1 Department of Chest Medicine, Taipei Veterans General Hospital, 112 Taipei, Taiwan
2 School of Medicine, National Yang Ming Chiao Tung University, 112 Taipei, Taiwan
3 Institute of Clinical Medicine, National Yang Ming Chiao Tung University, 112 Taipei, Taiwan
Academic Editor: Yoh Dobashi
Abstract
The molecular profiling of tumors is fundamental in the management of advanced non-small-cell lung cancer (NSCLC). A tissue specimen obtained from biopsy is needed for diagnosis and mutation analysis. However, this may not be feasible for some metastatic sites, such as central nervous system (CNS) lesions, particularly for repeated biopsy. Liquid biopsy with plasma is an emerging tool for molecular testing and could be a surrogate method if tissue cannot be obtained. However, the use of plasma is limited for the detection of mutations arising from intracranial lesions. Cerebrospinal fluid (CSF) was recently demonstrated to be an alternative material for genetic testing in patients with NSCLC having CNS metastasis. In this review, we discuss recent advancement in the use of CSF as a medium of liquid biopsy in patients with NSCLC.
Keywords
- Non-small-cell lung cancer
- Cerebrospinal fluid
- Liquid biopsy
- Cell free tumor DNA
Central nervous system (CNS) metastases are common in patients with lung cancer
and are associated with significant morbidity and mortality. The incidence is
approximately 25%–30% in patients with non-small-cell lung cancer (NSCLC),
particularly in oncogene-addicted NSCLC, and
Leptomeningeal metastasis (LM) is a late and lethal complication of various
solid tumors, defined as the seeding of tumor cells into the CSF and
leptomeninges [9]. LM incidence in patients with NSCLC is approximately 3%–5%
and reaches up to 9.4% in patients with EGFR mutation [10, 11]. The
prognosis of patients with NSCLC having LM is poor, with a median survival of
Liquid biopsy with plasma is an emerging tool for molecular testing and could be a surrogate method if tissue cannot be obtained [15]. However, the use of plasma cell–free DNA may be limited for the detection of mutations arising from intracranial lesions and may have limited performance in patients with isolated CNS progression [16, 17]. CSF, by contrast, circulates throughout the CNS and may provide a window to investigating intracranial lesions. The CNS includes two anatomic compartments, namely the brain parenchyma and leptomeningeal space. The blood–brain barrier and blood-CSF barrier are the two barrier systems for each compartment [18]. CSF enters the parenchyma along paravascular spaces that surround penetrating arteries and the brain interstitial fluid is cleared along paravenous drainage pathways [19]. Because these two compartments are not anatomically isolated, CSF-based liquid biopsy of CNS metastatic lesions could be investigated in clinical practice. Research indicates that CSF harbors clinically relevant genomic alterations in patients with CNS malignancies and could be considered for liquid biopsies to monitor tumor evolution in the CNS [20]. Different tumor components in the CSF are subjects to be analyzed, such as circulating tumor cells (CTCs), cell-free tumor DNA (ctDNA), extracellular vesicles (EVs), and RNA. In this article, we reviewed relevant English-language journals articles indexed in PubMed (using the search terms “CSF”, “CTCs”, “ctDNA”, “RNA”, “EVs” and “lung cancer”) from January 2000 to August 2021.
The presence of malignant cells in CSF remains the gold standard for diagnosing LM; however, the sensitivity of a single lumbar puncture is low and repeated sampling is often needed [12, 21]. Some technical issues such as inadequate CSF volume for the analysis or delayed processing may lead to false negative results [22]. Recently, novel assays have been developed to detect and quantify CTCs in CSF. CellSearch assay and immunoflow cytology methods have been the two most common techniques applied.
