1 Department of Chemoradiotherapy, Ningbo No 2 Hospital, 315000 Ningbo, Zhejiang, China
2 Department of Breast Surgery, Ningbo No 2 Hospital, 315000 Ningbo, Zhejiang, China
Abstract
Breast cancer stem cells (BCSCs) are instrumental in treatment resistance, recurrence, and metastasis. The development of breast cancer and radiation sensitivity is intimately pertinent to long non-coding RNA (lncRNA). This work is formulated to investigate how the lncRNA MIR155HG affects the stemness and radioresistance of BCSCs.
Effects of MIR155HG knockdown on BCSCs were gauged in MCF-7 and MDA-MB-231 cell lines. MIR155HG expression was manipulated in cells, followed by an assessment of stemness, DNA damage repair, apoptosis, cell cycle, and the Wnt signaling pathway under radiation conditions. The interaction between nuclear factor kappa B (NF-κB) subunit RelA and MIR155HG was examined using a dual-luciferase reporter assay. To examine the binding interaction between RelA and MIR155HG promoter, chromatin immunoprecipitation was performed.
Breast cancer-derived stem cells exhibited a high level of MIR155HG. Knockdown of MIR155HG reduced stemness, enhanced radiosensitivity, induced apoptosis, and arrested cells in the G1 phase. Mechanistically, MIR155HG knockdown repressed Wnt/β-catenin signaling and mediated apoptosis-related protein expressions. NF-κB subunit RelA transcriptionally activated MIR155HG, thereby contributing to radioresistance in BCSCs.
NF-κB regulates MIR155HG transcriptionally to activate the Wnt pathway, thus enhancing stemness and radioresistance in BCSCs. Targeting MIR155HG may enhance the susceptibility of cancer stem cells to radiation-induced cell death, potentially improving therapeutic outcomes. These findings underscore MIR155HG as a promising therapeutic target for breast cancer.
Keywords
- breast cancer
- cancer stem cells
- nuclear factor-kappa B
- MIR155 host gene
Breast cancer is a highly prevalent malignant tumor arising from breast tissue in women globally. According to statistics from the World Health Organization, breast cancer ranks first among malignant tumors in women, with new cases of roughly 2.3 million and deaths of around 685,000 annually [1]. Breast cancer has a complicated etiology that involves genetics, hormone levels, lifestyle, and environmental exposures [2]. Various treatment modalities exist, such as radiation, endocrine therapy, chemotherapy, surgery, and targeted therapy [3], of which radiotherapy is instrumental in controlling local tumor and reducing recurrence rates. However, radiotherapy has been confirmed to stimulate tumor stem cells to secrete cytokines or enhance DNA damage repair mechanisms, leading to increased self-renewal capacity and radioresistance [4]. Furthermore, tumor stem cells extracted from glioblastoma xenografts and live tissues are associated with radioresistance and tumor cell regeneration [5]. Stem cells from breast cancer display significantly higher invasion ability, proliferation capacity, and radioresistance compared to non-stem cells [6]. The presence of breast cancer stem cells (BCSCs) provides a possible explanation for the current treatment resistance, recurrence, and metastasis, indicating that inhibiting these stem cells-related signaling pathways could be promising in reversing chemoradiotherapy resistance.
Long non-coding RNA MIR155 host gene (lncRNA MIR155HG) is located on chromosome 21 of the human genome [7]. MIR155HG is transcribed into precursor miRNA, which is subsequently processed into mature miR-155. Numerous biological systems, including the immune system, inflammatory reactions, cell proliferation, and differentiation, depend on MIR155HG [8, 9]. Morman[10] showed that an early and persistent event in T cell activation is the transcriptional activation of MIR155HG. MIR155HG expression is abundant in various cancers, such as natural killer/T-cell lymphoma [11], cervical cancer [12] and melanoma [13]. In Hodgkin’s lymphoma, aberrant B-cell receptors may generate MIR155HG [14]. Under specific conditions, MIR155HG can be processed to produce miR-155, accelerating the progression of glioma [15]. According to earlier research, the MIR155HG/miR-155 axis promotes the epithelial-mesenchymal transition, which is thought to have carcinogenic properties in gliomas [16]. By sponging miR-185, overexpressed MIR155HG in glioblastoma can increase ANXA2 expression, and ANXA2 subsequently binds to the phosphorylated STAT3 and the MIR155HG promoter for stimulating MIR155HG expression and promoting tumor growth and development [17]. Additionally, it has been proved that MIR155HG activates the Wnt signaling pathway to enhance cell radioresistance [18, 19]. However, its functional role in regulating BCSCs and radiosensitivity has not yet been documented.
We discovered that BCSCs have high expression of lncRNA MIR155HG
through experimental investigation and bioinformatics analysis. Our findings
suggested that nuclear factor kappa B (NF-
AnWei-sci (Shanghai, China) provided human breast cancer cell lines MCF-7 (AW-CELLS-H0213) and MDA-MB-231 (AW-CELLS-H0217). Short tandem repeat (STR) analysis was leveraged to validate the purity and ensure cell line stability. All cells underwent mycoplasma testing and were mycoplasma free. Cells were cultured in Dulbecco’s Modified Eagle Medium (C0891-100 mL) supplemented with 10% fetal bovine serum (C0251), 100 µg/mL streptomycin (C0222), and 100 U/mL penicillin. All reagents were provided by Beyotime (Shanghai, China).
MCF-7 and MDA-MB-231 cells were initially sorted for CD44+ cells using magnetic cell sorting (MACS; Miltenyi Biotech, Auburn, CA, USA) before executing the side population (SP) sorting in order to separate CD44+/CD24– cell populations [20]. In MACS, cells were revived in 0.1% phosphate-buffered saline (PBS; C0221A)-bovine serum albumin (BSA, ST2254) from Beyotime (Shanghai, China). They were then labeled with phycoerythrin (PE)-conjugated CD24 antibody (555428, BD-PharMingen, San Diego, CA, USA) and fluorescein isothiocyanate (FITC)-conjugated CD44 antibody (555478, BD-PharMingen, San Diego, CA, USA)-phosphate-buffered saline (PBS; C0221A, Beyotime, Shanghai, China). Employing the “DOUBLE POSITIVE SORT” program on the AutoMACS system (Miltenyi Biotech, Auburn, CA, USA), cell suspension was filtered and sorted. The positive and negative fractions for SP were evaluated and sorted with a FACSCalibur flow cytometer (Becton Dickenson, Mountain View, CA, USA).
