1 College of Pharmacy, National & Local Joint Engineering Research Center of Targeted and Innovative Therapeutics, IATTI, Chongqing University of Arts and Sciences, 402160 Chongqing, China
2 Tongji Medical College, Huazhong University of Science and Technology, 430074 Wuhan, Hubei, China
3 Institute for Viral Hepatitis, Chongqing Medical University, 400016 Chongqing, China
†These authors contributed equally.
Abstract
Background: Humankind have been struggling with colorectal cancer (CRC)
for long period with its rapid progression and invasive metastasis. By
hyperactivating IL-6/STAT3 signaling, CRC facilitates the capacity of
angiogenesis to plunder massive nutrients and develops gradually under harsh
condition. Methods: The Cancer Genome Atlas database was analyzed for
acquiring interferon-
Keywords
- colorectal cancer
- IFITM10
- angiogenesis
- STAT3 activation
Colorectal cancer (CRC) stands as one of the most common malignant tumors in human, ranking third in global incidence and second in mortality [1]. Although there has been great progress in the treatment of advanced CRC, the five-year survival rate of patients is still less than 15% [2]. Approximately 30–40% of CRC patients who have undergone surgical resection of primary tumors will eventually develop metastases several years later [3]. Metastasis of important organs and postoperative recurrence are still the main causes of death. With the study of the pathogenesis of CRC and the improvement of genetic testing capabilities, more and more molecular markers have been found [2, 4, 5]. However, effective therapies that may eliminate residual tumor cells to prevent metastasis are still missing [3]. Therefore, it attaches great significance for CRC to study the intrinsic tumorigenesis mechanism and find efficient therapeutic targets.
Interferon-induced transmembrane proteins (IFITMs) are widely found in mammals, and human IFITMs, first reported in 1984, represent a group of proteins inducible by interferon (IFN) [6]. IFITMs has comprise five members, specifically IFITM1, IFITM2, IFITM3, IFITM5 and IFITM10 [7]. Functionally, IFITMs mainly plays a role in immune signal transduction, control of cell proliferation, promotion of homotype cell adhesion, tumorigenesis, antiviral activity, promotion of bone matrix maturation and mineralization, and germ cell development [8, 9, 10]. Among them, IFITM1, IFITM2 and IFITM3 play an important role in antiviral invasion and serve as tumor markers [10, 11, 12]. IFITM3 overexpression can be used as a diagnostic marker in esophageal cancer patients. A study demonstrated that patients exhibiting IFITM3 overexpression had a significantly decreased five-year survival rate and increased recurrence rate, as indicated by Kaplan–Meier analysis [13]. IFITM3 is also found to be highly expressed in acute leukemia with the evidence that patients with high IFITM3 expression experience significantly shorter event-free survival (EFS) and overall survival (OS) than those with low expression [14]. And in colon cancer patients, IFITM1 can be an independent prognostic factor because IFITM1 expression was significantly higher in colorectal cancers with poor differentiation and worse overall survival [11], while IFITM5 mutations lead to V-type osteogenesis imperfecta. In addition, IFITM10 and cathepsin D are considered as molecular markers of breast cancer because it has been reported that custom small interfering RNAs targeting the Cathepsin D (CTSD)-IFITM10 fusion junction reduced expression of the fusion transcript and reduced breast cancer cell proliferation [15, 16]. Recent studies have demonstrated a significant correlation between elevated levels of IFITM10 expression and both early diagnosis and T stage classification in gastric cancer [17]. However, the correlation between IFITM10 level and CRC tumorigenesis has not been reported. Therefore, exploring the function of IFITM10 in CRC is of great significance to better understand the function of IFITM family proteins.
In this study, the expression of IFITM10 was analyzed using The Cancer Genome Atlas (TCGA) database through various bioinformatic methods, and the findings underscore the importance of aberrant IFITM10 expression in CRC, shedding light on its role in disease progression and patient outcomes. Combined with our in-depth analysis of IFITM10 associated functions and pathways, we have confirmed that IFITM10 was overexpressed in CRC and down-regulation of IFITM10 in CRC cells inhibit tumorigenesis both in vitro and in vivo. Further study we found that IFITM10 promote CRC tumorigenesis depends on IL-6/JAK/STAT3 pathway. Together, our study provides deeper understanding of the mechanisms behind CRC tumorigenesis and reveals the potential of IFITM10 as a diagnostic and therapeutic target of CRC.
The expression profiles of 33 distinct tumor types were analyzed, as cataloged by the Cancer Genome Atlas (TCGA, https://www.cancer.gov/ccg/research/genome-sequencing/tcga) initiative. It is important to note that this research adhered to the guidelines outlined by TCGA, which resulted in the waiver of ethical approval and the need for informed consent from the patients involved.
The mRNA expression of IFITM10 were compared between normal tissues and tumor tissues (data were acquired from TCGA, https://www.cancer.gov/ccg/research/genome-sequencing/tcga).
GSEA method in conjunction with the “clusterProfiler” package (version 3.17.0) were performed to run the set enrichment analysis [18]. The analysis involved the utilization of Hallmark MSigDB gene sets as the reference gene sets for the enrichment analysis.
In this study, all animal procedure was approved by the Ethics committee of
Chongqing University of Arts and Sciences (CQWU2022001 and 14/03/2022).
BALB/c-Nude mice (6–8 weeks) were bought from Huafukang Bio-Technology (Beijing,
China). Xenografts were constructed by subcutaneously injecting with 1
HEK293 were purchased from Procell Life Science&Technology Co., Ltd. (CL-0001, Wuhan, China), SW-480 and HCT-116 cells were obtained from Nanjing Cobioer Biosciences Co., Ltd. (Nanjing, China). HEK293, SW-480, NCM356, Lovo, RKO cells were cultured in DMEM (PM150210A, Pricella, Wuhan, China) medium with 10% fetal bovine serum (FBS, 1645210, Pricella, Wuhan, China) under 37 °C and 5% CO2 conditions, HCT-116 were cultured in McCoy’s-5A (PM150710, Pricella, Wuhan, China) with 10% FBS under 37 °C and 5% CO2 conditions. Human Umbilical Vein Endothelial Cells (HUVECs) (CTCC-0804-PC, Meisen, Jinhua, China) were cultured in Endothelial Cell Growth Medium 2 (CTCC-002-031, Meisen, Jinhua, China) under 37 °C and 5% CO2 conditions. For all the cell lines that were used in this study has been validated through STR profiling. Meanwhile, no mycoplasma was tested out among all cell lines.
