1 The Affiliated Eye Hospital, Nanjing Medical University, 210029 Nanjing, Jiangsu, China
2 The Fourth School of Clinical Medicine, Nanjing Medical University, 211166 Nanjing, Jiangsu, China
†These authors contributed equally.
Abstract
Dendrobium spp. has traditionally been used to improve visual function, and related formulations, such as Dendrobium Glow in dark pills, are currently used in the management of dry eye disease (DED). This study investigated the positive effects and underlying mechanisms of 3-O-methylgigantol, an active compound from Dendrobium spp., against DED using network pharmacology and in vitro experimental validation.
Active compounds and targets were identified through database screening, and network analysis was used to identify the key compounds and targets. Molecular docking and dynamic simulations were performed to verify the binding of 3-O-methylgigantol to the epidermal growth factor receptor (EGFR). An in vitro DED model was established using hyperosmotic sodium chloride (550 mOsm) in human corneal epithelial cells (HCECs). The effects of 3-O-methylgigantol, with or without the EGFR inhibitor (erlotinib), on cell viability, apoptosis, reactive oxygen species, and inflammatory cytokines were assessed.
Network pharmacology predicted 3-O-methylgigantol as a key active compound and EGFR as a core target. Molecular simulations confirmed stable binding. In vitro experiments showed that 3-O-methylgigantol significantly increased HCEC viability and reduced apoptosis, reactive oxygen species accumulation, and inflammatory cytokine release under hyperosmotic conditions. The EGFR inhibitor erlotinib attenuated these protective effects.
3-O-methylgigantol, an active compound from Dendrobium spp., alleviates hyperosmolarity-induced corneal epithelial cell damage, oxidative stress, and inflammation by activating the EGFR signaling pathway. This compound may represent a potential therapeutic candidate for DED management, demonstrating its efficacy in restoring tear film homeostasis in in vitro models.
Graphical Abstract
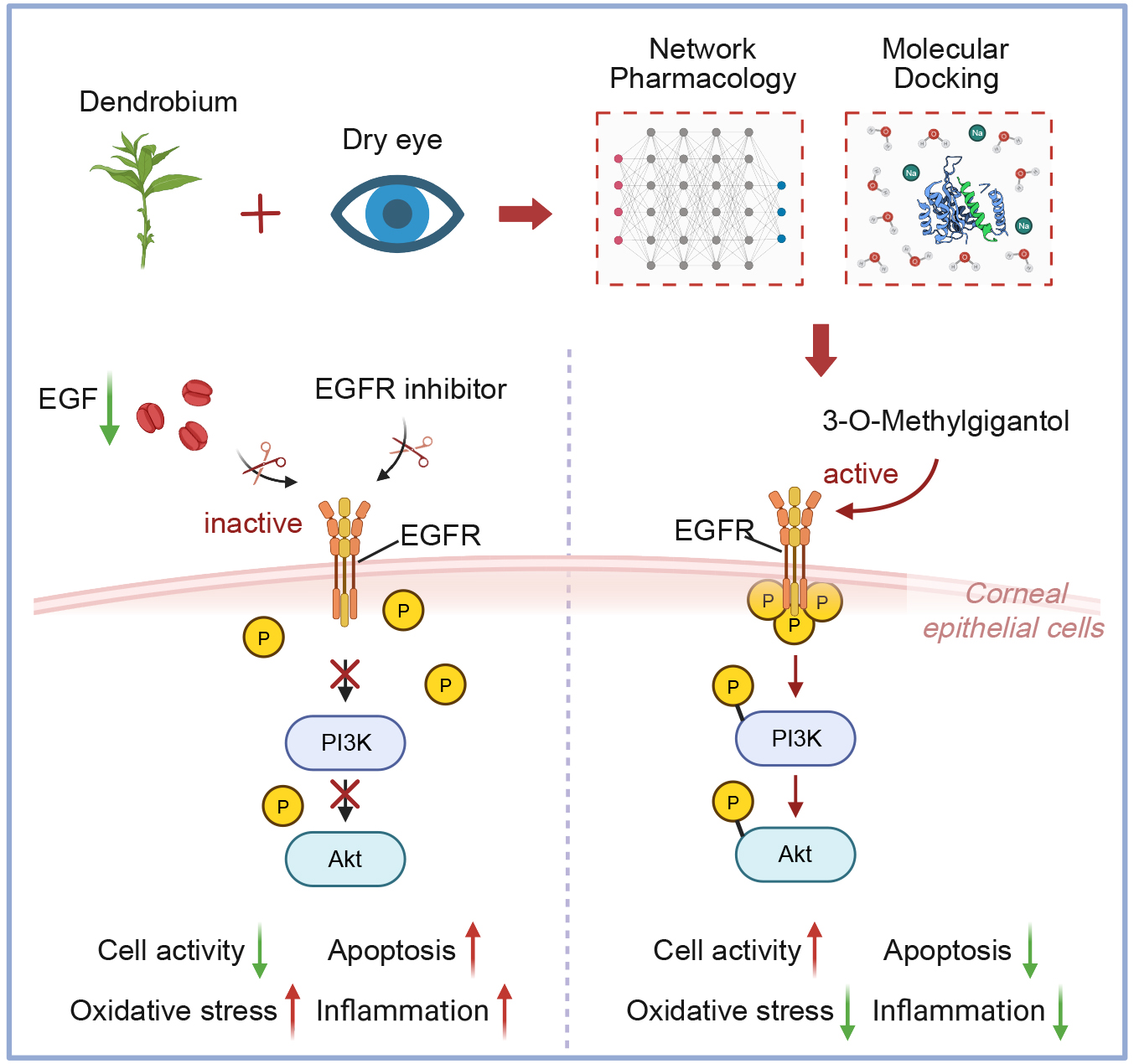
Keywords
- Dendrobium spp.
- traditional Chinese medicine
- 3-O-methylgigantol
- dry eye
- network pharmacology
Dry eye disease (DED) is a multifactorial ocular surface disorder primarily characterized by the loss of tear film homeostasis. It is accompanied by a range of ocular symptoms, including dryness, burning, stinging, foreign body sensation, fluctuating vision, and photophobia, among others [1]. Severe cases may result in loss [2]. With the shift toward modern lifestyles, including the prolonged use of video display terminals and smartphones, the prevalence of DED has increased, posing a growing public health concern and economic burden [3].
