1 Center for Clinical Laboratories, The Affiliated Hospital of Guizhou Medical University, 550004 Guiyang, Guizhou, China
2 Guizhou Center for Disease Control and Prevention, 550004 Guiyang, Guizhou, China
3 Department of Hematology, The Affiliated Hospital of Guizhou Medical University, 550004 Guiyang, Guizhou, China
4 Department of Clinical Biochemistry, School of Clinical Laboratory Science, Guizhou Medical University, 550004 Guiyang, Guizhou, China
†These authors contributed equally.
Abstract
A long non-coding RNAs (LncRNAs) called antisense noncoding RNA in the INK4 locus (ANRIL), has emerged as substantial regulators of cell survival in acute myeloid leukemia (AML). However, its speciffc and potential mechanism is uncertain in AML. In this research, we investigated the role of ANRIL in cell proliferation, apoptosis, and the underlying mechanism in AML cells.
ANRIL expression was quantified by real-time quantitative polymerase chain reaction (RT-qPCR). Kasumi-1 cells were transfected with LV-ANRIL plasmid to upregulate ANRIL expression, with or without co-transfection with a G Protein-Coupled Receptor Kinase 2 (GRK2) siRNA. Additionally, these cells were transfected with sh-ANRIL plasmid to inhibit ANRIL expression, with or without co-transfection with a GRK2 overexpression plasmid. Cell proliferation and apoptosis were determined using the cell counting kit-8 (CCK8) and flow cytometry. Protein expression levels of phosphatidylinositide 3-kinases (PI3K), protein kinase B (AKT), phosphorylated-Akt (p-AKT), Bcl-2-associated protein x (BAX), B-cell leukemia/lymphoma 2 protein (BCL-2), proliferating cell nuclear antigen (PCNA), cleaved caspase-3, and GRK2 were detected by western blot. The RNA-binding protein immunoprecipitation (RIP) assay was conducted to investigate the interaction between ANRIL and GRK2.
ANRIL expression was increased in Kasumi-1 cells. ANRIL upregulation expression promoted cell proliferation and inhibited apoptosis. Furthermore, its upregulation led to increased expressions of PI3K, AKT, p-AKT, PCNA, and BCL-2, and decreased expression of BAX in Kasumi-1 cells. Additionally, transfection with GRK2 siRNA attenuated the promoting effect of LV-ANRIL on Kasumi-1 cells proliferation and the PI3K/AKT pathway, increased BAX and cleaved caspase-3 expressions, and decreased BCL-2 and PCNA expressions. GRK2 overexpression reversed the inhibitory effect of sh-ANRIL on cell proliferation and the PI3K/AKT pathway. Furthermore, it promoted BCL-2 and PCNA expressions, and inhibited BAX and cleaved caspase-3 expressions. RIP assay confirmed the physical interaction between ANRIL and GRK2.
The GRK2 protein-mediated ANRIL, increasing Kasumi-1 cell proliferation and inhibiting apoptosis by activating the PI3K/AKT/BCL-2 pathway.
Keywords
- acute myeloid leukemia
- ANRIL
- GRK2
- proliferation
- apoptosis
- PI3K/AKT/BCL-2 pathway
Acute myeloid leukemia (AML) is a heterogeneous disease affecting myeloid hematopoietic stem or progenitor cells. It is marked by the aberrant proliferation of primitive and immature myeloid cells in the both the bone marrow and peripheral blood. AML is the most common acute leukemia in adults [1, 2]. The treatment modalities predominantly comprise chemotherapy and hematopoietic stem cell transplantation, with daunorubicin (DNR) being a crucial chemotherapeutic agent [3]. Nevertheless, the long-term survival rates for patients with AML remain low because of poor tolerance to chemotherapy, post-transplantation complications, and high mortality associated with treatment [4]. AML-M2, a common subtype with a median survival of less than 2 years and a five-year survival rate below 40% [5]. Thus, identifying novel molecular targets for AML-M2 is central to improving the overall survival (OS).
Long noncoding RNAs (lncRNAs) are RNA transcripts synthesized by RNA polymerase II, typically exceeding 200 nucleotides in length [6]. LncRNAs are crucial in tumor growth, differentiation, and development by interacting with DNA, RNA, and proteins [7, 8]. ANRIL, a lncRNA located at the INK4 locus, was first identified by Pasmant et al. [9] in the genetic study of melanoma nervous system tumor family. Interestingly, ANRIL imbalance is associated with the progression of numerous cancers, such as breast tumors, hepatic cancer, and AML [10, 11, 12, 13, 14]. Nevertheless, the underlying mechanisms remain elusive. Recent investigations have reported that ANRIL can silence p15 expression by binding and recruiting Polycomb-group protein 2 (PRC2) [15], suggesting that its role in interacting with proteins to participate in tumor development. Based on previous studies, the AML-M2 cell line Kasumi-1 with high expression of ANRIL was applied as a target cell line to probe the influence of lncRNA ANRIL on the growth and apoptosis of Kasumi-1 cells.
Our previous studies elucidated that the downregulation of ANRIL expression in Kasumi-1 cells brought about inhibition of cell proliferation and enhance apoptosis via the phosphatidylinositol-3-kinase (PI3K)/protein kinase B (AKT) pathway [16], which highlighted that regulation of ANRIL may pave the way for novel treatment of acute myeloid leukemia subtype M2 (AML-M2). Nevertheless, the effects of upregulation of ANRIL expression on the biological characteristics of AML-M2 and its mechanism still require further clarification and validation. Accordingly, the current study set out with a series of experiments to verify the role and possible mechanism of upregulation of lncRNA ANRIL expression on the proliferation and apoptosis of Kasumi-1 cells. To provide foundational theories for clarifying the function and mechanism underlying ANRIL in AML-M2.
Kasumi-1 cells (cat: TCHu202), HL-60 (cat: TCHu 23), THP-1 (cat: SCSP-567), HEL (cat: TCHu 71), TF-1 (cat: TCHu139), and U-937 (cat: TCHu159) were obtained from the Chinese Academy (Shanghai, China). SKM-1 cells were obtained from the Pricella Biotechnology Co., Ltd. (Wuhan, China). Fetal bovine serum (Siji Qing) was purchased from Tianhang Biotechnology Co., Ltd (cat: 13011-8611, Zhejiang, China). RPMI 1640 medium was acquired from Gibco (cat: 11875093, GrandIsland, NE, USA). Penicillin and streptomycin were sourced from Suolaibao Technology Co., Ltd (cat: P1400, Beijing, China). Reverse transcription (RT) and RT-quantitative real-time polymerase chain reaction (qPCR) reagents were purchased from Takara (cat: RR037A, Osaka, Japan). The Cell Counting Kit-8 (CCK-8) reagent was purchased from MedChemExpress (cat: HY-K0301, Newark, NJ, USA). Annexin V-APC/7-amino-actinomycin D reagent was purchased from Procell (cat: P-CA-107, Wuhan, China). Primary antibodies against glyceraldehyde-3-phosphate dehydrogenase (GAPDH, 1:5000), phosphatidylinositide 3-kinases (PI3K, 1:1500), protein kinase B(AKT, 1:1500), phosphorylated-Akt (p-AKT, 1:1500), Bcl-2-associated protein x (BAX, 1:1000), B-cell leukemia/lymphoma 2 protein (BCL-2, 1:1000), proliferating cell nuclear antigen (PCNA, 1:1500), G Protein-Coupled Receptor Kinase 2 (GRK2, 1:1500), and horseradish peroxidase (HRP, 1:5000)-labeled ancillary antibodies were procured from Sanying Biotechnology Co., Ltd (Wuhan, China). The cleaved caspase-3 antibody was purchased from Cell Signaling Technology (cat: 9664S, Danvers, MA, USA). The electrochemiluminescence (ECL) reagent was purchased from Millipore (cat: WBKLS0100-2, Bedford, MA, USA). The ANRIL gene lentivirus was purchased from Genesis Co., Ltd (Shanghai, China). Both GRK2 overexpression plasmid (pCDH-CMV-MCS-EF1-copGFP-T2A-Puro) and siRNA sequences were acquired from Aiji Biotechnology Co., Ltd (Guangzhou, China).