The IFITM10 shRNA was applied to inhibit the IFITM10 expression in HCT-116 and
SW480 cell line. We used the following sequences: shIFITM10#1
(5
For protein analysis preparation, the cells were lysed utilizing a commercial lysis kit (P0013B, Beyotime, Shanghai, China) supplemented with a protease cocktail inhibitor (HY-K0013, MCE, Shanghai, China). Protein levels were quantified by BCA assay (P0009, Beyotime, Shanghai, China). Proteins were separated by sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and then were transferred onto polyvinylidene difluoride (PVDF) membranes (IPVH00010, EMD Millipore, Shenzhen, China). The PVDF membranes were then blocked using 5% non-fat milk in TBS buffer at room temperature (RT) for 30 minutes. Following this, the membranes were incubated overnight with primary antibodies, including anti-IFITM10 (1:1000, A22196, ABclonal, Wuhan, China), anti-p-STAT3 (1:1000, AP0070, ABclonal, Wuhan, China), anti-STAT3 (1:1000, 60199-1-Ig, Proteintech, Wuhan, China), anti-VEGF (1:1000, A12303, ABclonal, Wuhan, China) and anti-GAPDH (1:2000, GB11002, Servicebio, Wuhan, China). After three washes with TBST, the membranes were further incubated with secondary antibody (the HRP Goat Anti-Rabbit IgG (1:5000, AS014, ABclonal, Wuhan, China) and Anti-Mouse IgG (1:5000, AS003, ABclonal, Wuhan, China) at RT for 1.5 hour. Finally, the blots were imaged using Tanon 5200 Multi imager (Tanon, Shanghai, China).
Tube formation was performed as previously described [19]. In brief, 50 µL
Matrigel (BD Biosciences, FranklinLakes, NJ, USA) was added into a prechilled 96-well plate
before being placed in a 37 °C CO2 incubator for 30 minutes. P5
HUVECs were seeded on Matrigel with 2
HCT-116 cells with or without IFITM10 inhibition were seeded at a density of 5
CRC cells (with or without IFITM10 inhibition) were seeded onto coverslips and subsequently fixed with 4% Paraformaldehyde. Following fixation, the cells were blocked with 1% FBS for 1 hour, and incubated with indicated primary antibodies (anti-IFITM10, 1:200, ABclonal, Wuhan, China; anti-STAT3, 1:200, Proteintech, Wuhan, China) at 4 °C overnight. After the primary antibody incubation, the cells were treated with secondary antibodies (Beyotine, Shanghai, China) and incubated for 1 hour. Finally, the cell nuclei were stained with DAPI (RM02978, ABclonal, Wuhan, China) for 15 minutes at room temperature. Cells were imaged using a confocal laser scanning microscope (BX53, Olympus Corporation, Tokyo, Japan).
Total RNA was extracted using RNA Easy Fast Tissue/Cell kit (DP451, TIANGEN,
Beijing, China). The synthesized cDNA was obtained using ReverAid First Strand
cDNA Synthesis Kit (K1622, Thermo scientific, Shanghai, China). cDNA was then
mixed with Genious 2X SYBR Green Fast qPCR Mix (RK21206, Abclonal Technology,
Wuhan, China) and the following primers: VEGFA forward:
5
Kaplan-Meier analysis (log-rank test) was performed to examine the correlation
between the expression level of IFITM10 and indicated outcomes in the TCGA
dataset, including overall survival (OS), progression-free interval (PFI), and
disease-specific survival (DSS). Survival curves with a significance threshold of
*p
Statistical analyses were conducted using Prism V.9.0 (GraphPad Software, Inc.,
San Diego, CA, USA). Data are presented as mean
We analyzed the levels of IFITM10 mRNA in various types of cancer using data from the TCGA database (Fig. 1A). Among the paired 23 tumor types examined, IFITM10 was found to be significantly overexpressed in 8 of them (Fig. 1B). Interestingly, it is worth mentioning that only the prostate adenocarcinoma (PRAD) exhibited a significant decline in IFITM10 expression. Additionally, when comparing unmatched groups (Fig. 1C) and paired samples from the same patients (Fig. 1D), we observed that the expression of IFITM10 was considerably higher in CRC tumors compared to pericancerous tissues. Further, colorectal cancer cell lines and normal colon gland cells were applied to investigate the protein levels of IFITM10. Consistent with previous results, IFITM10 were higher in CRC cells compared to normal colon gland cell line (Fig. 1E). These results support the notion that IFITM10 may play a crucial role in tumorigenesis and tumor development of colorectal cancer.
