1 Institut Hospitalo-Universitaire Mediterranée Infection, 13005 Marseille, France
2 Microbe Evolution Phylogeny and Infection (MEPHI), Aix Marseille University, Institut de recherche pour le développement (IRD), Assistance publique hôpitaux de Marseille (AP-HM), 13005 Marseille, France
3 Parasitology and Entomology Unit, Infectious Diseases and Microbiology Department, Institut de Recherche Biomédicale des Armées (IRBA), 13005 Marseille, France
4 Vecteurs - Infections Tropicales et Méditerranéennes (VITROME), Aix Marseille University, Institut de recherche pour le développement (IRD), Service de Santé des Armées (SSA), Assistance publique hôpitaux de Marseille (AP-HM), 13005 Marseille, France
5 National Reference center for Malaria, 13005 Marseille, France
Academic Editor: Josef Jampílek
Abstract
Background: Despite the fact that the clinical efficacy of
hydroxychloroquine is still controversial, it has been demonstrated in
vitro to control SARS-CoV-2 multiplication on Vero E6 cells. In this study, we
tested the possibility that some patients with prolonged virus excretion could be
infected by less susceptible strains. Method: Using a high-content
screening method, we screened 30 different selected isolates of SARS-CoV-2 from
different patients who received azithromycin
Keywords
- SARS-CoV-2
- Hydroxychloroquine
- Azithromycin
- In vitro model
- Vero E6
- WGS
- Africa
In December 2019, a novel coronavirus named SARS-CoV-2 emerged in Wuhan, China,
in the province of Hubei [1, 2, 3]. SARS-CoV-2 spread rapidly around the world and
the number of cases and deaths has increased rapidly
(https://ourworldindata.org/coronavirus-data). Since then, finding effective
treatments and vaccines has remained a major global challenge. Several drugs with
an antiviral effect have been tested in vitro and in vivo, with
drug repurposing being one of the strategies applied. Many drugs have shown an
inhibition in vitro such as certain antimalarial drugs (chloroquine,
hydroxychloroquine), ivermectin, macrolides (azithromycin, spiramycin), several
protease inhibitors and some RdRp (RNA-dependent RNA polymerase) inhibitors such
as remdesivir and sofosbuvir [4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18]. The combination of azithromycin and
hydroxychloroquine showed synergistic effects in vitro [19] at
concentrations of 5
Nasopharyngeal samples were collected at the IHU Méditerranée Infection