Hyperosmolarity of the tear film and ocular surface inflammation are considered core pathological mechanisms underlying DED [3, 4]. Hyperosmolarity can directly damage epithelial cells on the ocular surface or indirectly cause harm by inducing inflammatory responses. This inflammatory response involves immune cell infiltration and release of proinflammatory cytokines, chemokines, and matrix metalloproteinases (MMPs), particularly MMP-9 [5]. These factors act synergistically on the ocular surface and the lacrimal glands, leading to tissue damage and functional impairment [6]. Damage and inflammation mutually reinforce each other, creating a “vicious cycle” that results in a chronic progressive disease course [7]. Current therapeutic approaches for DED aim to alleviate symptoms, restore ocular surface homeostasis, and interrupt this vicious cycle [8, 9]. Artificial tears typically provide only temporary relief and have low bioavailability [10], resulting in limited efficacy for moderate-to-severe DED. Anti-inflammatory drugs such as cyclosporine and corticosteroids may cause side effects, including increased intraocular pressure and corneal toxicity, with long-term use [11, 12]. Emerging therapies, such as secretagogues [13] and local androgens [14, 15], are still under investigation [16]. Therefore, developing new safe and effective treatments that target multiple pathological mechanisms, particularly active compounds derived from natural products, has significant clinical potential.
Dendrobium (Dendrobium spp.) refers to a group of plants in the Orchidaceae family with medicinal value [17]. It was first documented in *Shennong Bencao Jing* (The Classic of Herbal Medicine), in which it was classified as a superior herb. According to Traditional Chinese Medicine, Dendrobium is believed to nourish the yin, clear heat, benefit the stomach, generate fluids, and improve vision [18]. It is commonly used to treat conditions related to “yin deficiency”, such as heat disorders, dry mouth and thirst, insufficient stomach yin, and blurred vision. Notably, traditional uses of Dendrobium specifically highlight its “vision-enhancing” or eyesight-improving effects. Among its formulations, Dendrobium Glow in dark pills is widely used to treat DED [19]. Modern phytochemical studies have revealed that Dendrobium contains a variety of bioactive compounds, including polysaccharides, alkaloids, bibenzyls [20], phenanthrenes [21], sesquiterpenes, flavonoids, and phenolic compounds [22]. These components contribute to Dendrobium’s broad pharmacological activities and may modulate multiple pathogenic pathways [23, 24], showing promise as potential therapeutic agents for DED.
Network pharmacology integrates high-throughput omics data, bioinformatics, and network analysis techniques. Its core concept is to analyze the complex interactions between drugs, especially multicomponent drugs, and the body or diseases at the system level within the framework of biological networks. This approach is particularly well-suited for studying traditional Chinese medicine (TCM) formulas, which often involve multiple components, targets, and pathways [25]. The primary objective of this study was to combine network pharmacology and in vitro modeling to elucidate the tear film–stabilizing effects of 3-O-methylgigantol, a key phytochemical found in Dendrobium, for the treatment of DED.
The Encyclopedia of Traditional Chinese Medicine (ETCM) database [26] (http://www.tcmip.cn/ETCM/), HERB database [27] (http://herb.ac.cn/), and relevant literature [28, 29] were used to identify the active components of Dendrobium. The structures and SMILES identifiers were retrieved from the PubChem database [30] (https://pubchem.ncbi.nlm.nih.gov/). Active compounds were further filtered using the SwissADME database [31] (https://www.swissadme.ch/) by applying the criteria of high gastrointestinal absorption (GI absorption) and drug-likeness, defined by satisfying at least two of the five established rules (Lipinski, Ghose, Veber, Egan, and Muegge) to obtain relevant pharmacologically active compounds [32, 33].
The potential Dendrobium bioactive compound targets were identified through consensus predictions across multiple databases, including SwissTargetPrediction [34] (https://www.swisstargetprediction.ch/), TargetNet [35] (http://targetnet.scbdd.com/), and Super-PRED [36] (https://prediction.charite.de/). Targets with probability scores exceeding 50% were further validated using the UniProt database [37] (https://www.uniprot.org/), retaining only human protein-coding genes. After deduplication, a curated set of potential Dendrobium targets was established for subsequent mechanistic analyses.
Relevant targets were obtained from the GeneCards database [38]
(https://www.genecards.org/Search/) by searching for the keywords “Dry Eye” and
“keratoconjunctivitis sicca”. Genes with a score of
A high-confidence interaction score threshold (“High Confidence
We used the Enrichr platform (https://maayanlab.cloud/Enrichr/) [39] to perform a biological function analysis of the targets involved in Dendrobium treatment of DED using Gene Ontology Biological Process (GO_BP) terms. Pathway analysis was conducted using the MSigDB Hallmark gene sets.
Molecular docking between the protein and small-molecule compounds was performed
using the Discovery Studio software (version 2019; Dassault Systèmes Biovia,
San Diego, CA, USA). The protein-ligand complex then underwent a 100-nanosecond
molecular dynamics simulation using GROMACS 2022. CHARMM36 and GAFF2 force fields
were used to parameterize the protein and ligand, respectively. The system was
solvated with explicit TIP3P water molecules in a cubic periodic box with a 1.2
nm boundary, using the Particle Mesh Ewald method for long-range electrostatics
and the Verlet cutoff scheme. Energy minimization was conducted prior to the
production runs under NPT ensemble conditions. The system was equilibrated with
100,000 steps each of NVT and NPT simulations (
SV40-immortalized human corneal epithelial cells (HCECs, obtained from ATCC CRL-11135) were cultured in DMEM supplemented with 10% FBS (AusGeneX, Molendinar, QLD, Australia; Lot No. FBS500-S) and 1% penicillin/streptomycin (Thermo Fisher Scientific, Inc., Waltham, MA, USA; Cat No. 15140-122) at 37 °C under 5% CO2. This cell line was validated using STR profiling and tested negative for Mycoplasma sp. To simulate hyperosmotic conditions, cells were treated with NaCl to achieve an osmolarity between 312 and 550 mOsm. In some experiments, cells were incubated with 3-O-methylgigantol (0–10 µM) (EvitaChem, Linyi, Shandong, China; Cat No. EVT-8342849) and Erlotinib (0–10 µM) (Selleck Chemicals, Houston, TX, USA; Cat No. S7786).
Cell viability was measured using a CCK-8 kit (Biosharp Life Sciences, Hefei,
Anhui, China; Cat No. BS350A). Cells were seeded at 1
Intracellular ROS levels were measured using 2′,7′-dichlorofluorescin diacetate (DCFH-DA) (Beyotime Institute of Biotechnology, Shanghai, China; Cat No. S0035S). The cells were incubated with 10 µM DCFH-DA at 37 °C for 30 min and observed under a Varioskan LUX microplate reader (Thermo Fisher Scientific, Inc., Waltham, MA, USA). The fluorescence intensity of each cell type was then measured.