The CellSearch platform uses epithelial cell adhesion molecule (EpCAM) antibody–conjugated ferroparticles and an immunomagnetic selection system to capture tumor cells in CSF. CTCs are defined as epithelial staining positive cells which lack CD45 expression [18]. Conversely, the immunoflow cytometry technique uses fluorescently labeled antibodies against membrane-bound proteins of epithelial tumor cells, such as EpCAM. Subsequently, CTC enumeration is performed using the fluorescence–activated cell sorting system. Furthermore, leukocytes and other hematopoietic cells are labeled to select CTCs [18]. To date, many studies have used these two assays to detect CTCs in the CSF of patients with solid tumors and LM and have mainly focused on patients with NSCLC (Table 1, Ref. [23, 24, 25, 26, 27, 28, 29, 30, 31, 32]). Overall, sensitivity is significantly higher with CTCs (75%–100%) than with conventional cytology (44.4%–65.3%) [23, 24, 25, 26, 27, 28, 29, 30, 31, 33]. Moreover, the CTC percentage in CSF is correlated with overall survival [25, 32]. However, the application of these assays in clinical practices has some limitations. Previous studies have focused only on patients with LM, and the use of CTCs in patients with parenchymal–only CNS metastasis remains unknown. Furthermore, the difference in experimental procedures and cutoff values between studies entail a need to conduct large prospective trials for validation. CTCs may have low EpCAM expression due to the epithelial-to-mesenchymal transition [34]. This leads to the underestimation of the CTC number, and key subpopulations may be overlooked.
| Study | Platform for sequencing | Condition of patient population | Number (Total/LC) | Main finding | Molecular application |
| Subirá et al., 2012 [23] | Flow cytometry immunophenotyping | Epithelial-cell solid tumors with suspected LM | 78/23 | CTC exhibited higher sensitivity compared with cytology (75.5% vs. 65.3%) | NA |
| Nayak et al., 2013 [24] | CellSearch assay | Solid tumors with suspected LM | 51/21 | CTC showed exhibited sensitivity compared with cytology (100% vs. 66.7%) | NA |
| Subirá et al., 2015 [25] | Flow cytometry immunophenotyping | Epithelial-cell solid tumors with LM | 144/35 | CTC exhibited higher sensitivity compared with cytology (79.8% vs. 50%) | NA |
| The percentage of CSF EpCAM+ cells predicted overall survival | |||||
| Tu et al., 2015 [26] | CellSearch assay | LC with MRI confirmed LM | 18/18 | CTC exhibited higher sensitivity compared with cytology (77.8% vs. 44.4%) | NA |
| Milojkovic Kerklaan et al., 2016 [27] | EpCAM-based flow cytometry | Epithelial-cell solid tumors with clinical suspicion of LM but a negative or inconclusive MRI | 29/8 | CTC exhibited higher sensitivity compared with cytology (100% vs. 61%) | NA |
| Lv, et al., 2016 [33] | Immunofluorescence in situ hybridization | LC with LM | 16/16 | Sensitivity of CTC: 75% | NA |
| Jiang et al., 2017 [28] | CellSearch assay and next-generation sequencing | LC with suspected LM | 21/21 | CTC showed exhibited sensitivity compared with cytology (95.2% vs. 57.1%) | Concordant rate with primary tumor: 89.5% |
| 1/14 had EGFR p.T790M mutations | |||||
| Lin et al., 2017 [29] | CellSearch assay | Epithelial-cell tumors with suspected LM | 95/31 | Sensitivity of CTC: 93% | NA |
| Cut-off for CTC positivity: 1 CTC/mL | |||||
| Nevel et al., 2020 [32] | CellSearch assay | LC with LM | 16/16 | CTC ( |
NA |
| Torre M, et al., 2020 [30] | CellSearch assay | Solid tumor with suspected LM | 20/4 | Sensitivity of CTC: 88.9% | NA |
| Cutoff for CTC positivity: 1 CTC/mL | |||||
| Van Bussel, et al. 2020 [31] | EpCAM-based flow cytometry | NSCLC with suspected LM | 81/81 | Sensitivity of CTC: 88.9% | 4/5 had EGFR mutation in CTC |
| Cutoff for CTC positivity: 0.9 CTC/mL | |||||