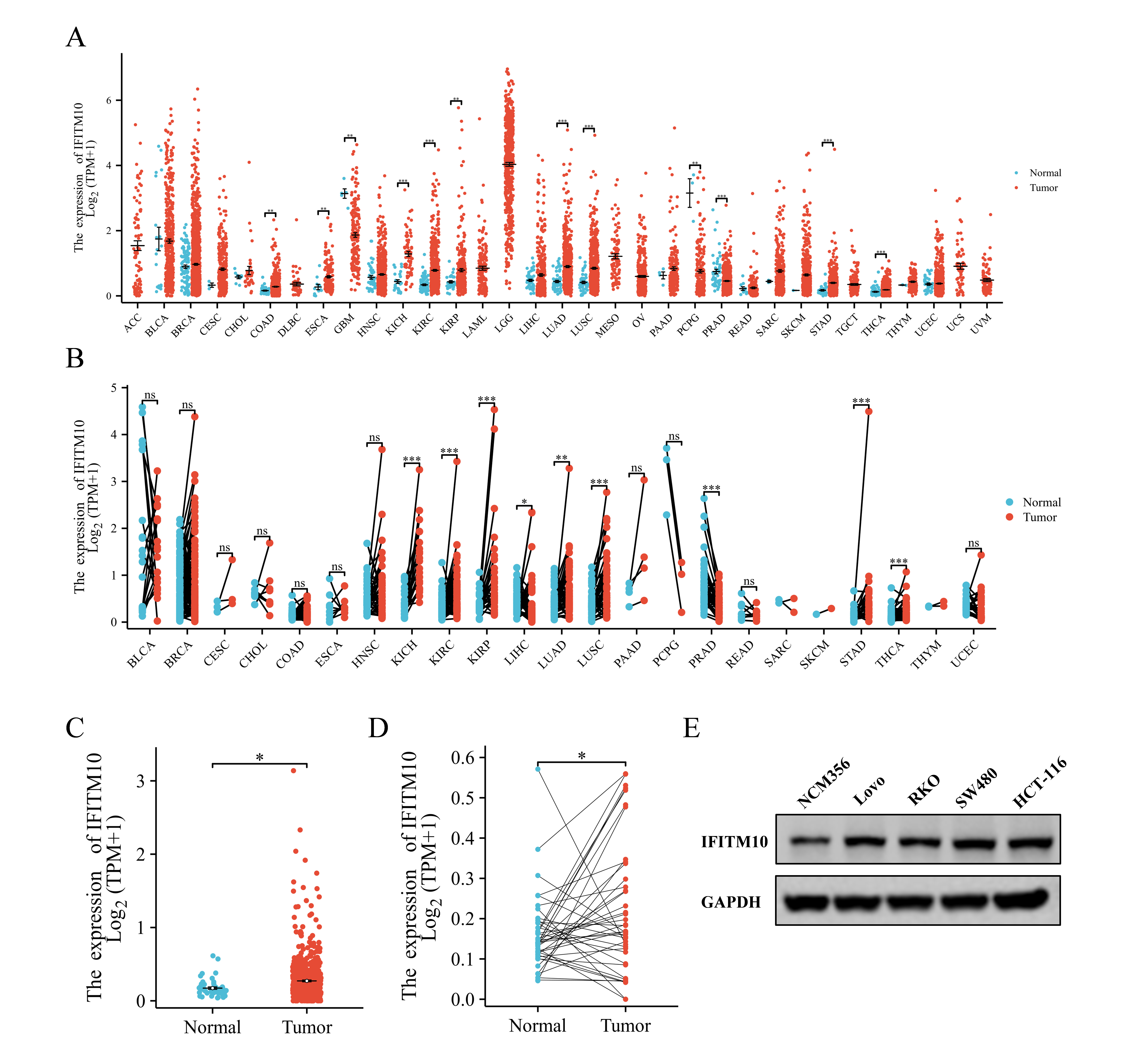 Fig. 1.
Fig. 1.
Interferon-
We found that IFITM10 is upregulated in CRC, however, little was known about the role of IFITM10 in CRC. Next, we performed GSEA with a public dataset and found that upregulation of IFITM10 was associated with angiogenesis (Fig. 2A). We further utilized IFITM10-targeted shRNA to knockdown IFITM10 in CRC cells (HCT-116 and SW480). Western blot showed the efficacy of IFITM10-1 knockdown (Fig. 2B). A mouse transplanted tumor model was then constructed using indicated cells with 5 mice per group, tumor growth curves was depicted and data showed slower tumor growth in the shIFITM10 groups, compared to control group (Fig. 2C). After 28 days, the mice were executed, and xenografts were collected and weighed. Xenografts formed by shIFITM10 cells are smaller, pale in the appearance, and lighter in weight (Fig. 2D,E). IHC staining showed that IFITM10 knockdown inhibited VEGFA expression in xenograft (Fig. 2F). CD31 immunohistochemical staining showed a decrease in tumor vascular density after IFITM10 inhibition (Fig. 2G). Furthermore, we detected the expression level of VEGFA in indicated xenografts. And data showed that VEGFA were reduced while IFITM10 was inhibited (Fig. 2H).
