1 School of Pharmacy, Faculty of Medicine and Health, University of Sydney, Sydney, NSW 2006, Australia
Abstract
The first and second phases of the human microbiome project provided a view of mucosal surfaces and the skin of humans that mapped an abundant and complex ecosystem (microbiota) that is composed of bacteria (bacteriobiota), fungi (mycobiota), viruses (virobiota), enteric phages, archaea, protists, and helminths. Intestinal dysbiosis describes an adverse shift in microbial homeostasis in the gut that enhances intestinal epithelial permeability, translocating toxins that may lead to endotoxemia. Numerous intestinal and extra-intestinal illnesses have been linked to gut dysbiosis, including inflammatory bowel disease, infections, food allergies, asthma, diabetes, obesity, multiple sclerosis, autism, periodontitis, and colorectal cancer. The gut-lung axis is a bidirectional communication network between the lungs and the intestines mediated by bacterial elaborated products (e.g., butyrate), immune cells and neural pathways influencing health and disease at both sites. This review has focused on the gut-lung axis and the role that probiotics, prebiotics and postbiotics may play on the amelioration of respiratory symptoms that may result from viral and/or bacterial lung infections. Clinicians have for some time focused on treating inflammatory lung disorders such as asthma and chronic obstructive pulmonary disease by encouraging beneficial effects on the intestinal microbiome through the gut–lung axis with orally administered probiotics and pre- and/or postbiotics. The purpose is to restore gut microbial homeostasis. Developing novel delivery platforms to administer probiotics directly to the airways or as adjunctive systemic modulators is a plausible and increasingly supported hypothesis, with careful strain selection, formulation to preserve viability, targeted delivery, and rigorous safety and efficacy testing before clinical use. It is posited that such adjunctive treatments may significantly influence the lung microbiota epigenome by positively impacting the balance of microorganisms within the lung, restoring eubiosis and consequently health.
Keywords
- microbiota
- dysbiosis
- probiotics
- prebiotics
- butyric acid/short-chain fatty acids
- respiratory tract infections
- asthma
- chronic obstructive pulmonary disease
- cystic fibrosis
- intestinal permeability
- immunology
The intestinal and lung mucosal surfaces share structural similarities, a consequence of their embryonic origin [1]. The initial phases of the human microbiome project (HMP) revealed a diverse and intricate system of bacteria, fungi, viruses, phages, archaea, protists and helminths inhabiting human mucosal surfaces and skin [2, 3]. Consequently the “microbiota” ecosystem has progressed research in microbiome-host interactions in health and disease [4], that has focused on the gut-brain axis [5], gut-skin axis [6], gut-liver axis [7], gut-lung axis [8] and gut-joint axis [9].
Genome investigation technologies such as high-throughput sequencing have demonstrated substantial intra-individual microbiome variation (i.e., at the phylum level) at different anatomical sites (Fig. 1, Ref. [2, 10]). In addition, studies have shown inter-individual variations at the same anatomical sites as reported by different investigative groups [2, 11, 12, 13, 14]. This has been achieved by employing a competitive response with pathogens (e.g., Clostridium perfringens), through niche exclusion [15], the production of antimicrobial peptides, participation in metabolism and energy production [16], bile salt metabolism, short chain fatty acids (SCFAs) production and utilization, synthesis of vitamins, neurotransmitters, as well as xenobiotic degradation [17]. Accordingly, the intestinal microbiome has been reported to exert significant influence on the host’s health [18] from birth prompting the development of the immune system to achieve immunological and metabolic tolerance [19].
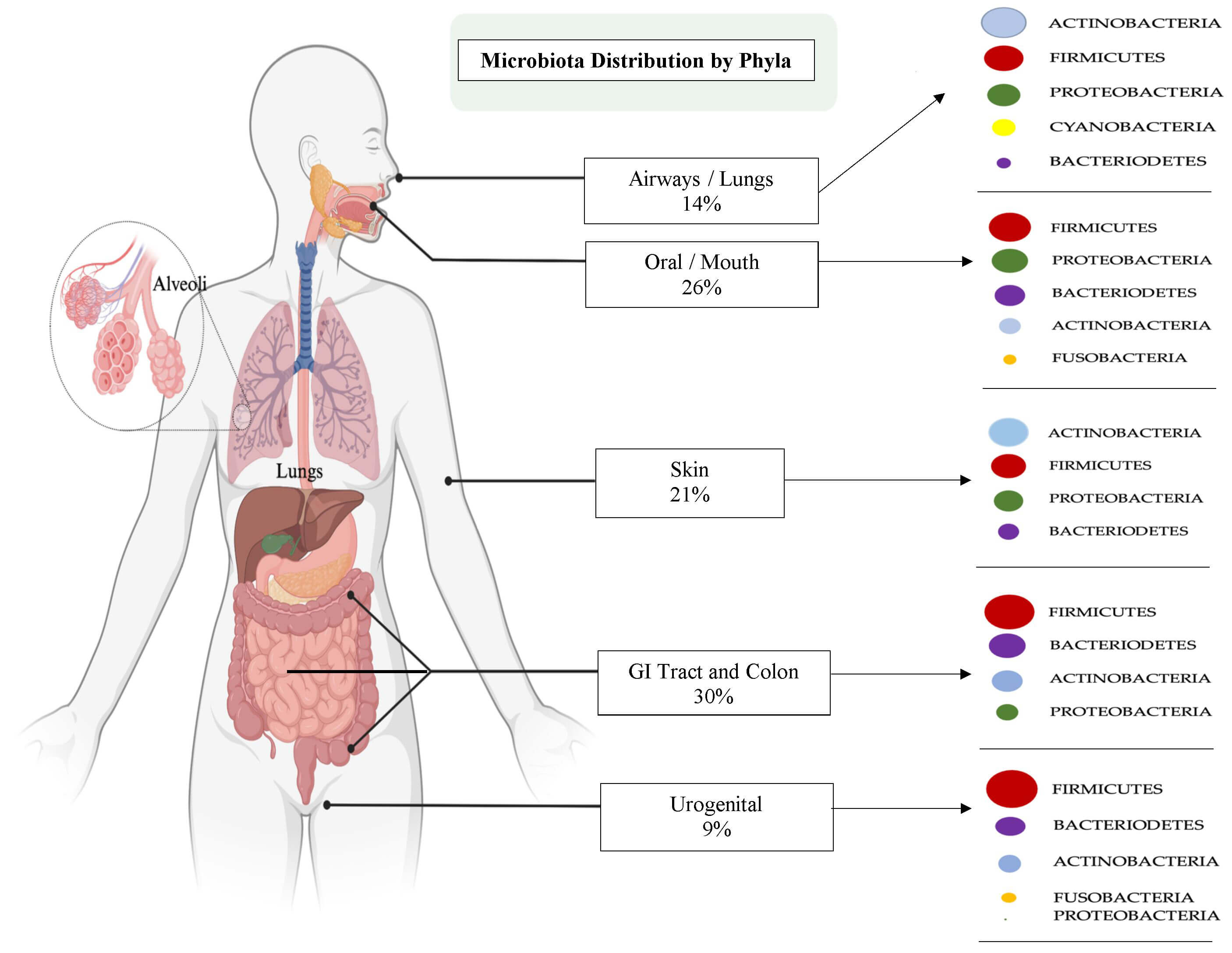 Fig. 1.
Fig. 1.
Human microbiota/microbiome distribution across anatomical sites displaying microbial phyla with percentage allocations in different structural sites. [Adapted from Cho and Blaser (2012)] [2] and Ziki Ma et al. [10]. Figure created in BioRender.com by A Byun and L Vitetta 2025.
Therefore, a posit has been advanced that there are a multitude of intestinal and extra-intestinal illnesses linked to gut dysbiosis [20, 21, 22] related to an overgrowth of pathobionts [23]. Pathobionts are commensal microbes that may cause disease when subject to environmental or host pressure [23]. These can include inflammatory bowel diseases (IBD), infections, food allergies, asthma, diabetes, obesity, multiple sclerosis, autism, periodontitis, and colorectal cancer [24]. The narration for this review will focus on the gut-lung axis, bacteria-based epigenetics, and the role that functional foods (e.g., probiotics, prebiotics) may have on the mitigation of respiratory symptoms that may ensue from viral and or bacterial infections of the lung. Furthermore, the exploration of a novel delivery platform for the administration of heat-killed probiotics as adjunctive medicines for the management of respiratory infections is advanced.
Epigenetic regulation encompasses multiple mechanisms that includes DNA modifications that describe stable changes to cellular functions [25]. The biochemical signals that control DNA-protein interactions progress phenotypic changes without producing genetic mutations in the infrastructure of the nucleotide sequence [26, 27, 28]. Epigenetic marks are superimposed on the genome that then shapes transcriptional actions and the cellular identity [28, 29]. Epigenetic modifications can also include acetylation, methylation, phosphorylation, and ubiquitination and constitute a dynamic regulatory layer that controls chromatin structure and thereby gene expression. Histone acetylation is a central and well-characterized epigenetic event governed by the opposing activities of histone acetyltransferases (HATs), which add acetyl groups to lysine residues on histone tails, and histone deacetylases (HDACs), which remove the acetyl groups. Acetylation neutralizes positive charges on histones, relaxes nucleosome packing and increases accessibility of the transcriptional machinery, promoting gene activation. Conversely, deacetylation restores chromatin compaction and contributes to transcriptional repression. Moreover, the HATs–HDACs balance is highly context dependent and integrates upstream signaling, metabolic state, and developmental cues to produce cell-type specific transcriptional programs. Perturbations of this balance are implicated in diverse lung pathologies including cancer and immune disorders because they can broadly rewire gene networks controlling proliferation, differentiation, apoptosis, and inflammatory responses [25, 28] (Fig. 2).