as part of COVID-19 diagnosis and patient follow-up. The study was approved by
the ethics committee of the University Hospital Institute Méditerranée
Infection (No.: 2020–029). With regards to both the French and the local
situations, we defined two periods of time during the pandemic: the first
consisting in the arrival of the virus and the lockdown between February and May
2020, and the second during the summer of 2020. Patients 6 to 15 were selected
according to the persistence of their infection (defined by two positive PCR
tests ten days apart). As stated above, all these patients were from the first
wave of pandemic, between late February and May 2020 in our area. As controls we
analysed a group of randomly chosen non-persistent control patients, patient 1 to
patient 5 for the first wave of pandemic, and patient 16 to patient 20 for the
second wave of pandemic, July–August 2020 in our area. Information on the sample
collection, name of the strains, and treatments are summarised in Table 1. All
patients received azithromycin that was, in most cases, combined with
hydroxychloroquine [27]. For persistent patients, when it was possible, we
evaluated the susceptibility of two strains, the first isolated upon admission
and the second isolated during evolution under treatment. The IHUMI-3 isolate was
among the first strains isolated in the laboratory and used as a control, as in
our previous experiments [19]. Viral isolation was performed following the
procedure described [29]. After isolation, the viruses were harvested and frozen
at –80
| Patientnumber | Strain | Month of sampling | Persistence | Day of the latest isolate after onset | RT-PCR Ct values | Treatment | Clade | Access number |
| Patient 1 | IHUMI-11 | March 2020 | No | NA | 29 | AZT, HCQ | 20A/15324T | EPI_ISL_568957 |
| Patient 2 | IHUMI-15 | March 2020 | No | NA | 23 | HCQ | 20B | EPI_ISL_568913 |
| Patient 3 | IHUMI-240 | March 2020 | No | NA | 22 | AZT, HCQ | 20C-5 | EPI_ISL_569338 |
| Patient 4 | IHUMI-243 | March 2020 | No | NA | 29 | AZT, HCQ | 20A/15324T | EPI_ISL_569338 |
| Patient 5 | IHUMI-597 | March 2020 | No | NA | 20 | / | 20A/25563T | https://doi.org/10.35081/8ytr-4812 |
| Patient 6 | IHUMI-215 | March 2020 | Yes | 8 | 23 | AZT, HCQ | 20A/25563T-1B | EPI_ISL_2286971 |
| IHUMI-611 | 32,2 | 20A/25563T-1B | ||||||
