1 Department of Orthopedics, Shanghai Fifth People’s Hospital, Fudan University, 200240 Shanghai, China
†These authors contributed equally.
Abstract
Background: Bone tissue engineering offers a new approach for the treatment of bone defects, with angiogenesis being critical to the survival and development of tissue-engineered bone. Mineralized osteoblasts (MOBs) have been reported to promote vascular formation by endothelial cells (ECs) through the secretion of exosomes containing a variety of angiogenic factors. The aim of the present study was to investigate the effect of miR-423-5p contained within exosomes derived from MOBs (MOB-Exos) on EC angiogenesis. Methods: The Cell Counting Kit-8 (CCK-8), scratch wound healing, Transwell migration, and tube formation assays were conducted to assess the in vitro effects of MOB-Exos on EC proliferation, migration, and tubule-forming capabilities. The miR-423-5p level in MOB-Exos was quantified using quantitative polymerase chain reaction (qPCR). Co-culture experiments were used to study the exosomal transport of miR-423-5p and its angiogenic effects. High-throughput sequencing was used to identify differentially expressed genes, and a dual luciferase reporter assay to determine whether CXCL10 was a direct target gene for miR-423-5p. Furthermore, the in vivo effect of MOB-Exos-derived miR-423-5p on angiogenesis was evaluated using a subcutaneous xenograft model. Results: MOB-Exos significantly promoted the in vitro proliferation, migration, and tubule formation of ECs. A high level of miR-423-5p was found in MOB-Exos and promoted the angiogenesis of ECs. The CXCL10 gene was significantly downregulated in ECs upon miR-423-5p mimic transfection. Dual luciferase reporter assay confirmed the direct binding of miR-423-5p to the CXCL10 gene. miR-423-5p derived from MOB-Exos upregulated expression of the vascular markers CD31 and vascular endothelial growth factor (VEGF) in vivo, thus underscoring its angiogenic potential. Conclusion: This study found that miR-423-5p derived from MOB-Exos could potentially enhance EC angiogenesis via the regulation of CXCL10. Therefore, exosomes are promising therapeutic candidates for clinical bone defects.
Keywords
- angiogenesis
- exosomes
- miR-423-5p
- CXCL10
- osteoblasts
- endothelial cells
Bone is an important organ in the musculoskeletal system and plays major roles
in load-bearing, locomotion, and the protection of internal organs. Normally,
bone tissue is in a balanced state of constant resorption and remodeling, thereby
allowing it to adapt to growth and mechanical loading. Trauma, inflammation, and
the removal of bone tumors can cause the loss of bone tissue [1]. Although bone
tissue can self-regenerate, severe bone defects (
Exosomes range from 30 to 200 nm in diameter and include proteins, nucleic acids, and other bioactive substances with important effects on cell-cell and inter-organ communication [8, 9]. Bone mesenchymal stem cell (BMSC)-derived exosomes promote angiogenesis by endothelial cells (ECs) [10, 11]. Matrix metalloprotein 2 (MMP2) found within exosomes secreted by mature osteoblasts (OBs) can promote EC angiogenesis [12]. In addition, exosome miR-139-5 secreted by senescent OBs can accelerate the aging and regeneration of ECs [13]. Because of their complex structural properties, exosomes are widely used in tissue engineering to carry drugs and active molecules. BMSC-derived exosomes combined with hydrogel promote the differentiation of BMSCs and cartilage repair [14]. The implantation of an adipose stem cell-derived exosome composite with polylactic acid-hydroxyacetic acid co-polymer scaffolds into cranial defects significantly enhanced bone regeneration in mice [15]. Placental mesenchymal stem cell (MSC)-derived exosomes combined with chitosan hydrogel were also shown to enhance vascular development and angiogenesis, resulting in effective treatment of posterior limb ischemia in a mouse model [16]. Furthermore, MSC-derived exosomes encapsulated with silk fibroin hydrogel increased the retention time of exosomes in tissues and promoted blood perfusion in ischemic hind limbs [17].
Among the exosome components, miRNAs have drawn considerable attention due to their regulatory functions in various biological processes. Accumulating evidence shows that exosomal miRNAs are closely involved with angiogenesis. Exosomal miR-126 derived from endothelial progenitor cells stimulates the angiogenic activity of ECs [18]. In contrast, exosomal miR-9 derived from nasopharyngeal carcinoma cells inhibits the migration of ECs [19]. These results indicate that exosomal miRNA may participate in angiogenesis. Previous studies have also shown that miRNA-423-5p is strongly related to tumorigenesis, drug sensitivity, tumor prognosis, and cardiovascular disease [20, 21, 22]. Furthermore, exosomal miRNA-423-5p derived from human adipose MSCs significantly enhances angiogenesis by ECs [23]. However, it is not known whether OB-derived exosomal miRNA-423-5p is associated with angiogenesis, or whether it targets any of the genes associated with angiogenesis.
Our previous studies have shown that exosomes promote EC angiogenesis [12, 24]. However, it is not known whether miR-423-5p derived from mineralized OB exosomes (MOB-Exos) can regulate the vascularization of ECs. The aim of the present study was therefore to investigate the angiogenic effect of miR-423-5p derived from MOB-Exos, and whether this may be a potential strategy for the clinical repair of bone tissue defects.
The endothelial cell line bEnd.3 and the OB cell line MC3T3-E1, derived from Mus musculus (mouse), were obtained
from the cell bank of the Chinese Academy of Sciences. Both cell lines underwent
authentication via STR profiling and were confirmed negative for mycoplasma
contamination. bEnd.3 cells were maintained in Dulbecco’s Modified Eagle Medium
(DMEM; Cat SH30022.01, Hyclone, Salt Lake City, UT, USA) supplemented with 10%
fetal bovine serum (FBS, Cat 10437-028; Gibco, Los Angeles, CA, USA). MC3T3-E1
cells were cultured in Minimum Essential Medium
Three experimental groups were established: a blank control, MOBs alone, and MOBs treated with GW4869, an inhibitor of exosome release (Cat: HY-19363, MedChemExpress, Lawrenceville, NJ, USA). In all three groups, MC3T3-E1 cells were cultured in six-well plates for 24 h. GW4869 (10 µM) was added to the MOBs+GW4869 group and the cells incubated overnight. Transwell chambers with a pore size of 0.4 µm (Cat 3422, Corning, Corning, NY, USA) were then added to the wells and ECs were seeded into the upper compartment of the chamber. After two days of co-culture, the ECs were harvested by trypsin digestion (Beyotime, Nanjing, China) and RNA was extracted from the cell pellets to evaluate the miR-423-5p content.
After induction for 21 d, the osteogenic medium was replaced by DMEM containing
10% exosome-free FBS and the cells cultured for a further 2 d. MOB-Exos were
then isolated using Exoquick reagent (Cat 4478359, Invitrogen, Waltham, MA, USA).
The culture medium was initially centrifuged at 3000
The detection of nanoparticles was carried out using a nanoparticle tracking
analysis instrument (Particle Metrix, Munich, Germany). A standard solution of
100 nm polystyrene microspheres (Thermo, Waltham, MA, USA) was diluted
250,000-fold with ultrapure water, and 1 mL of this diluted standard solution was
then used to automatically calibrate the instrument prior to testing. Exosome
samples were diluted 500-fold to 1.0
Exosome samples were adjusted to the appropriate concentration or viscosity. Approximately 15 µL of exosome sample was absorbed onto copper mesh and left for 1 h. They were then dried with filter paper and stained with 15 µL of a 2% uranyl acetate solution (Thermo, Waltham, MA, USA). The stained exosomes were subsequently observed with a transmission electron microscope (JEM-1400, JEOL, Tokyo, Japan).
MOB-Exos (200 µg) were mixed with PKH-67 dye (Cat MKCK8884, Sigma, St. Louis, MO, USA) at room temperature. A 1% BSA solution was used to stop the labeling procedure, and the stained exosomes were then added to EC culture medium. After incubation for 4 h, ECs were fixed with 4% neutral paraformaldehyde, stained with 4’,6-diamidino-2-phenylindole (DAPI), and the images saved and merged by ImageJ for Mac 1.51 (https://imagej.net/ij/).
Cell proliferation was assessed using the Cell Counting Kit-8 (CCK-8; Cat CK04,
Dojindo, Sendai-shi, Japan) assay. Each experimental group of bEnd.3 cells was
seeded into 96-well plates at 2
bEnd.3 cells were inoculated into 6-well plates at a density of 5
bEnd.3 cells were first diluted to 1
A pipette tip and Matrigel were placed in a 4 °C refrigerator the day
before the experiment. The Matrigel was used to coat 96-well plates. bEnd.3 cells
were diluted to 4
After cell lysis with Trizol reagent (Beyotime, Nanjing, China), samples were
mixed with chloroform at a 1:5 ratio and then shaken vigorously for 15 sec.