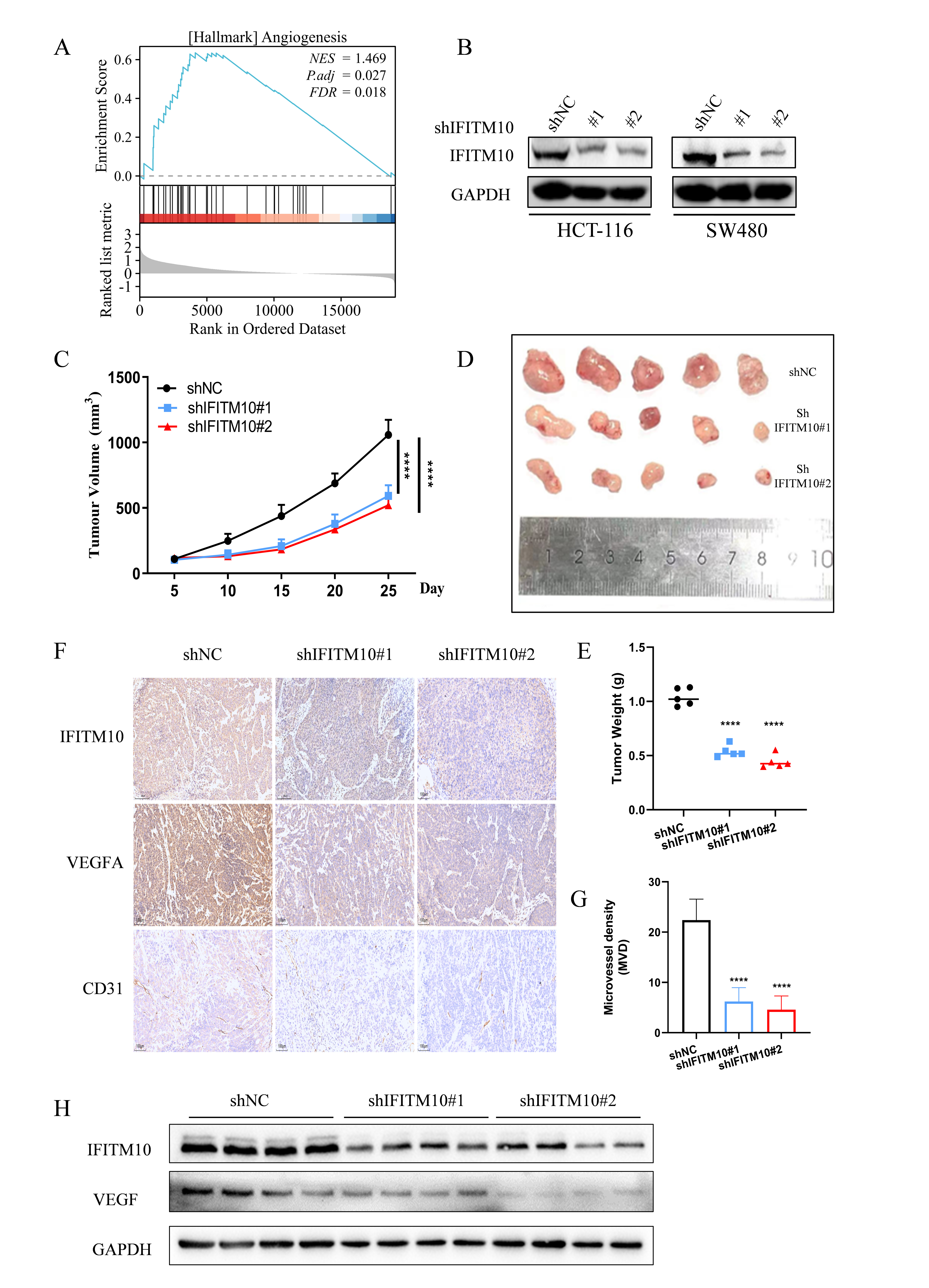 Fig. 2.
Fig. 2.
Inhibition of IFITM10 restrained tumor-induced angiogenesis
in vivo. (A) Gene Set Enrichment Analysis (GSEA) indicated enrichment
of IFITM10-associated Differentially Expressed Genes (DEGs) in the angiogenesis
signaling pathway. (B) HCT-116 and SW480 cells were transfected with indicated
shRNA, and the IFITM10 protein expression was analyzed. (C) The growth curve of
mice xenografts formed by shCON, shIFITM10#1 and shIFITM10#2 cells (n = 5). (D)
Mice xenografts formed by HCT-116 with or without IFITM10 intervention. (E) Tumor
weights after sacrificing the mice (n = 5). (F) Immunohistochemistry (IHC)
staining of xenografts using antibodies against IFITM10, VEGFA, and CD31. Scale
bar: 100 µm (G) Microvascular densities of the xenografts in
indicated groups, quantified by CD31 (n = 5). (H) Western blot results of
indicated xenografts. ****p
We next explored the role of IFITM10 during angiogenesis in vitro. The single gene co-expression heat map (from the TCGA database) revealed a positive correlation between IFITM10 and VEGFA (Fig. 3A). Meanwhile, we detected VEGFA secretion with IFITM10 inhibition by ELISA assay. Results showed that IFITM10 knockdown reduced VEGFA secretion levels in both HCT116 and SW480 cells (Fig. 3B,C). Then, tube formation test was performed using HUVECs, which were incubated in specified conditioned medium (CMs). Results showed that less tubular formation was observed in the shIFITM10 group (Fig. 3D–F). In addition, CCK8 assay showed that, the proliferation capacity of HUVECs were restrained with IFITM10 inhibition CMs compared to control CMs (Fig. 3G,H). Taken together, inhibition of IFITM10 restrained tumor-induced angiogenesis both in vivo and in vitro.
