1 Department of Biochemistry, Cancer Biology, Neuroscience and Pharmacology, School of Medicine, Meharry Medical College, Nashville, TN 37208, USA
Abstract
Chemokines bind to specific chemokine receptors, known as cell surface G protein-coupled receptors, constructing chemokine axes which lead to cell migration and invasion in developmental stage, pathophysiological process, and immune reactions. The chemokine axes in the tumor microenvironment are involved in tumor growth, angiogenesis, cancer stem-like cell properties, metastasis, and chemoresistance, modifying tumor immune contexture and cancer progression. Clinical features, including tumor state, grade, lymph node metastasis, and cancer subtypes, are related to the specific chemokine axes, which play a significant role in immune contexture and cell to cell interaction in the tumor microenvironment, followed by altered cancer prognosis and overall survival. The present review summarizes the role of chemokine axes in breast cancer, based on data obtained from cell line and animal models and human tumor samples. This review provides information that understand the important roles of each chemokine axis in breast cancer, probably offering a clue of adjuvant therapeutic options to improve the quality of life and survival for patients with breast cancer.
Keywords
- chemokines
- breast
- cancer
The family of chemokines called chemotactic cytokines are small and secreted proteins that exert cellular signal pathways through cell surface G protein-coupled chemokine receptors [1]. Chemokines consist of 4 subfamilies according to the number of amino acids inserted between the first cysteine (C) motifs as follows: C (XCL1-2), CC (CCL1-28), CXC (CXCL1-17) and CX3C (CX3CL1). Except for orphan chemokines CCL18 and CXCL14, each chemokine recognizes its specific chemokine receptor (XCR1, CCR1-10, CXCR1-8, and CX3CR1) to create the unique chemokine axis as shown in Table 1, which controls angiogenesis, regulates immune network, and modulates cellular functions. In breast cancer, chemokines recruit immune cells into the tumor microenvironment through the chemokine axis between tumor and immune cells [2], developing immune contexture which changes cancer progression and prognosis.
| Chemokine axis family | Chemokine axis subfamily |
|---|---|
| The XCL-XCR axis | The XCL1/2-XCR1 axis |
| The CCL-CCR axes | The CCL3/5/7/8/14/15/16/23-CCR1 axis |
| The CCL2/7/8/13/16-CCR2 axis | |
| The CCL5/7/11/13/14/15/24/26/28-CCR3 axis | |
| The CCL17/22-CCR4 axis | |
| The CCL3/4/5/8/11/14/16-CCR5 axis | |
| The CCL20-CCR6 axis | |
| The CCL19/21-CCR7 axis | |
| The CCL1/16-CCR8 axis | |
| The CCL25-CCR9 axis | |
| The CCL27/28-CCR10 axis | |
| The CXCL-CXCR axes | The CXCL6/7/8-CXCR1 axis |
| The CXCL1/2/3/5/6/7/8-CXCR2 axis | |
| The CXCL4/9/10/11-CXCR3 axis | |
| The CXCL12-CXCR4 axis | |
| The CXCL13-CXCR5 axis | |
| The CXCL16-CXCR6 axis | |
| The CXCL11/12-CXCR7 axis | |
| The CXCL17-CXCR8 axis | |
| The CX3CL-CX3CR axis | The CX3CL1-CX3CR1 axis |
Orphan chemokines: CCL18, CXCL14. XCL-XCR, Lymphotactin-X-C motif chemokine receptor; CCL-CCR, Chemokine (C-C motif) ligand-Chemokine (C-C motif) receptor; CXCL-CXCR, Chemokine (C-X-C motif) ligand-Chemokine (C-X-C motif) receptor.
This review has described functional roles of chemokine axes in breast cancer based on literature data about effects of chemokines and chemokine receptors in cell lines, tumor-bearing animal models, and patients with breast cancer. Molecular subtypes of breast cancer are defined in large part by expression levels of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2), as follows: luminal A (ER+/PR+/HER2-), luminal B (ER+/PR+/HER2+), HER2-enriched (ER-/PR-/HER2+), and basal-like (triple-negative, ER-/PR-/HER2-). Breast cancer cell models present in vitro results of each chemokine and chemokine receptor in breast cancer cell lines, which contain breast subtype cell lines, treatments including antibody (Ab), knockdown (KD), antagonists, overexpression, and inhibitors, cellular functions, and signaling (Table 2, Ref. [3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104]). Breast cancer animal models show in vivo results of each chemokine and chemokine receptor in tumor-bearing animals, which contain breast subtype cell lines, animal models, treatments including Ab, KD, knockout (KO), antagonists, overexpression, and inhibitors, functional parameters, signaling, and immune contexture (Table 3, Ref. [3, 5, 12, 13, 16, 19, 20, 25, 26, 27, 28, 30, 31, 32, 33, 34, 35, 36, 39, 41, 42, 43, 47, 53, 54, 59, 60, 64, 65, 66, 68, 69, 71, 74, 77, 78, 81, 83, 85, 87, 88, 89, 90, 91, 93, 96, 97, 100, 102, 103, 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 115, 116, 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132, 133, 134, 135, 136, 137, 138, 139, 140, 141, 142, 143, 144, 145, 146, 147, 148]). Finally, human breast cancer samples show clinical results of each chemokine and chemokine receptor in patients with breast cancer, which contain biomarker, clinical correlation, immune contexture, prognosis, and overall survival (Table 4, Ref. [5, 8, 9, 13, 14, 17, 23, 32, 34, 35, 36, 43, 44, 47, 49, 50, 52, 53, 62, 64, 74, 82, 88, 94, 98, 100, 101, 102, 107, 111, 117, 131, 140, 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168, 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181, 182, 183, 184, 185, 186, 187, 188, 189, 190, 191, 192, 193, 194, 195, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 207, 208, 209, 210, 211, 212, 213, 214, 215, 216, 217, 218, 219, 220, 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233, 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246, 247, 248]). These results obtained from cell lines, animal models, and human tumor samples contribute to a clear understanding on the critical roles of each chemokine axis in breast cancer progression, probably offering a clue of adjuvant therapeutic options to target the specific chemokine axis using either agonists or antagonists as well as antibodies for patients with breast cancer.
