1 Department of Molecular Cell Biology, Weizmann Institute of Science, 76100 Rehovot, Israel
2 Everaard Goodman Faculty of Life Sciences, Bar-Ilan University, 52900 Ramat-Gan, Israel
3 Center of Animal Biotechnology and Gene Therapy (CBATEG) and Department of Biochemistry and Molecular Biology, School of Veterinary Medicine, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain
4 CIBER de Diabetes y Enfermedades Metabólicas Asociadas, Instituto de Salud Carlos III, 28029 Madrid, Spain
Abstract
We have previously demonstrated that ovarian tumor (Otu) domain-containing ubiquitin aldehyde-binding protein 2 (Otub2), a deubiquitinating enzyme, exerts anti-apoptotic effects in primary human islets. The present study aims to further elucidate the molecular mechanisms underlying the role of Otub2 as a regulator of insulin secretion and β-cell function.
Otub2 overexpression or silencing was employed to study its effects on cultured MIN6 cells and dispersed human islets. To evaluate its in vivo effects, Otub2 knockout (KO) mice were employed, as well as a pancreata-specific Otub2 overexpression model. RNA sequencing was performed on pancreatic tissue from Otub2-KO and control mice to study its effects on gene expression patterns. Co-immunoprecipitation followed by mass spectrometry identified Otub2-interacting proteins.
Overexpression of Otub2 inhibited NF-κB activity and enhanced glucose-stimulated insulin secretion (GSIS) in cultured MIN6 cells and primary human islets. Otub2 KO mice exhibited impaired glucose tolerance and upregulation of NF-κB target genes. Conversely, selective in vivo overexpression of Otub2 in pancreata of C57BL wild-type mice resulted in significantly lower (~30%) blood glucose levels, post glucose injection, compared to control mice. Transcriptomic analysis of KO pancreata revealed downregulation of K+ transporter-related genes and upregulation of oxidative phosphorylation genes, consistent with defective insulin secretion. Mass spectrometry identified the voltage-gated potassium channel subunit Kv9.3 as a major Otub2 binding partner, along with paternally expressed 3 (Peg3) and calcium/calmodulin dependent protein kinase II delta (Camk2d) proteins known to promote NF-κB signaling and β-cell apoptosis.
Otub2 is a critical regulator of β-cell function, acting through modulation of NF-κB signaling and K+ channel-associated complexes. By deubiquitinating components such as Peg3 and Camk2d, Otub2 may protect β-cells from cytokine-induced apoptosis and sustain insulin secretory capacity. These findings position Otub2 as a potential therapeutic target for preserving β-cell function in diabetes.
Keywords
- Otub2
- pancreatic beta cells
- NF-κB
- apoptosis
- potassium channels
Type-1 and Type-2 diabetes mellitus involve selective and progressive loss of
pancreatic
To improve
Otub2, a family member of cysteine proteases having a deubiquitinase activity
[13], was a highly significant ‘hit’ in the above screens. Silencing of Otub2
expression increased caspase-3/7 activity in primary human islets treated with a
mixture of cytokines (TNF-
MIN6 murine pancreatic
pEGFP-c1 plasmid (Addgene, https://www.addgene.org/) harboring G418 resistance was used for creating stable MIN6 cell lines that overexpress Otub2 (mouse isoform 2) with a Flag tag (instead of EGFP). The pEGFP-c1 vector, in which the Flag tag was cloned instead of the EGFP sequence (pFlag), served as a control. pEGFP-c1 plasmids which contained the original EGFP sequence (p-EGFP-c1) and EGFP-Otub2 sequence (pEGFP-Otub2) were used for transient transfection of MIN6 cells. To create these constructs, the Otub2 cDNA was amplified by PCR using the 5′ primer (5′-CAGTCCGGAGACACTATGAGTGAAACATCTTTCAACC-3′); and the 3′ primer (5′-CGCGGATCCGCGGTAGTCAGTGTTTCTCGGCTGC-3′). The PCR product was digested using BspE1 and BamHI restriction enzymes and ligated into the pEGFP-c1 plasmid. An additional construct was generated using the following primers containing Flag tag: 5′ primer (5′-CTAGCATGGACTACAAAGACGATGACGACAAGT-3′) and 3′ primer (5′- CCGGACTTGTCGTCATCGTCTTTGTAGTCCATG-3′). The primers were hybridized and ligated into pEGFP-c1 digested with NheI and BspEI restriction enzymes. The EGFP cDNA was then excised from the plasmid and replaced by a Flag tag, creating the pFlag-Otub2 construct. pFlag plasmid was generated after digestion of pFlag-Otub2 construct with BspE1 and BamHI restriction enzymes (excising Otub2 out of the construct) and used as a control.
Polyclonal anti-Flag and anti-GAPDH antibodies were from Sigma (St. Louis, MO, USA). Polyclonal anti-Otub2 antibodies were from Novus Biologicals (Centennial, CO, USA). Polyclonal anti-GFP antibodies were from Abcam (Waltham, MA, USA). Alexa488 conjugated goat anti-mouse secondary antibodies were from Life Technologies (Waltham, MA, USA).
The cytokine mixture referred to as ‘1x-cytomix’ consisted of 3 nM
TNF-
Isolated human islets (~80% purity confirmed by dithizone staining) were provided by the European Consortium for Islets Transplantation (Islet for Basic Research program) through a Juvenile Diabetes Research Foundation Award 31-2008-413. Isolated human islets were grown in CMRL 1066 medium supplemented with 10% (v/v) FBS, 2 mM L-glutamine, 100 units/mL penicillin, 100 µg/mL streptomycin, 0.25 µg/mL amphotericin, and 40 µg/mL gentamycin, and were cultured at 37 °C in a 5% CO2 humidified atmosphere. Islets media was replaced every other day. Intact human islets were dispersed by incubation at 37 °C (4 minutes) with 1 mg/mL Trypsin/EDTA by pipetting the cells, and by passing them twice through a 21G needle. Trypsinized islets were washed and resuspended with CMRL 1066 medium supplemented with 10% FBS. Cells were used within 48 hours following dispersion. Human islets studies received Ethics approval from the Bioethics and Embryonic Stem Cell Research Oversight Committee (ESCRO) of the Weizmann Institute of Science.
MIN6 cells or human islets were seeded in 96 well plates (30,000 cells/well or 1000 dispersed human islets cells/well) in 100 µL medium (DMEM for MIN6 cells and CMRL for human islets), and immediately transfected with sequences of non-targeting siRNA or Otub2 siRNA (to a final concentration of 25 nM) using DarmaFECT-4 transfection reagent (Dharmacon, Waterbeach, UK) for MIN6 cells and Darma-FECT-1 transfection reagent (Dharmacon) for human islets, according to manufacturer’s instructions.