Peripheral blood samples were collected from 31 patients with AML. Their basic information and medical characteristics were recorded. AML diagnosis was confirmed according to the French American British classification standard. The control group consisted of peripheral blood from 13 healthy individuals. This project was approved by the Affiliated Hospital of Guizhou Medical University, and informed consent was obtained from all patients. The ethical approval number is 2016 (65).
The cells of the human AML line (Kasumi-1) were provided by the Chinese Academy. Kasumi-1 cells were cultured in RPMI 1640 complete medium with 10% fetal bovine serum and 1% penicillin/streptomycin in 5% CO2 at 37 °C, and were mycoplasma-free (Pulilai Gene Technology Co., Ltd., Mycoplasma Detection Kit, cat: E2036, Beijing, China). The medium was changed and the cells sub-cultured every 2 days. The cell line was authenticated shortly before use by the Short Tandem Repeat (STR) validation, carried out by Fuheng Biotechnology Co., Ltd (Shanghai, China).
Total RNA extraction was extracted from the human AML cells following the manufacturer’s instructions. The RNA was reverse-transcribed into complementary DNA (cDNA) using Prime Script RT kits (cat: RR036A, Takara, Osaka, Japan). ACTB (F: 5′-GCGTGACATTAAGGAGAAGC-3′, R: 5′-CCACGTCACACTTCATGATGG-3′) was used as the internal control. The expression patterns of ANRIL (F: 5′-ATAAGCCTCATTCTGATTCAACAGC-3′ and R: 5′-AGCAGTACTGACTCGGGAAAG-3′) and GRK2 (F: 5′-GCGCTCACTCCTTTCAA-3′, R: 5′-GAGATCTGGAGGCACCTGCTT-3′) were detected by RT-qPCR at the following cycling condition: 95 °C for 30 s, 1 cycle; 95 °C for 5 s, 60 °C for 30 s, 40 cycles. Gene expression was calculated using the 2-ΔΔCt method.
Kasumi-1 cells were seeded at a concentration of 5
According to the instructions of the RIP Kit (cat: FI8709, FITGENE, Guangzhou, China), Kasumi-1 cells lysates were collected. And extracted 50 µL protein A/G agarose beads into a 1.5 mL centrifuge tube, resuspended protein A/G agarose beads with 100 µL NT-2, added 5 µg of target antibody or IgG antibody, and incubated at room temperature for 1 hour. The magnetic bead antibody complex was subsequently washed and used for spare. Extracted 100 µL of cell lysate supernatant into 900 µL of protein A/G and antibody complex, and incubate overnight at 4 °C. The bead protein complexes were subsequently washed. RNA was extracted from the complex, and ANRIL expression was quantitatively analyzed by RT-qPCR.
Cells were adjusted to a density of 1
Cells were adjusted to a density of 5
Total cell lysates were prepared using RIPA lysis buffer (cat: R0010, Solarbio,
Beijing, China) consisting of a protease inhibitor cocktail. After mixing with
5
Kasumi-1 cells lysates were collected. According to the instructions of the Nuclear and Cytoplasmic Extraction Kit (cat: P0028, Beyotime, Shanghai, China). Added 200 µL of cytoplasmic extraction reagent A and 10 µL cytoplasmic extraction reagent B every 20 µL of cell precipitation. Subsequently, centrifuged the mixture at 12,000 g for 10 minutes and transfered supernatant into a tube to obtain the extracted cytoplasmic. For precipitation, added 50 µL of nuclear extraction reagent. Then centrifuged the mixture at 12,000 g for 10 minutes. Sucked the supernatant into the tube, which was the nuclear. RNA in nuclear and cytoplasmic was extracted following the manufacturer’s instructions. The RNA was reverse-transcribed into complementary DNA (cDNA) using Prime Script RT kits (cat: RR036A, Takara, Osaka, Japan). GAPDH (F: 5′-GGAGCGAGATCCCTCCAAAAT-3′, R: 5′-GGCTGTTGTCATACTTCTCATGG-3′) was designated as a positive control of cytoplasmic, and U6 (F: 5′-GTGCAGGGTCCGAGGT-3′, R: 5′-CTCGCTTCGGCAGCACA-3′) was used as positive control of nucleus.
The GEPIA2 database (http://gepia2.cancer-pku.cn/#index) was utilized to assess ANRIL expression in AML specimens. The Multi Experiment Matrix database (https://neic.no/mem-multiexperimentmatrix/) was utilized to predict potential downstream interacting proteins of ANRIL. The RPISeq database (http://pridb.gdcb.iastate.edu/RPISeq/#) was performed to predict the interaction probability between ANRIL and GRK2. The RNAfold software (SnapGene 6.1, GSL Biotech, San Diego, CA, USA) was adopted to predict the secondary structure of ANRIL. Moreover, the catRAPID database (http://service.tartaglialab.com/) was carried out to predict the interaction site between ANRIL and GRK2.
Statistical analysis were conducted using SPSS 2.0 (IBM Corp., Chicago, IL,
USA). Each experiment was conducted in triplicate, with the data recorded as the
mean
First, the GGEPIA2 database was utilized to compare ANRIL expression between the AML patients and individuals without leukemia. High ANRIL expression was observed in AML patients (Fig. 1A). ANRIL expression was further validated using peripheral blood specimens from the patients and human AML cell lines, including THP-1, HEL, Kasumi-1, HL-60, U937, SKM-1, and TF-1. ANRIL was substantially overexpressed in AML patients and Kasumi-1 cells, compared with healthy controls (Fig. 1B,C). Kasumi-1 cells were selected for the subsequent experiments. The cells have been validated for mycoplasma and Short Tandem Repeat (STR), no cross contamination of human derived cells or mycoplasma contamination (Supplementary Fig. 1A,B).
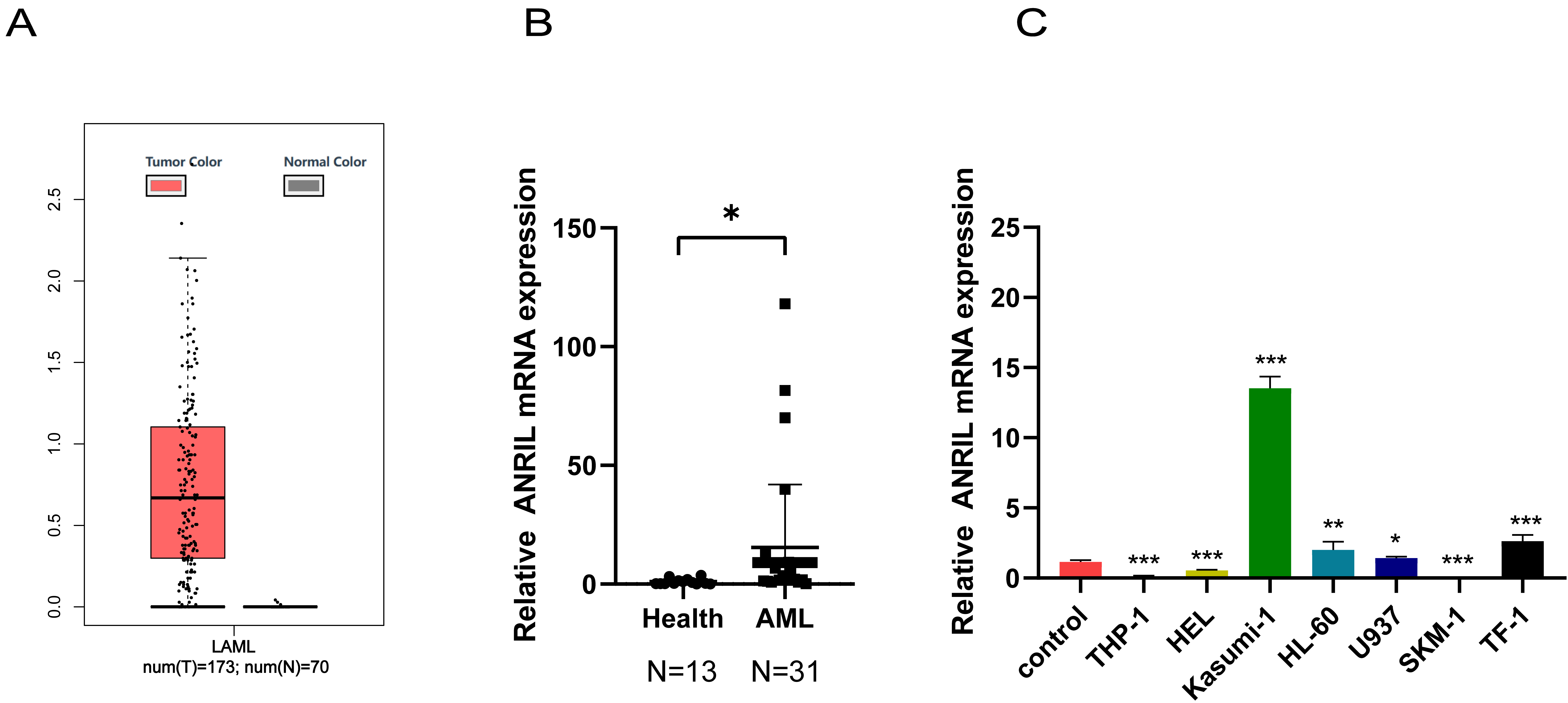 Fig. 1.