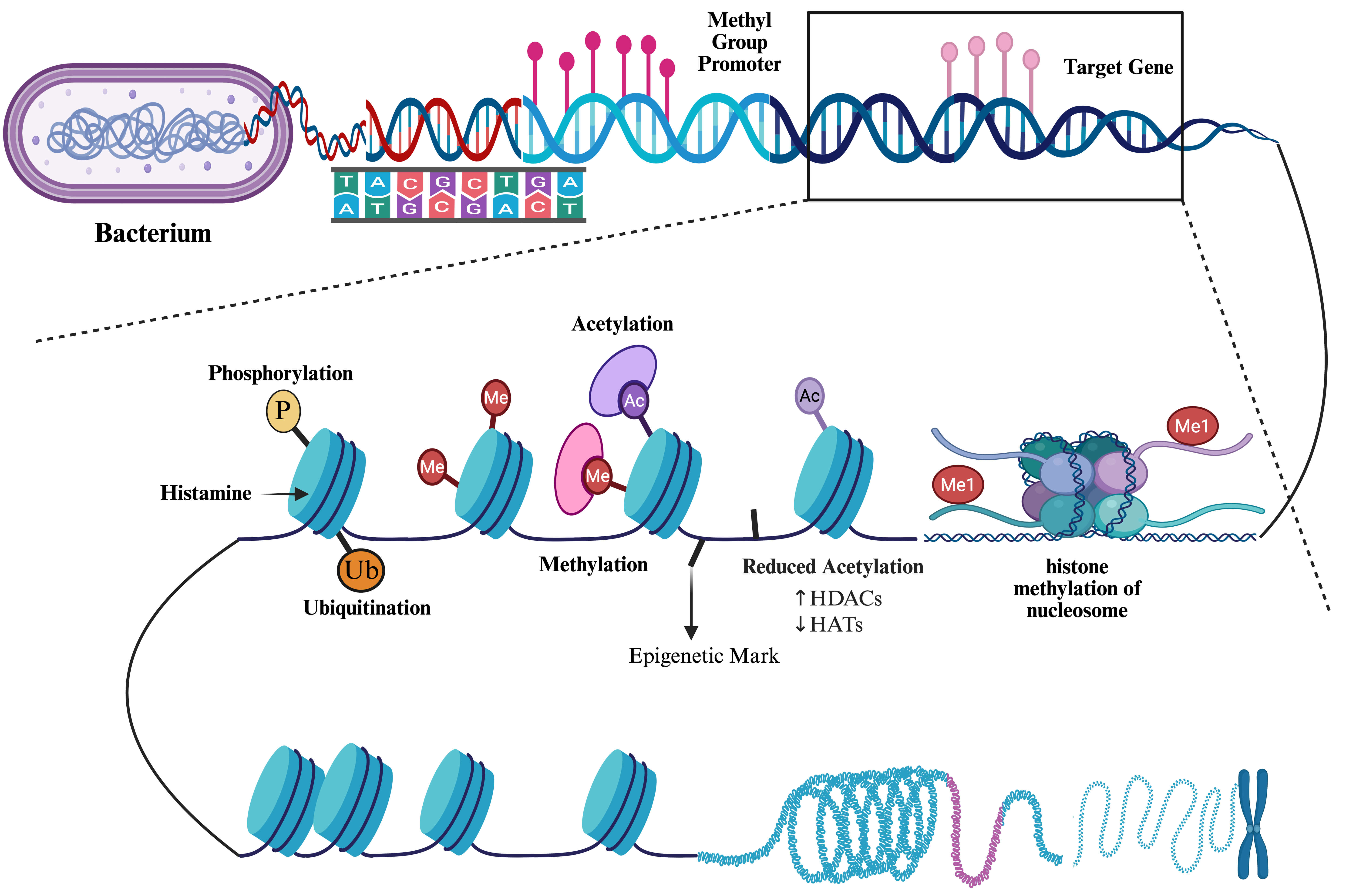 Fig. 2.
Fig. 2.
Diagrammatic representation of bacterial regulatory epigenetic modification mechanisms. [Created in BioRender.com by Vitetta L 2025].
Clinical knowledge of epigenetics in lung diseases is further complicated through the reduced activity of histone deacetylases (HDACs) [30] the enzymes that remove acetyl groups from histone proteins, increasing chromatin compaction and reducing accessibility to the transcriptional machinery [31]. It is reported that by tightening DNA–histone interactions, HDAC activity represses gene transcription and thereby influences fundamental cellular processes such as proliferation, differentiation, apoptosis, and stress responses [31]. Furthermore, HDACs are implicated in numerous diseases, notably cancer, where altered acetylation dynamics contribute to dysregulated growth and survival pathways [32]. Whereas, histone acetyltransferase (HAT) activity acetylates histone lysine, relaxing the chromatin structure and facilitating transcription of proinflammatory genes in the airways [33]. In asthma, upregulated HAT activity skews the epigenetic balance toward enhanced expression of cytokines, chemokines, and other mediators that drive airway inflammation and hyperresponsiveness [33]. Inhaled corticosteroids exert part of their anti-inflammatory effect by restoring acetylation balance, in part through suppression of HAT-driven transcriptional activation. Additional therapeutic strategies therefore aim either to restore physiological acetylation dynamics (e.g., employing HDAC inhibitors or HAT modulators) or to target specific acetylation-dependent pathways, with the goal of achieving selective modulation of pathogenic transcriptional states while minimising off-target effects.
Notwithstanding, DNA methylation comprises the most pervasive active mechanism serving as a fundamental biochemical action that controls gene regulation [34, 35]. Several epigenetic mechanisms influence genetic processes such as DNA modifications (e.g., CpG methylation and demethylation), histone modifications (e.g., acetylation and deacetylation), and RNA that integrate gene regulatory networks with environmental cues [35]. Of these DNA methylation comprises the most stable modification that can persist following cell divisions to the next generation. DNA methyltransferases (DNMTs) are indispensable enzymes whose activities establish and maintain methylation biochemical processes that overall ensure genome fidelity and continuance of epigenetic regulation [35, 36].
Epigenetic-like DNA, RNA/microRNA methylations and histone modifications significantly influence gut bacterial cellular functions such as metabolism, bacterial persistence, replication cycles, bacteriophage susceptibility, virulence and genome plasticity [37, 38, 39, 40, 41]. Methylation reactions in bacteria have been reported to commonly target N6-adenine and C5-cytosine residues [40, 41], and with N4-cytosine methylation also observed in many bacterial taxa [40]. Beyond classical methylation reactions reported, novel DNA modifications such as phosphonothioate based alterations [42] such biochemical reactions influence cellular redox potential status [43] thereby expanding the repertoire of heritable, non-sequence-based regulatory entities [42]. These modifications, together with RNA methylation, can produce stable, heritable phenotypes that alter gene expression without changes to the genome sequence of bacterial species. Moreover, these post-replicative modifications [43] enable rapid, reversible tuning of gene expression in response to local environmental pressures. Such responses allow bacteria to adapt growth, survival/persistence, and pathogenic strategies on a timescale that is more rapid than those through de novo mutations. This adaptive plasticity links bacterial DNA/RNA modification systems to organismal functions and, increasingly to disease processes in mammalian hosts.
Epigenetic changes can have a significant impact on the balance of microorganisms in any anatomical site. The interplay between the microbiome and the epigenome is intricate and mutually influential, especially in the gut [44]. In the lungs a dysbiotic microbial environment may contribute to the development and progression of lung diseases [45]. Modulation of the microbiota in the intestines through epigenetic changes that can be provoked by probiotics, prebiotics, synbiotics SCFAs, and micronutrients [44, 46] have been postulated to represent potential therapeutic strategies that protect against lung diseases [44, 45].
Many bacterial species are subject to epigenetic changes describing DNA modifications as gene biochemical regulatory actions. That is where epigenetic signals control DNA–protein interactions and can cause phenotypic changes in the absence of a mutation. A recent review [47] appropriately summarized how epigenetic DNA methylations in bacteria can protect bacterial genomes, promote chromosome replication and segregation, and nucleoid organization, and control bacterial cell cycles as well as repair bacterial genome DNA and regulate transcriptional activities. Moreover, DNA methylation has been shown to control the reversible switching of gene expression [48]. This versatile action is a phenomenon that generates phenotypic cell variants [49]. Hence, the development of epigenetic bacterial lineages is important as it facilitates the adaptation of bacterial populations to severe or changing environmental conditions and modulates the interaction of prokaryotic pathogens with their eukaryotic hosts [34, 49].
Reports cite epigenetic regulation and control as being increasingly recognized as a potent mechanism through which the microbiota influence host physiology occurring through multiple potential mechanisms [44, 47]. That is, in addition to environmental inputs (e.g., nutrition), epigenetic control includes (i) microbial biosynthesis or metabolism which influence the availability of chemical donors for DNA methylation, histone modifications or chromatin remodelling; (ii) regulation of epigenetic-modifying enzyme expression and/or activity; or (iii) activation of host cell intrinsic processes that direct epigenetic pathways such as microRNA pathways [47].
Research on the benefits of probiotic bacteria from major genera such as Lactobacillus has progressed from observational and interventional studies to the identification of the underlying molecular mechanisms that exist [34]. A regulatory mechanism of chromatin structure and gene expression is histone acetylation. Recent in vitro studies that have explored gene expression reported that probiotic bacteria such as Lactobacillus rhamnosus and Lactobacillus fermentum modulate host epigenetic signatures of intestinal epithelial cells through global histone acetylation independent of the recruitment of transcriptional activators and via Escherichia coli challenge [50].
A recent comprehensive screening investigation of existing datasets with chronic lung diseases utilising data from biological samples has further emphasised the importance in deciphering the relationship between the lung microbiome, epigenetics and its modifications in the prevention of chronic pulmonary disease or its progression [51]. An appreciation of the role of the lung microbiome in maintaining local eubiosis is recognized. Epigenetic microbial alterations can impact and skew the lung microbial cohort toward dysbiosis, significantly contributing to the development and progression of lung diseases [51].
Pseudomonas is a bacterial pathobiont that can cause respiratory
infections in individuals with compromised immune systems [52]. As it occurs in
severe asthma, cystic fibrosis or chronic obstructive pulmonary disease (COPD).
P. aeruginosa is an opportunistic pathogen that frequently infects the
lungs, particularly in immunocompromised patients. In individuals with cystic
fibrosis, it colonises the thickened mucus lining of the airways. In the lungs
the mucus provides a supportive, nutrient-rich environment for the bacterium’s
survival without disturbing the local mucosal airway epithelial cell scaffold
[53]. The persistent presence of this pathobiont in the respiratory tract of the
lungs can lead to disruption of the composition of the respiratory microbiome.