MIN6 cells or human islets were seeded (500,000 cells/well in 6-well plates for MIN6 cells, and 1000 islets/well in 96-well plates for human islets) and 24 hours post-seeding, the cells or human islets were transfected with pFlag or pFlag-Otub2 vectors (3 µg/well for a 6-well plate and 0.2 µg/well for a 96-well plate) using the transfection reagent Lipofectamine 2000 (Thermo-Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s instructions.
MIN6 cells were seeded in 12-well plates and 24 hours post-seeding, cells were transfected using jetPEI transfection reagent (Polyplus Transfection, Illkirch, France), with pEGFP-c1 vectors (1 µg/well) harboring G418 resistance to calibrate for the cells’ G418 resistance. pEGFP-c1 vector in which Flag tag was cloned instead of EGFP sequence (pFlag) served as control; and pEGFP-c1 with Flag-Otub2 cDNA clone without EGFP sequence (pFlag-Otub2) was utilized to overexpress Otub2. Non-transfected cells served as a control and 48 hours after transfection media were replaced with media containing elevated concentrations of G418. After the calibration process was complete and the proper concentration of G418 required for selection was established, MIN6 cells were seeded in 6-well plates (150,000 cells/well) to create stable cell lines, 24 hours post-seeding, cells were transfected with pEGFP-c1 constructs (3 µg/well), and 48 hours thereafter, the media were replaced with media containing 400 µg/mL G418 for the selection process. After completion of selection, the media were replaced with media containing 200 µg/mL G418 for maintenance.
MIN6 cells were seeded in 96-well plates (30,000 cells/well) and transfected with siRNA and 48 hours later, cells were treated with a cytokine mixture, referred to as 1x-cytomix. For stably transfected cell lines, cells were treated with 1x-cytomix 24 hours post-seeding and 24 hours later caspase 3/7 activity was examined using Sensolyte homogeneous RH110 caspase 3/7 assay kit (AnaSpec, Fremont, CA, USA). Human islets were dispersed and seeded in 96-well plates (1000 islets/well). Islets were transfected with p-Flag or pFlag-Otub2 constructs that stably express Flag-Otub2. The rest of the experiment was similar to the assay in MIN6 cells as described above.
NF-
Stably transfected MIN6 cells were seeded in 96-well plates (30,000 cells/well). After 48 hours, cells were incubated for 1 hour in Krebs-Ringer bicarbonate HEPES buffer (KRBH), containing 124 mM NaCl, 5.6 mM KCl, 2.5 mM CaCl2, and 20 mM HEPES, pH 7.4, at 37 °C, followed by incubation for 1 hour in KRBH with 0 mM or 20 mM glucose. Insulin concentration in the culture medium was determined using insulin detecting HTRF kit (Cisbio, Codolet, France) according to the manufacturer’s instructions.
Following treatment in 24-well plates, cells were harvested, and total RNA was extracted using the PerfectPure RNA kit (5-prime, Montreal, Canada). RNA was quantified using nano-drop. cDNA was generated by the cDNA Reverse Transcription kit (Applied Biosystems, Waltham, MA, USA), following the manufacturer’s instructions. qRT-PCR was carried out using an ABI-Prism 7300 instrument (Applied Biosystems, Waltham, MA, USA), utilizing SYBR Green PCR mix (Invitrogen, Carlsbad, CA, USA) and specific primers (100 nM final concentration). Expression levels of Actin or Hypoxanthine Phosphoribosyltransferase 1 (HPRT) were used to normalize mRNA levels. Primers used are given herein (see Table 1).
| Gene | Origin | Forward primer | Reverse primer |
| iNOS | Mouse | GCCCTGCTTTGTGCGAAGTG | AGCCCTTTGTGCTGGGAGTC |
| MCP-1 | Mouse | AGGTGTCCCAAAGAAGCTGTA | ATGTCTGGACCCATTCCTTCT |
| IL-1 |
Mouse | TGCCACCTTTTGACAGTGATG | TGATGTGCTGCTGCGAGATT |
| IP-10 | Mouse | ATGACGGGCCAGTGAGAATG | TCAACACGTGGGCAGGATAG |
| Otub2 | Mouse | AACCGAGCTGACTTCTTCCG | CGTCGACGTACTCTACCTGC |
| Glut2 | Mouse | TTTTCAGCCAAGGACCCCGT | GCCCAAGGAAGTCCGCAATG |
| Nkx6.1 | Mouse | AACACACCAGACCCACGTTCT | ATCCCCAGAGAATAGGCCAAG |
| MafA | Mouse | CAAGGAGGAGGTCATCCGAC | TCTCCAGAATGTGCCGCTG |
| HPRT | Mouse | GCAGTACAGCCCCAAAATGG | GGTCCTTTTCACCAGCAAGCT |
| Actin | Mouse | GGCCAACCGTGAAAAGATGA | CACAGCCTGGATGGCTACGT |
iNOS, nitric oxide synthase 2; MCP-1, C-C motif chemokine ligand 2; IP-10, C-X-C motif chemokine ligand 10; Otub2, ovarian tumor (Otu) domain-containing ubiquitin aldehyde-binding protein; Glut2, glucose transporter 2; Nkx6.1, NK6 homeobox 1; MafA, MAF BZIP transcription factor A; HPRT, hypoxanthine phosphoribosyltransferase 1.
RNA was extracted from the pancreas and liver of mice using the PerfectPure RNA
kit (5-prime, Montreal, Canada) following the manufacturer’s instructions. An
Agilent 4200 TapeStation System (Agilent Technologies, Santa Clara, CA, USA) was
used to assess RNA quality. RNA-seq libraries were generated by the MARS-seq
protocol [16]. Libraries were sequenced by the Novaseq 6000 (Illumina, San Diego,
CA, USA), using SP mode 100 cycles kit (Illumina, San Diego, CA, USA). Alignment
of sequences to the genome and count matrix determination were performed by the
UTAP pipeline (Weizmann Institute, Rehovot, Israel). Normalization of libraries,
low count genes filtration, and calculation of differentially expressed genes
were performed using the edgeR and Limma packages in R
(https://www.r-project.org/). Gene Ontology (GO) and MsigDB pathways enrichment
analysis were performed using the Camera method from the Limma package in R. The
enrichment results are given in p-values, with –log10(p-value)
Protein A-agarose beads were washed with 0.1 M Tris-HCl (pH 8.0, 4 °C),
followed by incubation with Flag or GFP antibodies in 0.1 M Tris-HCl (pH 8.0) for
4 h at 4 °C. Supernatants (centrifuged at 20,000
Cells were seeded and treated as indicated. Treated cells were washed three
times with PBS and were harvested in lysis buffer (25 mM Tris-HCl [pH 7.4], 10 mM
sodium orthovanadate, 10 mM pyrophosphate, 100 mM sodium fluoride, 10 mM EDTA, 10
mM EGTA, and 1 mM phenylmethylsulphonyl fluoride). Lysates were centrifuged
(20,000
All in vivo experiments were carried out in accordance with relevant guidelines and regulations. Experiments were approved by the Animal Care and Use Committee of the Weizmann Institute of Science (application number 23980116-3). Mice were housed under standard light/dark conditions and were given access to food and water ad libitum. The study is reported in accordance with the ARRIVE guidelines.
