1 State Key Laboratory of Wheat Improvement, Shandong Agricultural University, 271018 Tai’an, Shandong, China
Abstract
Pattern-triggered immunity (PTI), the first line of defense in plants, is activated when host pattern recognition receptors (PRRs) detect pathogen-associated molecular patterns (PAMPs) or damage-associated molecular patterns (DAMPs) released during microbial invasion. Over the past decade, major advances have clarified PTI’s complex signaling network, revealing its role as a dynamic regulator of plant immune responses. This review summarizes recent progress in PTI signal transduction, emphasizing the core mechanisms: (1) kinase cascades, including MAPKs and calcium-dependent protein kinases (CDPKs), that amplify defense signals; (2) calcium ion (Ca2+) dynamics acting as rapid secondary messengers; and (3) reactive oxygen species (ROS) bursts that mediate antimicrobial and signaling. The roles of lipid signaling, crosstalk between immune signaling and plant hormones, and scaffold proteins are also discussed. Furthermore, we explore the interplay between PTI and effector-triggered immunity (ETI), highlighting synergistic modules, such as the enhanced disease susceptibility 1–phytoalexin deficient 4–activated disease resistance 1 (EDS1-PAD4-ADR1) which integrate immune signals to enhance pathogen resistance. By linking molecular mechanisms to systemic immune regulation, this review provides a comprehensive framework for understanding plant defense strategies and identifies emerging opportunities to engineer disease-resistant crops through modulation of immune pathways.
Keywords
- plant immunity
- pattern recognition receptors
- signal transduction
- protein kinases
- host-pathogen interactions
- plant disease resistance
Plant pathogens employ various strategies to invade their hosts, including direct penetration of the cuticle, exploitation of natural openings (e.g., stomata, hydathodes, lenticels, and styles), or entry through wounds caused by pests, mechanical injury, frost, or natural fissures. Some pathogens develop specialized structures, such as haustoria, to breach plant cells and extract nutrients, ultimately leading to plant death. To counter these threats, plants have evolved multilayered defense mechanisms, with their immune system serving as a cornerstone of pathogen resistance [1]. Current research shows that plant immunity operates through a dual surveillance system [2], comprising two distinct branches: (1) pattern-triggered immunity (PTI), initiated by cell surface-localized pattern recognition receptors (PRRs) that detect conserved pathogen-associated molecular patterns (PAMPs); and (2) effector-triggered immunity (ETI), mediated by intracellular nucleotide-binding leucine-rich repeat (NB-LRR) proteins encoded by resistance genes (R genes). These NB-LRR proteins, characterized by nucleotide-binding (NB) and leucine-rich repeat (LRR) domains, recognize pathogen effectors and activate robust immune responses [1, 2, 3].
The plant immune system is described by the “zigzag model”, which outlines the relationship between PTI and ETI and the iterative signaling between plants and pathogens. PTI provides basal resistance, while ETI strengthens defenses against effectors that suppress immunity. Pathogen evasion of PTI triggers stronger ETI responses, escalating defense mechanisms. This model is widely used to analyze the complexity of immune signaling [1].
Upon detecting pathogens, plants activate multifaceted defensive arsenal to combat invasion. These strategies include: (1) Abscisic acid (ABA) regulates the closure of stomata to limit pathogen entry. Recent studies have shown that ABA enhances stomatal defense by promoting the local biosynthesis of salicylic acid (SA). Moreover, the ABA-mediated regulatory process may involve crosstalk between multiple hormones, which requires further investigation [4, 5, 6]; (2) the production and secretion of antimicrobial compounds to inhibit pathogen proliferation [7], and (3) the activation of mitogen-activated protein kinase (MAPK) signaling cascades, which coordinate ion flux and reactive oxygen species (ROS) production to directly impair pathogens [8]. These mechanisms collectively enhance plant resilience, illustrating the sophistication of their immune responses. Navarro et al. [9] demonstrated significant transcriptional overlap between PTI and ETI induced genes, indicating some shared regulatory nodes in defense signal transduction between PTI and ETI [8].
Recent transformative discoveries in PTI signaling lack a cohesive framework. This review synthesizes the molecular architecture of PTI, focusing on Arabidopsis thaliana, and outlines key signaling pathways, receptor complexes, and downstream responses. It also explores the interplay between PTI and ETI, providing new insights into their synergistic roles in layered plant defense (Fig. 1).
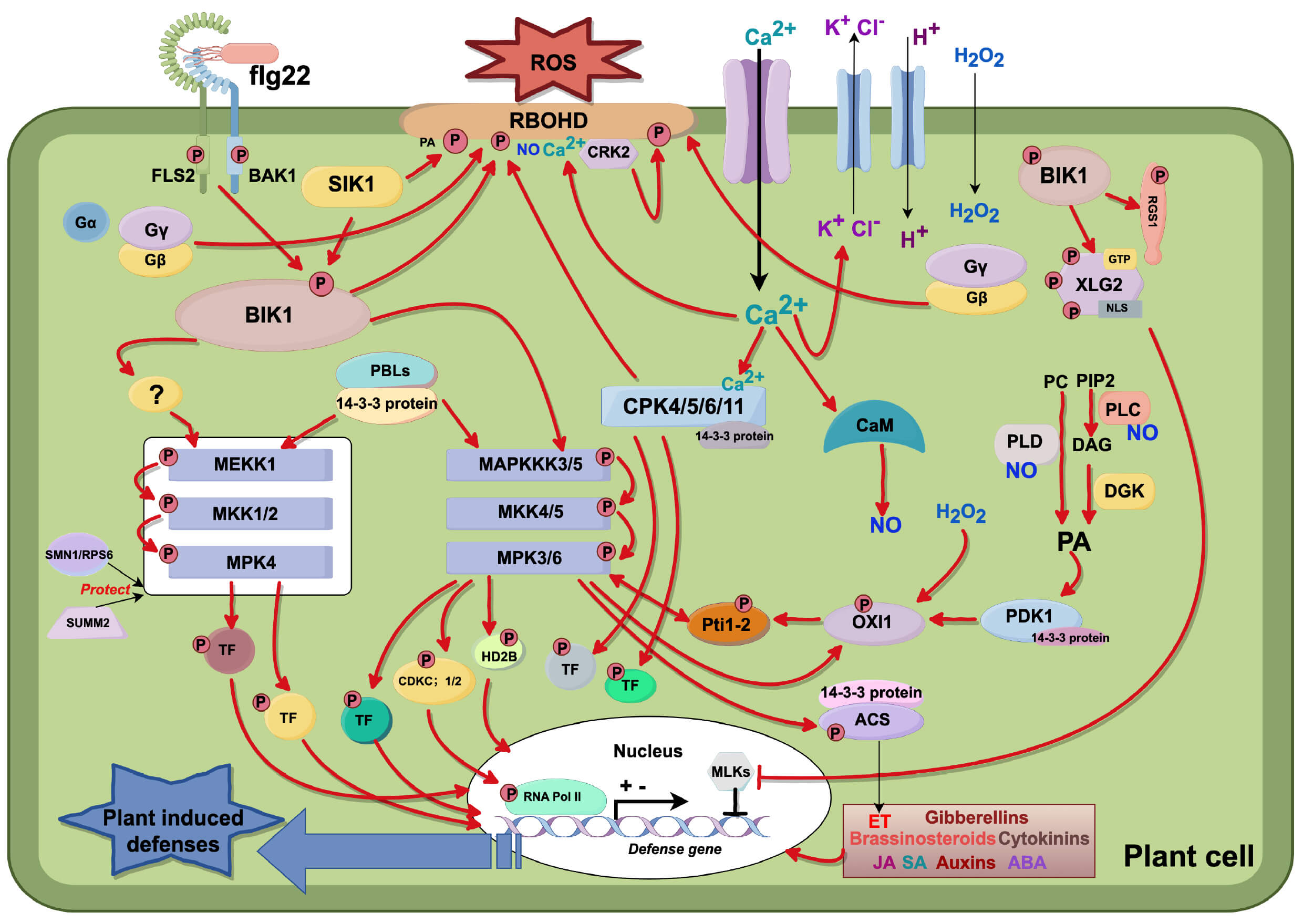 Fig. 1.
Fig. 1.
