1 Biological Sciences, Brock University, St. Catharines, ON L2S 3A1, Canada
2 Internal Medicine Department, St. Francis Medical Center, Monroe, LA 71201, USA
Abstract
Drosophila melanogaster has been used as a model system to identify and characterize genetic contributions to development, homeostasis, and to investigate the molecular determinants of numerous human diseases. While there exist many differences at the genetic, structural, and molecular level, many signalling components and cellular machineries are conserved between Drosophila and humans. For this reason, Drosophila can and has been used extensively to model, and study human pathologies. The extensive genetic resources available make this model system a powerful one. Over the years, the sophisticated and rapidly expanding Drosophila genetic toolkit has provided valuable novel insights into the contribution of genetic components to human diseases. The activity of Notch signalling is crucial during development and conserved across the Metazoa and has been associated with many human diseases. Here we highlight examples of mechanisms involving Notch signalling that have been elucidated from modelling human diseases in Drosophila melanogaster that include neurodegenerative diseases, congenital diseases, several cancers, and cardiac disorders.
Keywords
- Drosophila melanogaster
- human disease modelling
- neurodegenerative diseases
- cancer
- cardiomyopathy
Drosophila melanogaster is an excellent model system to study molecular mechanisms and cellular pathways [1]. The common fruit fly has contributed an immense amount of knowledge to the fields of molecular biology and genetics [2]. Owing to the high degree of conservation at the genomic level between Drosophila and vertebrates, this model system has helped researchers discover novel genes and elucidate their contribution to organismal development, cell communication, and tissue homeostasis [1, 2, 3]. Most of the major signalling pathways involved in cell survival, division, apoptosis, and differentiation are highly conserved between mammals and Drosophila [2]. The genome of Drosophila is fully sequenced and extensively annotated. The Drosophila genome, which contains ~143.9 million base pairs, is much smaller than the human genome, which contains 3 billion base pairs [4]. The Drosophila genome consists of 4 chromosomes; 3 autosomal and 1 set of sex chromosomes (X and Y) [5, 6]. This makes it relatively easy to manipulate the Drosophila genome, which directly led to the development of a large library of genetic tools and depositories of transgenic lines that allow researchers to study the contributions of different genes and their products to signalling pathways, developmental processes, and disease mechanisms. Over the years, many large genetic screens have been performed in Drosophila to characterize individual genes, their products, and their interactions [1, 2, 7, 8, 9, 10, 11]. Another advantage of this model is the ease of tracking phenotypic changes associated with disruptions in different cellular pathways, which have been well catalogued due to the massive genetic screens conducted to date [2, 7, 8, 9, 10, 11]. One of the first examples of these large-scale screens was one that identified the role of patterning genes necessary for embryogenesis by conducting large-scale mutagenesis with simultaneous tracking of phenotypic changes in the fly embryo [7, 8, 9, 10, 11]. In addition, Drosophila melanogaster has been used to conduct large-scale drug screens to identify potential therapeutics [12, 13, 14, 15, 16, 17]. The Notch signalling pathway, first discovered in Drosophila, is involved in cell-to-cell communication, tissue homeostasis, and stem cell fate decisions during development, and in adult organisms [18, 19, 20, 21, 22]. This pathway and its major players have been associated with several human diseases including several cancers, neurodegenerative disorders, cardiac disorders, and congenital diseases [23, 24, 25, 26, 27]. In this review, we will highlight research that have leveraged the power of Drosophila melanogaster as a model system to uncover and characterize mechanisms that contribute to human diseases associated with defects in the Notch signalling pathway, which was not extensively reviewed previously.
The Notch receptor is produced in the endoplasmic reticulum (ER) and is cleaved
by a Furin (S1 cleavage) in the Golgi and post translationally modified by
several proteins. Including glycosyltransferases, to produce a heterodimeric
protein containing an extracellular ligand binding domain (NECD) and a
transmembrane intracellular domain (NICD) [3, 28, 29]. Upon activation by Notch
ligands located on neighbouring cells, the Notch receptor is cleaved in a
stepwise manner by two proteases including ADAM10 (human)/Kuzbanian
(Drosophila) (S2 cleavage) and the gamma secretase complex (
Ligands of the Notch pathway are also transmembrane proteins, which span the plasma membrane. In Drosophila there are 2 ligands: Delta and Serrate and one Notch receptor. Mammals have 4 Notch receptors (Notch1-4) and 5 ligands; 3 Delta-like (DLL1, DLL3, DLL4) and 2 Serrate-like (Jagged1 and Jagged2) (Fig. 1) [1, 3, 28, 29].
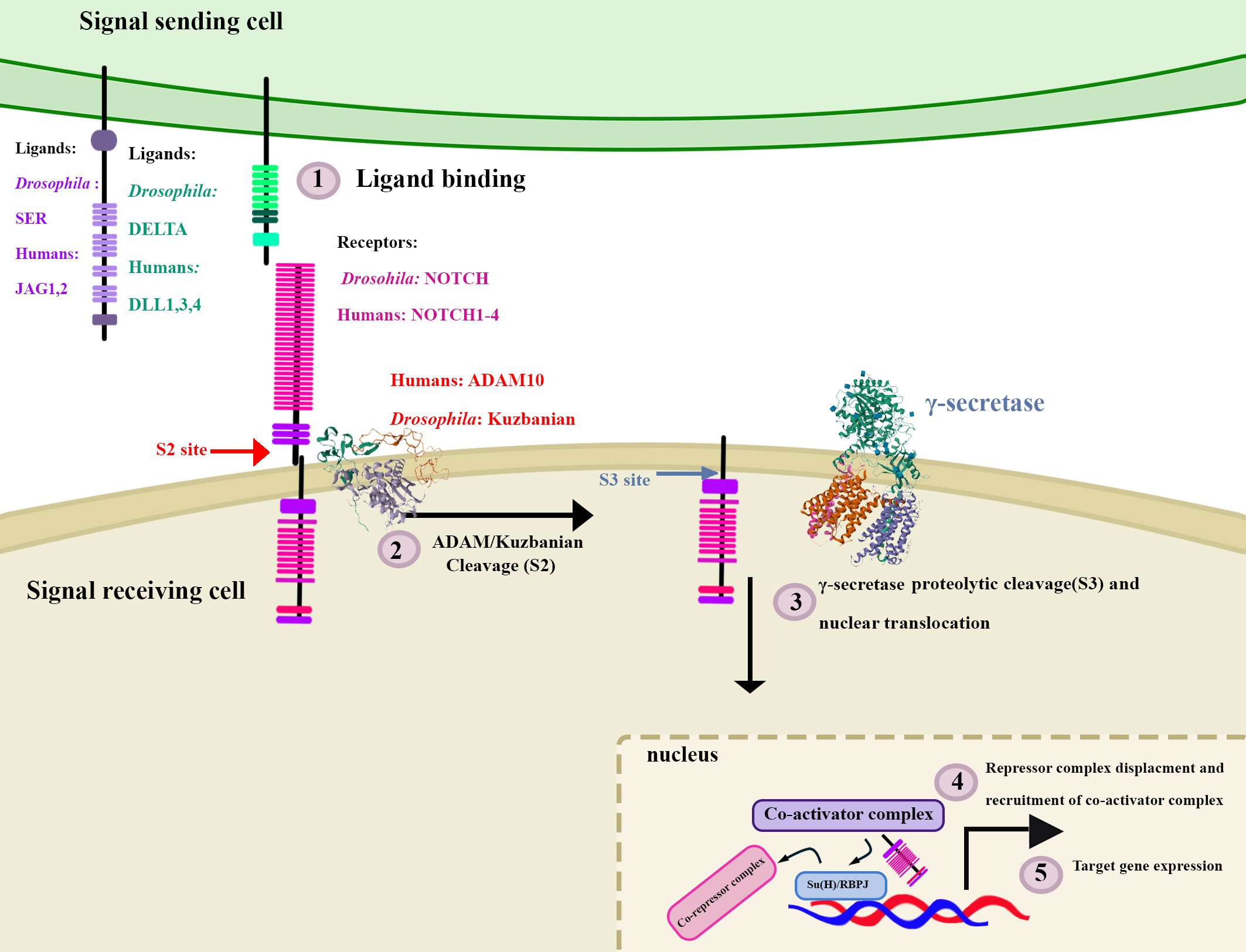 Fig. 1.