Apoptosis was assessed using calcein-AM and propidium iodide (PI) staining. The calcein-AM (Aladdin Biochemical Technology Co., Ltd., Shanghai, China; Cat No. C273362) stock solution contained 1 mM calcein-AM in dimethyl sulfoxide (DMSO). The stock solution was diluted to 1–50 µM with PBS. The stock solution of PI (Solarbio Science & Technology Co., Ltd., Beijing, China; Cat No. C0080) was diluted with PBS to 10–50 µM. The cells were incubated with calcein-AM and PI at room temperature for 15 min and observed under a fluorescence microscope. Calcein-positive cells were considered live, and PI-positive cells were considered apoptotic.
The concentrations of pro-inflammatory cytokines, including interleukin-6
(IL-6), interleukin-1 beta (IL-1
Biotinylation of 3-O-methylgigantol was achieved using a two-step click chemistry strategy. First, the alkynyl intermediate was prepared by dissolving 3-O-methylgigantol (1.0 eq) in anhydrous acetone, followed by the addition of anhydrous K2CO3 (2.0 eq) as a base. Propargyl bromide (1.2 eq) was added dropwise with stirring, and the mixture was refluxed for 2–4 h. Upon completion, the inorganic salts were filtered, and the filtrate was concentrated under reduced pressure and purified using silica gel column chromatography to yield alkynyl-3-O-methylgigantol. Subsequently, copper-catalyzed azide-alkyne cycloaddition (CuAAC) was performed by co-dissolving the obtained alkynyl intermediate and biotin-azide (1.1 eq) in a DMSO/water mixture (1:1, v/v). CuSO4 (0.1 eq) and sodium ascorbate (0.5 eq) aqueous solutions were added sequentially to generate the catalytic Cu(I) species in situ. The reaction mixture was stirred at room temperature in the dark for 1–2 hours. After confirming reaction completion by TLC, the mixture was extracted with ethyl acetate. The crude product was purified by preparative HPLC or column chromatography, then dried under vacuum to yield the final 3-O-methylgigantol-biotin conjugate.
A biotin-streptavidin pull-down assay was performed to verify the direct interaction between 3-O-methylgigantol and EGFR. HCECs lysates were prepared in a non-denaturing lysis buffer supplemented with protease and phosphatase inhibitors. Lysates (1 mg protein) were incubated with 20 µM biotin-3-O-methylgigantol overnight at 4 °C. Lysates were pre-incubated with a 10-fold excess (200 µM) of unlabeled 3-O-methylgigantol for 2 h prior to probe addition for competitive inhibition. Biotinylated complexes were captured using Pierce Streptavidin Magnetic Beads (Thermo Fisher Scientific, Inc., Waltham, MA, USA; Cat No. 88817) for 2 h at 4 °C. After five washes with lysis buffer, the bound proteins were eluted by heating in sodium dodecyl sulfate (SDS) loading buffer and analyzed by western blotting using a specific anti-EGFR antibody (Cell Signaling Technology, Inc., Danvers, MA, USA; Cat No. 4267; 1:1000 dilution).
Total protein was extracted from cells using radioimmunoprecipitation assay
(RIPA) lysis buffer (Thermo Fisher Scientific, Inc., Waltham, MA, USA; Cat No.
89900) supplemented with protease and phosphatase inhibitor cocktails. The
protein concentration was determined using a BCA protein assay kit. Equal amounts
of protein (20 µg) were separated by 10% SDS-PAGE and transferred onto
PVDF membranes (Millipore). Membranes were blocked with 5% bovine serum albumin
(BSA) in TBST for 1 h at room temperature to minimize nonspecific binding.
Subsequently, the membranes were incubated overnight at 4 °C with the
following primary antibodies from Cell Signaling Technology, Inc. (Danvers, MA,
USA): p-EGFR (Tyr1068, #3777; 1:1000), EGFR (#4267; 1:1000), p-AKT (Ser473,
#4060; 1:1000), AKT (#4691; 1:1000), and
Study outcomes were quantified using mean
A total of 92 compounds from Dendrobium were identified using the ETCM database [26], HERB database [27], and relevant literature [28, 29] (Fig. 1A). After screening based on parameters, such as high gastrointestinal absorption and drug-likeness, requiring at least two positive indicators, and supplementation with Dendrobium compounds reported in the literature to have high content and pharmacological activity, a total of 43 active compounds were selected (Fig. 1A). The basic information on the active compounds is presented in Supplementary Table 1.