| Patient 7 | IHUMI-364 | March 2020 | Yes | 4 | 21 | AZT, HCQ | 20A/15324T | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-599 | 29,1 | |||||||
| Patient 8 | IHUMI-284 | March 2020 | Yes | 4 | 30 | AZT | 20A/A0268G-2 | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-538 | 20,5 | 20A/A0268G-2 | ||||||
| Patient 9 | IHUMI-713 | March 2020 | Yes | 3 | 31 | AZT, HCQ | 20A/25563T-1 | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-800 | 23,1 | 20A/25563T-1 | ||||||
| Patient 10 | IHUMI-684 | March 2020 | Yes | 4 | 21,6 | AZT, HCQ | 20A/25563T | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-743 | 20,4 | 20A/25563T | ||||||
| Patient 11 | IHUMI-598 | March 2020 | Yes | 4 | 20,5 | AZT, HCQ | 20C-5 | EPI_ISL_569337 |
| IHUMI-801 | 20,7 | 20C-5 | ||||||
| Patient 12 | IHUMI-717 | March 2020 | Yes | 2 | 21,2 | AZT, HCQ | 20B-1a | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-742 | 19,1 | 20B-1a | ||||||
| Patient 13 | IHUMI-624 | March 2020 | Yes | 2 | 16,1 | AZT, HCQ | 20A/25563T-1b | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-719 | 17,7 | |||||||
| Patient 14 | IHUMI-288 | March 2020 | Yes | 5 | 23 | AZT, HCQ | 20C-4 | https://doi.org/10.35081/8ytr-4812 |
| IHUMI-614 | 26 | |||||||
| Patient 15 | IHUMI-880 | April 2020 | Yes | 3 | 19 | HCQ | 20B | EPI_ISL_568909 |
| IHUMI-990 | 21,4 | 20B | ||||||
| Patient 16 | IHUMI-2122 | July 2020 | Unknown | NA | 17,8 | AZT | Marseille 1 | EPI_ISL_569023 |
| Patient 17 | IHUMI-2123 | July 2020 | Yes | NA | 17,7 | AZT, HCQ | Marseille 1 | EPI_ISL_569029 |
| Patient 18 | IHUMI-2137 | August 2020 | Yes | NA | 14,7 | AZT, HCQ | Marseille 5b | EPI_ISL_569119 |
| Patient 19 | IHUMI-2177 | August 2020 | No | NA | 25,1 | AZT | Marseille 1A | EPI_ISL_1745715 |
| Patient 20 | IHUMI-2178 | August 2020 | Unknown | NA | 21,6 | AZT, HCQ | Marseille 1A | EPI_ISL_569088 |
| NA, not applicable because only one strain was obtained. | ||||||||
All 30 strains from 20 patients (Table 1) and the IHUMI-3 control strain were
screened using the high-content screening procedure for the combination of
hydroxychloroquine and azithromycin (Sigma-Aldrich) at 5
Simultaneously with the antiviral assays, 500
Of the 30 strains (plus the IHUMI-3 control strain) screened 72 hours after
viral infection by SARS-CoV-2 on the high-content screening, with or without
treatment by the combination of hydroxychloroquine and azithromycin both at 5
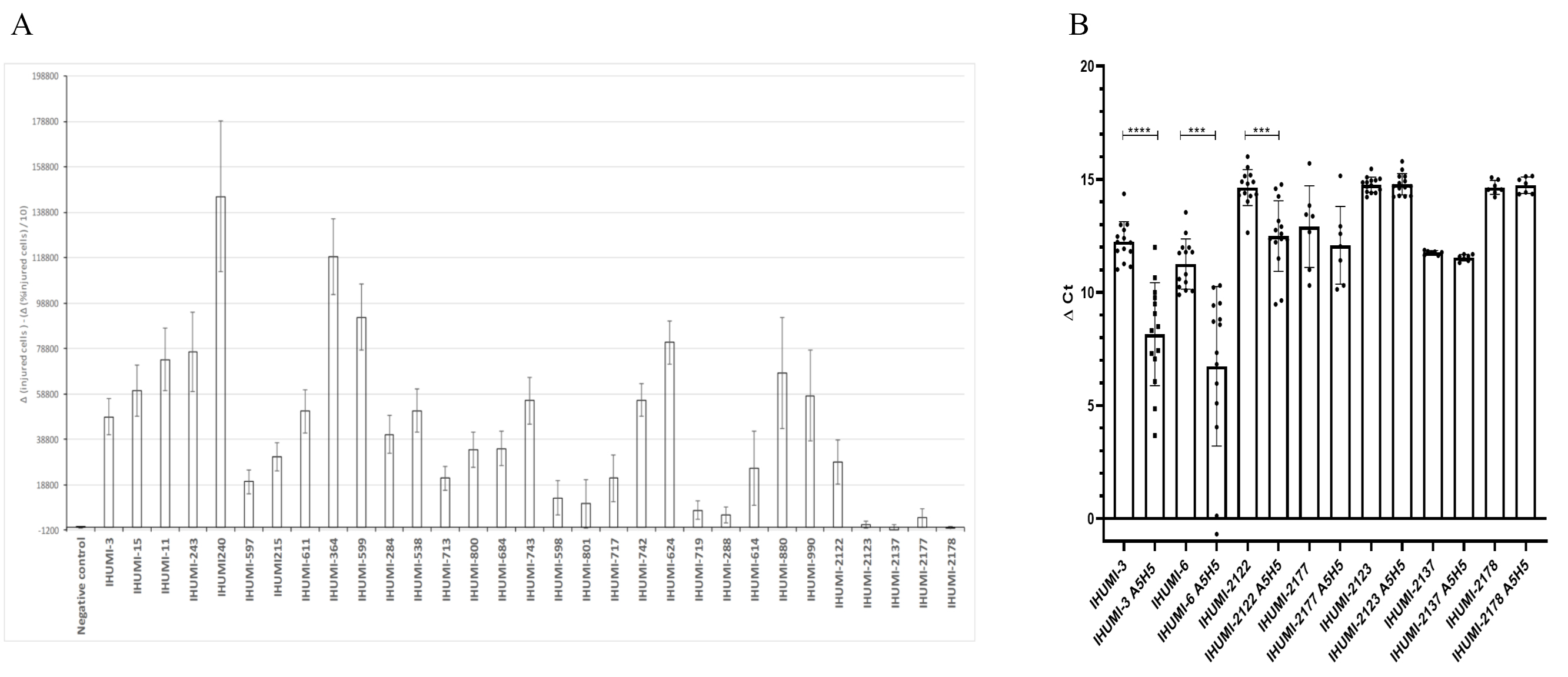 Fig. 1.
