptor | 4.75 | 5.17E-03 |
| Recep | MRGPRX2 | XM_013994496.2 | MAS related GPR family member X2 | 2.48 | 8.52E-03 |
| Recep | NPSR1 | XM_021079326.1 | neuropeptide S receptor 1 | 10.66 | 2.55E-04 |
| Recep | NPY5R | XR_002346654.1 | neuropeptide Y receptor Y5, transcript variant X2 | 1.75 | 7.10E-03 |
| Recep | OLR1 | NM_213805.1 | oxidized low density lipoprotein receptor 1 | 3.93 | 7.19E-05 |
| Recep | SLC40A1 | XM_013984335.2 | solute carrier family 40 member 1, transcript variant X2 | 1.69 | 1.29E-03 |
| Type: neuropeptide (Neurop), prohormone receptor (Recep) or Connecting pathway gene (Connect). Symbol, Accession, and Name: NCBI gene symbol, transcript accession number and transcript name; Diff: The expected difference between MIA and Control pigs; P-Value: the significance of the difference between Control and MIA pigs. | |||||
| Type | Symbol | Accession | Name | Diff | P-Value |
| Connect | HSPG2 | XM_021095484.1 | heparan sulfate proteoglycan 2, transcript variant X5 | 13.73 | 8.30E-03 |
| Neurop | AVP | NM_213952.2 | arginine vasopressin | 2.75 | 8.07E-03 |
| Neurop | HCRT | NM_214156.2 | hypocretin neuropeptide precursor | 2.97 | 3.39E-05 |
| Neurop | PYY | XM_021066091.1 | peptide YY, transcript variant X1 | 2.76 | 9.85E-05 |
| Neurop | RLN3 | XM_013994666.2 | relaxin 3, transcript variant X3 | 1.53 | 8.07E-04 |
| Recep | ADCYAP1R1 | XM_021078471.1 | ADCYAP receptor type I, transcript variant X3 | 32.07 | 2.72E-07 |
| Recep | GIPR | XM_021094579.1 | gastric inhibitory polypeptide receptor, transcript variant X2 | 3.02 | 1.89E-03 |
| Recep | NTSR2 | XM_021087826.1 | neurotensin receptor 2 | 1.32 | 1.35E-03 |
| Type: neuropeptide (Neurop), prohormone receptor (Recep) or Connecting pathway gene (Connect). Symbol, Accession, and Name: NCBI gene symbol, transcript accession number and transcript name; Diff: The expected difference between Control and MIA pigs; P-Value: the significance of the difference between Control and MIA pigs. | |||||
| Type | Symbol | Accession | Name | Diff | P-Value |
| Connect | DPP4 | NM_214257.1 | dipeptidyl peptidase 4 | 2.19 | 6.44E-03 |
| Connect | PAM | XM_021084567.1 | peptidylglycine alpha-amidating monooxygenase, transcript variant X17 | 0.14 | 9.23E-03 |
| Neurop | IAPP | XR_002344089.1 | islet amyloid polypeptide, transcript variant X2 | 0.58 | 5.91E-03 |
| Neurop | POMC | XM_021085834.1 | proopiomelanocortin, transcript variant X1 | 4.75 | 1.52E-04 |
| Neurop | PYY | XM_021066091.1 | peptide YY, transcript variant X1 | 3.27 | 5.82E-04 |
| Recep | AVPR1B | XM_003130445.3 | arginine vasopressin receptor 1B | 3.47 | 3.11E-03 |
| Recep | GHRHR | NM_214035.2 | growth hormone releasing hormone receptor | 4.29 | 4.07E-03 |
| Recep | GNRHR2 | NM_001001639.2 | gonadotropin-releasing hormone (type 2) receptor 2 | 0.54 | 4.69E-03 |
| Recep | OLR1 | NM_213805.1 | oxidized low density lipoprotein receptor 1 | 0.33 | 3.40E-03 |
| Type: neuropeptide (Neurop), prohormone receptor (Recep) or Connecting pathway gene (Connect). Symbol, Accession, and Name: NCBI gene symbol, transcript accession number and transcript name; Diff: The expected difference between Control and MIA pigs; P-Value: the significance of the difference between Control and MIA pigs. | |||||
In the hippocampus, 19 transcript isoforms were over-expressed in MIA relative
to control pigs at (Table 2), whereas 8 transcript isoforms were under-expressed
in MIA relative to control pigs at P-value
In general, differentially expressed transcript isoforms (P-value
The immune response of the mother to infection during gestation can impact the developing fetus and have long-lasting effects on the offspring after birth. The association between MIA and social behavior disorders such as autism spectrum disorders and schizophrenia spectrum disorders have been established in humans and biomedical models [31, 32, 33]. Pigs from gilts inoculated with the viral challenge of PRRSV during gestation exhibited lower sociability and locomotor activity compared to pigs from control gilts [25, 34, 35, 36]. The previous MIA effects are related to the susceptibility of developing brain regions that modulate behavior such as the hippocampus and amygdala. A gene expression study in the hippocampus of pig fetuses exposed to MIA detected increase the expression of three cytokines [37]. Our prior study of gene expression levels in the amygdala uncovered a few members of the neuropeptide system (POMC, VIPR2, PENK, PTH1R, CRHR2) that were affected by MIA [15]. The previous studies confirm that MIA can impact the expression levels of genes associated with behavior. However, the limited changes detected at the gene level and the multiple transcript isoforms that genes in the neuropeptide system can produce prompt us to investigate the impact of MIA at the transcript isoform level in the hippocampus and amygdala. Limitations in previous analyses have been addressed in the present study.