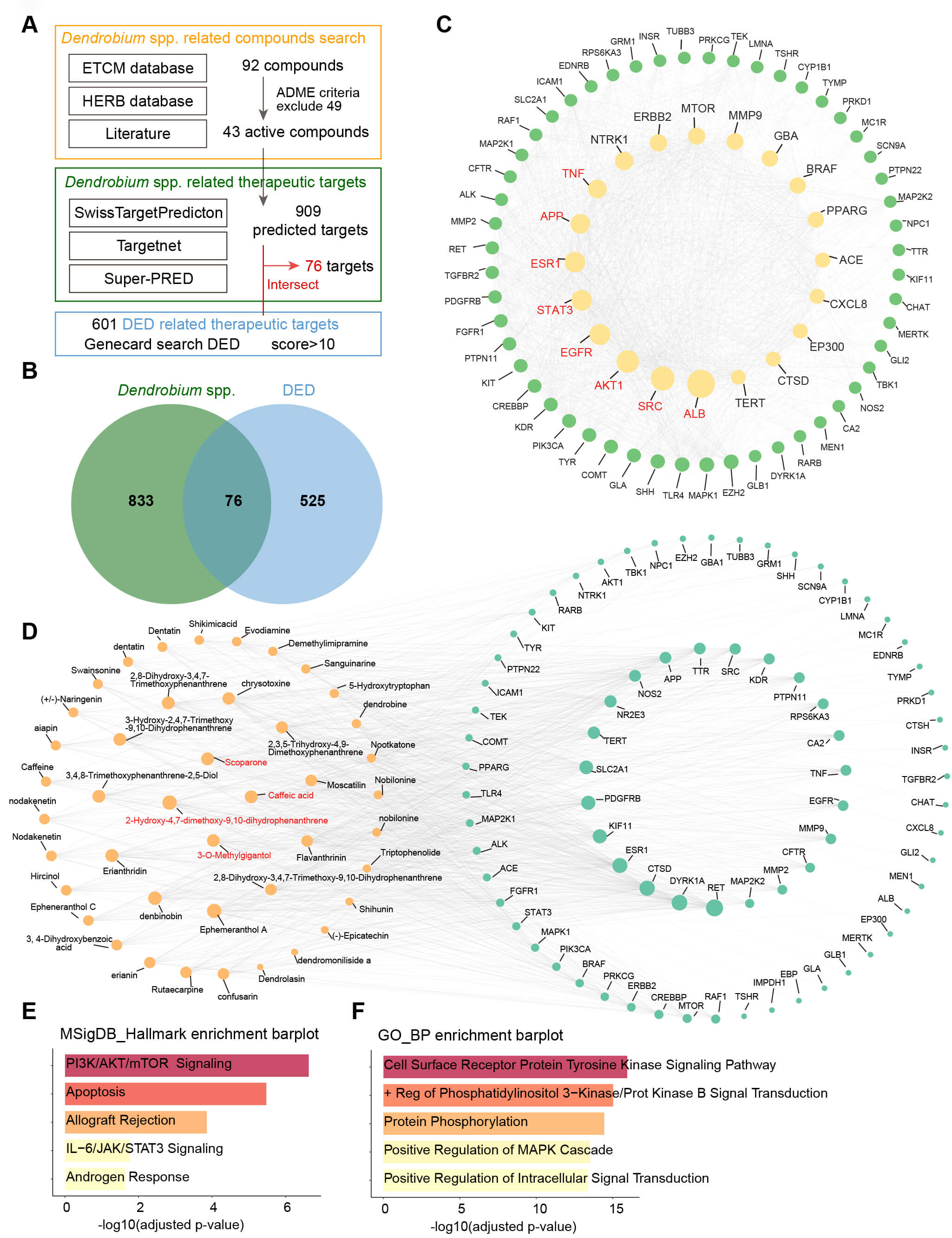 Fig. 1.
Fig. 1.
Network pharmacology analysis of Dendrobium in the treatment of dry eye disease (DED). (A) Databases used for screening Dendrobium spp.-related and DED-related therapeutic targets. (B) Venn diagram illustrating the intersection (76 common targets) between Dendrobium spp.-related targets and DED-related targets, representing potential Dendrobium targets for treating DED. (C) The Protein-Protein Interaction (PPI) network of the 76 common targets. The top 8 key therapeutic targets, identified based on degree values, are highlighted in red. (D) The Dendrobium active compound-target network. The 4 key active compounds, identified based on degree and betweenness centrality, are highlighted in red. (E,F) Functional and pathway enrichment analyses of the potential targets using Gene Ontology Biological Process (GO_BP) and MSigDB_Hallmarks databases.
The Swiss Target Prediction, Super-PRED, and TargetNet databases were used to predict 909 targets for the 43 active compounds in Dendrobium (Fig. 1A). A total of 6787 DED-related therapeutic targets were retrieved from the GeneCards database, and 601 targets with relevance scores greater than 10 were selected as highly associated with DED (Fig. 1A). By intersecting the 909 predicted targets of Dendrobium active compounds with 601 DED-related therapeutic targets, 76 common targets were identified. These were considered as potential targets through which Dendrobium may exert therapeutic effects on DED (Fig. 1B).
The top eight key targets based on degree values were APP, AKT1, SRC, EGFR, ESR1, STAT3, ALB, and TNF, which were identified as the primary targets of Dendrobium in the treatment of DED (Fig. 1C). Detailed information of the protein-protein interaction (PPI) network is presented in Supplementary Table 2. Additionally, a network of Dendrobium active compounds and potential therapeutic targets was constructed, and the key interaction data are provided in Supplementary Table 3. The top 10 compounds based on intermediate and betweenness centrality (BC) values intersected, yielding four key active compounds: Scoparone, Caffeic acid, 2-Hydroxy-4,7-dimethoxy-9,10-dihydrophenanthrene, and 3-O-methylgigantol, which were considered to be the main active compounds in the treatment of DED (Fig. 1D).
Biological functions and pathway predictions for the 76 Dendrobium targets for DED treatment were generated using the Enrichr database. MSigDB_Hallmark analysis revealed that these targets were enriched in the PI3K/AKT/mTOR and IL-6/JAK/STAT3 signaling pathways, as well as in apoptotic processes (Fig. 1E). Similarly, Gene Ontology Biological Process analysis indicated enrichment in the activation of cell surface tyrosine kinase signaling and positive regulation of PI3K signaling (Fig. 1F). These analyses allowed us to identify the key biological pathways and molecular mechanisms through which Dendrobium may exert therapeutic effects in the treatment of DED.
Molecular docking was performed using Discovery Studio to analyze the interactions within the Dendrobium active compound–potential therapeutic target network, and the lowest binding affinities were recorded. The results showed that 3-O-methylgigantol exhibited the lowest docking binding affinity with EGFR, at –32.2651 kcal/mol (Fig. 2A). Additionally, the top eight active compound–therapeutic target pairs with the lowest binding affinities were selected for visualization of the molecular docking results (Fig. 2A–H). The binding affinities of these active compounds with their respective targets were all below –5 kcal/mol, indicating that the key active compounds of Dendrobium exhibited strong binding affinities to critical targets in DED patients (Fig. 2A–H).