Fig. 1.Initial screening of the 31 selected SARS-CoV-2 isolates and a
combination of hydroxychloroquine and azithromycin at 5
Concerning the IHUMI-3 and IHUMI-2122 strains used as controls, we observed
consistent viral inhibition compatible with the results previously observed in
SARS-CoV-2 isolates. In contrast, concerning IHUMI-2123, IHUMI-2137 and
IHUMI-2178, we observed a displacement of susceptibility to hydroxychloroquine
(Fig. 2) confirming a specific pattern of reduced susceptibility for these
isolates. For low concentrations of hydroxychloroquine (
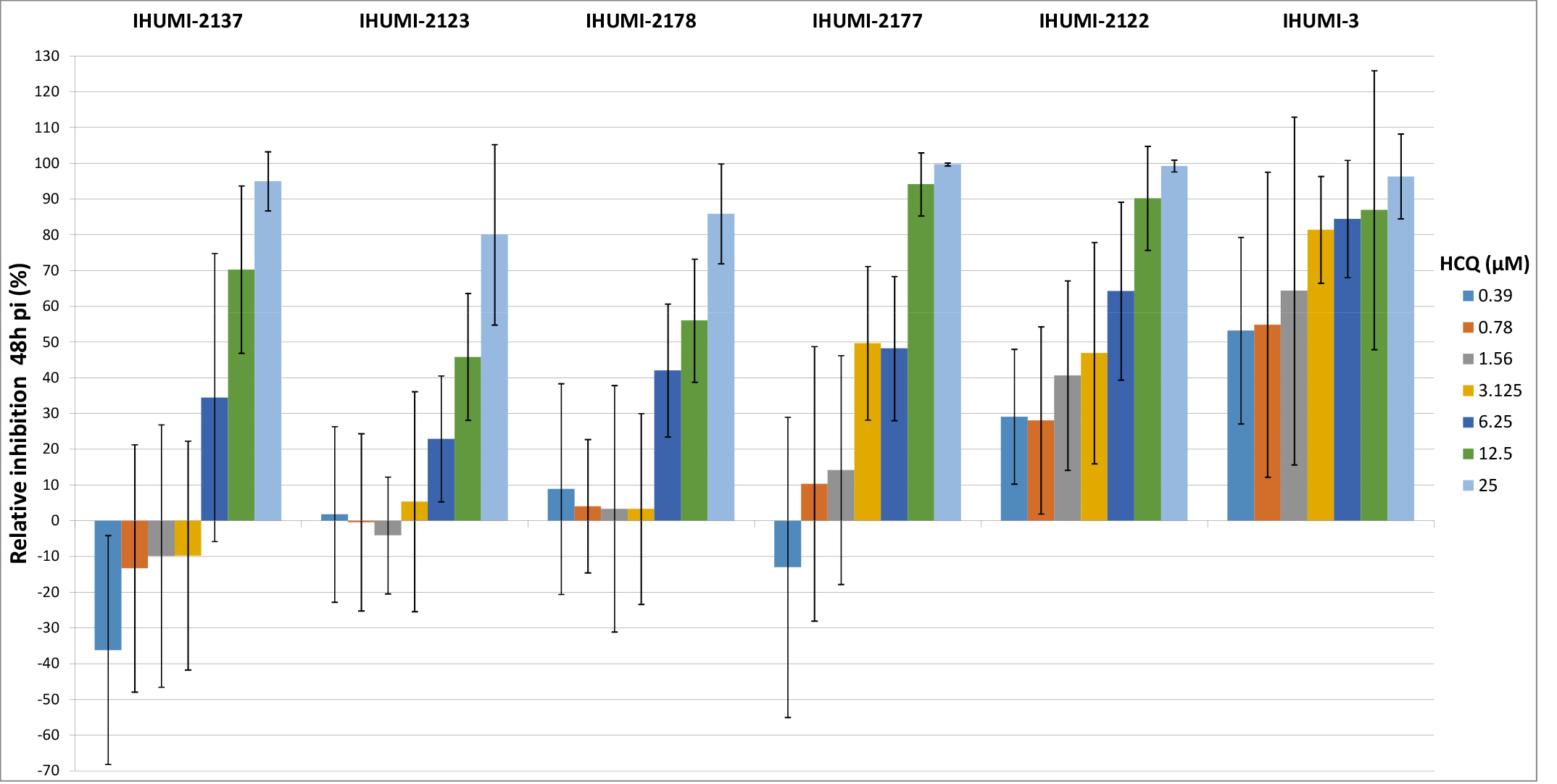 Fig. 2.