This disturbance results in changes to its normal functionality increasing
susceptibility to respiratory infections [54]. In cystic fibrosis, changes in the
It is generally accepted that the lungs and the gut are two independent systems that originated from one common embryonic organ, the foregut [58]. These two distinct anatomical sites share an embryological origin that structurally contain mucosa-lined luminal surfaces that show specific anatomic and biochemical features [59], resulting in markedly different microbial communities and immune system interactions [60]. Currently emerging research shows that there is a gut-lung axis that very much allows for bidirectional crosstalk interactions between the intestinal microbiota and the lung microbiota. The lungs and intestines, with large surface areas exposed to the external environment, are subject to immune homeostasis that determines continued health [61]. This proposition is dependent on a varied and balanced microbiome that includes microbial components and metabolites (e.g., SCFAs) that are important for the priming, maturation, and metabolic equilibrium of the immune system in both anatomical sites [62].
Environmental factors such as diet, psychosocial stressors, physical activity, and the use of prescribed antibiotics significantly affect the stability and composition of the gut and lung microbiomes [63, 64, 65]. Dysbiosis of the intestinal microbiome has been extensively studied and the data from association studies show that changes in the abundance and evenness of the gut microbiome can affect the susceptibility to diseases in distant organs, as so happens with pulmonary diseases. Inflammatory diseases of the gut (e.g., IBD), especially in patients diagnosed with ongoing IBD and irritable bowel syndrome (IBS), present a higher prevalence of pulmonary disease. Intestinal microbiome adverse changes have been reported to occur in various chronic lung disorders, such as asthma [66], COPD [67], and cystic fibrosis (CF) [68]. Moreover, the prescription of antibiotics in early life has been implicated [69] and reported to significantly increase the risk of developing atopy and specifically exacerbating asthma [70], and subsequently correlated to a reduced abundance of gut bacteria [71]. The intestinal microbiota also participates in a protective role against common viral and pulmonary infections [72], through the regulation of the innate and adaptive immune responses [73].
The relationship between the intestinal cohort of bacteria and common respiratory diseases involves gut derived metabolites such as SCFAs that stimulate protective mechanisms by increasing macrophage/dendritic cell progenitor cells [46]; as well as being directly involved in modulating the production of lipopolysaccharides (LPS). The bacterial metabolites SCFAs have key roles in the migration of immune cells via the systemic circulation, priming myeloid cells in the bone marrow promoting haematopoiesis, thereafter, migrating to the lungs supporting an anti-inflammatory environment [46].
The lungs are populated with alveolar macrophage progenitor cells that activate Th2 effector cells, which mediate immune responses through the production of local cytokines [46]. In essence, the changes in the abundance of intestinal bacteria that affects gut bacterial metabolites is linked to changes in immune responses that progress pro-inflammatory sequelae.
The host–microbe connections underpin the bidirectional crosstalk that exists between the intestines and the lungs involving physiological and pathological connections [74]. The pulmonary cohort of bacteria has been reported to modulate microbial communities in the intestines with a subsequent influence on intestinal signaling [75]. In the lungs, infections such as Mycobacterium tuberculosis have been reported to dysregulate the immune system, resulting in alteration in the intestinal microbiome. Studies with patients diagnosed with tuberculosis (TB) versus healthy controls the phyla of Firmicutes, Proteobacteria, and Verrucomicrobia were reduced in abundance whereas the Actinobacteria, Bacteroidetes, and Fusobacteria phyla expressed an increased abundance [76]. In patients with de novo infections or recurrent TB infections the changes in gut bacterial cohort showed decreased abundance in Bacteroidetes, the genus Prevotella and Lachnospira and enrichment in Actinobacteria and Proteobacteria [77]. The dysbiosis reported provides evidence for a disequilibrating in the production of microbial elaborated metabolites such as SCFAs. In a reverse intestine to lung cross-talk study it was shown that lung colonisation in the intensive care unit was driven by the translocation of Pseudomonas aeruginosa from the intestines [78].
Alterations in the gut microbiome in patients with asthma have been observed.
Current evidence points to the existence of the gut-lung axis from birth as the
microbial composition affects the risk of developing asthma in infants [79]. In
the infant gut microbiome, a significant increase in
In adult patients with mild-moderate asthma with well-controlled symptoms,
strong associations between the forced expiratory volume in 1 second (FEV1)
and the differences in gut microbiota compositions at the phylum level were
observed [84]. The ratio of Bacteroidetes to Firmicutes
relative abundance was lower in subjects with asthma [84] which was in contrast
to the findings by Wang et al. [85], who reported no significant
differences at the phylum level between non-severe asthma, severe asthma, and
healthy patients. These differences may be due to variation in the environmental
exposure of the patients in different geographical locations. Unlike the
intestines, the lungs are anatomically exposed to the external environment as it
is the passage for continuous breathing. An analysis of the lung microbiota of
162 patients with chronic respiratory conditions across different care centres in
the United Kingdom revealed that there were significant differences in the
Despite the discrepancies in the current studies, the data suggests that the differences in the microbiota gut compositions were associated with specific clinical respiratory presentations [84, 85]. Both studies [84, 85] have found that the asthmatic group had a higher proportion of bacteria from the Lachnospiraceae family than the non-asthmatic group. Furthermore, separating asthmatic patients according to their gut bacterial community structures revealed that the microbiota compositions were associated with lung function [84]. One particular group with the lowest community diversity but with high Prevotella abundance presented low levels of lung function [84]. Higher relative abundances of Prevotella in the gut may be associated with a more severe clinical presentation of asthma [85]. Again, contradicting results were reported by Sampaio et al. [89] where Prevotella was more abundant in patients without asthma compared to those with type 2 inflammatory asthma. However, it should be noted that the sample size of this study was very small (n = 28 asthma, n = 29 control) and consisted of children and young adults [89].
The transient nature of the lung microbiota due to constant environmental exposure may inevitably affect the gut and hence it is difficult to conclusively link the phenotypic symptoms of asthma with the diversity of the gut microbiota. It is well recognised though that intestinal dysbiosis has been significantly linked to asthma development, with severity linked to the gut-lung axis, affecting immune system maturation, promoting inflammation, and increasing allergies [90]. Moreover, low diversity of specific beneficial bacteria (i.e., Bifidobacterium, Faecalibacterium, Roseburia) may be linked to protective effects against asthma outcomes [90], while certain microbial imbalances increase asthma risk [91]. This connection suggests that early-life microbial exposure and restoring the intestinal microbial balance with probiotics or other interventions could better help manage asthma.
Pulmonary involvement in IBD is often reported in the literature [67, 92, 93, 94, 95] and possible reasons include the presence of chronic inflammation in the intestines which in turn affect the lungs, possible due to the common embryonic origins that the lungs and intestines share [67]. An association between new onset IBD and COPD reflects microbiome dysbiosis and intestinal endothelial barrier dysfunction that may occur in the gut of patients with COPD [93]. However, despite the bidirectional nature of the gut-lung axis, no reciprocal association of increased risk of COPD in IBD patients was observed as reported in a systematic review [93]. When patients were assessed for a more general obstructive lung disease (OLD), there was a 60% increased risk for individuals with new onset IBD [92]. The risk of OLD diagnosis further increased by 40% after IBD diagnosis for all age groups [92]. These results support the outcome that induced colitis in mice was causal for lung damage [94].
To determine whether a dysfunction in the gut microbiota affects the lungs, mice were treated with specific antibiotics, such as vancomycin and ampicillin, two weeks prior to COPD induction [96]. Antibiotic treatment reduced the extent of COPD pathogenesis, indicating that the composition of the gut microbiota may result in COPD development [96]. To further explore this implication, the gut microbiota of patients with different severity of COPD were compared [97]. Over the course of one year, patients with declining lung function (reduction in FEV1) were more abundant with Firmicutes [97]. Upon comparing patients with and without COPD, no significant differences in the relative abundances of the 20 most prevalent genera were observed [98]. However, COPD patients had differences in some of the least abundant taxa where there was a reduction in Veillonella genus and an increase in an unclassifiable genus from the Clostridia class [98]. Separately, species such as Streptococcus sp000187445, S. vestibularis were found to negatively correlate with FEV1 [99]. Furthermore, COPD-associated members of Lachnospiraceae was inversely related to the predicted forced vital capacity (FVC) and FEV1, whilst Desulfovibrio piger_A, CAG-302sp001916775 positively related to the lung function [99].
COPD often involves emphysema and in patients with computed tomography-verified
emphysema had significantly reduced bacteria from the Lachnospiraceae
ND3007 group and Eubacterium halli [98]. These groups produce SCFAs
which progress and maintain intestinal homeostasis [100]. It is possible that a
decrease in SCFA production led to the dysregulation of inflammation. SCFAs,
especially propionate, can inhibit IL-17 production by intestinal
What is of clinical importance with COPD is intestinal dysbiosis [105]. An imbalance of intestinal microbes is closely linked to COPD through the gut-lung axis where gut bacteria and their metabolites (e.g., butyrate) influence lung inflammation, potentially improving or worsening COPD. Furthermore, lung issues can alter the intestinal microbiota exacerbating lung diseases. In patients diagnosed with COPD patients that express intestinal dysbiosis often exhibit changes such as lower levels of Bacteroidetes and higher levels of Firmicutes, that result in increased gut permeability and inflammatory sequelae that affects the lungs via immune cell activities and circulating substances [105, 106]. These results stress the importance of dietary intake (e.g., fiber, fermented foods) and the administration of probiotics as potential management strategies.