Fig. 1.Summary of Notch signalling in Drosophila and humans.
Notch signalling is a highly conserved signalling pathway. In Drosophila
there is one receptor protein, NOTCH, and two ligand protein, DELTA and SERRATE
(SER). In humans there are four receptor proteins; NOTCH1, NOTCH2, NOTCH3 and
NOTCH4 which are largely similar in structure to the Drosophila NOTCH
receptor. There are five ligands, two of which are more similar to SER, JAGGED1
(JAG1) and JAGGED2 (JAG2), while the other 3 are more similar in structure to
DELTA and are named Delta-like ligands 1, 3 and 4 (DLL1, DLL3, DLL4). In both
Drosophila and humans, Notch signalling is activated upon ligand
binding, which is followed by a series of proteolytic cleavage events at the S2
and S3 sites facilitated by ADAM10/Kuzbanian and
Notch signalling is terminated via phosphorylation of the PEST domain and concomitant poly-ubiquitination of the NICD, which results in its subsequent proteasomal degradation [3, 28, 29]. Notch activity can drive transcriptional programs that control differentiation, proliferation, or cell death in a context-dependent manner [1, 28].
As previously mentioned, the
While a direct link between Notch and memory impairment in AD has yet to be
established, there is evidence that Notch and APP compete for proteolytic
cleavage by PSEN1 in neurons, with APP metabolites down-regulating Notch receptor
and NICD target genes, and NICD down-regulating PSEN1, and hence,
reducing A
Owing to the essential role that Notch plays during the development of
Drosophila and the well-characterized phenotypes of Notch-loss-of
function mutants, Drosophila serves as an excellent model to conduct
drug- screens to develop a safe and effective AD therapeutic treatment that
decrease
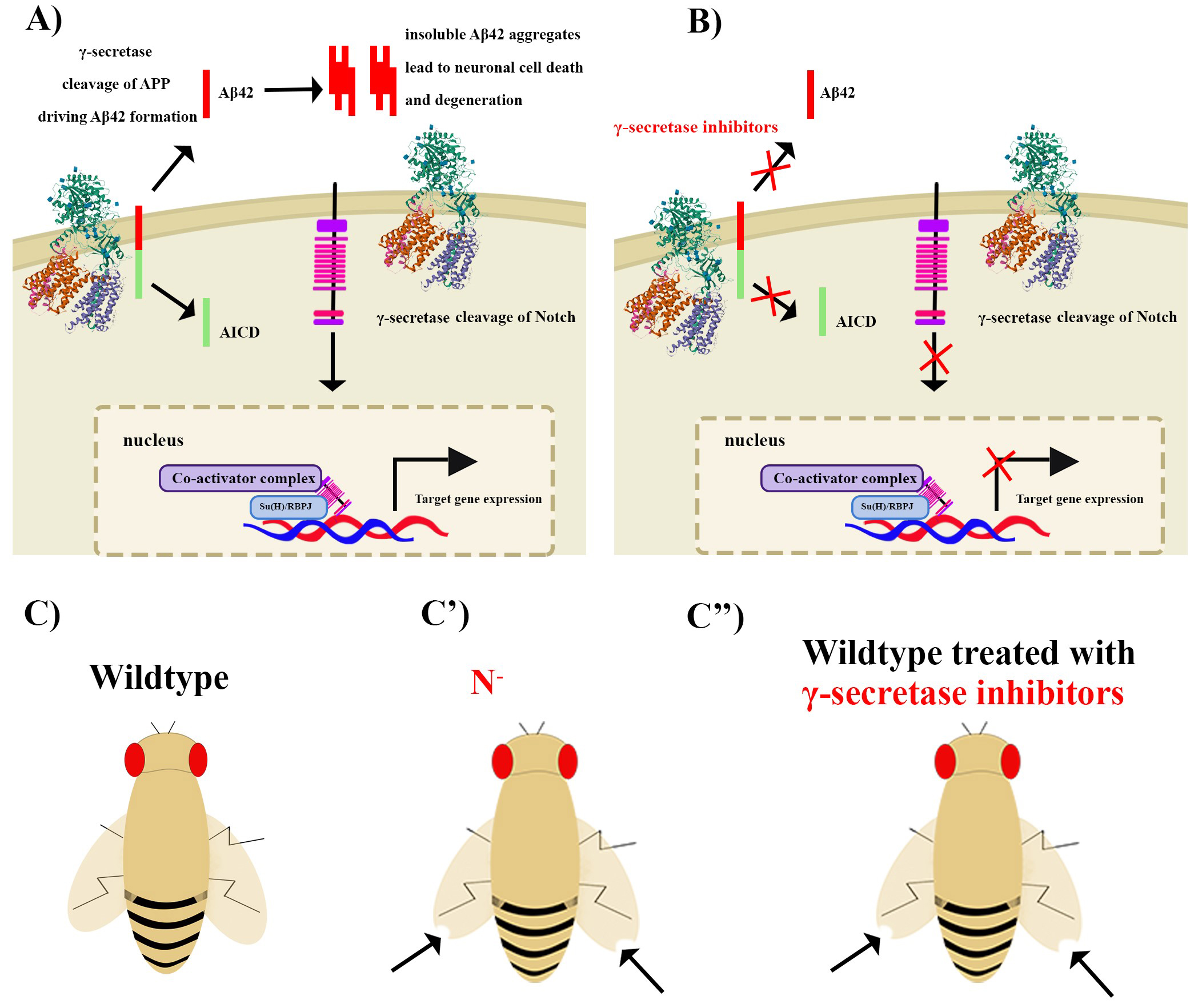 Fig. 2.