PTI signal transduction mechanism. (1) Initial Perception and
Early Signaling: Recognition of flg22 by the flagellin-sensing 2–BRI1-associated
receptor kinase 1 (FLS2–BAK1) complex triggers receptor transphosphorylation,
releasing phosphorylated botrytis-induced kinase 1 (BIK1) and inducing Ca2+
influx. This activates membrane ion fluxes (K+/Cl– efflux, H+
influx), resulting in membrane depolarization and apoplastic alkalization. (2)
ROS Production: Ca2+ activates respiratory burst oxidase homolog D (RBOHD)
via BIK1 and calcium-dependent protein kinases (CDPKs). RBOHD activity is
modulated by phosphatidic acid (PA), Ca2+, nitric oxide (NO), and kinases
(serine/threonine-protein kinase I1 (SKI1)/cysteine-rich receptor-like kinase 2
(CRK2)), generating extracellular hydrogen peroxide (H2O2). Calmodulin
(CaM) stimulates NO synthesis, while phospholipase C (PLC)/phospholipase D
(PLD)/diacylglycerol kinase (DGK) pathways produce PA, reinforcing ROS/NO
signaling loops. (3) Downstream Kinase Activation: PA activates
3-phosphoinositide-dependent protein kinase 1 (PDK1), which phosphorylates
oxidative signal-inducible 1 (OXI1). H2O2-activated OXI1 phosphorylates
Pto-interacting protein 1–2 (Pti1-2) to modulate mitogen-activated protein
kinases (MAPKs). MAP kinase 6 (MPK6) phosphorylates OXI1/Pti1-2, forming feedback
loops. (4) Heterotrimeric G Proteins: Activation of the FLS2–BAK1 complex
dissociates G
Substantial advances have elucidated the role of PAMPs (also known as Microbe-Associated Molecular Patterns (MAMPs)) in triggering plant immunity. These microbial molecules activate conserved defense pathways, enabling plants to detect and respond to potential invasions. For instance, flg22, a conserved 22-amino acid N-terminal fragment of bacterial flagellin, is a key inducer of plant immunity [1, 10]. In Arabidopsis, flg22 is recognized by the leucine-rich repeat (LRR) receptor kinase Flagellin-Sensing 2 (FLS2), leading to the rapid transcriptional activation of over 1100 defense-related genes [1, 11, 12]. This response includes the upregulation of receptors, signaling kinases, and antimicrobial compounds. Chitin, a major component of fungal cell walls, is as a critical fungal PAMP. In Arabidopsis, the receptor chitin elicitor receptor kinase 1 (CERK1) recognizes chitin, triggering downstream responses such as mitogen-activated protein kinase (MAPK) activation, ROS bursts, and defense gene expression [13]. Additionally, the chitin elicitor-binding protein (CEBiP) in rice senses chitin, initiating immune signaling and enhancing resistance to fungal pathogens [14]. Many proteins secreted by pathogens also act as PAMPs, and induce plant immune responses, including cell wall-degrading enzymes (CWDEs), harpins, and glycoprotein elicitors [15].
Damage-associated molecular patterns (DAMPs) are endogenous danger signals released by plants during pathogen attack. They are host-derived molecules released or exposed upon cellular damage caused by pathogen enzymes (e.g., cellulases and pectinases) or physical disruption, such as cutin monomers, cell wall fragments (e.g., oligogalacturonides and cellulose-derived oligomers), and endogenous peptides (e.g., systemin and plant elicitor peptides). These signals amplify immune responses and activate defense pathways that overlap with PTI [16].
PRRs are plasma membrane-localized proteins that serve as the frontline sensors of plant immunity. They include receptor-like kinases (RLKs) and receptor-like proteins (RLPs) with extracellular domains. They contain various ligand-binding extracellular domains (e.g., leucine-rich repeats (LRRs) and lysin motifs (LysM)) that recognize MAMPs/DAMPs to initiate immune responses upon ligand binding [17, 18]. The first PRR discovered in both plants and animals is the Xa21 protein, which confers resistance in rice against Xanthomonas oryzae [19]. Subsequently, FLS2 (the bacterial flagellin receptor) and EFR (the bacterial elongation factor Tu receptor) were identified in Arabidopsis [20].
Several PRR-MAMP pairs have been identified, including the PRRs FLS2 and EFR, which recognize the MAMPs flg22 and elf18, respectively [20, 21, 22]. PRRs assemble into immune receptor complexes with other proteins, which are essential for MAMP sensing and downstream signal transduction [23]. For example, in Arabidopsis, recognition of flg22 by FLS2 induces the rapid formation of such a complex. This triggers mutual transphosphorylation between FLS2 and its co-receptor BAK1, ultimately leading to the dissociation of the kinase BIK1 from the FLS2-BAK1 complex [24].
With ongoing research, an increasing number of PRRs and PRR–MAMP pairs have been identified. These topics have been comprehensively reviewed by DeFalco and Zipfel [25], and are therefore not discussed in this paper.
In Arabidopsis, approximately 41% of expressed proteins undergo phosphorylation under various conditions. The process is dynamically regulated by protein kinases and phosphatases, and predominantly occurs on serine, threonine, and tyrosine residues [26]. Phosphorylation determines the functional properties of proteins [27], and serves as a key regulatory mechanism for signaling pathways activated by external stimuli, such as environmental stressors, as well as internal cues like plant hormones [28, 29]. In the PTI immune pathway, MAMP recognition triggers a series of phosphorylation cascades. For example, Rayapuram et al. [30] identified several precisely mapped and highly reliable phosphopeptide sites associated with immune signaling. Their work also showed decreased phosphorylation levels of specific phosphopeptides following MAMP treatment, suggesting that certain protein phosphatases play important roles in PTI signal transduction.
MAMP recognition rapidly activates multiple protein kinases, including mitogen-activated protein kinases (MAPKs), calcium-dependent protein kinases (CDPKs), and AGC protein kinases. These kinases play crucial roles in regulating target proteins to coordinate immune responses [30]. CDPKs will be discussed in detail in Section 4.
In Arabidopsis, AGC kinases—a group of serine/threonine protein kinases, named after the protein kinase A, G, and C families—play crucial roles in various biological processes. The Arabidopsis genome encodes 39 AGC kinases, which are classified into five major sub-families: AGCVI, AGCVII, AGCVIII, AGC other, and PDK1 homologs [31]. Among these, the AGCVIII subfamily is the largest, consisting of 23 kinases that are further divided into four groups: ACG1, ACG2, ACG3, and ACG4 [32, 33]. Notably, plant AGC kinases have been implicated in immune responses and the regulation of pathogen-induced MAPK cascades [34].
OXI1 (AGC2-1), an AGC protein kinase, is activated by H2O2 and cellulase in Arabidopsis. The activation of MAPK3 and MAPK6, which is critical for mediating signaling responses to H2O2 and flg22 [35, 36], depends on OXI1. Wild-type plants exhibit robust MAPK3/6 activation upon H2O2 or cellulase treatment; however, this activation is significantly reduced in OXI1mutants, underscoring OXI1’s essential role in amplifying MAPK3/6 signaling [37]. Furthermore, OXI1 expression and kinase activity are rapidly upregulated in Arabidopsis roots in response to PAMPs, cellulase, and H2O2. Additionally, OXI1 governs resistance against the pathogens Hyaloperonospora arabidopsidis and Pseudomonas syringae, highlighting its broader role in plant immunity [37, 38, 39].
In Arabidopsis, OXI1 transcriptional induction is significantly reduced in AtRBOHD mutants or following treatment with the NADPH oxidase inhibitor diphenyleneiodonium (DPI) [39]. RBOHD generates ROS in response to PAMPs and biotrophic pathogens, demonstrating that ROS produced during plant-pathogen interactions mediate OXI1 upregulation [40, 41].
OXI1 modulates the activity of a subset of protein kinases, including PTI 1-1, PTI 1-2, PTI 1-3, and PTI 1-4, which are structurally related to the tomato PTI kinase involved in hypersensitive responses [42, 43]. RNA interference experiments show that PTI 1-2 activation in Arabidopsis depends on OXI1 during exogenous PA, xylanase-mediated PA, and H2O2 activation, whereas PDK1 is specifically essential for PA-dependent signaling. Additionally, PTI 1-2 enhances ROS promoter expression, linking it to redox signaling pathways [42].
In vitro studies demonstrate that OXI1 phosphorylates PTI 1-4, a kinase that interacts with MPK3 and MPK6. In turn, MPK3 and MPK4 phosphorylate both OXI1 and PTI 1-4, forming a dynamic signaling network. Within this network, OXI1-mediated phosphorylation of PTI 1-4 enables its interaction with MPK6, thereby regulating the cascade. MPK6 phosphorylates both Pti1-4 and OXI1, establishing a regulatory feedback loop [43]. Notably, OXI1 activity is further modulated by PDK1, a regulatory mechanism common among AGC kinases. Phosphatidic acid (PA) activates PDK1, which subsequently phosphorylates OXI1, suggesting that additional upstream kinases may contribute to OXI1 regulation [42].
Recent study has revealed that OXI1 and its closest homolog, AGC subfamily 2 member 2 (AGC2-2; which shares 75% amino acid sequence identity with OXI1), play key roles in the secretion of camalexin and help Arabidopsis resistance against Pseudomonas syringae and Botrytis cinerea [44]. These kinases can phosphorylate three pleiotropic drug resistance (PDR) transporters in vitro: PDR6, PDR8, and PDR12. Notably, physical interactions between PDR6 and both OXI1 and AGC2-2 have been confirmed. Direct phosphorylation of PDR6 by these kinases controls its efflux activity, thereby enhancing pathogen resistance in Arabidopsis. However, it remains unclear whether OXI1 and AGC2-2 directly modulate the function of PDR8 and PDR12, or influence the distribution of other secondary metabolites. Further investigation is needed to clarify these potential regulatory mechanisms [44].
The Arabidopsis genome encodes 20 MAPKs, 10 MAPK kinases (MAPKKs), and 60 MAPKK kinases (MAPKKKs), forming a tiered signaling network [45]. The MAPK cascade, an evolutionarily conserved module in eukaryotes, translates extracellular stimuli into intracellular responses via phosphorylation cascades [45]. This cascade operates through hierarchical mechanisms [46, 47, 48]: (1) Receptor activation: Plasma membrane receptors perceive stimuli, triggering the activation of MAPKKKs. (2) MAPKKK-MAPKK signaling: Activated MAPKKKs phosphorylate and activate downstream MAPKKs. (3) MAPKK-MAPK activation: MAPKKs (dual-specificity kinases) phosphorylate MAPKs at conserved threonine and tyrosine residues within their activation loop, enabling full kinase activity. (4) Effector regulation: Activated MAPKs (serine/threonine kinases) phosphorylate diverse cytoplasmic and nuclear targets, including transcription factors, enzymes, and other kinases, to orchestrate cellular responses [48].
