1 Department of Lung Cancer Surgery, Tianjin Medical University Cancer Institute and Hospital, National Clinical Research Center for Cancer, Key Laboratory of Cancer Prevention and Therapy, Tianjin’s Clinical Research Center for Cancer, 300060 Tianjin, China
2 Department of Thoracic Oncology, Jilin Cancer Hospital, 130012 Changchun, Jilin, China
3 Public Experimental Center, Changchun University of Chinese Medicine, 130017 Changchun, Jilin, China
Abstract
Turmeric-derived exosome-like nanoparticles (TELNs) are nanoscale vesicles of plant origin with therapeutic potential. However, the specific efficacy and mechanisms of TELNs in inhibiting non-small cell lung cancer (NSCLC) remain unclear. This study investigated the effects of TELNs on NSCLC by epigenetically regulatiing histone acetyltransferase human males absent on the first (hMOF) and histone H4K16 acetylation (H4K16ac).
TELNs were isolated from turmeric using differential centrifugation and characterized by nanoparticle tracking analysis (NTA), transmission electron microscopy (TEM), and zeta potential measurements. Cellular uptake was assessed via PKH26 labeling. In vitro assays evaluated the effects of TELNs on apoptosis (annexin V/PI staining, JC-1 mitochondrial depolarization, caspase-3 cleavage) and proliferation (CCK-8). The in vivo efficacy of TELNs was examined in A549 xenografts. Bioinformatics and molecular docking analyses revealed the interaction of curcumin in TELNs with hMOF, while RNA interference validated the role of hMOF in TELN-mediated apoptosis and migration suppression.
TELNs exhibited exosome-shaped morphology and efficient uptake by A549 cells. Treatment with TELNs induced apoptosis and reduced tumor volume by 58.1%. Mechanistically, TELNs upregulated hMOF expression and H4K16ac levels. RNA interference confirmed that knockdown of hMOF weakened the effect of the TELNs. Molecular docking suggested curcumin in TELNs may interact with hMOF.
This study reveals a novel epigenetic mechanism wherein TELNs suppress NSCLC by activating hMOF/H4K16ac. Curcumin within TELNs increases hMOF levels, thus positioning TELNs as a potential nanotherapeutics with the capacity for epigenetic modulation. Our findings underscore the potential of TELNs in NSCLC treatment and highlight hMOF as a therapeutic target.
Keywords
- exosome
- curcumin
- non-small cell lung cancer
- epigenesis
- histone acetylation
Non-small cell lung cancer (NSCLC) has the highest incidence and mortality rates of all cancer types [1]. Current therapeutic strategies include surgery and radiotherapy. However, surgical resection is unsuitable for frail patients, and incomplete tumor removal may lead to recurrence and metastasis. Radiotherapy induces tumor cell apoptosis by damaging DNA molecules via X-rays or proton beams [2]. However, it also exerts detrimental effects on adjacent healthy tissues and may cause pulmonary fibrosis in the long term [3]. Compared to conventional chemotherapeutic agents, natural products offer low toxicity, high biocompatibility, and the ability to suppress NSCLC through multi-pathway modulation [4].
Plant-derived exosome-like nanoparticles (PELNs) are nanoscale vesicles secreted by plant cells, ranging in diameter from 30 to 160 nm [5]. PELNs share structural similarities with mammalian exosomes, and their unique composition enhances antitumor efficacy [6]. These nanoparticles carry anti-cancer components such as microRNAs and flavonoids, significantly inhibiting tumor growth by inducing apoptosis and cell cycle arrest [7, 8]. The phospholipid bilayer of PELNs maintains stability under physiological conditions and preserves biocompatibility benefits over synthetic nanocarriers [9]. Turmeric (Curcuma longa L.), a traditional Chinese herb, has been used historically to treat abdominal masses and malignancies [10]. Recent research attributes its anticancer properties to curdione and curcumin, which exhibit anti-metastatic and anti-angiogenic effects [11, 12]. Yang et al. [13] successfully isolated turmeric-derived exosome-like nanoparticles (TELNs) and demonstrated their inhibitory effect on human colorectal cancer cells.
Although TELNs show potential for cancer treatment, their effects on NSCLC and the underlying mechanisms remain unclear. In this study, TELNs were successfully isolated, and their mechanism of action against NSCLC was demonstrated for the first time. Our results demonstrate that TELNs affect the expression and activity of the histone acetyltransferase hMOF. This subsequently increases the acetylation of histone H4 at lysine 16 (H4K16ac), thereby suppressing NSCLC epigenetically. We initially investigated the in vitro and in vivo antitumor efficacy of TELNs, as well as the specific regulation of hMOF. Molecular docking also revealed the potential interaction between TELNs and hMOF. Our findings establish, for the first time, that TELNs exert anti-NSCLC effects via hMOF/H4K16ac-mediated epigenetic regulation. This provides a foundation for the use of natural nanoparticles to develop therapies based on epigenetic reprogramming.
Cell Counting Kit-8 (CCK-8, AC11L054) was purchased from Life-iLab Biotech Co.,
Ltd. (Shanghai, China). The Annexin V/PI staining kit (FAK012.20) was purchased
from NeoBioscience Technology Co, Ltd. (Shenzhen, China). JC-1 staining kit
(AC13021) was sourced from Acmec Biochemical Technology Co., Ltd. (Shanghai,
China). The BCA kit was purchased from Beyotime Biotechnology Co., Ltd.
(Shanghai, China). Dulbecco’s Modified Eagle Medium (DMEM), Roswell Park Memorial
Institute (RPMI) 1640 medium, fetal bovine serum (FBS) and trypsin were acquired
from Gibco (USA). Anti-hMOF antibody (ab200660), anti-H4K16ac antibody
(ab109463), anti-histone H4 antibody (ab7311), anti-procaspase-3 antibody
(ab32150), and anti-cleaved caspase-3 antibody (ab32042) were purchased from
Abcam (Waltham, MA, USA). Anti-
Human lung cancer A549 and H1299 cells were purchased from Procell Biotechnology (Wuhan, China) and cultured in DMEM supplemented with 10% FBS and 1% Penicillin-Streptomycin, under an atmosphere of 5% CO2 at 37 °C. All cell lines were validated by STR profiling and tested negative for mycoplasma.
TELNs were isolated through multiple centrifugation steps and density gradient
centrifugation as described earlier [14]. Fresh turmeric was washed three times
with distilled water. The roots were then crushed in a juicer. Subsequently, the
mixture was centrifuged at 1000 g for 10 minutes, followed by 3000 g for 20
minutes, and finally at 10,000 g for 30 minutes to collect the supernatant. The
supernatant was filtered through a sterile 0.45 µm filter and placed in a
fixed-angle rotor for centrifugation at 100,000 g for 1 h. The pellet was
resuspended in 1 mL of PBS and subjected to sucrose gradient centrifugation at
different concentrations (15%, 30%, 45%, and 60%) at a speed of 150,000 g for
2 h. The layer at the sucrose concentration of 30%–45% was collected, and an
equal volume of PBS was added to wash away the sucrose. Finally, the mixture
underwent centrifugation at 150,000 g for 1 h to collect the pellet, which was
then resuspended in PBS as TELNs. The protein concentration of TELNs was measured
with the bicinchoninic acid (BCA) assay. To verify that TELNs were successfully
isolated, their structure was observed by transmission electron microscopy (TEM).