Fig. 2.The indirect link between Notch signalling and Alzheimer’s
diseas (AD) can be leveraged in Drosophila to conduct drug screens in
search of Notch sparing
Adams-Oliver syndrome (AOS) is a congenital disorder characterized by a range of anomalies, including congenital skin defects, particularly affecting the scalp (known as aplasia cutis congenita), as well as limb abnormalities often involving the loss of digits [81, 82]. Some individuals with AOS also exhibit nervous system and cardiac/vascular irregularities [81, 82]. Many genetic mutations have been identified in the NOTCH1 and DLL4 genes in AOS patients, implying that the condition’s dominant inheritance is linked to haploinsufficiency [27, 83, 84].
In 2012, Hassed et al. [82] reported the discovery of two autosomal
dominant variants within the RBPJ gene, found in separate families
(Family 1 and Family 2), with direct associations to Adams-Oliver Syndrome in
humans. RBPJ/Su(H) is crucial for regulating gene expression during various
developmental processes [1, 3, 28, 29, 85]. These mutations in the RBPJ gene
adversely affect its ability to bind to DNA, leading to pathogenic effects [82].
Specifically, in Family 1, the mutation involved an A to G transition at position
188 (c.188A
Interestingly, a recent study identified similar missense mutations in previously described Drosophila Su(H)T4 and Su(H)O5 alleles [86, 87]. Su(H)T4, in particular, carries an E137V mutation at the same highly conserved glutamic acid residue as human RBPJ E63G [87]. This study analyzed phenotypes in two different tissues that are sensitive to dose changes in Notch signalling; the wing and macrochaetae in sensory bristles [87]. Flies heterozygous for either Su(H)T4 or Su(H)O5 allele show the typical Notch loss-of-function phenotype, the “Notched wing”, although these alleles are much less penetrant than Notch loss-of function alleles [86, 87]. However, both Su(H)T4 and Su(H)O5 alleles showed varying phenotypes in the sensory bristles where a small subset of heterozygous flies showed a decrease in macrochaetae, which is seen in flies with a lower number of copies of Hairless (Notch antagonist), and a small subset showed a slight increase macrochaetae, which is observed in flies carrying a Notch loss-of-function mutation [87]. This report shows that both Su(H)T4 and Su(H)O5 alleles enhance the Notched wing phenotype and increased macrochaetae in flies carrying one Notch-loss-of function allele and enhance the decrease of macrochaetae in flies carrying one Hairless loss-of function allele, suggesting that these alleles exacerbate the effect of low amounts of both the activator (Notch) and co-repressor (Hairless) [87]. Although Su(H)T4 exhibits both gain-of-function and loss-of-function activities in different contexts, this study provides evidence that Su(H) AOS-like mutants, similar to RBPJ mutants, suffer from defective DNA binding while maintaining an affinity for Notch co-activator and co-repressor proteins [87]. This suggests a sequestration mechanism where mutated variants of Su(H) bind to essential protein complexes, including Notch and Hairless, but no longer bind to DNA effectively, thus sequestering Notch or Hairless and exerting a dominant negative effect, and therefore Notch target genes are misregulated (Fig. 3) [87]. Su(H) binds to Hairless, a co-repressor, in the absence of Notch [1]. This study revealed that mutant Su(H) protein acts in a context-dependent manner to either enhance the Notch loss-of-function phenotype (Notched wing) or the Hairless loss-function phenotype (decreased macrochaetae cells in sensory bristles) depending on the genetic background of the tissue [88]. This finding indicates that tissue-specific genetic modifiers play a pivotal role in determining the severity and presentation of AOS. This mirrors the complex nature of AOS in humans, where various genetic modifiers can greatly affect tissue-specific outcomes [87].
 Fig. 3.
Fig. 3.Molecular mechanism underlying Adams-Oliver Syndrome (AOS) learned from Drosophila. RBPJ Mutations found in human (AOS) patients and Drosophila Su(H) exert a dominant negative effect. Experiments performed using Drosophila showed that this is due to a sequestration mechanism, where normally Notch intracellular domain (NICD) binds to Su(H) and facilitates the recruitment of the co-activator complex (left), the mutant variant does not bind to DNA, preventing target gene expression (right).
Overall, the Drosophila model system provides valuable insights into the study of Adams-Oliver Syndrome (AOS). It mirrors the sequestration mechanism observed in human RBPJ mutants, highlighting the model’s utility in unraveling the molecular basis of AOS and shedding light on the genetic complexities of this condition. Moreover, the Drosophila study emphasizes the significant influence of genetic background of the Su(H)T4 allele on tissue-specific phenotypic outcomes [87].
Carcinomas are malignancies that arise from epithelial cells and are the most common type of human cancer. Under stress, these epithelial cells can undergo a few processes adapt to the stressors [89, 90]. These include maladaptive processes such as metaplasia and dysplasia, that give rise to neoplastic cancerous cells [90]. Metaplasia is the reprogramming of stem cells leading to the replacement of one cell type by another that can adapt to a new stress [90]. This process becomes maladaptive when the new cells are not able to recapitulate the function of the original cells but rather are more resistant to the stressor [90]. If the stressor persists metaplasia can progress into dysplasia, which is disordered growth of precancerous cells and is not considered a true adaptive response [90, 91, 92]. It is characterized by loss of uniformity of cell size and shape (pleomorphism), disrupted tissue orientation and nuclear changes [90, 93]. Mild to moderate dysplasia can be reversible if the stressor is stopped, while severe dysplasia is irreversible and can progress to neoplasia (carcinoma in situ) [94]. One of the classic examples showing this progression is Barret’s esophagus, where normal esophageal epithelial cells are replaced by intestinal epithelium (Metaplasia) due to continuous reflux of gastric acid into the esophagus [91, 92, 93, 94, 95]. The persistent insult of gastric acid reflux leads to progression into dysplasia and increases the risk of esophageal adenocarcinoma [92]. Neoplastic cells become hyperproliferative and detach from the extracellular matrix which allows them to migrate and further divide to spread throughout a particular organ [91, 96, 97]. These cancerous cells can also invade nearby organs by breaking through tissue barriers and eventually invade the circulatory system and metastasize throughout the rest of the body [93].
In the early 1900s, a genetic study performed using Drosophila larvae identified a genetic mutation that demonstrated that tumours were inheritable, providing strong evidence that there exists an underlying genetic contribution to the development of cancer, which at the time was not yet clear [98, 99]. Early allograft experiments showed that transplanted tumours can spread and invade organs of the transplant recipient [100]. Drosophila has been used to model many of the hallmarks of cancer [101], which include: sustained and unlimited proliferation, escape from programmed cell death, resistance to inhibitors of proliferation, invasion, and metastasis [101]. The larval imaginal discs are sac-like structures that give rise to the adult’s external organs including the wings, eyes, legs, and genitalia [102]. The Drosophila wing disc has been instrumental as a model system in uncovering molecular pathways involved in development and has also been used extensively to model aspects of human disease and pathology [99]. The wing disc has been particularly useful in modeling tumours arising from epithelial cells. Studies using the eye and wing imaginal discs have uncovered cellular mechanisms underlying key steps in tumour initiation and progression, including the synergistic role of JNK signalling in proliferation, invasion, and metastasis [99, 103, 104]. Furthermore, large-scale genetic screens have identified perturbations that cause loss of cell adhesion and cell polarity, which are critical drivers of tumour growth and invasion [105, 106]. Here we will highlight evidence that outlines the role of Notch signalling in tumorigenesis and tumour progression from studies in Drosophila.