Fig. 2.Exploration of effect-dose of hydroxychloroquine. The range
used from 25
We first conduced a global genome-to-genome comparison on the couple of strains isolated in persistent patients and could not detect any modifications (Table 1). We also analysed 20 genomes to place them in a phylogenetic tree. Regarding the quality score on the next clade, all strains received a good quality score (Supplementary File 1). We were able to detect that all the strains from the second wave have ten or more amino-acid changes in their genome, compared to strains from the first wave. In contrast, all the strains of the first period had fewer than ten amino-acid mutations, with one exception (patient 12). All viruses in those studies had the D614G mutation in the spike, described elsewhere as potentially increasing the infectious effects [32]. The phylogenetic tree was reconstructed by integrating all IHUMI strains and evolutionary relationships were revealed (Fig. 3). All five strains from the second period belonged to separate clades, provisionally named Marseille 1 and Marseille 5 [33]. Specifically, the strains with reduced susceptibility to hydroxychloroquine were from Marseille clades 1 (IHUMI-2123 and IHUMI-2178) and 5 (IHUMI-2137). All strains belonging to the Marseille 1 clade were positioned close to genomes originating from Africa, specifically Senegal and Gambia. The phenotype with reduced hydroxychloroquine in Marseille 1 genotype was shared by the IHUMI-2123 and IHUMI-2178 isolates, possibly IHUMI-2177, but not by the IHUMI-2122 isolate. IHUMI-2137 grouped within the Marseille 5 clade. Meanwhile, the IHUMI-2123, IHUMI-2178 and possibly IHUMI-2177 isolates with a reduced susceptibility to hydroxychloroquine, did not present mutations as compared to the IHUMI-2122 isolate without reduced susceptibility.
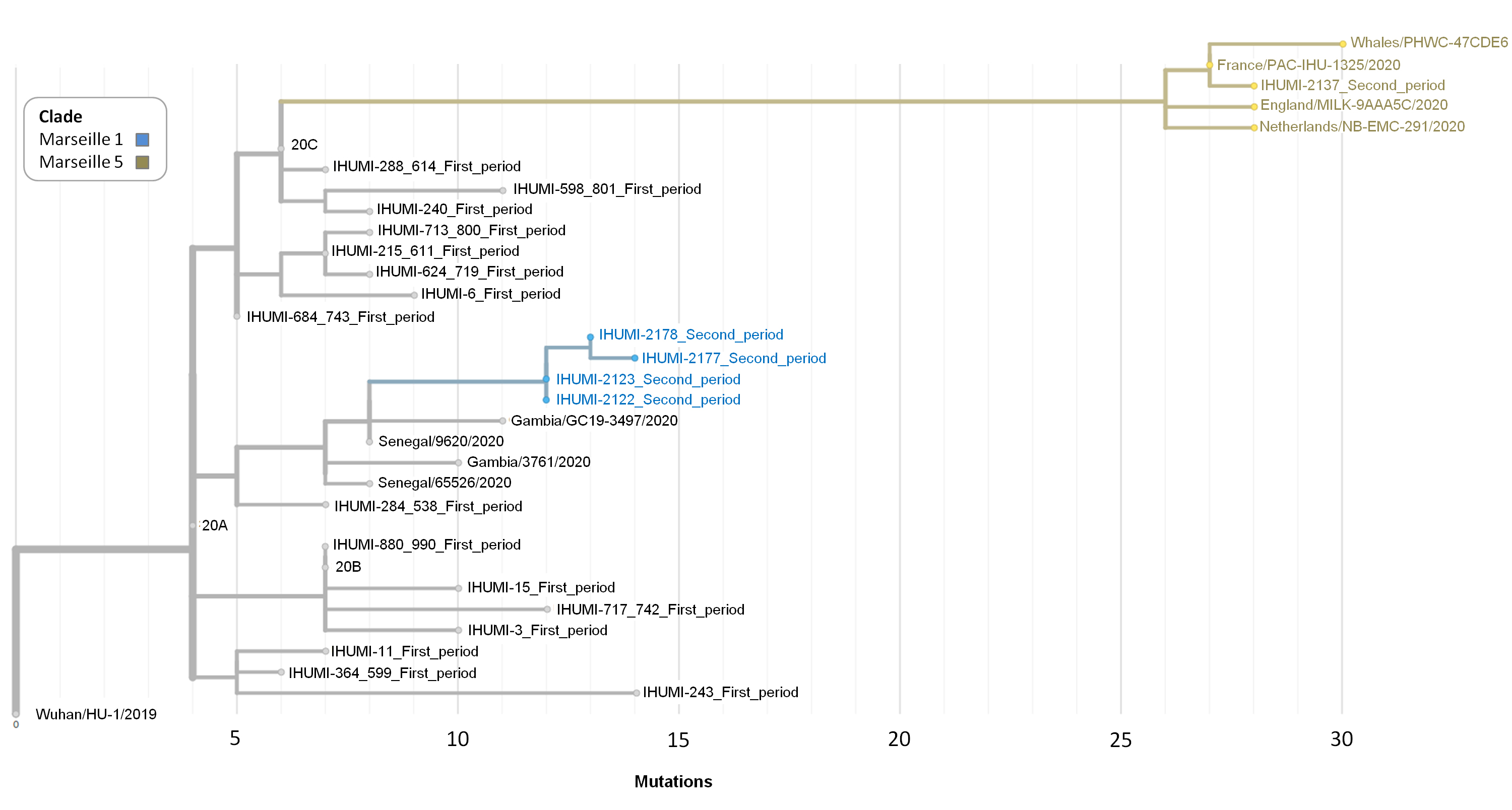 Fig. 3.