Using a capillary pipette, one drop of the prepared suspension sample (5 µL
of TELNs) was placed onto a copper grid coated with a support film. Excess liquid
was absorbed with filter paper. Subsequently, a 3% phosphotungstic acid solution
was added, and staining took place for 5 min. The staining solution was removed
by filter paper, the sample was dried, and the morphology of TELNs was then
examined and photographed by TEM (Jeol1230, JEOL Ltd., Peabody, MA, USA). The
particle size distribution of TELNs was characterized by nanoparticle tracking
analysis (NTA, PMX 110, ZetaView, Inning am Ammersee, Germany). Dynamic light
scattering (DLS, ZS90, Malvern Panalytical Ltd., Worcestershire, UK) was utilized
to measure the
PKH26-labeled TELNs were first prepared [15]. Briefly, 2 µM PKH67 dye and
1 mL Diluent C were mixed as a working solution. TELNs (50 µg) were then
incubated with the working solution for 20 min at room temperature, and the
labeled-TELNs were ultracentrifuged (100,000 g, 2 h) to remove free PKH26 dye.
Subsequently, A549 cells (1
The cytotoxicity of TELNs against A549 cells was analyzed by CCK-8 assay [15].
Specifically, A549 cells (5
TELNs-induced apoptosis was assessed by Annexin V/PI staining [15]. Specifically, after incubation with TELNs (0, 10, 50, and 100 µg/mL) for 24 h, cells were stained with the kit for 15 min and then analyzed by flow cytometry. Annexin V+/PI– cells were considered to be early apoptotic cells, while Annexin V+/PI+ cells were considered late apoptotic cells.
A549 cells were seeded into 6-well plates (2
Female nude mice (6-week-old) with a BALB/c genetic background were obtained
from Liaoning Changsheng Biotechnology Co., Ltd. All animal experiments were
conducted according to the guidelines of the Experimental Animal Welfare Ethics
Committee of Changchun University of Chinese Medicine (2025936). The mice
underwent a one-week acclimatization period before the establishment of the A549
hormonal nude model. Mice were anesthetized with isoflurane (induction
concentration 3%, maintenance concentration 1%). After confirmation of
anesthesia, 200 µL of an A549 cell suspension (5
A SteadyPure Universal RNA extraction kit (AG21017, ACCURATE BIOTECHNOLOGY, Changsha) was used to obtain total RNA from the tumor and A549 cells. All-in-One cDNA Synthesis SuperMix (TransGen Biotech Co., Ltd., Beijing, China) was used for reverse transcription. The quantitative real-time PCR mixture was formed with SYBR Green qPCR Master Mix for the Real-Time PCR System (QuantStudio 3, Thermo Fisher Scientific, Waltham, MA, USA). Relative changes in histone acetyltransferase human males absent on the first (hMOF) and GAPDH (Glyceraldehyde-3-Phosphate Dehydrogenase) mRNA levels were then measured. The primers [18] are shown in Table 1.
| Primer name | Sequence |
| hMOF forward | 5′-GCTGGACGAGTGGGTAGACAA-3′ |
| hMOF reverse | 5′-TGGTGATCGCCTCATGCTCCTT-3′ |
| GAPDH forward | 5′-ATCACTGCCACCCAGAAGAC-3′ |
| GAPDH reverse | 5′-ATGAGGTCCACCACCCTGTT-3′ |
PCR, Polymerase Chain Reaction; hMOF, human males absent on first; GAPDH, Glyceraldehyde-3-Phosphate Dehydrogenase.
A549 cells were transfected with siRNA oligonucleotides using Lipofectamine 2000 (Thermo Fisher Scientific, Waltham, MA, USA). The hMOF siRNA (5′-GAAAGAGAUCUACCGCAATT-3′) [19], Alexa Fluor 488 modified negative control siRNA (Cat no. 1027292), and unmodified negative control siRNA (Cat no. 1027280) were obtained from QIAGEN (USA). After transfection with siRNA, the cells were harvested for further analysis.
A549 cells were cultured in 6-well plates at a density of 5
(Ⅰ) Curcumin target search. (ⅰ) Single-drug search: the SMILES chemical formula for the active ingredient of curcumin (CID: 969516) was obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). The results were imported into the Swiss Target Prediction database (https://www.swisstargetprediction.ch/) to identify the target, and the target name was converted to gene name in the UniProt protein database (https://www.uniprot.org/). (ii) Herbal search: the target (sequence number: MOL000090) was retrieved from the TCMSP online database (https://www.tcmsp-e.com/), using curcumin as the keyword in the Chemical name entry. Its structure file was downloaded and imported into the PharmMapper database (https://www.lilab-ecust.cn/pharmmapper/) to retrieve the target moiety (Job ID: 250112151602). Target names were then converted to gene names in the UniProt protein database (https://www.uniprot.org/). The results from (i) and (ii) were combined to obtain the curcumin targets. (Ⅱ) Target search in NSCLC. Using “non-small cell lung cancer” as the keyword, NSCLC-related targets were identified in the GeneCards (https://www.genecards.org/), OMIM (https://omim.org/), and Disgenet (https://disgenet.com) databases. The results were combined to remove duplicates and identify the targets. Finally, the intersecting targets for curcumin (Ⅰ) and NSCLC (Ⅱ) were determined.
The intersecting targets were imported into the STRING protein interactions database (http://string-db.org/), with the species set to homo sapiens and the Interaction score set to 0.400. The interaction proteins were screened, and the PPI network was constructed, with the results visualized using Cytoscape 3.10.3 software.
(I) Intersecting target proteins were mapped to each term of the Gene Ontology
(GO) database (http://www.geneontology.org/), and the protein number in each term
was determined. GO terms that satisfied a p-value
(II) The intersecting targets were mapped to pathways using the Kyoto
Encyclopedia of Genes and Genomes (KEGG) database (https://www.genome.jp/kegg/).
Significantly enriched KEGG pathways were identified as having a p-value
Curcumin and hMOF protein were selected for molecular docking analysis. The 3D structure for the active ingredient of curcumin (CID: 969516) was obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). and that of hMOF protein (ID: pdb_00004dnc) from the PDB database (http://www.rcsb.org/). Water and ligand were removed from the receptor with PyMOL 3.1 (Schrödinger Inc., New York City, NY, USA), then converted to pdbqt format using Auto Dock 4.2.6 (Scripps Research Institute, Claremont, CA, USA). Appropriate positions were selected for docking. Binding energies were calculated and the results were visualized using PyMOL 3.1 (Schrödinger Inc., New York City, NY, USA).