Specifically, a landmark study in the cancer biology field was aimed at developing a Drosophila-based genetic model of metastatic cancer using the FLP/FRT recombinase system [106]. Flippase (FLP) is a site-directed DNA recombinase isolated from Saccharomyces cerevisiae that directs recombination between two FLP recombination target (FRT) sites [107, 108, 109]. FLP recombination can be induced in somatic tissues to generate mosaic clones, where some cells can be wildtype while others carry a mutation of interest [107, 108, 109, 110]. Generating somatic mosaics is particularly useful when studying loss-of-function mutations of essential genes that would result in lethality if completely deleted in the animal. This system has also been widely used to generate germline clones that carry the mutation of interest when recombination is used in germline tissues [111]. The FLP/FRT system can be used to delete, insert, or invert genes depending on the placement and orientation of the FRT sites (Supplementary Fig. 1) [107, 108, 109]. This study employed the FLP/FRT recombinase system to introduce gain-of function or loss-of-function mutations into a variety of oncogenes and tumour suppressor genes, while simultaneously labelling transgenic cells carrying a mutation of interest with green fluorescent protein (GFP)-based reporters to track the position of mutant cells and to quantify cell migration and invasion across tissue boundaries [106]. Using this approach, it was determined that increased activity of the Ras signalling pathway induces tumour formation, however, these tumours were non-invasive [106]. When these perturbations were combined with a genetic deficiency of the gene scribble, an essential cell polarity gene, the resulting tumours were capable of invading other tissues including the hemolymph, leg discs, and tracheal vasculature [106]. As in mammalian tumours, these fly tumours exhibited reduced expression of cell adhesion proteins, including E-cadherin, whose loss is necessary for the metastatic behaviour of these tumours [106]. As this study modelled metastatic tumours in flies by perturbing a known oncogenic pathway and a cell polarity determinant, this provided opportunities for other researchers to study metastatic tumours in Drosophila. In particular, this study demonstrates not only the utility of a combinatorial genetic approach in uncovering interactions between genes in driving tumour progression, but also underscores the utility of Drosophila as a model system that allows for complex perturbations of multiple genes in the context of a developing organism.
The first example of cooperative tumorigenesis in Drosophila was reported in the early 2000s [105]. In this study a creative genetic tool was employed to deplete a cell polarity determinant, Scribble, in a subset of epithelial cells in the eye disc to more closely model mammalian cancerous tissue. Genetic deficiency of scribble, which is also a known tumour suppressor, in a subset of epithelial cells in the eye disc was sufficient to cause loss of cell polarity and limited over proliferation, however, was not sufficient to induce hyperproliferation and tumorigenesis despite upregulation of expression of the cell cycle regulator, cyclin E; a critical determinant of proliferation [105]. This report showed evidence that this was due to the action of JNK signalling in neighbouring cells which induced cell death [94]. Notch overexpression in scribble mutant cells was sufficient to prevent cell death induced by signals from neighbouring wildtype cells and induce the development of neoplastic tumours in the fly eye disc [105]. This report supports previous observations in mammalian cancerous tissue regarding the importance of cell polarity to cell survival and revealed cooperativity between the Notch- and Ras-signalling pathways, which enables escape of cell death programs in tumour-initiating cells and concomitant formation of neoplastic amorphous tumours [105].
Another study conducted a genetic screen to identify modifiers of neoplastic tumours in the adult eye and larval eye discs induced by eye-specific hyperactivation of Notch signalling, identified mir-8 as a tumour suppressor that was sufficient to significantly reduce eye tumour size and block metastasis that occurs upon overexpression of Delta and the Eyeful family of genes (Pipsqueak and Lola) [112]. The significant reduction of tumour size upon overexpression of mir-8 implies that mir-8 reduces Notch signalling activity, thereby reducing Notch-mediated hyperproliferation. Using in vitro luciferase-based reporter assays, this study demonstrated the direct binding of mir-8 to a sequence in the 3’ untranslated region (UTR) of the mRNA of the Drosophila Notch ligand Serrate, which results in the reduction of Serrate protein levels without affecting levels of Serrate mRNA, which was further confirmed using genetic rescue experiments of mir-8 gain-of-function mutants by Serrate overexpression [112]. This study clearly demonstrated that mir-8 targets Serrate, a Notch ligand, and inhibits its translation, thereby inhibiting Notch-induced hyperproliferation and tumorigenesis [112]. These findings motivated subsequent investigation into whether a similar mechanism exists in prostate cancers, where elevated Jagged1 expression is correlated with aggressive metastatic tumours [113]. Interestingly, expression of Jagged1; a mammalian Serrate homologue, is inversely correlated with the expression of homologues of mir-8, including of a set of human microRNAs: microRNA-200c-141, in prostate cancers cells (PC-3) [112]. Given that the expression of the miRNA200c~141 cluster in both PC3 (metastatic prostate cancer cells), and in clinical samples of late-stage prostate cancer, is significantly reduced compared to non-cancerous prostate cells (PNT1A cells), this study aimed to investigate whether miRNA200c~141 can inhibit Jagged1 translation. Evidence from overexpression of mir200c, mir-141, or overexpression of both in PC-3 cells in combination with direct binding assays established that Jagged1 is a direct target of mir200c~141 [112]. Importantly, the translational inhibition of Jagged1 by mir200c~141 inhibits cell growth of PC-3 cells involving a mechanism similar to that identified in Drosophila [112]. Furthermore, this study outlined a novel molecular interaction conserved between Drosophila and humans that underlies hyperproliferation and metastasis in genetically induced tumours in the Drosophila eye and human prostate cancer cells, further underscoring the value of Drosophila as a model system in identifying mechanisms of tumorigenesis and tumour progression. Specifically, this study represents an experimental paradigm that makes use of the power of Drosophila as a model system to investigate molecular genetic interactions in vivo and to test their conservation in mammalian disease models in building a foundational understanding for future targeted therapeutic screens.
In addition, a recent study focusing on Drosophila eye development
provided the first evidence of a synergistic interaction between the Notch
pathway and MEF2, a transcription factor that is crucial for myogenesis, in the
context of hyperproliferation [114]. Using the adult eye, larval eye and wing
disc, this study investigated how this synergy between Notch signalling and MEF2
contributes to cell proliferation in different tissues and developmental stages.