The mouse strain C57BL/6NTac-Otub2tm1a(EUCOMM)Wtsi/WtsiH was ordered from the Wellcome Trust Sanger Institute, London, UK, as part of the EUCOMM Mutant Mouse Project. One heterozygous male and two heterozygous female mice on a C57BL/6NTac background were ordered from the European Mouse Mutant Archive (EMMA) mouse repository. EUCOMM vector contains 5′ and 3′ homology arms, facilitating homologous recombination, and a targeting cassette that disrupts gene function, flanked by flipase recognition target (FRT) recombination sites to allow removal by Flp recombinase. Further, exons 3/4 of Otub2 were flanked by a pair of loxP recombination sites, inducing a frame shift upon removal of the exons and leading to complete gene inactivation. For genotyping, mouse tail genomic DNA preparations were extracted using REDExract-NAmp tissue PCR kit (Sigma), followed by amplification reactions performed with oligonucleotide pairs (see Table 2) specific for the foreign or wild-type sequences, to amplify ~200–900 bp fragments.
| Forward | Reverse | Product size | |
| Wild-type 1 | GATGGTCAGCCTTGTTAGCA | CCGTTCAGTCAGGTCCCTAG | 940 bp |
| Wild-type 2 | GATGGTCAGCCTTGTTAGCA | CTTGAGGGAACAGGGCATGT | 970 bp |
| Tm1a allele 1 | GAGGACAGCTTGGGAGAGAT | CCACAACGGGTTCTTCTGTT | 635 bp |
| Tm1a allele 2 | AGGCGCATAACGATACCACGAT | CCACAACGGGTTCTTCTGTT | 204 bp |
Otub2 knock-out and wild-type (WT) mice were sacrificed by cervical dislocation. An incision was made from the upper abdomen downwards to expose the liver and intestines. The duodenum was exposed by shifting the intestines to the right, and the pancreas was isolated by pulling the intestines carefully from the duodenum downwards. Spleen was then removed, and the pancreas was detached from the large intestine to complete its isolation.
Adeno-associated viral vectors (AAVs) that can stably express Flag or Flag-Otub2
were generated by inserting the Flag or Flag-Otub2 inserts into the
pAAV-RIP-VEFG-WPRE vector, after eliminating the VEGF coding sequence, as
described [18, 19]. The pAAV-RIP-Flag (for control purposes) and
pAAV-RIP-Flag-Otub2 vectors were used for production of the AAV vectors. To
ensure selective expression of Otub2 in pancreatic
Mice were fasted overnight with water access ad libitum. Mice were intraperitoneally injected with glucose (1.8 g/kg body weight). Blood samples were taken at timed intervals (0–120 minutes) from a tail vein, and Glucose levels were monitored using MediSense Optium Blood Glucose test strips (Abbott Laboratories, Abbott Park, IL, USA). Glucose levels were plotted versus the time points, and the area under the curve (AUC) was calculated using the computeAUC function from the PharmacoGx package in R.
For immunofluorescence, pancreata were fixed for 24 h in 4% paraformaldehyde
and then transferred to 70% ethanol until embedding in paraffin by a standard
protocol (by using automated tissue processing) as follows: pancreata were first
dehydrated by immersing them sequentially for 45 minutes each in a series of
ethanol-water mixtures (70%; 95% (
Statistical analysis was performed in R. Differences between experimental conditions were determined by a two-tailed Student’s t test, unless otherwise mentioned in the figure legends.
To characterize the effects of Otub2 on
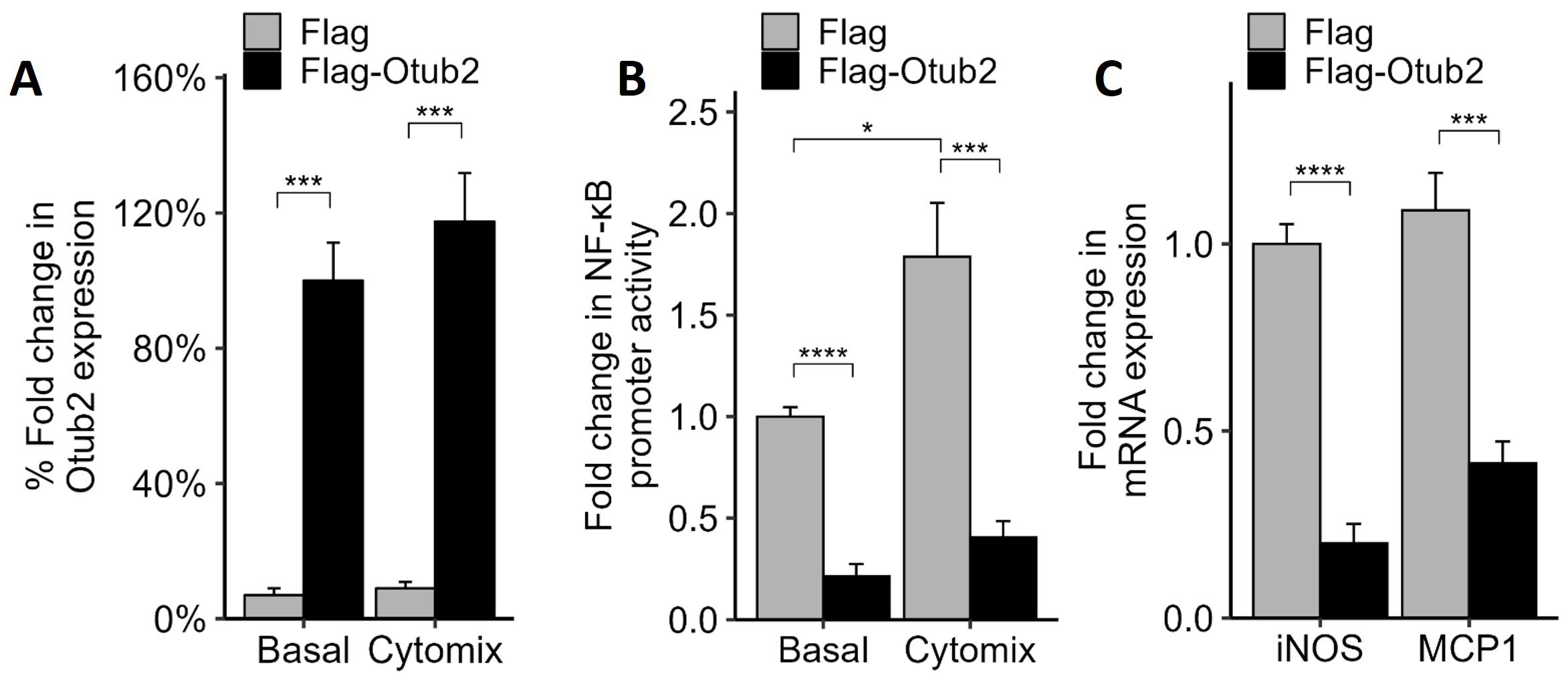 Fig. 1.