Following the directions of the maker, TRIzol reagent (R0016, Beyotime, Shanghai, China) was utilized to extract RNA. With a reverse transcription kit (D7168S, Beyotime, Shanghai, China), 500 mg RNA was reversely transcribed into cDNA (10 µL). qPCR was conducted to quantitate lncRNAs MIR155HG and RelA on the Step One Plus Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) using SYBR Premix Ex Taq™ II (Q331-02/03, Vazyme, Nanjing, China) in three independently repeated experiments. Expression analyses were achieved by 2-ΔΔCt method. Sangon Biotech (Shanghai, China) created primers, with the sequences below: lncRNA MIR155HG forward (F), 5′-ACGGTTGTGCGAGCAGAGAATCTA-3′ and reverse (R), 5′-CTCATCTAAGCCTCACAACAACCT-3′; RelA F, 5′-CTTCCTCAGCCATGGTACCTCT-3′ and R, 5′-CAAGTCTTCATCAGCATCAAACTG-3′; Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) F, 5′-CCATGGGGAAGGTGAAGGTC-3′ and R, 5′-AGTGATGGCATGGACTGTGG-3′.
MCF-7 and MDA-MB-231 stem cells were divided into the following groups: Blank, short hairpin RNA negative control (shNC), short hairpin RNA targeting MIR155HG (shMIR155HG), vector, RelA, shNC+vector, shMIR155HG+vector, shNC+RelA, and shMIR155HG+RelA. GenePharma (Shanghai, China) supplied shMIR155HG and shNC. The pcDNA3.1-RelA (RelA) and control empty vector plasmids (vector) were acquired from GeneChem (Shanghai, China). Cell transfection with 1 µg shRNA or 2 µg plasmid was completed utilizing Lipofectamine 2000 (11668019, Invitrogen, Carlsbad, CA, USA) (48 h, 37 ℃) for later trials. The sequences were as follows: shMIR155HG-1, 5′-GCAGATAACTTGTCTGCATTTCAAGAGAATGCAGACAAGTTATCTGCTTTTTT-3′; shMIR155HG-2, 5′-GCATTCACATGGAACAAATTTCAAGAGAATTTGTTCCATGTGAATGCTTTTTT-3′; shNC, 5′-GCACCCAGTCCGCCCTGAGCAAATTCAAGAGATTTGCTCAGGGCGGACTGGGTGCTTTTT-3′.
Equal densities of cells were planted onto 60 mm cell culture dishes, and
cultured for 4–10 h (37 ℃) to promote adhesion. Following that, cells received
X-ray radiation (0, 2, 4, and 6 Gy) utilizing an RS2000 X-ray biological research
irradiator (Rad Source Technologies, Suwanee, GA, USA) [21]. To facilitate colony
development, cell incubation was conducted for 10–14 days. The colonies
underwent fixation (methanol, 322415, Sigma-Aldrich, St. Louis, MO, USA) and
staining (0.1% crystal violet; C0121, Beyotime, Shanghai, China). At least fifty
cells in a colony were counted. Surviving fraction = the number
of colonies/(cells inoculated
Using lysis buffer (KGC4901; Keygene, Jiangsu, China), total protein from the
cells was determined via BCA protein assay kit (P0010; Beyotime, Shanghai,
China). For sodium dodecyl sulfate polyacrylamide gel electropheresis (SDS-PAGE,
KGC4901; Keygene, Jiangsu, China), 40 µg of protein per lane was loaded and
separated on 10% gels, followed by being transferred onto polyvinylidene
fluoride (PVDF) membranes (KGC4806; Keygene, Jiangsu, China). Membranes
experienced 1-h blockage at room temperature (RT) using 5% non-fat milk in
Tris-Buffered Saline with Tween-20 (TBST, ST825; Beyotime, Shanghai, China), and
cultivation with primary antibodies (4 ℃, overnight) obtainable
from Abcam (Cambridge, UK) except RelA (65 kDa; 1:1000, AM06378SU-N, Origene,
Rockville, MD, USA), incorporating
Cells were seeded at equal densities onto glass slides and incubated (37 ℃,
4–10 h) to allow for attachment. After that, cells underwent exposure to 2 Gy
X-rays for 24 h. Immunofluorescence staining was carried out employing a DNA
damage detection kit (
With the goal to create spheres, single-cell suspensions (30,000 cells/well)
were cultivated in serum-free Minimum Essential Medium (MEM, 11058021) with 20
ng/mL bFGF, 20 ng/mL EGF, and B27 (all from Thermo Fisher Scientific, Waltham,
MA, USA) in 100 mm ultra-low attachment plates (4615, Corning Inc., Corning, NY, USA). The
development of spheres was captured by an inverted microscope (200
Following 24-h exposure to 2 Gy X-rays, cells experienced color development by Annexin V-FITC and PI (FITC Annexin V Apoptosis Detection Kit, C1062S, Beyotime, Shanghai, China), followed by cell cycle analysis exploiting TaliTM Cell Cycle Kit (A10798, Thermo Fisher Scientific, Waltham, MA, USA). Measurements of apoptosis and cell cycle were completed utilizing a flow cytometer, follows by result evaluation with CellQuest software (version 5.1, BD Biosciences, San Jose, CA, USA).
Transcription factor-binding motifs were predicted using the JASPAR database (https://jaspar.elixir.no/). MIR155HG-wild type (WT) and mutant (MUT) sequences (wild and mutated sequences of the binding site between MIR155HG and RELA) were cloned into a luciferase reporter vector (16147, Promega, Madison, WI, USA). With Lipofectamine 2000, MIR155HG-WT or MIR155HG-MUT and RELA were co-transfected into MDA-MB-231 and MCF-7 stem cells. Overnight cell lysis was followed by luciferase activity assessment employing a luminometer (Bio-Rad, Hercules, CA, USA) and a Dual-Luciferase Reporter Assay Kit (E1910, Promega, Madison, WI, USA).
ChIP Assay Kit (17-295, Millipore, Temecula, CA, USA) was purchased in advance. Briefly, cells were lysed on ice for 10 min. To extract 200–1000 bp DNA fragments from cell lysates, cells underwent seven times of 5-second pulses on ice with a Sonicator 3000 (Misonix, Farmingdale, NY, USA). An agarose bead mixture, ChIP buffer, and a protease inhibitor cocktail were used to pre-clear cell lysates for 1 h on ice. The lysates were treated (12 h, 4 ℃) with either normal rabbit IgG (2729, 1:50) or RelA antibody (8242, 1:100) from Cell Signaling Technology (Danvers, MA, USA). The supernatant containing 60 µL Protein A Agarose/Salmon Sperm DNA was added and spun (1 h, 4 °C). Wash buffers containing low salt (150 mM), high salt (500 mM), LiCl, and TE buffer were applied to wash the beads. After elution of the complexes using elution buffer, 5 M NaCl was used to reverse the cross-links, and the mixture experienced 4-h incubation (65 ℃). Following RNase A treatment, samples were collected for qPCR to gauge the levels of the MIR155HG promoter. The primer sequences used were F 5′-GGTCTCCAGCTGATTCGGTC-3′ and R 5′-CCAGGAGCGTCTCCTTGGTT-3′.
