1 Department of Biochemistry and Molecular Biology, College of Pharmacy, Hebei University of Chinese Medicine, 050200 Shijiazhuang, Hebei, China
2 Hebei Key Laboratory of Chinese Medicine Research on Cardio-cerebrovascular Disease, 050091 Shijiazhuang, Hebei, China
3 Hebei Higher Education Institute Applied Technology Research Center on TCM Development and Industrialization, 050091 Shijiazhuang, Hebei, China
†These authors contributed equally.
Abstract
This study aims to examine the roles and mechanisms of action of bellidifolin (BEL) in alleviating doxorubicin-mediated cardiotoxicity using network pharmacology and experimental validation .
Mice with doxorubicin-induced cardiotoxicity were randomly assigned to control, model, BEL, and dexrazoxane (DEX) groups. Echocardiography, histological staining, network pharmacology, and molecular validation were employed to assess cardiac function and myocardial injury. Immunohistochemical staining, western blotting, and RT-qPCR were used to confirm predicted targets and fibrosis biomarkers.
In vivo experiments demonstrated that BEL significantly improved cardiac function, as indicated by enhanced Ejection Fraction (EF) and Fractional Shortening (FS) compared to the model group (p < 0.01). BEL also notably reduced myocardial injury markers, including creatine kinase MB isoenzyme (CK-MB) and lactate dehydrogenase (LDH) (p < 0.01), and alleviated doxorubicin-induced myocardial fibrosis. Network pharmacology identified 61 common target genes for BEL and cardiotoxicity. Proteinprotein interaction (PPI) network analysis highlighted 16 core genes, including transforming growth factor (TGF)-β1. Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) enrichment analyses revealed that BEL’s action pathways were primarily linked to the PI3K-AKT signaling pathway. Molecular docking and dynamic simulations showed a strong binding affinity between BEL and the core target TGF-β1. In vivo validation confirmed that BEL significantly downregulated the expression of TGF-β1, α-smooth muscle actin (SMA), collagen I (Col I), and collagen III (Col III) in myocardial tissue (p < 0.01 or p < 0.05), while activating the PI3K-AKT signaling pathway (p < 0.01 or p < 0.05).
BEL presents as a promising therapeutic candidate for cardiotoxicity, likely through its anti-fibrotic effects via the reduction of TGF-β1, α-SMA, Col I, and Col III expression, alongside regulation in the PI3K-AKT signaling pathway.
Keywords
- doxorubicin
- cardiotoxicity
- endomyocardial fibrosis
- network pharmacology
- molecular dynamics simulation
Doxorubicin (DOX) is a widely used antibiotic for treating various malignant
tumors [1]. While DOX is highly effective in its antitumor and antibacterial
properties, increasing evidence highlights its potential cardiotoxicity [2].
DOX-induced cardiotoxicity (DIC) commonly manifests as myocardial cell damage,
fibrosis, and cardiac insufficiency, which may progress to heart failure,
significantly impacting patient quality of life and prognosis [3]. The
pathological mechanisms underlying DIC involve a cascade of molecular events,
including excessive oxidative and nitrosative stress, mitochondrial dysfunction,
apoptosis, impaired autophagy, and the dysregulation of key signaling pathways
such as AMPK, Nrf2, TGF-
In Inner Mongolia, Gentianella acuta (G. acuta) is commonly used as a medicinal tea for preventing cardiovascular diseases [7]. As documented in the Mongolian Herbal Medicine Records, G. acuta is traditionally used to treat conditions such as angina pectoris, jaundice, and cholecystitis, and it is widely employed in clinical practice to alleviate myocardial fibrosis and cardiovascular injury [8, 9, 10, 11, 12]. Bellidifolin (BEL), an active xanthone molecule derived from G. acuta, has the molecular formula C14H10O6 and a molecular weight of 274.22 g/mol [13]. Previous studies have demonstrated that BEL can mitigate myocardial fibrosis through multiple mechanisms [14, 15].
Network pharmacology, based on systems biology principles, analyzes biological
networks to identify signal nodes for multi-target drug design [16]. Molecular
docking and molecular dynamics simulations can verify the binding modes and
interaction strengths between drugs and targets at the molecular level. Combined
with in vivo experiments, these methods offer an efficient approach to
elucidating the pharmacodynamic basis of natural products [17]. While prior
studies confirmed BEL’s role in improving myocardial fibrosis, its specific
molecular mechanism remains unclear. Based on bioinformatics, BEL’s therapeutic
effect may be associated with the fibrosis-related target TGF-
BEL (Lot: AFDE3105, Purity: 98%, Chengdu Alfa Biotechnology Co., Ltd., China)
was used in this study. In the preliminary experiment, two gradient doses of BEL
(25 mg/(kg
Sixty specific-pathogen-free (SPF) male C57BL/6 mice (8 weeks old, body weight 18–22 g) were obtained from Beijing Sibeifu Biotechnology Co., Ltd. (License No.: SCXK (Beijing) 2019-0010). All mice were housed individually in cages with free access to food and water, in a well-ventilated environment with a 12-hour light/dark cycle, a temperature of 20–25 °C, and relative humidity maintained at 40%–70%. The experiment was approved by the Animal Welfare Committee of Hebei University of Chinese Medicine (Approval No.: DWLL202212032).
The mice were randomly divided into four groups (15 mice per group) using a
random number table: Control, Model, BEL, and DEX groups. The Model, BEL, and DEX
groups were intraperitoneally injected with 5 mg/kg DOX to establish a myocardial
injury model, while the Control group received the same dose of normal saline
via intraperitoneal injection once a week for 4 weeks. Echocardiography
and myocardial tissue pathological staining were used to assess the success of
modeling. In addition to DOX injection, the BEL group received 50
mg/(kg
Throughout the experiment, the mice’s mental state, activity levels, water intake, and food intake were observed and recorded daily. Body weight measurements were taken weekly to adjust the gavage volume accordingly.
ECG detection: After drug administration, mice were anesthetized via intraperitoneal injection of pentobarbital sodium (For sodium pentobarbital: It was prepared into a 3% solution using sterile normal saline, and administered via intermittent and slow intraperitoneal injection at a standard dose of 50 mg/kg body weight). Afterward, they were placed supine on the experimental platform, and their extremities were subcutaneously linked to the BL-420S biological function experimental system. The standard lead II ECG was monitored for 60–120 seconds.
Echocardiography detection: The evaporator of the anesthesia machine was connected to an air pump, with the output gas flow rate set at 300–500 mL/min. Mice were placed in an anesthesia induction chamber with 3–4% isoflurane until fully anesthetized (approximately 2–3 minutes). Once the mice were fully anesthetized, the concentration was reduced to a maintenance level of 1–1.5%. The mice were removed from the induction chamber, their heads and noses fixed in an anesthetic mask, and positioned supine on the detection table with limbs attached to electrode pads. After depilating the chest, the ultrasonic probe was placed closely against the mice’s chest wall. Cardiac ultrasound was performed using a small animal ultrasound imaging system, where Left Ventricular Posterior Wall thickness in systole (LVPWs), Left Ventricular Posterior Wall thickness in diastole (LVPWd), Left Ventricular Volume in systole (LVVols), Left Ventricular Volume in diastole (LVVold), Left Ventricular Internal Diameter in systole (LVIDs), Left Ventricular Internal Diameter in diastole (LVIDd), Fractional Shortening (FS), and Ejection Fraction (EF) were measured. Turn off the evaporator after the procedure is completed, and the mice were kept breathing pure oxygen for approximately 5–10 minutes to facilitate their rapid recovery.