Fig. 1.
Effects of OTUB2 overexpression on NF-
To examine the effects of Otub2 on NF-
The reduced NF-
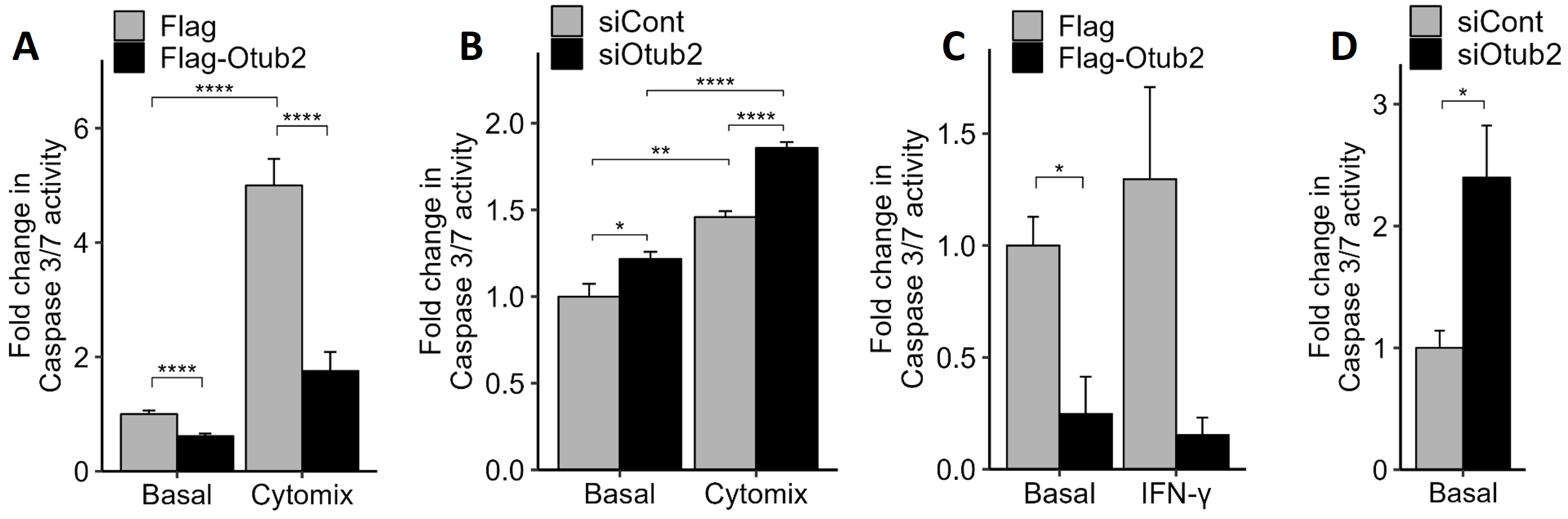 Fig. 2.
Fig. 2.
Effects of Otub2 on caspase 3/7 activity. (A) Control, Flag
over-expressing MIN6 cells or Flag-Otub2 over-expressing cells were treated with
1x-cytomix for 24 h (or remained untreated). Apoptosis was assayed by caspase-3/7
activity measurements. (B) MIN6 cells, transfected with the indicated siRNAs,
remained untreated or were treated with 1x-cytomix for 24 h. Apoptosis was
assayed by caspase-3/7 activity measurements. Control-siRNA transfected cells
served as control. (C,D) Dispersed human islets transiently transfected for 48 h
with pFlag (control) or pFlag-Otub2 constructs, remained untreated or were
treated with IFN-
Even stronger effects were observed in dispersed human islets. An approximate
5-fold decrease in caspase 3/7 activity was observed in human islets that were
transfected with the Flag-Otub2 construct, when compared with Flag-transfected
islets (Fig. 2C). Similar results were observed upon treatment of dispersed human
islets with IFN-
To determine whether overexpression of Otub2 affects
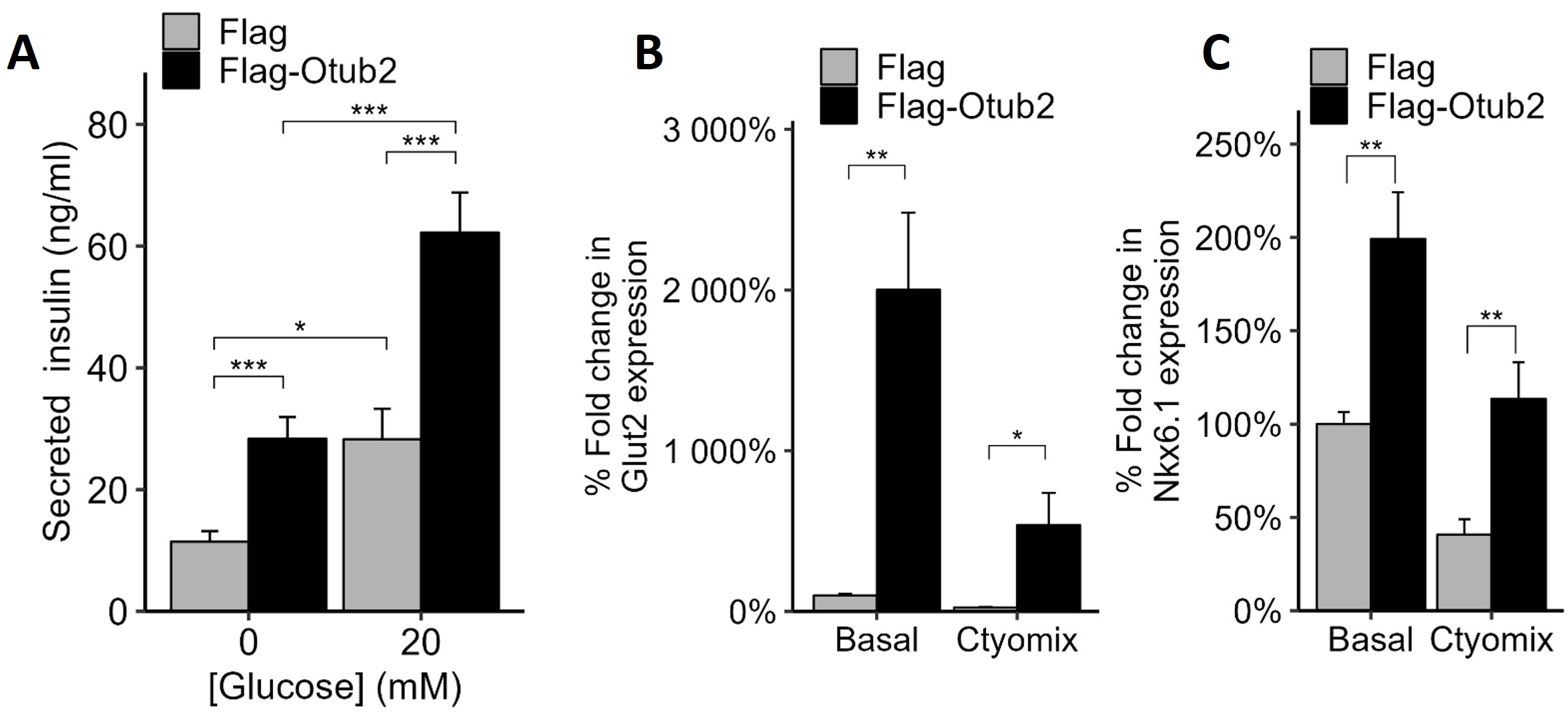 Fig. 3.