In Arabidopsis, PRRs detect PAMPs and activate downstream MAPK cascades that control immune responses. Two primary MAPK pathways are involved, including the MEKK1-MKK1/2-MPK4 cascade [49, 50] and the MAPKKK3/5-MKK4/MKK5-MPK3/6 cascade [51, 52]. Activation of these cascades triggers defense responses, including ROS bursts, stomatal closure, and transcriptional activation of immune-related genes. PRRs, positioned upstream of MAPK signaling, are key regulators of this process. For example, the Arabidopsis receptor FLS2 recognizes the bacterial flagellin-derived peptide flg22 and forms a complex with the co-receptor BAK1, which then activates downstream MAPK cascades to start plant immune responses [53, 54].
In the MAPKKK3/5-MKK4/5-MPK3/6 pathway, MAPKKK3 and MAPKKK5 regulate the activation of the MKK4/5-MPK3/6 module, which strengthens plant resistance to bacterial and fungal pathogens [51, 52, 55, 56]. Studies reveal that in Arabidopsis MAPKKK3/MAPKKK5 double mutants, flg22-induced MPK3/6 activation is reduced to ~45%, while activation by chitin, elf18, and pep2 is reduced to ~10–20%. This shows that MAPKKK3/5 are critical regulators of immune responses to microbial patterns (e.g., flg22, chitin, and elf18) and plant-derived signals (e.g., pep2). However, the remaining MPK3/6 activity in mutants suggests other MAPKKKs may also contribute to this pathway. Notably, single MAPKKK3 or MAPKKK5 mutations only partially reduce MPK3/6 activation, indicating overlapping functions between MAPKKK3 and MAPKKK5 in PTI [51, 52].
The MEKK1-MKK1/2-MPK4 cascade is another well-studied MAPK pathway [49, 50]. Initially, this pathway was proposed to negatively regulate defense responses, as its disruption leads to constant autoimmune activity, including spontaneous cell death and overactivation of defense genes [57, 58]. However, recent studies show that this module also plays a positive role in defense responses. SUMM2, a NLR protein, monitors the integrity of the MEKK1-MKK1/2-MPK4 cascade. Disruption of the cascade (e.g., in MEKK1 or MKK1/MKK2 mutants) activates SUMM2-dependent autoimmunity, masking the module’s normal role in basal defense. In MEKK1 SUMM2 or MKK1 MKK2 SUMM2 double mutants, where SUMM2-mediated autoimmunity is removed, plants become more susceptible to pathogens. This shows that the MEKK1-MKK1/2-MPK4 cascade actively promotes basal immunity [58]. Thus, this cascade not only fine-tunes immune responses but is also protected by SUMM2, which monitors its function and triggers backup defenses if the pathway is disrupted. Additionally, the SMN1 gene encodes RPS6, a toll/interleukin-1 receptor (TIR)-type NLR protein. Together with SUMM2, SMN1/RPS6 forms a dual surveillance system that monitors the integrity of the MEKK1-MKK1/MKK2-MPK4 cascade [59, 60].
MAPKs play a crucial role in plant immunity, especially in transcriptional regulation. Study shows that many MAPK targets are transcription factors, highlighting their importance in immune signaling [16]. Specifically, MPK3, MPK4, and MPK6 directly phosphorylate and activate the WRKY transcription factor 33 (WRKY33) both in vitro and in vivo. MPK3/6 regulate immune-related genes, including ACS2 and ACS6 (involved in ethylene biosynthesis) and camalexin biosynthesis genes, via WRKY33 phosphorylation [61, 62, 63]. Notably, mutations in WRKY33’s MPK3/6 phosphorylation sites stop it from activating camalexin biosynthesis genes, confirming that WRKY33 is a target of MPK3/6. Phosphorylation enhances WRKY33’s transcriptional activity, allowing it to bind its own promoter and directly upregulate camalexin biosynthesis genes [62, 64]. Additionally, MPK3/6 are needed for the accumulation of pipecolic acid (PIP), a precursor of N-hydroxypipecolic acid (NHP), which is an important component of systemic acquired resistance (SAR). PIP biosynthesis depends entirely on AGD2-like defense response protein 1 (ALD1) [65, 66]. WRKY33 directly regulates ALD1 transcription, increasing PIP synthesis [67]. Thus, MPK3/6 likely promote PIP production by regulating ALD1 through WRKY33, linking local immune signaling to systemic defense.
WRKY33 also interacts with MPK4, an important MAP kinase in Arabidopsis immune signaling. When MPK4 is activated by Pseudomonas syringae infection or the microbial peptide flg22, it phosphorylates MKS1 (MAP kinase substrate 1), a scaffold protein bound to WRKY33. This phosphorylation causes the MKS1-WRKY33 complex to dissociate from MPK4, allowing WRKY33 to translocate to the nucleus. There, WRKY33 directly binds to the promoter of PAD3 (phytoalexin deficient 3), activating its transcription and promoting the production of camalexin, an antimicrobial compound that is critical for defense against pathogens [68, 69].
MPK3 further interacts with the transcription factor VIP1 (virus-interacting protein 1), showing its broad role in transcriptional regulation. When flg22 is detected, MPK3 phosphorylates VIP1, causing it to move into the nucleus and activate the expression of PR1 (pathogenesis-related 1) [70]. PR1 is a common marker gene of plant disease resistance, and its activation indicates that MPK3 directly controls VIP1 localization and function, amplifing immune responses by activating defense-related genes [70].
MPK4 also targets the immune suppressor ASR3 (altered seed germination 3), a plant-specific trihelix transcription factor, adding another layer of immune regulation [71]. The Study shows that MPK4 quickly phosphorylates ASR3 at Thr189 post-flg22 perception. Phosphorylated ASR3 enhances its DNA-binding ability, suppressing the expression of many flg22-responsive genes. This negative regulation is consistent with the enhanced disease resistance observed in ASR3 mutants when infected with lethal bacterial pathogens. In contrast, overexpression of either wild-type or phosphomimetic ASR3 in transgenic plants impairs PTI responses, showing that ASR3 dampens immune signaling [71].
The ethylene response factor (ERF) family, which has over 120 members in Arabidopsis, is one of the largest groups of plant transcription factors involved in stress responses [72]. A key part of immune signaling is the interaction between ERF104 and the mitogen-activated protein kinase MPK6. MPK6 directly phosphorylates ERF104, causing it to separate from the protein complex. Once released, phosphorylated ERF104 activates expression of defense-related genes. Notably, the ERF104-MPK6 interaction is quickly modulated upon pathogen perception, and the ERF104/MPK6 complex dissociates within 5–15 minutes after flg22 treatment. This process depends on MPK6 kinase activity, ethylene production and signaling pathways. Transcriptomic analysis reveals that the overexpression of ERF104 upregulates multiple stress-responsive genes, indicating that MPK6-mediated phosphorylation is essential for ERF104’s role in regulating immunity [73].
Additionally, MPK3 and MPK6 phosphorylate ERF6, enhancing its protein stability [74]. In experiments of Botrytis cinerea infection, phosphomimetic ERF6 (e.g., T178D variant) strongly activates defensin genes (PDF1.1 and PDF1.2), leading to enhanced resistance to Botrytis cinerea. This activation happens independently of ethylene signaling, showing that ERF6 functions differently from typical ethylene-dependent defense pathways. These findings show that ERF6 acts as a phosphorylation-dependent effector in the MPK3/6 signaling cascade, directly linking kinase activity to antifungal defense gene expression [74].
In Arabidopsis, PAMP-triggered activation of MAPK cascades leads to phosphorylation of BES1, a key transcription factor in brassinosteroid (BR) signaling. Specifically, MPK6 directly interacts with and phosphorylates BES1, which is essential for effective immune responses [75]. BES1 loss-of-function mutants exhibit reduced resistance to the bacterial pathogen Pst DC3000, indicating that BES1 is required for plant immunity. A phospho-deficient BES1 variant (S286A/S137A) fails to undergo PAMP-induced phosphorylation and does not restore pathogen resistance in BES1 mutants [75]. These findings show that MPK6-mediated phosphorylation of BES1 at specific sites (S286/S137) is critical for coupling BR signaling to immune activation, providing a molecular connection between growth and defense.
In addition to transcription factors, MAPKs also interact with other substrates regulating biotic stress, growth, and development. We focus on those that are critical to immune signaling and pathogen resistance.
The interaction between VQ motif-containing proteins (VQPs) and MAPK signaling in Arabidopsis highlights the complex regulatory mechanisms that control plant immune responses. VQPs are a family of proteins containing the VQ motif, which function in diverse physiological processes in plants. Stress-activated MAPKs, including MPK3, MPK4, MPK6, and MPK11, interact with VQPs to modulate immune signaling pathways [76, 77]. MPK3/6-targeted VQPs (MVQs) are a subgroup of VQPs that are regulated by MAPK phosphorylation. For example, MVQ1 suppresses gene expression mediated by WRKY transcription factors, such as the NHL10 promoter, when PAMPs are detected [78]. MPK6 phosphorylates MVQ1, which changes its stability and promotes its degradation. Notably, non-phosphorylated MVQ1 exhibits greater stability than its phosphorylated form. This suggests that after PAMP infection, MAPK-mediated phosphorylation accelerates MVQ1 degradation, thereby alleviating its suppression of WRKY-dependent immune responses and reactivating defense-related gene transcription. This mechanism shows an important regulatory pattern: MAPKs modulate the stability of VQPs, potentially changing the composition of WRKY-VQP transcriptional regulatory complexes and fine-tuning defense gene expression [78]. Thus, plant immune signaling is regulated not only by the activating or suppressing specific transcription factors, but also by controlling their stability and interaction networks, enabling coordinated immune responses.