Each experiment was independently repeated three times. Statistical studies were
carried out with the aid of GraphPad Prism 8.0 (GraphPad Software, Boston, MA, USA). Data were displayed as mean
Through flow cytometry, we first separated stem cells from breast cancer cell
lines, and detected CD44+/CD24– cells
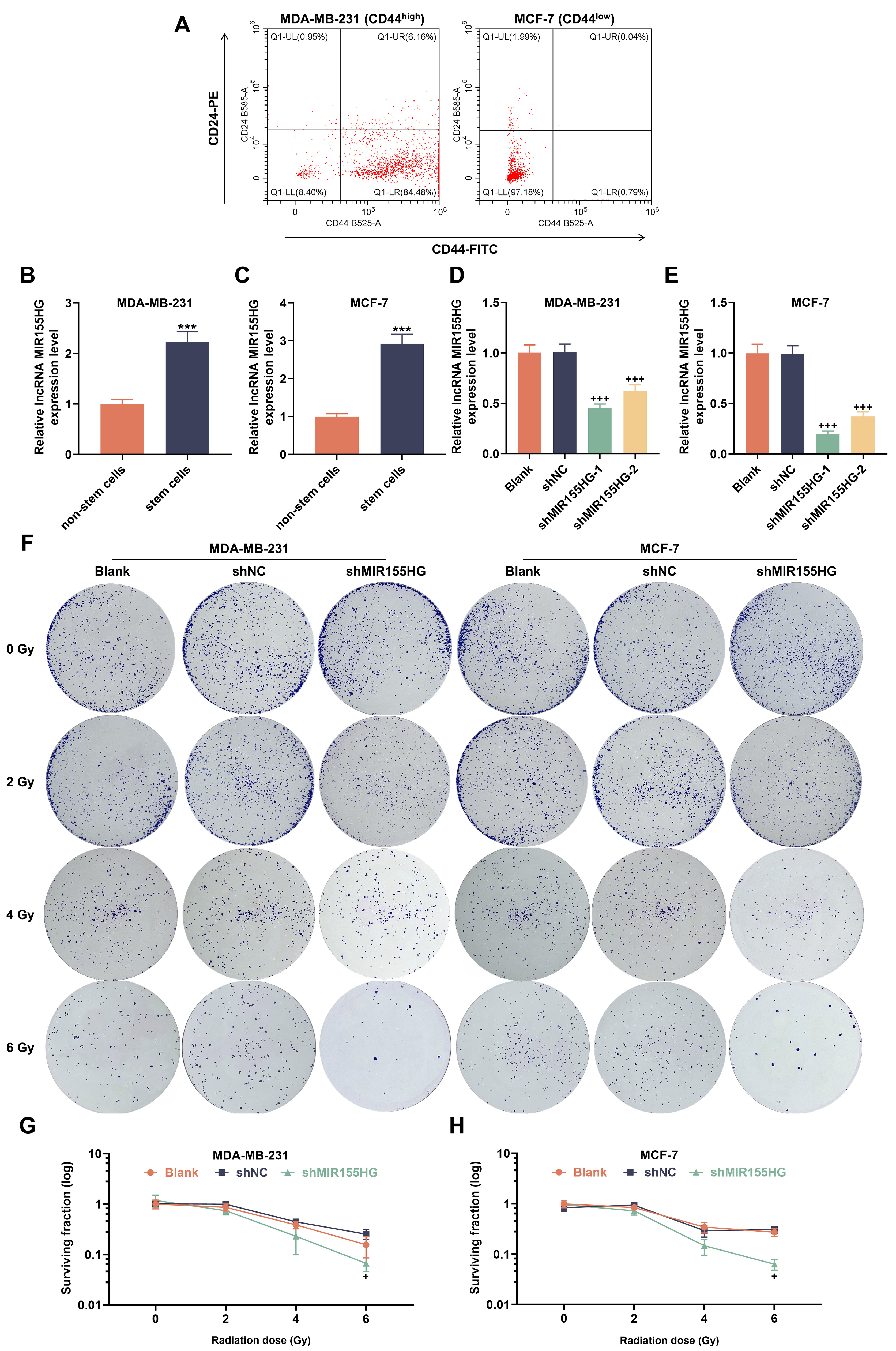 Fig. 1.
Fig. 1.
Long non-coding RNA MIR155HG is highly expressed in
Breast cancer stem cells (BCSCs), and knockdown of MIR155HG reduces
cellular radioresistance. (A) FITC-CD44- and PE-CD24-stained MDA-MB-231 and
MCF-7 cells (flow cytometry). (B,C) MIR155HG messenger RNA (mRNA) levels
in MDA-MB-231 and MCF-7 stem and non-stem cells (reverse
transcription quantitative polymerase chain reaction (RT-qPCR)). (D,E)
MIR155HG mRNA levels in short hairpin RNA targeting MIR155HG
(sh-MIR155HG)-transfected MCF-7 and MDA-MB-231 stem cells (RT-qPCR). (F)
Representative images of MIR155HG-silencing MDA-MB-231 and MCF-7 stem
cells under radiation (colony formation assays). (G,H) Survival curves of
MIR155HG-silencing MDA-MB-231 and MCF-7 stem cells under radiation.