Fig. 3.
Effects of OTUB2 stable over-expression on
Another gene of interest is NK6 homeobox 1 (Nkx6.1), a known transcription
factor required for
To identify proteins that interact with Otub2, immunoprecipitations (IPs) with
Otub2 antibodies were carried out using MIN6 cells transiently overexpressing
Otub2 (and appropriate controls), followed by mass spectrometry analysis. Several
proteins were enriched at least 100-fold in immunoprecipitates derived from
Flag-Otub2 overexpressing cells (Table 3, Supplementary Table 1)
compared to controls. A similar trend was observed when GFP-Otub-2 Min6 cells
were used. Top hits were Peg3 and Camk2d, which enhance the NF-
| Protein | Gene | Ratio | Ratio | Function |
| OTUB2/FLAG | OTUB2/GFP | |||
| OTUB2 | Otub2 | 145.09 | 179.16 | |
| PEG3 | Peg3 | Inf | Inf | Acts synergistically with TRAF2 through activation of NF- |
| KCC2D | Camk2d | Inf | 133.39 | Calcium/calmodulin-dependent kinase |
| KCNS3 | Kcns3 | Inf | 29.11 | Voltage-gated potassium channel subunit Kv9.3 |
| SSRD | Ssr4 | Inf | 8.37 | Binds calcium to the ER membrane and regulates the retention of ER resident proteins |
| SSRG | Ssr3 | 2.53 | 3.35 | Binds calcium to the ER membrane and regulates the retention of ER resident proteins |
| AT1A1 | Atp1a1 | Inf | 3.62 | Sodium/potassium-transporting ATPase |
| ATPG | Atp5c1 | Inf | 2.25 | Mitochondrial membrane ATP synthase |
| AT2A3 | Atp2a3 | 3.09 | 4.56 | Catalyzes the hydrolysis of ATP coupled with the transport of calcium |
PEG3, paternally expressed gene 3; TRAF2, TNF receptor associated factor 2; KCC2D, calcium/calmodulin dependent protein kinase II delta; KCNS3, potassium voltage-gated channel modifier subfamily S member 3; SSRD, signal sequence receptor subunit 4; ER, endoplasmic reticulum; SSRG, signal sequence receptor subunit 3; AT1A1, ATPase Na+/K+ transporting subunit alpha 1; ATPG, ATP synthase F1 subunit gamma; AT2A3, ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 3.
The effects of Otub2 on
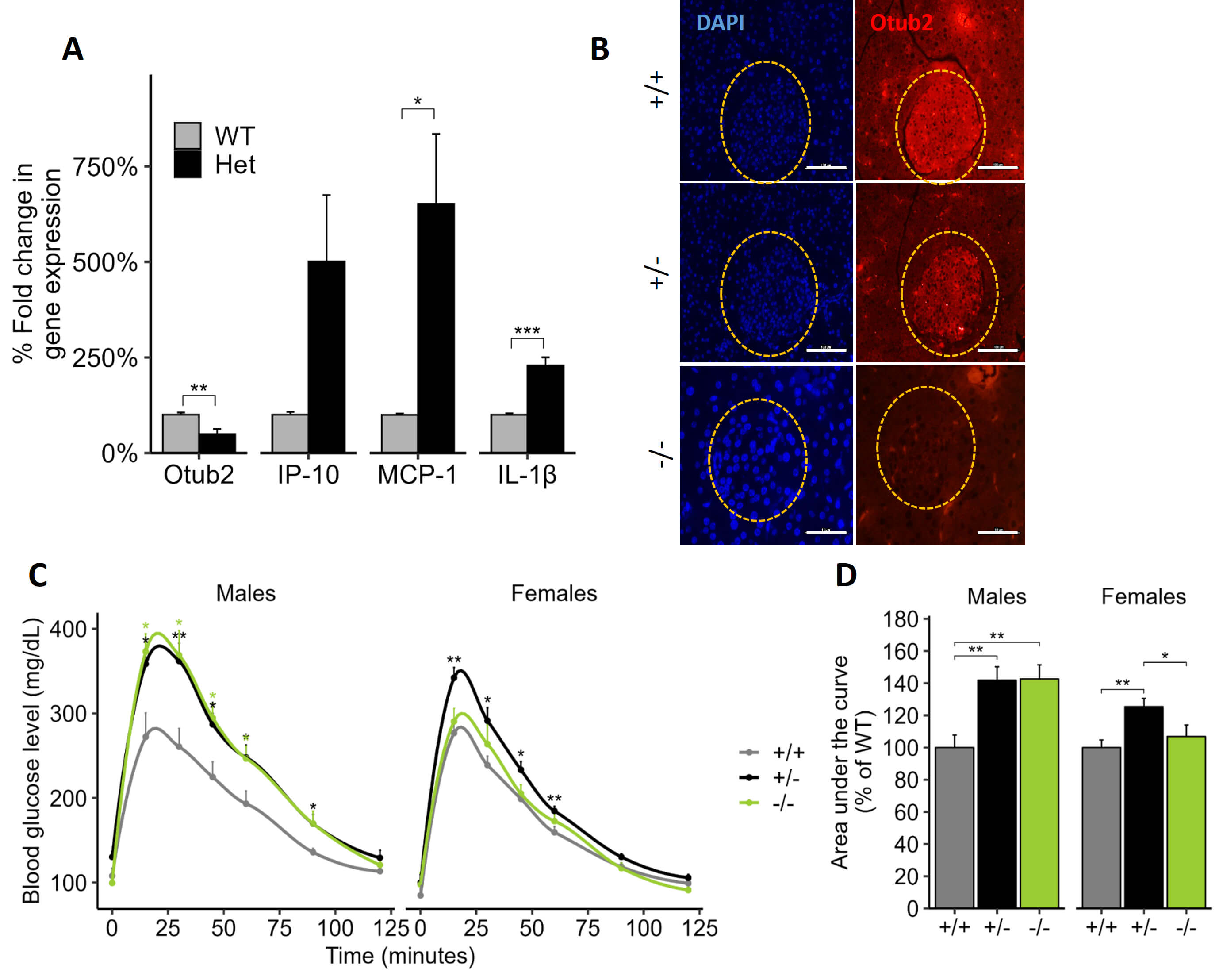 Fig. 4.