Pathogen infection triggers ethylene production in plants, which plays an important role in immunity. When plants are attacked, pathogens induce the expression of ACS genes, such as ACS6 in Arabidopsis, which encode key enzymes in ethylene biosynthesis [79, 80]. Early study revealed that MPK6-mediated phosphorylation stabilizes ACS2 and ACS6, increasing their levels and enhancing ACS activity in cells [81]. In liquid-cultured seedlings, B. cinerea-induced ethylene biosynthesis is significantly reduced in MPK3/MPK6 double mutants but not in single mutants, indicating that MPK3 also phosphorylates ACS2 and ACS6 [82]. Li et al. [83] showed that WRKY33 directly activates ACS2 and ACS6 downstream of the MPK3/MPK6 signaling cascade during pathogen invasion. Therefore, MPK3 and MPK6 regulate ACS2 and ACS6 by phosphorylation and by promoting their gene expression, providing dual-layer regulation of ethylene synthesis during immune responses.
The AT-hook motif nuclear localized (AHL) gene family is evolutionarily conserved across land plants, and Arabidopsis has 29 members (AHL1–AHL29). AHL proteins play roles in plant growth, development, and differentiation. For example, AHL27 and AHL29 affect petiole growth [84], AHL22 influences flowering [85], and AHL18 shapes root architecture [86]. Chromatin phosphoproteomic studies [87] have identified AHL13—a nuclear DNA-binding protein containing an AT-hook motif—as a substrate of MAPK signaling. Although AHL13 mainly interacts with MPK6 in vivo, it can be phosphorylated in vitro by MPK3, MPK4, and MPK6. AHL13 knockout mutants exhibit impaired PAMP-triggered immunity, including reduced ROS burst, attenuated defense gene induction, and compromised PTI responses.
Phosphorylation of AHL13 at S109 and S376 is critical for its stability [87]. Since MPK3, MPK4, MPK6, and AHL13 are expressed throughout plant tissues, and AHL13 phosphorylation at these sites is widespread, this suggests that MAPK-mediated regulation of AHL13 stability is a conserved mechanism important for plant defense responses [88].
In eukaryotes, RNA polymerase II (RNA Pol II) is essential for protein-coding gene transcription. RNA Pol II contains a C-terminal domain (CTD) with heptad repeats (Tyr1-Ser2-Pro3-Thr4-Ser5-Pro6-Ser7). Phosphorylation of the CTD by various kinases regulates transcription initiation, elongation, and termination [89]. Notably, PAMP perception triggers rapid but transient phosphorylation of serine residues in these CTD repeats. In Arabidopsis, activation of the flg22-induced MAPK cascade leads to MPK3-mediated phosphorylation of CDKC;1 and CDKC;2, which enhances their kinase activity on the CTD. This process is balanced by CPL3-mediated CTD dephosphorylation [90]. Together, this MAPK cascade-CDKC-RNA Pol II CTD phosphorylation module amplifies RNA Pol II activity and upregulates defense-related gene expression during immune responses to combat pathogens.
Arabidopsis encodes 18 histone deacetylases (HDACs) [91, 92], categorized into four groups. Group IV, including HD2A-D and HD2B, is specific to plants [91]. Among these, HD2B is phosphorylated by the mitogen-activated protein kinase MPK3 at residues T249 and S266, both in vitro and in vivo. MPK4 and MPK6 can also phosphorylate HD2B less efficiently. Functional studies reveal that HD2B knockout mutants are more susceptible to both pathogenic and non-pathogenic strains of Pseudomonas syringae, whereas overexpression of HD2B enhances plant resistance [93]. Thus, MPK3 directly targets HD2B, which regulates numerous defense genes, highlighting the important role of chromatin remodeling through HDAC activity and kinase signaling in plant immune responses.
The ubiquitination cascade, involving E1 ubiquitin-activating enzymes, E2 ubiquitin-conjugating enzymes, and E3 ubiquitin-ligases enzymes, mediates the tagging of target proteins with ubiquitin. U-box-type E3 ligases, in particular, play important roles in plant immunity [94, 95]. In Arabidopsis, the U-box ligases PUB22, PUB23, and PUB24 negatively regulate PAMP-triggered immunity [96] and reduce PAMP-induced signaling [97]. Notably, PUB22 undergoes self-ubiquitination and proteasomal degradation when exposed to the bacterial PAMP flg22. However, MPK3, a key immune signaling kinase, specifically interacts with and phosphorylates PUB22, which disrupts its oligomerization, blocks self-ubiquitination, and stabilizes the ligase. This MPK3-mediated stabilization enhances PUB22’s ability to suppress immune responses, creating a feedback loop that fine-tunes immune signaling [98].
MPK3/MPK6 interact with LIP5, a protein associated with multivesicular bodies (MVB) [99, 100]. Pathogen-activated MPK3/6 signaling phosphorylates and stabilizes LIP5, leading to its accumulation during infection and promoting pathogen-responsive vesicle trafficking. Although LIP5 mutants show normal flg22-triggered defenses, they are more susceptible to pseudomonas syringae due to defective MVB formation [100]. Thus, MPK3/6 stabilize LIP5 to activate the MVB pathway, thereby enhancing plant immunity.
Recognition of fungal or bacterial PAMPs triggers a rapid increase in cytosolic Ca2+ levels, a hallmark of early plant immune signaling [101, 102, 103, 104, 105, 106]. Studies using transgenic aequorin plants and BIK1 mutant show that flg22 induces rapid and strong luminescence, indicating Ca2+ influx in wild-type plants. In contrast, BIK1 mutants exhibit only 50% of this response, highlighting BIK1’s critical role in amplifying Ca2+ signals during immune activation [107]. Mechanistically, BIK1 phosphorylates and activates the calmodulin (CaM)-gated channels CNGC2 and CNGC4 upon flg22 perception, thereby increasing cytosolic Ca2+ concentrations. This phosphorylation-dependent activation directly links BIK1 kinase activity to Ca2+ mobilization, underscoring its essential role in initiating Ca2+-dependent defense responses [105, 108].
Moreover, CDPKs act as central calcium sensors, regulating numerous Ca2+-stimulated protein kinase activities to coordinate responses to both internal signals and environmental cues [107, 109]. For example, CDPKs are rapidly activated upon detection of biotrophic pathogen invasion or herbivore attack, triggering localized defense mechanisms [107]. Genetic studies underscore the critical role of CDPKs in plant immunity, as many are essential for effective defense responses. Their activity increases dynamically following pathogen infection, further highlighting their importance in immune signal transduction [110].
The structure of CDPKs is well defined. They have an N-terminal Ser/Thr kinase domain responsible for autophosphorylation and a C-terminal EF-hand domain that binds Ca2+ to activate the kinase. Additionally, CDPKs can also autophosphorylate Tyr, classifying them as dual-specificity kinases with both Ser/Thr and Tyr kinase activity [104, 111, 112]. In Arabidopsis, there are 34 distinct CDPK isoforms, showing their functional diversity [109]. For example, fungal infection rapidly upregulates AtCDPK1 expression. Functional analyses demonstrate that AtCDPK1 loss-of-function mutants are more susceptible to pathogens compared to wild-type plants, confirming its critical role in innate immunity [113].
Upon flg22 perception, CPK4, CPK5, CPK6, and CPK11 are rapidly activated, displaying distinct molecular weights and activation kinetics compared to tobacco CDPKs, which remain persistently active during the Avr9-Cf9 immune interaction [114, 115, 116]. CDPKs also regulate ROS bursts, which are critical for plant immunity [110]. For example, ectopic expression of AtCPK1 in tomato protoplasts [117] or NtCDPK2 in Nicotiana benthamiana [118] induces ROS bursts. Genetic evidence in Arabidopsis underscores their role: flg22-induced oxidative bursts are significantly reduced in cpk5/cpk6 double mutants, cpk5/cpk6/cpk11 triple mutants, and cpk4/cpk5/cpk6/cpk11 quadruple mutants, demonstrating their collective contributions to ROS generation. This mechanism may involve direct phosphorylation of RBOHD, highlighting the role of CDPKs in the local activation of RBOHD [110, 114].
The receptor-like cytoplasmic kinase (RLCK) PBL1, encoded by the CCE5 gene, plays a pivotal role in calcium signaling during plant immunity. CCE5 mutants exhibit reduced cytosolic Ca2+ elevation in response to PAMPs [105, 119]. This Ca2+ influx activates membrane transporters, driving H+ influx along with Cl–, K+, and NO3– efflux. These ion fluxes cause membrane depolarization and extracellular alkalization within one minute of flg22 or elf18 treatment, marking early immune signaling events [106]. Cytosolic Ca2+ homeostasis is tightly regulated by channels, such as Ca2+-permeable glutamate receptor-like (GLR) channels [120]. Arabidopsis mutants lacking GLR2.7, GLR2.8, and GLR2.9 show a ~25% reduction in flg22, elf18, and Pep1 induced Ca2+ influx and impaired resistance to Pseudomonas syringae, indicating that GLRs contribute to PAMP-induced Ca2+ influx during PTI [121]. In contrast, cyclic nucleotide-gated channel CNGC2 mutants lack Ca2+ elevation in response to endogenous danger signals (e.g., AT PEP3 and bacterial lipopolysaccharides (LPS)), but maintain flg22-induced responses, highlighting pathway-specific roles for Ca2+ channels in immune signaling [122].