After completing the ECG and echocardiography, the mice were fasted for 12 hours
with free access to water. Blood was collected via eyeball enucleation
into clean test tubes, and serum was separated by centrifugation. The supernatant
was stored at –80 °C for subsequent biomarker assays. The operator
pressed the mouse’s head and neck with the thumb and index finger, while grasping
the mouse’s tail or hind limbs with the other hand. Then, the operator quickly
pulled the hindquarters backward and upward with force to dislocate the cervical
vertebrae. The chest cavity was then rapidly opened, and a 1.0 mm
After 4 weeks of administration, the mice’s body weight was recorded, and the abdomen was incised to expose the heart. The abdominal aorta was ligated, and residual blood in the heart was flushed with pre-cooled saline. The tissue was blotted dry with filter paper before weighing to record heart weight.
Heart tissues were fixed with 4% paraformaldehyde (G1101, Wuhan Servicebio Technology Co., Ltd., Wuhan, China), processed through routine paraffin embedding, sectioned, and stained with HE and Masson. Pathological alterations in myocardial tissue were examined microscopically.
The heart tissue of the mice was stained with Sirius red to assess fibrotic changes in the myocardial tissue under a microscope. ImageJ 1.51j software (National Institutes of Health, Bethesda, MD, USA) was used for quantitative analysis of the collagen area, allowing evaluation of the degree of fibrosis.
Following serum collection, the levels of LDH (lactate dehydrogenase), CK-MB (creatine kinase MB isoenzyme), ALT (alanine aminotransferase), UREA (urea), and CREA (creatinine) were measured using an automatic biochemical analyzer (Chemray 800, Rayto Life and Analytical Sciences Co., Ltd., Shenzhen, Guangdong, China).
Target identification for BEL involved retrieving data from multiple sources, including Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, https://www.tcmsp-e.com/load_intro.php?id=43), Traditional Chinese Medicine Integrated Database (TCMID, https://bidd.group/TCMID/), and other chemical databases (Chemical professional database, http://www.organchem.csdb.cn/scdb/default.asp). The 3D structure and Canonical Simplified Molecular Input Line Entry System (SMILES) of BEL were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and used in the Swiss Target Prediction database (http://www.swisstargetprediction.ch/) for further exploration. After eliminating duplicate targets, species were restricted to “Homo sapiens” using the UniProt database (https://www.uniprot.org/), and the target names were standardized into gene names to build the BEL target database.
For disease-related target identification, keywords like “cardiotoxicity”,
“anthracycline-induced cardiotoxicity”, and “doxorubicin-induced
cardiotoxicity” were used to retrieve targets from the GeneCards database
(https://www.genecards.org/), with a relevance score cutoff of
The intersection of BEL targets and cardiotoxicity-related targets was
extracted, representing the potential action targets of BEL in treating
cardiotoxicity. These targets were imported into the STRING database
(https://string-db.org/) for PPI analysis, selecting the “multiple proteins”
mode with the species limited to “Homo sapiens” and a confidence score of
Further functional analyses, including GO and KEGG enrichment analyses, were
performed to explore the main biological functions and signaling pathways
involved in the treatment of DIC by BEL. The targets were input into the DAVID
2021 platform (https://davidbioinformatics.nih.gov/) for biological information annotation,
with “OFFICIAL_GENE_SYMBOL” as the identifier, “gene list” as the list
type, and “Homo sapiens” as the species. The threshold was set at p
As a ligand, the 2D structure of compound BEL was retrieved from the PubChem database and stored in “sdf” format. For selecting PDB files of core gene-related proteins, the RCSB PDB database (https://www.rcsb.org/) was utilized, with the following selection criteria: experimental method—X-ray crystallography (the primary method for determining protein tertiary structures), target species—Homo sapiens (human-derived proteins were chosen to accurately represent in vivo structural characteristics and avoid binding mode discrepancies due to species differences), resolution—0.5–2.5 Å (proteins with a resolution within this range provide detailed information on amino acid side chains and hydrogen bond networks, ensuring docking accuracy), presence of ligands in the structure (facilitating direct identification of the docking region and minimizing invalid structures without target pockets), pH range—7.35–7.50 (representing the physiological pH of the human body, simulating the native conformation of proteins in vivo). The proteins were screened according to these criteria and stored in “PDB” format. The online molecular docking software CB-Dock2 (https://cadd.labshare.cn/cb-dock2/php/blinddock.php) was then used to evaluate the binding activity between the compound and the target gene. In the molecular docking results, a Vina score lower than –5 kJ/mol indicates good docking between the protein and the ligand, with a lower Vina score reflecting a more stable ligand-receptor binding.
Based on the molecular docking results, YASARA 10.3.16 (YASARA Bioinformatics GmbH, Vienna, Austria) was used for molecular dynamics simulations. The protonation state of the residues was corrected by adding hydrogens and optimizing bond lengths and angles. Periodic boundary conditions (PBC) were applied, counterions were added to neutralize the system, and the physiological pH was set to 7.4. The appropriate force field was selected for the simulation. Through energy minimization and equilibrium simulations, steric clashes between atoms were resolved, and the energy of the initial structure was minimized. The predefined MD macro file was executed, and equilibrium simulations in the canonical ensemble (NVT) and isothermal-isobaric ensemble (NPT) were conducted for 100 ps each, at a temperature of 298 K and a pressure of 1 bar. Subsequently, a 100 ns molecular dynamics simulation of the complex system was performed, with conformations saved every 10 ps. Using YASARA’s built-in tools, parameters such as RMSD and RMSF were calculated to assess system stability. The binding free energy between the ligand and receptor was analyzed, and key interaction residues were identified through binding energy decomposition.
Paraffin sections prepared as described in section 1.5 were subjected to
sequential dewaxing, hydration, and antigen retrieval. After a 3% hydrogen
peroxide treatment to block endogenous peroxidase activity, the sections were
sealed. The primary antibodies targeting TGF-
The TRIzol method was utilized for the extraction of total RNA from mouse myocardial tissue, and cDNA was synthesized through reverse transcription. Real-time quantitative PCR was performed using SYBR qPCR Master Mix (G3325, Wuhan Servicebio Technology Co., Ltd., Wuhan, China), with primer sequences presented in Table 1. The PCR program included pre-denaturation at 95 °C for 30 seconds, followed by 40 cycles of denaturation at 95 °C (10 seconds), annealing at a temperature ranging from 55 to 65 °C (10 seconds), and extension at 72 °C (30 seconds). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) acted as the internal control gene, and relative gene expression was determined using the 2-ΔΔCt method.