Fig. 4.
Effects of OTUB2 knockdown on pancreatic NF-
Next, glucose tolerance tests (GTT) were performed in Otub2+/+, Otub2+/-, and Otub2-/- mice. As shown in Fig. 4C, basal glucose levels of Otub2+/+, Otub2+/-, and Otub2-/- male and female mice were approximately similar. However, following glucose injection, blood glucose levels, at all-time points, were significantly higher in Otub2+/- and Otub2-/- male mice, when compared to Otub2+/+ animals (Fig. 4C). Accordingly, the area under the curve (AUC) was ~40% greater in Otub2+/- and Otub2-/- male mice compared to WT controls (Fig. 4D). The effects of Otub2 deletion on the female mice were a bit more complex. There were no differences in GTT between Otub2+/+ and Otub2-/- female mice, but the Otub2+/- animals exhibited a slight, yet significant higher GTT response (Fig. 4C) that was also evident by a modest, yet significant higher (approximately 20%) AUC (Fig. 4D). These results suggest a positive role for Otub2 in the improvement of islet functionality under physiological conditions in an in vivo setting, with possible compensatory pathways in KO female mice.
To complement and validate the above findings that were based upon Otub2 KO mouse models, we used a second, independent model in which Otub2 was selectively overexpressed in the pancreas of C57BL wild-type mice. To introduce the Otub2 gene into the pancreas, we made use of adeno-associated viral constructs (AAVs) that are considered a favorable delivery system to express exogenous proteins due to their low immunogenicity and excellent safety profile [29]. C57BL wild-type mice were subjected to pancreatic intraductal administration of AAV8 constructs containing Flag (control) or Flag-Otub2 sequences (Supplementary Fig. 2A). As shown in Supplementary Fig. 2B, islets derived from mice, subjected to intraductal injection of AAV8-Flag-Otub2, showed a strong fluorescent signal, while a much weaker signal was observed in mice injected with AAV8-Flag viral control vector. We further validated these findings by extracting RNA from the pancreases and performing qRT-PCR (Supplementary Fig. 2C). These results verified the efficacy of increasing Flag-Otub2 expression in pancreases, by intra-ductal AAV8 injection.
Glucose-tolerance tests in these mice revealed that basal glucose levels in
AAV8-Flag-Otub2 mice were almost identical to the AAV8-Flag control animals,
however, blood glucose levels of Flag-Otub2 mice were significantly lower
(approximately 30%) at 15–45 minutes post glucose injection (Fig. 5A).
Accordingly, the AUC decreased by 20% in AAV8-Flag-Otub2 injected mice compared
with control animals (Fig. 5B). These findings further support our conclusion that
Otub2 exerts favorable functional effects on
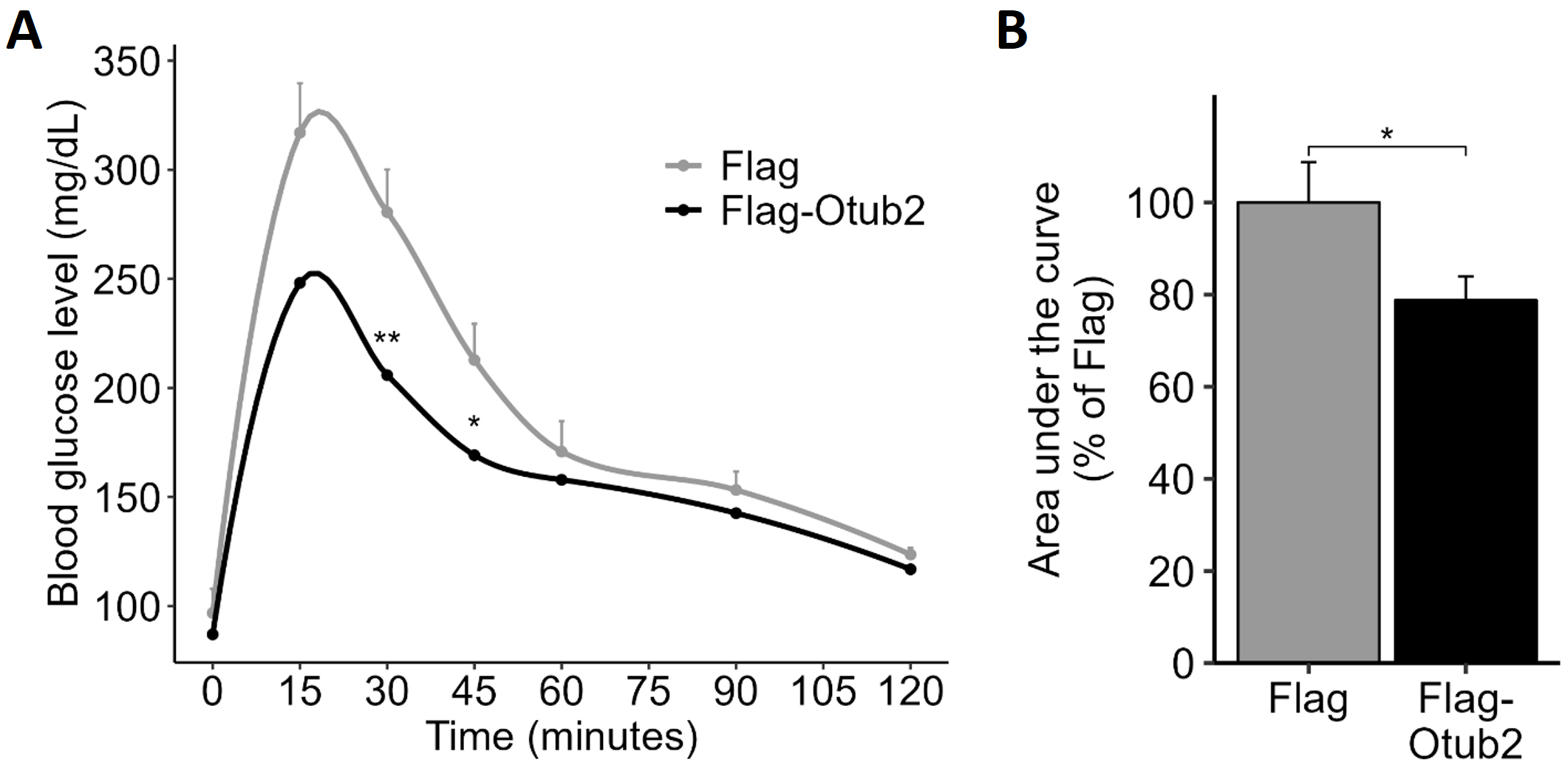 Fig. 5.