Cells were lysed using RIPA lysis buffer (89900, Thermo Fisher Scientific,
Waltham, MA, USA) containing a 1% protease inhibitor mixture (87785, Thermo
Fisher Scientific, Waltham, MA, USA). The protein concentration was subsequently
determined using a BCA protein assay kit (P0012, Beyotime Biotechnology Co.,
Ltd., Shanghai, China). Protein samples were separated by 10% sodium dodecyl
sulfate polyacrylamide gel electrophores (SDS-PAGE) gel electrophoresis
(XP00100BOX, Thermo Fisher Scientific, Waltham, MA, USA) and transferred to
polyvinylidene fluoride (PVDF) membranes (Millipore). The membrane was then
blocked for 1 h in Tris-buffered saline (TBS) containing 5% skim milk at room
temperature, followed by overnight incubation at 4 °C with the primary
antibody: Anti-hMOF antibody (1:1000; ab200660, Abcam), anti-H4K16ac antibody
(1:1000; ab109463, Abcam), anti-histone H4 antibody (1:1000; ab7311, Abcam),
anti-procaspase-3 antibody (1:1000; ab32150, Abcam), anti-cleaved caspase-3
antibody (1:1000; ab32042, Abcam), anti-
Results are presented as the mean
To obtain TELNs, turmeric was juiced and then subjected to differential
centrifugations and density gradient centrifugation (Fig. 1A). The physical
properties of the TELNs were subsequently characterized. The size distribution of
TELNs was measured using nanoparticle tracking analysis (NTA). TELNs showed a
particle size of 157.2 nm (Fig. 1B), and their
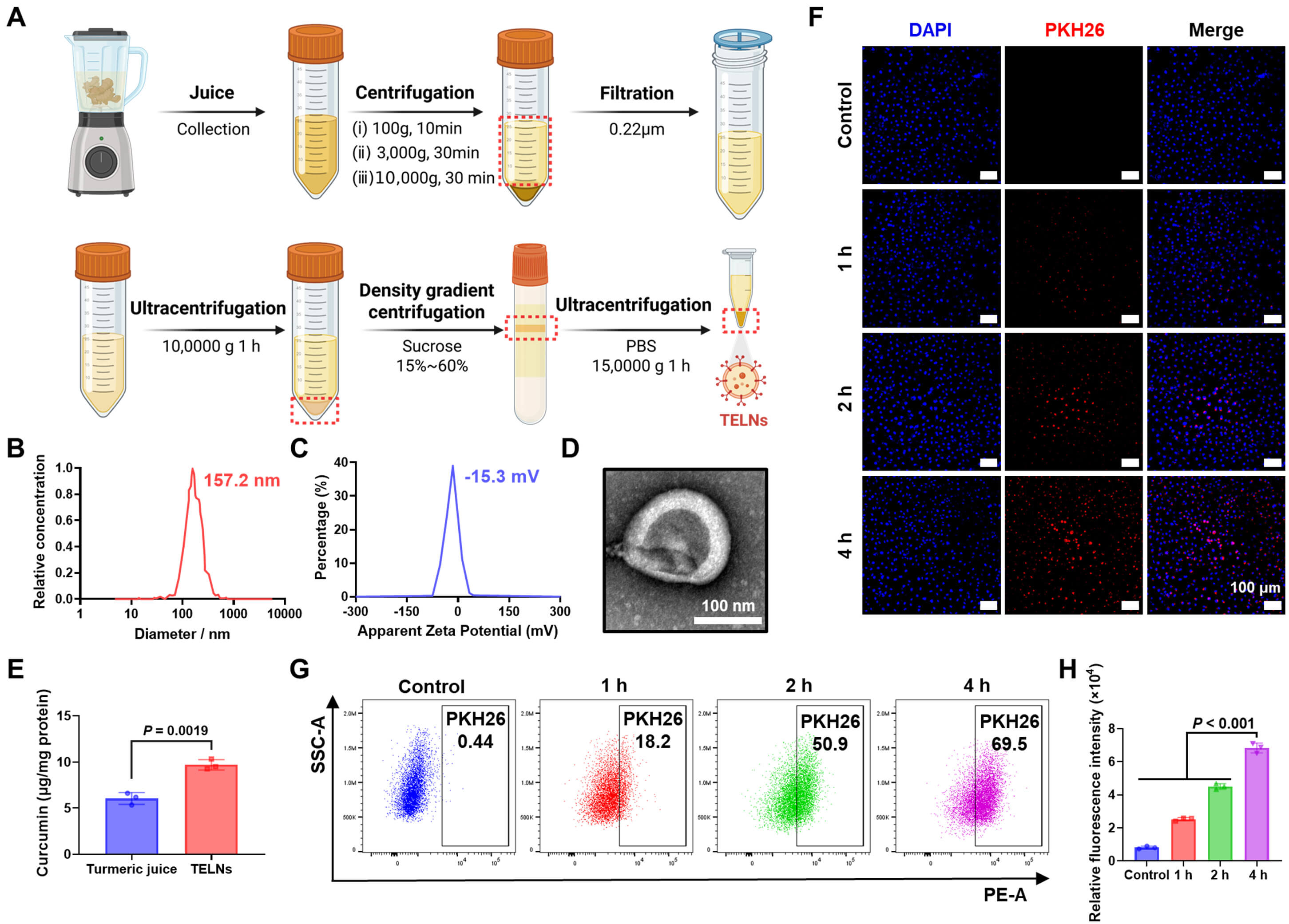 Fig. 1.
Fig. 1.
Extraction, characterization, and uptake analysis of
turmeric-derived exosome-like nanoparticles. (A) Procedure for extracting TELNs.
(B) Nanoparticle tracking analysis (NTA) of TELNs, showing a particle size of
157.2 nm. (C) Zeta potential analysis of TELNs, indicating a
We first assessed the effect of TELNs on apoptosis in NSCLC. Flow cytometry analysis revealed the apoptosis rate of A549 cells treated with 100 µg/mL TELNs was 4.54–fold higher compared to the control group, with late-stage apoptosis increasing from 1.02% to 29.60% (Fig. 2A,C). Additionally, 100 µg/mL TELNs increased the apoptosis rate of H1299 cells by 45.46% (Supplementary Fig. 5). Consistent with these observations, CCK-8 assays revealed dose-dependent anti-NSCLC activity of TELNs, with a 45.25% reduction in cell viability at 100 µg/mL (Fig. 2E). The same phenomenon was also observed in H1299 cells, where TELNs demonstrated anticancer activity. At a dose of 100 µg/mL, the viability of H1299 cells decreased by 44.67% (Supplementary Fig. 6). The JC-1 probe is commonly used to detect the membrane potential of mitochondria, which is closely related to apoptosis. When apoptosis occurs, the mitochondrial membrane is destroyed, and JC-1 changes from an aggregated state to a dispersed state. Thus, enhanced green fluorescence (JC-1 monomers) implies increased apoptosis. TELNs significantly induced mitochondria-related apoptosis in A549 cells (Fig. 2B,D). The expression of cleaved caspase-3 protein was examined by Western blot assay to corroborate the presence of apoptosis. As a crucial executor of apoptosis, caspase-3 translocates from the mitochondria and induces apoptosis. TELNs significantly increased the ratio of cleaved caspase-3 to pro caspase-3 in A549 cells (Fig. 2F,G). Similarly, TELNs also increased the ratio in H1299 cells (Supplementary Fig. 7). Additionally, real-time quantitative PCR revealed that hMOF mRNA expression increased 2.01–fold in A549 cells (Fig. 2H) and 1.82–fold in H1299 cells (Supplementary Fig. 8) following treatment with TELNs. Collectively, these findings establish TELNs as inducers of apoptosis in A549 cells, with hMOF being a critical factor.