Plasma membrane-localized Ca2+-ATPases, such as ACA8 and ACA10 [123], and vacuolar membrane-localized ACA4 and ACA11 [124], play essential roles in regulating Ca2+ influx. While some of these transporters are well understood, many ion channels mediating Ca2+ signaling in immunity remain unexplored. Interestingly, inhibition of serine/threonine kinases and MAPKKs reduces cytosolic and chloroplast stromal Ca2+ oscillations, suggesting that MAPK cascades may influence Ca2+ signaling [125]. During PTI, Ca2+ sensors decode cytosolic Ca2+ fluctuations and interact with MAPKs to orchestrate defense gene activation. For example, the Ca2+ responsive transcription factor CAMTA3 binds to target gene promoters to suppress defense responses [126]. However, MPK3 and MPK6 phosphorylate CAMTA3, leading to its degradation. This MAPK-mediated elimination of CAMTA3 lifts its suppression of defense pathways, enhancing plant resistance [127].
ROS, such as superoxide (O2–) and hydrogen peroxide (H2O2), are rapidly generated in the apoplast following pathogen recognition, marking an early event in PTI [128]. These ROS bursts typically occur within 2–3 minutes of PAMP detection, and peak around 10–15 minutes [129]. Plasma membrane NADPH oxidases and cell wall-associated peroxidases both contribute to ROS bursts. NADPH oxidases transfer electrons from cytosolic NADPH/NADH to apoplastic oxygen to generate O2–, which is then converted to H2O2 by superoxide dismutase. Notably, ROS, particularly H2O2, enhance immune signaling by promoting Ca2+ influx, thereby elevating cytosolic Ca2+ levels [128]. PAMP or DAMP induced ROS bursts raise cytosolic Ca2+ levels, indicating a feedback loop between ROS and Ca2+ signaling. These ROS bursts are critical for the second peak or sustained plateau of Ca2+ concentrations observed during immune activation [103].
Arabidopsis has ten respiratory burst oxidase homolog (RBOH) genes, containing FAD/NADPH binding domains, six transmembrane regions, and a functional oxidase domain. Among these, RBOHD—localized to the plasma membrane—mediates PAMP-triggered ROS bursts [103, 130, 131]. Upon PAMP detection, the receptor kinases EFR and FLS2 form a complex, and BIK1 directly interacts with RBOHD, and phosphorylates its N-terminus at Ca2+-independent sites to initiate ROS production [108, 132, 133]. This early phosphorylation is distinct from regulation by calcium-dependent protein kinase (CDPK). Phosphorylated RBOHD is essential for PAMP-induced ROS bursts. The EF-hand motifs in RBOHD’s N-terminus are critical for regulation, as demonstrated by EF-hand mutation analysis [134]. Ca2+ influx induces conformational changes in these motifs, enabling phosphorylation by CDPKs such as CPK4, CPK5, CPK6, and CPK11, which amplify ROS bursts in Arabidopsis [114, 134]. Notably, CPK5 phosphorylates RBOHD at Ser148, Ser163, and Ser347, enhancing ROS production and immune signal propagation [135].
Regulation of ROS bursts in Arabidopsis involves both activation and suppression. For positive regulation, SIK1, a MAP4K in Arabidopsis, has dual functions [136]. It phosphorylates BIK1 to stabilize it, enhancing its role in immune signaling, and also directly senses the bacterial PAMP flg22 to promote phosphorylation of RBOHD’s N-terminal region, boosting RBOHD activity and ROS production [136]. CRK2 pre-forms a complex with RBOHD and phosphorylates its C-terminal S703 residue during pathogen attack, further regulating PAMP-induced ROS generation [137]. Conversely, some protein kinases act as negative regulators. For example, CPK28 phosphorylates BIK1, triggers its degradation, and indirectly suppresses ROS bursts [138, 139]. CIPK14 is another negative regulator; over expression of CIPK14 suppresses ROS, while CIPK14 mutants display hyperactive ROS responses upon PAMP detection [140]. PBL13 directly binds to RBOHD’s C-terminus and phosphorylates it at S862 and T912. The E3 ubiquitin ligase PIRE interacts with PBL13 and RBOHD, promoting RBOHD ubiquitination and reducing its activity, thereby decreasing plant disease resistance [141]. Additionally, the Arabidopsis translation initiation factor binding protein eIF4E1 (CBE1) may negatively regulate RBOHD accumulation at the translational level, impairing ROS-dependent immune response [142].
The role of MAPKs in PAMP-induced ROS bursts remains unclear, and key
experimental findings suggests these pathways operate independently. In
Arabidopsis, atrbohD mutants show similar MPK3/MPK6 activation
to wild-type plants upon flg22 treatment, indicating that ROS production does not
depend on MAPK signaling [143, 144]. It was found that ROS production was not
reduced in MPK3 or MPK6 mutants, although MPK3 mutants had a delayed response,
suggesting that MAPKs may fine tune ROS dynamics but not drive them [103].
Furthermore, overexpression of the phosphatase AP2C1, which dephosphorylates and
inactivates MPK3/MPK6, does not suppress flg22-mediated ROS production, further
supporting the independence of these pathways [145]. In anp2/anp3 double
mutants or
Nitric oxide (NO) plays a significant role in plant-pathogen interactions, enhancing plant immunity through multiple mechanisms [149]. Plants rapidly produce NO upon recognition of PAMPs or DAMPs during pathogen invasion [150, 151]. NO works with ROS together to amplify both local and systemic defense responses [152]. In Arabidopsis, NPR1 and TGA1 are redox-regulated signals that interact in their reduced state, enhancing TGA1’s DNA-binding ability and activating pathogenesis-related PR gene expression [153]. NO regulates the NPR1/TGA1 signaling network, further boosting plant immune defenses [154]. PAMP-induced Ca2+ signaling is also linked to NO production, though the molecular mechanisms remain unclear [155]. For example, the cation channel CNGC2 is required for lipopolysaccharide (LPS)-triggered NO synthesis, while CNGC2 mutants suppress LPS-induced NO production in Arabidopsis [156]. The Ca2+-binding protein CML24, activated by pathogen-induced Ca2+ influx, mediates downstream NO production, bridging Ca2+ and NO signaling pathways [157].
Phosphatidic acid (PA), a key intermediate in lipid biosynthesis, integrates environmental cues into plant immune responses [158]. PA and its associated enzymes—phospholipase C (PLC), phospholipase D (PLD), and diacylglycerol kinase (DGK)—play important roles in plant immune defense [159]. PA influences immune outputs by modulating ROS bursts and CDPK activity [16, 160]. In tomato cells, PA levels increase in response to xylanase, flg22, and fungal elicitors, along with elevated levels of its precursor, diacylglycerol pyrophosphate (DGPP), and enhanced PI-PLC and PLD activity. This indicates that PLC and PLD are required for PA accumulation and subsequent immune responses [161, 162]. In Arabidopsis, DGK phosphorylates diacylglycerol (DAG) to produce PA, and inhibitors of DGK block flg22-induced PA production and ROS bursts, highlighting DGK’s positive role in plant immunity [163]. PA also enhances 3-phosphoinositide-dependent kinase 1 (PDK1) activity, further promoting immune defense responses [42]. Notably, PAMP-triggered PA synthesis via PLD and PLC/DGK pathways partially depends on NO, suggesting cross-talk between lipid and redox signaling pathways [164]. Thus, NO and PA operate within interconnected networks to enhance plant immunity. NO influences Ca2+-mediated PA synthesis, while PA enhances ROS generation and kinase cascades that, in turn, promote NO production. This reciprocal regulation ensures robust and coordinated defense responses against pathogens.
The heterotrimeric G protein system in Arabidopsis consists of a single
canonical G
G
The current model proposed by Pandey describes that the dynamic G protein cycle
is central to Arabidopsis immune signaling, and outlines its structured
sequence from quiescence to activation [170]. In the resting state, the
heterotrimeric G protein complex—including either the canonical G
Arabidopsis heterotrimeric G proteins coordinate immune responses
through both cytoplasmic and nuclear pathways. Upon flg22 detection, BIK1
phosphorylates XLG2, causing it dissociate from G
14-3-3 proteins are key regulators of plant immunity, acting as dynamic scaffolds to mediate protein-protein interactions and modulate the activity, stability, or localization of their target proteins. In Arabidopsis, the 14-3-3 gene family includes 15 members, which are typically activated by interacting with phosphorylated proteins to amplify and coordinate signaling pathways [177, 178]. 14-3-3 proteins interact with various immune-related molecules, such as CDPKs, PA, PDK1, and 1-Aminocyclopropane-1-carboxylate synthase (ACS) [16].
In plant defense responses, 14-3-3 proteins interact with multiple members of the Receptor-like Cytoplasmic Kinases (RLCK) subfamily VII, such as PBL19, PBL20, and PBL37, as well as subfamily XII members PBL40 and BSK1. These RLCKs regulate plant resistance by phosphorylating downstream targets like MAPKKK5 or MPK15 [179]. Additionally, 14-3-3 proteins interact with PBL19, PBL20, and BIK1 within PRR receptor complexes, supporting rapid signal propagation [179, 180].