n = 3. ***p
One day after 2 Gy X-ray irradiation, we detected the expression of
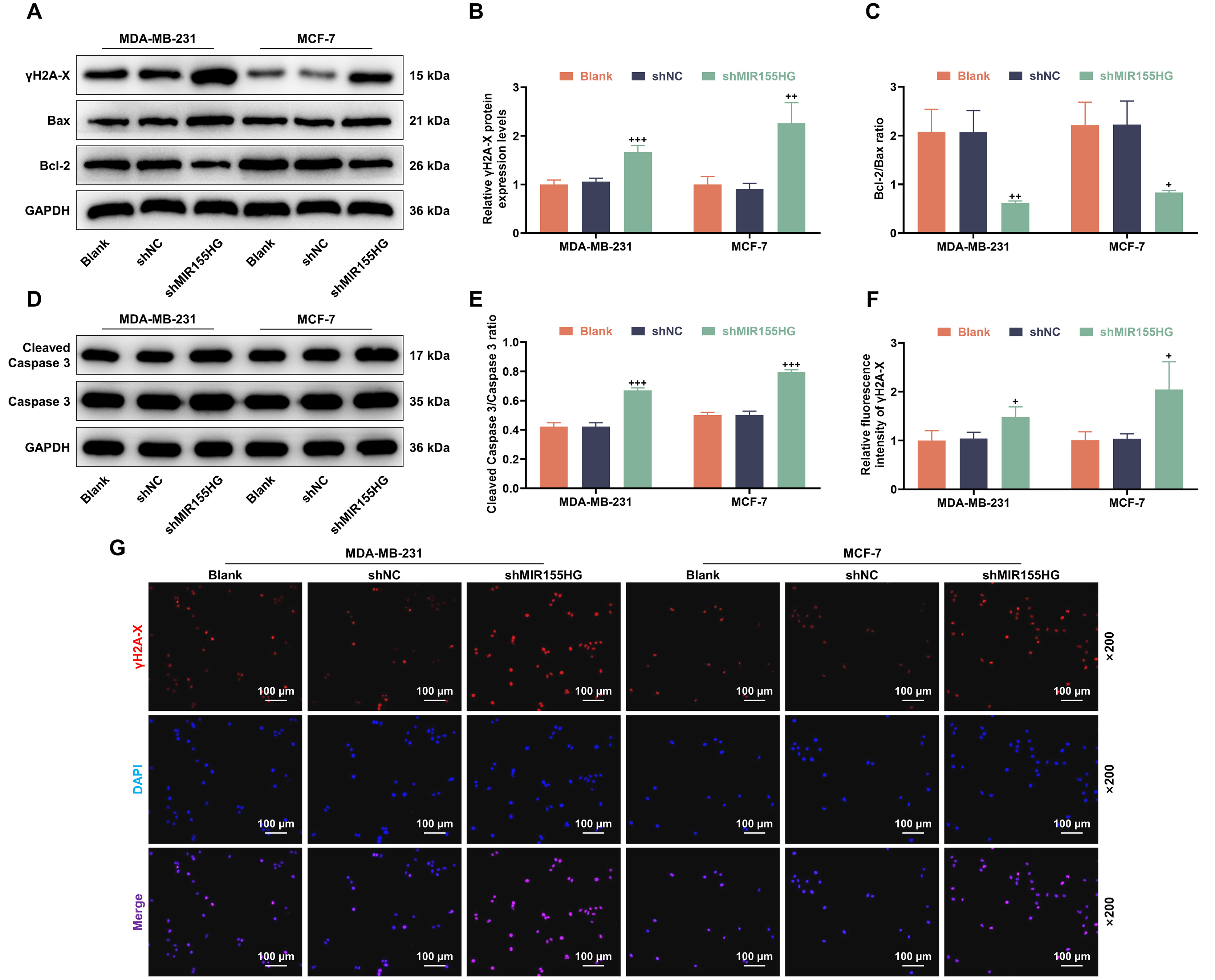 Fig. 2.
Fig. 2.
Knockdown of MIR155HG reduces DNA damage repair and
promotes apoptosis protein expression in BCSCs. (A–E) Phosphorylated histone
variant H2A.X (
Next, following 2 Gy X-ray irradiation, we assessed the development of tumor
spheres in MDA-MB-231 and MCF-7 stem cells. The tumor sphere-forming ability was
weaker in sh-MIR155HG group than shNC group (p
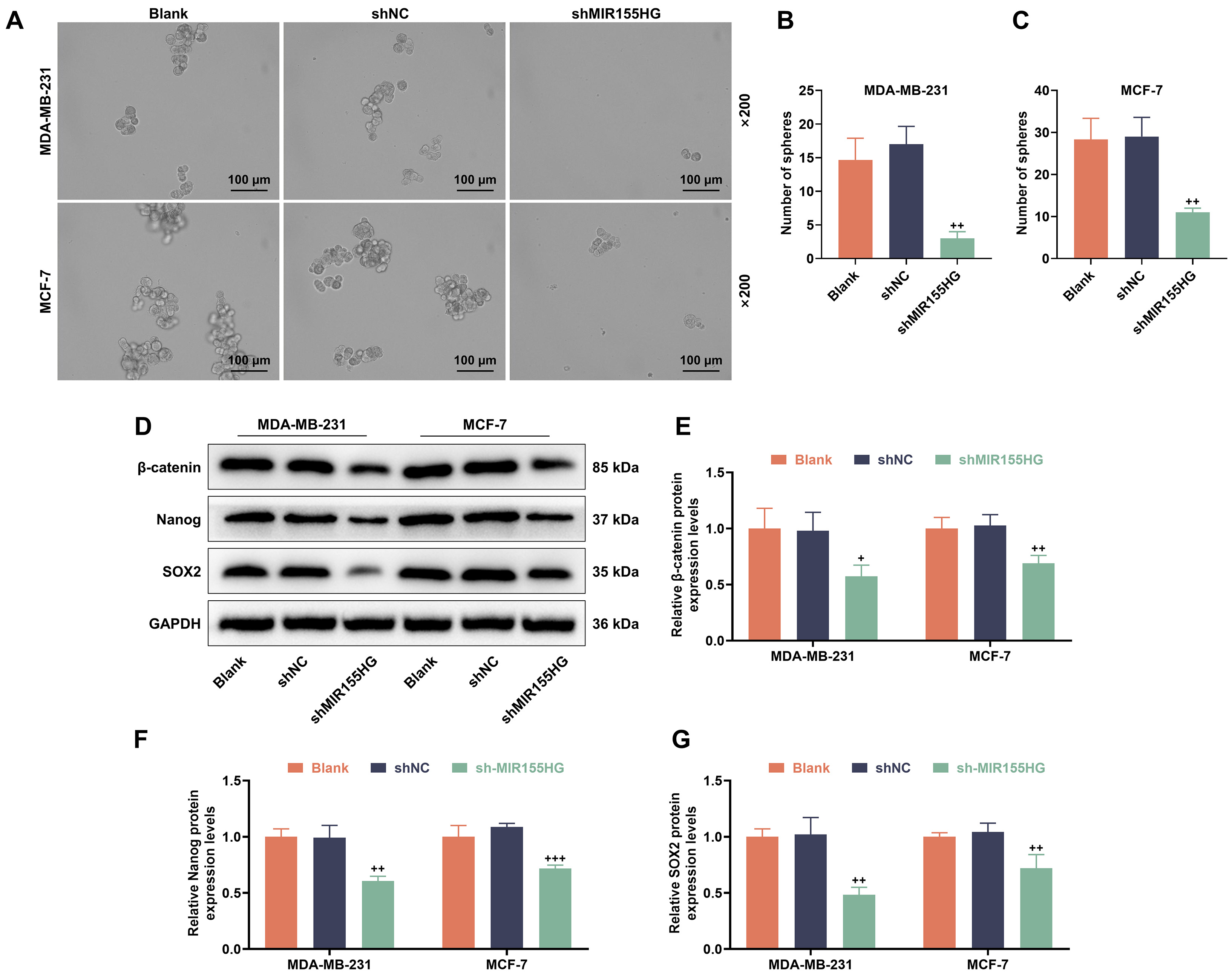 Fig. 3.