| Primer | Sequence (5′-3′) | Length/bp | Annealing Temperature | |
| TGF- |
Forward | ACTGGAGTTGTACGGCAGTG | 121 | 60 °C |
| Reverse | GGCTGATCCCGTTGATTTCC | |||
| Forward | TCAGGGAGTAATGGTTGGAATG | 348 | 60 °C | |
| Reverse | CCAGAGTCCAGCACAATACCAG | |||
| Col I | Forward | GAGAGGTGAACAAGGTCCCG | 153 | 60 °C |
| Reverse | AAACCTCTCTCGCCTCTTGC | |||
| Col III | Forward | GTGGCAATGTAAAGAAGTCTCTGAAG | 191 | 60 °C |
| Reverse | GGGTGCGATATCTATGATGGGTAG | |||
| M-GAPDH | Forward | CCTCGTCCCGTAGACAAAATG | 133 | 60 °C |
| Reverse | TGAGGTCAATGAAGGGGTCGT | |||
TGF-
Radio Immunoprecipitation Assay (RIPA) lysis buffer (G2002, Wuhan Servicebio
Technology Co., Ltd., Wuhan, China) was utilized for the lysis of myocardial
tissue, and centrifugation was performed to harvest the supernatant. A BCA assay
kit (G2026, Wuhan Servicebio Technology Co., Ltd., Wuhan, China) was used to
determine the protein concentration. After separating protein samples by
SDS-PAGE, they were transferred onto PVDF membranes. The membranes were blocked
in 5% non-fat milk, incubated with primary antibodies (TGF-
Statistical analysis was conducted using SPSS 27.0 (IBM Corp., Chicago, IL, USA), while GraphPad Prism 8.0.2 (GraphPad Software, San Diego, CA, USA) was used to
generate statistical graphs. Data normality was tested, and comparisons of
normally distributed data were performed using one-way analysis of variance
(ANOVA). For homogeneous variances, post-hoc testing was done using the least
significant difference (LSD) method. For heterogeneous variances, the Dunnett’s
T3 test was applied. Data are presented as mean
During the experiment, compared to the blank control group, mice in the model
group exhibited reduced water and food intake, yellowing fur, and lethargy, among
other symptoms. These conditions improved to varying degrees in the treatment
groups when compared to the model group. Echocardiography results revealed that
the cardiac contour and internal structure in the blank control group were
normal, with the ventricular wall showing a regular contraction/relaxation
rhythm. In contrast, the model group displayed left heart enlargement, abnormal
ventricular wall motion, and decreased EF and FS, along with increased values of
LVIDs, LVPWs, LVVols, LVIDd, LVPWd, and LVVold (p
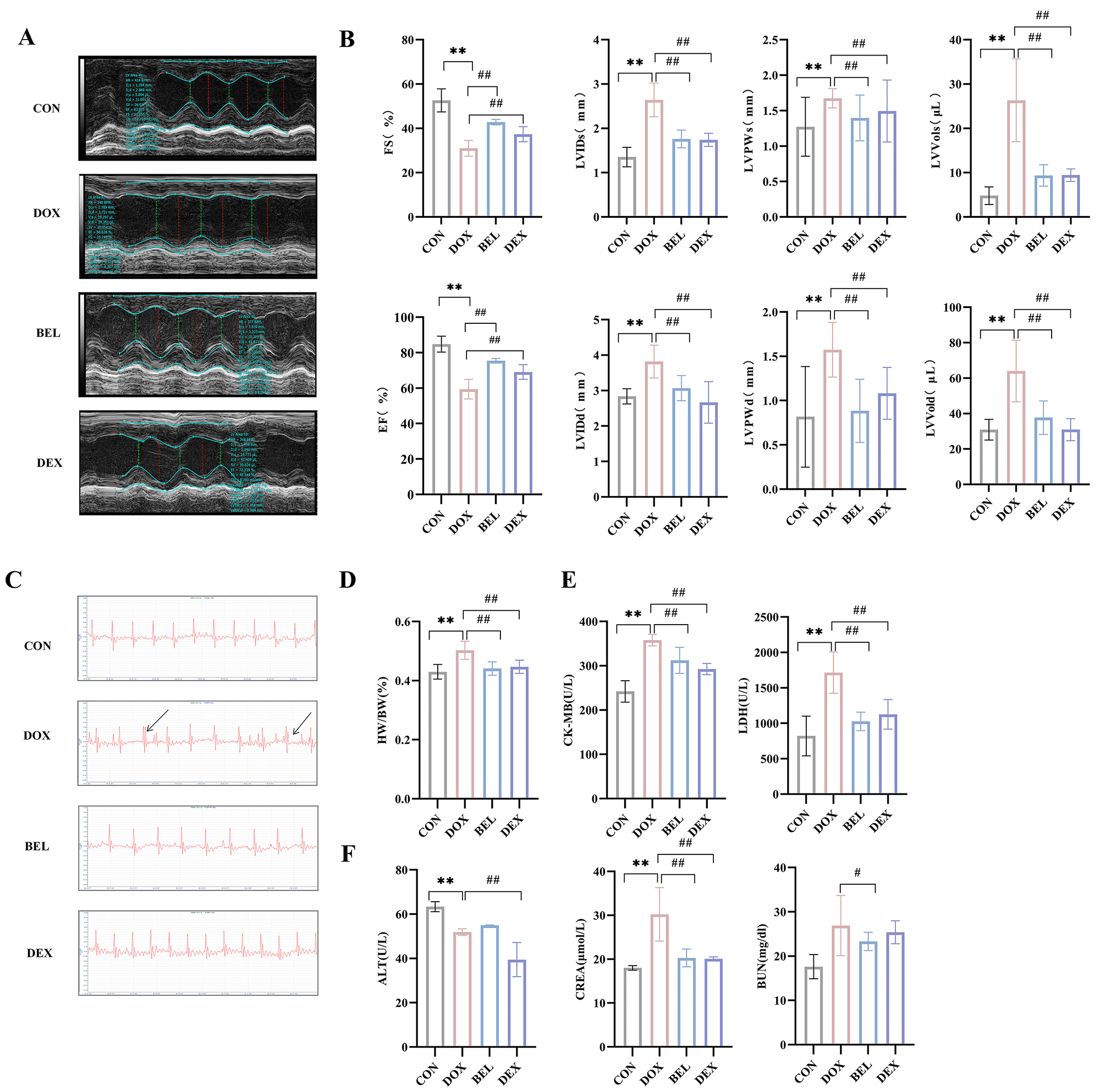 Fig. 1.
Fig. 1.
Effects of bellidifolin (BEL) on cardiac function, myocardial
injury, and hepatorenal damage in mice. (A) Echocardiography of mice. (B)
Cardiac function indices of mice. (C) Electrocardiogram of mice (The area pointed
to by the arrow corresponds to the electrocardiographic features of arrhythmia).
(D) Heart weight ratio of mice. (E) Serum indices of myocardial injury. (F) Serum
indices of hepatorenal damage. (Data presented as mean
Myocardial HE staining results (Fig. 2A,B) revealed that, compared to the control group, mice in the model group exhibited varying degrees of eosinophilic changes, marked swelling of myocardial cells, significant rupture of myocardial striations, fibroblast proliferation, considerable thickening of myocardial cells compared to normal cells, deep-stained cytoplasm, pyknosis of myocardial cell nuclei, progressive nuclear disappearance, and visible apoptosis and necrosis, accompanied by inflammatory cell infiltration in necrotic areas. In contrast, each treatment group showed significant improvements compared to the model group.