Fig. 5.
Effects of intra-ductal injection of AAV8-Flag-Otub2 vector on
glucose tolerance. (A) C57BL WT, 10 weeks old, male mice, were anesthetized and
subjected to retrograde pancreatic intraductal AAV administration. After two
weeks of recovery and overnight fasting, mice were subjected to an
intraperitoneal (i.p.) glucose tolerance test (GTT; 1.8 g D-glucose per kg body
weight). Blood samples were taken at the indicated time points (0–120 minutes),
and glucose levels were determined by a Glucometer. (B) Area under the curve of
the GTT. Data represent means
To reveal the genes whose expression is affected upon Otub2 deletion, RNA was
extracted from the pancreases of Otub2-/-, Otub2-/+, and
Otub2+/+ (WT) mice, and RNAseq analysis was performed. The expression levels
of Otub2 were indeed below detection in most Otub2-/- and Otub2+/-
pancreases (Supplementary Fig. 3A). Both heterozygous and homozygous
pancreases showed significantly different patterns of gene expression compared to
WT pancreases, as shown by the volcano plots (Supplementary Fig. 3B). Of
note, more than 20 genes were significantly up- or down regulated, both in
heterozygous and homozygous mice when compared to wild-type animals
(Supplemental Table 2). Enrichment analysis of curated signatures
employing the MsigDB database and the Camera method [30] revealed several
gene-sets that were significantly upregulated (Supplementary Fig. 3C).
Most significant were the MOOTHA_VOXPHOS gene families [31], involved in
oxidative phosphorylation. Other relevant families were WANG_NFKB_TARGETS,
which includes NF-
To gain a deeper insight into the gene networks affected by Otub-2 depletion,
pathway enrichment analysis was carried out using the GO database. Such analysis
revealed that potassium ion transport was significantly affected in
Otub-2-/- and Otub-2-/+ mice (Fig. 6A,B), with several genes related
to this pathway being down-regulated in OTUB2 KO animals (Fig. 6B). One is Ank2
that encodes the ankyrin-2 (AnkB) protein, which is essential for localization
and membrane stabilization of potassium channels. Other relevant genes were
(potassium voltage-gated channel subfamily C member 3) KCNC3, encoding a
voltage-dependent potassium channel involved in insulin secretion [35], and
Cacna1a, encoding the
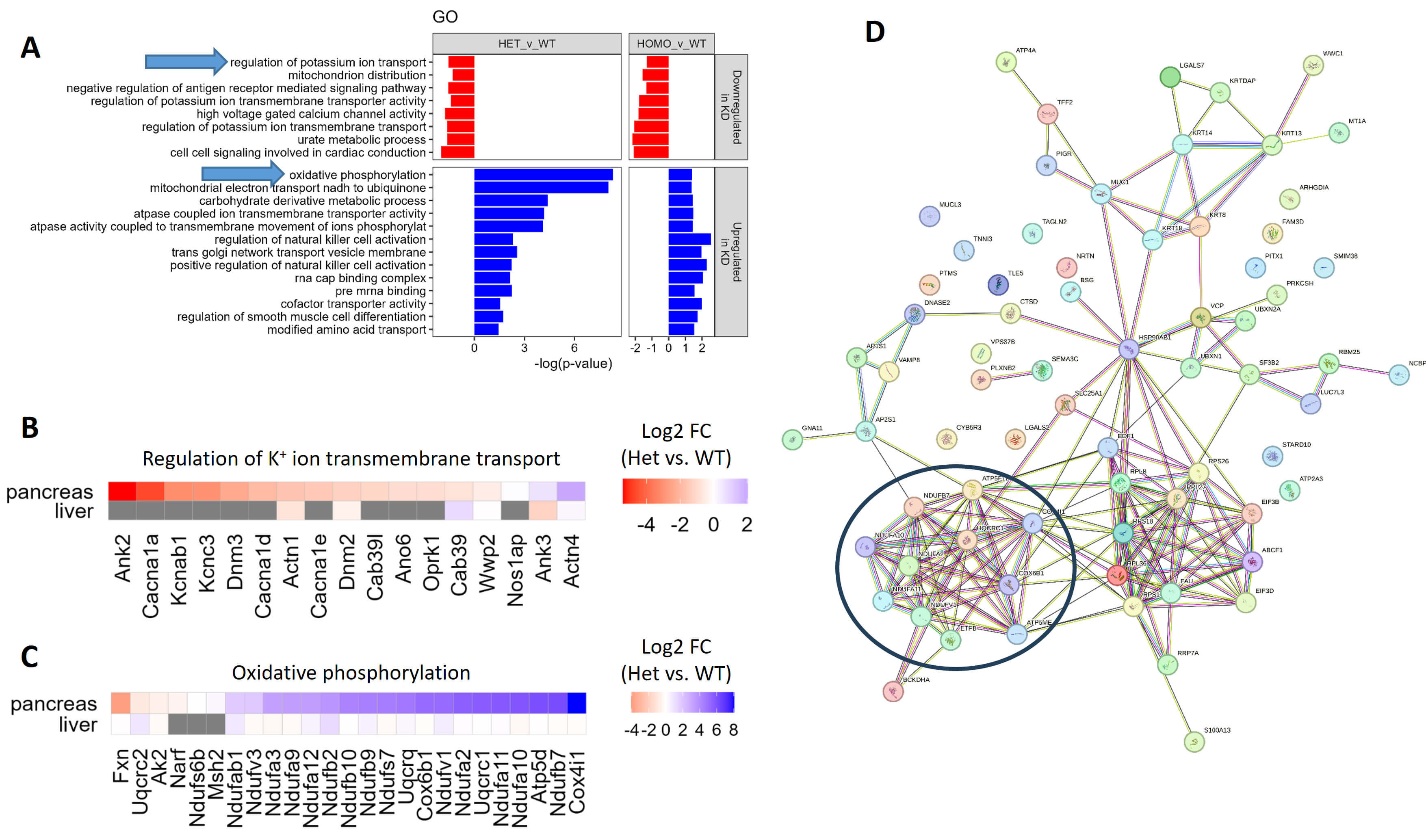 Fig. 6.