Following centrifugation at 12,000 rpm for 10 min, the upper aqueous phase
containing RNA was transferred to a new tube and mixed with an equal volume of
isopropyl alcohol. After the same centrifugation process, the white RNA
precipitate at the bottom of the tube was washed with 75% alcohol. The mRNA was
reverse transcribed using TB Green® Premix Ex Taq™
II (Cat RR820L, TakaraBio, Takara Bio Inc., Kyoto, Japan), while miRNA was
reverse transcribed using the miRcute miRNA First-strand cDNA Synthesis Kit
(KR211, Tiangen, Beijing, China). PCR conditions for mRNA amplification were as
follows: initial denaturation at 95 °C for 30 sec, followed by 40 cycles
of 95 °C for 45 sec and 60 °C for 34 sec. For miRNA
amplification, PCR conditions included an initial denaturation step at 95
°C for 15 min, followed by 40 cycles of 94 °C for 20 sec and 60
°C for 34 sec. Relative expression levels were calculated using the
2
| RNA | Primer sequence | |
| U6 | Forward: | 5 |
| Reverse: | 5 | |
| Forward: | 5 | |
| Reverse: | 5 | |
| CXCL10 | Forward: | 5 |
| Reverse: | 5 | |
| mmu-miR-423-5p | Forward: | 5 |
| Reverse: | 5 | |
| mmu-miR-423-5p mimic | Forward: | 5 |
| Reverse: | 5 | |
| mmu-miR-423-5p inhibitor | Single-stranded | 5 |
Proteins from cells and exosomes were extracted with protease and phosphatase inhibitors (Servicebio, Guangzhou, China). The protein solution was mixed with loading buffer and heated to 100 °C for 20 min, separated by sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE), and then transferred onto polyvinylidene difluoride membranes (Millipore, Burlington, MA, USA). After blocking, membranes were incubated overnight at 4 °C with primary antibodies against Glyceraldehyde 3-phosphate dehydrogenase (GAPDH, 1:10,000 dilution; cat. AC002; ABclonal, Waltham, MA, USA), CD63 (1:1000 dilution; cat. no. 10112; CST, Danvers, MA, USA), CD81 (1:1000 dilution; cat. 10037; CST, Danvers, MA, USA), vascular endothelial growth factor (VEGF, 1:1000 dilution; cat. no. A12303; ABclonal, Waltham, MA, USA), or C-X-C motif chemokine ligand 10 (CXCL10, 1:500 dilution; cat. 10937-1-AP; Proteintech, Chicago, IL, USA). Subsequently, the membranes were washed and incubated with secondary antibody (Servicebio, Guangzhou, China) for 1 h. Protein bands were visualized using an electrochemiluminescence (ECL) detection kit (Beyotime, Nanjing, China). Image analysis was performed using ImageJ software version 1.51 for Mac to quantify the gray values of protein blots, with GAPDH serving as the internal reference. GraphPad Prism 9.0 (https://www.graphpad.com/updates/prism-900-release-notes) was used to normalize the relative intensity of target protein bands in the experimental groups against the average intensity observed in the control group.
To identify the predicted direct binding site, a plasmid containing either wild-type CXCL10 3’UTR or a mutant form was co-transfected into HEK293T cells, along with either a miR-423-5p mimic or a scrambled negative control (NC). Luciferase activity in the transfected cells was subsequently quantified using a Dual Luciferase Reporter Assay System (E1910, Promega, Madison, WI, USA). The complete binding site is shown in the Supplementary Material.
Animal experiments were authorized by the Committee for the Ethics of Animal Experiments, East China Normal University (Approval No. m20210910). Experimental animals were obtained from the Experimental Animal Center, Minhang Campus, East China Normal University. MOBs were transfected with miR-423-5p mimics, inhibitors, or negative control (NC), and the exosomes subsequently isolated. Matrigel was then mixed with ECs that had been pretreated with exosomes derived from the different experimental groups. Each experimental group consisted of 5 female nude mice (BALB/c strain, age 7–8 weeks). These were injected with the Matrigel mixture into the dorsal region after anesthetization with 1% pentobarbital sodium (200 mg/kg). After two weeks, mice were euthanized by placing them in a euthanasia box with carbon dioxide until the cessation of vital signs, as observed by the absence of toe reflex and respiratory and cardiac activity. The Matrigel plugs were collected and analyzed by immunofluorescence and immunohistochemistry staining.
RNA-seq analysis was conducted by Lianchuan Biotechnology (Hangzhou, China).
Briefly, RNA extracted from bEnd.3 cells underwent quality assessment using a
Bioanalyzer 2100 instrument (Agilent Technologies, Santa Clara, CA, USA). Only
those samples with a RIN value
Statistical analyses were conducted using GraphPad Prism 9.0
(https://www.graphpad.com/updates/prism-900-release-notes). Student’s two-tailed
t-test was performed to evaluate the difference between two independent
samples, while one-way ANOVA was used to evaluate differences between multiple
groups. Two-way ANOVA was used to analyze two independent variables. All data are
presented as the mean
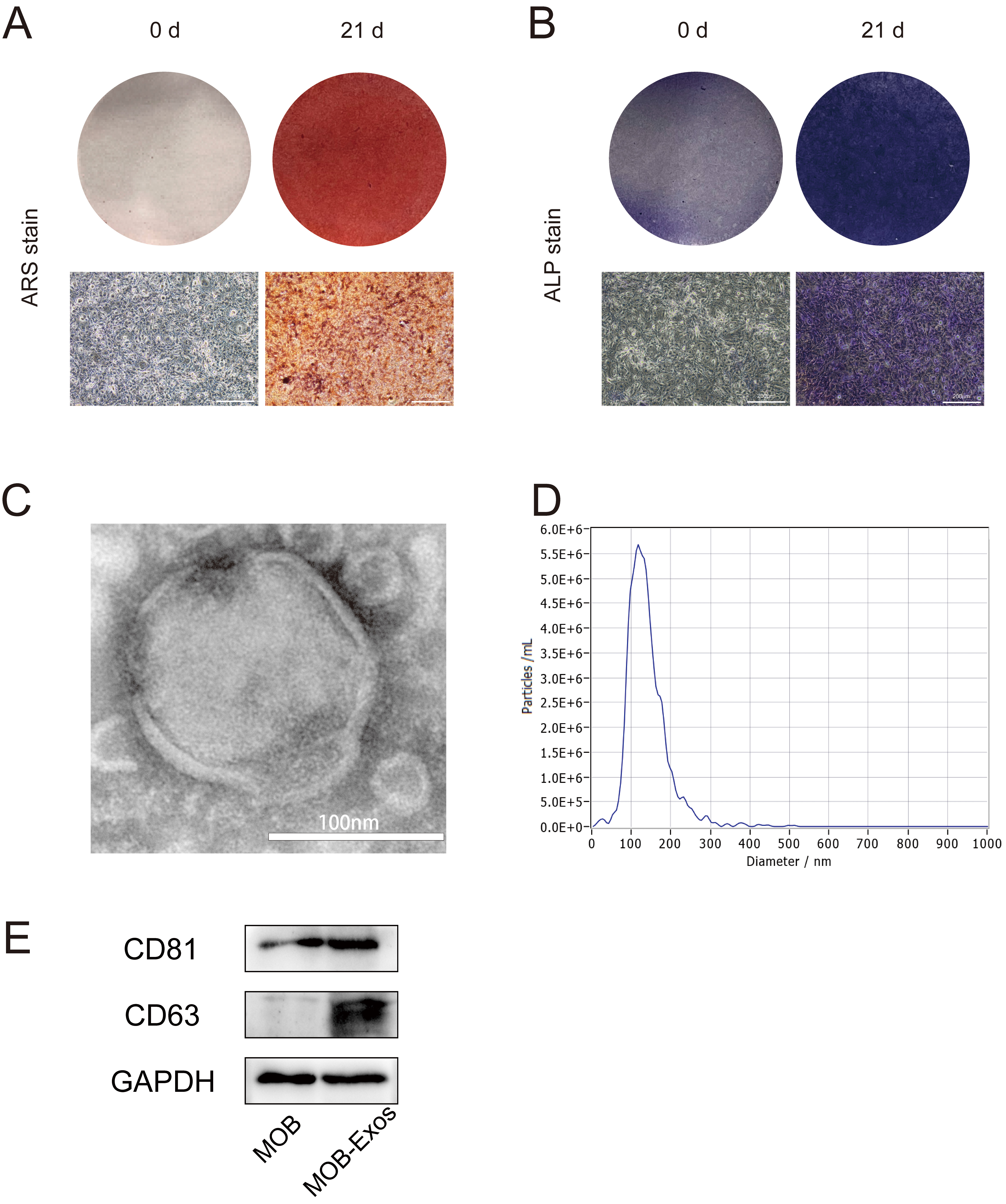
Exosomes were extracted from the MC3T3-E1 cell line following osteogenic induction for 21 d. The presence of mature MOBs was confirmed by Alizarin Red S staining (Fig. 1A) and ALP staining (Fig. 1B). Transmission electron microscopy revealed the characteristic hollow spherical morphology typical of exosomes (Fig. 1C), consistent with previous reports. Nanoparticle tracking analysis further characterized MOB-Exos and revealed a particle diameter ranging from 88.6 to 195.0 nm, with an average of 139.4 nm and a peak at 120.6 nm (Fig. 1D). Western blotting confirmed the presence of exosome-specific markers CD63 and CD81 (Fig. 1E), thus verifying the successful isolation of exosomes from MOBs.