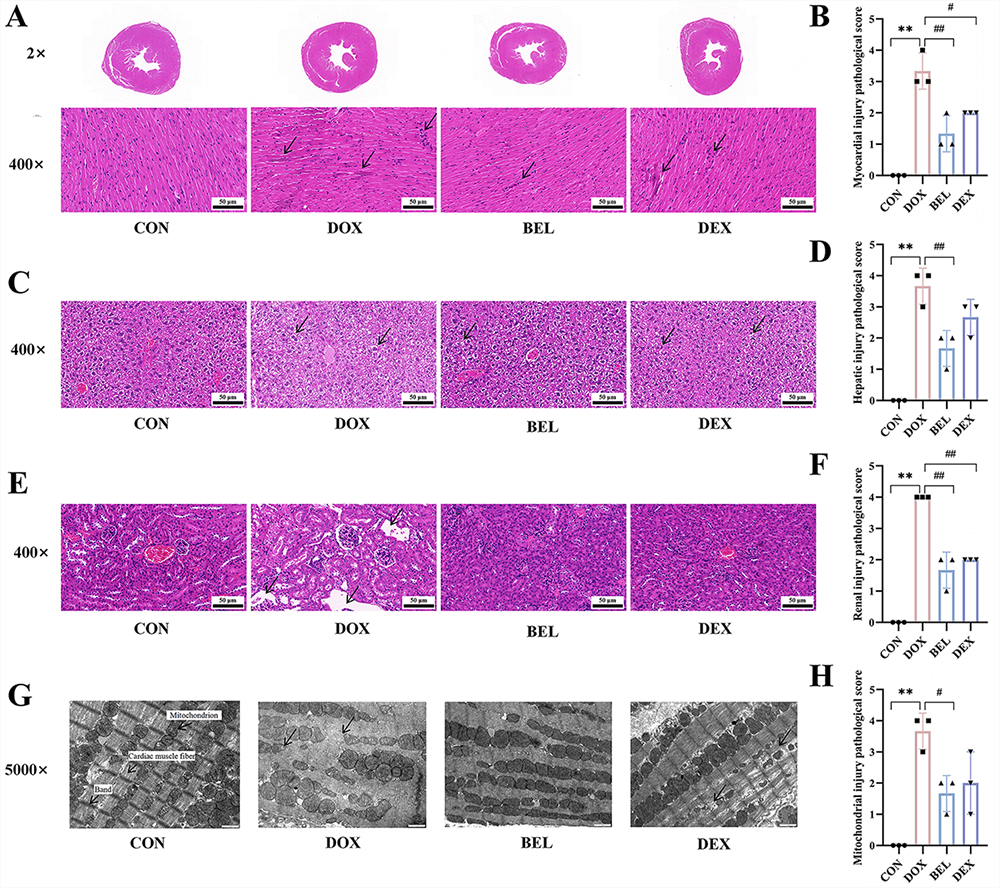 Fig. 2.
Fig. 2.
BEL improves pathological changes in the myocardium, liver, and
kidneys of cardiotoxic mice. (A,B) HE staining results of myocardial tissue
(400
Liver HE staining results (Fig. 2C,D) indicated that liver tissue from mice in the model group exhibited extensive inflammatory cell infiltration around the portal vein, dilation of hepatic lobular venous lumens, and liver damage. The inflammatory infiltration in liver tissue was markedly reduced in all treatment groups compared to the model group.
Kidney HE staining results (Fig. 2E,F) showed that compared to the control group, renal tissue in the model group exhibited obvious tubular epithelial edema, swollen cells, vacuolated cytoplasm, visible connective tissue hyperplasia, and lymphocyte infiltration. Pathological damage in the renal tissue was significantly alleviated in all treatment groups compared to the model group.
To further assess myocardial injury, the microstructure of the mouse myocardium was observed by transmission electron microscopy (TEM). As shown in Fig. 2G,H, myocardial mitochondria in the blank control group were neatly arranged, numerous, with intact mitochondrial membrane structures, clear cristae, and no rupture or obvious hypertrophy. In contrast, myocardial mitochondria in the model group were disorganized, condensed, scattered, with dissolution and disappearance of surrounding myofilaments, mitochondrial cristae rupture, hypertrophy, hyperplasia, and severe vacuolization. The administration groups showed significant improvements in mitochondrial structure compared to the model group.
Myocardial Masson staining results (Fig. 3A,B) revealed that myocardial cells appeared pink, while collagen fibers were stained blue-purple. Compared to the blank control group, myocardium in the other groups exhibited varying degrees of injury. In the model group, pathological sections showed disorganized myocardial cell arrangement, significant accumulation of blue-purple fibrous collagen, and destruction of myocardial structure. In contrast, the administration groups exhibited more organized myocardial cells with reduced collagen fiber accumulation compared to the model group. The Masson staining results indicated a strong correlation between cardiotoxicity and the development of myocardial fibrosis. To further assess collagen expression in myocardial tissues, Sirius red staining and Sirius red polarized light staining were performed. The results were consistent with the Masson staining, showing that collagen expression in myocardial tissues correlated with the severity of fibrosis, as shown in Fig. 3C–F.
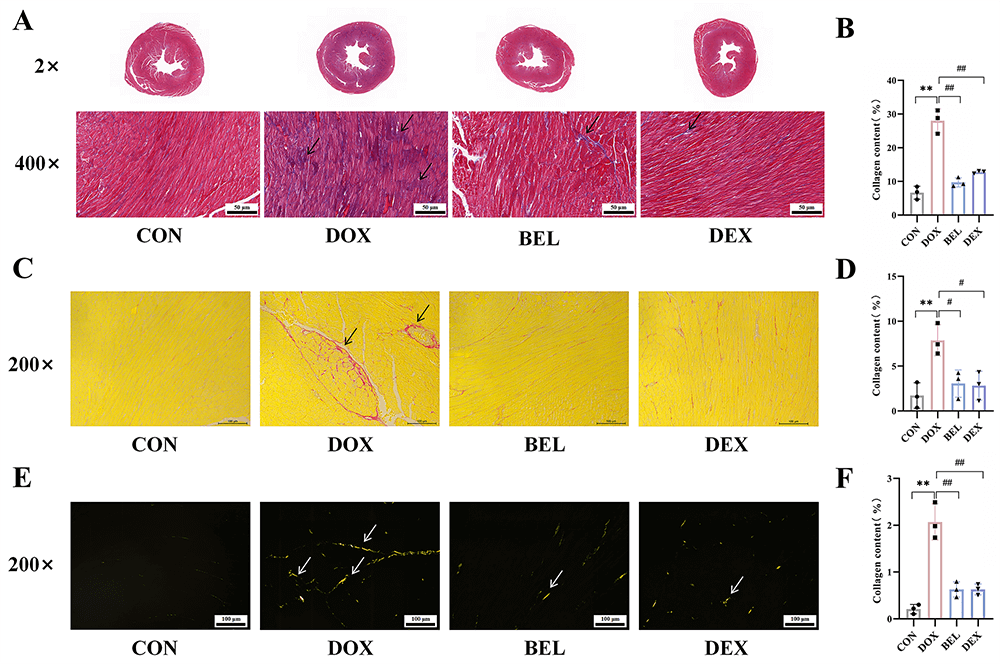 Fig. 3.
Fig. 3.
BEL ameliorates myocardial fibrosis in cardiotoxic mice. (A,B)
Masson staining results of myocardial tissue (400
After merging and removing duplicates from the collected targets, the targets were imported into the UniProt database for standardization. A total of 180 targets corresponding to BEL and 1478 targets related to DIC were retrieved. The intersection of these targets resulted in 61 potential targets for BEL in the treatment of DIC, as shown in Fig. 4A.