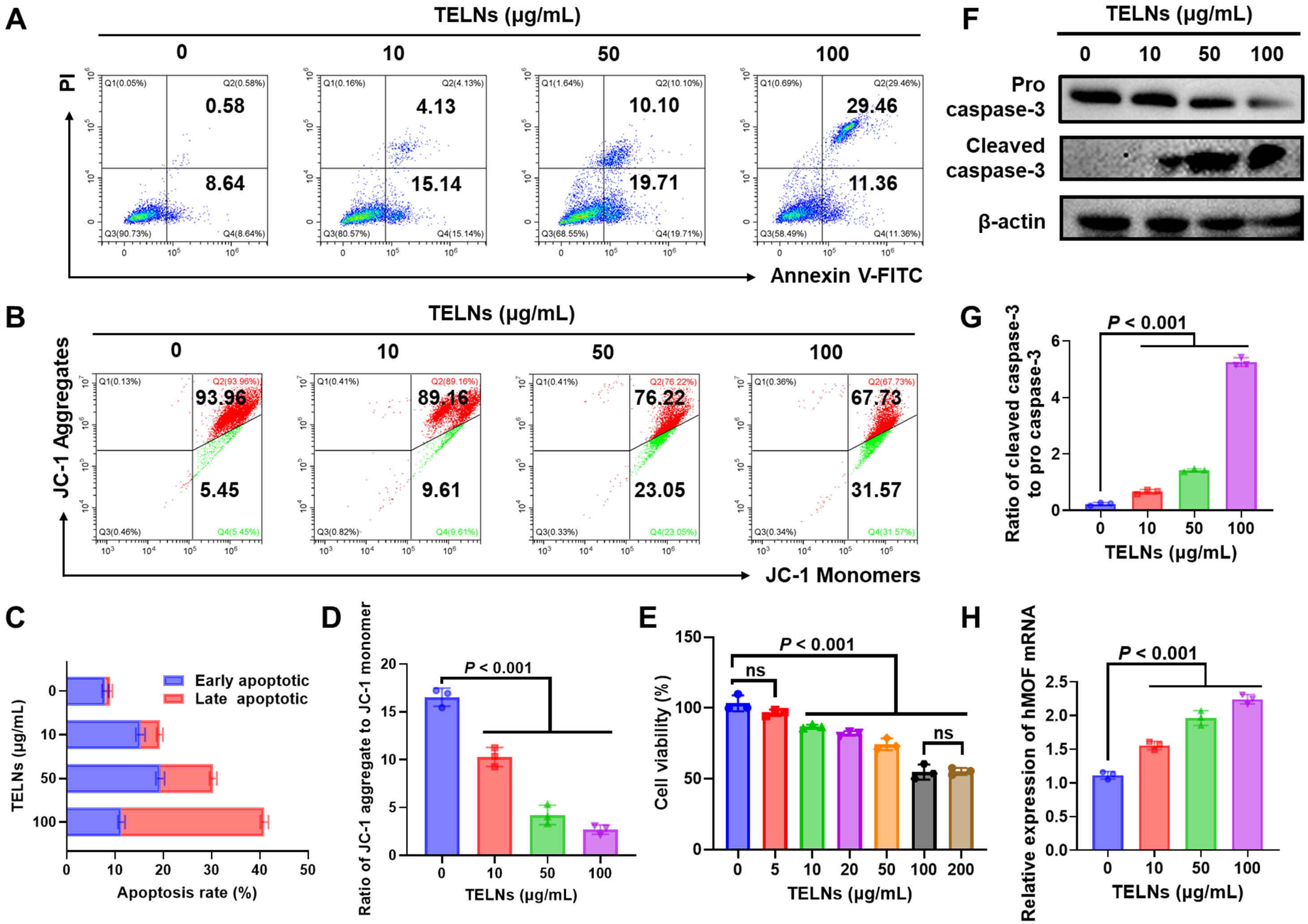 Fig. 2.
Fig. 2.
Turmeric-derived exosome-like nanoparticles induce apoptosis in non-small cell lung cancer. (A) Apoptosis of TELN-treated A549 cells, as measured by flow cytometry. (B) Mitochondrial membrane potential of A549 cells, as detected by flow cytometry. (C) Apoptosis rate of A549 cells (n = 3). (D) Ratio of JC-1 aggregate to JC-1 monomer (n = 3). (E) Cytotoxicity of TELNs against A549 cells, as measured by CCK-8 assay (n = 3), ns indicates that the results are not significant. (F) Western blot analysis of pro and cleaved caspase-3 expression in A549 cells after treatment. (G) The ratio of cleaved caspase-3 to pro caspase-3 (n = 3). (H) hMOF mRNA levels analyzed by real-time PCR (n = 3). JC-1, 5,5,6,6′-tetrachloro-1,1′,3,3′ tetraethylbenzimi-dazoylcarbocyanine iodide; CCK-8, Cell Counting Kit-8; hMOF, human males absent on first; PCR, Polymerase Chain Reaction.
An A549 xenograft nude mouse model was established to evaluate the antitumor efficacy of TELNs in vivo. Compared to the control group, treatment with TELNs resulted in a 58.1% reduction in tumor volume, confirming its potent growth-inhibitory effects (Fig. 3A,B). In tests conducted every three days, treatment with TELNs significantly slowed tumor growth (Fig. 3C). H&E staining of the tumor tissue revealed distinct morphological differences, with PBS-treated tumors displaying uniform cell size, whereas TELN-treated tumors exhibited marked cellular disorganization (Fig. 3D). TUNEL staining showed a significant increase in apoptosis after treatment with TELNs (Fig. 3E,F). Since hMOF is known to catalyze histone H4K16 acetylation (H4K16ac), downstream epigenetic modulation was also examined.