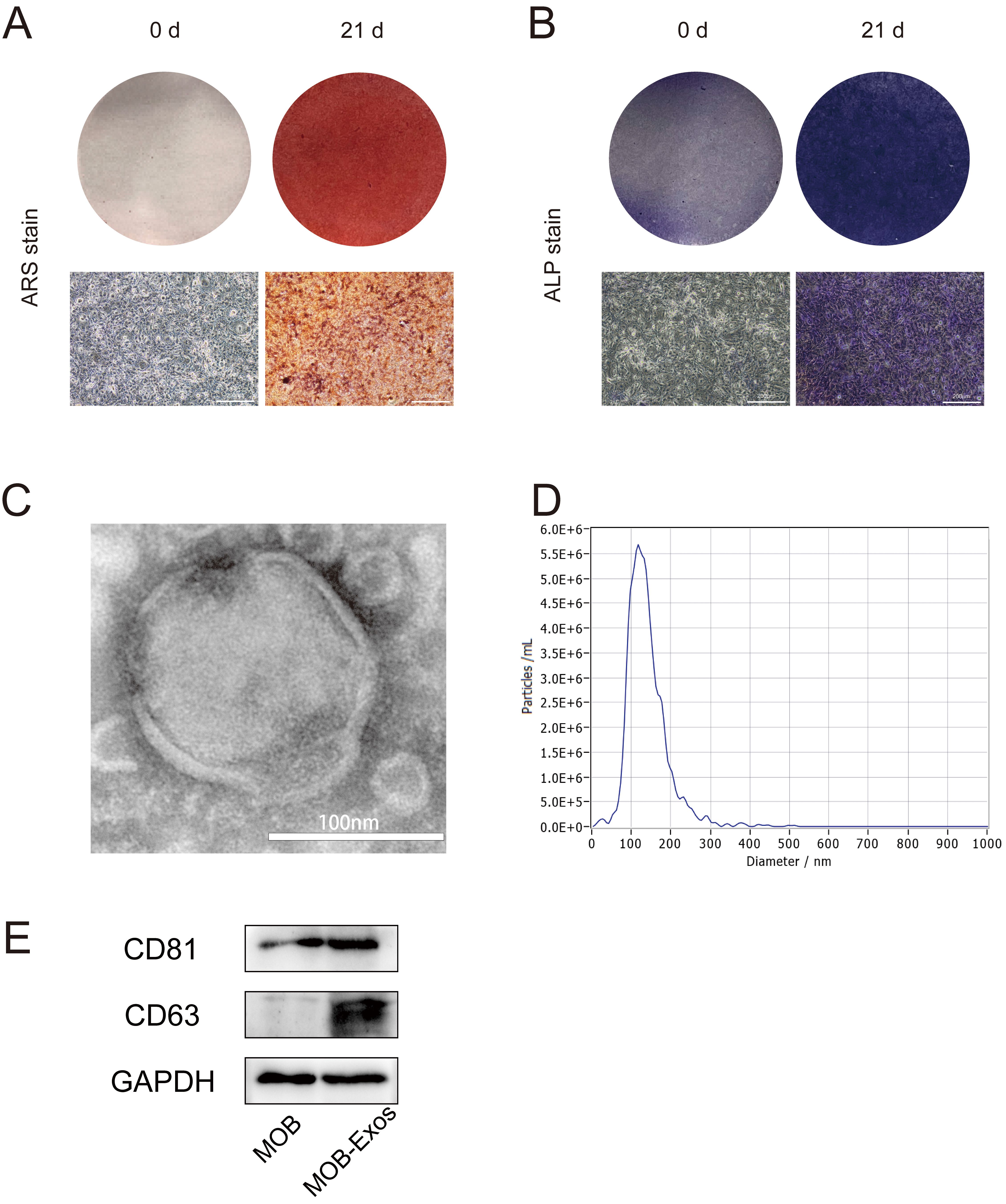 Fig. 1.
Fig. 1.
Identification of exosomes derived from MOBs. (A) ARS staining, and (B) ALP staining showed successful osteogenic induction of MC3T3 cells (scale bar: 200 µm) (C) Transmission electron microscopy images showing exosomes (scale bar: 100 nm). (D) Nanoparticle tracking analysis showing exosome size distribution. (E) Western blot analysis showing the expression of exosomal makers CD63 and CD81 in MOB-Exos. MOBs, Mineralized osteoblasts; ARS, Alizarin Red S; ALP, Alkaline Phosphatase; MOB-Exos, exosomes derived from MOBs.
MOB-Exos were labeled with PKH67 and cultured with ECs to study their
internalization. As shown in Fig. 2A, the strong green fluorescent signal
surrounding the nucleus indicated internalization of PKH67-labeled MOB-Exos. A
series of functional assays was performed to evaluate the effect of MOB-Exos on
vascularization. Results from the CCK-8 assay demonstrated that MOB-Exos
significantly promoted EC proliferation compared to the control group treated
with an equal volume of 1
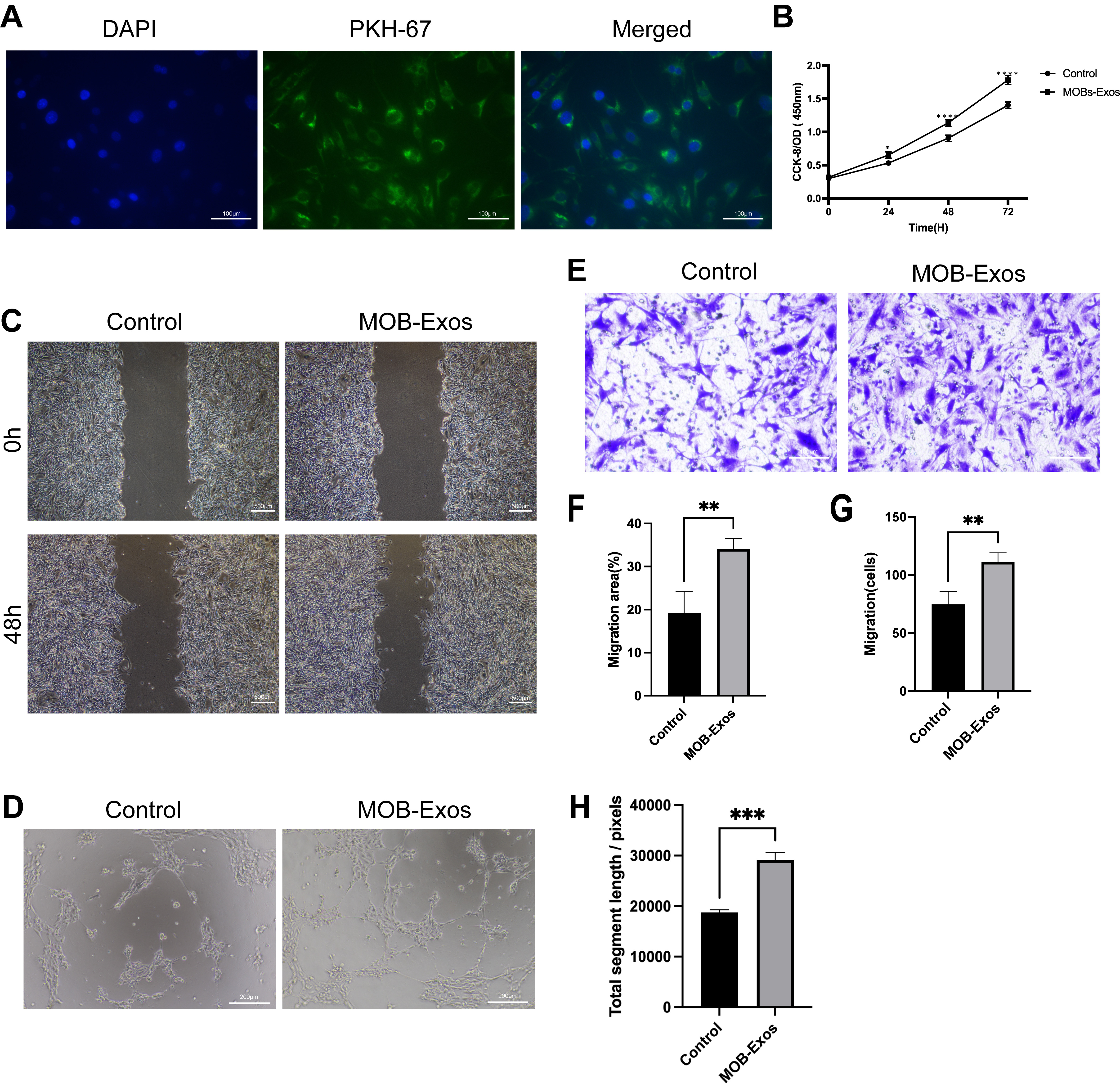 Fig. 2.
Fig. 2.