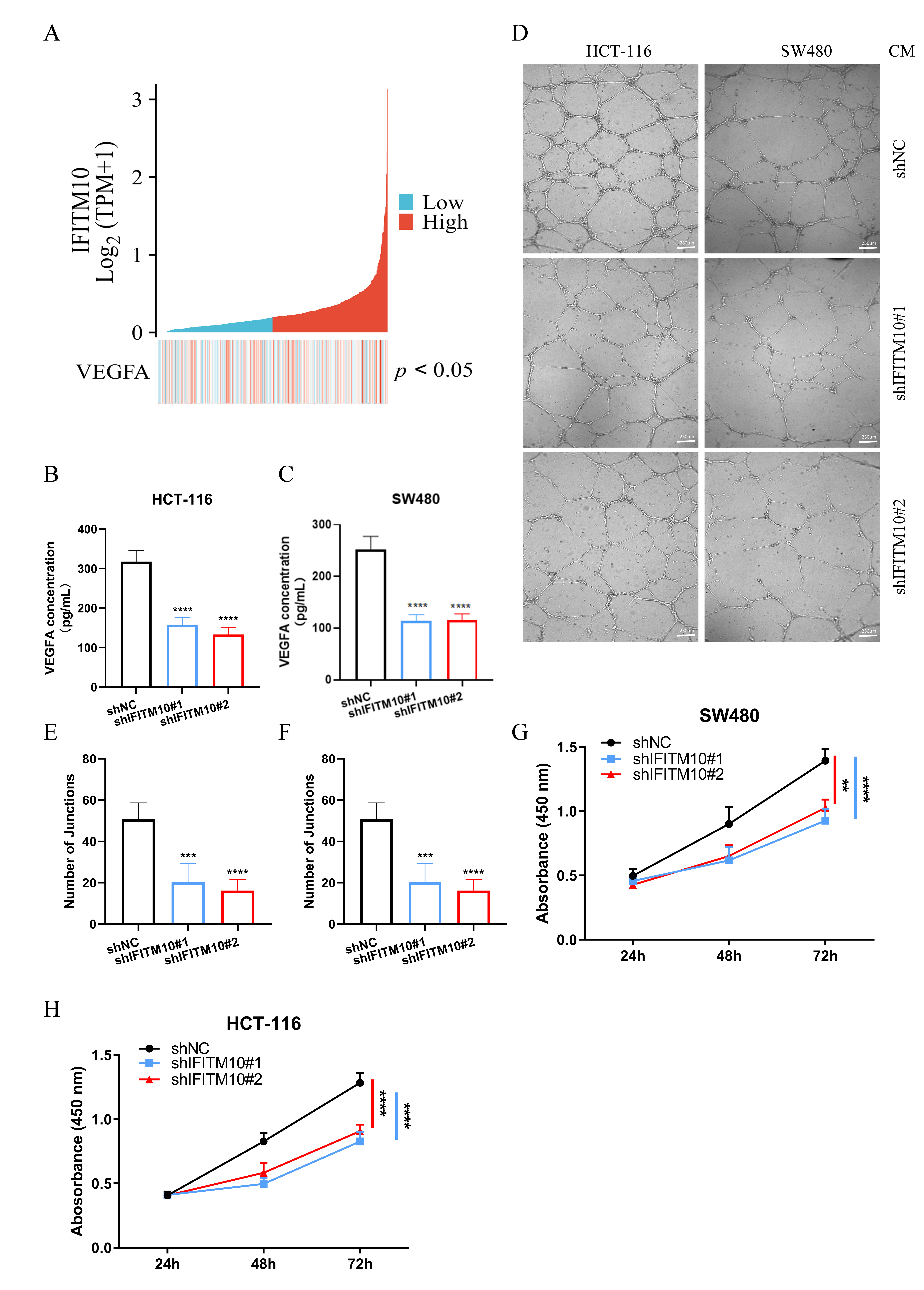 Fig. 3.
Fig. 3.
Inhibition of IFITM10 restrained tumor-induced angiogenesis
in vitro. (A) The single-gene co-expression heatmap in the TCGA
database shows a positive correlation between the expression of IFITM10 and VEGF
in CRC. (B,C) Detection of VEGFA concentration of conditioned medium obtained
from indicated cells by enzyme-linked immunosorbent assay (ELISA) (n = 3). (D–F)
Tube-formation assay was performed with Human Umbilical Vein Endothelial Cells
(HUVECs) incubating with indicated conditioned mediums (CMs) (n = 3). (G,H) The proliferation capacity
of HUVECs detected by CCK-8 assay. Scale bar: 250 µm. **p
To further investigated the molecular mechanism of IFITM10 during angiogenesis, GSEA enrichment analysis was performed. Results showed that IFITM10 upregulation may trigger IL-6/STAT3 signaling (Fig. 4A). Next, we found a positive correlation between IFITM10 and STAT3 with single-gene co-expression analysis (Fig. 4B). Thus, we hypothesized that IFITM10 may activate angiogenesis process through STAT3 activation. To validate this hypothesis, western blot assays were performed. Our data showed that with IFITM10 intervention, the phosphorylation of STAT3 was impaired (Fig. 4C). The extent of nuclear translocation of STAT3 reflects its functional status and its transcriptional activity. Next, we conducted a protein fractionation assay to separate nuclear and cytosolic proteins and found that IFITM10 facilitated the movement of STAT3 from the cytoplasm into the nucleus (Fig. 4D). Furthermore, an immunofluorescence assay demonstrated that IFITM10 knockdown resulted in a reduced aggregation of STAT3 in the nucleus (Fig. 4E). Together, these results indicate that IFITM10 promotes the activation of STAT3 signaling pathway.