Metastatic potential was increased in cells where MEF2 and a constitutively
active form of the Drosophila NICD were co-expressed which was
demonstrated by their migration from the anterior compartment of the wing across
the anterior-posterior (AP) boundary [114]. Evidence was also provided to further
demonstrate that the synergistic relationship between NOTCH and MEF2 contributes
to the loss of apio-basal cell polarity, which was shown by the mislocalization
of Discs large (Dlg) to the apical surface rather than the basolateral junctions
that it is normally localized to [114]. This study demonstrated that the
overgrowth phenotype induced by overexpression of NOTCH and MEF2 is dependent on
JNK activation resulting from the increased expression of a JNK ligand,
Eiger, which was identified as a common transcriptional target of NOTCH
and MEF2 [114]. In addition, NOTCH and MEF2 synergy also leads to the altered
expression of cell adhesion molecules, including
2.4.3.1 The Drosophila Digestive System
The fly digestive system is composed of a long tubular structure as in mammals. The anterior foregut is analogous to the mammalian esophagus and is where ingested food travels through to the temporary storage organ, the crop [115, 116]. From the crop, food travels to the midgut, which is subdivided into 3 compartments; the anterior midgut, the middle midgut (which is a low pH region that contains Fe/Cu cells), and posterior midgut [115, 116]. The midgut is functionally analogous to the mammalian small intestine, as it is where the majority of nutrient absorption takes place [116]. As in mammals, the final stages of absorption (mainly of water and electrolytes) occur prior to excretion [116, 117]. In Drosophila these final stages of nutrient absorption prior to excretion occur in the hindgut, which is functionally analogous to the mammalian large intestine [116, 117]. From the hindgut, food and waste products are transported to the rectum and anus for excretion [116, 117]. In mammals, the renal system is responsible for filtration of the hemolymph, the reabsorption of water and solutes, and aids in excretion of nitrogenous wastes in the form of urine [116, 117]. In Drosophila, Malpighian tubes are the functional equivalent of the mammalian kidney that aid in water and solute reabsorption from the hemolymph, and which produce solid nitrogenous compounds that are released in the hindgut and excreted through the anus along with other waste products from the intestinal tract [116, 117]. Despite the structural differences that exist between the mammalian digestive system and its Drosophila counterpart, there is a high degree of functional conservation between these systems, making Drosophila a suitable model for studying the molecular and cellular determinants of digestive system development and homeostasis, regeneration, and the interplay with innate immune responses.
2.4.3.2 Drosophila as a Model System to Study Diseases of the Digestive System
The Drosophila midgut is composed of cell types that are similar in function to the mammalian digestive system. The molecular mechanisms underlying organ homeostasis are highly conserved [116, 118]. In this organ, the balance between stem cell renewal and differentiation is critical to the survival of the fly. Thus, Drosophila provides an excellent model system to study mechanisms that contribute to stem cell homeostasis and differentiation [116, 117, 118]. The fly midgut is also an interface separating the external environment (the midgut lumen) and internal organs, making the midgut an excellent model system to study innate immune responses and their impact on the digestive system [119, 120]. The Drosophila intestinal tract consists of four types of cells; intestinal stem cells (ISCs), enteroblasts (EBs), secretory enteroendocrine cells (EEs), and absorptive enterocytes (ECs) [121]. As a fly ages the proliferation rate of ISCs is reduced, which is reflected in the ratio of cell numbers between ISCs and the other intestinal stem cells [122, 123]. These age-induced changes make this organ a suitable model system for studying the mechanisms underlying stem cell homeostasis, tumorigenesis, and ageing [123]. The regulation of homeostasis and differentiation of ISCs is driven by conserved signalling pathways between Drosophila and mammals including; JAK-STAT, Wnt, and Notch signalling [124]. Notch signalling is necessary for stem cell differentiation in mammals and Drosophila. Notch signalling regulates stem cell differentiation and renewal in a context-dependent manner, which explains its complex context-dependent role in cancer pathogenesis [125, 126, 127]. In Drosophila, Notch signalling is necessary for the differentiation of ISCs into absorptive cells (ECs), while in mammals Notch signalling is required for the differentiation of ISCs into absorptive cells as well as the proliferation of ISCs [118, 124, 127]. Due to the differences outlined above, it is critical for researchers to confirm the conservation of mechanisms discovered in Drosophila in mammalian model systems. While the differential role of Notch signalling in ISC proliferation between Drosophila and mammals is an important difference to be noted, its conserved role in ISC differentiation presents an opportunity to study the mechanisms underlying ISC differentiation, which when misregulated can lead to tumorigenesis [128, 129]. In addition, Notch signalling can act as a tumour suppressor or oncogene in human stem-cell derived tumours in a context-dependent manner, therefore mechanisms discovered in Drosophila ISCs can provide insight into the potential tumour suppressing role of Notch signalling such as in hepatocellular carcinomas [130, 131, 132]. Furthermore, Drosophila offers an advantage to studying the underlying mechanisms of the above processes in vivo where interactions between neighbouring tissues and organs contribute to these signalling pathways, in contrast to cell culture-based models, which are limited in their ability to recapitulate key mechanisms involved in inter-tissue/organ communication.
2.4.3.3 Studying Mechanisms underlying Cancers of the Digestive System in Drosophila
A recent study used the Drosophila midgut intestinal stem cell niche to
study the effects of intercellular reactive oxygen species (ROS) levels on the
survival of tumour cells derived from ISCs in the context of hyperproliferation
(tumour growth) and metastasis (cell detachment and migration) [128]. This study
provided evidence that altering ROS levels alone does not significantly alter the
proliferation rate of ISCs [128]. To model tumour initiation and growth
Notch was genetically depleted using RNAi silencing in ISCs to induce
hyperproliferation [128]. When tumorigenesis was induced by Notch
depletion in ISCs, ROS levels were moderately elevated compared to their wildtype
counterparts [128]. Reduction of ROS levels, using overexpression of antioxidant
enzymes including CncC, Catalase, or SOD2, in these tumours resulted in decreased
tumour formation and decrease of hyperproliferation of the tumour like-ISCs,
suggesting that ROS production is necessary for tumorigenic ISC
hyperproliferation [128]. ISC survival has been previously shown to be dependent
upon attachment to the extracellular matrix (ECM) while metastatic tumours detach
from the ECM and migrate to distant tissues and invade other organs [133]. To
model metastasis, ECM adhesion was depleted by genetically silencing the gene
myospheroid (mys), which encodes
Another study used the fly gut to study interactions between intestinal stem cell-derived tumours and their microenvironment [129]. This study employed genetic blockade of Notch signalling using a heat-inducible RNAi system to generate intestinal stem cell tumours, demonstrating that these tumour cells were able to proliferate continuously in the absence of differentiation [129]. However, when Notch signalling was restored through removal of RNAi (by returning flies to 18 °C), these tumours differentiated into enteroendocrine cells, which contrasts with the normal program of differentiation in the fly midgut, where Notch activation promotes differentiation of enteroblasts into enterocytes [129]. This study provided evidence that Notch depletion is not necessarily sufficient for the initiation of tumorigenesis in all animals, suggesting that additional signalling cues are necessary for tumorigenesis in some contexts [129]. It was specifically shown that, in addition to Notch depletion-driven loss of differentiation capacity of ISCs, a stress signal, facilitated by JNK signalling and concomitant cytokine release, which induces ISC division, is required for tumour initiation in some contexts [129]. Additional signals, epidermal growth factor receptor (EGFR) activation, further drive tumour growth and displacement of intestinal enterocytes [129]. Signals from these invading ISCs induce JNK signalling activity and expression of cytokines, including UPD2 and UPD3, in enterocytes leading to enterocyte apoptosis [129]. In summary, this study outlined synergistic interactions between Notch and JNK signalling in tumour initiating ISCs as well as the contribution of paracrine signalling between tumorigenic ISCs their niche, via epidermal growth factor receptor (EGFR) and JNK signalling that facilitate tumour growth, detachment, and further proliferation [129]. These findings underscore the importance of signalling integration within cells and between different cells in the context of cancer development and progression.