The present study validated our working hypothesis that a significant mode of action of MIA is through disruption of the levels of transcript isoform expression, which in turn influences the function and activity of molecules in the neuropeptide system. Our systematic study of over 200 transcript isoforms produced by genes in the neuropeptide system enabled us to expose various MIA effects within gene and across brain regions that are subsequently exemplified. Transcript isoforms were identified in the hippocampus and hypothalamus for most neuropeptide and receptor genes known in mammals [16, 17, 38, 39]. Transcript isoforms from neuropeptide Y receptor Y6 (NPY6R), insulin (INS) and urocortin 2 (UCN2) were identified in fewer than 5% of the pigs in both regions. Transcript isoforms from neuropeptides glucagon (GCG), urocortin 3 (UCN3), and urotensin 2B (UTS2B) were identified in the hippocampus of fewer than 5% of the pigs but were identified in a higher percentage of the amygdala samples. Similarly, transcript isoform from insulin-like 3 (INSL3), secretin receptor (SCTR) and urotensin 2 (UTS2) were identified in the amygdala of fewer than 5% of the pigs. Still, it was identified in a higher percentage of the hippocampus samples. These genes with low transcript isoform numbers also have very low or no expression in the human brain [40].
Our study at the transcript isoform level enabled the detection of strong MIA effects that could be obscured in analyses at the gene level when the transcript isoforms from the same gene are differentially affected by MIA. This is the case with two transcript isoforms of PYY that were differentially expressed in the same direction in both brain regions but with different significance levels. Neuropeptide PYY has been associated with feeding patterns, and the comorbidity of autism spectrum disorders is alterations in feeding behavior [41]. The detection of significant MIA effects on the patterns of isoforms from neuropeptide gene PYY offers fresh insights into these molecules’ role in MIA-associated disorders not previously established.
The original MIA-isoform associations detected in the present study can be corroborated by reports of gene associations with MIA comorbidities. PYY influences appetite, and the PYY transcript isoforms detected in the present study contain the expected active PYY peptide sequence that is conserved across vertebrates [42]. The predicted PYY transcript isoform sequence (XM_021066091.1) has a more extended reading frame than the curated sequence (NM_001256528.1), explaining the different magnitude of the MIA effect on the two transcript isoforms.
Another example of the innovative insights into the impact of MIA gained by the study of transcript isoform patterns is the receptor VIPR1 that had one transcript isoform over-expressed and another transcript isoform under-expressed in the amygdala of MIA relative to control pigs. The distinct impact of MIA across transcript isoforms may have hindered the ability of studies to detect the role of VIPR1 in MIA; however, our finding is endorsed by reports of significant differential expression of VIPR1 in the amygdala of rat lines selected for high or low ethanol consumption. This condition can disrupt inflammatory signals [43]. Similarly, transcript isoforms that presented opposite MIA profiles in the same region included GNRHR2 and ADCYAP receptor type I (ADCYAP1R1), relaxin/insulin-like family peptide receptor 1 (RXFP1) in the hippocampus and IGF1 in the amygdala.
Among the genes, including transcript isoforms, presented opposite MIA effects across brain regions G protein-coupled receptor 182 (GPR182) and connecting gene MHC class I antigen 2 (SLA-2). Although limited information is available on the role of the previous genes in MIA, our findings are corroborated by reports that GPR182 regulates the leukotriene biosynthesis pathway [44], and leukotriene are lipid mediators involved in immune response [45]. SLA-2 also participates in the immune response, and associations with MIA-associated autism have been published [46, 47].