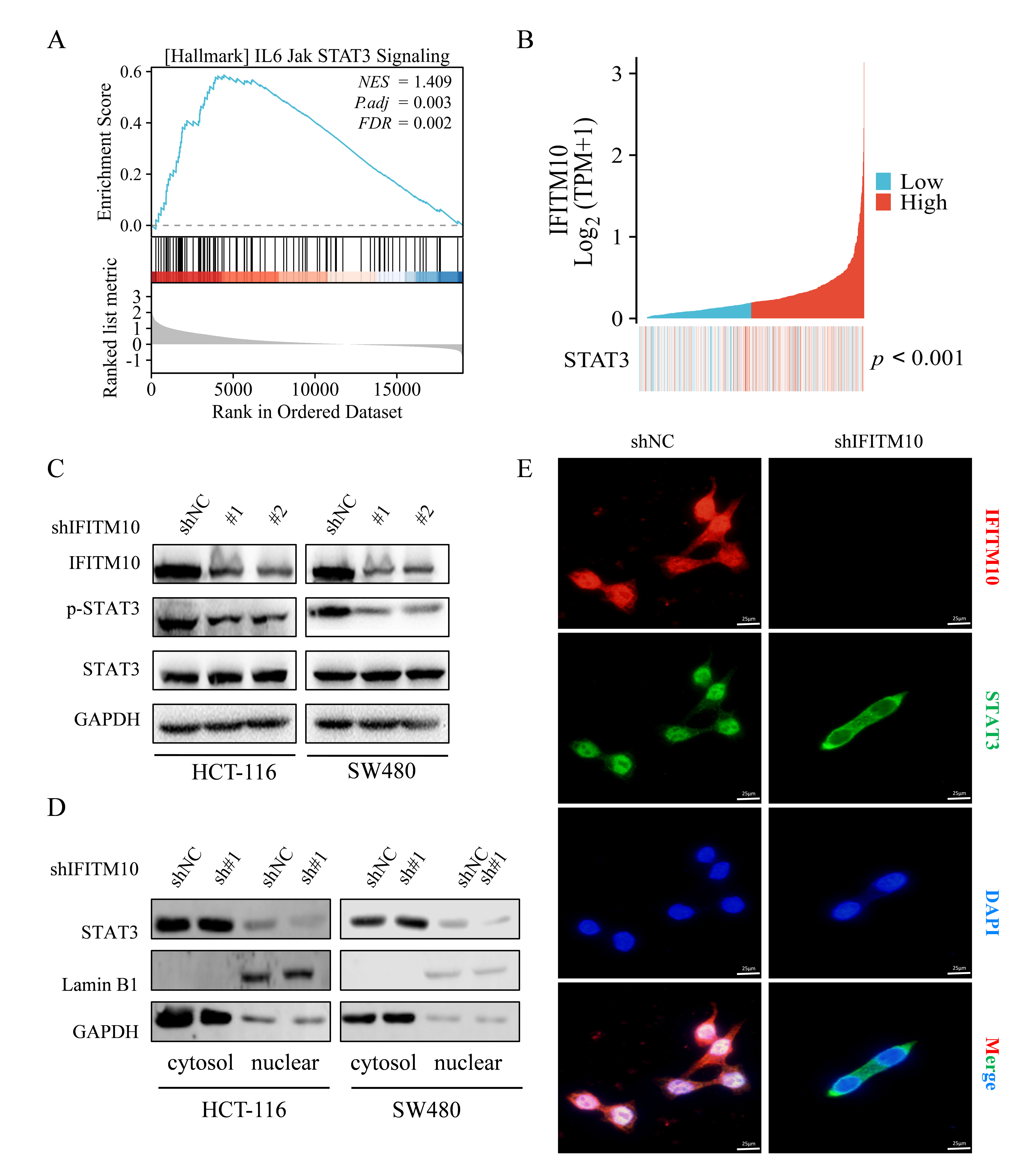 Fig. 4.
Fig. 4.
IFITM10 promotes the activation of STAT3 signaling pathway. (A) GSEA analysis indicated an association between IFITM10 expression and activation of the IL-6/JAK/STAT3 pathway. (B) The single gene co-expression heat map revealed significant relationships between IFITM10 and STAT3. (C) Western blotting was used to detect changes in STAT3 phosphorylation upon IFITM10 intervention in HT-116 and SW480 cell lines. (D) Nuclear and cytoplasmic proteins were separated and analyzed by western blotting after IFITM10 intervention in HT-116 and SW480 cell lines. (E) An immunofluorescence assay was conducted to assess changes in the nuclear localization of STAT3 in response to IFITM10 intervention. Scale bar: 25 µm.
The transcriptional activity of STAT3 plays critical roles in both VEGFA secretion and angiogenesis. We have showed that IFITM10 promoted angiogenesis, VEGFA expression, and STAT3 activation. We then determined whether IFITM10-mediated angiogenesis was dependent on the STAT3 activation. By western blot assay, we found that IFITM10 overexpression increased STAT3 phosphorylation and upregulated VEGFA, but those phenomena were reversed with STATTIC application (a pharmacological inhibitor of STAT3 activation) (Fig. 5A). Further, consistent results were obtained through VEGFA secretion detection by ELISA assay (Fig. 5B,C). We next incubated HUVECs with indicated CMs. Results showed that, compared to control group, culturing with CMs that obtained from IFITM10-overexpressing cells, HUVECs exhibited better migratory ability and this effect was inhibited with STAT3 inhibition (Fig. 5D–F). Therefore, we concluded that the angiogenic function of IFITM10 is mediated by STAT3 activation.
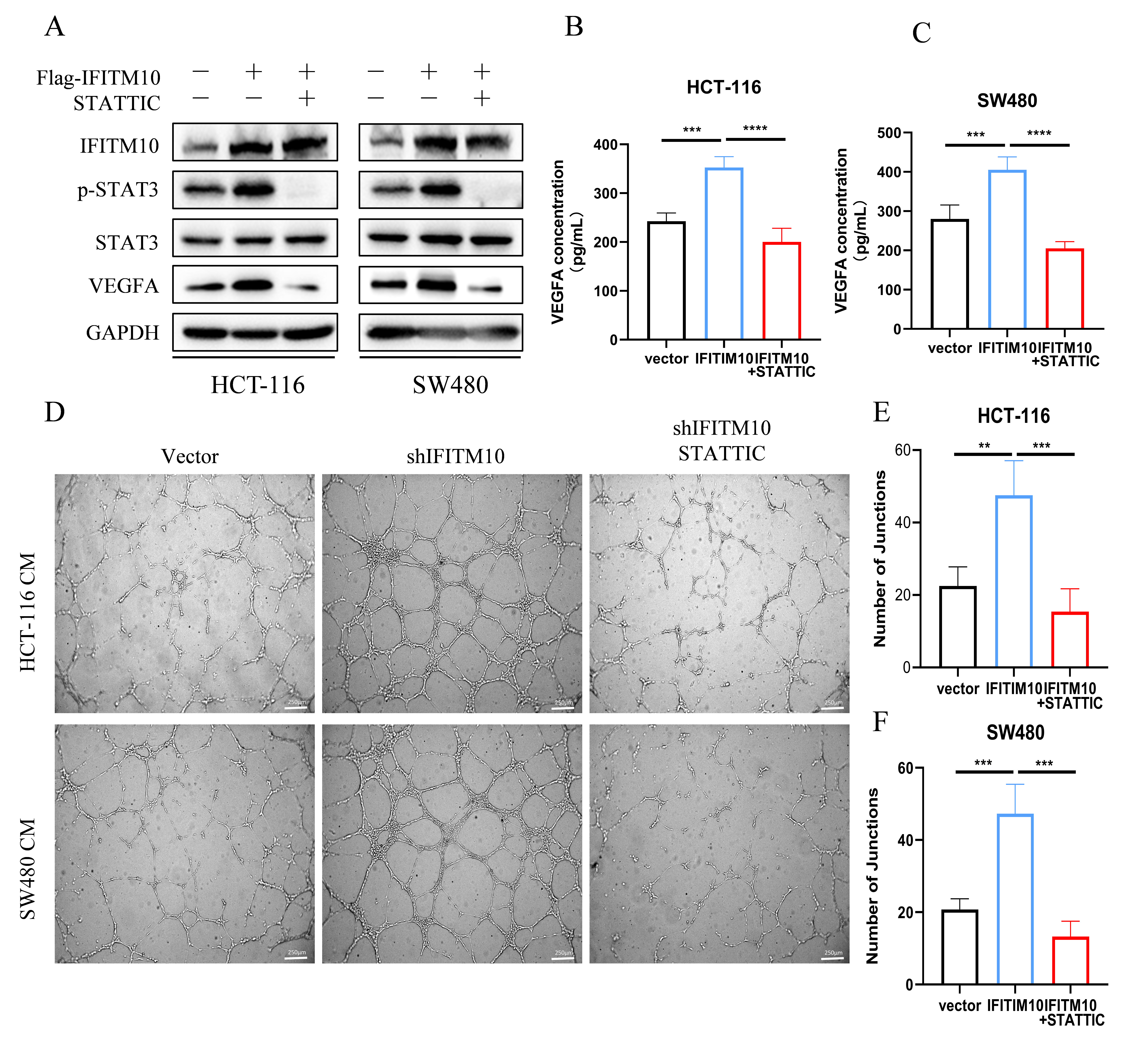 Fig. 5.