Gastrointestinal (GI) symptoms and gut microbiota dysbiosis are observed in patients with cystic fibrosis (CF). These differences in the composition of the intestinal microbial communities are observed from early childhood [107, 108]. The diversity of the microbiome composition plateaus at around 2 years of age for children with CF (cwCF) [108]. Generally, decreased relative abundance and diversity of GI microbiome in children [107, 108] is continued as adults [109, 110]. Significant increases in Firmicutes and Actinobacteria were observed in the gut of adults with CF (pwCF) whilst there was a decrease in Faecalibacterium, Roseburia, and Bifidobacterium, groups considered to be ‘healthy gut bacteria’ [109]. Enrichment of Adlercreutzia, Ruminococcaceae, Lachnospiraceae, Tyzzerella, and Candidatus soleaferrea in paediatric CF gut microbiota were directly associated with FEV1 [107]. CF inflammatory marker calprotectin positively correlated with Acidaminococcus, Allisonella, Eubacerium coprostanoligenes group, Howardella, Lachnospiraceae UCG-010, Mogibacterium, Olsenella, Sutterella, uncultured Lachnospiraceae, and uncultured Porphyromonadaceae [107]. Fusobacterium, which has been linked to colorectal cancer (CRC) [111, 112, 113] was more abundant in cwCF [107] but not in pwCF [114]. A possible reason for its absence in adults may be due to the continual development of the infant microbiome with aging. Price et al. [108] reported that that majority of the taxa changed in relative abundance to shift towards “a healthy-like” microbiome with age. However, genera such as Prevotella_7, Akkermansia, Bifidobacterium and Blautia increased to resemble “a CF-like” microbiome [108].
Nonetheless, the risk of colorectal cancer in pwCF was elevated [115], despite
the cluster distinction of the gut microbiome of pwCF from CRC and healthy
samples [114]. In addition to CF-driven microbiota changes, the greater exposure
to antibiotics for pwCF may further account for these differences. Early
antibiotic exposure can have a significant long-term impact on the gut microbiota
[108]. Within the cwCF, as expected, children who used antibiotics had a
decreased
As with asthma and COPD, cystic fibrosis has also been linked to intestinal dysbiosis as a causative factor [122]. Mechanistically a defective Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) protein progresses thick mucus, dehydration and adverse changes in the intestinal environment with overall reductions in beneficial bacteria (e.g., Bacteroides) and increased abundance of dysbiosis triggering bacteria (e.g., E. coli, Staphylococcus). These adverse microbiome changes are known to drive inflammatory processes, disrupting the intestinal epithelial barrier with links to issues such as small intestinal bacterial over growth (SIBO), distal intestinal obstruction syndrome (DIOS), and an increase in cancer risk [123]. It is reported that new therapies such as CFTR modulators and probiotics show promise for restoring gut balance and as such improving cystic fibrosis outcomes [124].
The intestinal microbiota has been extensively studied, with first reports appearing in the Western literature circa the 1840s [125]. The lung microbiota has alternatively had a relatively recent history, established in 2010 by Hilty et al. [126], and successfully challenged the then-established misconception of a “sterile lung”. However, pulmonary manifestations have been reported over the last 50 years in patients with IBD [127, 128]. Connections between the intestines and the lungs have only now been recognised and the establishment of a gut-lung axis has only recently been addressed.
Reports from studies on healthy individuals and those with disordered microbial communities in the lungs, detail a core microbiome in the lungs that includes bacteria from the Pseudomonas, Streptococcus, Proteus, Clostridium, Haemophilus, Veillonella, and Porphyromonas genera [126, 129]. In the lungs, the greater proportion of bacteria are aerobic or obligate anaerobes such as those from the Clostridium, Veillonella and Porphyromonas genera [59].
The ecological habitat and bacterial niches that exist in the lungs express a dynamic environmental flux that is very much dependent on bacterial migratory movements from the nasal cavity, paranasal sinuses, pharynx, and supraglottic portion of the larynx. Notwithstanding the upper respiratory tract displays significant topographical differences in microbial composition of dominant taxa that inhabit different anatomical locales, such as the nasal cavity and nasopharynx (i.e., species from Moraxella, Staphylococcus, Corynebacterium, Haemophilus, and Streptococcus groups), and the oropharynx with a high abundance of species from the Prevotella, Veillonella, Streptococcus, Leptotrichia, Rothia, Neisseria, and Haemophilusgroups [130]. As so happens with colonisation of the infant gut after birth [131], commensal bacteria in the lungs are essential for the regulated development of local immune homeostasis. In the lungs, as in the intestines [132], the recognition of microorganisms by the innate immune system initiates a signalling cascade in the respiratory tract from microbial encounters with amniotic fluid, placenta and the vagina that progresses the development of the lung microbiota, and related responses toward immune tolerance, equilibrium and maturation.
Eubiosis to dysbiosis in the intrapulmonary microbiota significantly alters the structure, abundance, and diversity of the commensal cohort increasing susceptibility to infections by pathobionts with disease progression [73]. Disruptive interactions between lung microbes and immune mucosal barriers can result in irregular local inflammatory responses (e.g., bronchopulmonary dysplasia). Innate immune response signalling can be elicited by internalising particles from ambient air such as toxins, allergens, microbes and endogenous debris [132]. As with the intestinal epithelia, the lung epithelial cells are also part of multiple mechanisms of non-haematological and haematological interactions with the intrapulmonary microbiota primarily acting as a permeability barrier that continually senses microbes and responds to their migratory effects that may lead to infections [133]. Effective clearance of endogenous particles and bacteria is the first site of interplay between the non-haematological structures of the lung (i.e., epithelia and mucus cilia) and the commensal lung cohort. Chronic infections induce the production of increased mucus levels by lung epithelial cells. High levels of mucus can facilitate the growth of bacteria by eliciting low oxygen concentrations and elevating temperatures in lower respiratory tract areas, promoting selective stability of specific bacterial species that shape intrapulmonary immunity [134].
In the intestines, macrophages that encounter resident bacteria generate signals
that lead to changes in the phenotype of macrophages corresponding to adaptive
immunity effects. Similarly in the lungs, there exists a connection between
adaptive immunity and the intrapulmonary microbiome that drives adaptive immunity
associated cell involvement [75]. Murine models with neutrophil infiltration due
to bacterial infections show high levels of IL-6 and TNF-
A healthy intrapulmonary microbiome is in a transient state of flux and is influenced by adjacent body parts and the external environment [138]. As reported for a dysbiotic intestinal microbiome, abundance and diversity changes in the status of the intrapulmonary microbiome from eubiosis to dysbiosis has links associated with disease progression [138]. In respiratory diseases, the microbiota is much more likely to be long-lasting and reside in the respiratory tract and lungs, further confirming that the lungs can also harbour an unstable infective disease state of the pulmonary microbiota. Several studies indicate the existence of a relay network between the intestinal microbiome, metabolites such as SCFAs and the intrapulmonary microbiota suggestive of a microbiota-gut-lung-microbiota axis. Perturbations originating in the composition of the intestinal microbiota can induce acute and chronic effects on the pathophysiology of lung diseases [138].
Chronic inflammatory lung diseases have a bidirectional impact on the gut
microbiome. As respiratory viral infections exert an inflammatory response from
the immune system, this may also indirectly affect the gut. Viral respiratory
infections post-allogeneic haematopoietic stem cell transplantation is more
likely to result in a lower respiratory tract infection (LRTI) [139]. However,
patients with higher abundance of butyrate-producing bacteria were inversely
associated with LRTI development from the initial infection [139]. Similar
observations were made in patients with coronavirus disease (COVID-19) [140, 141, 142].
Differences in the gut microbiome composition was observed upon comparison of the
gut microbiome of antibiotic-naïve patients with COVID-19 and patients
without COVID-19 [142]. This difference was significant when the patients were at
a severe stage of infection, with depletions in Bifidobacterium
adolescentis, Ruminococcus bromii, F. prausnitzii and an
enrichment of Bacteroides ovatus, Bacteroides dorei and
Bacteroides thetaiotaomicron was noted [142]. As a result, these
patients had lower levels of SCFAs, such as butyric acid [142]. A cross-sectional
study found that patients with COVID-19 or H1N1 influenza had significantly
reduced gut microbiota richness and diversity compared to healthy controls [143].
Patients with COVID-19 presented a discernible reduction in the abundance of the
Ruminococcaceae family and several genera within the
Lachnospiraceae family compared to healthy controls [143]. These
alterations in butyrate-producing bacteria observed in COVID-19 patients differed
from those reported in individuals with H1N1 infection [143]. Furthermore, the
reduction in butyrate-producing bacteria negatively correlated with
proinflammatory biomarkers, C-reactive protein (CRP), procalcitonin or D-dimer
levels [143]. Similarly, patients with elevated D-dimer levels
experienced more severe COVID-19 and had significant depletion of SCFAs [144].
The severity of the COVID-19 infection was negatively correlated with the
abundance of Faecalibacterium prausnitzii [140, 141], which is a
well-known butyrate producing species [145]. These observations were not limited
to lung infections, but was also noted in patients hospitalised due to abdominal,
urinary tract, and skin infections [146]. However, whether these infections were
viral or bacterial was not specified [146]. Additionally, amongst post-acute
COVID-19 syndrome (PACS) patients, those with significantly decreased lung
function, longer hospital stays, and higher rates of intensive care unit
admissions had reduced
Vaccines were also shown to alter the composition of gut microbiota. COVID-19
vaccines shifted
Respiratory infections, such as respiratory syncytial virus (RSV) infections and
influenza, increase the relative abundance of Bacteroidetes whilst
decrease Firmicutes in the gut [150]. Groves et al. [150]
suggested that this change was driven by the loss of appetite as a result of the
infection. Although the viral load was not directly related to the weight change,
depletion of CD8+ T cells resulted in an increased viral load and weight loss
[150]. CD8+ T cells are responsible for viral clearance in a respiratory viral
infection [151] and is modulated by the gut microbiota [152]. This modulation by
the microbiota may be through its metabolites as oral butyrate administration in
mice increased IFN-
The gut microbiota is not the only system to be altered in patients with respiratory viral infections [144, 162, 163, 164, 165, 166, 167]. Nasopharyngeal samples collected from infants with RSV in their first year of life had differing composition of the lung microbiota compared to infants without infection [168]. Increased abundance in Haemophilus, Streptococus, Moraxella genera and lower abundance of Dolosigranulum, Corynebacterium was associated with severe disease [168]. Similar increases in Haemophilus and Streptococcus was found in other studies that characterised the nasopharyngeal microbiota of infants under 2 years [169] and 1 year [161] of age with RSV infections. Interestingly, the study of the lung microbiota during RSV infection was more common than that of the gut and produced more consistent results as previously outlined [157, 161]. The consistency of these findings suggests that the changes in the lung microbiota due to RSV infections may be more promising as future biomarkers.