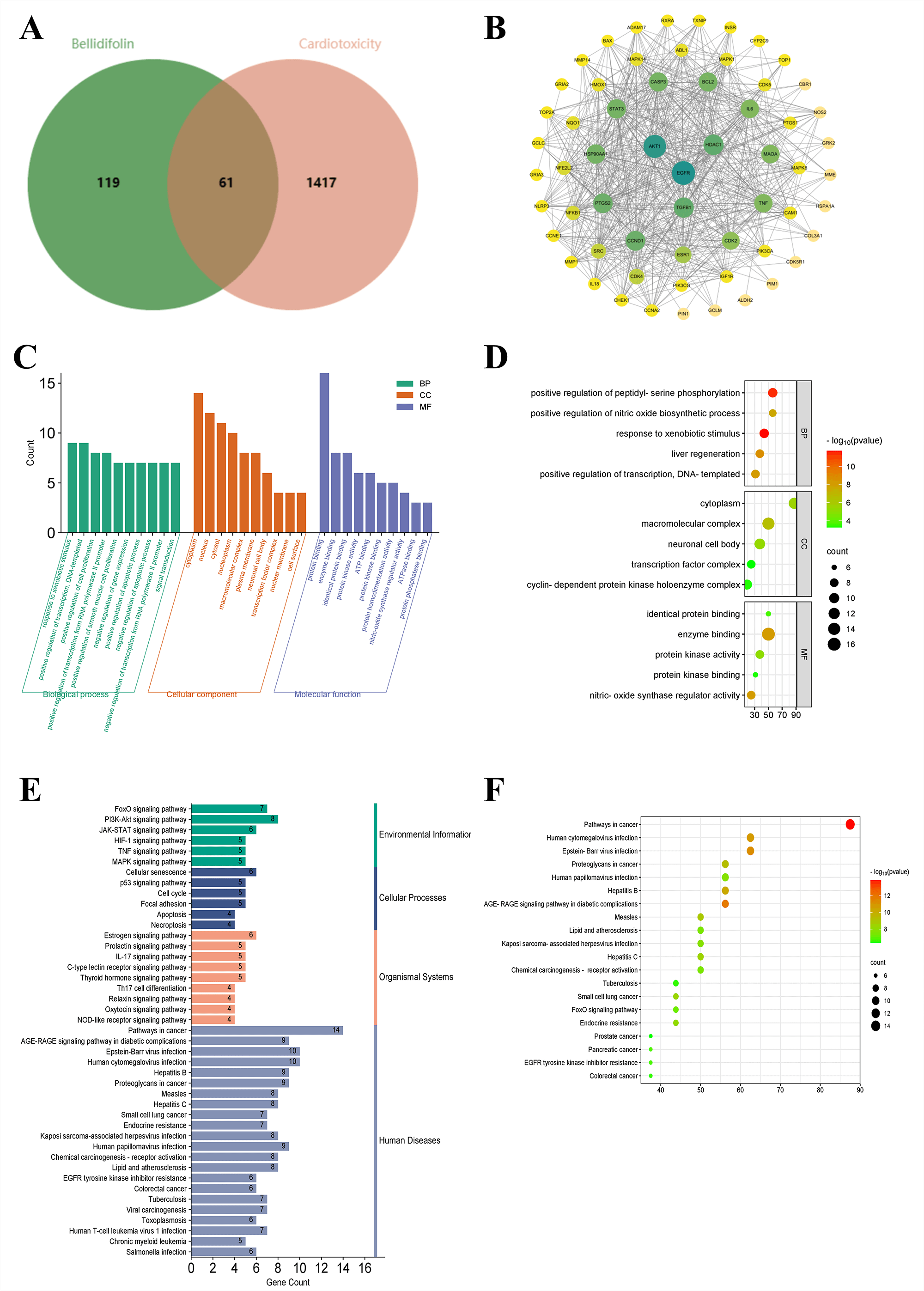 Fig. 4.
Fig. 4.
Network pharmacology results of BEL intervention in doxorubicin-induced cardiotoxicity. (A) Venn diagram of intersections between BEL targets and cardiotoxicity targets. (B) PPI network diagram. (C) Bar chart of GO analysis. (D) Bubble chart of GO secondary analysis. (E) Bar chart of KEGG. (F) Bubble chart of KEGG.
As depicted in Fig. 4B, the PPI network of BEL interfering with DOX-induced cardiotoxicity contained 66 nodes and 828 edges, where nodes represented proteins and edges denoted their mutual relationships. Targets with Degree, Closeness, and Betweenness values higher than the average were identified as key targets for BEL intervention in DIC. The specific details of all targets are presented in Table 2.
| Uniport ID | Protein target | Hub genes | Degree | Closeness | Betweenness |
| P31749 | RAC-alpha serine/threonine-protein kinase | AKT1 | 48 | 0.8428571 | 207.92517 |
| P00533 | Epidermal growth factor receptor | EGFR | 45 | 0.7972973 | 219.73232 |
| P10415 | Apoptosis regulator Bcl-2 | BCL2 | 44 | 0.7972973 | 131.657 |
| P42574 | Caspase-3 | CASP3 | 43 | 0.7763158 | 132.9298 |
| P40763 | Signal transducer and activator of transcription 3 | STAT3 | 43 | 0.7866667 | 135.45213 |
| P05231 | Interleukin-6 | IL6 | 42 | 0.7763158 | 122.438065 |
| Q9UNG2 | Tumor necrosis factor ligand superfamily member | TNF | 42 | 0.7763158 | 121.26485 |
| P35354 | Prostaglandin G/H synthase 2 | PTGS2 | 40 | 0.75641024 | 143.20937 |
| P24385 | G1/S-specific cyclin-D1 | CCND1 | 38 | 0.72839504 | 145.6201 |
| P03372 | Estrogen receptor | ESR1 | 37 | 0.72839504 | 88.119705 |
| P07900 | Heat shock protein HSP 90-alpha | HSP90AA1 | 37 | 0.72839504 | 136.71877 |
| P12931 | Proto-oncogene tyrosine-protein kinase Src | SRC | 36 | 0.7108434 | 55.051365 |
| P01137 | Transforming growth factor beta-1 proprotein | TGF- |
35 | 0.70238096 | 153.86162 |
| Q13547 | Histone deacetylase 1 | HDAC1 | 31 | 0.6781609 | 159.10611 |
| P24941 | Cyclin-dependent kinase 2 | CDK2B1 | 28 | 0.64835167 | 108.45579 |
| P11802 | Cyclin-dependent kinase 4 | CDK4 | 27 | 0.6413044 | 64.967896 |
Notes: Degree reflects the number of connections a node has, Betweenness
measures the bridging role of a node within the shortest paths, Closeness
represents the reciprocal of the average distance from the node to all other
nodes. These three indicators are commonly used to assess the topological
properties of each node. Higher values indicate more important nodes.