Fig. 3.Phylogenetic tree of whole genomes from IHUMI strains including closely related genomes available from GISAID. Mutation scales are compared to the Wuhan reference genome.
To the best of our knowledge, this is the first time that three SARS-CoV-2 strains have been shown to have a profile of reduced susceptibility to hydroxychloroquine in vitro. Susceptibility to azithromycin was not tested independently, as its effect alone in vitro is limited. In this study, the high-content screening technology, first applied to the high-throughput culture of giant viruses of amoeba then to SARS-CoV-2 [30, 34], was used for the first time to rapidly screen for the susceptibility to drugs of a large panel of viruses. Although other research will be needed to clearly confirm that the procedure can be sufficiently standardised to provide efficient large screening of strains, it has been shown to be efficient at detecting isolates with reduced susceptibility. However, although highly time-consuming and susceptible to many confounding factors, as presented below, when it comes to the fine determination of susceptibility, dose-effect determination using molecular biology remains necessary. Indeed, in vitro sensitivity assays carried out on the same virus can provide divergent results, reflecting great discrepancies due to several key determinants in the experiments. First of all, the cell lines used may harbour different permissivity levels, resulting in differences in viral titres, although those used in these assays needed to be permissive. For SARS-CoV-2, the entry step involves the ACE2 receptor and two independent host protease pathways, TMPRSS2 or the cathepsins B/L that activate the spike viral protein. The virus may not use these two pathways in the same manner, and the expression level of these receptors mediating virus entry are differentially expressed according to the cell lines [35]. For example, VeroE6 engineered to express greater amounts of TMPRSS2 has been used elsewhere, resulting in 100-fold higher titres of SARS-CoV-2 [36]. Inversely, viral titres provided by SARS-CoV-2 infected Calu-3 cells (continuous human lung epithelial cell line) are lower than in Vero cells [36]. It could make sense for the sensitivity assays to use the cells physiologically closest to those of the replication site in vivo. From this perspective, primary cells derived from organ explants were used for sensitivity assays and seemed to present a relevant approach. However, variable effects which are donor-dependent on the sensitivity for some tested drugs should be expected due to differences in viral replication and gene expression [37]. Thus, this approach could be a “false” good idea and testing molecules in a coarse model such as in Vero E6 that has genetic defects in terms of interferon production could help provide evidence of such an effect. Secondly, the multiplicity of infection reported for the drug concentration is not standardised. It seems obvious that the higher the MOI, the lower the relative drug concentration, and the more likely the virus can replicate. This MOI is not even mentioned in some studies. Finally, the time of end point evaluation and the method used for assessing viral replication also varies according to the study from one hour to 120 hours [38, 39]. In addition, the assessment of viral replication by PCR or fluorescent assay or visual inspection to monitor cell viability may not have the same sensitivity. For example, when it comes to visual inspections, some permissive cells such as human intestinal epithelial Caco-2 do not produce cytopathic effects after SARS-CoV-2 infection and thus cannot be evaluated using such a method [40]. As a result, it is risky to draw conclusions on the basis of a single sensitivity test, especially when testing a virus with a high genomic variability.