Substantial evidence has long indicated that influenza infections can induce
alterations in the gut microbiota. Sublethal infections with influenza A virus
(IAV) led to minimal changes in either
In a separate study consisting of a larger sample size and assessing patients over 60 years old with influenza-like-illness (ILI), Bacterioidetes and Proteobacteria were more abundant compared to the control [166]. It was hence reported that the relative abundance of Ruminococcus torques was positively associated with ILI and proinflammatory responses, while the co-occurrence of Echierichia and Shingella was negatively associated with certain beneficial taxa such as butyrate producers [166]. R. torques, an intestinal mucous degrader [173] associated with IBD [174] was also enriched in patients with COVID-19 [144], suggesting a possible role in contributing to the gastrointestinal symptoms during respiratory infections. The impact that IAV has on the gut microbiota composition leading to damaged intestinal barrier and decreased production of SCFA was also observed in a murine model [175]. SCFA supplementation 2 days post infection resulted in symptoms indicative of reduced inflammation and partially restored the damaged intestinal barrier [175]. Thus, IAV infection disrupts SCFA-producing commensals which may have clinically relevant implications as diminished SCFA levels affect immunomodulation and epithelial integrity. However, these clinical symptoms have not yet been thoroughly investigated in the context of IAV infections especially in humans.
The gut microbiome modulates the host immune response against bacterial lung
infections. Gut microbiota-depleted mice had higher bacterial load and higher
mortality rate compared to control mice post intranasal S. pneumoniaeinfection [176]. Faecal microbiota transplantation (FMT) to the
microbiota-depleted mice resulted in a comparable TNF-
Observations that lung diseases, such as COPD and asthma, regularly occur with chronic inflammatory intestinal diseases, such as IBD or IBS, has progressed the posit that the gut–lung axis presents an intimate connectivity in health and disease. In support, epidemiological studies have reported that at least 50% of adults with IBD and 33% of patients with IBS have pulmonary inflammation involvement or with impaired lung function and this without a previous history of acute or chronic respiratory diseases. Notwithstanding, patients diagnosed with COPD were reported to be 2–3 times more likely to be diagnosed with IBD [180]. Furthermore, in patients diagnosed with asthma there were reported functional and structural alterations in the intestinal mucosa suggestive of gut dysbiosis [181]. Likewise, patients with COPD typically had an increase in intestinal permeability [97, 182].
Intestinal dysbiosis not only describes adverse shifts in the intestinal microbiome but the consequent effects of increased intestinal permeability that is associated with numerous metabolic disorders and extra-intestinal diseases [183]. Increased intestinal barrier permeability has been associated with an exaggerated pro-inflammatory response as so happens with IBD. Pro-inflammatory cytokines and chemokines are present in elevated levels in mucosal tissue and/or in peripheral blood, suggesting a monocyte/macrophage stimulatory effects by enteric bacteria and/or their constituents (e.g., LPS) [184, 185]. Studies with murine models of IBD have reported that organisms such as Citrobacter rodentium and Helicobacter hepaticus can progress increased intestinal barrier permeability and mucosal dysbiosis with translocation of bacterial endotoxins (e.g., LPS) that trigger local endotoxemia [24].
There are numerous mechanisms that have been reported associated with intestinal microbial dysbiosis [24]. An often-cited example is the overgrowth of Enterobacteriaceae from the phylum Proteobacteria that resides in the gut near the intestinal epithelial barrier. This is attributed to the relative higher oxygen tolerance by these bacteria due to gas diffusion across the intestinal epithelium. Under normal physiological conditions Enterobacteriaceae are present in low levels. An increased abundance of Enterobacteriaceae with pathogenic members from this family (e.g., Escherichia coli) signals significantly enhanced local inflammatory responses, as observed in IBD [186]. Moreover, studies have provided evidence that an inflammatory tissue environment is conducive to perturbations of the intestinal microbiota often characterized by bacterial species endowed genetically with the capability of utilizing nutrients more abundantly found in the inflamed gut. This causes significant disturbances observed in the intestinal bacteria community structure [24].
Recently, there has been an increased understanding and consequently research
interest in the gut–lung axis and the link to respiratory diseases [187]. In an
investigation of the intestinal microbiota make-up in infants and adolescents
diagnosed with asthma it was reported that there was observed reduced levels of
A. muciniphila and F. prausnitzii critical gut commensals that
elaborate SCFAs associated with a healthy gut [188]. Consequently, the
researchers also showed that the levels of inflammatory factors, including CRP,
TNF-
Probiotics have a long history of use and have provided contentious results for
their anti-inflammatory and immunomodulatory effects in attenuating intestinal
inflammation and atopic diseases. The first documented randomised double–blind
placebo–controlled study with Lactobacillus gasseri demonstrated
efficacy in school children with asthma and allergic rhinitis [189]. There was a
significant increase on pulmonary function and peak expiratory flow rate, and the
clinical symptom scores for asthma and allergic rhinitis decreased in the
probiotic-treated group as compared to the controls [189]. In addition,
inflammatory markers TNF-
The administration of probiotics that can induce positive effects on adverse intestinal microbiome profiles has a long history of use, that is underpinned by the fact that probiotics are safe yet not a panacea to ameliorate all metabolic diseases in the gut [191]. Probiotic formulations present as single or multiple bacterial species with variable concentrations expressed as colony forming units making it difficult to translate to clinical practice. Formulations usually contain bacterial species from the Lactobacillus and Bifidobacterium genera that have been studied in different scenarios in clinical studies [191]. Notwithstanding some clinical studies have reported efficacy in upper respiratory tract infections [192], reduced occurrence of allergies in children [193] and immune response to oral rotavirus vaccine [194], results that propounds the idea of adopting further investigations with probiotics in lung diseases.
There is therefore biological plausibility for investigating inhaled probiotic
formulations delivered directly to the lungs, as a posited strategy for the
treatment of respiratory infections. Probiotics are defined as “live
microorganisms which, when administered in adequate amounts, confer a health
benefit on the host” [195]. Probiotic bacteria are conventionally administered
orally to improve gut health, but their beneficial effects are not restricted to
the gastrointestinal tract. Prophylactic oral L. rhamnosus GG (LGG)
decreased non-infective lung inflammation and fibrosis in rodents by modulating
immune responses and improving the diversity of the intestinal microbiota [196, 197]. The lung function of rats improved with reduced pro-inflammatory cytokines
(IL-1
Probiotics were administered to investigate whether these formulations could
restore the immunomodulatory role of the gut microbiome. To model gut dysbiosis,
antibiotic-treated mice were subsequently infected with IAV [201]. This
intervention led to significantly reduced CD4 and CD8 T cell responses as well as
lower levels of virus-specific antibody levels, resulting in elevated IAV titres
9 days post infections [201]. Specifically, oral treatment of neomycin led to a
significant reduction in the gut microbial biomass which was associated with the
absence of CD8 T cell responses in the lung [201]. Murine maternal
supplementation with Gram positive probiotics such as L. rhamnosus and
B. animalis subsp. lactis resulted in increased IAV clearance in
neonates and improved survival [202]. Perinatal exposure to the two probiotics
resulted in increased IAV antigen-experienced IFN-
Oral probiotic administration in humans has also demonstrated comparable efficacy, suggesting its potential to modulate immune responses similarly to the findings observed in animal models. A recent randomised controlled trial (RCT) by Lau et al. [204] studied whether the treatment with a symbiotic preparation consisting of B. adolescentis, B. bifidum, B. longum and galacto-oligosaccharides, xylo-oligosaccharides and resistant dextrin (SIM01) were able to promote growth of these strains in the gut and also increase SCFA levels to alleviate the symptoms of PACS. Patients reported that clinical symptoms such as fatigue, memory loss and difficulty in concentration were reduced without associated adverse events [204]. SIM01 administration was able to increase the bacterial diversity and richness compared to the placebo group at 6 months [204]. The targeted Bifidobacterium genus had significant increases in relative abundance in the SIM01 group along with Roseburia intestinalis, R. hominis, F. prausnitzii and A. munciniphila which led to consequent increases in SCFA production [204]. Oral LGG treatment in participants who had household-exposure to someone with confirmed COVID-19 reduced the risk of developing COVID-19 symptoms [162]. However, the incidence of COVID-19 diagnosis had no difference to placebo groups [162]. In a RCT involving children with upper respiratory tract infection (viral or bacterial not specified), oral administration of a commercially available mixture of B. breve M-16V, B. lactis HN019, L. rhamnosus HN001 reduced fever duration by 2 days compared to the placebo group [205]. Adverse effects reported included diarrhoea, constipation and abdominal pain but its distribution was not significant between the treatment and placebo groups indicating the safety of oral probiotics in adolescents [205]. These results strongly support the oral administration of probiotics to alleviate the duration and or the symptoms of respiratory infections.
Although the mechanism of these oral-to-lung protective effects remain uncertain, they are attributed to the complex physiological and immunological network between the gastrointestinal and respiratory tracts. Translational interactions between gut-associated immune cells trigger immunomodulation indirectly in the lungs, resulting in local anti-inflammatory and antimicrobial responses.
Mechanistically intestinal-derived immune factors exert profound effects on pulmonary immunity through both direct and indirect pathways. Direct communication occurs via the migration of activated immune cells through the blood and lymphatic systems, enabling effector populations primed in the intestines to influence distal sites such as the lungs [206]. Exemplified by T helper 17 (Th17) cells and regulatory T (Treg) cells that are elaborated in the intestinal environment that can then traffic to the pulmonary mucosa, where modulation of the local immune responses occur [206]. Furthermore, the indirect communication is reported to be mediated by systemically circulated microbial components and metabolites such as short-chain fatty acids (e.g., SCFAs) [206, 207]. Among these, SCFAs produced by gut commensal bacterial species (e.g., the Clostridia cluster IV and XIVa group of gut commensals produce butyrate) and lipopolysaccharides (LPS) derived from Gram-negative bacteria (e.g., Escherichia coli) are key modulators of the gut immune tone. SCFAs influence the differentiation and function of immune cells, including those within the bone marrow, thereby shaping both innate and adaptive responses. LPS, conversely, can act as a potent immunostimulant, priming inflammatory pathways that extend beyond the gut and in this instance to the lungs. This combined effort exemplifies the bidirectional nature of the gut–lung axis, whereby intestinal immune education and elaboration, and microbial metabolites orchestrate pulmonary homeostasis and the resolution of airway inflammatory responses [207].