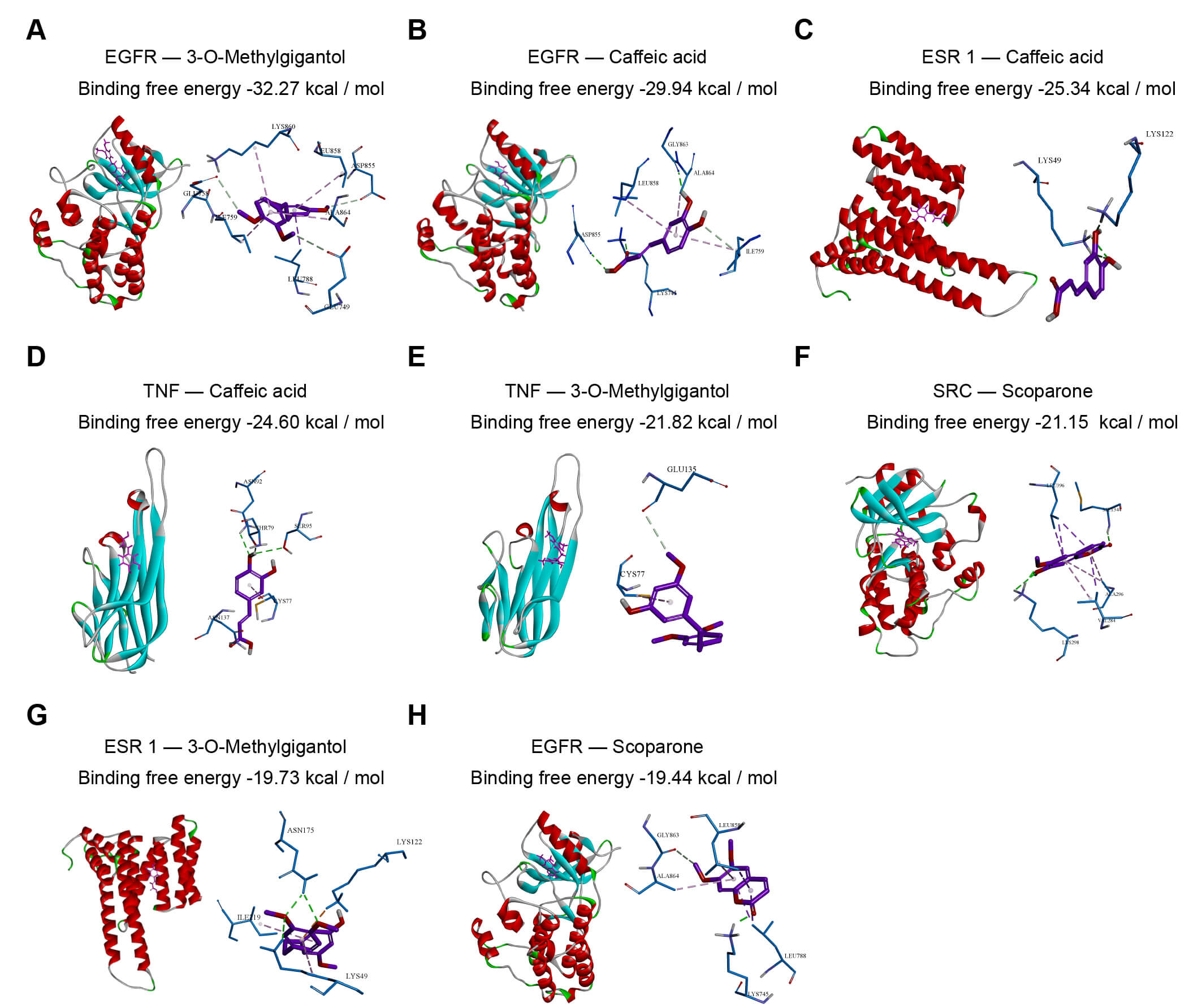 Fig. 2.
Fig. 2.
Molecular docking of key Dendrobium active compounds with predicted targets. Molecular docking was performed using Discovery Studio 2019. The figure displays the binding modes and lowest binding energies for the top 8 compound-target pairs, ranked from lowest to highest energy: (A) EGFR – 3-O-methylgigantol, (B) EGFR – Caffeic acid, (C) ESR1 – Caffeic acid, (D) TNF – Caffeic acid, (E) TNF – 3-O-methylgigantol, (F) SRC – Scoparone, (G) ESR1 – 3-O-methylgigantol, and (H) EGFR – Scoparone. Binding affinities less than –5 kcal/mol indicate strong interactions. EGFR, Epidermal Growth Factor Receptor; ESR1, Estrogen Receptor 1; TNF, Tumor Necrosis Factor; SRC, Proto-oncogene Tyrosine-protein Kinase Src.
Despite its high centrality across multiple targets, EGFR has been prioritized for molecular dynamics (MD) simulations owing to its biological and physical significance. Biologically, as an upstream cell surface receptor, EGFR initiates intracellular signaling cascades and is a highly accessible target for pharmacological interventions. Physically, it exhibited the lowest binding energy in docking screening, indicating the most stable interaction.
The binding stability of the 3-O-methylgigantol-EGFR complex was investigated using Gromacs, with simulations performed in triplicate. RMSD is a good indicator of the conformational stability of the protein-ligand complex and of the deviation of atomic positions from their initial positions. Lower RMSD values signified restricted backbone fluctuations, reflecting persistent ligand-binding positions. As shown in Fig. 3A, the complex system reached equilibrium after 80 ns, with fluctuations around 2 Å across the three replicates, indicating that the small molecule exhibits high stability upon binding to the target protein.