| CSF, cerebrospinal fluid LM, leptomeningeal metastasis; LC, lung cancer; EpCAM, epithelial cell adhesion molecule; NA, not available; NSCLC, non-small cell lung cancer; CTC, circulating tumor cell; MRI, magnetic resonance imaging; EGFR, epidermal growth factor receptor. | |||||
In addition to the identification and quantitation of CTCs in CSF, researchers have attempted to analyze the genetic alteration of CTCs in CSF. Using next-generation sequencing (NGS), Jiang et al. [28] observed that the genetic profiles of CSF CTCs were highly concordant with primary tumors (17/19, 89.5%). In addition to other resistant mutations, the EGFR exon 20 p.T790M resistant mutation was identified in 7.1% of patients. Furthermore, van Bussel et al. [31] exhibited a high EGFR mutation detection rate (4/5, 80%) in CTCs from CSF of patients with EGFR-mutant NSCLC patients. Ruan et al. [35] further sequenced the CSF CTCs from patients with lung adenocarcinoma into a single cell level and found that metastatic-CTCs were enriched for metabolic and cell-adhesion pathways. Furthermore, interpatient and intrapatient heterogenicities were observed. These novel techniques break new ground in the diagnosis and treatment of LM in patients with NSCLC.
The constituents of cell-free DNA extracted from plasma are mostly the germline DNA from white blood cells, which make ctDNA analysis challenging due to high background noise. On the other hand, cell-free DNA extracted from CSF of patients with cancer is thought to be enriched for ctDNA due to the significantly lower number of normal cells in CSF compared with that in blood. Using mutation allele-specific amplification, Swinkels et al. [36] first identified KRAS mutation in CSF of two patients with NSCLC with LM. Subsequent studies, using either polymerase chain reaction (PCR) or NGS, have demonstrated that CSF ctDNA is a feasible material for mutation detection, particularly in patients with LM, with a detection rate as high as 100% [17, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]. Published studies on ctDNA in CSF of patients with NSCLC with CNS metastasis are summarized in Table 2 (Ref. [17, 20, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53]). Most studies have focused on patients with EGFR-mutant NSCLC with LM due to the relatively high number of these patients [10]. Li et al. [39] eported that among patients with EGFR mutant NSCLC patients with LM, the ctDNA of CSF supernatant had a unique genetic profile and high allele fractions compared with CSF pellets or plasma. Furthermore, in patients with brain metastasis, the detection of ctDNA mutation is higher in CSF than in plasma, ranging from 50% to 63% [16, 20, 51]. These results reveal that CSF ctDNA is a feasible material for mutation testing.
| Study | Platform for sequencing | Condition of patient population | Number | Pattern of CNS metastasis | Mutation detection rate of CSF | Resistant mutation |
| Swinkels et al., 2000 [36] | Mutation-allele specific amplification (PCR) | NSCLC | 2 | LM | KRAS mutation 100% detected in CSF (2/2) | NA |
| Pentsova et al., 2016 [20] | Targeted sequencing | Solid tumors | 41, NSCLC:11 | 32 BM | 63% (20/32) of patients with BM | EGFR p.T790M: 50% (2/4) post-TKI |
| 9 LM | 75% (3/4) of patients with LM | |||||
| Marchiò et al., 2017 [37] | Targeted sequencing | NSCLC | 2 | LM | KRAS mutations 100% detected in CSF (2/2) | NA |
| Fan et al., 2018 [38] | Targeted sequencing | EGFR-mutant NSCLC | 11 | LM | 100% | EGFR p.T790M: 0% |
| Li et al., 2018 [39] | Targeted sequencing | EGFR-mutant NSCLC | 28 | LM | 100% | EGFR p.T790M: 30.4% (7/23) post-TKI |
| MET copy number gain:47.8% (11/23) post-TKI | ||||||
| Xu et al., 2018 [40] | Mutation-allele specific amplification (PCR) | NSCLC | 49 | LM | 92.3% (39/42) in patients EGFR mutation | EGFR p.T790M: 3.85% (1/26) in post-EGFR-TKI patients |