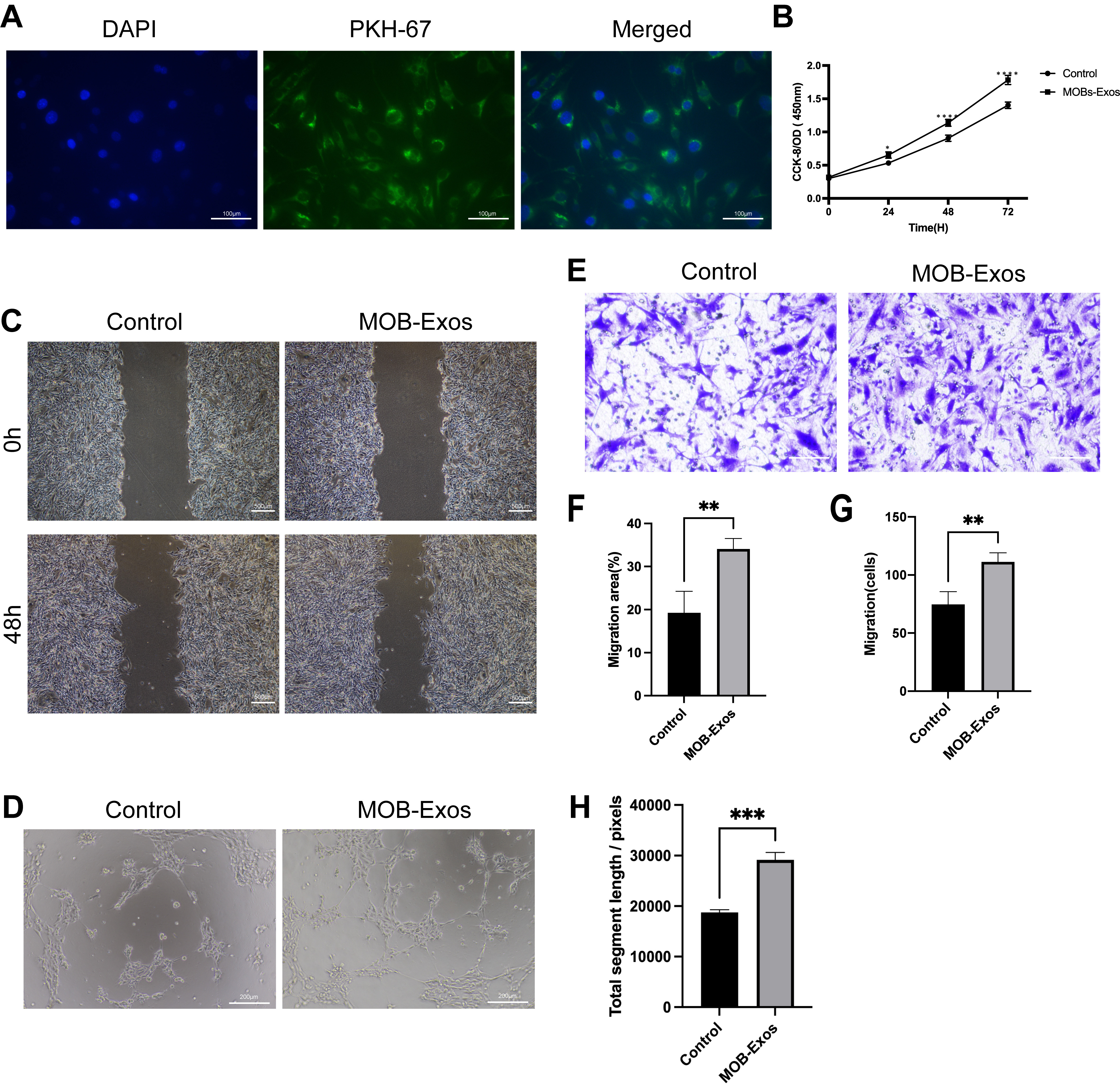
MOB-Exos promoted endothelial cell (EC) proliferation,
migration, and tube formation. (A) Internalization of PKH67-labelled MOB-Exos by
ECs (scale bar: 100 µm). (B) The proliferation rate of ECs as
determined by the Cell Counting Kit-8 (CCK-8) assay. (C) Scratch wound (scale
bar: 500 µm) and (E) Transwell assays (scale bar: 100
µm) were performed to evaluate the migration of ECs. (D) Tube
structure formation assay performed on ECs (scale bar: 200 µm).
Results are presented as the mean
Exosomes are extracellular vesicles containing bioactive substances that
participate in the process of cell-cell communication. miR-423-5p was highly
expressed in MOB-Exos (p
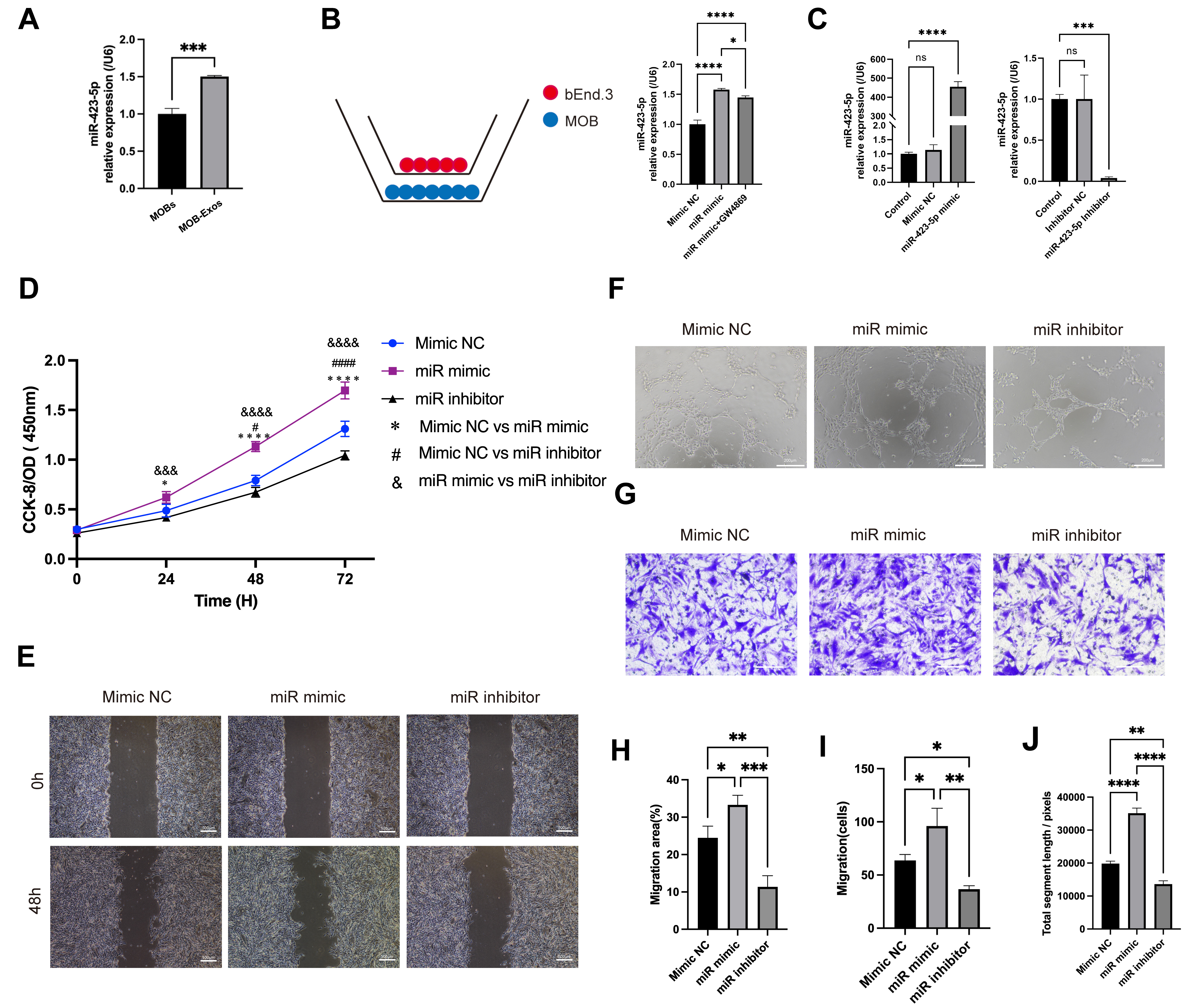 Fig. 3.
Fig. 3.