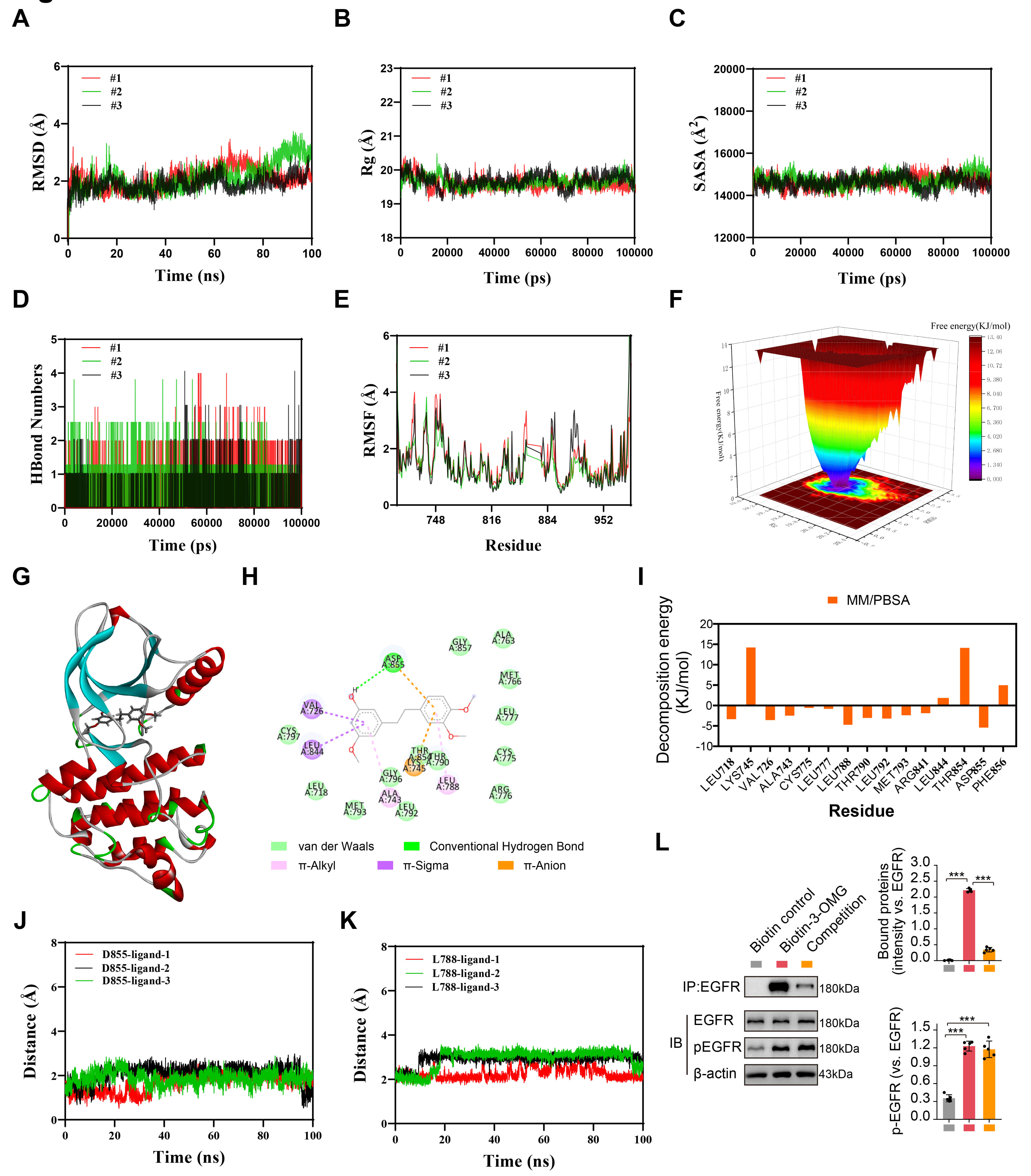 Fig. 3.
Fig. 3.
Molecular dynamics (MD) simulation and validation of the
3-O-methylgigantol–EGFR interaction. A 100 ns MD simulation was conducted using
Gromacs 2022 in three independent replicates. Analysis included: (A) RMSD. (B)
Radius of Gyration (Rg). (C) SASA. (D) Number of hydrogen bonds. (E) RMSF. (F)
Free Energy Landscape (FEL) profile. (G) 3D and (H) 2D representations of the
complex conformation at the minimum binding free energy. (I) MM/PBSA analysis
identifying key amino acid residues contributing to the binding free energy.
(J,K) The distances between 3-O-methylgigantol and the ASP855 (J) and LEU788 (K)
residues within the EGFR active site. (L) HCEC cell lysates were incubated with
biotinylated 3-OMG (Biotin-3-OMG) or biotin control, followed by precipitation
with streptavidin-conjugated beads. For the competition group, lysates were
pre-incubated with an excess of unlabeled 3-OMG prior to the addition of
Biotin-3-OMG. The presence of EGFR in the pull-down precipitates was detected by
immunoblotting. Phosphorylation levels of EGFR (Tyr1068) and total EGFR were
analyzed by Western blotting. Data are presented as means
We used two metrics, the radius of gyration (Rg) and solvent-accessible surface area (SASA), to assess the molecular stability. Rg measures protein compactness and reflects tertiary structure. The SASA quantifies the surface area of a protein that is accessible to the solvent during binding. The SASA is commonly used in simulations to study protein folding, unfolding, and structural stability. Further analysis revealed that the Rg and SASA of the complex remained stable throughout the simulation, indicating that the small molecule–target protein complex did not undergo significant expansion or contraction during movement (Fig. 3B,C). Hydrogen bonds play a crucial role in the binding between the ligand and protein. Fig. 3D shows the number of hydrogen bonds formed between the small molecule and the target protein over time. Throughout the simulation, this count fluctuated between 0 and 4, stabilizing at approximately 2 in most frames, which was consistent across the three replicates. This consistent presence indicated strong hydrogen bonding interactions within the 3-O-methylgigantol-EGFR complex (Fig. 3D).
RMSF quantifies the positional variance in protein residues and maps local flexibility along the polypeptide chain. As shown in Fig. 3E, the RMSF values of the complex are relatively low (mostly below 4 Å), indicating low flexibility and high stability. The RMSF profiles of the three replicates were nearly identical with minor variations across a few regions.
The free energy landscape (FEL) uses a visualization of free energy changes to
more intuitively explore the molecular energy landscape and protein-ligand
interactions. The energy minima represent stable states, whereas the maxima
indicate barriers to conformational change. This method can predict
ligand-binding affinities and elucidate molecular recognition mechanisms. In the
conformations in which the energy minima occurred in the complex (Fig. 3F–H),
van der Waals interactions were formed between the small molecule and the
receptor at residues GLY857, ALA763, MET766, LEU777, CYS775, ARG776, CYS797,
LEU718, MET793, LEU792, GLY796, THR854, and THR790. Additionally, the receptor
residue ASP855 formed a conventional hydrogen bond with small molecules.
Furthermore, ASP855 and LYS745 exhibited
Based on the binding conformation of the complex, the binding free energy of 3-O-methylgigantol to EGFR was calculated using MM/PBSA. The binding energies of the three replicates were –12.054, –12.240, and –11.921 kcal/mol, respectively, with a standard deviation of 0.131 kcal/mol. respectively, with a standard deviation of 0.547 kJ/mol. This narrow range underscores the reliability of the calculated binding affinities (Fig. 3I). Further analysis identified key amino acids that contributed significantly to the binding of 3-O-methylgigantol within the complex. Residues ASP855 and LEU788 of EGFR exhibited high contribution values (Fig. 3I), suggesting they may play important roles in the catalytic process.
We monitored the dynamic distances between 3-O-methylgigantol and two key
residues to validate the ligand’s stability. The hydrogen bond formed between the
polar group of the ligand and the acidic side chain of ASP855 remained stable in
all three replicate simulations, exhibiting minimal fluctuations throughout the
simulations (Fig. 3J). Similarly, the hydrophobic
In conclusion, 3-O-methylgigantol binds stably to EGFR with a low binding free energy, suggesting that it may exert its effects via EGFR.