However, using oral probiotics for respiratory conditions entails reliance on the yet unclear workings of the gut-lung axis. Furthermore, probiotics transiting through the gastrointestinal tract must survive in, or protected from, stomach acid to ensure the right dose can be delivered. Gastric food content also affects residence time of probiotics in the upper gastrointestinal tract. These factors may render the efficacy of oral probiotics slow and/or variable. A more efficient and rapid delivery of a probiotic to the lungs can be envisaged by the direct administration of a probiotic into the respiratory tract and to the lungs to induce local immunomodulation.
While inhaled probiotics have not been investigated as extensively as oral formulations, research has shown that administering probiotics directly to the respiratory tract before pathogen exposure can have beneficial effects. However, these findings come solely from animal studies. Intratracheally and intranasally instilled probiotics in mice protected against subsequent acute bacterial and viral infections, reduced pathogen load, and increased survival. Murine models were the most prevalent and intranasal instillation of the Lactobacillus species were common. Table 1 (Ref. [195, 208, 209, 210, 211, 212]) highlights the anti-inflammatory and antibacterial effects of live and heat-killed probiotics.
| Pathogen | Probiotics/Postbiotics | Route and Murine Model | Main outcomes | Reference |
| P. aeruginosa PAO1 | Mixture of live | Intratracheal C57BL/6 | ↓ bacterial load | [195] |
| L. fermentum | ↓ IL-6 | |||
| L. zeae | ↓ TNF- |
|||
| L. paracasei | ||||
| at dose 107 CFU/mouse | ||||
| at 18 h or 42 h and 18 h before infection. | ||||
| P. aeruginosa PAO1 | Mixture of live | Intranasal C57BL/6J | Improved survival | [208] |
| L. rhamnosus | ↓ bacterial load | |||
| L. fermentum and | ↓ pro-inflammatory | |||
| L. fermentum or | cytokines | |||
| Mixture of live | ↓ chemokines | |||
| L. paracasei | ↑ anti-inflammatory | |||
| L. salivarius | cytokine IL-10 in | |||
| L. brevis | BALF | |||
| at unspecified dose 18 h before infection. | ||||
| Influenza virus H1N1 A/PR8/34 | Heat-killed | Intranasal BALB/c | Improved survival and respiratory immune responses | [209] |
| L. rhamnosus GG | ||||
| unspecified dose to mice for | ↓ infection symptoms | |||
| 72 h consecutively before infection. | ||||
| Influenza virus H1N1 A/PR8/34 | Live or heat-killed | Intranasal BALB/c | Improved survival | [210] |
| L. rhamnosus | ↓ viral load | |||
| at dose 108 CFU/mouse/day for | ↓ tissue damage | |||
| 48 h consecutively before infection. | ↓ oedema in lungs | |||
| ↓ pro-inflammatory | ||||
| cytokines | ||||
| ↓ chemokines | ||||
| ↑ anti-inflammatory | ||||
| cytokine IL-10 in | ||||
| BALF | ||||
| Improved antiviral response | ||||
| Influenza virus H1N1 A/PR8/34 or H3N2 | Live | Intranasal BALB/c | ↓ weight loss and | [211] |
| A/Philippines/82 | L. plantarum | ↓ lung viral load | ||
| at dose 107 or 108 CFU/mouse for | ↓ IL-6 and TNF- | |||
| 96 h before infection or | BALF | |||
| 108–109 CFU/mouse at infection. | Modulated dendritic cells and macrophages | |||
| Influenza virus | Heat-killed | Intranasal | IAV | [212] |
| H1N1 A/PR/8 | Bacillus subtilis DSM 3244 | C57BL/6 (H1N1) BALB/c (RSV) K18-hACE2 | 100% survival when challenged with IAV | |
| RSV | at dose 1.5 × 109 CFU/mouse for 21, 14, 7 days before infection. | ↑ neutrophils | ||
| SARS-CoV-2 (Omicron variant BA.1) | T cells recruitment | |||
| ↑ IFN |
||||
| IL-1 |
||||
| CCL5 | CXCL10 | ||||
| in BALF | ||||
| RSV | ||||
| No obvious weight loss and complete clearance of RSV viral load | ||||
| ↑ CD4 | CD8 T cells | ||||
| SARS-CoV-2 | ||||
| Significantly decreased SARS-CoV-2 viral load by day 4 post infection | ||||
| ↑ serum IgG | ||||
| pulmonary IgA |
Note: BALF, Bronchoalveolar Lavage Fluid; CCL5, CC motif Chemokine receptor 5;
CD4, Cluster of Differentiation 4; CD8, Cluster of Differentiation 8; CXCL10, CXC
motif Chemokine Ligand 10; CFU, Colony Forming Units; IAV, Influenza A Virus; IL,
Interleukin; IFN, Interferon; RSV, Respiratory Syncytial Virus; SARS-CoV-2,
Severe Acute Respiratory Syndrome Coronavirus 2; TNF, Tumour Necrosis Factor;
The idea of combining nanomedicine technologies with dry powder inhalers into a loosely bound submicron sized microparticle (i.e., usually ~1 to 5 µm) is a feasible posit [213]. The idea is that on inhalation the particle agglomerates and disperses into the nanoparticle constituents in the airway surface liquid, enabling deep lung deposition. Delivering and accumulating the nanoscale drug carriers in the lung tissue would constitute a strategy that bridges the aerodynamic requirements for inhalation with the functional advantages of nanoparticles, given that otherwise the drug would be exhaled or poorly aerosolized [213].
There are a number of advantages with nanoparticles such as (i) enhanced solubility and dissolution with increases in particle surface area, carrying poorly soluble payloads in amorphous and solubilized matrices that can accelerate dissolution at the epithelial surface [214]. It is also reported that nanoparticles (ii) exhibit enhanced stability in dry form. This is especially important stabilizing labile molecules (e.g., peptides, proteins, nucleic acids), protecting the payload from hydrolysis and aggregation storage especially when compare to liquid aerosols [215]. Furthermore, nanoparticles as nanocarriers of lipid, polymeric or inorganic molecules can (iii) be target tissue specific for delivery with controlled release of the drug [214, 215]. These functional gains have been demonstrated in preclinical and formulations investigations of inhalable nano-powders [214].
Direct respiratory delivery of probiotics is a promising scientific pursuit. However, it remains under-studied in humans. Advancing this research posit requires further rigorous safety testing, sensitive lung microbiome methods, and clinical studies that link local microbiome changes to beneficial immune and clinical outcomes. The challenges encountered raise numerous concerns around (i) low biomass and contamination risk that makes reliable lung microbiome profiling difficult; technical problems in distinguishing resident bacteria from transient colonizers with strict sampling and negative control requisites; (ii) further safety barriers relative to the introduction of live microbes into already inflamed and immunocompromised airways that would raise regulatory and ethical concerns; and (iii) formulation and delivery loopholes that relate to include keeping probiotic strains viable in aerosols, with the avoidance of bronchospasm that then achieves deposition of the drug payload in target airway regions.
At present, only a limited number of studies have explored the respiratory route
for probiotic administration in humans as outlined in Table 2 (Ref. [216, 217, 218, 219]). A notable study used a nasal spray of containing 5 billion of
Bacillus subtilis ANA4 and B. clausii ANA39 suspended in 0.9%
NaCl (LiveSpo Navax) was developed to be tested on children with RSV [216]. The
children did not experience any negative side effects upon spraying the LiveSpo
Navax spray such as choking, signs of nasal mucosa irritation, symptoms of local
bacterial infection, vomiting or diarrhoea indicating the safety and suitability
of the dosage form [216]. In terms of efficacy, patients using the LiveSpo Navax
no longer had runny noses by day 6 compared to 17.5% of patients in the control
group [216]. In a Phase 2 trial with children with severe pneumonia due to RSV
and bacterial co-infections, significant reductions in RSV and bacterial
co-infections were noted at day 3 of LiveSpo Navax treatment [217]. Similar to
their previously reported study, these reductions were accompanied with notable
decreases in pro-inflammatory cytokines such as IL-6, IL-8 and TNF-
| Disease | Treatment | Administration | Main outcomes | Reported adverse events | Reference |
| Route | |||||
| Patient profile | |||||
| n = number | |||||
| RSV | LiveSpo Navax: | Intranasal (spray) | Day 3: | No abnormal changes to | [216] |
| Bacillus subtilis ANA4 | Children | ↓ percentage of patients with moist rales, fast pulse in treatment group | —breath rate | ||
| B. clausii ANA39 | Treatment Gp | —heart rate | |||
| 5 × 109 CFU/dose in | (n = 46) | RSV viral load | —body temperature | ||
| 0.9% NaCl | Placebo Gp | ↓ |
—No choking upon | ||
| Dose: t.i.d./6 days | (n = 40) | ↓ IL-6 | IL-8 | TNF- |
spraying | ||
| Day 6: No patients in treatment group with runny nose | —No signs of nasal | ||||
| mucosa irritation | |||||
| Treatment Gp recovered in 4 days compared to 5 days in Placebo Gp | —No local bacterial | ||||
| infection | |||||
| —No emesis | |||||
| —No diarrhoea | |||||
| Pneumonia due to RSV bacterial co-infection | LiveSpo Navax | Intranasal (spray) | Day 3: | N/A | [217] |
| Dose frequency not specified | Children | ↓ RSV and bacterial co-infection | (Abstract only) | ||
| Total recruited | ↓ H. influenzae | ||||
| (n = 120) | S. pneumoniae | ||||
| ↓ IL-6 | IL-8 | TNF- |
|||||
| ↑ IgA | |||||
| Restoration of nasal microbiota | |||||
| ↑ Corynebacterium | |||||
| Bacillus | |||||
| ↓ Streptococcus | |||||
| Haemophilus | |||||
| Moraxella | |||||
| Allergic rhinitis (Positive skin prick test to birch or grass pollen with wheal diameter |
L. rhamnosus SP1 | Intranasal (spray) | No significant differences in Mini-RQLQ | TNSS | PNIF | —No severe adverse events reported | [218] |
| L. paracasei 101/37 Lactococcus | Adults Treatment Gp (n = 13) | Treatment group: | |||
| lactis L1A | Placebo | No significant differences in FeNo | specific IgE and total IgE | —2 patients: burning sensation | ||
| 1.9 × 1010 CFU/dose | Gp (n = 11) | —1 patient: itching | |||
| Dose: b.i.d./21 days | ↑ TNF- |
—1 patient: pain upon spraying | |||
| MIP-1 |
|||||
| IL-8 | IL-10 | |||||
| Chronic rhinosinusitis with previous endoscopic sinus surgery | Lactococcus lactis W136 | Intranasal sinus (irrigation in 0.9% saline) | SNSS: | Well tolerated | [219] |
| 1.2 × 109 CFU/dose | progressive improvement over treatment period | —No acute infections during Treatment | |||
| Dose: b.i.d./14 days | Adults Treatment | Greatest improvement in nasal congestion | Reported | ||
| Gp (n = 24) | post-nasal drip | —headaches | |||
| No Placebo Gp | need to blow nose | —migraines | |||
| SNOT-22: | —nasal congestion | ||||
| ↑ in score over treatment | —dental infection | ||||
| period | —throat pain | ||||
| POSE: | —cold sore | ||||
| Improvement aligned with SNSS and SNOT-22 | —gastroenteritis | ||||
| UPSIT-40: | —nasal allergy | ||||
| Stable no deterioration | —shoulder pain | ||||
| Nasal microbiome: | but… | ||||
| no change in a-diversity | did not differentiate between before trial and during trial | ||||
| ↓ Dolosigranulum | |||||
| pigrum |
Note: Gp, Group; b.i.d., twice per day; t.i.d., three times per day; SNSS, Total
Sino Nasal Symptom Score; SNOT 22, Sino-Nasal Outcome Test-22; POSE,
Perioperative Operative Sinus Endoscopy Score; UPSIT-40, University Pennsylvania
Smell Identification Test-40;
A recent review [221] has narrated a synthesis of the published research evidence based on the nature of the studies namely, clinical studies or human samples, in vivo or animal models, in situ, in vitro and/or in silico concluding that probiotics may have adverse effects. What is of importance is the administration of a probiotic formulation to an immune compromised patient with a chest infection and the risks that probiotic treatments may ensue. In such vulnerable patients, the modulation of the lung microbiota with probiotic formulations can present several potential risks to health and limitations to treatment efficacy. In particular opportunistic infections, excessive immune stimulation, transfer of antibiotic resistance genes, and challenges in effective delivery to the lungs [222]. Notwithstanding expert opinions do not recommend stopping research into probiotics, rather, there is encouragement for rigorous, targeted research to better understand probiotic specific effects, safety, and mechanisms of action [223]. Specifically current ideas focus on the effects of probiotics that are highly specific to strains and the individuals administered to [223, 224].