| Chemokines | Cell lines | Treatment | Function | Signaling | References |
| CCL2 | BT549 | KD | Invasion |
MMP9 |
[14, 15] |
| MDA-MB-231 | Ab; KD | Proliferation |
NOTCH |
[16, 18, 24, 26, 27, 28, 29] | |
| MDA-MB-468 | Ab | Invasion |
|||
| MDA-MB-361 | Mammosphere |
NOTCH |
[16] | ||
| BT474 | Ab; KD | Viability |
Ki67 |
[23, 24] | |
| MCF10CA1d | Growth |
PCNA |
[15, 20] | ||
| BT-20 | |||||
| HCC1937 | PCNA | ||||
| MCF10A | Invasion |
Vimentin |
[21] | ||
| DCIS.com | Growth |
E-cadherin |
[22] | ||
| SUM225 | Growth |
||||
| MCF-7 | Viability |
pAkt |
[4, 14, 16, 17, 18, 19] | ||
| T47D | Viability |
||||
| ZR-75-1 | Migration |
[4] | |||
| A3250 | KD | Proliferation |
[30] | ||
| 4T1 (m) | Apoptosis |
Caspase-3 |
[18] | ||
| PyVmT carcinoma cells | KD | Viability |
Caspase-3 |
[23, 31] | |
| Met-1 (m) | Monocytes |
Migration |
[25] | ||
| CCL3 | MCF-7 | Migration |
[4] | ||
| T47D | Migration |
||||
| ZR-75-1 | Migration |
||||
| CCL4 | MCF-7 | Migration |
[4] | ||
| T47D | Migration |
||||
| ZR-75-1 | Migration |
||||
| CCL5 | MDA-MB-231 | Proliferation |
pAkt |
[5, 6, 7] | |
| Trastuzumab-resistant BT-474 | KD | Chemoresistance |
[8] | ||
| MCF-7 | Proliferation |
GLUT-1 |
[4, 6, 9, 10, 11] | ||
| T47D | Migration |
||||
| ZR-75-1 | Migration |
||||
| Tamoxifen-resistant MCF-7 | KD | Chemoresistance |
pSTAT3 | ||
| 4T1 (m) | KD | Proliferation |
[12] | ||
| CCL8 | MDA-MB-231 | Migration |
[13] | ||
| MDA-MB-468 | |||||
| E0771 | |||||
| CCL18 | MDA-MB-231 | Migration |
[34] | ||
| CCL19 | MDA-MB-231 | Proliferation |
E‐cadherin |
[38] | |
| MCF-7 | |||||
| PyVmT-CCR7 | Proliferation |
[39] | |||
| CCL20 | MDA-MB-231 | Ab; KD | Viability |
ALDH |
[34, 35, 36, 37] |
| EV | Viability |
uPA activity | |||
| BT549 | Invasion |
MMP2 | |||
| HCC38 | |||||
| SUM159 | Chemoresistance |
pPKCζ | |||
| CCL21 | MDA-MB-231 | Migration |
TAP-1 |
[39, 40, 41, 42, 43, 44, 45, 46] | |
| SKBR-3 | HLA-1 | ||||
| HCC1428 | Migration |
E-cadherin | |||
| MCF7 | Migration |
HLA-1 | |||
| MCF10A | Cell spreading |
||||
| 4T1 (m) | Migration |
||||
| PyVmT-CCR7 | Proliferation |
||||
| CCL25 | MDA-MB-231 | Proliferation |
MMP1/9/11/13 |
[48, 49] | |
| MCF7 | Migration |
||||
| CCR27 | MDA-MB-231 | Migration |
pErk |
[50] | |
| CXCL1 | MDA-MB-231 | Migration |
Vimentin |
[4, 60, 61] | |
| BT549 | |||||
| MCF-7 | Vimentin | ||||
| T47D | Migration |
||||
| ZR-75-1 | Migration |
||||
| BCSC-105/BCSC-608 | Proliferation |
CXCR7 |
[62] | ||
| Ab | Proliferation |
||||
| SUM149 | KD | Invasion |
Fibronectin |
[58] | |
| 4T1 (m) | Migration |
Vimentin |
[60] | ||
| PyMT cells | Proliferation |
[63] | |||
| Ab | MSC-CM induced Migration |
||||
| CXCL2 | SUM149 | KD | Invasion |
Fibronectin |
[58] |
| CXCL3 | SUM149 | KD | Invasion |
Fibronectin |
[58] |
| MCF-7 | Migration |
[4] | |||
| T47D | Migration |
||||
| ZR-75-1 | |||||
| CXCL4 | MDA-MB-231 | CXCL12-induced migration |
[67] | ||
| CXCL447–70 | Proliferation |
[68] | |||
| CXCL5 | PyMT cells | Proliferation |
[63] | ||
| Ab | MSC-CM induced Migration |
||||
| CXCL7 | MDA-MB-231 | Migration |
pFak |
[53] | |
| MCF-7 | Migration |
VEGF |
[4, 51, 52] | ||
| T47D | Migration | ||||
| ZR-75-1 | Migration | ||||
| MCF10A | CXCL7; EV | Invasion | |||
| CXCL8 | Patient-derived BC cells | Mammosphere formation |
[4, 54, 55, 56, 57, 58] | ||
| MCF7/HER2-18 | pHER2 | ||||
| MDA-MB-231 | Invasion |
||||
| KD | Proliferation |
Cyclin D1 | |||
| SK-Br-3 | Invasion |
||||
| BT549 | Invasion |
||||
| KD | Proliferation |
Cyclin D1 | |||
| BT20 | Invasion |
||||
| MCF-7 | CXCL8; EV | Migration |
|||
| Ab | Chemoresistance |
||||
| T47D | Migration |
||||
| ZR-75-1 | Migration |
||||
| SUM149 | KD | Invasion |
Fibronectin | ||
| CXCL9 | MDA-MB-231 | Migration |
MMP9 |
[69, 70] | |
| 66.1 (m) | |||||
| CXCL10 | MDA-MB-231 | Migration |
ALDH1+ cells |
[70, 71] | |
| MCF-7 | [4] | ||||
| T47D | Migration |
||||
| ZR-75-1 | |||||
| 66.1 (m) | Migration |
[69] | |||
| 4T1 (m) | p65 |
[72] | |||
| CXCL11 | MDA-MB-231 | Migration |
ALDH1+ cells |
[70, 71] | |
| MCF-7 | Proliferation |
[73] | |||
| 66.1 (m) | Migration |
[69] | |||
| CXCL12 | MDA-MB-231 | Migration |
RhoA |
[40, 43, 46, 75, 76, 77, 78, 79, 80, 81, 82] | |
| EV | Proliferation |
||||
| Ab | Transendothelial migration |
||||
| SUM-159 | Migration |
RhoA | |||
| MDA-MB-468 | Migration |
RhoA | |||
| MCF-7 | Proliferation |
[73, 78, 80, 83, 84, 85, 86, 87, 88] | |||
| KD; Ab | Proliferation |
||||
| SKBR3 | Proliferation |
pHER2 | |||
| T47D | Migration |
||||
| BT474/BT547 | Migration |
||||
| SUM-149 | Migration |
||||
| 4T1 (m) | Proliferation |
pSTAT3 | |||
| MVT1 (m) | Migration |
||||
| CXCL13 | MDA-MB-231 | CXCL13 ligand + CXCR5 EV | Migration |
Vimentin |
[98] |
| T47D | |||||
| MDA-MB-231 | Ab | Proliferation |
pErk |
[99] | |
| CXCL14 | MDA-MB-231 | EV | Proliferation |
[100] | |
| CXCL16 | MDA-MB-231 | Migration |
[101] | ||
| MCF-7 | |||||
| CXCL17 | MDA-MB-231 | KD | Proliferation |
[102] | |
| SKBR3 | Migration |
||||
| MCF-7 | pErk | ||||
| CX3CL1 | T47D | Proliferation |
pErk |
[104] | |
| XCR1 | MDA-MB-231 | EV | Proliferation |
pErk |
[3] |
| CCR2 | MDA-MB-231 | Antagonist | Migration |
[19] | |
| MCF-7 | |||||
| T47D | |||||
| MCF10CA1d | KO | Growth |
ALDH activity |
[20] | |
| SUM225 | EV | CCL2-induced growth, invasion |
E-cadherin |
[22] | |
| DCIS.com | KO | CCL2-induced growth, invasion |
PCNA | ||
| CCR4 | MDA-MB-231 | EV | Proliferation |
[32] | |
| KD | Proliferation |
||||
| CCR5 | MDA-MB-231 | EV | Proliferation |
[5, 6, 7, 33] | |
| Antagonist | CCL5-inuced calcium flux |
CCL5-induced pAkt | |||
| CCR5 EV MDA-MB-231 | KD | Migration |
|||
| MCF-7 | Antagonist | CCL5-inuced glucose uptake |
|||
| Hs578T | CCL5-inuced calcium flux |
||||
| SUM-159 | Invasion |
||||
| CCR7 | MDA-MB-231 | Ab; KD | Proliferation |
VEGF-C |
[38, 45, 46, 47] |
| MCF-7 | CCL19 induced proliferation, migration & invasion |
||||
| 4T1 (m) | Proliferation |
EpCAM | |||
| CCR9 | MDA-MB-231 | Ab | CCL25-induced proliferation, migration, invasion |
CCL25-induced MMP1/9/11/13 |
[48, 49] |
| CCR10 | MDA-MB-231 | KD | CCL27-induced migration/invasion |
CCL27-induced pErk and MMP7 |
[50] |
| CXCR1 | SUM159 | Ab; Antagonist | Viability |
ALDH |
[59] |
| CXCR2 | Patient-derived BC cells | CXCR1/2 antagonist | CXCL8-induced mammosphere formation |
[56] | |