14-3-3 proteins relieve kinase autoinhibition to promote signaling. In MAPKKK5, autoinhibitory interactions between its N- and C- terminal regions block PBL19 access. 14-3-3 proteins (GRF6/8) bind PBL19 and MAPKKK5’s C-terminus, allowing PBL19 to phosphorylate and relieve autoinhibition. 14-3-3 proteins also scaffold the interactions between PBL19 and MEKK1, activating the MAPK cascade. Therefore, 14-3-3 proteins accelerate immune responses, such as ROS bursts and defense gene expression [180].
A summary of the key kinases, their substrates, and regulatory roles in PTI signaling is provided in Table 1 (Ref. [1, 16, 24, 37, 39, 42, 43, 49, 51, 52, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 73, 74, 75, 90, 91, 98, 99, 100, 105, 108, 110, 114, 119, 133, 136, 137, 139, 140, 141, 146, 170, 179, 180]).
| Kinase | Type/family | Substrates/interacting proteins | Functions | Reference |
| FLS2 | LRR-RLK (PRR) | flg22; BAK1 | Perceives flg22; recruits BAK1 to form active complex and initiate PTI signaling | [1, 24] |
| BAK1 | SERK-RLK | FLS2; BIK1 | Co-receptor with FLS2; phosphorylates BIK1 and mediates FLS2 ubiquitination | [24] |
| BIK1 | RLCK VII | RBOHD; RGS1; CNGC2/4 | Phosphorylates RBOHD to trigger ROS burst; phosphorylates RGS1 to activate G-proteins; activates CNGC channels | [105, 108, 133, 170] |
| SIK1 (MAP4K) | MAP4K | BIK1; RBOHD | Phosphorylates and stabilizes BIK1; directly phosphorylates RBOHD to enhance ROS burst | [136] |
| CRK2 | CRK-RLK | RBOHD; BIK1 | Directly phosphorylates RBOHD; amplifying ROS signaling | [137] |
| CDPKs (CPK4/5/6/11) | CDPK | RBOHD | Ca2+-dependent phosphorylation of RBOHD; enhancing early ROS burst | [110, 114] |
| CPK28 | CDPK | BIK1 | Phosphorylates BIK1 to promote its ubiquitination and degradation; forming negative feedback on ROS | [139] |
| CIPK14 | CIPK | - | Negatively regulates PTI signaling, suppressing ROS production | [140] |
| PBL13 | RLCK | RBOHD (S862, T912) | Negatively phosphorylates RBOHD; limiting ROS levels | [141] |
| PBL1 | RLCK | - | Contribute PAMP-induced Ca2+ influx, regulating early calcium signaling | [119] |
| PDK1 | AGC-PDK1 | OXI1 | Phosphorylates and activates OXI1 in response to phosphatidic acid | [42] |
| OXI1 | AGC | PTI1-2/4; MPK3/6 | Phosphorylates PTI1 kinases and contributes to the activation of MPK3/6 in response to H2O2 or PA, thereby forming positive feedback loop that amplifies ROS production and cell death | [37, 39, 42, 43] |
| PTI1-2/ PTI1-4 | RLCK VII | - | Substrate of OXI1; involved in ROS burst and cell death regulation | [42, 43] |
| PBL19 | RLCK VII | MAPKKK5 | Phosphorylates and activates MAPKKK5; initiating MAPK cascade | [179, 180] |
| MEKK1 | MAPKKK | MKK1/2; SUMM2 | Phosphorylates MKK1/2 to activate MPK4; induces backup immunity via SUMM2 upon cascade disruption | [16, 49, 58] |
| MAPKKK3/5 | MAPKKK | MKK4/5 | Phosphorylates MKK4/5 to activate MPK3/6 in response to multiple PAMPs | [51, 52] |
| MAPKKK5 | MAPKKK | MAPKK4/5; 14-3-3 proteins | Forms complex with PBL19 and 14-3-3 proteins to relieve autoinhibition and activate MAPKK4/5 | [51, 180] |
| MKK1/2 | MAPKK | MPK4; SUMM2 | Phosphorylates and activates MPK4; cascade monitored by SUMM2 for stability | [49, 58] |
| MAPKK4/5 | MAPKK | MPK3/6 | Phosphorylates MPK3/6 to regulate defense gene expression via multiple transcription factors and chromatin modifiers | [51, 52] |
| MPK3/6 | MAPK | WRKY33; VIP1; ERF104; ERF6; HD2B; BES1; PUB22; RNA Pol II CTD; LIP5 | Phosphorylates multiple TFs and nuclear proteins to drive defense gene expression, chromatin remodeling, stabilize PUB22 and enhance Pol II activity, stabilize LIP5 to activate the MVB pathway, enhancing immunity | [61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 73, 74, 75, 90, 91, 98, 99, 100] |
| MPK4 | MAPK | MKS1; ASR3; RPS6/SMN1 | Phosphorylates MKS1 to release WRKY33; phosphorylates ASR3 to negatively regulate defense; monitored by RPS6/SMN1 | [57, 59, 60, 71] |
| ANP1/2/3 | MAPKKK | - | It required for PAMP-induced ROS burst, functioning in a pathway partially independent of MPK3/6 | [146] |
AGC, AGC kinase family; ANP1, Arabidopsis NPK1-related protein kinase 1; ASR3, altered seed germination 3; BAK1, BRI1-associated receptor kinase 1; BES1, BRI1-EMS-suppressor 1; BIK1, botrytis-induced kinase 1; PBL1, PBS1-like protein 1; CDPK, calcium-dependent protein kinase; CIPK, calcineurin B-like interacting protein kinase; CIPK14, CBL-interacting protein kinase 14; CNGC2, cyclic nucleotide-gated channel 2; CPK28, calcium-dependent protein kinase 28; CRK-RLK, cysteine-rich receptor-like kinase; CTD, C-terminal domain; ERF104, ethylene response factor 104; ERF6, ethylene response factor 6; flg22, flagellin 22 peptide; FLS2, Flagellin-Sensing 2; HD2B, histone deacetylase 2B; LIP5, LYST-interacting protein 5; LRR-RLK, leucine-rich repeat receptor-like kinase; MAP4K, mitogen-activated protein kinase kinase kinase kinase; MAPK, mitogen-activated protein kinase; MAPKK, mitogen-activated protein kinase kinase; MAPKKK, mitogen-activated protein kinase kinase kinase; MEKK1, MAPK/ERK kinase kinase 1; MKK1, MAPK kinase 1; MKS1, MAPK substrate 1; MPK3, mitogen-activated protein kinase 3; MVB, multivesicular body; OXI1, oxidative signal-inducible 1; PA, phosphatidic acid; PAMP, pathogen-associated molecular pattern; PBL13, PBS1-like protein 13; PDK1, 3-phosphoinositide-dependent protein kinase 1; PRR, pattern recognition receptor; PTI, pattern-triggered immunity; PTI1-2, PTO-interacting 1-like kinase 2; PUB22, Plant U-Box 22; RBOHD, respiratory burst oxidase homolog D; RGS1, regulator of G-protein signaling 1; RLCK, receptor-like cytoplasmic kinase; RNA Pol II, RNA polymerase II; ROS, reactive oxygen species; RPS6, ribosomal protein S6; SMN1, suppressor of MPK4 nuclear immunity 1; SERK-RLK, somatic embryogenesis receptor kinase–receptor-like kinase; SIK1, stress-induced kinase 1; SUMM2, suppressor of MKK1 MKK2 2; TF, transcription factor; VIP1, VirE2-interacting protein 1; WRKY33, WRKY transcription factor 33.
Salicylic acid (SA), ethylene (ET), and jasmonic acid (JA) are key hormones involved in plant defense, playing distinct roles against different types of pathogens [181]. These hormones are rapidly produced upon PAMP perception and work together with PTI to enhance defense responses. SA mainly defends against biotrophic or hemibiotrophic pathogens, while JA and ET are critical for combating necrotrophic pathogens [181, 182]. Besides these three hormones, other auxiliary hormones, such as brassinosteroids, auxins, abscisic acid (ABA), cytokinins, and gibberellins, also modulate immune outcomes, forming a complex regulatory network in plant immunity.
These hormones do not regulate plant immunity independently, but form dynamic networks that fine-tune immune responses through both antagonistic and synergistic interactions [16]. They transmit different signals and perform diverse defense functions. For example, in SA-JA interplay, SA signaling often suppresses JA pathways. The SA receptor NPR1 inhibits JA-mediated defenses, although their relationship also includes synergistic effects [183]. NPR1, the SA receptor acts as a key regulator of SA signaling and mediates its antagonistic crosstalk with the JA pathway [184, 185]. Conversely, the JA receptor COI1 represses SA signaling, establishing a bidirectional regulatory loop [186]. Interestingly, the SA receptors NPR3 and NPR4 have opposite functions to NPR1, and they activate JA signaling during ETI to induce combined SA and JA defenses that enhance plant immunity [187, 188, 189].
In SA-ABA interactions, the two signaling pathways are generally antagonistic, and ABA suppresses SA signaling, weakening plant immunity against certain pathogens [190]. For example, ABA suppresses SA accumulation, impairing resistance to pathogens such as Pseudomonas syringae pv. tomato [191]. However, SA and ABA also have synergistic effects. For instance, SA and ABA work together to enhance RNA silencing in viral defense, as shown in Arabidopsis resistance to Bamboo mosaic virus (BaMV) [192].