Fig. 1.
ANRIL overexpressed in AML patients and
Kasumi-1 cells. (A) GGEPIA2 database was utilized to compare the
differential expression of ANRIL between the AML patients and
individuals without leukemia. (B) RT-qPCR was performed to detect the expression
of ANRIL in AML patients. (C) RT-qPCR was adopted to detect the
expression of ANRIL in AML cell lines (n = 3) (*p
Recombinant lentivirus was adopted to establish the Kasumi-1
cells with upregulated ANRIL. The expression of green fluorescent
protein (GFP) in both groups were observed via fluorescence microscopy (Fig. 2A),
RT-qPCR analysis indicated a 7.5-fold increase in ANRIL expression in
the LV-ANRIL group, compared with the LV-ANRIL NC group (Fig. 2B). The CCK-8 assay demonstrated that upregulated ANRIL significantly
promoted Kasumi-1 cell proliferation (p
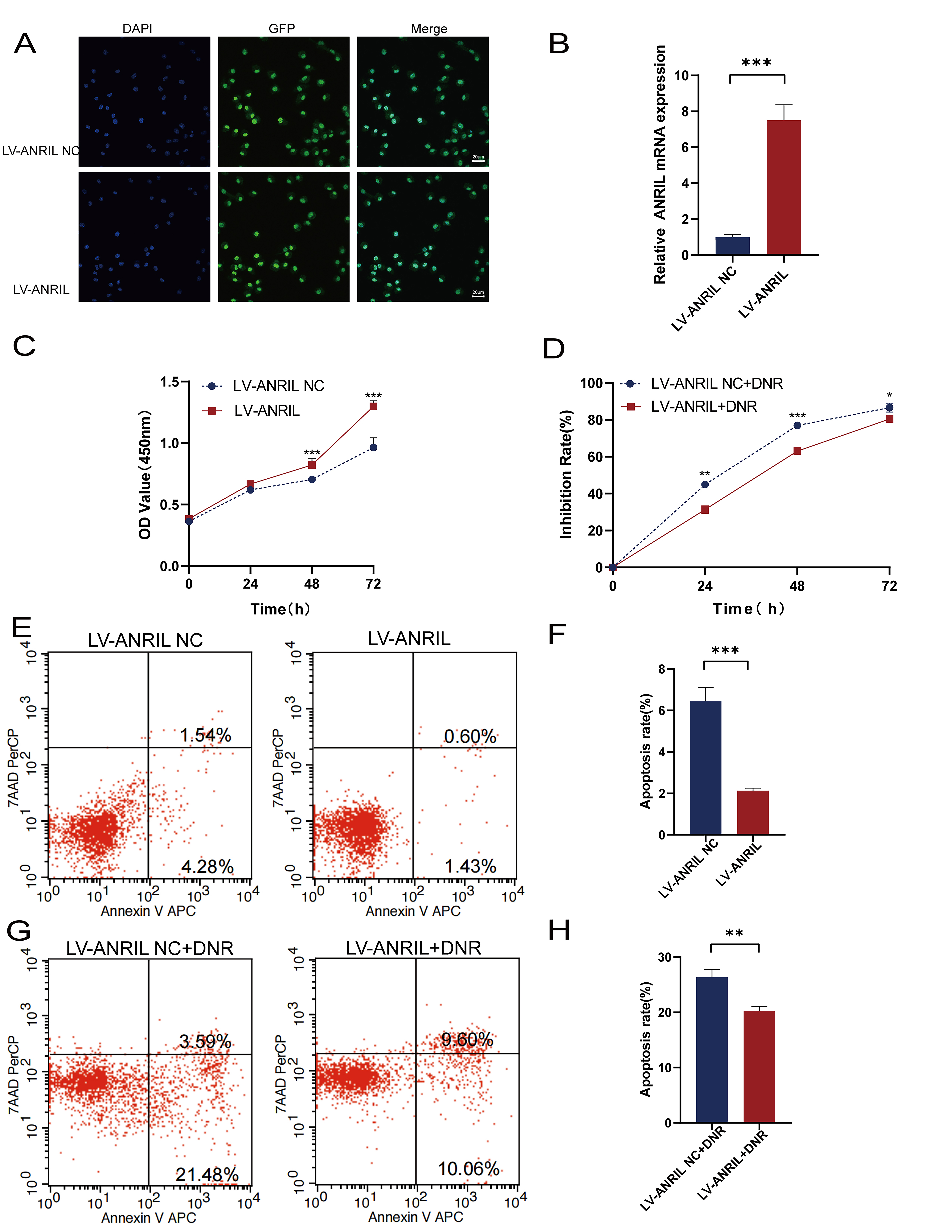 Fig. 2.
Fig. 2.
ANRIL upregulation promoted Kasumi-1 cell
proliferation and reduced the cytotoxic effect of DNR. (A) A fluorescence
microscope was utilized to observe green fluorescent protein (GFP) expression and
DAPI fluorescent staining (scale bar: 20 µm). (B) RT-qPCR was performed to
detect the overexpression efficiency of ANRIL in Kasumi-1 cells (n = 3).
(C) CCK-8 assay was carried out to detect the effect of upregulation of
ANRIL on the proliferation of Kasumi-1 cells (n = 3).
(D) CCK-8 assay was applied to detect the effect of upregulation of
ANRIL on the cytotoxic effect of DNR on Kasumi-1 cells (n = 3). (E,F)
Flow cytometry was adopted to detect the effect of upregulation of ANRIL
on Kasumi-1 cell apoptosis. (G,H) Flow cytometry was used to detect the apoptosis
rate of Kasumi-1 cells under DNR after ANRIL upregulation (n = 3).
(*p
The PI3K/AKT pathway regulates cell proliferation, apoptosis, migration, and
invasion [17]. Therefore, a western blot assay was conducted to identify the
expression patterns of the PI3K/AKT pathway and associated proteins after
increased ANRIL express in Kasumi-1 cells. ANRIL upregulation
significantly increased PI3K/AKT pathway activity in Kasumi-1 cells, evidenced by
enhanced protein expressions of PI3K, PCNA, and BCL-2 as well as AKTphosphorylation. Moreover, it reduced BAX expression, suggesting that
ANRIL upregulation may promote activation of the PI3K/AKT signaling
pathway, thereby altering the expression or activity of related molecules.