Fig. 3.
Knockdown of MIR155HG reduces stemness and inhibits
Wnt-related protein expression in BCSCs. (A–C) Stemness in
sh-MIR155HG-transfected MCF-7 and MDA-MB-231 stem cells after 2 Gy X-ray
irradiation for 24 h (Sphere formation assay). Magnification 200
To further assess whether MIR155HG regulates cellular apoptosis and
cycle induction in response to radiation, we examined apoptosis and cell cycle
profiles in MDA-MB-231 and MCF-7 stem cells after X-ray irradiation (2 Gy).
Sh-MIR155HG group exhibited increased apoptosis (p
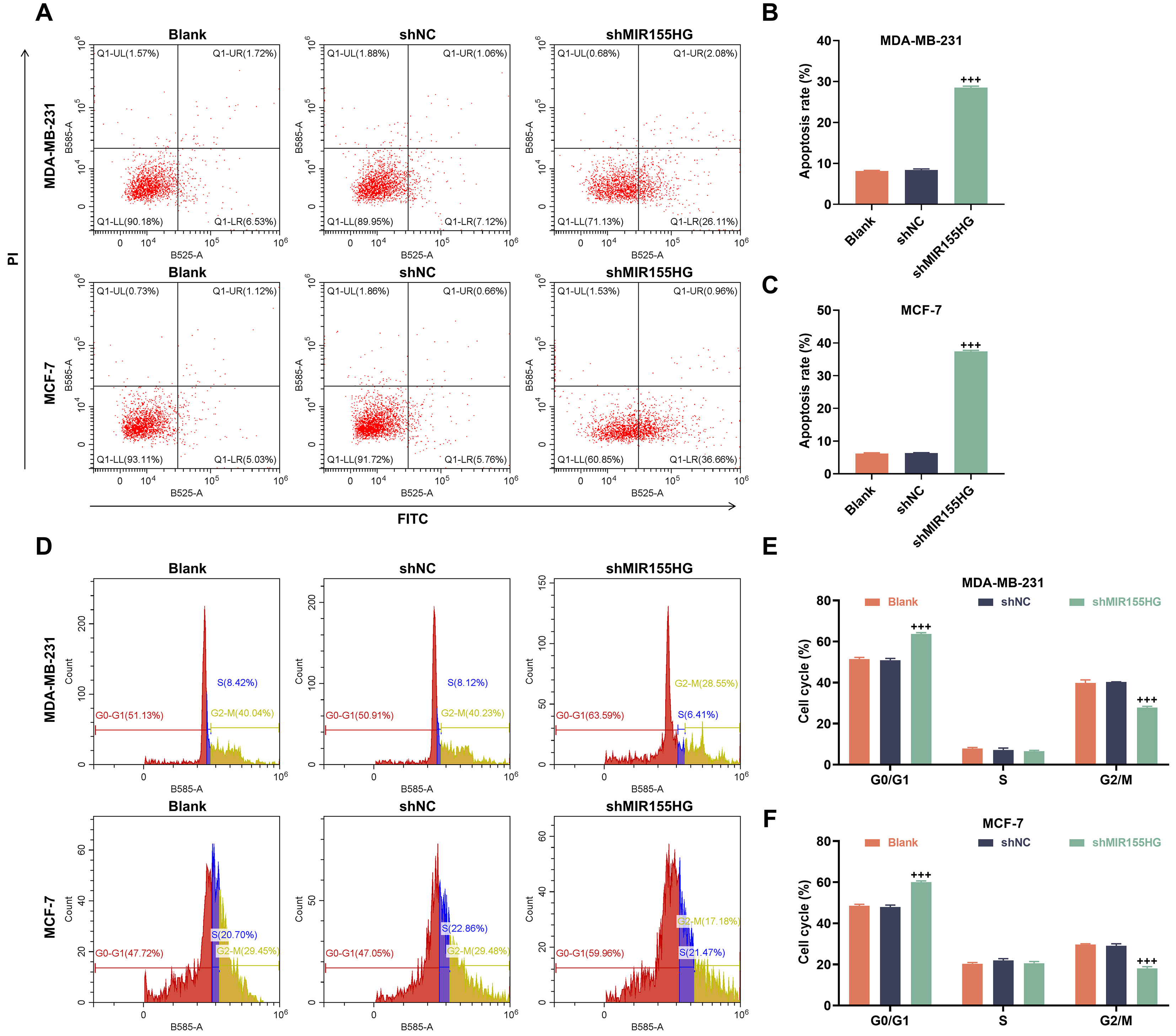 Fig. 4.
Fig. 4.
Knockdown of MIR155HG promotes apoptosis and induces G1
phase arrest in BCSCs. (A–C) Apoptosis in sh-MIR155HG-transfected MCF-7 and
MDA-MB-231 stem cells after 2 Gy X-ray irradiation for 24 h (flow cytometry).