| MCF7/HER2-18 | |||||
| MCF-7 | EV | Colony |
pAkt |
[64] | |
| BT474 | |||||
| SKBR3 | KD | Colony |
pAkt | ||
| MDA-MB-231 | pAkt | ||||
| MCF10A-CXCL17 | Inhibitor | Invasion |
VEGF |
[51] | |
| Cl66 (m) | KD | Proliferation |
[65, 66] | ||
| 4T1 (m) | |||||
| PyMT cells | Proliferation |
[63] | |||
| AB; Inhibitor | MSC-CM induced migration |
||||
| CXCR3 | MDA-MB-231 | CXCR3B EV | CSC population |
ALDH1+ cells |
[70, 71] |
| CXCR3B KD | CSC population |
ALDH1+ cells | |||
| Ab | CXCL9/10/11-induced migration |
||||
| 66.1 (m) | Antagonist AMG487; KD | Proliferation |
[69] | ||
| 4T1 (m) | Proliferation |
p65 |
[72, 74] | ||
| CXCR4 | MDA-MB-231 | KD | Proliferation |
[46, 75, 77, 78, 80, 89, 90] | |
| Antagonist; Ab | CXCL12-induced migration |
||||
| MCF-7 | Constitutively active CXCR4 | Cadherin11 |
[83, 91, 92, 93] | ||
| KD; Ab; Antagonist | Proliferation |
||||
| MCF7 (TAM-R) | Antagonist; KD | Proliferation |
|||
| BT-474 | Ab | CXCL12-induced invasion |
[94] | ||
| MCF10A | EV | Invasion |
pErbB2 |
[95] | |
| 4T1 (m) | KD | Proliferation |
[96] | ||
| AMD3465 | Migration |
pSTAT3 |
[97] | ||
| MVT1 (m) | AMD3100 | Migration |
[87] | ||
| CXCR5 | MDA-MB-231 | CXCR5 Ab in CXCL13 ligand + CXCR5 EV cells | Migration |
Vimentin |
[98] |
| T47D | |||||
| CXCR7 | MDA-MB-231 | EV | Proliferation |
[73] | |
| MCF7 | KD | Proliferation |
Cyclin B1 | ||
| BT474 cells | Proliferation |
||||
| 4T1 (m) | KD/Inhibitor (CCX771) | CXCL12-induced migration |
pSTAT3 |
[88] | |
| CXCR8 | 4T1 (m) | MCF7 | CXCL17-induced pErk |
[102] | |
| CX3CR1 | MDA-MB-436 | EV | pErk |
[103] |
BCSC, breast cancer stem cell; CM, conditioned media; CSC, cancer stem cells;
EpCAM, epithelial cell adhesion molecule; ERO1-
| Chemokines | Cell lines | Species & model | Treatment | Function | Signaling | Immune contexture | References |
| CCL2 | MDA-MB-231 | Female SCID; Orthotropic | Ab | Angiogenesis |
TAM |
[26, 28, 111] | |
| shCCL2 | |||||||
| Intratumoral siCCL2 | Tumor growth |
PCNA |
M2 Mφ | ||||
| TAM | |||||||
| FVB female mice; Lung metastasis | Inflammatory Mo |
[25] | |||||
| MCF10CA1d | Female athymic nude mice; Orthotropic | CCL2 EV fibroblasts | Tumor growth |
[20] | |||
| CCL2 KO fibroblasts | Tumor growth |
M2 Mφ | |||||
| Ab | Tumor growth |
Blood & tumoral CCL2 |
TAM |
[114] | |||
| A3250 | SCID mice; Orthotopic | KD | Tumor growth |
Lung & blood Mo |
[30] | ||
| MCF-7 | Female athymic mice; Orthotropic | Ab | Tumor growth |
[106] | |||
| Zebrafish embryos | Cell dissemination |
||||||
| 4T1 (m) | Female mice; Orthotopic | L-RNA aptamer inhibiting CCL2 | Angiogenesis |
TAM |
[113] | ||
| KO mice | Tumor growth |
TAM |
[144, 145] | ||||
| Ab | Tumor growth |
[112] | |||||
| Discontinued CCL2 Ab | Metastasis |
Ki67 |
Metastatic & tumoral Mo |
[115] | |||
| KD | Bone/lung metastasis |
[116] | |||||
| Aerobic exercise | Tumor growth |
Plasma CCL2 |
[146] | ||||
| XP265922 primary BC + CAF265922 cells | Female NOD/SCID/IL2R |
Fibroblast-specific CCL2 KD | Tumor growth |
NOTCH1 |
[16] | ||
| 67NR (m) | BALB/c mice; lung metastasis | Intranasal CCL2 | Tumor growth |
CD45+ cells |
[147] | ||
| Met-1 (m) | Female nude mice; lung metastasis | Ab | Metastasis |
Inflammatory Mo |
[25] | ||
| AT-3 (m) | C57BL/6 mice; Orthotopic | KO mice | Tumor growth |
Splenic MDSCs |
[112] | ||
| Ab | Tumor growth |
||||||
| PyMT tumor cells | FVB/N mice; Orthotopic | Ab | Tumor growth |
TAM |
[106] | ||
| Spontaneous models | Her2/neu-driven mammary carcinoma | KO | Survival |
[27] | |||
| CCL3 | MDA-MB-231 | Nude mice; Orthotropic | KO Mo | Tumor growth |
[105] | ||
| E0771 (m) | Mice; Orthotropic | Metastasis |
Mφ | ||||
| Met-1 (m) | FVB mice; Orthotropic | Ab | Metastasis |
||||
| CCL4 | D2F2/E2 tumors (m) | s.c. | pDNA-CCL4 vaccination | Tumor growth |
[121] | ||
| CCL5 | MCF-7 | Zebrafish embryos | Ab | Cell dissemination |
[106, 107] | ||
| PyMT tumor cells | FVB/N mice; Orthotopic | Ab; KO mice | Tumor growth |
TAM | |||
| 4T1 (m) | Female mice; Orthotopic | Aerobic exercise | Tumor growth |
Plasma CCL5 |
[146] | ||
| Ab; KO mice | Tumor growth |
CD4 |
[108] | ||||
| KD | Tumor growth |
Mφ |
[12] | ||||
| SCID mice | Tumor growth |
||||||
| 410.4 (m) | Antagonist Met-CCL5 | Tumor growth |
Mφ |
[148] | |||
| TAN primary tumor cells | TAN mice; Orthotopic | Primary tumor growth |
Mφ |
[109] | |||
| CCL8 | MDA-MB-231 | Nude mice; s.c. | Ab | Invasion |
[13, 110] | ||
| MCF10.DCIS | SCID mice; Orthotopic | KO mice | Delay in the latency for tumor onset | M2 Mφ | |||
| E0771 (m) | Female mice; s.c. | Ab; KO mice | Survival |
Vimentin |
TAM | ||
| CCL17 | 4T1 (m) | Orthotopic | Chemotoxin | Lung metastasis |
[120] | ||
| TARC-PE38 | |||||||
| CCL18 | MDA-MB-231 | SCID mice; Orthotopic | Intratumoral injection | Angiogenesis |
[34, 122] | ||
| BT-474 | |||||||
| CCL20 | MDA-MB-231 | Nude mice; Cardiac | Ab | Metastasis |
[35] | ||
| Nude mice; Orthotropic | EV | Tumor growth |
[36] | ||||
| CCL21 | SKBR-3 | Nude mice; s.c. | CCL21 + human lymphocytes (i.v.) | Tumor growth |
[42] | ||
| MCF7 | |||||||
| 4T1 (m) | Mut-CCL21 | Tumor growth |
[41] | ||||
| CCL27 | 4T1 (m) | Orthotropic | CCL27-PE38 | Lung metastasis |
[120] | ||
| CXCL1 | MDA-MB-231 | SCID mice, Orthotropic | THP1-M2 or THP1-M2/shCXCL1 cells | Tumor growth & lung metastasis: |
Vimentin |
[60] | |
| Chemokinostatin-1 | Tumor growth |
[123] | |||||
| PyMT mammary cancer cells | MMTV-PyMT+/- mice | TAM-shCXCL1, shCXCL1/2 | Tumor growth |
CD11b(+)Gr1(+) myeloid cells |
[60, 124] | ||
| CXCL4 | MDA-MB-231 | SCID mice; s.c. | CXCL447–70 | Tumor growth |
Tumoral mRNA levels: F4/80 |
[68] | |
| CXCL7 | MDA-MB-231 | SCID mice; Orthotropic | Ab | Tumor growth |
M2 Mφ |
[53] | |
| CXCL8 | MDA-MB-231 | Nude mice; s.c. | Angiogenesis |
[54] | |||
| SK-BR3 | |||||||
| CXCL9 | 4T1 (m) | Balb/c mice; Orthotropic | Ab | Tumor growth |
[126] | ||
| CXCL12 | Human BC | PDX | HER2 |
[127] | |||
| MDA-MB-231 | Lung metastasis | EV | Lung metastasis |
[81, 128] | |||
| Orthotropic | Tumor growth |
||||||
| Ab | Lymphatic vessel |
||||||
| MCF-7 | SCID; Orthotopic | ICI-inhibited tumor growth |
[131] | ||||
| Ab | Tumor growth |
[83] | |||||
| MVT1 (m) | Fibroblast CXCL12 cKO mice; Orthotopic | Tumor growth |
[87] | ||||
| Lung metastasis | Lung metastasis |
||||||
| 4T1 (m) | Female mice; Orthotopic | WT compared to mutant EV | Tumor growth |