Subsequently, Kasumi-1 cells with upregulated ANRIL expression were
treated with LY294002, a PI3K/AKT inhibitor. Compared with the untreated group,
protein expressions of PI3K, PCNA, and BCL-2 as well as AKT phosphorylation were diminished in the LY294002 treatment group. Additionally,
BAX expression was enhanced in the LY294002 treatment group, suggesting that it
could reverse activation of the PI3K/AKT/BCL-2 pathway induced by the upregulated
ANRIL expression, restore BAX protein expression, and inhibit PCNA
protein expression (Fig. 3A,B). Flow cytometry analysis suggested that the
apoptosis rate in LY294002 treated group was (11.04
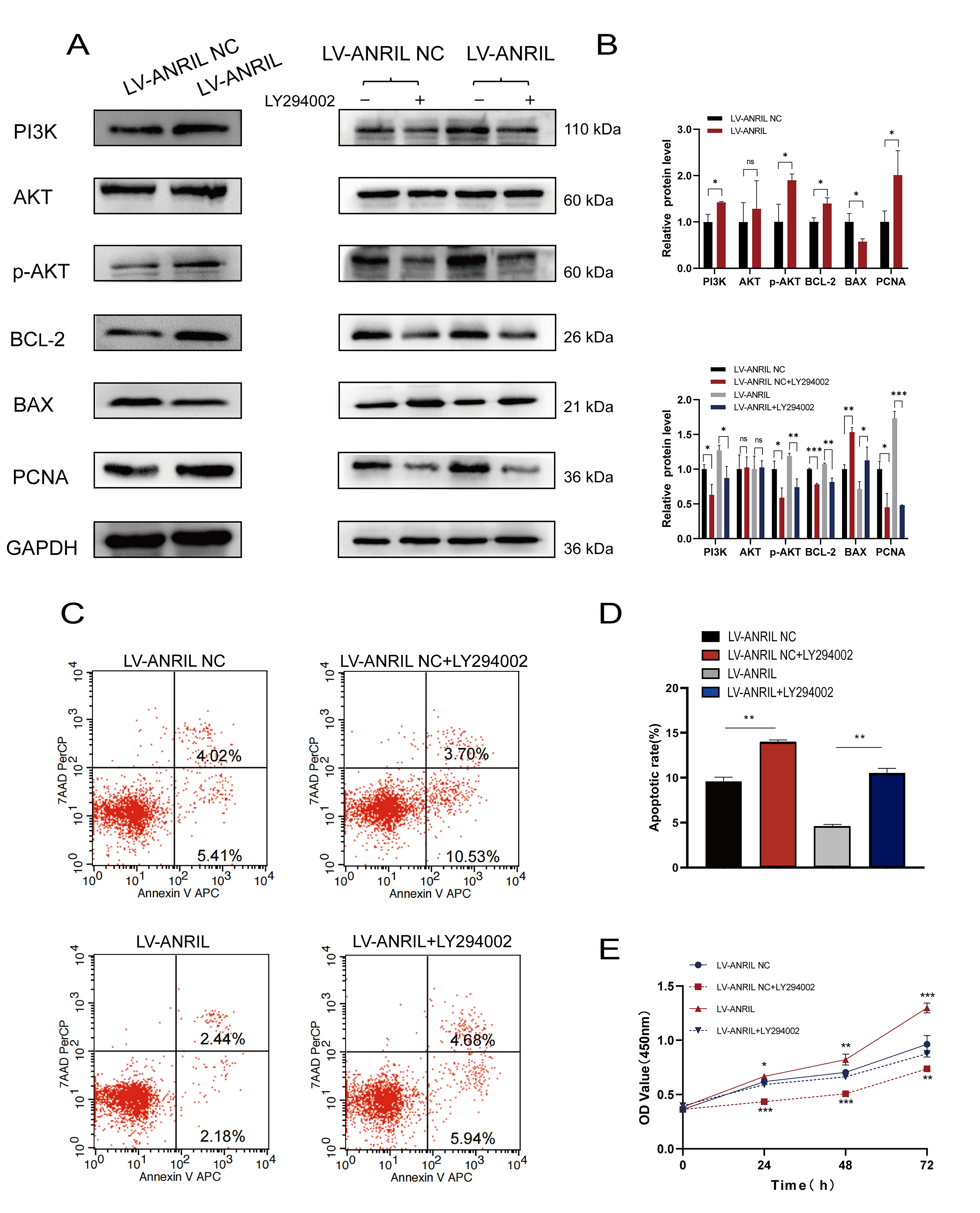 Fig. 3.
Fig. 3.
ANRIL promoted proliferation and inhibited apoptosis of
Kasumi-1 cells via PI3K/AKT pathway. (A,B) Western Blot assay was
performed to detect the expression of related proteins after upregulation of
ANRIL and LY294002 (n = 3). (C,D) Flow cytometry was utilized to detect
the apoptosis of cell treated with LY294002 after ANRIL
upregulation. (n = 3). (E) CCK-8 assay was carried out to detect cell
proliferation after ANRIL upregulation (n = 3) (*p
ANRIL activates the PI3K/AKT/BCL-2 pathway, enhancing Kasumi-1 cell growth and inhibiting apoptosis, however, its underlying remains unclear.
Karyoplasm isolation experiments suggested that ANRIL is distributed in
both the cytoplasm and nucleus of Kasumi-1 cells (Fig. 4A). We used
bioinformatics to predict potential downstream proteins interacting with
ANRIL, and observed an interaction with GRK2 (Fig. 4B,C). The RIP assay
demonstrated significantly enriched ANRIL by the anti-GRK2 antibody,
compared with non-specific IgG control (p
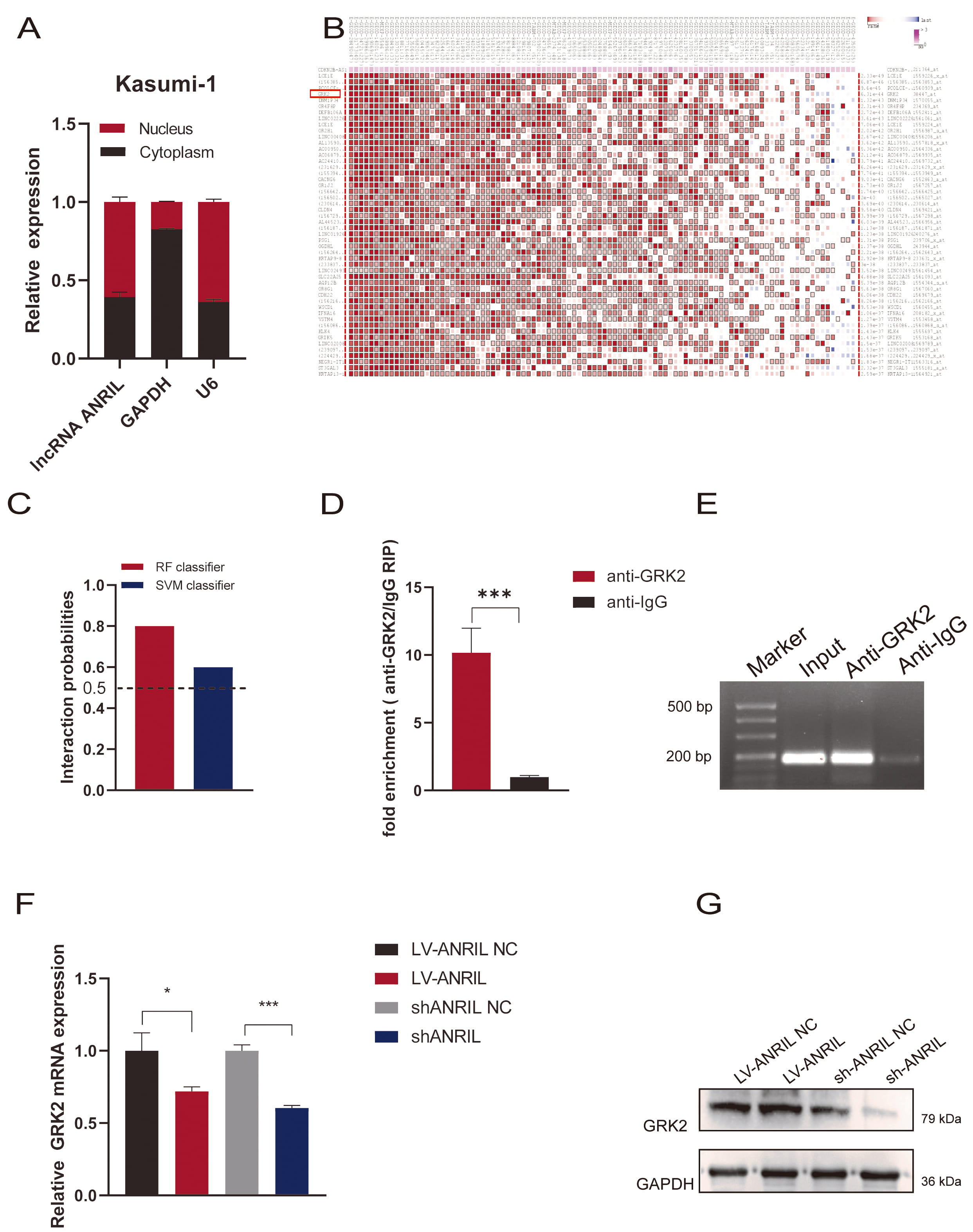 Fig. 4.