| Ge et al., 2019 [41] | Targeted sequencing | NSCLC | 29 | LM | 66.67% overall | NA |
| Huang et al., 2019 [51] | Droplet digital PCR | EGFR-mutant NSCLC | 35 | 20 BM | 50% (5/10) of patients with BM | EGFR p.T790M: 13% |
| 15 LM | 75% (9/12) of patients with LM | |||||
| Zheng, et al., 2019 [44] | Targeted sequencing | ALK mutant NSCLC | 30 | LM | ALK fusions were detected in 83.3% (10/12) of samples | ALK p.G1202R and p.C1156F were found in one post-alectinib sample |
| Kawahara et al., 2019 [52] | Mutation-allele specific amplification (PCR) | LC | 26 | NA | 75% of valid samples had EGFR mutations | EGFR p.T790M: 33.3% (4/12) in valid samples |
| Ying et al., 2019 [42] | Targeted sequencing | NSCLC | 92 | LM | 81.5% overall, EGFR is the most common gene (58%) | EGFR p.T790M: 2.8% (2/72) |
| Zhao et al., 2019 [43] | Targeted sequencing | Malignancy | 35, LC:74% | LM | 100% | EGFR p.T790M: 33.3% (2/6) |
| Ma et al., 2020 [45] | Targeted sequencing | NSCLC | 21 | 10 BM | 95.2% (20/21) overall, 100% with LM | NA |
| 11 LM | 81.8% (9/11) LM patients had EGFR mutation | |||||
| Li, et al., 2020 [53] | Targeted sequencing | NSCLC | 94 | 24 BM | 100% overall, EGFR is the most common gene (84%) | NA |
| 70 LM | ||||||
| Aldea et al., 2020 [17] | Targeted sequencing | NSCLC | 12 | 1 BM | 83% | NA |
| 11 LM | ||||||
| Moushumi Suryavanshi et al., 2020 [47] | droplet digital PCR | EGFR-mutant NSCLC | 17 | LM | 94.1% | No EGFR p.T790M mutation was found post-TKI |
| Zhao et al., 2020 [46] | Targeted sequencing | Solid tumors | 58, LC: 42 | LM | 100% overall, EGFR mutation rate: 70.97% | NA |
| Chiang et al., 2021 [48] | Mutation-allele specific amplification (PCR) | EGFR-mutant NSCLC | 48 | LM | 68.8% (33/48), 80.5% in patients with abnormal cytologically finding | EGFR p.T790M: 14.6% |
| Choi et al., 2021 [49] | Nanowire-based cfDNA assay | EGFR-mutant NSCLC after third generation TKI | 11 | LM | 72.7% (8/11) | EGFR p.C797S: 36.3% (4/11) |
| MET amplification: 18.2% (2/11) | ||||||
| Zheng et al., 2021 [50] | Targeted sequencing | EGFR-mutant NSCLC | 80 | LM | ||
| cohort 1: pre-osimertinib (45) | Cohort 1: 93.3% (42/45) | Cohort 1: 21.4% (9/42) | ||||
| cohort 2: post-osimerinib (35) | Cohort 2: 97.1% (34/35) | Cohort 2: 21.7% (5/23) maintained EGFR p.T790M | ||||
| NSCLC, non-small cell lung cancer; LM, leptomeningeal metastasis; BM: brain metastasis; TKI, tyrosine kinase inhibitor; cfDNA, cell-free tumor DNA; PCR, polymerase chain reaction; EGFR, epidermal growth factor receptor; ALK, anaplastic lymphoma kinase. | ||||||
Acquired resistance at disease progression is often inevitable after the use targeted therapies. Tumor rebiopsy is required to obtain information regarding histologic or genetic changes. For patients with LM, tissue rebiopsy is not feasible, and CSF ctDNA could provide useful information for subsequent treatment in this scenario. EGFR exon 20 p.T790M mutation, which is the most common resistant mechanism after first- or second- generation EGFR-tyrosine kinase inhibitor (TKI), has an incidence rate of approximately 55%–64% [54, 55]. Despite the difference in the testing method used, the percentage of EGFR exon 20 p.T790M mutation was relatively low in CSF ctDNA, ranging from 0% to 50% [20, 38, 39, 40, 42, 43, 48, 51, 52]. Conversely, MET copy number gain was a relatively common resistance mechanism in this patient group [39, 56]. Zheng et al. [44] analyzed CSF ctDNA through targeted sequencing in patients with anaplastic lymphoma kinase (ALK)-rearranged NSCLC with LM, and ALK fusion was detected in 83.3% of the patients. ALK-resistant mutation was detected in one patient after resistance to alectinib. In addition to being capable of detecting a resistant mutation, serial CSF ctDNA analysis has the potential of longitudinally monitoring tumor burden in patients with LM [39, 44].