Prebiotics are “selectively fermented ingredient that allows specific changes
both in the composition and/or activity in the gastrointestinal microflora that
confers benefits upon host well-being and health” [225]. Common examples
approved by the United States Food and Drug Administration include alginate and
inulin [226] which are used to drug delivery. Alginate is frequently used for
stabilising and enhancing pulmonary drug delivery [227, 228, 229]. Encapsulation with
alginate is a promising approach as it does not compromise the antimicrobial
effect of antibiotics and is not toxic to lung cells [227]. More recently, dry
powder alginate oligosaccharide has been tested as a drug in patients with cystic
fibrosis [229] as in vitro studies demonstrated the potential to modify
the viscoelastic properties of CF sputum [230]. Though its desired effects were
not observed in the Phase 2 clinical trial, the trial demonstrated the safety of
inhaling alginate oligosaccharides [229]. Inulin is also popular in drug
formulation and delivery as it stabilises and protects drugs against the acidic
environment of the stomach, making them especially useful for oral delivery
[231, 232, 233]. Interestingly, inulin-enriched diet for 15 days prior to lethal
intranasal P. aeruginosa challenge increase the survival of mice by 33%
through
Postbiotics are not live bacteria, rather compounds produced by microorganisms during a growth phase or the fermentation of dietary fiber, the indigestible part of plant-based foods (e.g., vegetables, legumes) [239]. Furthermore, postbiotics are “formulations of inanimate microorganisms and/or their components that confers a health benefit on the host” [240]. These compounds include small molecules, peptides, enzymes, cell wall fragments, and other microbial products such as SCFAs (e.g., acetate, propionate, butyrate), bacteriocins, exopolysaccharides, enzymes and vitamins produced during microbial fermentation [241]. Moreover, postbiotics can also encompass larger molecules like polysaccharides and proteinaceous factors [241]. Hence heat-killed bacteria shown in Table 1 can be considered postbiotics. Interestingly, heat-killed probiotics were reported to be as good as their live counterpart in that regard [210]. Intranasal live and heat-killed L. rhamnosus CRL1505 were also effective against respiratory syncytial virus infection in BALB/c mice [242]. This phenomenon is well-documented for oral probiotics and is attributed to the immunomodulatory effects of bioactive intracellular or cell membrane components released upon the death of the probiotic [243]. Using non-viable probiotics is clinically safer because it eliminates the risk of live probiotics infecting or causing sepsis in immunocompromised patients. It would also facilitate the manufacture and storage of the formulations because there is no need to maintain a live probiotic load [243]. The immunomodulatory effects of non-viable probiotics administered via the respiratory route remain underexplored, underscoring the need for further investigation in this area.
SCFAs are a central subset of postbiotics and considered to be the most studied entity [241]. SCFAs can also be considered products of probiotic gut metabolism. SCFAs postbiotics have anti-inflammatory, intestinal barrier-supporting and metabolic-modulating effects in the gut [241]. Investigations with SCFAs are reported as safe, well-defined biotherapeutics or as supplements for chronic diseases, intestinal disorders, and immune modulation [241]. Strengthening factors such as standardization of products, doses, and administration without live organisms further improves the knowledge base for clinical practice approval. Reports on how actively mapping which specific postbiotic molecules produce what host response is observed, will correlate how the formulation, the dose, and the delivery route affect outcomes [241].
Although less extensively studied than probiotics in terms of their effects on the lungs, emerging research supports the direct use of postbiotics as a promising approach for managing common respiratory infections [244, 245]. Oral administration of a butyrate releaser, N-(1-carbamoyl-2phenyl-ethyl) butyramide (FBA) had a protective effect against SARS-CoV-2 in humans [246]. It reduced the number of SARS-CoV-2-infected cells and reduced angiotensin-converting enzyme 2 (ACE2) and transmembrane protease serine-2 TMPRSS2 expressions which were related to the pathogenicity of SARS-CoV-2 [246]. Emerging animal evidence from animal studies suggests that butyrate may have a therapeutic potential in treat pulmonary conditions, such as pulmonary fibrosis [247] and COPD [248]. However, pulmonary delivery of butyrate is yet to be explored.
In section 7.1., certain commensal microbiota were shown to be responsible for CD8 T cell regulation in mice [201]. Recently, the impairment in the immunoregulation observed in dysbiotic mice were also evident in infant humans with gut dysbiosis through the disruption of nuclear factor interleukin 3 (NFIL3)-dependent T cell programming [249]. Among the four genera that significantly distinguish healthy from dysbiotic human infant gut microbiota, Bifidobacterium is notable for its metabolite, inosine [249]. Oral administration of B. pseudolongum in mono-associated germ-free infant mice resulted in increased inosine levels in both lungs and faeces which subsequently led to 2-fold increase in effector CD8 T cells in the lungs [249]. Directly treating dysbiotic mice with IAV infection with inosine resulted a higher influenza-specific CD8 T cells levels and increased influenza-specific tissue resident memory CD8 cells [249]. The observed reduction in morbidity among inosine-treated dysbiotic mice suggests that similar immunomodulatory effects may be replicable in human infants, potentially leading to improved health outcomes.
SCFAs were found in millimolar concentrations in the cystic fibrosis airways due to hypoxia [250]. The immune responses vary depending on the type of SCFAs, its concentration and the cell line used in testing [250]. High concentrations of SCFAs (25–50 mM) reduced the levels of inflammatory cytokines such as GM-CSF, IL-6 in A549 and control cystic fibrosis bronchial epithelial (CFBE) cells [250]. In contrast, low acetate levels (0.5–2.5 mM) and IL-8 expression specifically in F508del-CFTR, suggests its potential role in CF airway inflammation [250]. SCFA concentration also influenced the growth of a common CF pathogen, P. aeruginosa [250]. High SCFA levels inhibited the pathogenic growth whilst low levels promoted it under aerobic conditions [250]. However, under microaerobic conditions mimicking the CF lungs environment, SCFAs significantly suppressed P. aeruginosa growth, highlighting their vital role in maintaining lung homeostasis and health [250]. Currently, the application of SCFA-based interventions in human clinical settings for cystic fibrosis patients remain unexplored.
Vancomycin induces gut dysbiosis by reducing the overall abundance and
diversity, especially decreasing the Clostridia class which produces
butyrate [251]. Oral supplementation with a SCFA cocktail consisting of butyrate,
acetate and propionate did not replace what is lost by vancomycin in the gut
[251]. However, it reduced airway allergic inflammation [197] evident through
significantly decreased IgE levels and histopathology during allergen exposure
[251]. Similar anti-inflammatory observations were made with older mice with
acute lung injury [252]. Oral SCFA treatment significantly reduced the pulmonary
IL-6, CXCL1, and GM-CSF levels in old mice [252]. However, the anti-inflammatory
effect was less profound in young mice and in fact IL-12 and IFN-
Interestingly, SCFAs failed to suppress TNF-
In multiple clinical trials, inosine pranobex (IP), a combination of p-acetamindo-benzoate salt of N-N dimethylamine-2-propanol and inosine in 3:1 molar ratio [254], demonstrated safe and efficacious effects in reducing influenza-like symptoms in adults [254, 255]. In the phase 4 trial, though patients in the treatment group had their influenza symptoms ending 2 days earlier than the placebo group, the difference in time to resolution of all flu-related symptoms were not statistically significant [254]. However, for a subgroup of patients under 50 without related ongoing disease and non-obese, the time to symptom resolution was significantly shorter. Similar results were reported in the phase 3 trial that investigated the efficacy of IP against COVID-19 [255]. On day-6 of treatment in COVID-19 patients, patients on IP displayed a higher clinical response (2-point improvement or being asymptomatic on the modified WHO ordinal scale) and higher clinical cure (asymptomatic on the modified WHO ordinal scale) [255]. Both trials reported minimal adverse events indicating the safety of IP [254, 255].