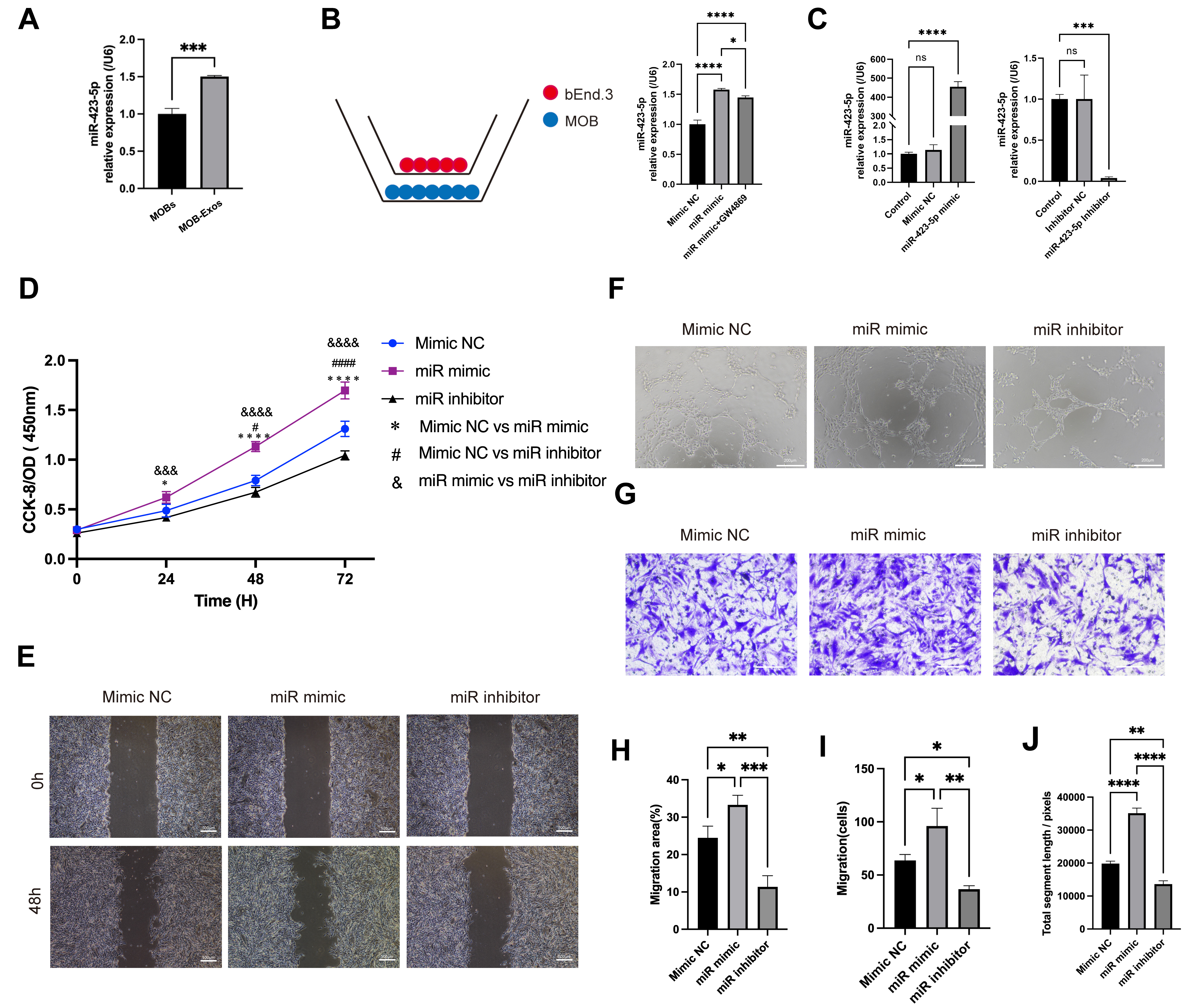
Exosomal miR-423-5p promoted the angiogenic ability of
ECs. (A) Quantitative polymerase chain reaction (qPCR) results showing an
increased level of miR-423-5p in MOB-Exos. (B) ECs were co-cultured with MOBs and
the miR-423-5p content of the ECs was subsequently evaluated in different
experimental groups. (C) Successful establishment of the mimic and inhibitor
systems. (D) CCK-8, (E) scratch wound (scale bar: 500 µm), (F) tube
formation (scale bar: 200 µm), and (G) Transwell migration assays
(scale bar: 100 µm) were performed to assess the angiogenic ability
of ECs. Results are presented as the mean
The specific impact of MOB-Exos-derived miR-423-5p on EC angiogenesis was further investigated to exclude the possible impact of other exosome components. ECs were transfected with miR-423-5p mimic, negative control (NC), inhibitor, or inhibitor NC. Subsequent qPCR confirmed the successful modulation of miR-423-5p levels in each group (Fig. 3C). Results from the CCK-8 assay revealed that overexpression of miR-423-5p significantly increased EC proliferation, whereas inhibition of miR-423-5p attenuated this effect (Fig. 3D). Scratch wound and Transwell migration assays further demonstrated that ECs transfected with miR-423-5p mimic showed increased migration, whereas ECs transfected with miR-423-5p inhibitor showed reduced migratory ability (Fig. 3E,G,H,I). Additionally, ECs that overexpressed miR-423-5p displayed increased total segment lengths of tube-like structures in angiogenesis assays, whereas inhibition of miR-423-5p reversed this effect (Fig. 3F,J). These results establish conclusively that miR-423-5p derived from MOB-Exos plays a pivotal role in promoting the angiogenic potential of ECs.
We next conducted RNA sequencing of the miR-423-5p overexpression group and the
control group to investigate changes in the EC transcriptome following
transfection with miR-423-5p mimic. The level of miR-423-5p was also assessed to
confirm the successful transfection of miRNA in ECs (Fig. 4F). In ECs that
overexpressed miR-423-5p, 108 genes were found to be upregulated (FC
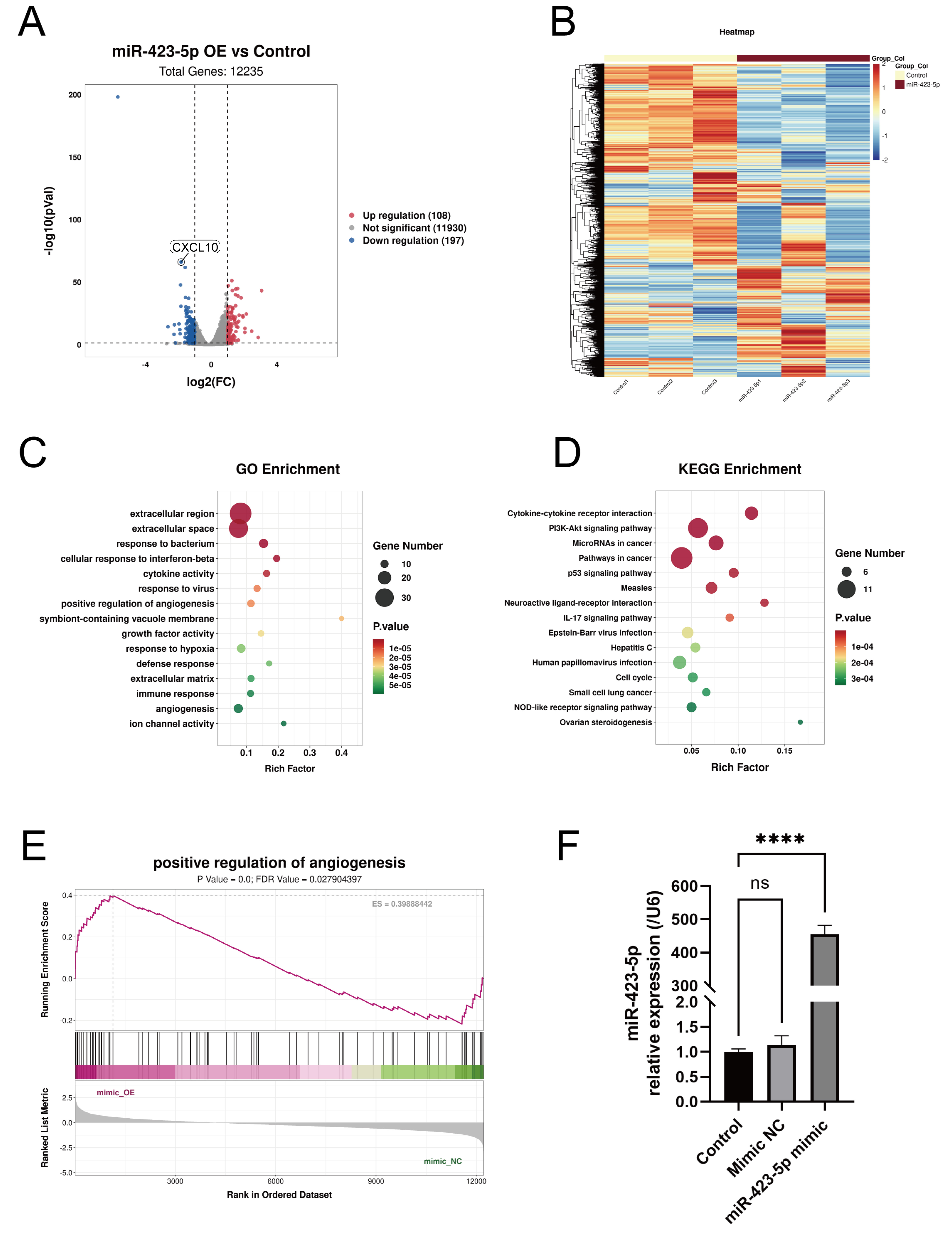 Fig. 4.
Fig. 4.