(D–F) Cell cycle in sh-MIR155HG-transfected MCF-7 and MDA-MB-231
stem cells after 2 Gy X-ray irradiation for 24 h (flow
cytometry). n = 3. +++p
By means of JASPAR analysis, we identified four potential binding sites for the
NF-
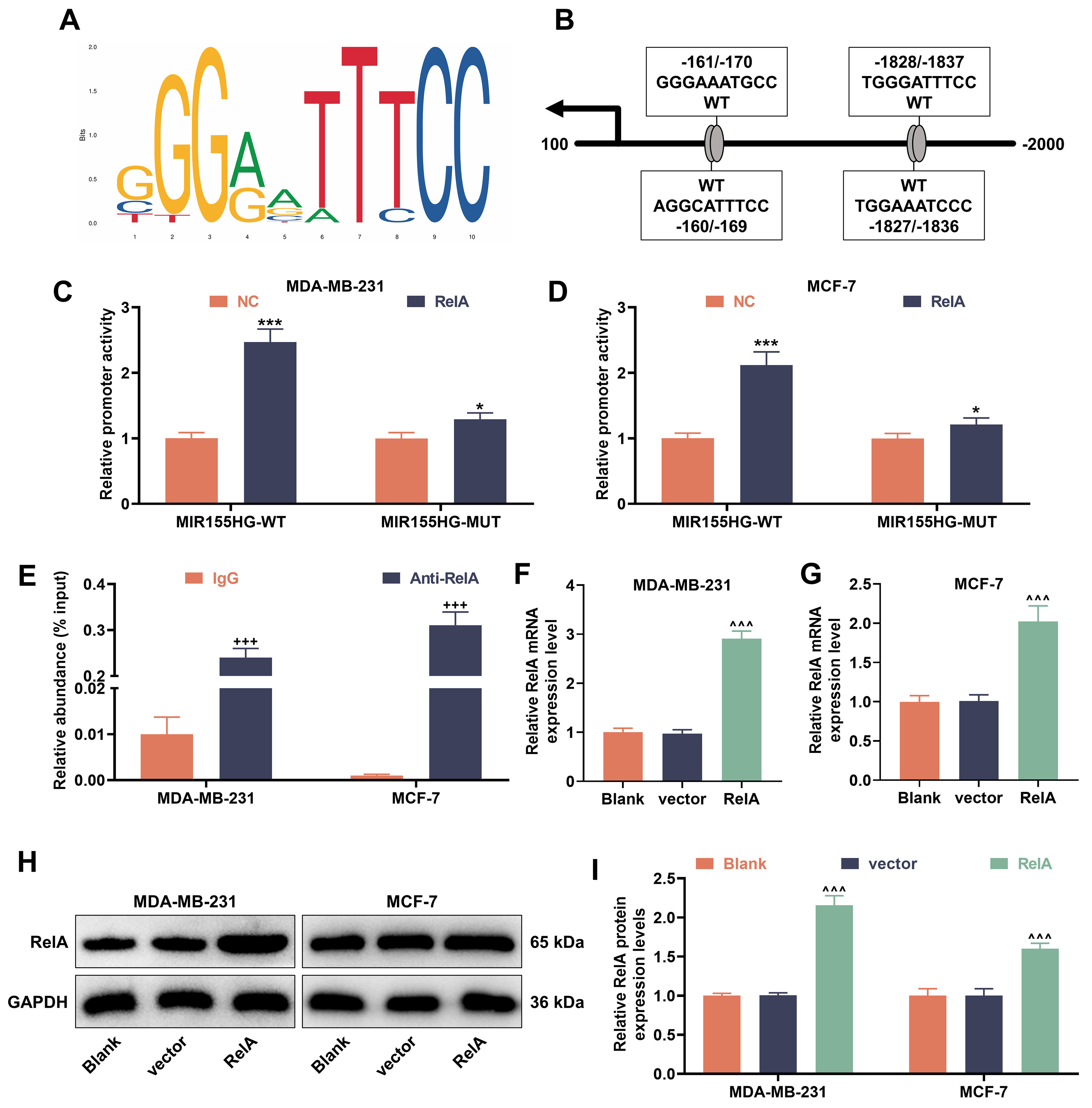 Fig. 5.
Fig. 5.
Nuclear factor kappa B (NF-
Following 2 Gy X-ray treatment for 24 h, colony survival rates were reduced in
the shMIR155HG+vector group, but were elevated in the shNC+RelA group (p
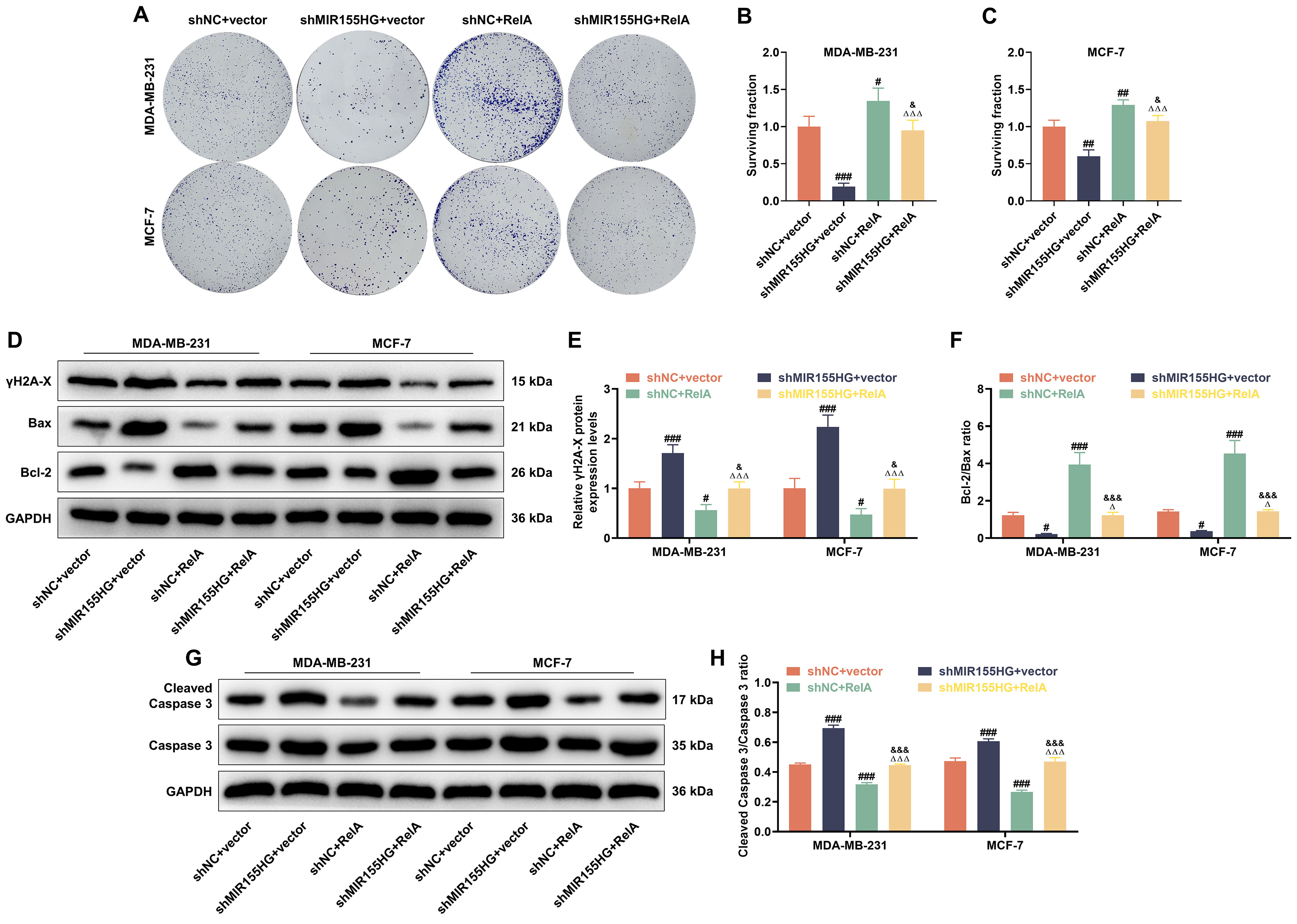 Fig. 6.