One of the most interesting perspectives should also be to test multiple viral
strains to check the concordance of the results. Currently, in vitro
assays essentially use one or two SARS-CoV-2 strains. Our work suggests it is
risky to draw conclusions on a single sensitivity test when testing a virus with
a high genomic variability. Indeed we observed heterogeneity in the
hydroxychloroquine antiviral activity screening of 30 strains and were able to
detect three strains with a lower susceptibility profile. For isolates from
patients during the first wave of the epidemic, persistence was clearly not
associated with a lowered susceptibility profile to hydroxychloroquine in
vitro. This confirmed the observation that persistence and severity are rather
associated with host factors, as suggested by recent genetic research on COVID-19
severity-associated factors [41, 42] or immunocompromised status [43, 44].
Moreover, genomic analyses did not reveal any modification in these isolates that
could explain persistence, neither in the sequence of the strain isolated upon
admission nor in that of the strain isolated during the course of the disease.
The less susceptible hydroxychloroquine strain, IHUMI-2123, which belongs to
Marseille 1 genotype, was isolated in early summer 2020 at the beginning of the
second wave, from a patient returning from Tunisia [33], a country in which
hydroxychloroquine was massively used [21]. We evidenced a close phylogenetic
proximity between all strains of the Marseille 1 clade (IHUMI-2123, IHUMI-2122,
IHUMI-2178, and IHUMI-2177) with strains isolated in Senegal and Gambia, two
countries which use hydroxychloroquine to treat patients with COVID-19 [45, 46].
We believe that it is possible that the widespread use of hydroxychloroquine in
these countries led to strains being selected with reduced, which were later
transmitted to the Marseille population. This observation is worthy of analysis
on a statistically larger number of strains of this genotype. Paradoxically, the
patients tested in Marseille hospitals during the early summer of 2020 and
infected by isolates of this genotype presented milder infections and lower
mortality than that observed during the first wave of the epidemic, despite the
fact that the viral loads in their respiratory secretions were higher. This
observation raises several questions that will be difficult to resolve, such as
“does the lowered susceptibility to hydroxychloroquine reduce the severity of
infection?” or “is it useful to use hydroxychloroquine in such cases or only in
patients with severity markers or risk factors such as anticoagulant lupus, which
is treated with hydroxychloroquine and the likely efficiency of which is
therefore not due to an antiviral effect?” [47, 48, 49]. Finally, the
hydroxychloroquine concentration to achieve 50% of viral inhibition was around
3.125
However, these genotypic and phenotypic variations could be frequent in the viral populations in the future and could apply to more drugs and need to be considered in the global repurposing strategy. Following the description by Korber et al. [32], we know that the spike population evolved between February and April 2020 and constituted a fast replacing situation by the G614. Recently, a major situation was reported in Denmark, where minks were infected with a strain presenting a few mutations, notably in the spike protein and associated with a selection pressure in a potential zoonotic transfer. Those aspects need to be carefully considered, in terms of testing and using antiviral compounds, but also in terms of epidemiology and vaccination strategy.
To summarise, we observed that the persistence of SARS-CoV-2 in some patients was unrelated to any decrease in the in vitro sensitivity of their strains to hydroxychloroquine. In contrast, we unexpectedly found, in some control strains, a decrease in sensitivity. This characteristic is linked to particular genotypes observed during the second wave of the epidemic in our region and the emergence of the first variants, which were certainly imported from Africa. Finally, it should be recalled that these simple cell models on Vero cells, even if they certainly reflect specific characteristics of the strains tested, are not necessarily relevant for use in humans. In vitro work on other cells such as Calu-3 lung cells [52] or using animal models such as ferrets [53] show a clear divergence of the effect of the drugs tested compared to the Vero model.
Conceptualization—PC, AL and BLS; software—JD, SA and AL; Investigation—CB, MLB, JA, PJ and MG; formal analysis—CB, MLB, JA; writing—original draft preparation—CB, MLB, JA, MG; writing—review and editing—PC, BP, AL and BLS.
The collection of strains was conducted according to the guidelines of the Declaration of Helsinki, and approved by the ethical committee of the University Hospital Institute Méditerranée Infection (No: 2020-029). Patient consent was waived due to the fact that diagnosis by culture was part of routine diagnosis procedure and patients’ data anonymized.
Not applicable.
This research was funded by a grant from the French State managed by the National Research Agency under the “Investissements d’avenir” (Investments for the Future) program with the ANR-10-IAHU-03 (Méditerranée Infection) reference.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://www.imrpress.com/journal/FBL/26/12/10.52586/5043.