SA-ET exhibit mutual antagonism, with each hormone’s biosynthesis pathways inhibiting the other. SA signaling via NPR1 inhibits ET responses by interacting with the ethylene response factor EIN3, blocking ET-dependent gene transcription [193]. Conversely, EIN3 and EIL1 suppress SA biosynthesis by binding to the SID2 promoter, increasing susceptibility to Pseudomonas syringae [194].
In summary, the complex interactions among plant hormones play crucial roles in plant immune responses. However, further research is needed to fully understand how these hormones precisely regulate plant immunity through their mutual interactions. Future studies using systems biology and single-cell approaches will unravel these complexities and provide new strategies for developing disease-resistant crop.
The classical zigzag model posits that ETI is an amplified version of PTI, forming a stepwise defense hierarchy [1]. However, recent emerging evidence reveals a more complex interplay, where PTI and ETI engage in bidirectional crosstalk, either enhancing or suppressing each other to fine-tune immune responses (Fig. 2).
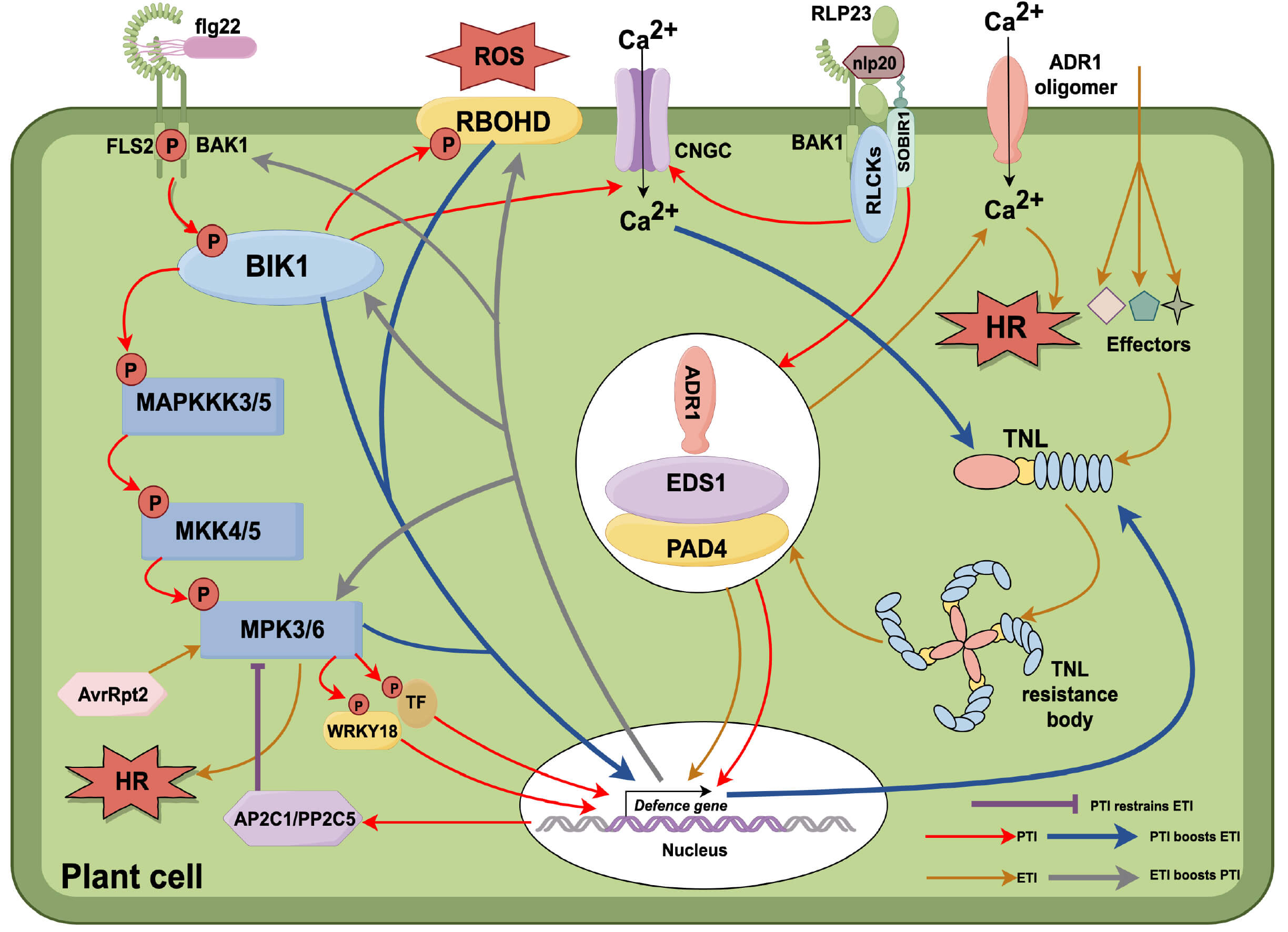 Fig. 2.
Fig. 2.
PTI–ETI interplay in plant immunity. (1) Recognition and activation. Receptor-like kinases (RLKs) and receptor-like proteins (RLPs) detect pathogen-/microbe-associated molecular patterns (PAMPs/MAMPs) to activate pattern-triggered immunity (PTI). Upon recognition of flg22 (flagellin 22 peptide), Flagellin-Sensing 2 (FLS2) and BRI1-Associated Receptor Kinase 1 (BAK1) transphosphorylate each other, releasing the cytoplasmic kinase botrytis-induced kinase 1 (BIK1). BIK1 phosphorylates Respiratory Burst Oxidase Homolog D (RBOHD) to trigger a reactive oxygen species (ROS) burst, promotes Ca2+ influx through cyclic nucleotide-gated channels (CNGCs), and phosphorylates Mitogen-Activated Protein Kinase Kinase Kinase 3/5 (MAPKKK3/5) to activate the mitogen-activated protein kinase (MAPK) cascade, which regulates defense-related transcription factors (TFs). Upon recognition of nlp20 (necrosis- and ethylene-inducing peptide 20), the RLP23–Suppressor of BIR1-1 (SOBIR1) complex recruits BAK1 and receptor-like cytoplasmic kinases (RLCKs), such as PBS1-like protein 31 and 13 (PBL31/PBL13), and signals through the enhanced disease susceptibility 1–phytoalexin deficient 4–activated disease resistance 1 (EDS1-PAD4-ADR1) module to activate defense genes. (2) TNL resistosomes. Toll/interleukin-1 receptor–nucleotide-binding–leucine-rich repeat (TIR-NBD-LRR, TNL) proteins detect pathogen effectors and form resistosomes that activate the EDS1-PAD4-ADR1 module, triggering effector-triggered immunity (ETI). ADR1 oligomerizes to form plasma membrane Ca2+ channels, increasing cytoplasmic Ca2+ levels and inducing the hypersensitive response (HR). (3) Shared hub. The EDS1-PAD4-ADR1 module functions in both PTI and ETI, acting as a key convergence point to coordinate immune responses. (4) Mutual reinforcement of PTI and ETI. ETI enhances PTI by upregulating key PTI signaling components, such as BIK1, BAK1, Mitogen-Activated Protein Kinase 3 (MPK3), and RBOHD. Conversely, PTI and its associated Ca2+ influx promote the expression of TNL genes, thereby strengthening ETI. (5) PTI-mediated suppression of ETI. Avirulence protein Rpt2 (AvrRpt2) can trigger HR through the MPK3/MPK6 pathway; however, PAMPs such as flg22 activate Mitogen-Activated Protein Kinase 3/6–WRKY transcription factor 18 (MPK3/6–WRKY18) signaling, which increases Arabidopsis Protein Phosphatase 2C 1 (AP2C1) and Protein Phosphatase 2C 5 (PP2C5) (PP2C family proteins), thereby suppressing ETI-induced HR. The figure was created with figdraw.com.
PTI and ETI exhibit synergetic effect and mutually reinforce each other. Ngou et al. [195] found that PTI-impaired Arabidopsis mutants are more susceptible to AvrRps4 infection, indicating that ETI alone is insufficient to combat pathogens without PTI. They also showed that PTI enhances ETI-mediated hypersensitive cell death (HR), which is aligned with the findings of Yuan et al. [196], which demonstrated that PRR mutants impair ROS production during ETI, highlighting the essential role of PRRs in ETI. Moreover, ETI signals can also enhance PTI outputs [195, 196, 197].
In nlp20-induced PTI, RLP23, EDS1, PAD4, and ADR1 play key roles. The co-receptor SOBIR1 bridges RLP23 to the EDS1-PAD4-ADR1 module, enabling signal transduction [198, 199]. EDS1, PAD4, and ADR1 are also crucial for TNL-mediated ETI. Thus, the EDS1-PAD4-ADR1 module acts as a convergence point for PTI and ETI signaling, coordinating transcriptional reprogramming and systemic immunity, and illustrating mechanistic overlap between PTI and ETI.
However, PTI can also suppress ETI. In Arabidopsis, flg22-induced PTI activates the MPK3/MPK6-WRKY18 pathway, which upregulates the phosphatases AP2C1/PP2C5. These phosphatases suppress Pst DC3000 AvrRpt2-induced MAPK activation, dampen AvrRpt2-triggered ETI and reduce ETI-mediated HR [200]. Flg22 also triggers a separate pathway, PIPs-RLK7-MPK3/MPK6-WRKY18/40/60-prePIPs, which is required for PTI-mediated ETI suppression. PIP2 treatment induces pre-PIP2 production, which suppresses AvrRpt2-induced MAPK activation through the MPK3/6-WRKY18/40/60-AP2C1/PP2C5 module, ultimately inhibiting ETI-mediated HR.