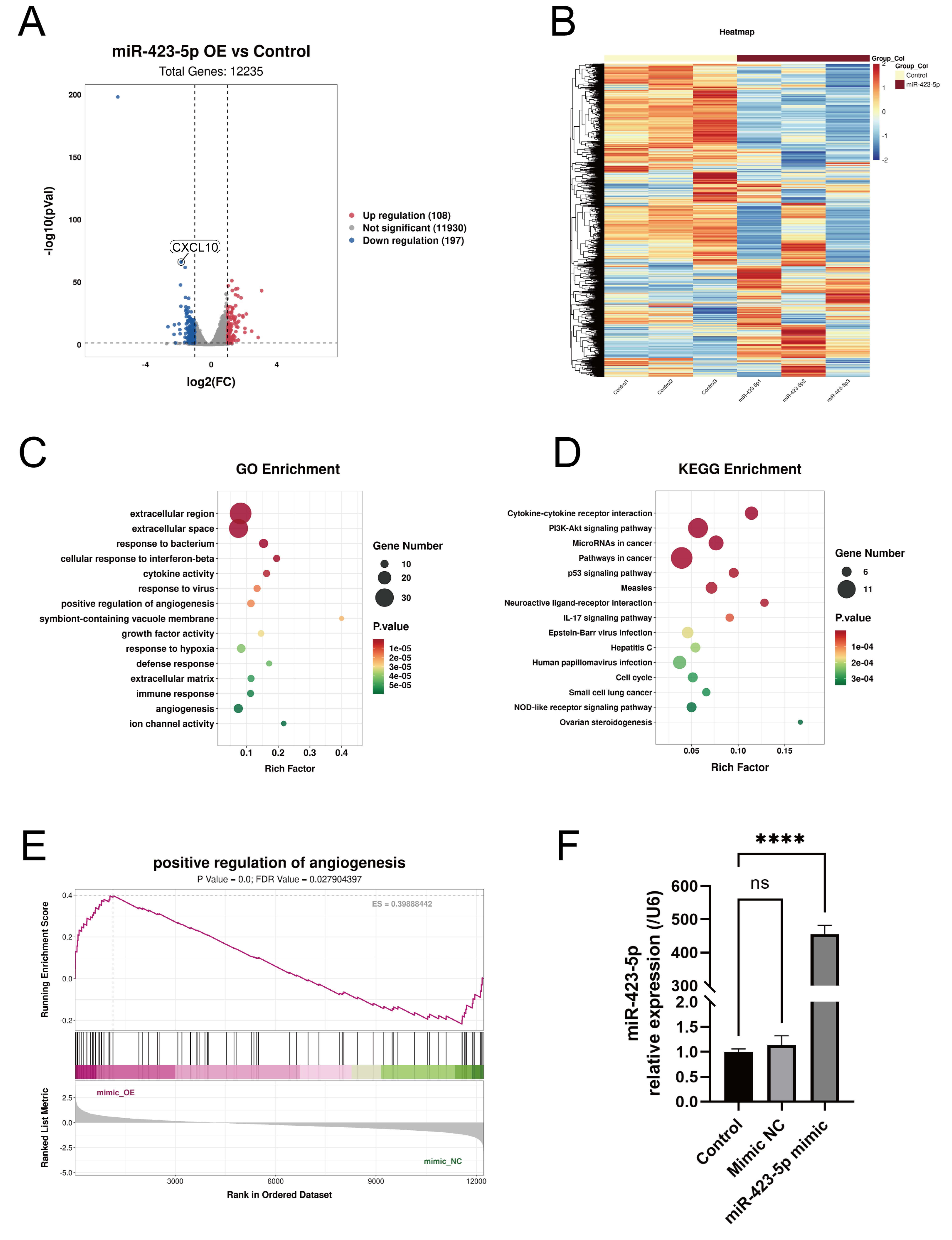
RNA sequencing of ECs with miR-423-5p overexpression compared to
the mimic NC group (n = 3). (A) Volcano plot and (B) heatmap showing the
differentially expressed genes. These were evaluated by (C) GO and (D) KEGG
enrichment analyses. (E) GSEA enrichment analyses showed that miR-423-5p was a
positive regulator of angiogenesis. (F) qPCR results showing the miR-423-5p level
in the three experimental groups. Results are presented as the mean
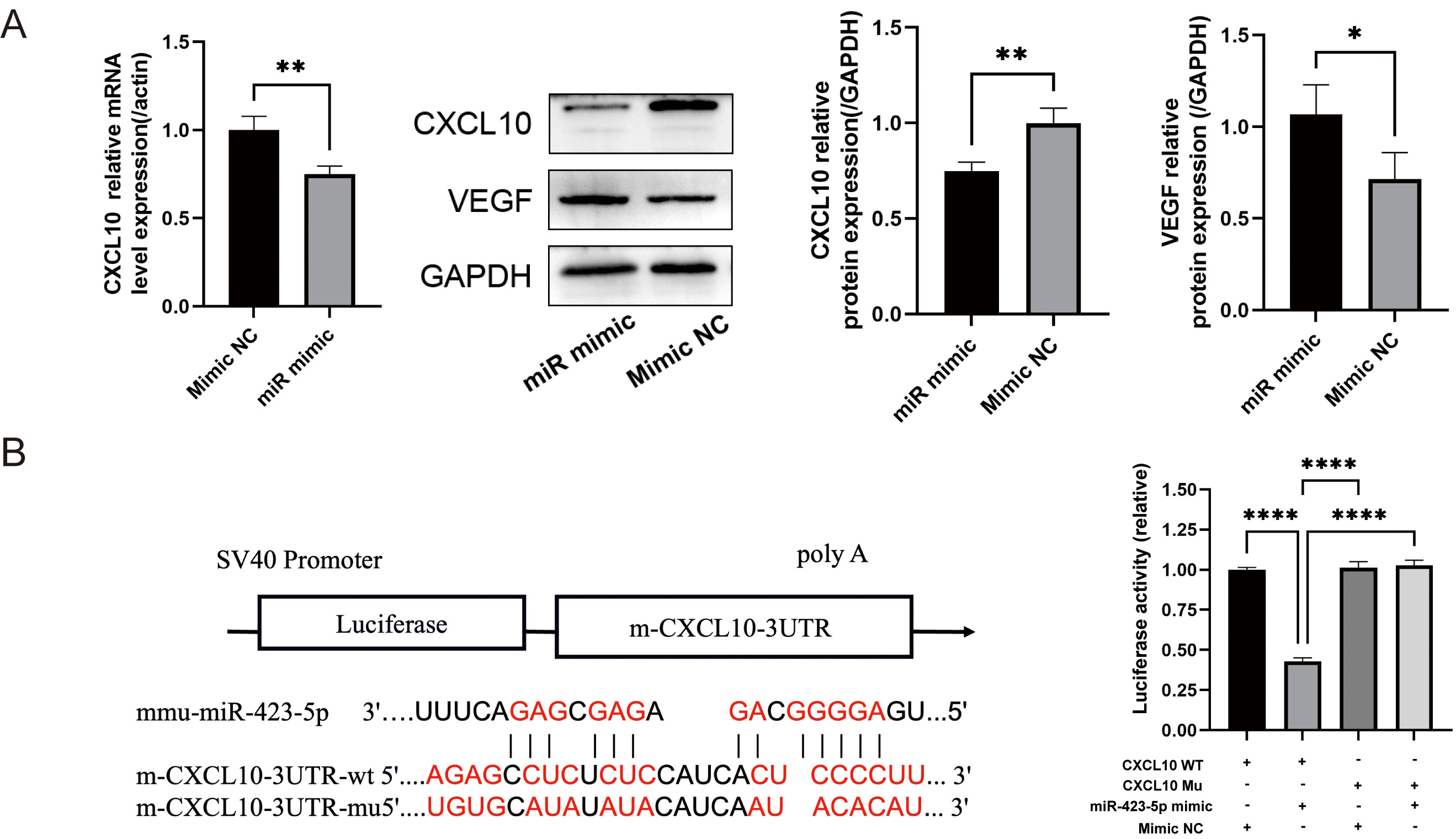
Western blot and qPCR analyses showed significant reductions in the mRNA and
protein levels of CXCL10 following transfection with miR-423-5p mimic, whereas
the VEGF protein level increased (Fig. 5A). A luciferase reporter assay was
performed to determine whether CXCL10 is a direct target of miR-423-5p.