DC in lymph nodes |
[129, 130] | ||
| CXCL12-EV stromal cells | Lung metastasis |
||||||
| MTLn3 (r) | SCID mice; Orthotopic | EV | Tumor growth |
TAM |
[132] | ||
| CXCL14 | MDA-MB-231 | SCID mice; Orthotopic | EV | Tumor growth |
[100] | ||
| 4T1 (m) | Female Balb/c mice; Orthotopic | EV | Tumor growth |
Myeloid cells |
[140] | ||
| CXCL17 | 4T1 (m) | Female Balb/c mice; Orthotopic | EV | Tumor growth |
[102] | ||
| CX3CL1 | Tg-neu mice | Intratumoral injections of Ad-CX3CL1 | Palpable tumors |
[104] | |||
| KO in Tg-neu mice | Delayed mammary tumor onset; Tumor number |
||||||
| KO in Tg-PyMT mice | Mammary tumor onset |
||||||
| XCR1 | MDA-MB-231 | EV | Tumor growth |
[3] | |||
| CCR1 | E0771 (m) | Mice; Orthotropic | KO Mo | Metastasis |
Mφ |
[105] | |
| CCR2 | SUM225 | Female NOD/SCID | EV | Invasion |
Ki67 |
CCL2+ fibroblasts |
[118] |
| MCF10CA1d | Female athymic nude mice; Orthotropic | KO | Tumor growth |
M2 Mφ |
[20] | ||
| MCF-7 | Antagonist | E2-induced tumor growth, Angiogenesis, Metastasis |
PCNA |
[19] | |||
| hDCIS cells | NOD-SCID IL-2r |
CCR2+ cells | Tumor formation |
PCNA |
[117] | ||
| KD | Tumor growth |
PCNA |
[118] | ||||
| 4T1 (m) | Female mice; Orthotopic | KD | Tumor growth |
M2 Mφ |
[20] | ||
| Aerobic exercise | Tumor growth |
Plasma CCR2 |
[146] | ||||
| PyVmT mammary carcinoma cells | FVB mice; Intraductal injection | KD | Tumor growth |
PCNA |
F4/80+ Mφ |
[31] | |
| E0771 (m) | Female CCR2 KO mice | KO mice | Tumor growth |
CD4 |
[119] | ||
| Mice; Orthotropic | KO Mo | Metastasis |
Mφ |
[105] | |||
| AT-3 (m) | C57BL/6 mice; Orthotopic | KO mice | Tumor growth |
Splenic MDSCs |
[112] | ||
| Spontaneous models | Her2/neu-driven mammary carcinoma | KO | Survival |
[27] | |||
| Antagonist CCX872 | |||||||
| CCR4 | MDA-MB-231 | Nude mice; Orthotropic | EV | Tumor growth |
[32] | ||
| KD | Tumor growth |
||||||
| CCR5 | MDA-MB-231 | SCID mice; lung metastasis | Antagonist maraviroc | The number and the size of pulmonary metastases |
[5] | ||
| Nude rats | Bone metastasis |
[33] | |||||
| 4T1 (m) | Female mice; Orthotopic | Aerobic exercise | Tumor growth |
Plasma CCR5 |
[146] | ||
| CCR7 | PyVmT-CCR7 | FVB mice; Orthotopic | Compared to CCR7 negative cells | Tumor growth |
Survival |
[39] | |
| 4T1 (m) | Female mice; Orthotopic | KD | Tumor growth |
[47] | |||
| CXCR1 | SUM159 | SCID mice; Orthotopic | Antagonist | Tumor growth |
ALDH |
[59] | |
| MDA-MB-453 | Tumor growth |
||||||
| HCC1954 | Tumor growth |
||||||
| CXCR2 | MDA-MB-231 | Female mice; Orthotopic | KD | Tumor growth |
[64] | ||
| PyMT mice | KO mice | Tumor growth |
Killing ability of Cxcr2 KO TANs |
TAN |
[125] | ||
| Cl66 (m) | Female mice; Orthotopic | KD | Tumor growth |
PCNA |
[65, 66] | ||
| CXCR3 | MDA-MB-231 | Lung metastasis | Antagonist | Metastasis |
[71] | ||
| 66.1 (m) | Lung metastasis (Antagonist-treated cells); Orthotopic | Antagonist AMG487 | Tumor growth |
[69] | |||
| 4T1 (m) | Orthotopic; Lung metastasis | Antagonist; KD; KO mice | Tumor growth |
Splenic CD3/CD4/CD8 |
[74] | ||
| CXCR4 | Human BC | HER2 BC PDX | Inhibitors: AMD3100; TN14003 | Tumor growth |
Ki-67 |
[127] | |
| TNBC PDX | Tumor growth |
Ki-67 |
|||||
| MDA-MB-231 | SCID mice; lung metastasis | CXCR4 Ab; KD; Inhibitors | Metastasis |
[43, 77, 89, 134] | |||
| SCID mice; Orthotopic | KD; AMD3100; CTCE-9908 | Tumor growth |
Ki-67 |
[78, 90, 135, 136] | |||
| MTLn3 | SCID mice; Orthotopic; | EV | Invasion |
VEGFA |
[133] | ||
| MCF-7 | Female SCID mice; Orthotopic | EV | Tumor growth |
E-cadherin |
[91, 131] | ||
| KD/Ab/AMD3100 | Tumor growth |
[83, 93] | |||||
| MCF7 (TAM-R) | AMD3100 | Tumor growth |
ABCG2 |
||||
| 4T1 (m) | Orthotopic; Lung metastasis | KD | Tumor growth |
[96] | |||
| Orthotopic | Vaccinia viruses | Tumor growth |
CD31 |
Bone marrow-derived endothelial and myeloid cells |
[139] | ||
| Orthotopic | Inhibitors | Tumor growth |
pAkt |
TAM |
[85, 97, 137] | ||
| E0771 (m) | Orthotopic | AMD3100 | Metastasis |
CD8 | |||
| MMTV-PyMT | Inhibitor: CTCE-9908 | Tumor growth |
VEGF |
[138] | |||
| CXCR6 | 4T1 (m) | Mice; s.c.; Orthotopic | KO mice | Tumor growth |
[141] | ||
| CXCR7 | MTLn3 | SCID mice; Orthotopic | EV | Invasion |
VEGFA |
[133] | |
| MDA-MB-435 | SCID mice; Orthotopic | EV | Tumor growth |
[88, 130, 142] | |||
| 4T1 (m) | Mice; s.c.; Orthotopic | KD/Inhibitor (CCX771) | Tumor growth |
pSTAT3 |
TAM | ||
| AT-3-FL (m) | Orthotopic; Lung metastasis | Endothelial CXCR7 cKO | Tumor growth |
[143] | |||
| E0771 (m) | |||||||
| CX3CR1 | MDA-MB-231 | SCID mice; Intracardiac | KO mice | Bone metastasis |
[103] | ||
| MDA-MB-436 | EV | Bone metastasis |
Ab, antibody; ALDH1, aldehyde dehydrogenase 1; CAF, carcinoma-associated
fibroblasts; CTC, circulating tumor cells; EV, expression vector; hDCIS, human
ductal carcinoma in situ; HTRA2, high temperature requirement protein A2; ICI,
ICI182780; KD, knockdown; KO, knockout; m, mouse; MDSC, myeloid-derived
suppressor cells; Mo, monocytes; PCNA, proliferating cell nuclear antigen; PDX,
patient-derived xenografts; TAM, tumor-associated macrophages; TAN,
tumor-associated neutrophils; TARC-PE38, TARC fusion with a truncated toxin PE38;
TIM, tumor-infiltrating M
| Chemokines | Biomarker | Clinical correlation | Immune contexture | Prognosis | Overall survival | References |
| CCL1 | Invasive; ER- | Grade | Treg |
[168, 175] | ||
| CCL2 | BC; Low differentiation; Invasive ductal BC; Early relapse; ER-; PR-; CAFs; BL; Claudin-low; LB; Advanced | Postmenopausal; Lymph node involvement; Grade; Tumor size; Nodal status; Angiogenic; AP-1+; Ki67 |
TAM |
X in stromal CCL2 | [14, 17, 23, 111, 117, 150, 153, 159, 161, 162, 163, 164, 165, 166, 167, 168, 169, 170, 171, 172] | |
| CCL3 | BC; Inflammatory BC; ER-; PR- | Grade; Ki67 |
[149, 150, 151, 152, 153, 154] | |||
| CCL4 | Inflammatory BC; ER-; Metastasis of LB | Grade; Ki67 |
[149, 151, 153, 155, 157, 176] | |||
| CCL5 | TNBC; Advanced; Inflammatory BC; BL; HER2; ER-; PR- | Stage; Histological grade; LN+; Microvessel density | CD163+ Mφ infiltration | O in BC, LA, LB | [5, 8, 9, 107, 152, 155, 156, 157, 158, 247, 248] | |
| CCL7 | BC; Poorly differentiated; TNBC; AA | Grade; Ki67 |
[153, 157, 159] | |||
| CCL8 | TNBC; AA; ER- | Grade; Ki67 |
X | [13, 153, 157] | ||
| CCL11 | HER2 | O | [157] | |||
| CCL13 | HER2 | Ki67 |
[153] | |||