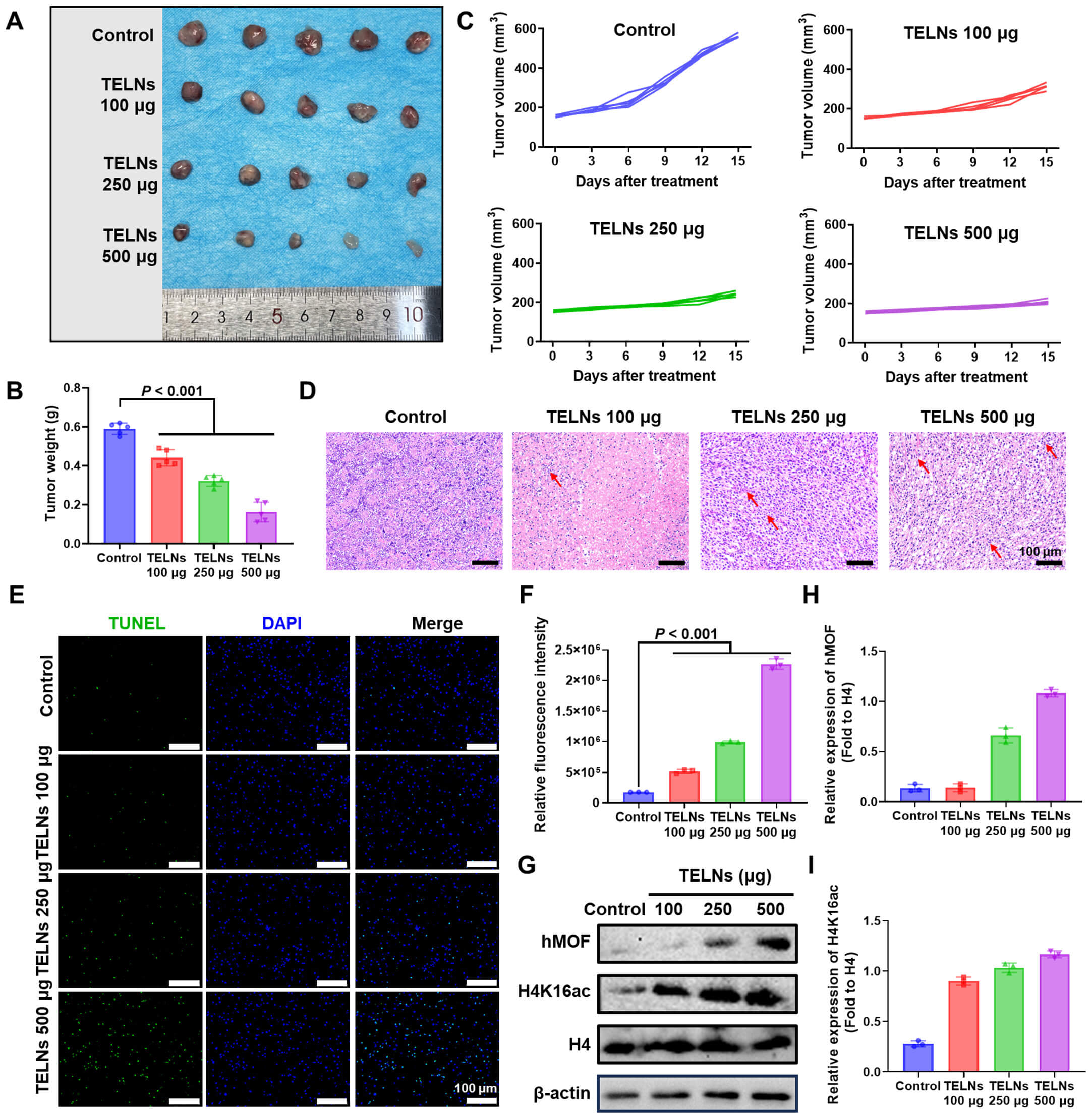 Fig. 3.
Fig. 3.
Turmeric-derived exosome-like nanoparticles inhibit non-small cell lung cancer progression in vivo. (A) Images of tumors from mice after treatment with TELNs (n = 5). (B) Tumor weight after treatment (n = 5). (C) Changes in tumor volume over time (n = 5). (D) H&E staining images of tumor tissue (scale bar: 100 µm; red arrows indicate the cellular disorganization). (E) TUNEL staining of tumor (green: TUNEL; blue: nuclei staining; scale bar: 100 µm). (F) Quantitative analysis of fluorescence intensity (n = 3). (G) hMOF and H4K16ac levels in tumor detected by Western blot assay. (H) The expression of hMOF quantified by ImageJ software (n = 3). (I) The expression of H4K16ac quantified by ImageJ software (n = 3). TUNEL, Terminal deoxynucleotidyl transferase dUTP nick end labeling; H4K16ac, histone H4K16 acetylation.
Western blot analysis of tumor tissues revealed that TELNs significantly increased the levels of hMOF and H4K16ac (Fig. 3G–I). This suggested that hMOf and H4K16ac may serve as the target through which TELNs promote apoptosis in NSCLC. These findings demonstrate that TELNs effectively inhibited NSCLC progression and upregulated hMOF and H4K16ac in vivo.
Building upon previous studies examining the efficacy of TELNs in NSCLC, we conducted further investigations to determine whether its activity depends on the induction of hMOF. RNA interference was employed as a knockdown method, and the AF488-labeled Negative Control demonstrated an siRNA transfection efficiency of 80.4% for this approach (Supplementary Fig. 9). sihMOF effectively reduced hMOF expression by 86.22% (Supplementary Fig. 10). hMOF knockdown substantially reduced JC-1 monomer formation, suggesting impaired apoptosis. Notably, treatment with TELNs partially rescued this effect, demonstrating the role of TELNs in hMOF regulation (Fig. 4A,B). Consistent with the above observations, direct assessment of apoptosis rates revealed that silencing of hMOF decreased cell death by 54.28%, while treatment with TELNs restored apoptosis by 13.13% (Fig. 4C,D). Furthermore, wound healing assays were performed to analyze the effect of TELNs on cell migration. Scratch closure rates were quantified at 12, 24, and 48 h. Knockdown of hMOF enhanced A549 cell migration by 31.70% in 48 h, whereas TELNs partially counteracted the pro-migration effects induced by hMOF downregulation (Fig. 4E,F). These findings indicate that TELNs inhibit NSCLC cell migration by upregulating hMOF expression. Western blot assay further demonstrated suppression of both hMOF and H4K16ac expression after hMOF knockdown. Crucially, TELNs restored H4K16ac levels, consistent with their ability to upregulate hMOF (Fig. 4G–I). Collectively, these data establish the hMOF-H4K16ac axis as a key pathway through which TELNs exert their suppressive effects on NSCLC.