A 3-O-methylgigantol-biotin conjugate probe (biotin-3-OMG) was synthesized via click chemistry to validate EGFR as a direct target of 3-O-methylgigantol (Supplementary Fig. 1). A pull-down assay was performed using a biotinylated probe (biotin-3-OMG). Immunoblotting revealed that biotin-3-OMG significantly enriched EGFR from cell lysates compared to that in the biotin control, indicating a physical interaction (Fig. 3L). Notably, this binding was specific, as it was markedly blocked by preincubation with excess unlabeled 3-OMG (Fig. 3L). Furthermore, 3-OMG treatment significantly upregulated EGFR phosphorylation (p-EGFR, Tyr1068) (Fig. 3L). Collectively, these findings demonstrate that 3-O-methylgigantol directly binds to EGFR and induces its activation.
Cell viability of human corneal epithelial cells (HCECs) treated with varying concentrations of hypertonic solutions was assessed using the CCK-8 assay to determine the optimal concentration. In our hypertonic stress experiment, after 24 h of incubation, cell viability decreased in a dose-dependent manner as the osmotic pressure increased from 312 to 550 mOsm (Supplementary Fig. 2). When the osmotic pressure reached 550 mOsm, the cell viability declined to approximately 50%; therefore, this concentration was selected for subsequent experiments. Following treatment with different concentrations of 3-O-methylgigantol (312–550 mOsm) for 24 h, the cell viability increased in a dose-dependent manner. At a 5 µM concentration, 3-O-methylgigantol significantly enhanced the cell viability (Fig. 4A) and mitigated the reduction in cell viability induced by hypertonic stress (Fig. 4B).
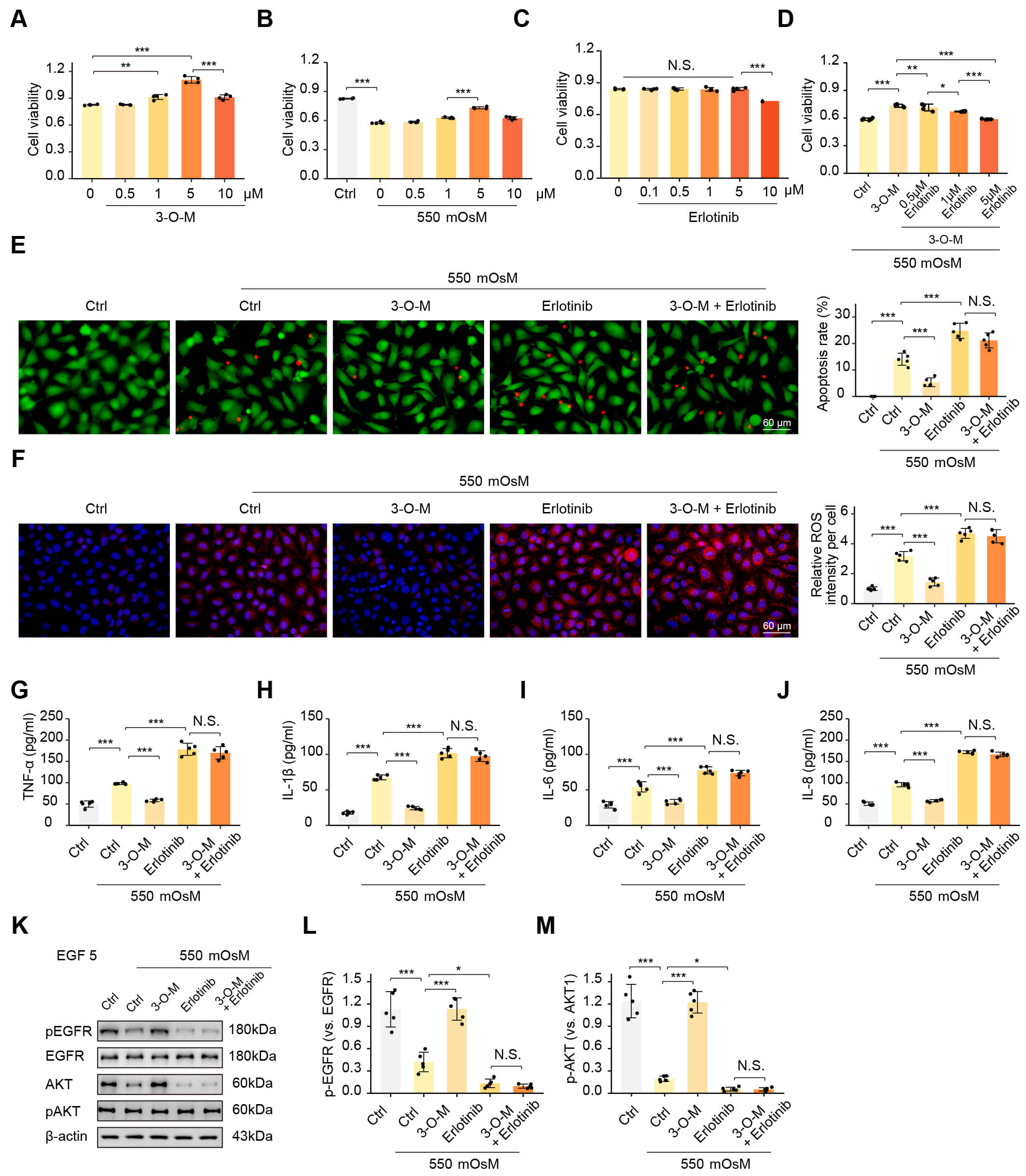 Fig. 4.
Fig. 4.
3-O-methylgigantol ameliorates hyperosmotic-induced cytotoxicity
in HCECs via the EGFR pathway. (A–D) Viability of Human Corneal Epithelial
Cells (HCECs) measured by the CCK-8 assay (n = 4 biological repeats): (A) HCECs
treated with different concentrations of 3-O-methylgigantol for 24 h under normal
conditions. (B) HCECs were treated with different concentrations of
3-O-methylgigantol for 24 h under 550 mOsM hyperosmotic stress. (C) HCECs were
treated with different concentrations of the EGFR inhibitor Erlotinib for 24 h
under normal conditions. (D) HCECs treated with 5 µM 3-O-methylgigantol
alone or combined with different concentrations of Erlotinib for 24 h under 550
mOsM stress. (E–J) HCECs treated with 5 µM 3-O-methylgigantol
To determine whether EGFR is a key mediator of 3-O-methylgigantol-induced HCEC activation, we used the EGFR inhibitor, erlotinib. Erlotinib binds to the ATP-binding site, preventing the activation of EGFR kinase by binding to ATP and transferring phosphate groups to substrates. First, to rule out potential off-target cytotoxicity, we evaluated the viability of HCECs treated with a concentration gradient of Erlotinib (0.1, 0.5, 1, 5, and 10 µM) under normal culture conditions. The CCK-8 assay indicated that Erlotinib exhibited no significant toxicity at concentrations up to 5 µM (Fig. 4C). Therefore, we performed dose-dependent blocking assays. Under hypertonic stress, HCECs were co-treated with 5 µM 3-O-methylgigantol and increasing concentrations of Erlotinib (0.1, 1.0, and 5.0 µM). Erlotinib attenuated the protective effect of 3-O-methylgigantol in a dose-dependent manner. While 0.1 µM showed partial inhibition, 5 µM Erlotinib completely abolished the 3-O-methylgigantol-mediated restoration of cell viability and exacerbated hypertonic damage (Fig. 4D). These findings confirm that the protective action of 3-O-methylgigantol is specifically dependent on EGFR signaling activity.