Our investigation of transcript isoforms from immune-related genes uncovered the over-expression of SLA-2 and under-expression of GPR182 in the amygdala of MIA relative to control pigs. This finding suggests that immune activation during development can have a prolonged effect on the modulation of immune responses to stressors later in life. This result is supported by MIA pigs’ differential behavior relative to controls when injected with the immunostimulant Poly (I: C) at two months of age [25]. The opposite patterns of transcript isoforms in the previously reviewed genes reinforce the need to study the effect of inflammation signals during gestation on the offspring transcriptome later in life. The study at the gene level would cancel the distinct profiles and result in an apparent null effect of MIA on this receptor.
Transcript isoforms of several neuropeptides, neuropeptide receptors, and
connecting pathway genes in the neuropeptide system were differentially expressed
(P-value
Similarly, the differential expression of SLC40A1 is consistent with reports that MIA in rats disrupts the transport of iron through the materno-fetal interface, and the iron exporter ferroportin/SLC40A1 is essential for iron homeostasis. The resulting hypoferremia altered dopamine function in the adult offspring in rats and is associated with schizophrenia spectrum disorders [49]. These results demonstrate the added value of studying the impact of MIA in neuropeptide systems at the transcript isoform level.
The effects of MIA on the transcript isoforms produced by PLCG1, IGF1 and neuropeptide Y (NPY) were similar in the hippocampus and amygdala. Two PLCG1 transcript isoforms and two IGF1 transcript isoforms were over-expressed in the hippocampus of MIA relative to control pigs. Two NPY transcript isoforms in the amygdala and one of them detected in the hippocampus were under-expressed in MIA relative to control pigs. The detected effect of MIA on PLCG1 is consistent with reports that PLCG1 participates in the interactome of schizophrenia [50]. Genetic deletions of PLCG1 in mice exhibited behavioral alterations, including hypoactivity and reduced anxiety [51]. The NPY transcript isoform profile detected in this study is also consistent with reports that the expression of NPY (a member of the neuroactive ligand-receptor interaction pathway) was altered in the dorsolateral prefrontal cortex of individuals with schizophrenia [52]. Likewise, IGF1 was identified in association with the risk to develop neuropsychiatric and neurodevelopmental disorders due to maternal immune activation secondary to influenza infection [53]. Our results are first to elucidate the pattern of transcript isoforms associated with MIA. Our results confirm the need to study neuropeptide and receptor mRNA abundance at the transcript isoform level. Studies at the gene level fail to account for variation between transcript isoforms.
In addition to our transcript isoform-level discoveries, we confirmed reports of an association between numerous genes and MIA or MIA-related disorders. Many of the transcript isoforms presenting significant MIA effects are annotated to genes in Tables 2-4 that have differentially expressed genes between MIA and control pigs have been associated with neurological disorders. Some MIA associations with genes detected in the present study were also established at the DNA variant level. For example, single nucleotide polymorphism in or near ADCYAP receptor type I (ADCYAP1R1), NPS receptor (NPSR1), estrogen receptor 1 (ESR1), SRSF protein kinase 2 (SRPK2) genes have been associated with autism spectrum disorders, Alzheimer’s disease behavioral traits, anxiety, impulsivity and attention deficit-related traits [21, 54, 55, 56, 57, 58, 59].
Among the MIA effects bridging the hippocampus and amygdala, our results offer original evidence that MIA can disrupt the vasoactive intestinal peptide (VIP) and pituitary adenylate cyclase-activating peptide (PACAP). Significant effects of MIA were detected on transcript isoforms of the neuropeptide receptor ADCYAP1R1 and vasoactive intestinal peptide receptor 1 (VIPR1) receptors and the adenylate cyclase-activating polypeptide 1 (ADCYAP1) gene that produces PACAP27 and PACAP38 neuropeptides. However, no differential expression was detected with the VIP ligand and VIPR2. While the ADCYAP1R1, VPAC1, and VPAC2 interact with receptor-activity-modifying proteins (RAMPs) [60, 61], none of the RAMPs were differentially expressed. The VIP/PACAP signaling pathway is important in brain development, psychiatric illness and behavior [62, 63]. Altered behavior was observed in mouse lines knockout for ADCYAP1 and VIP prohormone genes [64, 65], and abnormal social behaviors have been recorded in ADCYAP1R1 deficient mice [66, 67]. ADCYAP1R1 is a member of the circadian entrainment pathway with multiple transcript isoforms presenting different ligand binding properties and signal transduction, therefore modulating the response to a vasoactive intestinal peptide (VIP) and PACAP [68, 69]. Comparison of the pig ADCYAP1R1 protein sequences identified that the different transcripts corresponded to the third intercellular loop that generate differential responses to PACAP-27 [70]. Compared to normal zebrafish (Danio renio), larvae lacking the hop exon had increased anxiety-like behavior [71, 72]. ADCYAP1 is a member of the circadian entrainment pathway and considered a potential target for therapies to ameliorate drug abuse, migraine, stress-related psychopathologies and inflammatory diseases [69, 73, 74, 75, 76, 77, 78]. In addition to the anti-inflammatory effect of VIPR1 [64], mutant mouse strains with inactivated VIPR1 showed a hippocampus-dependent deficit in fear conditions amygdala amygdala-dependent fear conditioning [79].