| CCL14 | O | [155, 160] | ||||
| CCL15 | O | [151] | ||||
| CCL16 | X in LA; O in TNBC | [156] | ||||
| CCL17 | TNBC; AA | Ki67 |
X | [151, 153, 157] | ||
| CCL18 | BC, Advanced stage; Metastatic BC | Lymph node metastasis | X | [34, 151, 159, 168] | ||
| CCL19 | Aggressive | O; X in LA | [151, 155, 156, 157, 173, 178] | |||
| CCL20 | TNBC; AA; ER- | Ki67 |
[35, 36, 153, 157] | |||
| CCL21 | Metastatic BC | O | [151, 155, 157, 160, 173, 179] | |||
| CCL22 | BC; HER2 | Low grade | O | [151, 155, 157, 159, 175] | ||
| CCL23 | Grade | O | [153, 157] | |||
| CCL24 | [151, 155] | |||||
| CCL25 | TNBC; AA | [157] | ||||
| CCL26 | Inflammatory BC | [149] | ||||
| CCL27 | ||||||
| CXCL1 | BC; TNBC; ER-; Metastasis of BC | Grade, in stromal CXCL1; Ki67 |
CD133 |
X, in stromal CXCL1 | [62, 153, 188, 201, 202, 203, 204] | |
| CXCL2 | Metastasis of BC | O | [187, 188, 190, 203] | |||
| CXCL3 | Aggressive BC; Metastasis of BC | X | [187, 188, 189, 190, 203, 205] | |||
| CXCL5 | BC; ER- | Low metastasis in BC, LB | [153, 161, 168, 176, 188] | |||
| CXCL6 | ER-; Metastasis of BC | [187, 188] | ||||
| CXCL7 | Stage III | [53, 155, 187, 189, 190] | ||||
| CXCL8 | BC; Inflammatory BC; TNBC; Advanced stage; LA; LB; ER-; HER2+; PR-; CAFs; ER- | Lymph node metastasis Angiogenic; Grade; Stage; AP-1+; Ki67 |
CD68 |
X | [149, 150, 153, 155, 159, 166, 176, 187, 188, 189, 191, 192, 193, 194, 195, 196, 197, 198, 199, 200] | |
| CXCL9 | BC, TNBC, Low proliferative, Lymph node negative; HER2; ER- | Grade; Ki67 |
O | [153, 155, 159, 160, 187, 189, 190, 203, 209] | ||
| CXCL10 | BC, Poor differentiation; HER2; ER-; HR- | Stage; Grade; Ki67 |
TIL |
O | [153, 159, 187, 190, 203, 210, 211, 212] | |
| CXCL11 | BC, TNBC; HER2; ER- | Grade; Ki67 |
[153, 159, 203] | |||
| CXCL12 | BC, BL; Subtypes |
Stage; Grade; lymph node metastasis | Treg |
O | [82, 155, 160, 168, 187, 189, 190, 203, 216, 217, 218, 219, 220, 221, 222] | |
| CXCL13 | BC; ER-; Metastatic BC | Lymph node metastasis with CXCR5 coexpression; Ki67 |
O | [52, 98, 155, 160, 168, 187, 189, 190, 203, 209] | ||
| CXCL14 | Lymph node metastasis |
[100, 140, 155, 187, 189] | ||||
| CXCL16 | Stage in N-terminal CXCL16 | X in HER2 | [101, 156] | |||
| CXCL17 | ER- | Ki67 |
[102, 153] | |||
| CX3CL1 | Inflammatory BC; LB | Grade; Stage; Tumor size; Lymph node metastasis; PR+; Ki67 |
Stromal CD8 |
[149, 153, 245, 246] | ||
| CCR2 | Invasive ductal BC | [117, 173, 174] | ||||
| CCR4 | Lymph node metastasis; HER2 expression | [32, 173] | ||||
| CCR5 | BL; HER2 | [5] | ||||
| CCR6 | Pleura metastasis; Aggressive | [177, 178] | ||||
| CCR7 | BC; Metastatic BC; HER2+; TNBC; LB | Lymph node metastases |
CD68 |
[43, 44, 47, 177, 178, 179, 180, 181, 182, 183, 184, 185, 186] | ||
| CCR9 | Poorly differentiated | [49] | ||||
| CCR10 | Capsular invasion; Stage; Lymph node metastases | [50] | ||||
| CXCR1 | Invasive BC | [174] | ||||
| CXCR2 | High-grade; TNBC; ER-; PR-; Relapse |
TILs |
X in C1208T variation | [64, 174, 200, 206, 207, 208] | ||
| CXCR3 | ER- | Grade; Tumor size | [74, 213, 214, 215] | |||
| CXCR4 | BC; LA; LB; BL; HER2; Locally advanced BC; TNBC; Atypical ductal hyperplasia; Ductal carcinoma in situ; Invasive BC | Metastasis of lymph node & liver; Distant metastasis; Recurrence in HER2-, TNBC; Grade; ER-; PR-; tumor size & advanced TNM stage in TNBC | CXCR4+ Treg |
[43, 94, 131, 174, 177, 181, 182, 185, 213, 214, 216, 217, 218, 219, 221, 223, 224, 225, 226, 227, 228, 229, 230, 231, 232, 233, 234, 235, 236, 237, 238, 239, 240, 241, 242, 243] | ||
| CXCR5 | Lymph node metastasis; Stage | [199] | ||||
| CXCR6 | Stage | [101] | ||||
| CXCR7 | ER-; PR-; TNBC; ER+; PR+ | TNM stage; Grade | [88, 218, 244] | |||
| CXCR8 | Grade; Ki67 |
[102] | ||||
| CX3CR1 | Brain metastasis | [177] |
O, good; X, Poor; AA, African Americans; CAFs, cancer-associated fibroblasts;
EA, European Americans; ER-BC, ER-negative breast cancer; PD-1, programmed cell
death 1; TILs, tumor-infiltrating lymphocytes; BL, Basal-like; HER2, Human epidermal growth factor receptor 2; LB, Luminal B; LA, Luminal A; ER, Estrogen receptor; PR, Progesterone receptor; TNBC, Triple-negative breast cancer; LN+, Lymph node positive; NK, Natural killer cells; FOXP3, Forkhead box P3; TNM, Tumor-node-metasta.
Studies on the XCL1/2 in cell lines and animal models and human breast cancer samples are lacking. The XCR1 overexpression in MDA-MB-231 cells reduced cell proliferation and colony formation but increased migration and invasion. The XCR1-mediated signaling shows reduced phosphorylated extracellular signal-regulated kinase (pErk), phosphorylated Jun N-terminal kinase (pJNK), phosphorylated p38 (pp38), BH3-interacting domain death agonist (Bid), phosphorylated protein kinase B (pAkt), phosphorylated mammalian target of rapamycin (pmTOR), phosphorylated p70 ribosomal S6 kinase (pP70S6K), and phosphorylated eukaryotic translation initiation factor 4E binding protein 1 (p4EBP1) but induced microtubule-associated protein 1A/1B-light chain 3 (LC3) (Table 2) [3]. Animal models bearing XCR1-expressed breast cancer cells reduced tumor growth (Table 3) [3]. Although studies on XCR1 in patients with breast cancer are lacking, the XCL1/2-XCR1 axis shows a benefit for breast cancer, probably reducing tumor growth.
CCL3 induced migration in MCF-7 and ZR-75-1 cells but did not change the
migration of T47D cells (Table 2) [4], probably depending on cellular expression
levels of CCR1. Blocking CCL3 reduced metastasis and treatment of CCL3 KO
monocytes also decreased metastasis but caused no change in tumor growth,
reducing infiltration of M
In cell models (Table 2), CCL2 increased mammosphere, cell viability, migration,
invasion, and G2/M phase in cell cycle and decreased apoptosis in parallel with
increased
CCL5, CCL7, CCL13, CCL14, and CCL15 are described in sections of the CCL3/5/7/8/14/15/16/23-CCR1 axis and the CCL2/7/8/13/16-CCR2 axis. Studies on CCL11, CCL24, CCL26, and CCL28 in cell and animal models for breast cancer are lacking. CCL11 is highly expressed in HER2 subtype and shows a good prognosis (Table 4) [157]. CCL24 shows a poor survival in patients with breast cancer (Table 4) [151, 155]. CCL26 is highly expressed in inflammatory breast cancers (Table 4) [149]. Studies on CCL28 and CCR3 in cell and animal models for breast cancer and patients with breast cancer are lacking.