Fig. 6.
NF-
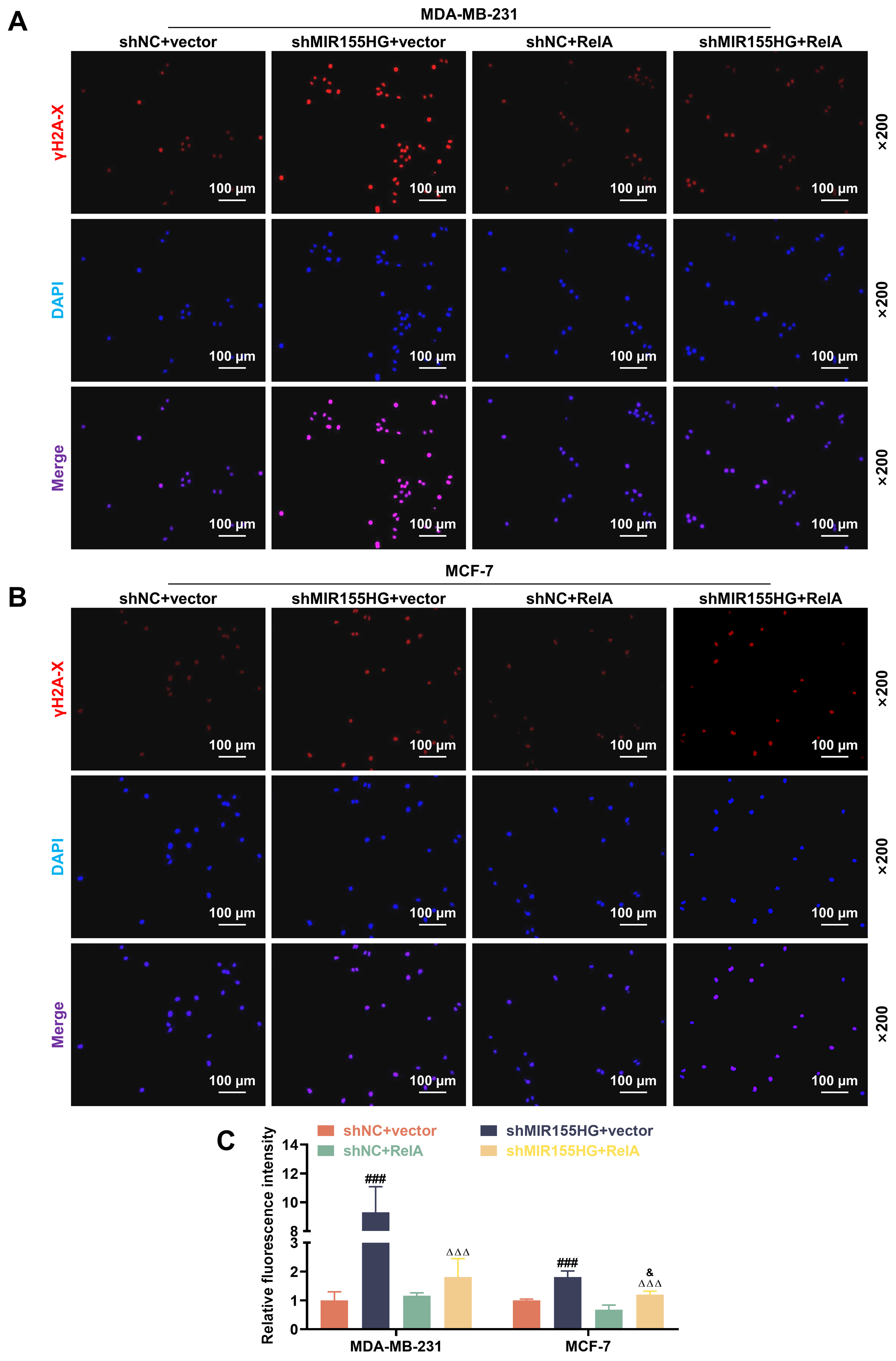 Fig. 7.
Fig. 7.
NF-
Sphere formation assays revealed that following 2 Gy X-ray treatment for 24 h,
tumor sphere was reduced in shMIR155HG+vector group yet increased in shNC+RelA
group (p
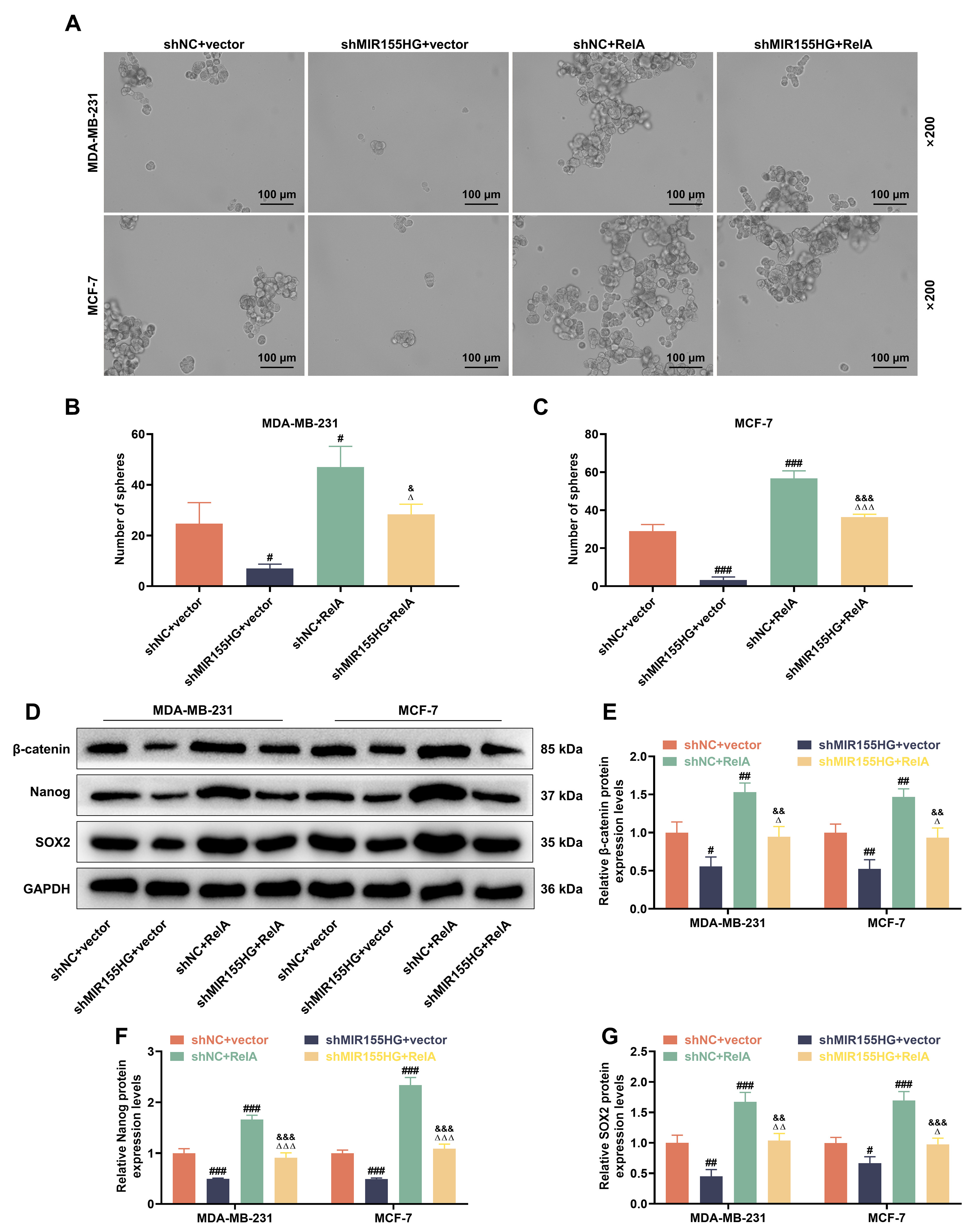 Fig. 8.