Peptidylglycine alpha-amidating monooxygenase (PAM) and ESR1 are genes connecting MIA-dysregulated neuropeptides and receptor transcript isoforms in the networks depicted in Fig. 1. PAM is responsible for the post-translational amidation of neuropeptides that can affect bioactivity, and this monooxygenase [80, 81] was over-expressed in MIA compared to control pigs in the hippocampus and amygdala. Our results are consistent with a report that PAM was over-expressed in the neocortex of 56-day old mice prenatally exposed to influenza [82]. ESR1 served as a connector between neuropeptides genes natriuretic peptide A (NPPA) and platelet-derived growth factor subunit A (PDGFA) and receptors RXFP1, VIPR1, and neuropeptide Y receptor Y2 (NPY2R) and was over-expressed in MIA relative to control pigs in the hippocampus and amygdala. Sequence variants of ESR1 have been linked to impaired social interaction [58, 83].
A review of the impact of MIA in the hippocampus indicates significant effects on the transcript isoforms from genes, including neuropeptide arginine vasopressin (AVP), calcitonins, hypocretin neuropeptide precursor (HCRT), IGF1, MAS related GPR family member X2 (MRGPRX2), PLCG1, relaxin 3 (RLN3), and SLC40A1. The detection of MIA associations with the previously enumerated transcript isoforms agrees with findings on behavior disorders, including reports that HCRT regulates sleep, feeding behavior, wakefulness, emotion, and stress response [41]. Similarly, RLN3 influences feeding behavior, stress responses, anxiety and memory [84], and AVP is a regulator of social behavior [85, 86]. IGF1 has been associated with autism spectrum disorder symptoms [87, 88, 89], with IGF1-treatment proposed as an autism therapeutic intervention [88].
The over-expression of SLC40A1 in the hippocampus of MIA relative to control pigs suggests that transcript isoform dysregulation may be a neuro-inflammatory response to the MIA signaling [89]. SLC40A1 has been implicated in the incidence of autism spectrum disorders [90]. MRGPRX2 is a receptor with multiple neuropeptide ligands, including cortistatin-14, and this receptor’s role was noted in a review of molecular mechanisms of autism spectrum disorder [91].
Whereas two calcitonin genes, calcitonin related polypeptide alpha (CALCA) and calcitonin related polypeptide beta (CALCB), have been uncovered in human and rodents, four calcitonin genes (CRSP1, CRSP2, CRSP3 and CALCB) are known in pigs [39]. The associated transcript isoforms can code for numerous neuropeptides. The distinct evolutionary path of the CALCA gene family between pigs and other biomedical species challenges the direct alignment of published associations. Nevertheless, an overarching profile can be identified between the over-expression of the calcitonin transcript isoforms in the hippocampus of MIA relative to control pigs and published work in other species and MIA-related phenotypes. The N-terminal procalcitonin (NPCT) neuropeptide, cleaved from calcitonin producing CALCA transcript isoform, participates in an inflammatory response and is a potential human biomarker and therapeutic target for Alzheimer disease [92]. Also, calcitonin gene-related peptide 1 (alpha-CGRP), a neuropeptide produced from a different CALCA transcript isoform that produces NPCT, plays an important role in social interactions with autism spectrum disorders [92, 93, 94].
Significant effects of MIA amygdala transcript isoforms were detected on AVPR1B, DPP4, GNRHR2, growth hormone-releasing hormone receptor (GHRHR), islet amyloid polypeptide (IAPP), OLR1, and POMC. Neuropeptide IAPP is associated with motivated behaviors [85]. Participants in the neuroactive ligand-receptor interaction pathway also detected an association with MIA at the gene level [15]. Similar to PYY, this neuropeptide has been linked to changes in feeding patterns that are also observed in autism spectrum disorder patients with the over-expression of IAPP transcript isoforms in the amygdala of MIA pigs, this protein aggregates in amyloid deposits and the detection of IAPP in the brain has been associated with cognitive decline. Also over-expressed in the amygdala of MIA relative to control pigs was PMCH. This result substantiates reports that dysregulation of PMCH is associated with the development of affective disorders [95].