PTI-ETI interplay involves complex signaling networks. While PAMPs/DAMPs trigger PRR-mediated immunity, feedback loops suppress excessive ETI, balancing immune responses. This balance maintains robust immunity without overactivation, tailoring immune responses to pathogen threats. To provide a structured overview of their distinct yet interconnected features, a comparison of PTI and ETI is summarized in Table 2 (Ref. [1, 2, 18, 35, 51, 68, 70, 73, 82, 89, 108, 109, 114, 115, 116, 117, 118, 154, 158, 195, 196, 197, 198, 199, 200, 201, 202]). Future work should explore spatiotemporal PTI-ETI dynamics and their evolutionary origins.
| Category | PTI | ETI | Reference |
| Activation mechanism | Triggered by membrane-localized PRRs upon perception of conserved PAMPs (e.g., flg22, elf18, chitin) | Initiated by intracellular NLRs recognizing specific pathogen effectors, often targeting host defense components | [1, 201] |
| Receptor type and location | PRRs such as FLS2, EFR, CERK1, which are plasma membrane-localized receptor kinases forming complexes with BAK1 | NLRs (TIR-NLRs, CC-NLRs) localized in the cytosol and nucleus, containing NB-ARC and LRR domains | [1] |
| Downstream signaling | Activates RLCKs (e.g., BIK1), MAPKs (e.g., MPK3/6), CDPKs, ROS burst, Ca2+ influx, and transcriptional reprogramming via transcription factors | Shares the same signaling modules as PTI but with greater intensity and prolonged duration; MPK3/6 shows extended activation in ETI | [35, 68, 73, 109, 115, 154, 200] |
| ROS and Ca2+ signaling | Transient ROS burst via BIK1/CRK2/CPK5 phosphorylation of RBOHD; Ca2+ influx via CNGC2/4 is rapid but short-lived | ROS burst is stronger and more sustained; ETI enhances RBOHD phosphorylation stability and Ca2+ elevation is amplified during hypersensitive response (HR) | [108, 114, 115, 116, 117, 118, 197, 198] |
| MAPK Dynamics | Transient activation of MPK3/6 typically peaks within 15–30 minutes | Sustained MPK3/6 activation in ETI, leading to prolonged TFs phosphorylation and robust transcriptional defense response | [51, 73, 195, 196, 197] |
| Transcriptional Output | Induces early immune genes, involves CTD phosphorylation of RNA Pol II | Activates a broader range of resistance genes, with stronger magnitude and longer duration | [2, 18, 70, 82, 89, 201] |
| Programmed cell death (PCD) | Rarely induces HR; ROS and Ca2+ responses are usually non-lethal and localized | Frequently triggers hypersensitive response (HR), especially via TIR-NLR signaling amplified by EDS1-PAD4-ADR1 | [1, 198, 199, 201] |
| Response strength & duration | Moderate and short-lived, often subsiding within 2–3 hours | Stronger, broader, and longer-lasting (6–12+ hours), with reinforced PRR stability and downstream signaling | [2, 158, 202] |
| Functional relationship | ETI often follows PTI and depends on early PTI signals such as BIK1, MAPKs, RBOHD, and transcriptional priming | ETI enhances PTI by stabilizing PRRs (e.g., FLS2/BAK1), prolonging MAPK activation, boosting BIK1 activation and reinforcing ROS signaling | [158, 195, 196, 197, 198, 199, 202] |
PRR, pattern recognition receptor; PAMP, pathogen-associated molecular pattern; flg22, flagellin 22 peptide; elf18, elongation factor Tu epitope 18; NLR, nucleotide-binding leucine-rich repeat receptor; TIR-NLR, Toll/Interleukin-1 receptor–NLR; CC-NLR, coiled-coil–NLR; NB-ARC, nucleotide-binding APAF1–R proteins–CED4 domain; LRR, leucine-rich repeat; FLS2, Flagellin-Sensing 2; EFR, elongation factor Tu receptor; CERK1, chitin elicitor receptor kinase 1; BAK1, BRI1-associated kinase 1; RLCK, receptor-like cytoplasmic kinase; BIK1, botrytis-induced kinase 1; MAPK, mitogen-activated protein kinase; CDPK, calcium-dependent protein kinase; ROS, reactive oxygen species; CRK2, cysteine-rich receptor-like kinase 2; HR, hypersensitive response; TF, transcription factor; CTD, C-terminal domain; RNA Pol II, RNA polymerase II; PCD, programmed cell death; EDS1-PAD4-ADR1, enhanced disease susceptibility 1–phytoalexin deficient 4– activated disease resistance 1.
Pattern-triggered immunity (PTI) forms a fundamental layer of plant innate immunity, regulated by a complex network of receptor-like kinases (RLKs), receptor-like cytoplasmic kinases (RLCKs), MAPKs, and CDPKs. These kinase pathways convert extracellular signals into intracellular responses, triggering rapid immune outputs such as reactive oxygen species (ROS) production, calcium influx, transcriptional reprogramming, and stomatal closure.
Although many components of PTI signaling are increasingly well understood, conventional bulk-tissue analyses often overlook the spatial and cellular complexity of immune responses. Recent advances in single-cell and spatially resolved transcriptomic studies have begun to reshape this understanding. For instance, Delannoy et al. [203] showed that different leaf cell types in Arabidopsis leaves exhibit distinct immune transcriptional programs and non-uniform activation patterns upon bacterial infection, revealing coordinated but variable responses across tissues. Similarly, Han and Tsuda [204] applied temporally resolved single-cell profiling to trace changes in immune activity over time, revealing dynamic response patterns and cell-type–specific defense states. These studies provide valuable insight into the spatial and temporal regulation of plant immunity.
Such findings highlight the necessity of moving beyond static, bulk-based models toward high-resolution approaches that reflect the dynamic and context-dependent nature of immune signaling. Integration of single-cell and spatial transcriptomics with complementary technologies—such as real-time biosensors, phosphoproteomics, optogenetics, and inducible perturbation systems—will be essential to dissect transient kinase activity and cell-to-cell signaling at high resolution. Moreover, systems-level modeling approaches can help explain how immune responses maintain specificity, stability, and adaptability in complex biological networks.
Together, embracing these emerging tools will deepen our mechanistic understanding of PTI and its interface with ETI, ultimately supporting the rational engineering of crops with durable and cell-type–specific resistance to evolving pathogens.
PTI, pattern-triggered immunity; PRRs, pattern recognition receptors; PAMPs, pathogen-associated molecular patterns; DAMPs, damage-associated molecular patterns; MAPKs, mitogen-activated protein kinases; CDPKs, Ca2+-dependent protein kinases; ACS, 1-aminocyclopropane-1-carboxylate synthase; ROS, reactive oxygen species; ETI, effector-triggered immunity; NB, nucleotide-binding; LRR, leucine-rich repeat; RBOHD, respiratory burst oxidase homolog D; PA, phosphatidic acid; NO, nitric oxide; PLC, phospholipase C; PLD, phospholipase D; DGK, diacylglycerol kinase; PDK1, phosphoinositide-dependent protein kinase 1; BIK1, botrytis-induced kinase 1; CDKC, cyclin-dependent kinase complex; PBLs, PBS1-like proteins; ET, ethylene; JA, jasmonic acid; SA, salicylic acid; ABA, abscisic acid; FLS2, Flagellin-Sensing 2; RLKs, receptor-like kinases; RLPs, receptor-like proteins; CPK, calcium-dependent protein kinase; AGC, protein kinase A, G, and C families; AtRBOHD, Arabidopsis thaliana Respiratory Burst Oxidase Homologue D; MAPKKs, MAPK kinases; MAPKKKs, MAPKK kinases; WRKY33, WRKY DNA-binding protein 33; VIP1, virus-interacting protein 1; PR1, pathogenesis-related 1; ASR3, altered seed germination 3; ERF, ethylene response factor; BES1, BRI1-EMS-suppressor 1; VQPs, VQ motif-containing proteins; MVQ1, MPK3/6-targeted VQ Motif-containing protein 1; ERF6, ethylene response factor; CTD, C-terminal domain; HDACs, histone deacetylases; PUB, plant U-box; MVB, multivesicular body; LIP5, lyst-interacting protein 5; RLCK, receptor-like cytoplasmic kinase; NADPH, nicotinamide adenine dinucleotide phosphate; CIPK14, calcineurin B-Like-Interacting Protein Kinase 14; GPA1, G protein alpha subunit; XLGs, extra-large G proteins; GAP, GTPase-accelerating protein; NPR, Natriuretic Peptide Receptor; EDS, Enhanced Disease Susceptibility; PAD, phytoalexin deficient; ADR, activated disease resistance.
Conceptualization was carried out by CH and FY; review framework design and thematic organization by CH, FY and JD; literature collection, analysis and interpretation by FY, JD, BL, ZZ and MZ; writing—original draft preparation by FY; writing—review and editing by JD, BL, ZZ, MZ and CH; supervision by CH; project administration by CH; and funding acquisition by CH. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
The authors thank the researchers whose work has been cited in this review for their valuable contributions to the field. We also appreciate the reviewers for their constructive comments and insightful suggestions, which have greatly improved the quality and clarity of this manuscript.
This research was funded by The Key Technology R&D Program of Shandong Province of China, grant number 2022CXPT007 and The Postdoctoral Science Foundation of China, grant number 2022M721980.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.