The results showed a marked decrease in firefly luciferase activity when
miR-423-5p was co-transfected with wild-type CXCL10 3
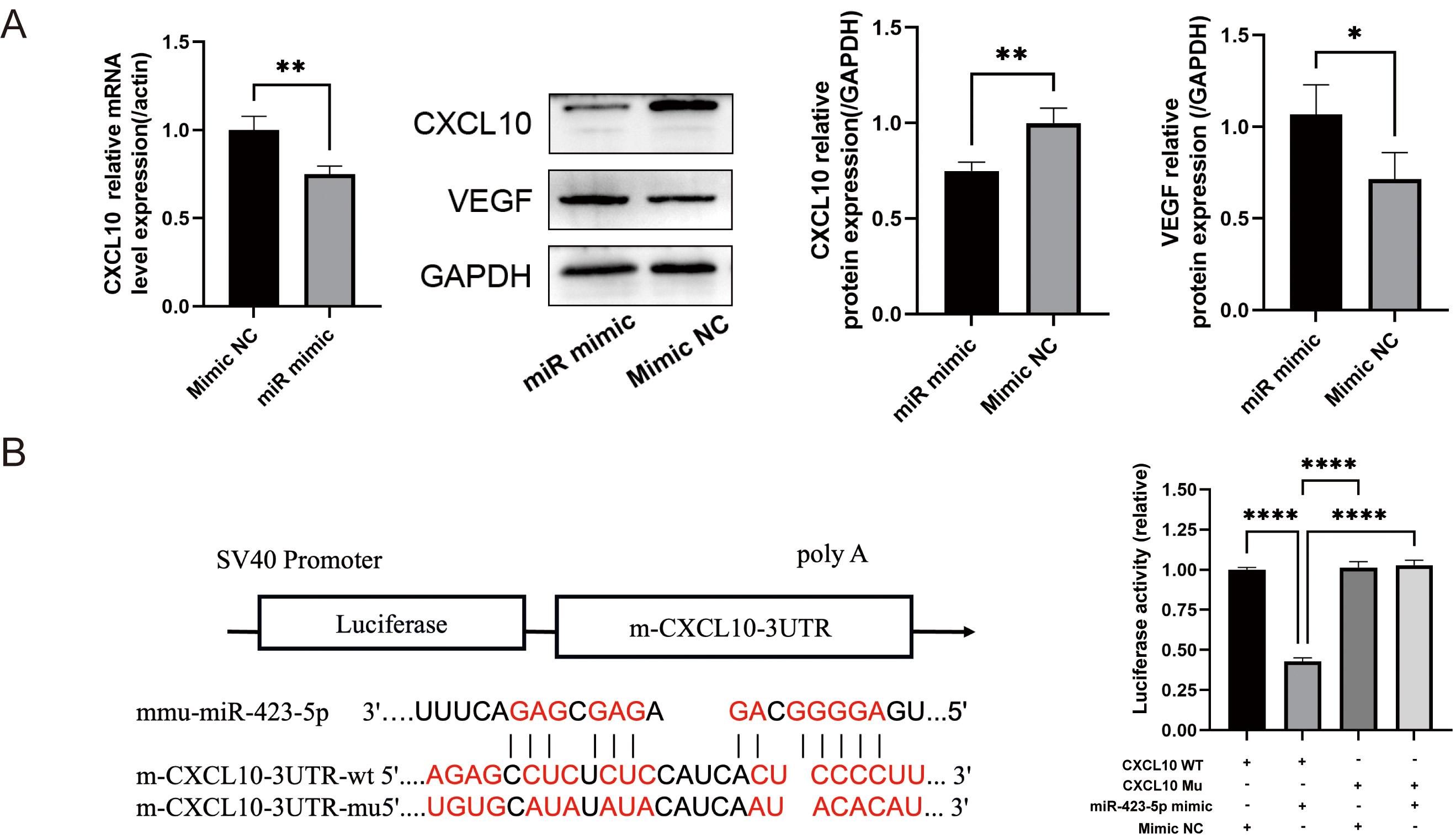 Fig. 5.
Fig. 5.
miR-423-5p directly targets CXCL10. (A) qPCR was used
to quantify CXCL10 mRNA, and Western blot for the analysis of C-X-C motif chemokine
ligand 10 (CXCL10) and vascular endothelial growth factor (VEGF)
proteins. (B) One of the predicted binding sequences of miR-423-5p to
CXCL10, together with results from a dual luciferase reporter assay
showing luciferase activity in the various experimental groups. Results are presented as the mean
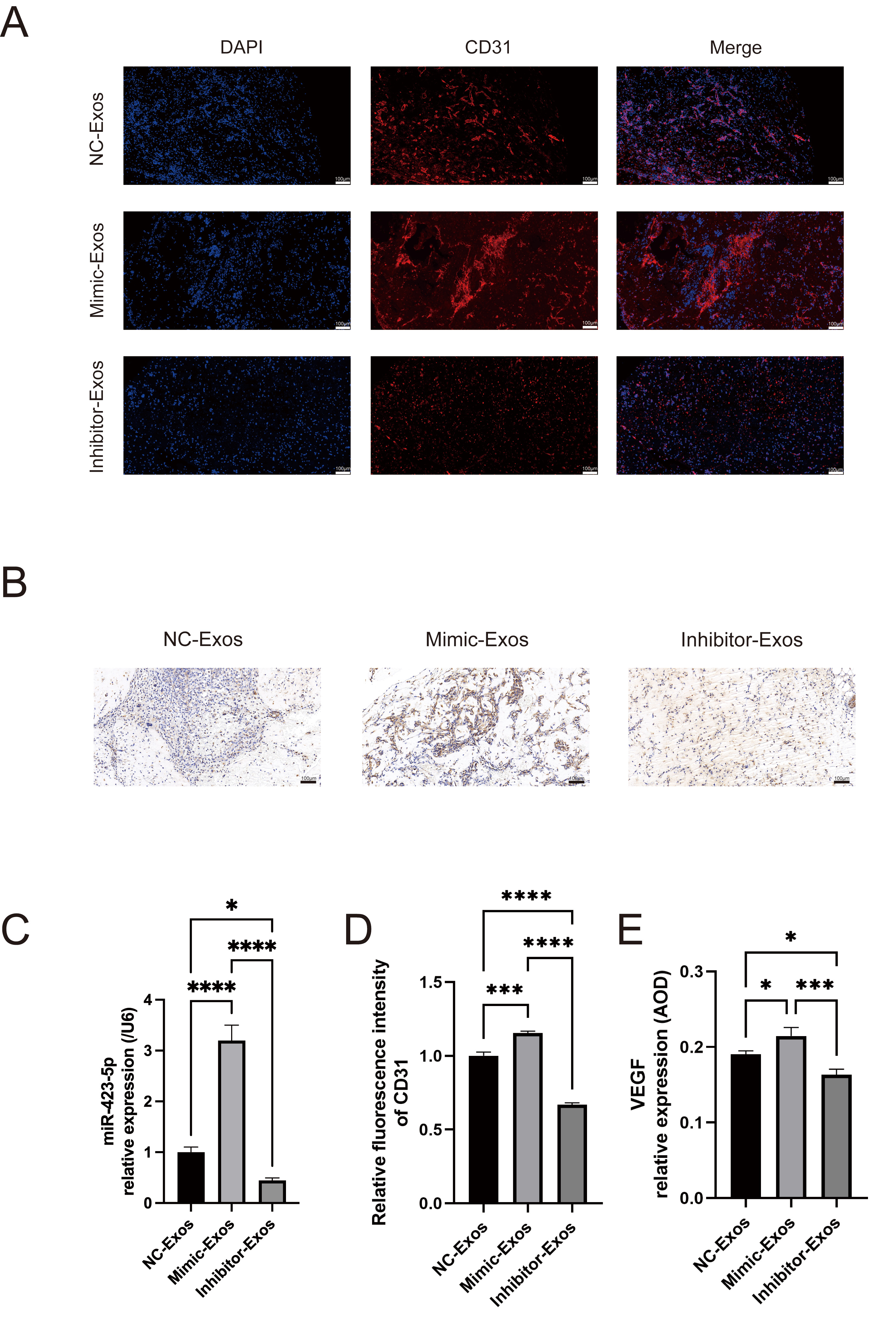
Matrigel plug assays were used to investigate the in vivo angiogenic role of MOB-derived exosomal miR-423-5p. MOBs were transfected with miR-423-5p mimics, inhibitors, or NC, and the exosomes subsequently isolated. Exosomes from the different groups were first cultured with ECs. Matrigel was then mixed with the ECs and injected into the dorsal region of nude mice. qPCR was subsequently performed to assess the level of exosomal miR-423-5p in the different experimental groups (Fig. 6C). Additionally, immunofluorescence and immunohistochemical staining were performed to evaluate the expression levels of CD31 and VEGF. The miR-423-5p mimic group showed significantly higher levels of CD31 and VEGF compared to the NC group. Interestingly, these effects were reversed after inhibition of miR-423-5p using a specific inhibitor (Fig. 6A,B,D,E). Taken together, these results demonstrate that exosomal miR-423-5p derived from MOBs plays a crucial role in promoting EC vascularization in vivo.