These reports highlight the utility of Drosophila in elucidating the potential underlying molecular mechanisms and genetic determinants that contribute to the development and progression of different human cancers. Studies such as the ones discussed here, highlight the importance of understanding the role of signalling integration within cells and across different tissues. In addition, these reports demonstrate the complexity of the mechanisms underlying tumour initiation and growth due to the context-dependent effects of signalling pathways, where activation of a particular signalling pathway can be oncogenic or suppress tumour growth depending on the context (Fig. 4, [134, 135]). Several studies have made use of the rapid life cycle and low cost of Drosophila to conduct drug screens and have successfully identified potential therapeutic agents for several types of cancer [15, 16, 136, 137, 138, 139, 140]. The response to cancer therapies varies from patient to patient, hence cancer research is now aiming to develop and implement personalized treatment protocols that are patient-specific [140]. Recently, Drosophila was used to generate patient-specific models of cancer, where transgenic lines carrying the patient’s specific mutations were generated and used to test candidate treatments, demonstrating the versatility of Drosophila as a model system [16]. Here, we have focused specifically on the role of Notch signalling, and we hope that the studies discussed provide useful examples of the application of Drosophila as a model system in studying genetic interactions, conducting drug screens, as well as cell signalling interactions, that contribute to disease progression, as well as providing a base of knowledge that can be used to identify novel therapeutic agents.
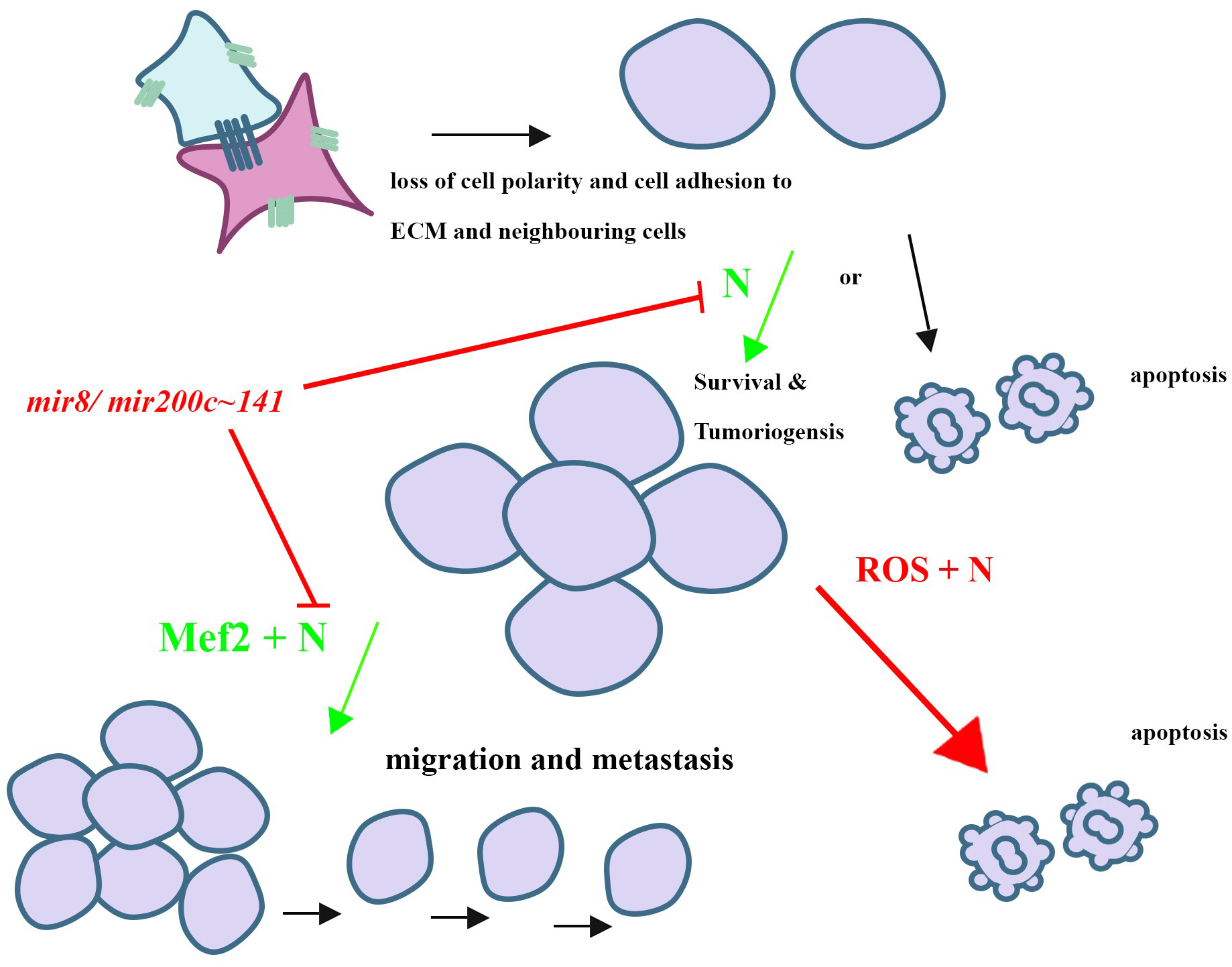 Fig. 4.
Fig. 4.Summary of the Notch-mediated mechanisms of tumorigenesis and cancer progression learned from Drosophila melanogaster. Notch signalling has been shown to play an oncogenic (green arrows) or tumour suppressor (red arrow) role depending on the cell type and environment in both mammals and Drosophila. One of the critical initial steps of tumorigenesis is the loss of cell polarity and adhesion to neighbouring cells as well as the extracellular matrix (ECM). Cells that lose cell polarity and/or adhesion often undergo apoptosis, however, if they escape apoptosis, they may start to over proliferate and form tumours. Notch signalling was shown to play a pro-survival role of these detached, aiding in their escape from apoptosis and tumour formation. In addition, MEF2, a transcription factor, has been associated with human cancers. Synergy between Notch signalling and MEF2 facilitates tumorigenic cells to metastasize in Drosophila and their co-expression was shown to be correlated with recurring breast cancers. mir-8 was found to play a tumor suppressor role by inhibiting the translation of Notch ligands reducing hyperproliferation induced by Notch signalling activity in Drosophila tumours. While Notch signalling is known to play an oncogenic role in several types of human cancer tissues, including T-cell leukemia, it is known that Notch signalling can also drive the expression of tumour suppressors, including Hes1 in thyroid carcinoma cells [134, 135]. Notch signalling was found to cooperate with reactive oxygen species signalling to promote apoptosis of tumorigenic cells in Drosophila. The human homologues of mir-8 were later found to inhibit Notch signalling in a similar manner, showing the utility of Drosophila as an excellent model system for the investigation of genetic interactions underlying human diseases such as cancer and for the discovery of potential therapeutic targets. ROS, reactive oxygen species.