Fig. 4.
ANRIL interacted with GRK2 in Kasumi-1 cells.
(A) Nucleocytoplasmic separation assay was carried out to detect the expression
of ANRIL in cytosolic and nuclear in Kasumi-1 cells (n = 3). (B)
Bioinformatics was adopted to predict potential downstream proteins interacting
with ANRIL. (C) RPISeq database was used to predict the possibility of
interacting between ANRIL and GRK2. (D,E) RIP assay was applied to
detect the interaction between ANRIL and GRK2 (n = 3). (F) RT-qPCR was
performed to detect the mRNA level of GRK2 after upregulation or
downregulation of ANRIL (n = 3). (G) Western Blot assay was adopted to
detect the expression of GRK2 protein after upregulation or downregulation of
ANRIL (*p
Researchers have established the role of GRK2 in regulating tumor cell proliferation and invasion [18]. Nonetheless, its role in the AML-M2 Kasumi-1 cells remains elusive. Consequently, we investigated the impact of GRK2 on Kasumi-1 cell proliferation and apoptosis by upregulating or downregulating GRK2 expression.
The GRK2 overexpression plasmid was transfected into Kasumi-1 cells, and GFP
expression was observed via fluorescence microscopy (Fig. 5A), and western blot
assay was performed to detect the effect of GRK2 overexpression transfection
(Fig. 5B). Three GRK2 siRNAs (siGRK2#1/2/3) were transfected into Kasumi-1
cells, with the interference efficiency determined through the western blot
assay. siGRK2#1 and siGRK2#2 displayed noticeable interference effects (Fig. 5C). To guarantee the precision and objectivity of the outcomes, two sequences,
namely siGRK2#1 and siGRK2#2, were used in subsequent experiments. The flow
cytometry assay indicated that the apoptosis rate of the oe-GRK2 group was
significantly lower in the oe-GRK2 group (4.73
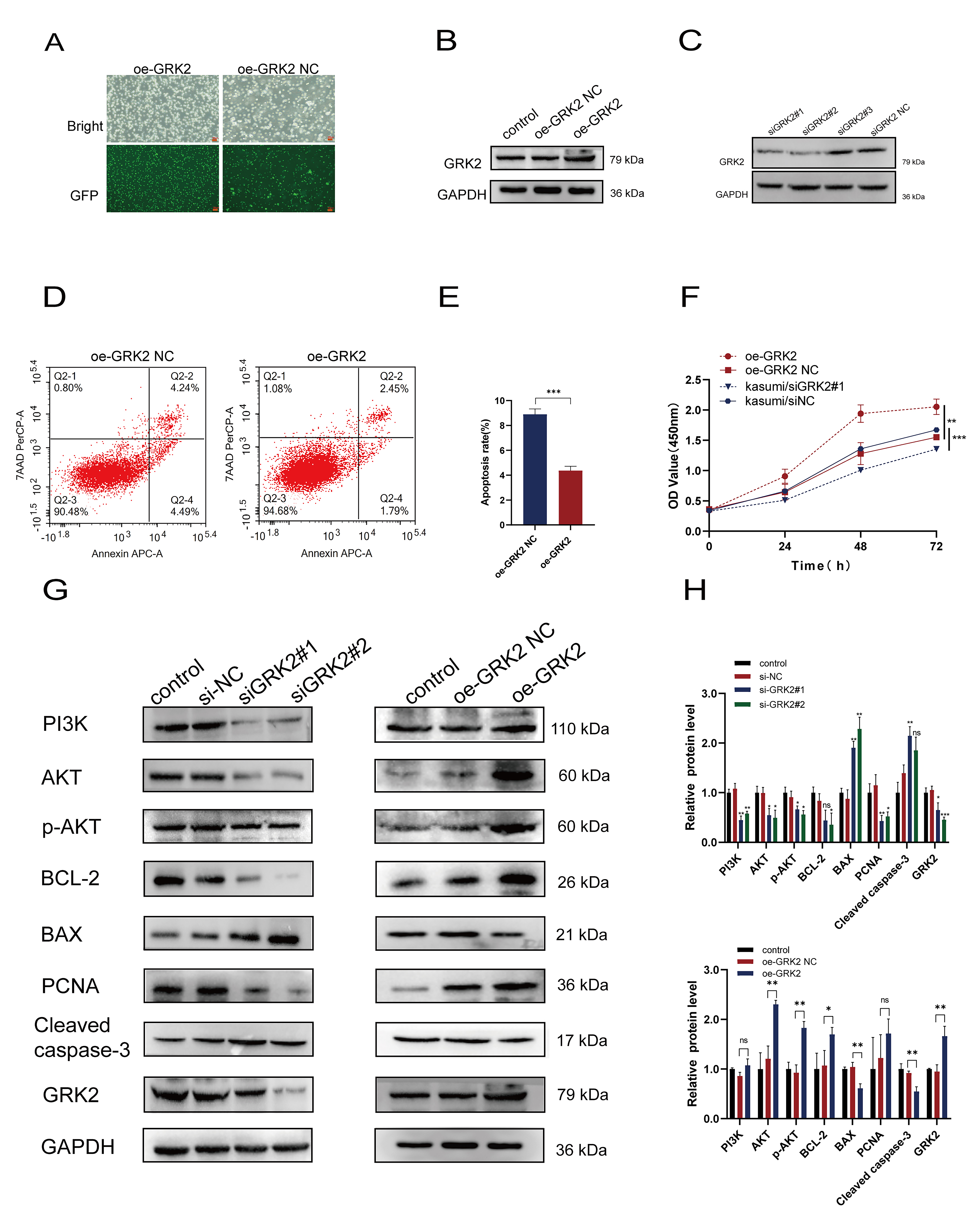 Fig. 5.
Fig. 5.
GRK2 promoted Kasumi-1 cell proliferation and
inhibited apoptosis via activation of the PI3K/AKT pathway. (A) A fluorescence
microscope was utilized to observe green fluorescent protein (GFP) expression
(scale bar: 100 µm). (B) Western blot assay was performed to detect the
effect of GRK2 overexpression transfection. (C) Western blot assay was performed
to detect the interference effect of GRK2 siRNA#1/2/3. (D,E) Flow cytometry was
adopted to detect the effect of upregulation of GRK2 on Kasumi-1 cell apoptosis
(n = 3). (F) CCK-8 assay was carried out to detect cell proliferation after
knockdown and upregulation of GRK2 (n = 3). (G,H) Western blot assay was applied
to detect the expression of related proteins after knockdown and upregulation of
GRK2 (n = 3) (*p
We elucidated the role of GRK2 in regulating the PI3K/AKT/BCL-2 pathway and
affecting Kasumi-1 cell proliferation and apoptosis. Next, we constructed
downregulated ANRIL co-transfection with upregulated GRK2
(sh-ANRIL+oe-GRK2) and upregulated ANRIL co-transfection with
GRK2 siRNA (LV-ANRIL+siGRK2) in Kasumi-1 cells for rescue experiments.