Consequently, key targets were selected based on the criterion that all three
indicators were higher than the average. AKT1, RAC-alpha serine/threonine-protein
kinase; EGFR, Epidermal Growth Factor Receptor; BCL2, B-cell lymphoma 2; CASP3,
Caspase-3; STAT3, Signal Transducer and Activator of Transcription 3; IL6,
Interleukin 6; TNF, Tumor Necrosis Factor; PTGS2, Prostaglandin G/H synthase 2;
CCND1, cyclin-D1; ESR1, Estrogen Receptor 1; HSP90AA1, Heat Shock Protein 90
Alpha Family Class A Member 1; SRC, Proto-oncogene tyrosine-protein kinase Src;
TGF-
GO functional annotation analysis (p
Pathway enrichment analysis (p
Molecular docking of BEL with the 16 core targets was performed to verify its
binding capacity to target proteins at varying degrees. Based on the
aforementioned criteria, the PDB files of AKT1 (7MYX), EGFR (8A27), BCL2 (8H7B),
CASP3 (7JI7), STAT3 (6TLC), IL6 (5GW9), TNF (6U66), PTGS2 (5F19), CCND1 (6P8E),
ESR1 (7UJO), HSP90AA1 (3O0I), SRC (3EAC), TGF-
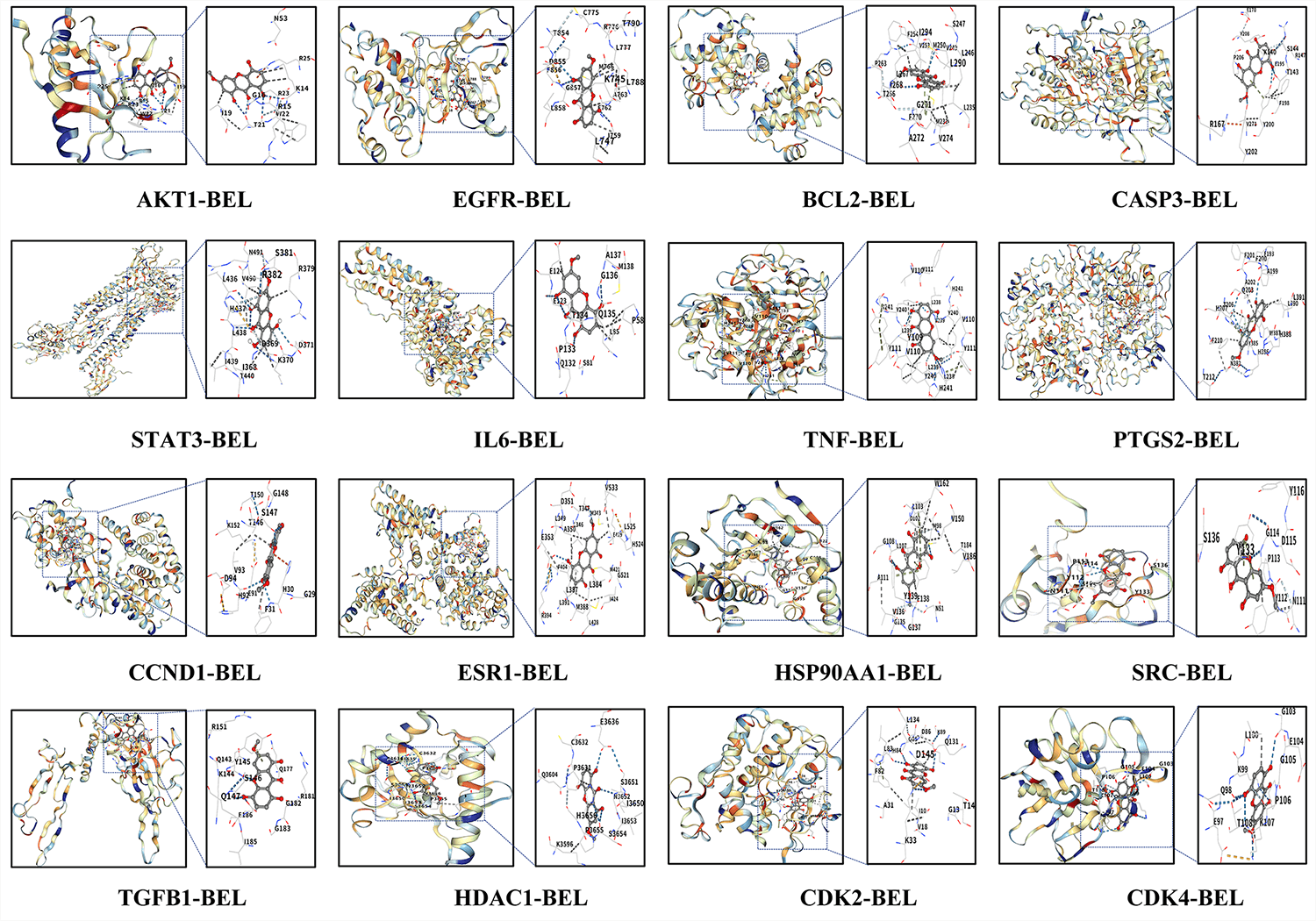 Fig. 5.
Fig. 5.
Molecular docking results of BEL and key target proteins. Note:
Hydrogen bonds (blue dotted line), weak hydrogen bonds (white dotted line),
hydrophobic interactions (gray dotted line), ionic interactions (yellow dotted
line), cation-
| Vina scores | Cavity size | Center | Size | |||||
| x | y | z | x | y | z | |||