Fig. 6.
RNA-seq analysis of pancreas from OTUB2 KO mice. Pancreases and livers were extracted from OTUB2+/+ (WT, n = 3), OTUB2+/- (Heterozygotes, n = 5) and OTUB2-/- (Homozygote, n = 3) mice. RNA was extracted, and RNA-seq was performed to determine differential gene expression between the homozygotes/heterozygotes and the WT mice. (A) Geneset enrichment of gene ontology (GO) terms, performed by the CAMERA method. Arrows indicate interesting GO terms. Analysis is based on gene expression in the pancreas. (B,C) Heatmaps representing the fold change (log2) in gene expression between heterozygotes and WT mice in the pancreases and livers. Genes related to potassium ion transport (B) and oxidative phosphorylation (C) are taken from the respective GO terms. (D) STRING analysis of genes significantly upregulated in Otub2+/- vs. Otub2+/+ pancreases. The black circle depicts a cluster of genes associated with oxidative phosphorylation.
Conversely, genes related to oxidative phosphorylation were mostly upregulated in heterozygous and homozygous Otub2 KO mice (Fig. 6A,C), in accordance with the increased enrichment of the MOOTHA_VOXPHOS geneset (Supplementary Fig. 3C,D). Independently, STRING analysis revealed a cluster of oxidative phosphorylation-related genes among those genes that were significantly upregulated in pancreata of Otub-2-/+ mice (Fig. 6D).
Given that
The alterations in the transcriptomic landscape of pancreata depleted of Otub2 were quite unique, as largely different alterations in transcriptomic landscapes were observed in livers of heterozygous and homozygous Otub2 KO mice when compared to livers of WT animals (Supplementary Fig. 4A,B). Still, pathway enrichment analysis revealed that oxidative phosphorylation genes, as well as components of the ETC complexes in the mitochondria, were enriched in livers as well (Supplementary Fig. 4C). Although these hepatic genes were not upregulated to the same extent as they did in the pancreas (Fig. 6C, Supplementary Fig. 3D), our results suggest that knockdown of Otub2 ubiquitously upregulates genes involved in oxidative phosphorylation and mitochondrial ETC.
We have previously shown that down-regulation of Otub2 by siRNAs in MIN6 cells
and human islets increased caspase-3/7 activity, reduced GSIS, and elevated
expression of NF-
Several lines of evidence support this conclusion. First, we showed that
overexpression of Otub2 decreases NF-
At the molecular level, earlier work has shown that the level of ubiquitination
of TRAF6, which is involved in cytokine-induced NF-
The physiological relevance of Otub2 as a regulator of
Several mechanisms could account for the impaired glucose tolerance in
Otub2+/- and Otub2-/- animals. Most likely, these effects could be
attributed to the inhibitory effects of Otub2 on NF-
Yet, Otub2 seems to exert additional effects that might be only partially
related to NF-
The beneficial effects of Otub2 under physiological conditions were further
illustrated when its overexpression in beta cells was studied. We found that
overexpression of Otub2 promotes insulin secretion in cultured
A broader perspective concerning the physiological functions of Otub2 was gained by analyzing alterations in the transcriptomic landscape of pancreata depleted of Otub2. While the use of whole-body depletion is a limitation of this study, it is obvious that the effects of Otub2 on the transcriptomic landscape seem to be specific, as different gene sets are affected when we compare liver to pancreata in wild type vs. Otub2 knockout animals.
Concerning the pancreas, the analysis highlighted a significant reduction in the expression of genes that regulate potassium ion channel transport, which directly influences insulin secretion [48]. Of interest, one of these family members, Kv9.3, which is expressed in human pancreatic islets [28], forms complexes with Otub2. This channel is involved in repolarization of excitable cells; blocking the activity of this delayed-rectifier potassium channel is expected to increase intracellular free calcium and promote GSIS [28]. Potassium channels need to be closed for proper insulin secretion, in a process initially mediated by ATP [48]. Therefore, activating mutations in potassium channels may induce adult and neonatal diabetes [49]. Indeed, impaired expression of the potassium channel regulator AnkB, observed in Otub2 KO mice, impairs insulin secretion and induces diabetes [50]. Similarly, downregulation of the dynamin genes Dynamin 2 (DNM2) and Dynamin 3 (DNM3) [51], observed in Otub2 KO mice, attenuates internalization of potassium channels on one hand, while inhibiting exocytosis of insulin granules [52, 53]. Hence, maintenance of higher concentrations of potassium channels at the plasma membrane of Otub2 KO mice might impair glucose-stimulated insulin secretion, which is the characteristic feature of these animals. This model, linking Otub2 to GSIS via regulation of potassium channels, still merits further experiments for its establishment.
In line with this hypothesis, knockdown of Otub2 promotes transcription of
oxidative phosphorylation pathways, with the NDUF (NADH:ubiquinone oxidoreductase
subunits) gene-family members being the targets. These proteins are components of
complex I in the mitochondrial electron transport chain [54]. Hence, increased
ATP production in the KO animals might be a compensatory mechanism utilized by
these mice to foster closure of the highly abundant potassium channels and thus
promote insulin secretion. Yet, overexpression of NDUF gene-family members,
particularly when it disrupts the balance of mitochondrial complex 1 activity,
can induce oxidative stress, which is detrimental for
Our findings highlight the role of Otub2 as an anti-apoptotic regulator of
The paper is listed as, “The Deubiquitinating Enzyme Otub2 Modulates Pancreatic Beta-Cells Function and Survival” as a preprint on bioRxiv at: https://www.biorxiv.org/content/10.1101/2024.09.30.615641v1.
The RNAseq generated during the current study is available in the Gene Expression Omnibus database under accession number GSE285973, and can be viewed in: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE285973. The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
MO, RI, SB-H, EC and YV performed the studies and analyzed the data. SS, SL, FB, YV, and YZ contributed to the design of the experiments and drafting the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
All in-vivo experiments were carried out in accordance with ARRIVE guidelines. Experiments were approved by the Animal Care and Use Committee of the Weizmann Institute of Science (application number 23980116-3). Human islets studies received Ethics approval from the Bioethics and Embryonic Stem Cell Research Oversight Committee (ESCRO) of the Weizmann Institute of Science (date: 21 May 2015). The study was conducted in accordance with the principles of the Declaration of Helsinki.
We thank Dr. Eythan Elhanany for insightful comments and discussions. We thank the JDRF award 31-2008-413 (ECIT Islet for Basic Research program) for providing Human islets.
This work was supported by grants from the Juvenile Diabetes Research Foundation International (17-2013-442) and the Israel Science Foundation (759/09). FB work was supported by grant from the Ministerio de Educación y Competitividad, Plan Estatal de I+D+I (SAF2014-54866-R), Spain.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/FBL44406.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