Downregulated GRK2 expression reversed the proliferation-promoting effects of
ANRIL overexpression on Kasumi-1 cells (Fig. 6A). Conversely,
upregulated GRK2 expression reversed the inhibitory effects of
ANRIL downregulation on cell proliferation (Fig. 6B). Flow cytometry
results suggested that the apoptosis rate was significantly lower in the
sh-ANRIL+oe-GRK2 group (1.20
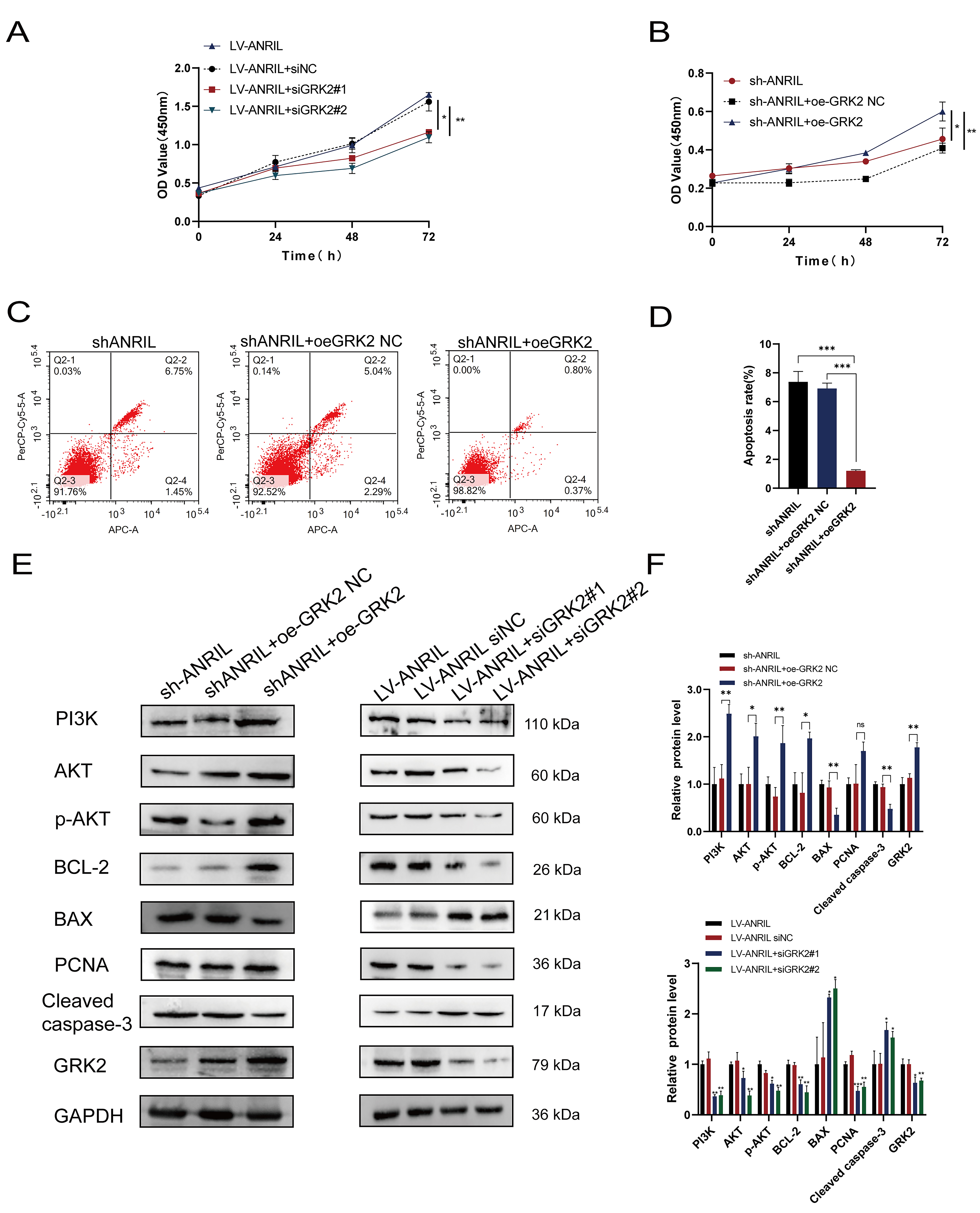 Fig. 6.
Fig. 6.
ANRIL regulated Kasumi-1 cell proliferation
and apoptosis by interacting with GRK2 protein. (A) CCK-8 assay was
carried out to detect the effect of co-transfection of LV-ANRIL and
siGRK2 on Kasumi-1 cell proliferation (n = 3). Compare the data of group siNC and siGRK2#1/2. (B) CCK-8 assay was
performed to detect the effect of co-transfection of sh-ANRIL and
oe-GRK2 on Kasumi-1 cell proliferation (n = 3). (C,D) Flow cytometry was
utilized to detect the effect of co-transfection of sh-ANRIL and
oe-GRK2 on Kasumi-1 cell apoptosis (n = 3). Compare the data of group (sh-ANRIL + oe-GRK2 NC) and (sh-ANRIL + oe-GRK2). (E,F) Western Blot assay was
applied to detect the effect of co-transfection of (sh-ANRIL and
oe-GRK2) and (LV-ANRIL and siGRK2) on the PI3K/AKT
signaling pathway (n = 3) (*p
LncRNAs can interact with DNA, RNA, and proteins through secondary space structures to regulate gene expression [19]. The RNAfold software predicted the secondary structure of ANRIL. ANRIL regions at positions 1550 to 1880 nt (Fig. 7A,a), 2140 to 2620 nt (Fig. 7A,c) and 200 to 500 nt (Fig. 7A,b) were folded to form a complex stem-loop structure, which may serve as a protein interaction site. The catRAPID database was utilized to predict the interaction points between ANRIL and GRK2. The heatmap illustrates that the 279 to 418 nt regions of ANRIL (Fig. 7A,b) have possible interaction site with the 439 to 640, 251 to 351 and 39 to 101 regions of the GRK2 amino acid sequence. Additionally, the interaction score was the highest with the 439 to 640 region (Fig. 7B), corresponding to the carboxyl-terminal Pleckstrin Homolgy (PH) domain of the GRK2 protein structure. Thus, the 279 to 418 nt regions of ANRIL may interact with the PH domain of GRK2 protein (Fig. 7C).