Fig. 5.
IFITM10 mediated angiogenesis depend on STAT3 activation. (A)
The expression level of indicated targets with/without IFITM10 intervene or
STATTIC. (B,C) The section level of VEGFA was detected by ELISA assay. (D–F)
Tube-formation assay was performed with HUVECs incubating with indicated CMs
through indicated treatment (n = 3). Scale bar: 250 µm, **p
To evaluate the diagnostic and prognostic significance of IFITM10 in CRC, we
conducted several analyses. Firstly, we utilized receiver operating
characteristic (ROC) curves to assess the ability of IFITM10 to differentiate CRC
diagnosis. The results showed that IFITM10 had a sensitivity and specificity for
diagnosing CRC, as indicated by the area under the curve (AUC) of 0.622 (Fig. 6A). Additionally, we performed Kaplan-Meier analysis to determine the predictive
value of IFITM10 for clinical outcomes. The analysis revealed that high IFITM10
expression was associated with worse overall survival (hazard ratio [HR]: 2.18,
p
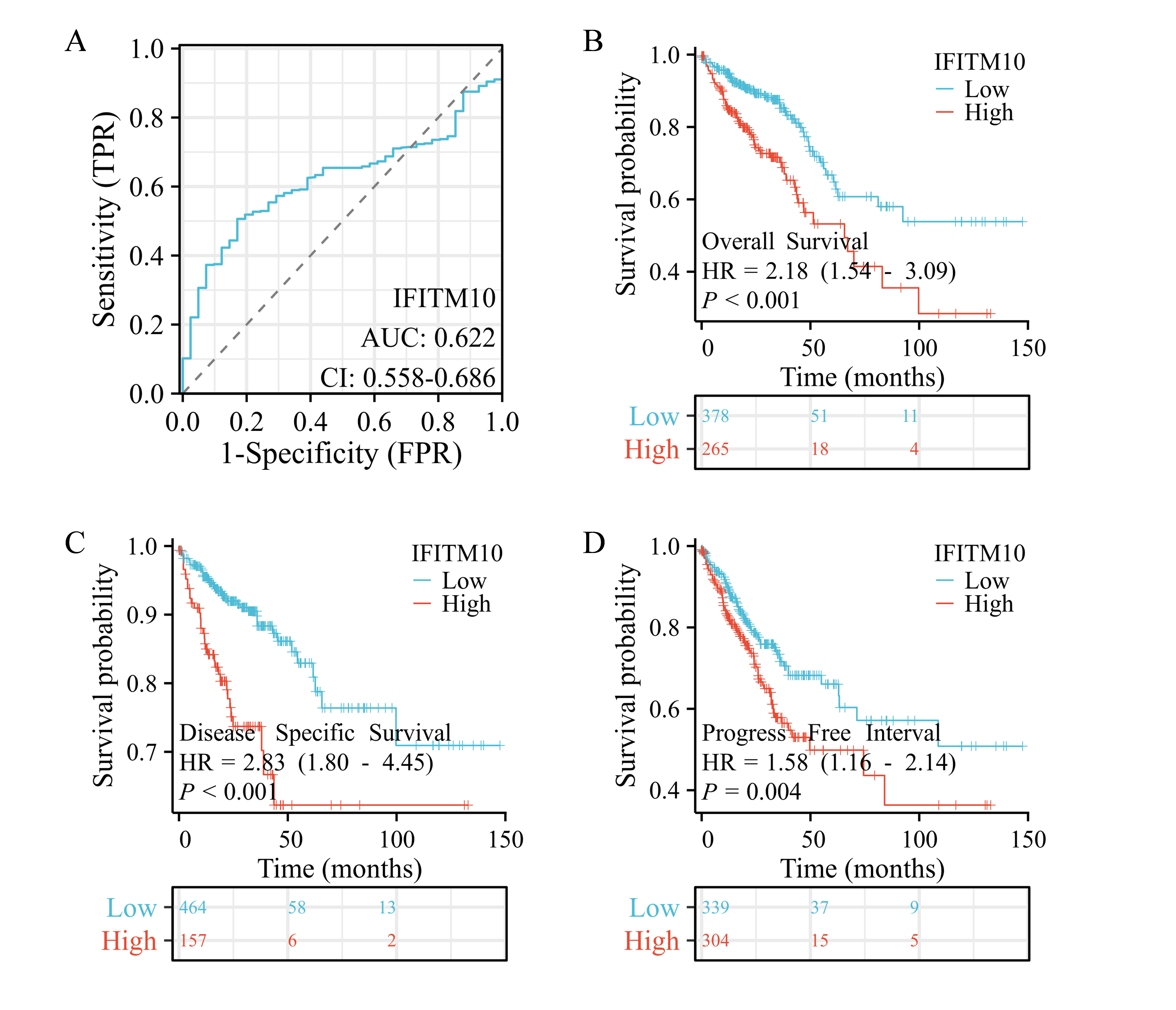 Fig. 6.