Different sequencing platforms may affect the mutation detection rate in patients with lung cancer. Studies have reported that droplet digital PCR (ddPCR) may have higher sensitivity in plasma EGFR mutation detection compared with amplification refractory mutation system PCR assays (ARMS) [57, 58]. However, few studies have directly compared different platforms for the CSF ctDNA analysis. Using the primary tumor testing results as reference, Xu et al. [40] reported that ddPCR and ARMS had similar sensitivity and specificity when used for CSF ctDNA analysis. Moreover, targeted sequencing with the NGS method exhibited a high mutation detection rate in CSF ctDNA (Table 2). In addition to sensitizing mutation, NGS can be used to identify the resistance mechanism after targeted therapy.
The prognostic value of the CSF ctDNA analysis in patients with NSCLC with CNS metastasis has also been examined. Li et al. [53] identified five molecular subtypes by using cluster analysis and found that the cluster containing high percentages of CDK4, TP53, MET, and CDKN2A was associated with the worst prognosis. In patient with EGFR-mutant NSCLC with LM, the mutation type detected in CSF ctDNA indicated a different response to the third-generation EGFR-TKI, osimertinib. Patients with EGFR exon 19 deletion had better intracranial progression-free survival compared to EGFR exon 21 p.L858R mutation. Moreover, the prognosis was better in patients harboring EGFR p.T790M mutation in CSF before osimeritnib use [50]. Thus, the ctDNA of CSF is a promising material for LM diagnosis, genomic alteration detection, and the treatment response supervision. The CSF ctDNA analysis should be considered a useful tool accompanied by cytological examination in patients with CNS metastasis, particularly in oncogene-addicted patients with LM.
Cells may communicate with each other through EVs, which comprise of proteins, genomic DNA, and various RNAs. EVs are divided into three groups based on their size and origin: exosomes, microvesicles, and apoptotic bodies [59]. Short noncoding RNAs regulate protein translation by binding to the 3’ untranslated regions of messenger RNAs and are generally called microRNAs (miRNAs). In addition to being secreted by the cells in EVs, miRNAs could be released by necrotic cells as free oligonucleotides [60]. Recent studies have investigated the role of EVs and circulating miRNAs in CSF, in addition to CTCs and ctDNA, as a potential biomarker of CNS tumors.