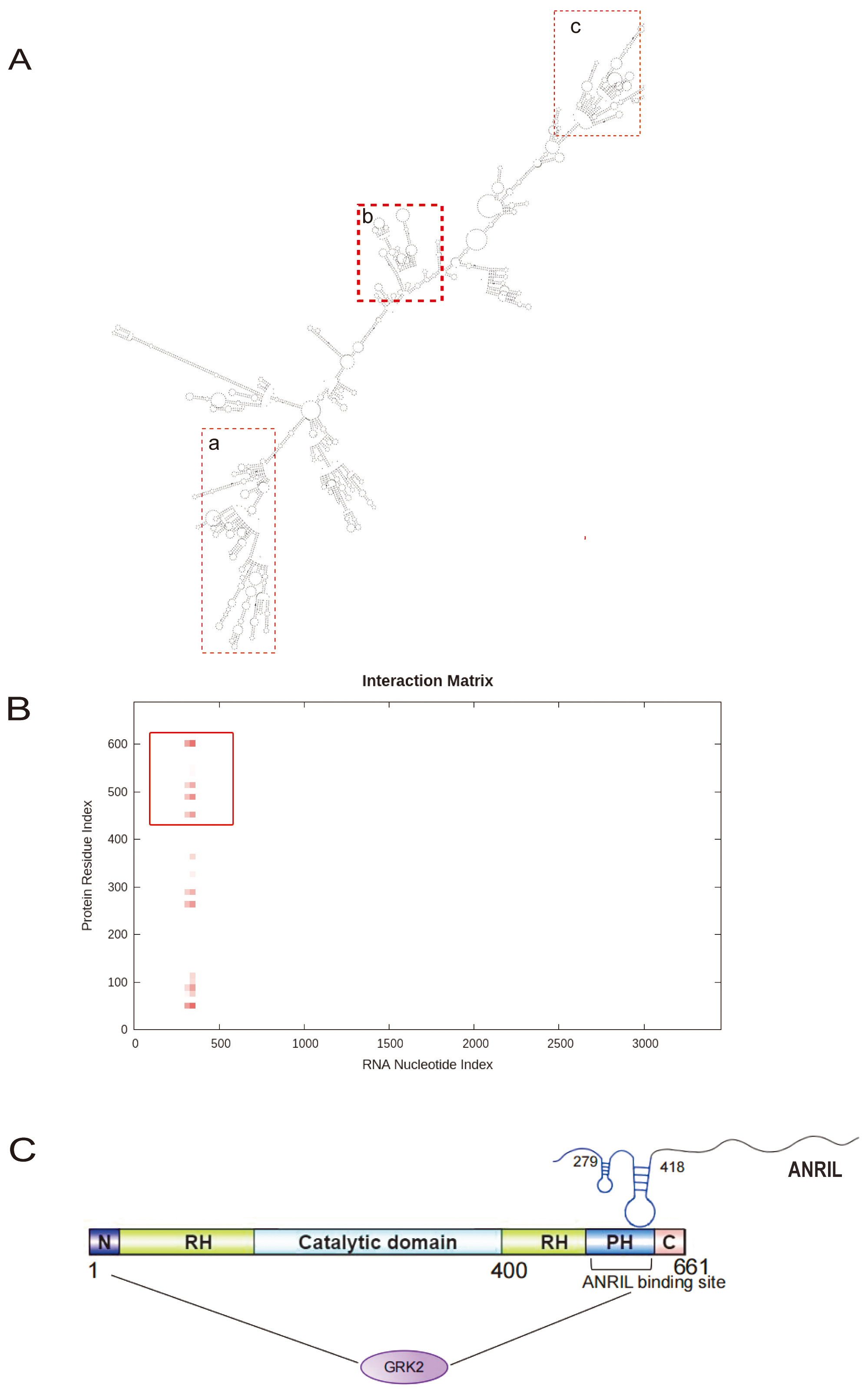 Fig. 7.
Fig. 7.
Predicted interaction sites between ANRIL and GRK2. (A) Secondary structure of ANRIL predicted by RNAfold software (SnapGene 6.1, GSL Biotech, San Diego, CA, USA). (A,a) Secondary structure of ANRIL regions at positions 1550 to 1880 nt. (A,b) Secondary structure of ANRIL regions at positions 200 to 500 nt. (A,c) Secondary structure of ANRIL regions at positions 2140 to 2620 nt. (B) Interaction sites between ANRIL and GRK2 predicted by the catRAPID database (http://service.tartaglialab.com/). (C) Interaction sites between ANRIL and GRK2. N, Amino-terminal domain; C, Carboxyl-terminal domain; PH, Pleckstrin Homolgy domain; RH, RGS Homology domain.
AML is a common hematological malignancy [20]. Novel drugs and treatment approaches have enhanced the clinical efficacy and quality of life. Nonetheless, several patients experience relapse after remission or develop drug resistance. The 5-year OS rate remains only 27% [21, 22], necessitating the need for novel therapeutic targets to improving clinical outcomes in AML. Emerging evidence underscores the pivotal function of lncRNAs in the pathophysiological mechanisms underlying hematological malignancies, highlighting them as pivotal biomarkers for leukemia diagnosis, prognosis, and therapeutic interventions [23]. ANRIL has been implicated as an oncogenic driver in AML. Wang et al. [13] reported that ANRIL interacts with miR-34a to upregulate histone deacetylase 1 expression, affecting the AML cell proliferation and apoptosis. Furthermore, Sun et al. [14] mentioned that ANRIL promotes AML cell survival by regulating the AdipoR1/AMPK/SIRT1 metabolism-related pathway. However, a comprehensive understanding of ANRIL expression and its functional implications in AML cells remains imperative.
Previously, we observed high ANRIL expression in peripheral blood and bone marrow samples from patients with AML. Downregulated ANRIL expression effectively induced apoptosis in AML-M2 Kasumi-1 cells [16]. Therefore, we demonstrated this phenomenon by upregulating ANRIL expression. The upregulation of ANRIL expression facilitated Kasumi-1 cell proliferation while impeding apoptosis. Simultaneously, it antagonized the cell-killing effect of the DNR. Therefore, ANRIL may play a pivotal role in AML occurrence and progression. This is consistent with previous studies [13, 14]. Next, we investigated the mechanism by which ANRIL affects the proliferation and apoptosis of Kasumi-1 cells.
The PI3K/AKT pathway controls cell proliferation, apoptosis, and tumor cell growth [24, 25, 26], and is abnormally stimulated in various cancers, including AML [27, 28]. PI3K serves as a critical regulator of tumor progression. The phosphorylation of downstream AKT affects the expression of several downstream effector molecules, thereby regulating cellular functions, such as metabolism, apoptosis, and proliferation of cells [29]. Of these effectors, the BCL-2 family consists of anti-apoptotic (BCL-2, BCL-x) and pro-apoptotic (BAX, BAK) proteins [30, 31]. BCL-2 plays an anti-apoptotic function and high BCL-2 expression has been associated with drug resistance in hematological malignancies [32]. By contrast, high BAX expression promotes apoptosis and indicates a good prognosis in AML patients [33]. Notably, PCNA serves as a tumor proliferation marker and is closely associated with tumor occurrence and development [34]. Strikingly, high PCNA expression has been reported in patients with relapsed AML [35]. In line with our findings, we found that BCL-2 and PNCA were highly expressed in AML [32, 35]. Herein, our findings demonstrated that ANRIL upregulation promoted PI3K, AKT, p-AKT, and BCL-2 expressions as well as PCNA, while inhibiting BAX protein expression in Kasumi-1 cells. Additionally, we utilized LY294002, a PI3K/AKT pathway inhibitor, to understand the association between ANRIL and the PI3K/AKT pathway. Treatment with LY294002 could reverse the proliferative and anti-apoptotic effects induced by ANRIL upregulation, emphasizing that ANRIL activates the PI3K/AKT/BCL-2 pathway to promote Kasumi-1 cell proliferation and inhibit apoptosis.
GRK2, a serine/threonine-protein kinases regulates diverse signaling pathways, including PI3K/AKT and MAPK, thereby influencing tumor cell proliferation, differentiation, and apoptosis [18]. Consistent with this research, our study elucidated that GRK2 promotes Kasumi-1 cell proliferation and inhibits apoptosis by activating the PI3K/AKT/BCL-2 pathway. Recently, some reports have found that ANRIL acts as a microRNA sponge [13, 36], or binds with proteins to regulate downstream responses. ANRIL was reported to modulate cancer cell survival by physically interacting with PRC2 [15]. Here, we found that ANRIL can physically bind to GRK2 protein. Interestingly, ANRIL upregulation elevated GRK2 protein expression without changing its mRNA levels. Thus, this interaction may modulate GRK2 protein synthesis or degradation by altering its structure or activity, potentially masking the ubiquitination or phosphorylation modification sites on GRK2 [37, 38]. Furthermore, GRK2 downregulated reversed ANRIL’s ability to promote Kasumi-1 cell proliferation and attenuate its activation of the PI3K/AKT/BCL-2 pathway, suggesting that ANRIL may promote the AML progression through the GRK2 protein. To validate this hypothesis, we intervened Kasumi-1 cells by upregulating GRK2 on the basis of ANRIL downregulation. Remarkably, GRK2 upregulation restored ANRIL’s effects on Kasumi-1 cell proliferation, apoptosis, and the PI3K/AKT pathway. Hence, the GRK2 protein mediated ANRIL, which activated the PI3K/AKT/BCL-2 pathway to promote Kasumi-1 cell proliferation and inhibit apoptosis.