Reports show that microbiota derived compounds from the metabolism of dietary sources such as SCFAs and inositol can regulate host epigenetics that affect DNA methylation and histone acetylation [49, 256]. SCFAs and inositol can also deliver enzymes that directly modify host proteins or trigger signaling pathways that can alter the host’s epigenetic machinery, influencing gene expression related to immune response and metabolism [257].
SCFAs produced by intestinal bacteria (i.e., butyrate, propionate, acetate) can act as inhibitors of class I/II HDACs raising the global histone acetylation and promoting open chromatin at certain target loci [258]. SCFAs can also influence one-carbon metabolism and methyl-donor availability, biochemical reactions that indirectly affect DNA methylation patterns and gene expression that are linked to inflammation and metabolism [258]. In combination with the activity of SCFAs gut bacteria encode methyltransferases that can modify host substrates or when delivered to host cells, alter host methylation environment [259, 260]. Moreover, bacterial methyltransferases can target nucleotides and proteins. Interactions with host chromatin or regulatory proteins can enact stable epigenetic changes that modify host transcriptional responses to infection or gut bacterial colonization [259].
In the gut intestinal pathogens influence host histone-modifying enzymes as a survival mechanism by delivering effectors that inhibit HDACs or alter HAT activity to suppress antimicrobial genes or skew inflammatory programs [261]. This epigenetic targeting pathogen survival mechanism is a documented immune-evasion strategy that can be exploited by both commensal bacteria for homeostasis and pathobionts for persistence [261].
Gut bacteria have also been reported to release outer membrane vesicles and deploy secretion systems specialized to translocate proteins, nucleic acids and small molecules into host cells [259, 261]. The net effect is that these compounds the host cell’s nucleus proximal machinery and signaling centres and there alter the state of chromatin and transcriptional outputs in the recipient cells.
Bacterial epigenetic actions can ensue functional consequences. As in (i) immune modulation where epigenetic reprogramming can dampen or amplify innate and adaptive immune responses that can then affect tolerance, inflammatory dispositions, and susceptibility to infections and auto-immunity. (ii) Metabolic effects that reshape expression of metabolic genes in intestinal epithelia cells, liver and adipose tissue that in concert influence host energy balance and risk of metabolic disease (e.g., diabetes). Microbial driven epigenetic physiological and chemical signals are essential factors during early life and at stages of chronic colonization shifts in the gut as they can influence epithelial differentiation, gut barrier integrity and long-term tissue homeostasis [258, 259, 260].
Given the constant migration of bacteria between the upper and lower respiratory tract sites, the lung microbiota is in a state of continuous flux. There is almost a consensus that by examining the diversity of the lung microbiome, new targets for therapeutic interventions can be identified. The gut-lung axis mediated by microbial components and metabolites such as postbiotics like SCFAs links intestinal health to respiratory health. Reviewing the clinical evidence of probiotics, fecal microbiota transplantation (FMT), and microbial metabolites modulating lung immunity and pathology in animal models and early clinical work, has provided technical challenges that help focus research on the lung microbiome. Given the complex nature of the interactions that exist between the microorganisms that inhabit the lung, host immunological defences and the environment, provide insight into the close physiological and pathological influences between the intestines and the lungs [74]. The reliance encountered is mainly associated with what the nature of the host-microbe cross-talk entails [74]. The cross-talk between intestinal and lung resident commensals necessitates an exchange of components and metabolites through the systemic circulation that indeed can then contribute significantly to the health of both sites [262].
The application of probiotics to influence the intestinal microbiome to manage disease processes has a long history [263], with formulations posited to have mutually beneficial relationships with humans pivotal in maintaining human health [263]. Consequently, members from the Lactobacillus genera have been advanced as natural immunobiotics [264]. Such immune-active actions have originated from the administration of Lactobacillus species in animal models and clinical trials that have reported alleviation of the symptoms of respiratory diseases such as asthma, respiratory tract infections, lung cancer and cystic fibrosis [264, 265]. Of further interest has been the role that Lactobacillus species may have in regulating respiratory mucosal immunity. Murine models have shown that when the lungs were infected with S. pneumoniae, nasal administration of heat-killed L. casei increased resistance to infection, decreased pulmonary bacteria load, and increased survival [266].
The crosstalk between the gut and lung teaches that with microbiome modifications progressed with FMT interventions it may be possible to influence the condition of the lungs. In emphysema-induced mice, FMT coupled with diet modifications resulted in reduced lung inflammation, limiting alveolar destruction [238]. Interestingly FMT treatments resulted in increases in the relative abundance of bacterial families such as Bacteroidaceae, Lachnospiraceae, and Ruminococcus in the gut that are involved in the production and metabolism of SCFAs [238]. Improving the intestinal microbiome improved the tone of the immune system and reduced inflammation in the lungs.
Oral administration of SCFAs also decreased inflammation and alveolar destruction in emphysema mice [238].
Massalha and colleagues [267] have investigated FMT for improving lung immunity with checkpoint inhibitors. Currently, a Phase 2 clinical trial is investigating whether FMT can be used as an adjuvant therapy for metastatic lung cancer and also analyse its effect on the gut microbiome [268].
FMT using post-COVID microbiota obtained from patients was performed on mice to investigate whether post-COVID symptoms could be induced [269]. Lung histology revealed that post-COVID FMT resulted in an increase in inflammatory cells, and an increase in a-smooth muscle actin expression, indicating physiological dysfunction [269]. Gut microbiota modifications affect the lungs but whether it can also affect the lung microbiome has not been studied. The difficulty in studying this possible change exists as the biomass in the lungs is significantly smaller than that of the gut.
Exploring direct lung–targeted approaches with inhaled probiotics, prebiotics, synbiotics or postbiotics share the same goals as gut-delivered strategies. The aim being to modulate the local microbiota and host immunity. Differences in delivery, dosing, safety and tolerability present technical challenges for efficacious treatments. Hence, drug delivery for lung conditions such as asthma and COPD is achieved by inhalation devices such as nebulisers, dry powder inhalers, and pressurised metered dose inhalers. Nebulisers are an inexpensive and easy device to deliver drugs into the airways. A recent study explored the possibility of delivering LGG via a vibrating mesh and jet nebuliser [270]. Although only 10% of the loaded dose was deliverable to the lungs [270], the advantage of a nebuliser is that it can deliver doses larger than those from dry powder inhalers and pressurised metered dose inhalers. Hence the lower delivery efficiency can be overcome by loading a higher dose to achieve the desired dose in the lungs.
Additionally, probiotic delivery to the lungs has employed dry powder formulations. The food industry has progressed dry processed powdered probiotics in a lyophilized form, that could be formulated to be inhalable. Numerous in vitro studies have reported promising inhalable results with spray dried powders of LGG, L. plantarum and L. acidophilus. Importantly also showing that they were individually safe against human lung-derived Calu-3 and A549 epithelial cell lines [271]. Moreover, L. plantarum in particular displayed antimicrobial properties by eradicating a common respiratory pathogen P. aeruginosa [271]. Similarly, a spray freeze dried L. rhamnosus GG powder was non-toxic against A549 human adenocarcinoma cell line whilst inhibiting P. aeruginosa growth [272].
An inhalable spray dried powder containing a blend of live L. plantarum BAA-793, L. acidophilus 4356, and LGG improved lung structure and function in murine models of COPD and bronchopulmonary dysplasia [273]. Nasal sprays is an alternative and promising drug delivery platform to the upper respiratory tract. A nasal spray containing Bacillus spores were successful in reducing RSV symptoms in children whilst demonstrating safety [216].
Currently, there is a paucity of human clinical studies investigating what constitutes the potential of direct respiratory delivery of probiotics to the respiratory tract. To therapeutically target the respiratory microbiome a precise requisite understanding of the respiratory microbiome is essential, the effects that clinical interventions may have on the equilibrium of the respiratory cohort of bacteria and what immunological effects the lung microbiome provides [274].
Respiratory infections of viral or bacterial origin are among the most common infectious diseases. The gut microbiota plays an important role in dictating the severity of these infections. The literature points towards a healthy gut equalling a healthy lung. Oral probiotics and postbiotics may be used to indirectly treat lung conditions.
The last two decades has been witness to an exponential increase in support and a rapid escalation in interest for the administration of probiotics, prebiotics, postbiotics and synbiotic (i.e., a probiotic + prebiotic) formulations that can act as mediators in health and disease [224]. The future challenges are extensive and complex. A recent report has advanced provisions into how industry and academia must adapt to probiotic research [224]. The posits are centred on how to maximize treatment success through targeted applications of probiotic strains such as narrated herewith with the clinical translation of inhaled probiotics to manage bacterial pathobiont and viral infections in the lungs. This being dependent on individual probiotic strain capabilities as well as the application of multiple advanced analytical technologies that can then further elucidate mechanistic understanding and fast-track the science of the microbiome in organ specific sites (e.g., the lungs).
The scientific pursuit of innovative treatments for lung diseases via the incorporation of nanoparticle technology of inhaled compounds is a plausible posit. Given that probiotics and postbiotics have shown high level of safety in the gut illustrates translational interest, albeit, without direct inhaled delivery. Direct pulmonary delivery would be the most efficient and effective delivery platform for postbiotic compounds and heat-killed probiotic bacteria to further investigate efficacy and safety in the management of lung infections.
Conceptualization and design of the review framework, LV, AB, PK, H-KC ; interpretation of findings across studies, LV, AB, PK, H-KC; creating figures or tables that summarise key concepts or evidence, LV, AB, PK, H-KC; writing—original draft preparation, LV, AB, PK; writing—review and editing, LV, AB, PK, H-KC. All authors have read and agreed to the published version of the manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
Figs. 1,2 were adapted from BioRender templates namely “Gut-Lung Axis” and “Bacterial Epigenetics” with authors of the manuscript adding referenced content [BioRender.com (2020)].
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.