| AKT1 | –6.5 | 84 | 14 | 7 | 16 | 20 | 20 | 20 |
| EGFR | –8.1 | 3691 | 18 | –10 | –13 | 30 | 28 | 31 |
| BCL2 | –7.0 | 3257 | 2 | –1 | –13 | 20 | 29 | 32 |
| CASP3 | –7.1 | 2136 | 3 | 3 | –24 | 20 | 32 | 20 |
| STAT3 | –7.6 | 607 | 14 | 49 | 66 | 20 | 20 | 20 |
| IL6 | –8.0 | 669 | –1 | 151 | –5 | 20 | 20 | 20 |
| TNF | –7.8 | 833 | –41 | 105 | –19 | 20 | 20 | 27 |
| PTGS2 | –9.3 | 1233 | 28 | 28 | 59 | 20 | 27 | 33 |
| CCND1 | –6.9 | 413 | 11 | 30 | 72 | 20 | 20 | 20 |
| ESR1 | –7.7 | 2644 | 12 | –21 | –24 | 20 | 32 | 20 |
| HSP90AA1 | –9.2 | 2115 | –6 | –10 | –25 | 26 | 20 | 20 |
| SRC | –6.1 | 54 | –11 | 8 | –5 | 20 | 20 | 20 |
| TGF- |
–6.8 | 198 | 97 | 54 | 52 | 20 | 20 | 20 |
| HDAC1 | –6.5 | 85 | 23 | 9 | 0 | 20 | 20 | 20 |
| CDK2 | –7.3 | 444 | 0 | –7 | –25 | 20 | 20 | 20 |
| CDK4 | –5.9 | 360 | 12 | 38 | 9 | 20 | 20 | 20 |
Notes: The Vina score is a scoring function used to evaluate the binding strength
in molecular docking. It calculates the interaction and binding free energies
between molecules to assign a score, with lower scores indicating stronger
binding. A Vina score
Based on the molecular docking results, the TGF-
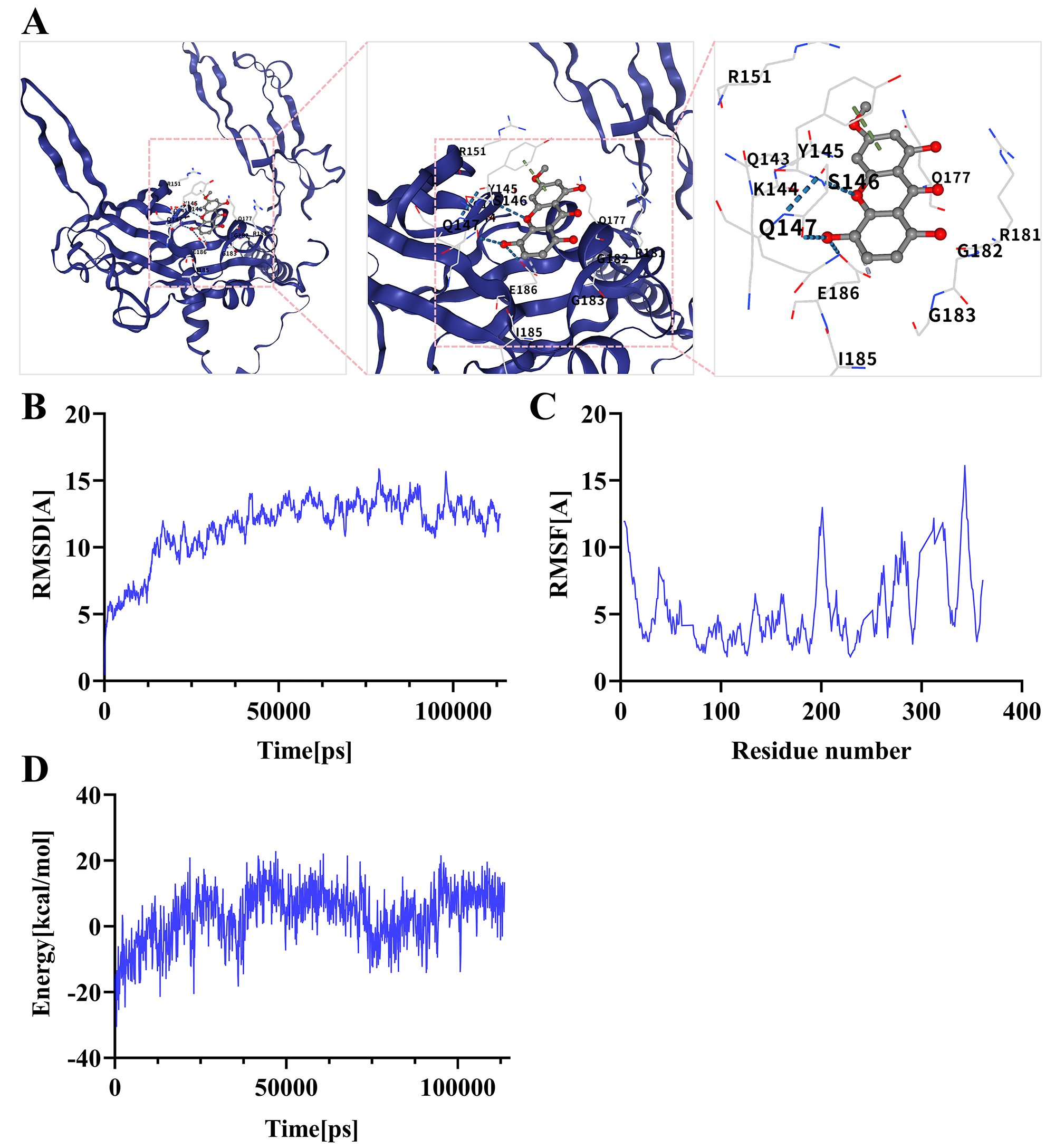 Fig. 6.
Fig. 6.
Molecular dynamics simulation results of BEL and key target
proteins. (A) Binding mode of the TGF-
The expression of TGF-
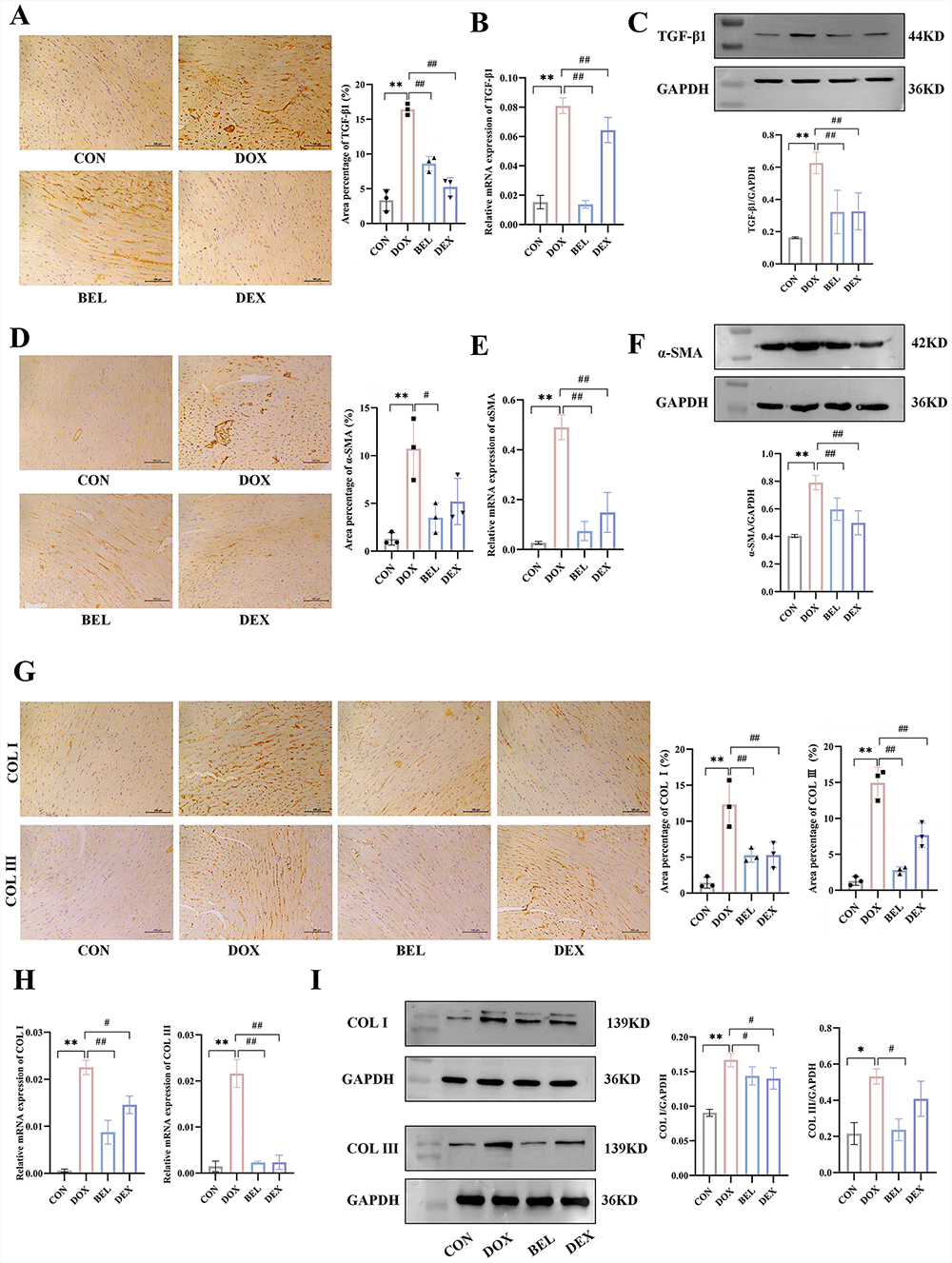 Fig. 7.
Fig. 7.