Teplyuk et al. [61] analyzed the miRNAs of CSF by using quantitative reverse transcription PCR and observed that the levels of miR-10b and miR-21 were significantly higher in the CSF of patients with glioblastoma and metastatic brain tumors compared with those with nonneoplastic conditions. Furthermore, they found that the longitudinal miRNA profiles could reflect disease activity. Using microarray analysis and quantitative real-time PCR, Pan et al. [62] analyzed CSF miRNAs in patients with lung adenocarcinoma with LM. Three miRNAs in CSF (miR-7975, miR7977, and miR7641) were significantly upregulated in patients with LM and the changes in these miRNAs correlated with disease course. Through function analysis, miR7977 was predicted to be involved in various pathways of cancer metastasis. In lung adenocarcinoma cells, miR7977 was found to be a key regulator of proliferation, migration, and invasion. However, some concerns regarding miRNA use in CSF exist. The major concern is specificity; differentiating identify the difference between cancer-derived and inflammation-derived miRNA profiles is difficult. Furthermore, differentiating between miRNAs types in CSF, such as EV-associated miRNAs or naked miRNAs, is difficult. Additionally, a universal guideline is lacking for the processing of CSF samples, miRNA extraction, measurements, and data analysis [63].
Circular RNAs (circRNAs) are a class of noncoding RNAs that comprise a circular loop with miRNA binding sites and that regulate gene expression by acting as a miRNA sponge to compete with endogenous miRNAs [64]. Furthermore, circRNAs are abundant in the brain [65], and circRNA-miRNA networks that comprise circ-TTC39C and circ-PCCA have been identified in the CSF of patients with neurodegenerative diseases [66, 67]. Multiple circRNAs have been discovered in the blood and tumor tissue of patients with lung cancer [68, 69]. The expression of CircRNA_102481 in exosomes was related to brain metastasis in patients with NSCLC [70]. Collectively, alterations in circRNAs can be detected in CSF and are currently being actively investigated for their potential as biomarkers for patients with lung cancer with CNS metastasis.
Lee et al. [71] measured nanoparticles in 472 CSF samples and found that the concentration of CSF nanoparticles significantly increased in patients with LM. The concertation changes of nanoparticles were examined in 33 patients with NSCLC after intrathecal chemotherapy. EVs was positively correlated with overall survival. The same group further investigated the subpopulation of small noncoding RNA from EVs in patients with NSCLC with LM. The functions of these noncoding RNA, particularly miR-21, in LM pathogenesis were documented [72]. Further studies are required to investigate the role of EVs as a novel pharmacological target for LM.
The use of CSF as a medium of liquid biopsy in patients with NSCLC with CNS metastasis is still evolving. In addition to having better diagnostic sensitivity, CTCs detection in CSF can provide information regarding tumor burden, particularly in patients with LM. The analysis of CSF ctDNA could identify driver oncogenes, monitor treatment response, and track the resistance mechanism [73]. The sequencing of RNA or EVs in CSF is new in this field and may lead to a better understanding of LM pathogenesis (Fig. 1). No study has compared these platforms with respect to detection rate and clinical usefulness. However, we believe that each platform could provide complementary information. The optimal use of these materials in CSF for diagnosis, tumor heterogenicity tracking and treatment guidance requires prospective studies for validation. It is hoped that the application of liquid biopsy by using CSF in clinical practice will improve outcomes in this patient group.