Studies on CCL17 in cell models for breast cancer are lacking and chemotoxin TARC-PE38 for CCL17 reduced lung metastasis in animal models (Table 3) [120]. CCL17 shows a poor prognosis and survival in patients with breast cancer, particularly African Americans. CCL17 is highly expressed in TNBC and African Americans and is related to the induced Ki67 (Table 4) [151, 153, 157]. Studies on CCL22 in cell and animal models for breast cancer are lacking. CCL22 is highly expressed in breast cancers and HER2 subtypes and is related to low grade, showing a good prognosis and unchanged or good survival (Table 4) [151, 155, 157, 159, 175]. In cell models, CCR4 expression increased CCL17-induced migration with no change in proliferation, while CCR4 KD decreased CCL17-induced migration with no change in proliferation (Table 2) [32]. In animal models, CCR4 overexpression increased tumor growth, lung metastasis, and angiogenesis, while CCR4 KD decreased these effects (Table 3) [32]. CCR4 has controversial survival effects between patients with breast cancer. Expression levels of CCR4 are related to lymph node metastasis and HER2 expression (Table 4) [32, 173]. A clinical trial (NCT06320392) is designed to explore whether CCR4-NOT Complex Subunit 7 contributes to metastasis in metastatic BC patients through NK cell resistance (https://clinicaltrials.gov/search?cond=breast%20cancer&intr=ccr4).
CCL3, CCL5, CCL8, CCL11, CCL14, and CCL16 are described in sections of the
CCR1/CCR2/CCR3 axis. CCL4 induced migration in MCF-7 and ZR-75-1 cells but had no
effects on migration in T47D cells (Table 2) [4], probably depending on cellular
expression levels of CCR5. The pDNA-CCL4 vaccination increased tumor growth and
tumor rate in mice bearing D2F2/E2 tumors (Table 3) [121]. CCL4 has a good
survival in patients with breast cancer. CCL4 is highly expressed in inflammatory
breast cancers and ER negative cancers and is related to metastasis of LB subtype
and grade (Table 4) [149, 151, 153, 155, 157, 176]. In cell models (Table 2), CCR5
expression had no effects on proliferation, but CC5 KD and antagonist reduced
proliferation, colony formation, migration, invasion, and CCL5-inuced calcium
flux, proliferation, glucose uptake, and intracellular ATP/pyruvate/G6P in
parallel with decrease of CCL5-induced pAkt, pmTOR, and pGSK-3
Although PITPNM3 is reported as a specific receptor for CCL18 [34], further studies require clarifying functional roles of chemokine receptor based on the similarity of CCR1-10. CCL18 induced migration in MDA-MB-231 cells (Table 2) [34] and intratumoral injection of CCL18 increased angiogenesis, lung/liver metastasis, vascular invasion, and lung weight (Table 3) [34, 122]. CCL18 is highly expressed in breast cancers, advanced stage cancers and metastatic breast cancers and is related to metastasis and lymph node involvement. CCL18 shows a poor prognosis and survival in patients with breast cancer, particularly with higher CCL18-positive TAM (Table 4) [34, 151, 159, 168].
CCL20 expression and treatment increased viability, migration, invasion, colony
formation, chemoresistance, stemness and tumorsphere in parallel with increased
uPA activity, MMP1, MMP2, MMP9, RANKL, OPG, ALDH, NANOG, OCT4, SOX2,
pPKC
CCL19 increased proliferation, migration, and invasion in parallel with
increased N-cadherin, vimentin, pAkt and MMP2/9 and decreased E-cadherin (Table 2) [38]. Although CCL19 induced migration in PyVmT-CCR7 cells, it has no effects
on proliferation (Table 2) [39]. Studies on CCL19 in animal models for breast
cancer are lacking. CCL19 is related to aggressive status and shows increased
risks in LA subtype, but a good prognosis and survival in patients with breast
cancer. Interestingly, patients with ER positive cancers showed a good survival
with plasma levels of CCL19 but a poor survival with tumoral levels of CCL19
(Table 4) [151, 155, 156, 157, 173, 178]. Although CCL21 had no effects on proliferation, it
increased migration, invasion, and colony formation and decreased apoptosis in
parallel with increased TAP-1, slug, vimentin, N-cadherin, pAkt, pErk, VEGF, and
BclX and decreased TGF-
Studies on CCL1 in cell and animal models for breast cancer are lacking. CCL1 is highly expressed in invasive cancers and ER negative cancers and is related to tumor grade. CCL1 increased Treg infiltration and showed a poor survival in patients with breast cancer (Table 4) [168, 175]. CCL16 is described in section of the CCL3/5/7/8/14/15/16/23-CCR1 axis [156]. Studies on CCR8 in cell and animal models for breast cancer are lacking. A clinical trial (NCT06387628) is designed to evaluate the efficacy and safety of LM108 (CCR8 Ab) plus toripalimab plusnab-paclitaxel or eribulin as first-line or post-line treatment in patients with metastatic TNBC (https://clinicaltrials.gov/search?cond=breast%20cancer&intr=ccr8). There is a human study about the safety of BAY3375968 (CCR8 Ab) alone or in combination with pembrolizumab in patients with advanced solid tumors including BC (NCT05537740, https://clinicaltrials.gov/search?cond=breast%20cancer&intr=ccr8).
In cell models, CCL25 increased proliferation, migration, and invasion and reduced cisplatin-induced apoptosis in parallel with increased MMP1/9/11/13. However, CCL25 had no effects on migration and invasion in MCF7 cells (Table 2) [48, 49]. Studies on CCL25 in animal models for breast cancer are lacking. CCL25 is highly expressed in TNBC and African Americans and shows a poor survival, particularly in African Americans (Table 4) [157]. CCR9 Ab reduced CCL25-induced proliferation, migration, and invasion in parallel with a decrease of CCL25-induced MMP1/9/11/13 (Table 2) [48, 49]. Studies on CCR9 in animal models for breast cancer are lacking. CCR9 is highly expressed in poorly differentiated breast cancers (Table 4) [49].
CCL28 is described in section of the CCL5/7/11/13/14/15/24/26/28-CCR3 axis. In cell models, CCL27 increased migration and invasion in parallel with increased pErk and MMP7 (Table 2) [50]. Chemotoxin CCL27-PE38 had no change in lung metastasis in mice bearing 4T1 cells (Table 3) [120]. In human breast cancer samples, CCL27 is highly expressed in inflammatory breast cancers (Table 4) [149]. CCR10 KD reduced CCL27-induced migration and invasion in parallel with decreased pErk and MMP7 in cell model (Table 2) [50]. Studies on CCR10 in animal models for breast cancer are lacking. CCR10 is related to capsular invasion, stage, and lymph node metastasis (Table 4) [50].
Studies on CXCL6 in cell and animal models for breast cancer are lacking. In
human breast cancer samples, CXCL6 is highly expressed in ER negative breast
cancers and is related to metastasis of breast cancers, showing unchanging or
good survivals (Table 4) [187, 188]. CXCL7 increased migration and invasion in
parallel with increased pFak, MMP13, and VEGF. However, CXCL7 had no effects on
migration in T47D and ZR-75-1 cells (Table 2) [4, 51, 52, 53]. In animal models,
CXCL7 Ab reduced tumor growth and lung metastasis with decreased M2 M
CXCL1 increased sphere formation, proliferation, migration, and invasion in
parallel with increased vimentin,
CXCL4 reduced CXCL12-induced migration and CXCL447-70 decreased cell
proliferation (Table 2) [67, 68]. In animal models, CXCL447-70 decreased
tumor growth in parallel with increased tumoral mRNA levels of F4/80, CD11c, and
IFN-
CXCL12 increased proliferation, migration, invasion, colony formation, S and
G2/M phases, and adhesion and decreased apoptosis and calcium flux in parallel
with increased RhoA, Rac1, BclX, pHER2, pEGFR, pSTAT3, pErk, and pAkt and
decreased Bmf. Some reports showed that CXCL12 decreased proliferation, unchanged
invasion, and apoptosis in MDA-MB-231 cells, and had no effects on migration in
T47D, BT474/BT547, and SUM-149 cells. Although CXCL12 Ab and KD had no change in
proliferation, it reduced transendothelial migration (Table 2) [40, 43, 46, 73, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88]. In animal models, CXCL12 increased tumor growth and ICI-inhibited tumor
growth, preferring human HER2 cancers to TNBC or LA subtype [127]. CXCL12
expression increased tumor growth in orthotropic model and reduced lung
metastasis in pulmonary metastasis model with increased survivals in mice bearing
MDA-MB-231 cells, while CXCL12 Ab decreased lymphatic vessel and lymph node
metastasis [81, 128]. Compared to CXCL12 mutant, CXCL2 WT reduced tumor growth
and lung metastasis in mice bearing 4T1 cells, showing increased DC in lymph
nodes and decreased splenic MDSC. Treatment of CXCL12-EV stromal cells decreased
lung metastasis in mice bearing 4T1 cells [129, 130]. CXCL12 Ab had no effects on
tumor growth in mice bearing MCF-7 cells but reduced these effects in CXCL12
positive CAF-induced tumors [83, 131]. Fibroblast CXCL12 cKO mice showed
decreased tumor growth, lung metastasis, and angiogenesis after MVT1 cell
orthotropic injection, leading to an increased survival [87]. These results
indicate the involvement of CXCL12+ fibroblasts in breast cancer progression. In
addition, CXCL12 expression had no change in tumor growth but increased invasion
and angiogenesis with augmented TAM infiltration in mice bearing MTLn3 cells
(Table 3) [132]. In human breast cancer samples, CXCL12 is highly expressed in
breast cancers and BL subtype and is related to tumor stage, tumor grade, and
lymph node metastasis with increased Treg infiltration. CXCL12 has a good
prognosis and survival in patients with breast cancers, particularly who have
high plasma levels of CXCL12 and CXCL12
CXCL13 in combination with CXCR5 expression increased migration in parallel with augmented vimentin, slug, snail, N-cadherin, RANKL, MMP9, and pSrc, particularly decreasing E-cadherin in T47D cells [98]. CXCL13 Ab reduced proliferation and increased apoptosis in parallel with decreased pErk and cyclin D1 and increased caspase-9 (Table 2) [99]. Studies on CXCL13 in animal models for breast cancer are lacking. In human breast cancer samples, CXCL13 is highly expressed in breast cancers, ER negative cancers, and metastatic breast cancers and is related to lymph node metastasis with CXCR5 coexpression and increased Ki67. CXCL12 has a good prognosis and both unchanged and good survivals in patients with breast cancers (Table 4) [52, 98, 155, 160, 168, 187, 189, 190, 203, 209]. CXCR5 Ab reduced migration in parallel with decreased vimentin, slug, snail, N-cadherin, and RANKL and increased E-cadherin in CXCL13-treated CXCR5 expression cells (Table 2) [98]. Studies on CXCR5 in animal models for breast cancer are lacking. In human breast cancer samples, CXCR5 is related to lymph node metastasis and tumor stage (Table 4) [199].