Expression of TGF-
Following the same verification methods as for TGF-
Given that TGF-
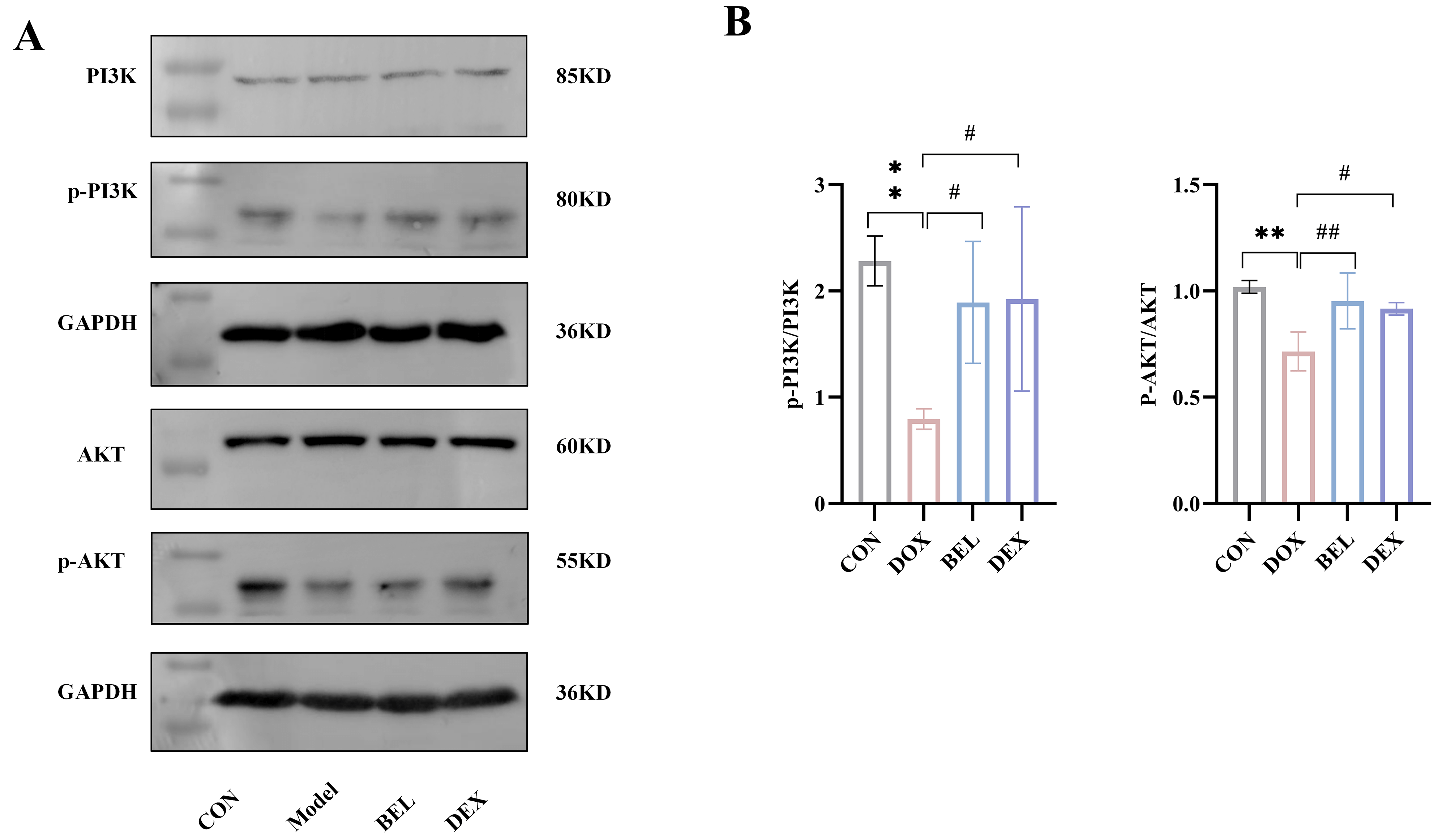 Fig. 8.
Fig. 8.
Expression of the PI3K-AKT signaling pathway in myocardial
tissue of mice. (A) WB detection strip chart. (B) Grayscale analysis of WB
detection. (Antibody dilution: PI3K = 1:1000. p-PI3K = 1:1000. AKT = 1:1000.
p-AKT = 1:1000) (CON vs. DOX: *p
Extensive research indicates that DIC significantly raises the mortality risk of cancer patients, thus limiting its widespread clinical use [20, 21]. As the global incidence of cancer continues to rise, DIC warrants greater attention. Early diagnosis and treatment of DIC are essential for improving clinical outcomes. However, current therapeutic options for DIC have limitations in addressing this complex condition [22]. This study aims to identify a potential therapeutic candidate for DIC and clarify its underlying mechanisms.
Network pharmacology analysis was employed in this study to explore the targets
and mechanisms of BEL in the treatment of DIC. The results revealed 61 drug
targets overlapping with DIC disease genes. Among these, 16 core targets were
identified through PPI network analysis, including AKT, TGF-
Chronic DIC is characterized by myocardial fibrosis, which significantly impairs
cancer patient prognosis [24]. DIC leads to the upregulation of fibrosis markers,
such as phospho-Smad3, collagen I, fibronectin, and
TGF-
Numerous studies have confirmed that the PI3K/Akt signaling pathway plays a key role in regulating the development, progression, and pathological formation of cardiac fibrosis by modulating cell survival, apoptosis, proliferation, myocardial contractility, and the transcription of associated genes, involving molecules such as mTOR, GSK-3, FoxO1/3, and NOS [30, 31, 32]. This study demonstrates that both BEL and DEX reduce collagen accumulation in myocardial tissue by regulating the PI3K-AKT signaling pathway. Additionally, TEM results revealed damage to the myocardial mitochondria in the model group, which may be attributed to oxidative stress and mitochondrial dysfunction—key pathogenic mechanisms of DIC. Both BEL and DEX were found to improve mitochondrial structure, providing another potential pathway for their cardioprotective effects.
Based on these findings, it can be inferred that BEL treats DIC by inhibiting myocardial fibrosis. The strength of this study lies in its identification of the potential mechanisms of DIC through network pharmacology analysis and the provision of a promising therapeutic drug. Future studies will focus on determining the optimal human dosage of BEL, evaluating its potential interactions with chemotherapeutic drugs, and conducting safety assessments to provide a solid foundation for its clinical application.
In this study, the primary mechanism by which BEL treats DIC appears to be its
anti-myocardial fibrosis effect, with TGF-
This study has several limitations: First, the in vivo drug metabolism
of BEL remains unclear. Second, additional cellular experiments are needed to
verify the biological functional modules involved. Third, while a 4-week acute
animal model was used, DIC may occur years after treatment in clinical settings.
Therefore, future research will focus on establishing a chronic DIC model to
investigate the long-term effects of BEL. Fourth, only male mice were used, and
future studies will examine potential differences in the ameliorative effects of
BEL in female mice. Fifth, the timing of BEL administration in this study was
aligned with the modeling period. however, further investigation is needed to
explore the effects of pre-modeling versus post-modeling administration. Sixth,
the effect of BEL on tumor cells and its potential interference with the
antitumor efficacy of DOX remain unclear. Seventh, pathway inhibition experiments
and small interfering RNA (siRNA) experiments targeting the TGF-
The authors confirm that the data supporting the findings of this study are available within the article. The data associated with this paper are available upon request to the corresponding author.
Conceptualization: XMZ, YL, AYL; Data curation: XMZ, AYL; Formal analysis: CD, AYL; Funding acquisition: YL, AYL; Investigation: XMZ; Methodology: XMZ, FG, YL; Project administration: YL, AYL; Resources: JLW; Software: ZYZ, FG; Supervision: JLW; Validation: XMZ; Visualization: XMZ, YLL; Writing—original draft: XMZ, ZYZ, AYL; Writing—review and editing: YL, AYL. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The animal study protocol was approved by the Animal Welfare Committee of Hebei University of Chinese Medicine (Approval No.: DWLL202212032). The study adhered to the guidelines set by the committee.
The authors would like to express our gratitude to all those who helped me during the writing of this manuscript. And thanks to all the peer reviewers for their opinions and suggestions.
This work was supported by the Hebei Natural Science Foundation [grant number: H2025423167]; the National Natural Science Foundation Youth Fund of China [grant number: 81500317].
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.