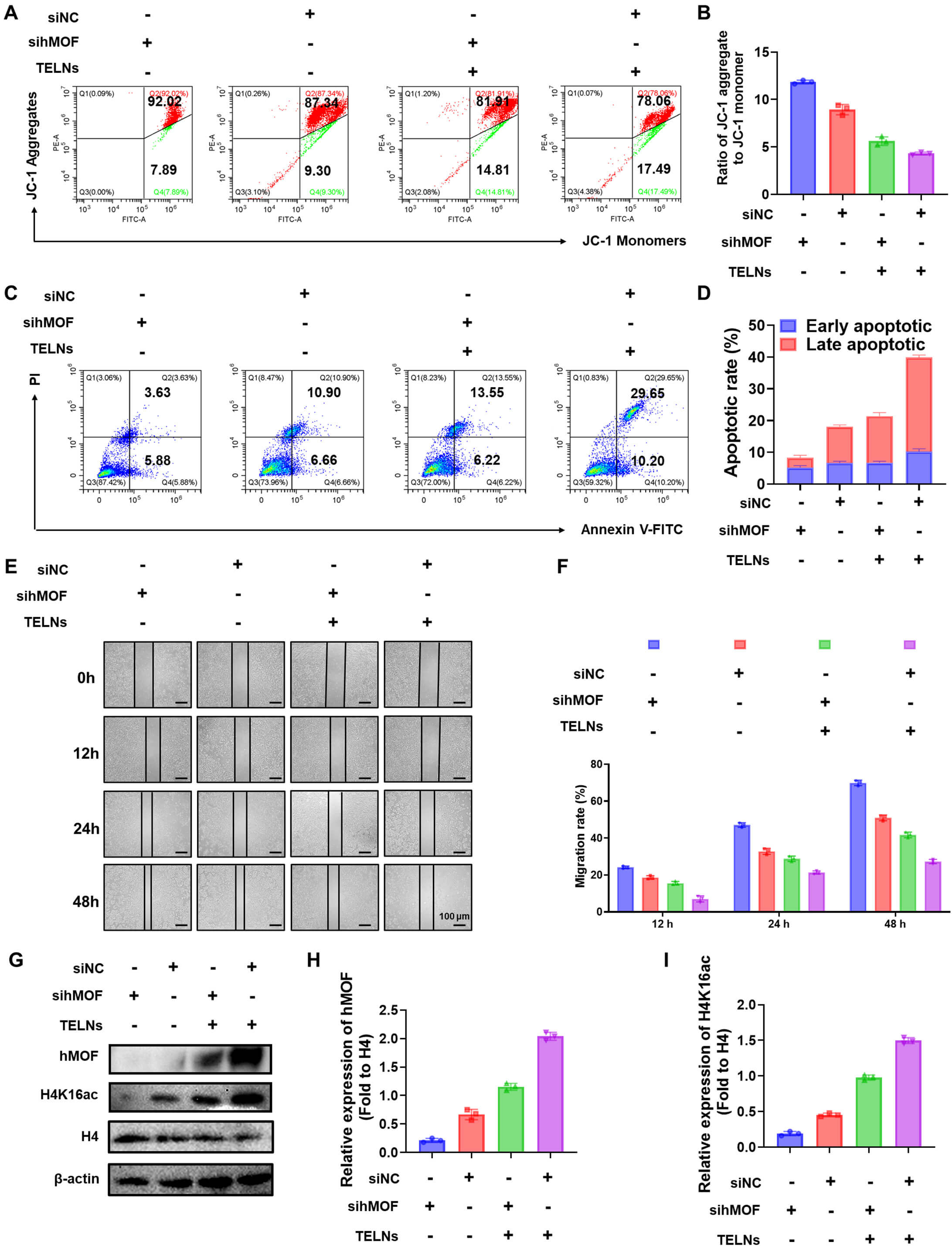 Fig. 4.
Fig. 4.
Turmeric-derived exosome-like nanoparticles inhibit non-small cell lung cancer by upregulating hMOF expression and histone H4K16 acetylation. (A) Mitochondrial membrane potential of A549 cells, as detected by flow cytometry. (B) Ratio of JC-1 aggregate to JC-1 monomer (n = 3). (C) Apoptosis of A549 cells, as measured by flow cytometry. (D) Early and late apoptosis rates of different groups (n = 3). (E) Wound healing assay for A549 cells observed under a microscope at 12 h, 24 h, and 48 h (scale bar: 200 µm). (F) Analysis of cell migration rate (n = 3). (G) Effect of TELNs on hMOF and H4K16ac levels in A549 cells, as detected by Western blot. (H) The expression of hMOF quantified by ImageJ software (n = 3). (I) The expression of H4K16ac quantified by ImageJ software (n = 3). “+”/ “-” indicate whether cells were treated with siRNA or TELNs. siNC, Negative Control siRNA; sihMOF, hMOF siRNA; FITC, Fluorescein isothiocyanate; H4, Histone H4.
As expected, TELNs possess multiple properties of turmeric. In a previous study, Wei et al. [20] demonstrated that TELNs contain a high concentration of curcumin. Curcumin is a natural compound extracted from turmeric that exhibits anticancer properties through its impact on epigenetic modifications [21, 22]. However, its regulatory role in hMOF remains unclear. Therefore, we hypothesized that epigenetic regulation by TELNs is related to the curcumin contained within them. We conducted a systematic bioinformatics analysis to elucidate the molecular mechanism underlying this effect. First, the chemical structure of curcumin (Fig. 5A) was used to search for interacting proteins, which were then compared with NSCLC-related proteins. The database identified 324 curcumin target proteins and 3782 NSCLC-associated proteins, with 191 overlapping targets highlighted by a Venn map (Fig. 5B). The protein-protein interaction (PPI) network was performed using the STRING database (Supplementary Fig. 11), with parameters set to Homo sapiens and the interaction score at 0.400. The network comprised 191 nodes and 920 edges, with an average node degree of 9.63, indicating extensive connections among the proteins. KEGG pathway enrichment analysis revealed these proteins were highly associated with cancer progression, VEGF signaling, PD-L1 immune checkpoint regulation, and the development of NSCLC (Supplementary Fig. 12). Furthermore, GO enrichment analysis (Fig. 5C–E) categorized the overlapping proteins into three functional subsets: (ⅰ) cellular component (CC): localized in the cytoplasm and nucleus; (ⅱ) biological process (BP): involved in histone remodeling, apoptosis, and response to reactive oxygen species (ROS); (ⅲ) molecular function (MF): enriched in DNA binding, histone H4K16 acetylation, and protease activity. The overlapping targets were visualized using Cytoscape software. This revealed significant enrichment of KAT8, KAT6A, KAT6B, KAT5, and KAT7 (all members of the hMOF histone acetyltransferase family), suggesting the anti-tumor activity of curcumin was closely linked to the regulation of histone H4K16 acetylation (Fig. 5F). We next investigated whether curcumin directly interacts with hMOF to modulate its function. Molecular docking analysis was performed using PDB software to simulate the binding between curcumin and KAT8. The calculated binding energy of –6.7 kcal/mol (significantly below the threshold of –5 kcal/mol) strongly supports a stable interaction between curcumin and hMOF (Fig. 5G). Collectively, these results demonstrate that curcumin in TELNs upregulates hMOF expression and H4K16ac in A549 cells. Mechanistically, molecular docking suggested curcumin may interact with hMOF. In brief, curcumin in TELNs promotes H4K16 acetylation by activating hMOF, thereby promoting apoptosis and inhibiting NSCLC. This reveals a new epigenetic regulatory mechanism.