CXCL14 expression reduced proliferation and invasion in MDA-MB-231 cells (Table 2) [100]. In animal models, CXCL14 expression reduced tumor growth, lung metastasis, and angiogenesis, showing decreased myeloid cells and Treg and increased CD8 infiltration in primary tumors and decreased TAM and Treg and increased CD8 infiltration in lung metastatic tumors (Table 3) [100, 140]. In human breast cancer samples, CXCL14 is related to lymph node metastasis and has good survivals in all breast cancers and BL/HER2/LA subtypes (Table 4) [100, 140, 155, 187, 189].
CXCL16 promoted migration, invasion, and F-actin polymerization in MDA-MB-231 and MCF-7 cells (Table 2) [101]. Studies on CXCL16 in animal models for breast cancer are lacking. In human breast cancer samples, CXCL16 increased the risk of cancer in HER2 subtype [156] and was related to stage (Table 4) [101]. CXCR6 Ab reduced CXCL16-induced migration, invasion, and F-actin polymerization in cell models (Table 2) [101]. CXCR6 KO mice showed no change in tumor growth but increased radiation-reduced tumor growth (Table 3) [141]. CXCR6 in human breast cancer samples is related to stage (Table 4) [101].
CXCL11 and CXCL12 are described in sections of the CXCL4/9/10/11-CXCR3 axis and the CXCL12-CXCR4 axis. In cell models, CXCR7 expression increased cell proliferation, while CXCR7 inhibitor and KD reduced proliferation and CXCL12-induced migration in parallel with decreased cyclin B1, Cdk4, pErk, EGF-induced pErk, pEGFR, pSTAT3, VCAM-1, MMP2, MMP9, and S-phase, and increased p21 and G0/G1 phase in cell cycle (Table 2) [73, 88]. In animal models, CXCR7 expression increased tumor growth but had no effect on invasion although it induced reduction in both metastasis and VEGF levels. On the other hand, CXCR7 inhibitor and KD caused no change in tumor growth or reduce tumor growth and metastasis in parallel with decreased pSTAT3, pErk, CD31, Ki67, cyclin D1, and MMP9 by downregulating TAM infiltration (Table 3) [88, 130, 142]. Interestingly, endothelial CXCR7 cKO treatment in lung metastasis model increased tumor growth and metastasis and showed no change in angiogenesis, leading to a poor survival (Table 3) [143]. In human breast cancer samples, CXCR7 is highly expressed in TNBC, ER/PR negative or positive cancers and is related to TNM stage and tumor grade, showing a poor survival (Table 4) [88, 218, 244].
In cell models, CXCL17 KD reduced proliferation and migration and CXCL17 increased pErk (Table 2) [102]. CXCL17 expression increased tumor growth in animal model (Table 3) [102]. In human breast cancer samples, CXCL17 is highly expressed in ER negative cancers and is related to Ki67 expression with a poor survival (Table 4) [102, 153]. CXCR8 KD decreased CXCL17-induced pErk in MCF7 cells (Table 2) [102]. Studies on CXCR8 in animal models for breast cancer are lacking. CXCR8 is related to tumor grade and Ki67 expression, showing no change in overall survival (Table 4) [102].
CX3CL1 increased proliferation in T47D cells by increasing pErk, pErbB1, and pErbB2 levels (Table 2) [98]. In animal models, intratumoral injections of Ad-CX3CL1 increased palpable tumors but had no change in angiogenesis. CX3CL1 KO showed unchanged or delayed mammary tumor onset, unchanged or decreased tumor number without alteration in tumor growth (Table 3) [103]. In human breast cancer samples, CX3CL1 is highly expressed in inflammatory breast cancers, LB subtype, and PR positive cancers and is related to tumor grade, tumor stage, tumor size, and lymph node metastasis with increased Ki67, stromal CD8, intratumoral DC, stromal NK, and TIL infiltration, showing both good and poor survivals (Table 4) [149, 153, 245, 246]. CX3CR1 expression increased pErk in MDA-MB-436 cells (Table 2) [103]. CX3CR1 expression increased bone metastasis in intracardiac injection models, while CX3CR1 KO mice showed reduced bone metastasis (Table 3) [103]. In human breast cancer samples, CX3CR1 is related to brain metastasis but has no change in survival (Table 4) [177].