Cardiomyopathy is a broad term describing a broad range of diseases of the heart muscle. These diseases all affect the function of the heart in regulating circulatory flow. Cardiomyopathies are broadly categorized based on which phase of the cardiac cycle they affect; either the filling phase (diastolic function) or the pumping phase (systolic function) [141, 142]. Impairment of diastole, where not enough blood enters the heart due to structural abnormalities (e.g., Hypertrophic, restrictive, or constrictive cardiomyopathies), results in insufficient blood delivered to the periphery and poor perfusion to the organs [142]. Impairment of systole, usually a result of the weakness of the heart muscle or excessive dilation (dilated cardiomyopathy), also causes insufficient blood delivery to the periphery and poor perfusion to the organs [142]. In both cases, an insufficient volume of blood is pumped out from the heart and instead accumulates in the lungs, resulting in fluid leakage into the parenchyma causing dyspnea (difficulty breathing) [142]. As blood continues to accumulate in the heart, there can be a point of overload in the right side of the heart, which ultimately results in a redistribution of blood to the periphery causing edema and swelling [142]. In addition, the volume-overloaded heart undergoes structural changes, i.e., ventricle dilation and ventricle wall weakening, which causes valvular insufficiency, and which interferes with the electrical conduction of the heart [142]; changes that manifest as dizziness, fatigue, fainting, heart murmurs, and arrhythmias [141, 142].
The fly heart is structurally distinct from the human heart. It is organized as a linear, rather than a chambered and convoluted, tube that is composed of a monolayer of cardiomyocytes, which comprise a single chamber [143, 144]. In contrast to the human heart, the fly heart lacks coronary blood vessels and rather relies on simple oxygen diffusion for cardiac perfusion [143, 144]. The Fly “Heart-beat” is made up of two types of pulses: retrograde and anterograde pulses that are generated by pacemaker cells located on the caudal and rostral sides of the heart tube [145]. Hemolymph enters the heart through ostia (openings) and flows towards the rostral side during retrograde beating, and leaves through a structure called the aorta (located in the thorax) [146]. Hemolymph flows through valves in the caudal circulatory system into the lumen of the tube and is propelled towards the rostral or caudal end [146].
Due to the structural differences between the fly and human heart, it is
difficult to model many aspects of cardiovascular disease such as arrhythmias,
ischemia, and diseases specific to coronary blood vessels. While in these
contexts, zebrafish and mice provide alternative model systems, models of dilated
cardiomyopathy have been well characterized in the fly [144, 146, 147, 148, 149, 150, 151].
Specifically, in 2006, a study used Optical Coherence Tomography (OCT) to image
the cardiac chambers in living, awake, adult flies for the first time [149]. This
report presented evidence that a mutation in
A genetic screen for dilated cardiomyopathy phenotypes identified a potential
atypical Notch signalling ligand (Wry) lacking a Delta/Serrate/LAG-2 (DSL)
domain, which is present in all other Notch ligands [148]. Using OCT, this group
identified a deletion that caused enlargement of cardiac chambers and
corresponding impairments to cardiac function [148]. Through the application of a
combination of tissue-specific GAL4 drivers, the temperature-inducible
GAL80
Overall, this study leveraged the genetic tools available in Drosophila to study cardiomyopathy, identified a novel gene necessary to the function of Notch in regulating homeostasis, and characterized the function of the protein product of the Wry gene, its interactions, and its role in cardiac function (Fig. 5). Thus, this study supports previous research that demonstrated a role for Notch signalling in cardiac function, and which provided the novel understanding that this role is not limited to developmental stages but is also important in regulating homeostasis in adult tissues.
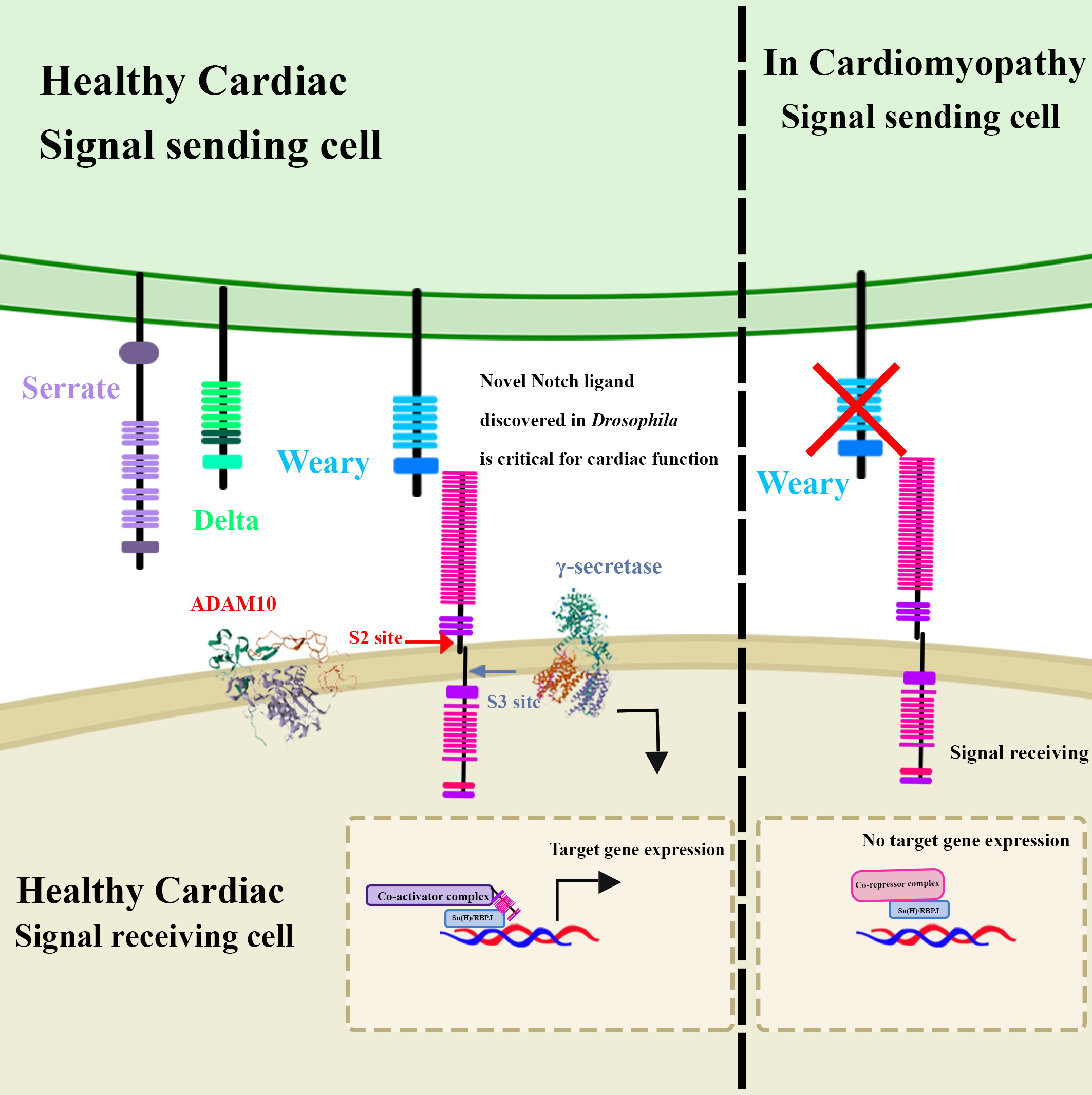 Fig. 5.