Among other transcript isoforms dysregulated by MIA in the amygdala, the detection of POMC is in agreement with our analysis at the gene level [15] and with reports of pre- and post-translational bioactive forms associated with behaviors observed in autism spectrum disorders [96]. AVPR1B has been associated with autism spectrum disorders and is a receptor involved in the AVP-mediated activation of the hypothalamic-pituitary-adrenal-axis [97, 98, 99]. Consistent with this known function, and AVPR1B transcript isoform was over-expressed in the amygdala of MIA relative to control pigs. The impact of MIA on the expression of a GHRH transcript isoform could relate to the established influence of this hormone neuropeptide in animal cognition [100, 101].
OLR1 (also known as lectin-like oxidized low-density lipoprotein receptor-1, LOX-1) participates in inflammatory responses [102] and agrees with these results OLR1 transcript isoform was over-expressed in the amygdala of MIA relative to control pigs. In the present study, DPP4 is a connecting pathway gene with a transcript isoform under-expressed in the amygdala of MIA relative to control pigs. The observed pattern agrees with the role of DPP4 in processing the neuropeptides substance P and NPY and immune response [103] and the proposed association with autism spectrum disorders [104]. In agreement with the under-expression of the neuropeptide insulin-like growth factor 2 (IGF2) transcript isoform in the amygdala of MIA relative to control pigs, IGF2 treatment can reverse behavior disorders such as anxiety-like phenotypes [105]. IGF2 has also been associated with autism spectrum disorders and similar phenotypes [105, 106].
Bioactive neuropeptides result from multi-tiered pre- and post-transcriptional and translational processes that can influence the profile of neuropeptides across brain regions and should be considered in understanding the impact of MIA on neuropeptide systems. This study focused on exposing the effect of MIA at the post-transcriptional stage. However, pre-transcriptional effects such as epigenetics and post-transcriptional effects such as post-translational modifications can further alter the neuropeptide and receptor system’s profile and activity. Methylation can alter glucocorticoid receptors’ expression in a tissue-specific manner [107]. High-throughput or candidate proteomic detection of neuropeptides is complex. The small size and complex post-translational modifications and differential cleavage in different tissues result in some neuropeptides’ differential occurrence, such as glucagon [108]. Our results demonstrated the impact of MIA on multiple isoform profiles corresponding to genes in the neuropeptide system that can modulate behavior disorders.
The present study offered additional insights into the impact of MIA on neuropeptide systems by exposing the prolonged effects on transcript isoforms in the hippocampus and amygdala. The analysis of transcript isoforms both uncovered transcription dysregulation that may have been hindered by the analysis of the system at the gene level, and aided in the corroboration of other gene-level findings. The present study detected multiple neuropeptides, neuropeptide receptors. It connected gene transcript isoforms that were significantly differentially expressed in response to the maternal signals elicited in response to infection during gestation. We confirmed that the present MIA model elicited by viral infection during gestation in pigs assisted in the identification of genes that have known associations with neurodevelopmental disorders, including schizophrenia spectrum disorders, autism spectrum disorders, and Alzheimer’s disease. This study exposed differences in MIA effect on transcript isoforms from the same gene within and across brain regions. This finding suggests that the identification of effective targets to ameliorate MIA and neurodevelopmental disorders necessitate discrimination among transcript isoforms. Our results advance the understanding of proposed neuropeptide-based antipsychotic and antidepressant therapies [109, 110] to ameliorate behavior disorders induced by MIA.
S.R-Z. and R.J. conceived and designed the experiments; M.K., H.R. and S.R-Z. performed the experiment and collected the data; B.S., P.Z. and K.K. analyzed the data; B.S., J.S. and S.R-Z interpreted the results. B.S. and S.R-Z. wrote the paper.
The Illinois Institutional Animal Care and Use Committee (IACUC) at the University of Illinois approved the experimental design and animal care protocols. The experiments are in compliance with the USDA Animal Welfare Act and the NIH Public Health Service Policy on the Humane Care Use of Animals.
We thank the anonymous reviewers for their valuable review of the article.
This study was supported by USDA NIFA AFRI grant number 2018-67015-27413 and by the NIH National Institute on Drug Abuse award number P30 DA018310.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://jin.imrpress.com/EN/10.31083/j.jin.2021.01.332.