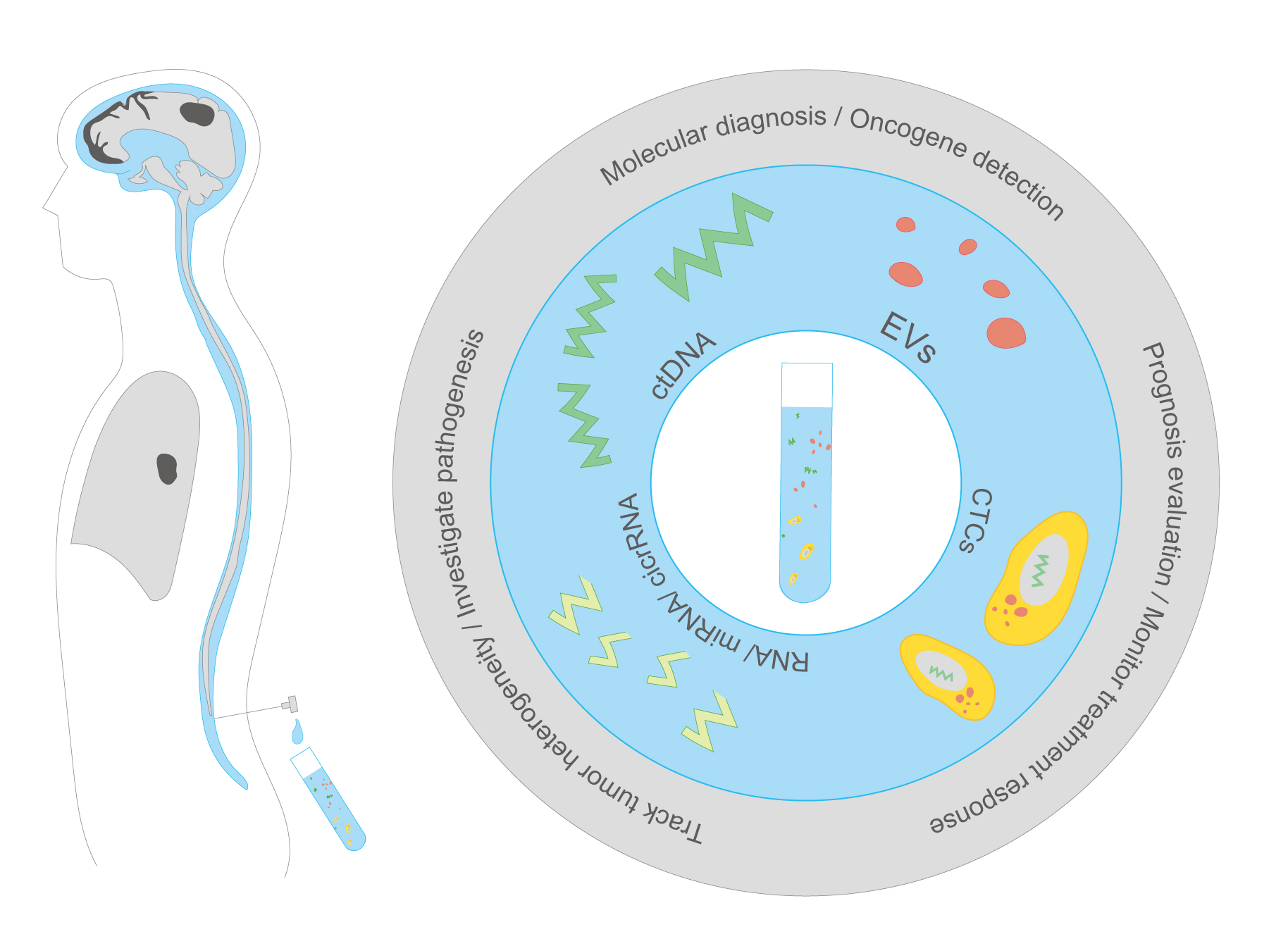 Fig. 1.
Fig. 1.Cerebrospinal fluid as a medium of liquid biopsy in non–small-cell lung cancer. miRNA, microRNA; circRNA, circular RNA; ctDNA, cell-free tumor DNA; EVs, extracellular vesicles; CTCs, circulating tumor cells.
CLC and CHC designed the study. HCH performed the illustration. CLC and YHL wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript.
Not applicable.
This manuscript was edited by Wallace Academic Editing.
This study was funded by Taipei Veterans General Hospital, Taiwan (V110B-008).
CLC and CHC has received honoraria from AstraZeneca, Boehringer Ingelheim, Pfizer, and Roche. Other authors declare no conflict of interest.
NSCLC, non-mall-cell lung cancer; CNS, central nervous system; EGFR, epidermal growth factor receptor; CSF, Cerebrospinal fluid; LM, Leptomeningeal metastasis; CTC, circulating tumor cells; EpCAM, epithelial cell adhesion molecule; ctDNA, cell-free tumor DNA; ALK, anaplastic lymphoma kinase; EV, extracellular vesicles; NGS, next-generation sequencing.