Fig. 6.
Predictive value of IFITM10 in colorectal cancer (CRC) diagnosis and prognosis. (A) Receiver operating characteristic curve analysis evaluated the diagnostic performance of IFITM10 in CRC. (B–D) Kaplan-Meier analyses compared overall survival, disease-specific survival, and progression-free interval between CRC patients with high and low IFITM10 expression.
Our research has revealed that IFITM10 plays a role in tumor angiogenesis in CRC, by activating STAT3 signaling, which also suggested that IFITM10 promotes the tumorigenesis of CRC through facilitating the tumor blood vessels generation which feed the tumor. These findings suggest that IFITM10 could be a promising therapeutic target for anticancer treatment, specifically targeting angiogenesis in CRC.
The expressions of IFITMs in different tissues have been well studied [7, 10, 20]. IFITM1, IFITM2, IFITM3, and IFITM10 are expressed in 34 types of normal tissues. Remarkably, when comparing the expression levels of these IFITMs between tumor tissues and their respective normal counterparts, a prevalent trend emerged: the majority of tumor tissues exhibited higher expression levels of IFITMs compared to those in normal tissues [10]. Our analysis of TCGA data aligns with the conclusions of another research group, indicating that IFITM10 is overexpressed in the majority cancers, including CRC [17]. Notably, we observed that IFITM10 expression does not decrease in any of the tumors. These results suggest that IFITM10 may facilitate tumorigenesis and tumor progression.
Our further study found that the loss of IFITM10 impairs cancer angiogenesis both in vitro and in vivo. In terms of mechanism, IFITIM10-mediated CRC angiogenesis is highly correlated with STAT3 activation, which is regulated by STAT3 phosphorylation. STAT3, as a key transcription factor, has been shown to regulate the transcription of vascular growth factors, including VEGF [21, 22]. Our qPCR results showed that IFITM10 up-regulated VEGF mRNA expression. Notably, when we inhibited STAT3 activity through drug inhibition, the upregulation of these genes by IFITM10 was attenuated. These findings suggest that IFITM10 regulates the expression of VEGF related genes through STAT3-mediated transcriptional regulation.
Existing comprehensive bioinformatics analysis has shown that IFITM1, IFITM2 and IFITM3 serve as a prognostic markers for kidney cancer, specifically in kidney clear cell carcinoma, suggesting an unfavorable prognosis. This correlation is attributed to increased immune cell infiltration, particularly in endothelial cells and cancer-associated fibroblasts [10, 13], IFITM1 emerges as a potential marker molecule for both rare squamous cell/adenosquamous carcinoma (SC/ASC) and common adenocarcinoma (AC), becauce the percentage of positive IFITM1 expression was significantly higher in SC/ASC patients with high tumor node metastasis (TNM) stage, lymph node metastasis, invasion, and no resection surgery compared to patients with low TNM stages, no lymph node metastasis, no invasion, and resection surgery [23]. Products of IFITM5 and IFITM10 cannot be used as markers of tumor prognosis because in a summary covers the mutations, expression, and functions of human IFITMs, along with an analysis of their varied expression levels in normal and diseased tissues showed that while FITM1, IFITM2, and IFITM3 share similar motif patterns and functions, IFITM5 and IFITM10 exhibit considerable diversity compared to them [10]. This difference may be related to the database used. By analyzing the diagnostic and prognostic value of IFITM10 in CRC, we assessed the clinical relevance of our findings. Our ROC curve analysis showed that IFITM10 was sensitive and specific for CRC diagnosis. In addition, consistent with Liu et al.’s study [10], our Kaplan-Meier analysis showed that high IFITM10 expression was associated with worse overall survival (OS), disease-free survival (DFS), and progression-free interval (PFI) in CRC patients. These results suggest that IFITM10 may serve as a prognostic biomarker for CRC.
In conclusion, our study provides important insights into the role of IFITM10 in the regulation of angiogenesis through the STAT3 signaling pathway in CRC progression. IFITM10 acts as a key regulator of CRC proliferation, highlighting its potential as a therapeutic target for CRC. Further study of the underlying molecular mechanisms and exploration of targeted treatment strategies for IFITM10 may pave the way for more effective treatment of CRC.
The data used to support findings of the study are available from the corresponding author upon request.
Conceptualization: XL and KO; Methodology: YL and MW; Software: JJ, FP, WL and DT; Validation: ZC, YL and MW; Formal analysis: YL, FP and DT; Investigation: JJ, FP, WL, YL and MW; Resources: ZC; Data curation: YL and MW; Writing—original draft preparation, YL and MW; Writing—review and editing, XL and KO; Supervision: XL, DT and KO; Project administration: DT and KO; Funding acquisition: YL and KO. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
All animal procedure was approved by the Ethics committee of Chongqing University of Arts and Sciences (CQWU2022001 and 14/03/2022).
Not applicable.
The study was supported by the Natural Science Foundation of Yongchuan (2023yc-jckx20079), District Science and Technology Research Program of Chongqing Municipal Education Commission (KJZD-M202101302, KJQN202301322), Natural Science Foundation of Chongqing, China CSTC (CSTB2023NSCQ-MSX0817).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