Fig. 5.Notch signalling activity was found to be critical for normal
cardiac function. Through a genetic screen, a novel protein, Weary, which
physically interacts with the Notch receptor and drives expression of Notch
reporter gene, was identified and characterized as a potential Notch ligand
necessary for cardiac function. Its deletion or the reduction of Notch signalling
in cardiac tissue causes cardiomyopathy in Drosophila. The ADAM10 structure was obtained from the RCSB PDB (https://www.rcsb.org/) of PDB ID 6BE6. The
One of the largest advantages of using Drosophila as a model system is the availability of a large and diverse genetic toolbox. FlyBase is a freely available database is compilation of information from literature and other databases about the Drosophila genome. Flybase provides information including, but not limited to; genomic architecture and expression data, links to reagents and stocks, known mutations, links to high throughput expression data at the protein and mRNA level at different developmental stages and across different tissues, and associations with human diseases and relevant publications [159]. Thus, Flybase is an invaluable tool for researchers working with Drosophila as it provides useful summaries of what is known about a gene of interest and available tools to study this gene [159]. The Berkeley Drosophila Genome Project (BDGP), which is integrated into Flybase, was a project that set out to sequence the Drosophila genome, annotate gene sequences, characterize their expression patterns using mRNA in situ hybridization, and to develop bioinformatics tools to make this data easily accessible to researchers [160, 161, 162, 163, 164, 165, 166]. The modEncode project is a more recent project of the BDGP that is designed to characterize both functional elements of the Drosophila genome and the expression patterns of gene products on a genome-wide basis using high-throughput transcriptional assays, which include RNA-sequencing [167]. Flyfish is another excellent resource that provides high resolution images of fluorescent in situ mRNA hybridizations across different stages of embryogenesis and larval stages [168, 169]. Another excellent resource is the Drosophila Genome Resource Center, which provides useful reagents, including cell lines, DNA plasmid vectors, etc. as well as protocols that make accessible established techniques for Drosophila researchers worldwide. In addition, Developmental Studies Hybridoma Bank (DSHB) provides affordable antibodies against Drosophila proteins. FlyCRISPR is an excellent database which identifies specific target sequences in an input DNA sequence provided by the user, which is helpful in designing sgRNAs [170, 171]. In addition, FlyCRISPR provides useful protocols, reagents and other resources for genomic editing using CRISPR [170, 171]. These resources and their corresponding links can be found in Table 1 (Ref. [168, 169, 170, 171]).
| Tool | Useful for obtaining | Link |
| FlyBase | Gene sequence | https://flybase.org/ |
| Genetic interactions | ||
| Expression data of products | ||
| Recent publications | ||
| Relevant reagents | ||
| Known mutations and mutant flylines | ||
| Associations of gene with signalling pathways | ||
| Associations of gene with human diseases | ||
| BDGP & modEncode | Expression data | https://www.fruitfly.org/index.html |
| cDNA clones | ||
| Expression vectors | ||
| FlyFish | Expression patterns and mRNA localization patterns | https://fly-fish.ccbr.utoronto.ca/ [168, 169] |
| FlyCRISPR | sgRNA for target sequences and protocols for genome editing using CRISPR | https://flycrispr.org/ [170, 171] |
| Drosophila Genomics Resource Center | Fly stocks, cDNA clones, vectors, and Drosophila cell lines | https://dgrc.bio.indiana.edu/Home |
| Protocols for caring for Drosophila lines and cell lines | ||
| Protocols for molecular assays | ||
| InteractiveFly | Summaries of gene function and interactions and contributions to development, cellular and tissue homeostasis taken from research publications | https://www.sdbonline.org/sites/fly/aimain/1aaintrb.htm |
| DSHB | Monoclonal and Polyclonal Antibodies | https://dshb.biology.uiowa.edu/ |
| Bloomington | Fly stocks | https://bdsc.indiana.edu/index.html |
Drosophila melanogaster has been used as a model system to characterize the contributions of many genes and their products during development, disease, and adult tissue homeostasis. In this review, we highlight the role of Drosophila in investigating and elucidating molecular mechanisms that contribute to Notch signalling-associated human diseases (Figs. 2,3,4,5). The high degree of conservation of the Notch signalling pathway between Drosophila and humans provides researchers with an opportunity to take advantage of the vast genetic toolbox available that can be used in combination with advanced imaging techniques that allow visualization of molecular interactions in real-time in the context of cells, tissues, organs, and the entire fly. However, while there exist many advantages to using Drosophila to model human disorders, it is important to mention that some limitations do exist, and that scientists working with Drosophila should be aware of. On a molecular level, despite the high degree of conservation of the Notch signalling pathway, Drosophila lack some of the human ligands and receptors in the Notch pathway. In addition, due to major anatomical structural differences between Drosophila and humans, certain aspects of human pathologies cannot yet be modelled, although there have been highly creative techniques and methods that have circumvented some of these issues to closely model human disease mechanisms. Regardless, modelling human pathologies in Drosophila has led to several landmark discoveries that highlighted novel potential therapeutic targets and biomarkers of several human diseases including cancers and cardiac disorders. Furthermore, findings from studies using Drosophila as a model system have elucidated underlying mechanisms and identified novel genetic contributors to human disease. One of the invaluable advantages that Drosophila offers as model system is that it provides the opportunity to investigate signalling crosstalk within cells and between different tissues and organs in vivo, while still being a cost-effective and efficient system to genetically manipulate and perform high throughput studies on. We anticipate that further advances will continue to be made in this field in the future, and that insights from these advances will contribute to the understanding and treatment of human disease.
All authors MarvM, AT, RDH, GF, MarkM, and AN contributed significantly to searching the literature and writing the original manuscript and meet the requirements of ICMJE. MarvM and AN contributed to literature search, writing and editing of all sections (‘1’–‘3’) of the review paper. AT wrote section ‘2.3’. RDH wrote section ‘2.2’. GF assisted with writing section ‘1’ and assisted with creating Figs. 1,2,3,5. MarkM wrote parts of sections ‘2.4’ and ‘2.5’. MarvM created Figs. 1,2,3,4,5. Funding was acquired by AN. All authors contributed to editing and writing the final manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable. No experiments were conducted by the authors of this review paper.
The authors thank the National Science and Engineering Research Council of Canada for funding this work. The authors also thank the editors and reviewers for their constructive feedback and valuable insights.
This research was funded by the National Science and Engineering Research Council of Canada (Grant #: RGPIN-2018-06781 NSERC).
The authors declare there are no conflicts of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.