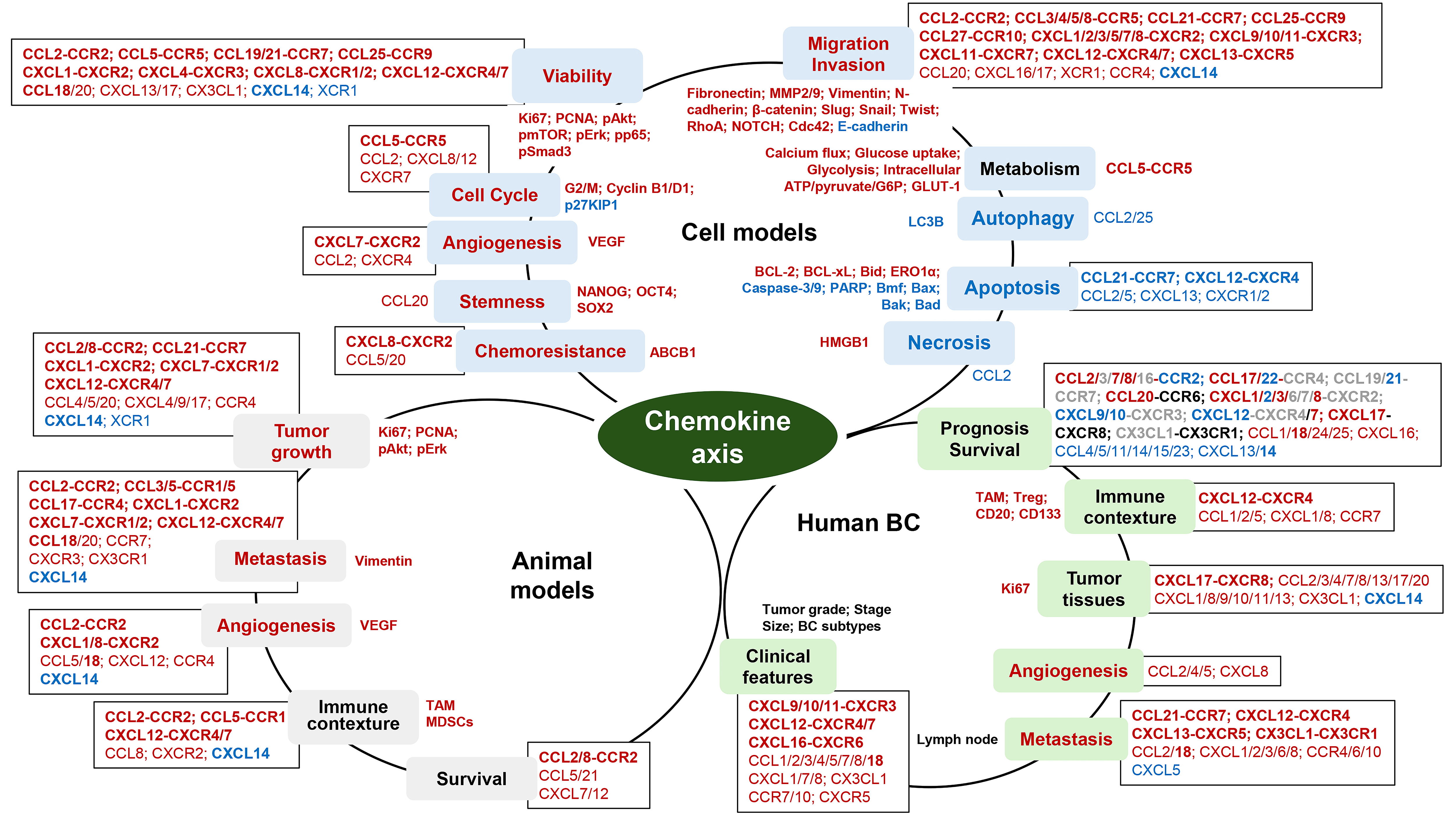
In breast cancer cell models, chemokine axes play a key role in cell migration and invasion, showing positive effects on cell viability, angiogenesis, cell cycle, stemness, and chemoresistance but negative effects on autophagy, apoptosis, and necrosis (Fig. 1). Because some breast cancer cells are less responsive to chemokines, significant roles of chemokine axes in breast cancer cells are likely to be quite dependent on signature of chemokines and chemokine receptors which breast cancer cells express. In addition, chemokine axes may be involved in cellular metabolism by increasing calcium flux, glucose uptake, glycolysis, intracellular ATP/pyruvate/G6P, and GLUT-1 (Fig. 1) which require cancer cell growth and spread. In animal models for breast cancer cells, chemokine axes play a crucial role in tumor growth, metastasis, and angiogenesis, as well as immune contexture in the tumor microenvironment by increasing TAM and MDSCs, which affect overall survivals (Fig. 1). Differently from single cell models, animal models may appear to have various chemokine axes between tumor cells, stromal cells, immune cells, and adipocytes in the breast tumor microenvironment, showing complex interaction which makes hard to simply explain with single chemokine axis. In human breast cancer, chemokine axes appear to play a critical role in immune contexture by recruiting immune cells, such as TAM, Treg, CD20, and CD133 cell, probably affecting prognosis and overall survivals (Fig. 1). Some chemokine axes are related to clinical features, such as tumor grade, stage, size, lymph node metastasis, and breast cancer subtypes, showing increased Ki67 in tumor tissues. Based on the integrated results from cell and animal models and clinical aspects, the CCL2-CCR2 axis, the CCL3/5-CCR5 axis, the orphan chemokine CCL18, the CXCL1/8-CXCR2 axis, and the CXCL12-CXCR4/7 axis play a significant role in chemokine-induced breast cancer progression. Interestingly, orphan chemokine CXCL14 looks like to block the harmful roles of some chemokine axes in breast cancer (Fig. 1), supporting benefits for breast cancer. In conclusion, chemokine axes are involved in breast cancer progression via tumor growth, migration, invasion, and angiogenesis in the tumor microenvironment, as well as systemic metastasis. Furthermore, inhibitors and antibodies to disrupt specific chemokine axes have diminished chemokine-derived breast cancer progression, providing the adjuvant therapeutic options to enhance the effectiveness of conventional therapeutic strategies.
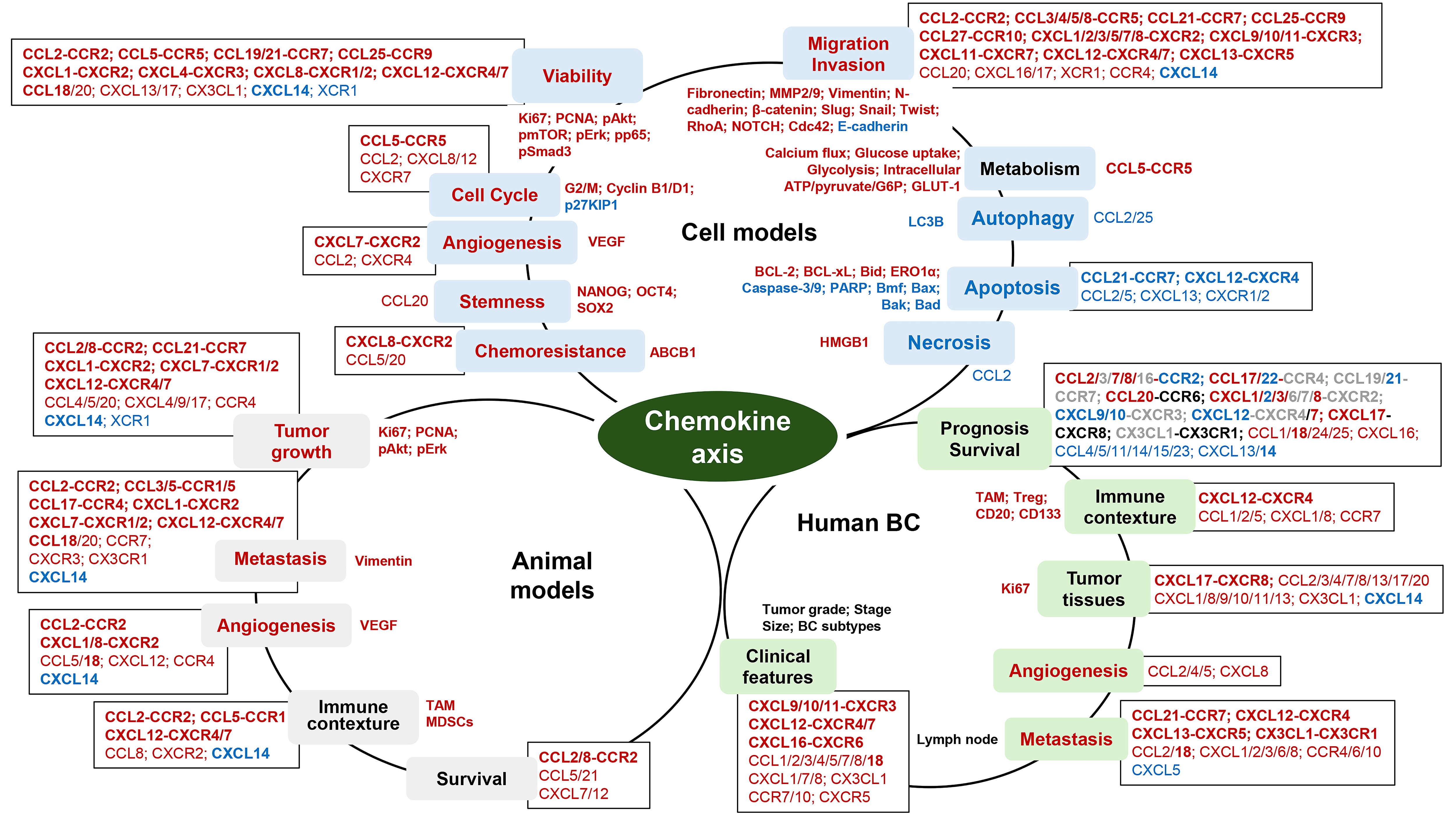 Fig. 1.
Fig. 1.
Functional roles of chemokine axis in cell and animal models and human breast cancers. The chemokine-induced signaling pathways are inside circles for each model, biological effects of chemokines are on the circular lines (color blocks), and the contributed chemokine axis and chemokines are outside circles of each model (square boxes). Red: positive effects; blue: negative effects. In box, bold: chemokine axis; roman: chemokine or chemokine receptor. In survival box, red: poor survival; blue: good survival; gray: controversial; black: no effects.
D-SS and SEA designed the research review. D-SS wrote original draft preparation. Both authors contributed to editorial changes in the manuscript. Both authors read and approved the final manuscript. Both authors have participated sufficiently in the work to take public responsibility for appropriate portions of the content and agreed to be accountable for all aspects of the work in ensuring that questions related to its accuracy or integrity.
Not applicable.
Not applicable.
This research was funded, in whole or in part, by National Institutes of Health (NIH) as the following grants: NCI SC1CA200519 (D.-S.S.), NCI P50CA098131 (D.-S.S.), U54MD007586 (S.E.A.), U54CA163069 (D.-S.S., S.E.A.), and ACS DICRIDG-21-071-01-DICRIDG (D.-S.S., S.E.A.).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.