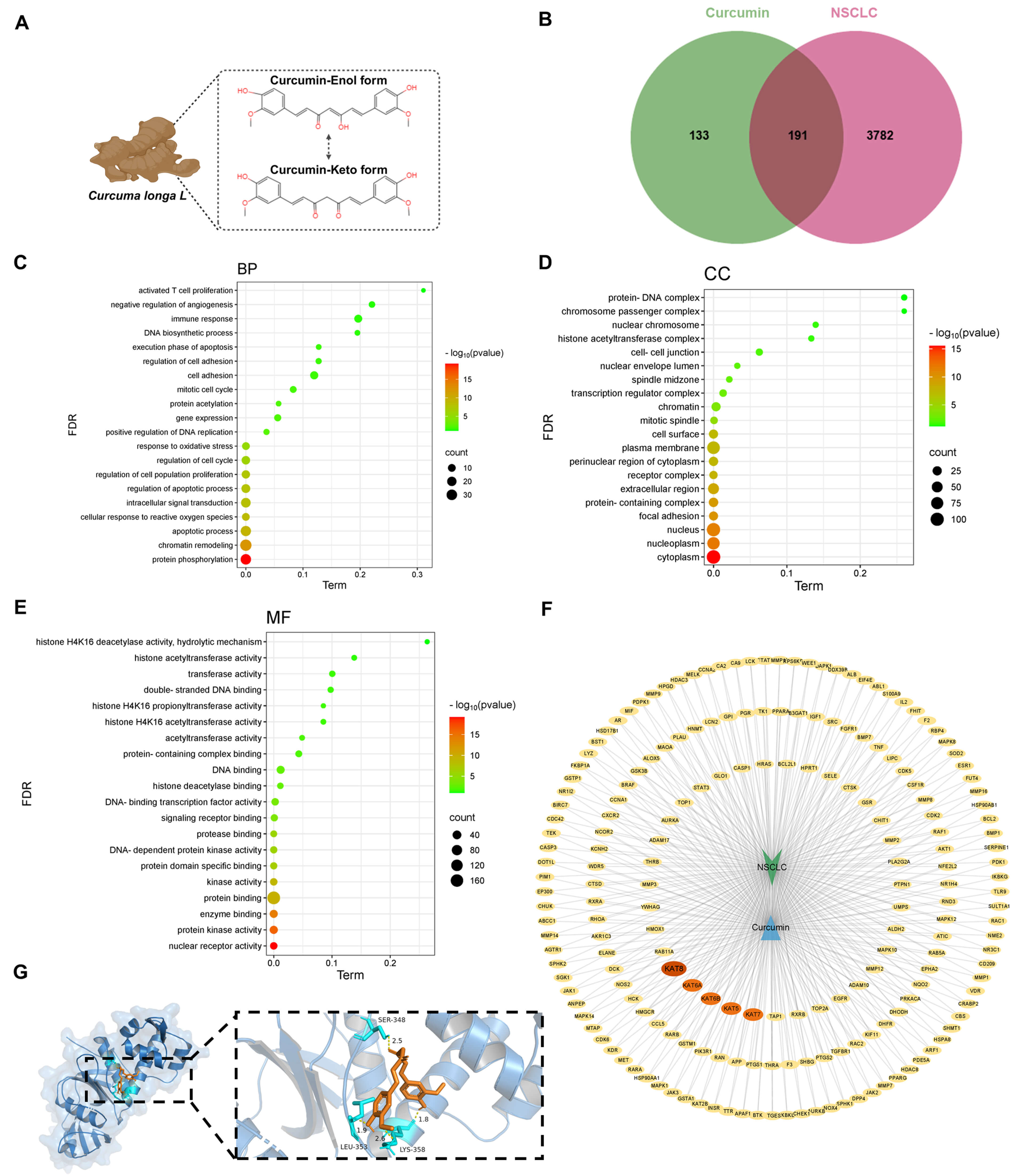 Fig. 5.
Fig. 5.
Epigenetic regulation by turmeric-derived exosome-like nanoparticles is due to curcumin. (A) Chemical structure of curcumin. (B) Venn map showing the intersection of curcumin and NSCLC targets. (C) GO enrichment analysis: biological process. (D) GO enrichment analysis: cellular component. (E) GO enrichment analysis: molecular function. (F) “Curcumin-NSCLC” target protein network. (G) Molecular docking simulation of curcumin with hMOF. BP, biological process; CC, cellular component; MF, molecular function; FDR, False Discovery Rate; NSCLC, non-small cell lung cancer.
This study successfully isolated TELNs and validated their anti-cancer efficacy both in vitro and in vivo. Moreover, a novel epigenetic mechanism was elucidated in which TELNs inhibit NSCLC through hMOF. Bioinformatics and experimental analyses identified hMOF as a pivotal target, with curcumin in TELNs enhancing the levels of both hMOF and histone H4 at lysine 16 (H4K16ac) (Fig. 6).
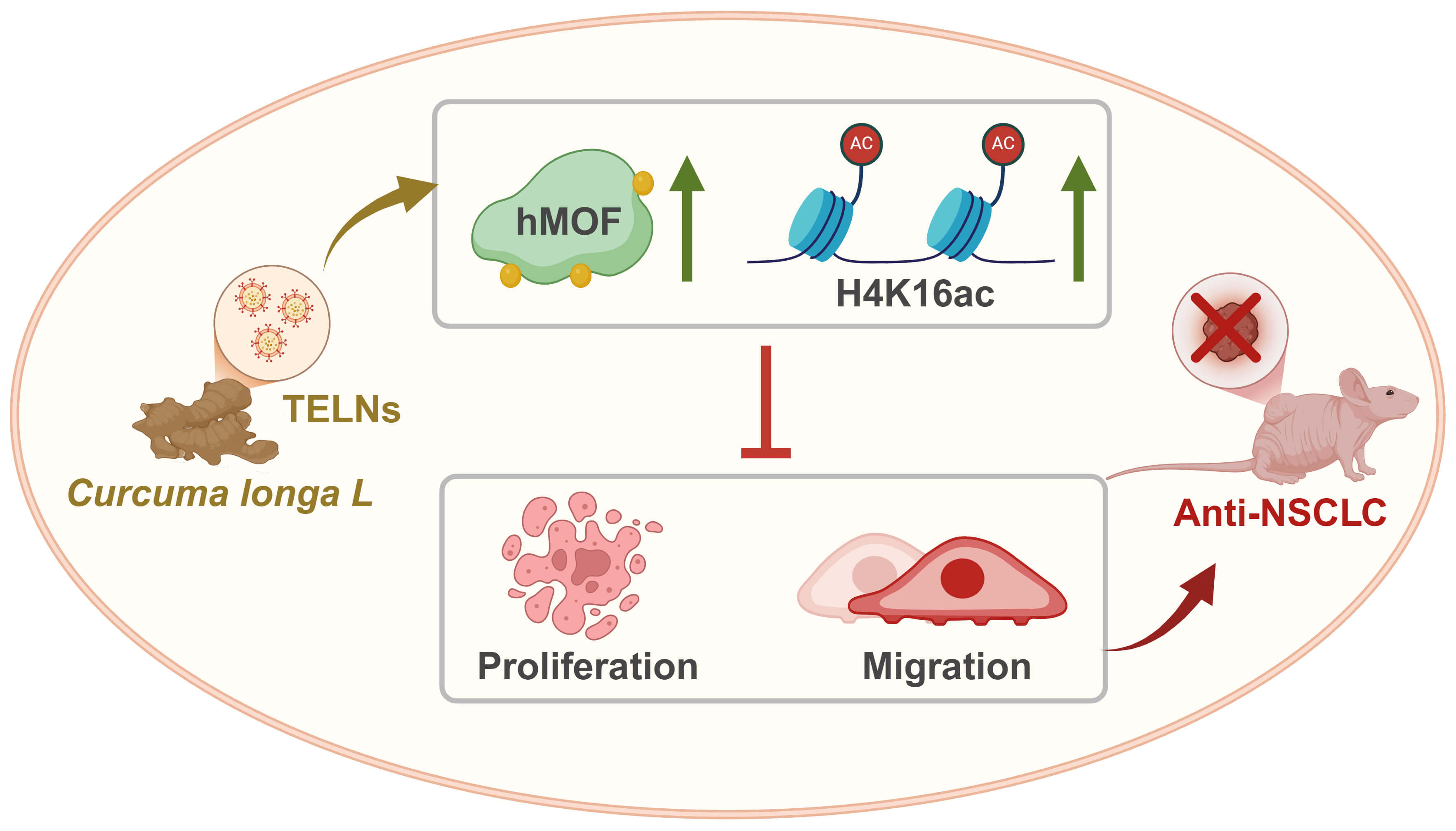 Fig. 6.