Fig. 8.
MIR155HG Knockdown reverses NF-
Flow cytometry analysis showed that following 2 Gy X-ray treatment for 24 h,
apoptosis was facilitated in stem cells in shMIR155HG+vector group, but dampened
in shNC+RelA group (p
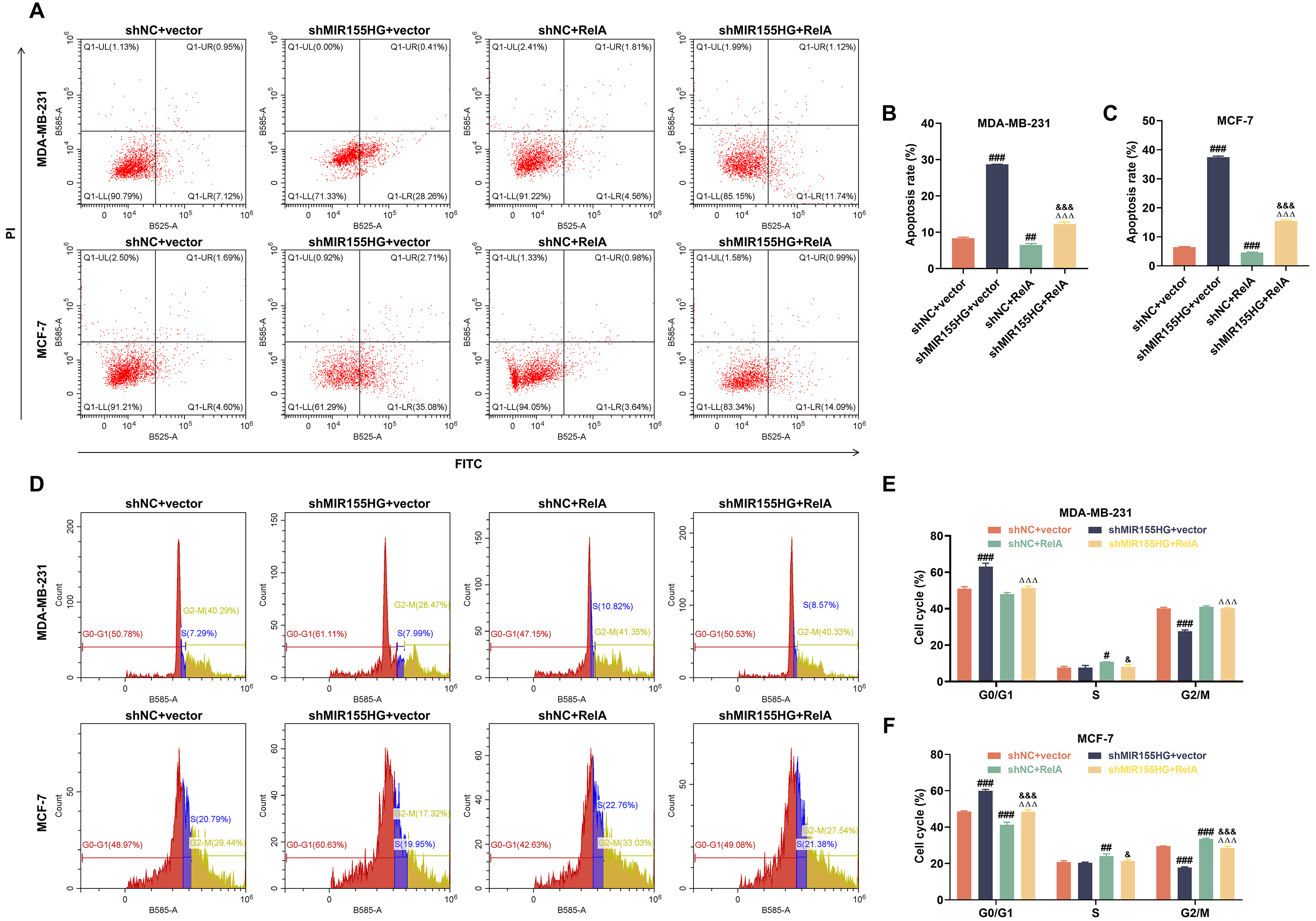 Fig. 9.
Fig. 9.
MIR155HG knockdown reverses NF-
This study revealed that BCSCs expressed MIR155HG at significant
levels, which enhanced cell resistance to radiation and DNA damage repair while
promoting stemness. The Wnt signaling pathway is crucial for cellular signal
transduction, influencing cell growth, differentiation, and stem
cell maintenance [22, 23]. Normally, the Wnt pathway regulates
cell fate decisions by modulating the stability and activity of
By attaching to particular gene promoter regions, the transcription factor
NF-
Of note, this study has some limitations. Despite using various experimental
methods to validate our findings, further verification using in vivo
models is necessary. Additionally, we need to collect and validate the expression
levels of MIR155HG and NF-
In summary, our findings indicate that NF-
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
Substantial contributions to conception and design: YBX. Data acquisition, data analysis and interpretation: GML, LY, CZR. Drafting the article or critically revising it for important intellectual content: YBX. Final approval of the version to be published: All authors. Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of the work are appropriately investigated and resolved: All authors. All authors contributed to editorial changes in the manuscript.
Not applicable.
Not applicable.
This work was supported by the Zhejiang Province Medical Project [2020KY842].
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.