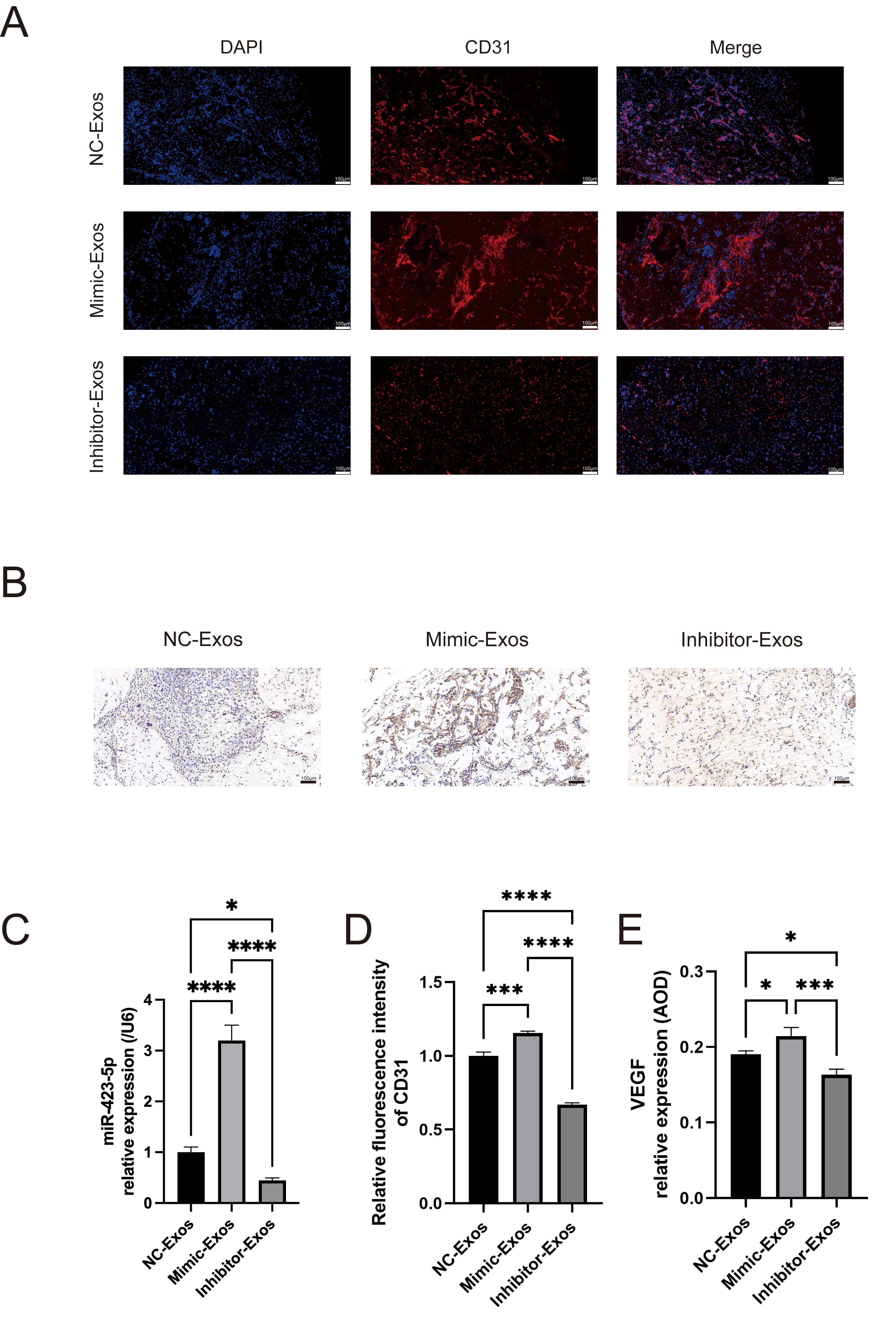 Fig. 6.
Fig. 6.
MOB-derived exosomal miR-423-5p promoted angiogenesis
in vivo. (A,D) Immunofluorescence of CD31 expression in Matrigel plugs.
(B,E) Immunohistochemical detection of VEGF. (C) The level of miR-423-5p in
NC-Exos, Mimic-Exos, and inhibitor-Exos was evaluated by qPCR. Scale bar: 100
µm. Results are presented as the mean
Bone tissue is a highly vascularized structure in which ECs provide the necessary oxygen and nutrients. Previous research has highlighted the angiogenic potential of exosomes derived from adipose stem cells [23, 25] and umbilical cord MSCs [26]. However, these studies focused mostly on the effect of exosomes in angiogenesis, while ignoring the bone tissue microenvironment. In tissue-engineered bone, ECs and OBs are closely related in terms of spatial location and physiological function. The identification of specific active substances affecting these cells may provide new therapeutic approaches for bone tissue engineering. Exosomes are important carriers for intercellular signaling and can deliver functional molecules required for biological processes. By isolating MOB-Exos and investigating their role in angiogenesis, this study also demonstrated a positive effect of MOB-Exos on vascularization. A high level of miR-423-5p was found in MOB-Exos, and co-culture experiments demonstrated the exosome-mediated transport of miR-423-5p into ECs. The angiogenic potential of miR-423-5p from MOB-Exos was further demonstrated by manipulating its expression in ECs using mimics, inhibitors, and negative controls. Animal experiments confirmed that exosomal miR-423-5p enhanced the in vivo angiogenic ability of ECs. Overall, the present study indicates that exosomal miR-423-5p derived from MOBs significantly promotes EC angiogenesis, thus highlighting its potential as a therapeutic for enhancing vascularization in bone tissue engineering applications.
Osteogenesis and angiogenesis are pivotal processes in tissue engineering, with signaling factors being important in this process. VEGF-A is a chemotactic molecule that induces EC migration and promotes the differentiation of OBs [27]. Basic fibroblast growth factor (bFGF) is another critical angiogenesis factor that facilitates the mobilization and recruitment of ECs, thereby promoting the formation of capillaries crucial for bone regeneration [28]. Platelet-derived growth factor from pre-osteoclasts promotes osteogenesis and angiogenesis through bone modeling [29]. Biomimetic nanocomposite scaffolds combining VEGF/bFGF and bone morphogenetic protein 2 (BMP2) have been shown to promote both angiogenesis and bone formation [30]. Additionally, bionic periosteum incorporating gelatine diselenide and calcium alginate activates the NO-cGMP signaling pathway through sustained release of nitric oxide (NO), thereby enhancing osteogenesis and angiogenesis [31]. These studies highlight the significance of signaling factors as innovative strategies for the advancement of bone tissue engineering.
Exosomes are recognized as crucial extracellular vesicles containing diverse signaling factors that play specialized roles in intercellular communication. Exosomes derived from various cell types exert significant influences on angiogenesis in ECs, and osteogenesis in OBs [32]. Indeed, exosomes from MSCs are known to enhance angiogenesis, thereby improving bone healing [33]. Similarly, exosomes from adipose-derived stem cells promote EC mobility and angiogenesis by modulating the expression of transient receptor potential cation channels [34]. Moreover, MOB-Exos have been reported to induce the differentiation of bone marrow stromal cells into OBs [35]. Exosomes derived from ECs can accelerate bone formation by stimulating angiogenesis [36]. However, the specific mechanisms by which MOB-Exos regulate ECs remain unclear. In the current study, MOB-derived exosomes were extracted, characterized, and shown to positively regulate angiogenesis by ECs.
To further investigate the specific components in MOB-Exos that affect ECs, this study focused on miRNAs. These non-coding RNA molecules bind to the 3’-untranslated region (3’-UTR) of target genes, thereby modulating gene transcription and regulating various physiological processes [37, 38]. miRNA has unique advantages in cell communication because of its small molecular weight and stable properties. Previous reports have linked exosome-derived miR-423-5p to biological processes such as tumorigenesis and metastasis [39, 40]. Building upon this prior knowledge, our study manipulated the miR-423-5p level by overexpressing or inhibiting miR-423-5p. Exosomal miR-423-5p from MOBs was found to significantly enhance the angiogenic capabilities of ECs, both in vitro and in vivo. These findings highlight the potential therapeutic use of exosome-delivered miRNAs in bone tissue engineering.
CXCL10 is a well-known anti-angiogenic chemokine that plays an important role in inhibiting angiogenesis [41, 42, 43]. Previous studies have shown that CXCL10 inhibits angiogenesis by ECs. Infection of ECs with the Rubella virus was shown to reduce angiogenesis via interferon-induced CXCL10 [42]. Furthermore, exosomal miR-21-5p enhanced EC vascularization by inhibiting CXCL10 [44]. In the present investigation, CXCL10 was identified as a novel downstream target gene of miR-423-5p. This miRNA interacts directly with the CXCL10 gene, resulting in the down-regulation of both CXCL10 mRNA and protein. Consistent with previous findings, our results showed that down-regulation of CXCL10 could increase angiogenesis in ECs. These results highlight the role of miR-423-5p in regulating CXCL10 and suggest that it has potential therapeutic utility in promoting vascularization. The latter is critical for the progress of tissue engineering and regenerative medicine.
Due to the complexity of their contents, many of the mechanisms involving exosomes in bone tissue repair are still unknown. The pro-angiogenic effects of specific active molecules other than miR-423-5p in exosomes require further investigation.
MOB-Exos have demonstrated the ability to enhance angiogenesis in ECs. This activity is potentially mediated by the transfer of miR-423-5p. It has been established that miR-423-5p directly targets CXCL10, leading to its down-regulation in ECs. These findings provide strong evidence for the potential therapeutic use of exosomes in bone tissue engineering applications.
The RNA-seq data in this research have been uploaded to the GEO database (https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE262133. The datasets used in this study are available from the corresponding author upon reasonable request.
FL and YH designed the experiment research. FL and HL performed the experiments. FL performed formal analyses and wrote the original manuscript. XC and YD contributed to the conception and design of the work. All authors have revised the manuscript and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Animal experimental was authorized by the Committee on the Ethics of Animal Experiments of East China Normal University (No.m20210910). All experiments were carried out in strict accordance with the Declaration of Helsinki.
We would like to express our gratitude to East China Normal University for their support in conducting animal experiments.
This work was funded by the Minhang District Natural Science Foundation (2023MHZ048).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.