Fig. 6.
Mechanism underlying the anti-cancer effect of turmeric-derived exosome-like nanoparticles. TELNs upregulate hMOF and histone H4K16 acetylation, thereby promoting apoptosis and inhibiting NSCLC. The green upward arrow indicates increased expression.
Plant-derived exosome-like nanoparticles (PELNs) are nanoscale vesicles secreted by plants and exhibit bioactivity and biocompatibility [23]. PELNs carry lipids, proteins, miRNAs, and other components, exhibiting a similar structure to mammalian exosomes. They also offer advantages such as large-scale production capacity and natural therapeutic components [24]. Researchers have highlighted the anticancer effects of PELNs, including ginger-derived nanoparticles (GDENs) that induce apoptosis in cancer through TRAIL-mediated processes [25], and grapefruit-derived vesicles that deliver chemotherapeutics to tumor sites [26]. However, the epigenetic regulatory mechanisms of PELNs have not been studied. Our study bridges this gap by isolating turmeric-derived exosome-like nanoparticles (TELNs) and investigating the epigenetic mechanism underlying their anti-NSCLC effects. Originally defined as the interaction between genotype and phenotype, epigenetics primarily involves chromatin structural modifications that regulate gene expression [27]. Epigenetic research has provided insights into NSCLC progression and the mechanisms underlying anti-NSCLC drugs [28]. Histone acetylation, a pivotal epigenetic modification, adds an acetyl group to lysine residues on histones. This process is regulated by both histone acetyltransferases (HATs) and histone deacetylases (HDACs), which maintain acetylation homeostasis [29]. MOF is a MYST family member of HATs and specifically acetylates histone H4 at lysine 16 (H4K16ac) [30]. Notably, the expression of hMOF differs significantly between NSCLC and normal tissues [31]. By modulating the H4K16ac modification, hMOF promotes transcriptional activation and DNA repair, thereby regulating NSCLC progression [32]. TELNs significantly inhibit NSCLC progression in vitro and in vivo. Treatment with 100 µg/mL of TELNs induced apoptosis in A549 and H1299 cells and reduced tumor volume by 58.1%. These effects were mediated through upregulation of the histone acetyltransferase hMOF and its product H4K16ac. Molecular docking suggested potential interaction of curcumin to hMOF, while hMOF knockdown experiments confirmed the effects of TELNs. This dual action—upregulating both the expression and activity of hMOF—makes TELNs a unique epigenetic regulator.
Curcumin is a natural polyphenol isolated from the rhizome of Curcuma longa [33]. Accumulating evidence has demonstrated multiple mechanisms by which curcumin inhibits NSCLC progression [34], including downregulation of cyclin expression, induction of A549 cell cycle arrest, and suppression of cell proliferation [35]. Additionally, it promotes apoptosis in multiple pathways [36]. Although curcumin demonstrates anticancer effects against NSCLC, its poor bioavailability and low water solubility significantly hinder its translational application [37]. However, TELNs can overcome the bioavailability limitations of curcumin, with their phospholipid bilayer protecting cargoes from degradation. The particle size of TELNs in the present study remained stable during testing. Moreover, TELNs have dual functions as drug carriers and epigenetic modulators. As plant-derived nanoparticles, TELNs not only exhibit anticancer effects, but also serve as drug delivery systems capable of carrying various medications. Liposomes loaded with curcumin (CURLipo) have been developed (via ethanol injection method) for comparison with TELNs [38]. CUPLipo exhibits similar properties to TELNs with regard to uptake efficiency by A549 cells (Fig. 1G,H; Supplementary Fig. 13) and stability (Supplementary Figs. 1,14). However, TELNs demonstrated superior anti-NSCLC efficacy (Supplementary Fig. 15). Furthermore, hemolysis assays revealed that TELNs exhibited superior safety compared to CURLipo (Supplementary Fig. 16). This indicates that TELNs, as natural curcumin delivery carriers, offer advantages over liposomes.
The present study demonstrates for the first time that TELNs suppress NSCLC by epigenetic regulation. TELNs overcome the bioavailability limitations of curcumin, thus serving as a natural nanotherapeutic platform. However, their clinical translation requires further evaluation in more complex tumor models. Additionally, as TELNs are natural products derived from turmeric, standardized protocols must be established to overcome batch-to-batch variability in translational applications.
This study explored the anti-cancer mechanism of turmeric-derived exosome-like nanoparticles (TELNs). TELNs were found to regulate the histone acetyltransferase hMOF and histone H4 at lysine 16 (H4K16ac). They significantly induced apoptosis in non-small cell lung cancer (NSCLC), upregulated hMOF expression, and enhanced H4K16 acetylation. These findings establish for the first time that TELNs exert anti-tumor effects through epigenetic regulation of hMOF/H4K16ac, thus providing a basis for plant-derived nanoparticle-based therapies in cancer treatment.
The datasets used in this article are accessible from the corresponding author DL upon request.
Conceptualization: WS; Methodology: WS, DL; Software: WS; Validation: WS, DL, ZZ; Formal Analysis: WS; Investigation: WS; Data Curation: DL; Writing–Original Draft: WS; Writing–Review & Editing: DL, ZZ; Visualization: WS; Supervision: DL, ZZ; Project Administration: DL, ZZ; Funding Acquisition: DL, ZZ; Resources: ZZ. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The experimental procedures involving animals were conducted according to the Experimental Animal Welfare Ethics Committee of Changchun University of Chinese Medicine (Approval number: 2025936), Jilin Province, China.
We acknowledged the support of Biorender, Fig. 1A and Fig. 6 were created in https://biorender.com.
The authors declare this study was funded by a grant from National Natural Science of China (No. 81803680) and a grant from Natural Science of Jilin Province (No. YDZJ202401082ZYTS).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/FBL47950.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