In summary, this research shed light on the biological importance of ANRIL and GRK2 in Kasumi-1 cells. It was found that ANRIL can physically bind to GRK2 protein, promote the expression of BCL-2 and PCNA proteins by activating the PI3K/AKT signaling pathway, and inhibit the expression of cleaved caspase-3 and BAX proteins, leading to Kasumi-1 cell proliferation. However, interference with GRK2 expression can reverse this phenomenon, and siGRK2 has a similar effect to the application of PI3K/AKT signaling pathway inhibitor LY294002. It can reverse the regulatory effect of upregulating ANRIL expression on Kasumi-1 cell proliferation, apoptosis, and PI3K/AKT signaling pathway, leading to an increase in cleaved caspase-3 and BAX protein expression, a decrease in BCL-2 and PCNA protein expression, and ultimately inducing Kasumi-1 cell apoptosis (Fig. 8). Altogether, ANRIL can serve as a potential therapeutic target for AML-M2, and specific small-molecule inhibitors may be developed for ANRIL, which might improve the status of clinical treatment for AML-M2. Nevertheless, the intricate regulatory mechanism underlying the interaction between ANRIL and GRK2 remains elusive. Future investigations should aim to delineate the specific association and binding sites between GRK2 and ANRIL, and further in vivo experiments are warranted to ascertain the therapeutic potential of ANRIL in AML.
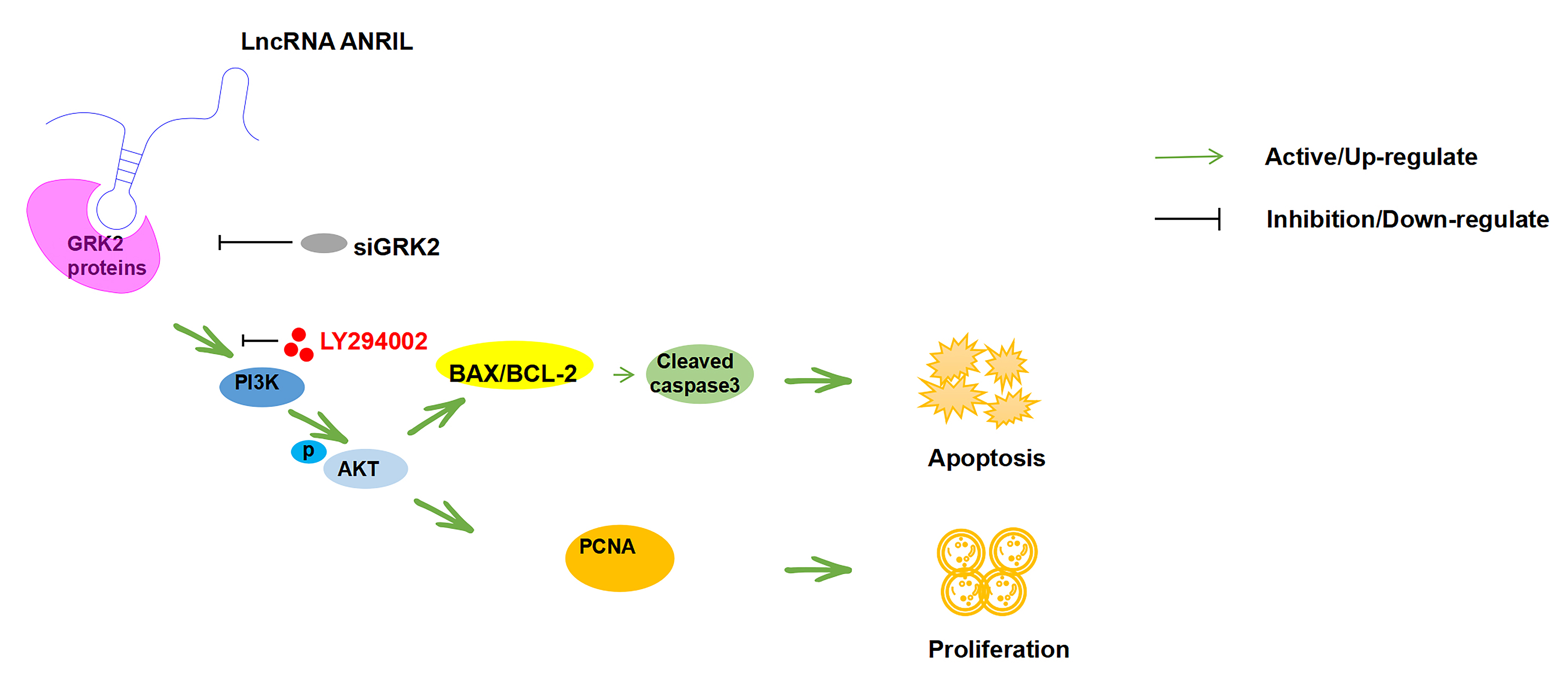 Fig. 8.
Fig. 8.
Mechanism of ANRIL in Kasumi-1 cells.ANRIL can physically bind to GRK2 protein, promote activation of the PI3K/AKT signaling pathway and further affect the proliferation and apoptosis of Kasumi-1 cells. Meanwhile, the PI3K/AKT signaling pathway inhibitor LY294002 and GRK2 siRNA can weaken the aforementioned effects of ANRIL upregulation on the Kasumi-1 cells. siGRK2, downregulation of GRK2; PI3K, phosphatidylinositide 3-kinases; AKT, protein kinase B; BCL-2, B-cell lymphoma-2; BAX, Bcl-2 assaciated X protein; BAX/BCL-2, the ratio of BAX to BCL-2; PCNA, proliferating cell nuclear antigen.
Our results showed that lncRNA ANRIL overexpress in AML patients and Kasumi-1 cells. ANRIL can physically bind to GRK2 protein, activate the PI3K/AKT/BCL-2 pathway, promote Kasumi-1 cell proliferation and inhibit apoptosis. However, the effect of ANRIL on Kasumi-1 cells was reversed by GRK2 siRNA, leading to an increase in cleaved caspase-3 and BAX protein expression, a decrease in BCL-2 and PCNA protein expression, and ultimately inducing Kasumi-1 cell apoptosis. Therefore, we confirmed that GRK2 protein-mediated ANRIL can enhance Kasumi-1 cell proliferation and prevent apoptosis by activating the PI3K/AKT/BCL-2 pathway. This research offers a foundational theory and novel ideas for elucidating the function of and mechanism underlying ANRIL in the clinical treatment of AML-M2.
AML, Acute myeloid leukemia; ANRIL, Antisense non-coding RNA in the INK4 locus; cDNA, complementary DNA; FBS, Fetal bovine serum; DNR, Daunorubicin; LncRNA, Long non-coding RNA; RT-qPCR, Reverse transcription-Real-time quantitative; PCR, Polymerase Chain Reaction; RNA, Ribonucleic Acid.
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
SQW, CSZ conceived and designed the research and wrote the manuscript; SQW, CSZ performed experiments; SXW, HH, FHD revised the manuscript and participated in the experimental guidance; JXX, HLH, JXZ, WZ, YMZ, CLH, YCL assisted in collecting peripheral blood and bone marrow samples. All authors have participated sufficiently in the work to take public responsibility for appropriate portions of the content and agreed to be accountable for all aspects of the work in ensuring that questions related to its accuracy or integrity. All authors read and approved the final manuscript. All authors contributed to editorial changes in the manuscript.
The study was approved by the medical ethics committee of the Affiliated Hospital of Guizhou Medical University, and informed consent was obtained from all patients. The ethical approval number is 2016 (65).
The authors gratefully acknowledge everyone who contributed to this paper, including those who helped with sample collection, clinical consultation, writing guidance. Thanks for the years lost in the experimental work but given us harvest.
The study was supported by research grants from the National Natural Science Foundation of China to Sixi Wei (Grant No.81660027), the Science and Technology Project of Guiyang City to Sixi Wei (Grant NO. 20161001021).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/j.fbl2910362.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.