Cell apoptosis was assessed using calcein-AM/PI staining, revealing that
3-O-methylgigantol alleviated hypertonic-induced apoptosis in HCECs, an effect
that was blocked by Erlotinib (Fig. 4E). DCFH-DA staining, used to measure
intracellular ROS levels, showed that 3-O-methylgigantol reduced
hypertonic-induced ROS accumulation in HCECs, and this effect was inhibited by
Erlotinib (Fig. 4F). ELISA assays measured the levels of TNF-
DED is characterized by a tear film imbalance that causes dryness, fatigue,
foreign body sensation, and pain. Untreated condition can cause inflammatory damage to
the cornea and the conjunctiva. Dendrobium exhibits antitumor,
immunomodulatory, antioxidant, anti-inflammatory, anti-aging, anti-cataract, and
digestion-promoting properties. Recently, its potential for the treatment of DED
has gained attention. In a DED rat model, Dendrobium water extracts
enhanced tear secretion, protected the ocular surface structure, increased goblet
cell numbers and mucin expression, and inhibited the activation of MAPK and
NF-
This study used network pharmacology to identify four key bioactive compounds
from Dendrobium scoparone, caffeic acid,
2-Hydroxy-4,7-dimethoxy-9,10-dihydrophenanthrene, and 3-O-methylgigantol,
considered core agents for the treatment of dry eye syndrome. Scoparone
and Caffeic acid exhibit strong anti-inflammatory and immunomodulatory effects by
inhibiting the NF-
3-O-methylgigantol, which is found in several Dendrobium species, including Dendrobium fimbriatum, is one of the primary active compounds. Although their specific bioactivity remains underexplored, stilbene analogs possess anti-inflammatory and antioxidant properties. Given that its parent compound, gigantol, exhibits various pharmacological effects, 3-O-methylgigantol is expected to have significant therapeutic potential, particularly in diseases related to inflammation and oxidative stress. To date, no studies have reported the role or mechanism of DED.
Network pharmacology analysis identified EGFR as the principal target. Molecular
docking and kinetic simulation validated the stable binding interaction between
3-O-methylgigantol and EGFR. Notably, this receptor is abundantly expressed in
corneal epithelial cells. Proper EGFR signaling is crucial for corneal epithelial
homeostasis, as it regulates cellular renewal, barrier function, and injury
repair. EGFR activation relies on ligands such as epidermal growth factor (EGF),
transforming growth factor-alpha (TGF-
Hyperosmolarity of the tear film is the core mechanism of DED [46] and is caused by either insufficient tear secretion or excessive evaporation [47]. It damages the epithelial cells on the ocular surface, triggering oxidative stress, inflammation, and apoptosis [48]. In our in vitro hyperosmolarity model, 550 mOsm/L was selected as the optimal concentration to simulate dry eye conditions. We found that 3-O-methylgigantol effectively alleviated corneal epithelial damage caused by hyperosmolarity and reduced reactive oxygen species accumulation and inflammatory cytokine levels. These experiments validate the potential therapeutic role of 3-O-methylgigantol in treating hyperosmotic corneal damage.
This study has several limitations. First, our investigation relied solely on a single SV40-immortalized HCEC line [49]. Although this cell line serves as a reproducible in vitro model, the immortalization processmediated by the SV40 Large T antigen involves the functional inactivation of p53 and retinoblastoma (Rb) tumor suppressor pathways [50]. This mechanism is known to dysregulate cell cycle checkpoints and apoptotic signaling, potentially resulting in inflammatory and survival responses that differ from those observed in primary corneal epithelial cells. Second, the study was conducted exclusively in vitro without in vivo validation of 3-O-methylgigantol’s tear film stabilization or corneal epithelial repair effects. Future studies should verify these findings using established animal models of dry eye to determine the therapeutic dose and confirm the absence of systemic or local adverse effects. Furthermore, small molecules frequently exhibit polypharmacology, including off-target binding, which may lead to unintended cytotoxicity or functional impairment of sensitive ocular tissues. Therefore, comprehensive and unbiased screening, such as high-throughput profiling, is essential to fully map the target profile of a compound and ensure both specificity and long-term safety prior to clinical translation.
This study comprehensively explored the therapeutic potential of Dendrobium and its active constituents in the treatment of DED using network pharmacology, molecular docking, and cellular assays. These results suggest that these compounds may protect the ocular surface by modulating inflammation, oxidative stress, and key signaling pathways such as EGFR/PI3K/AKT. Notably, 3-O-methylgigantol, the key focus of this study, was the first compound to alleviate hyperosmolarity-induced corneal epithelial damage via EGFR activation, providing important evidence for novel dry eye treatments.
DED, Dry Eye Disease; EGF, Epidermal Growth Factor; EGFR, Epidermal Growth Factor Receptor; ERK, Extracellular Signal-Regulated Kinase; HCECs, Human Corneal Epithelial Cells; BC, Betweenness Centrality; TNF-
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
JY and WB designed the research study. SZ, JL, and YL performed the research. CS and YZ conducted the ELISA experiments, contributing substantially to data acquisition. JL analyzed the data. SZ drafted the manuscript. YW and SW visualized the data and prepared the figures. All authors contributed to critical revision of the manuscript for important intellectual content. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
We thank the medical laboratory of the Eye Hospital, Nanjing Medical University, for providing the experimental equipment and space. We would like to thank Editage for English language editing. The graphical abstract was created in BioRender (https://www.biorender.com/). Yao, L. (2026) https://BioRender.com/xw5gpv9.
This research received no external funding.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/FBL47218.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.




