1 Graduate School of Bioscience and Biotechnology, Chubu University, 487-8501 Kasugai, Aichi, Japan
Abstract
Transient transformation is a convenient and less time-consuming method for investigating gene functions compared with production of stable transformants. Most systems for Agrobacterium-mediated plant transformation, regardless of transient or stable transformation, utilize a combination of Agrobacterium strains carrying a helper Ti-plasmid and a binary vector. However, the helper Ti-plasmids which are mega-plasmids with sizes of 200–250 kbp and are difficult to manipulate directly with conventional molecular cloning techniques due to their large sizes.
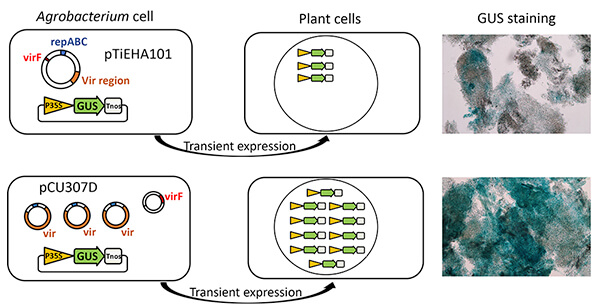
A small helper Ti-plasmid, pCU307D with a size of 46 kbp was constructed from pTiEHA101 which is commonly used for plant genetic engineering. They consist of the virulence (vir) region, the mutated replication origin from pTiEHA101, the gentamicin resistance marker and the replication origin for E. coli cells. The abilities of T-DNA transfer were examined by transient expression in Arabidopsis cultured cells.
Efficiencies to transfer T-DNA to plant cells by EHA101 and C58C1 carrying pCU307D were similar, although pCU307D had 2.8-fold higher copy numbers than EHA101. The construction processes of pCU307D eliminated the A281virF gene encoding F-box-like protein which was located outside of the vir region. The A281virF gene was cloned into a binary vector and it was introduced into C58C1 cells with pCU307D which was named as CU307DF. T-DNA transfer efficiencies by EHA101 or CU307DF were examined by GUS activities from transiently transformed T-DNA with the CaMV35S::NLS-GUS cassette in Arabidopsis cells. Co-culture with CU307DF carrying the GUS gene conferred 3.2-fold higher GUS activities, compared to that with its parental strain EHA101.
A new Agrobacterium strain CU307DF has higher capacity for T-DNA transfer, compared with its parental strain EHA101. pCU307D is as small as 46 kbp and can propagate in E. coli, and that may enable to add further modification to it for further improvement of T-DNA transfer by CU307DF.
Graphical Abstract
Keywords
- Agrobacterium
- Ti-plasmid
- T-DNA transfer
- Arabidopsis
- transient expression
Transgenic plant production is a powerful tool for analyzing gene functions. However, the expression levels in stable transformants depend on the regions of the genome where the transgenes of interest are integrated; therefore, the results complicate the interpretation of transgene expression effects when comparing the phenotypes of transgenic plants [1]. In contrast, transient transformation is a convenient and less time-consuming method for investigating gene functions [2]. The effects caused by transgenes can be compared because they are analyzed before integration into the genome to prevent position-related effects. Currently, agro-infiltration is the most commonly used method for transient expression in plant cells, even though some alternative methods including protoplast transfection, biolistic bombardment, and plant virus infection have been established. Transient expression of exogenous genes in plant cells is also a crucial step for plant genome editing, since it enables to omit a time-consuming step, production of stable transformants. In general genome editing systems, a sequence-specific nuclease such as CRISPR/Cas9 which is transiently expressed creates double-stranded DNA breaks (DSBs) in plant genome and short deletion adjacent to the DSB site is introduced during the process of DSB repairing [3]. However, the efficiencies of gene editing using transient expression of sequence-specific nucleases are not high enough to effortlessly find genome-edited individuals without any selective maker. Raising T-DNA transfer efficiencies is a crucial challenge to improve Agrobacterium-mediated genome editing.
Most systems for Agrobacterium-mediated plant transformation, regardless of transient or stable transformation, utilize a combination of Agrobacterium strains carrying a helper Ti-plasmid and a binary vector [4]. In the systems, the helper Ti-plasmids are disarmed by deletion of T-DNA regions from native Ti-plasmids, on the other hand, binary vectors in which desired genes to be delivered to plant cells as T-strands are inserted between the left border and the right border sequences of T-DNA. In general, the efficiencies of T-DNA transfer depend on the capacity of the helper Ti-plasmids which are mega-plasmids with the sizes of 200–250 kbp and are difficult to manipulate directly with conventional molecular cloning techniques due to their large sizes. Helper plasmids have been constructed by deleting most parts of the T-DNA region and their resulting sizes are 150–200 kbp. One of the smallest helper Ti-plasmid is pTiLBA4404 (Genbank accession: NZ_KY000037) is still 139,561 bp.
The virF gene was originally identified as a gene in the virulence region of nopaline type Ti-plasmid, pTiC58 which was not essential for virulence on most plant species, although it was critical on tomato and tobacco [5]. The virF gene encodes F-box protein involved in ubiquitin pathway. The gene product is translocated to plant cells on infection of Agrobacterium cells and affects on plant defense system [6, 7, 8]. Other type of Ti-plasmids also have genes encoding virF-like F-box proteins [8, 9].
In this study, we constructed a new helper Ti-plasmid with a size of only 46 kbp derived from a hyper-virulent Ti-plasmid, pEHA101. It is consisted of the virulence region, the mutated repABC gene to increase its copy numbers from pEHA101. In addition, a plasmid carrying the putative virF gene from pEHA101 which had been removed during the process to construct the new helper Ti-plasmid improved the efficiencies of T-DNA transfer by Agrobacterium cells to plant cells.
Arabidopsis thaliana Wassilewskija plants were used to produce calli. Root segments from Arabidopsis plants were incubated on the callus inducing medium (CIM) for four weeks to form calli. The obtained calli were then ground into small calli (approximately 1 mm in diameter) in a sterile plastic bag with the bottom of a glass bottle and suspended in 10 mL of MS medium; then, one mL of this callus solution was spread onto the CIM. The calli were subcultured every two-week. The calli were grown at 22 °C under continuous light conditions.
The Murashige-Skoog (MS) medium contained MS salts (FUJIFILM Wako Pure Chemical Corp., Japan), Gamborg’s B5 vitamins (Nacalai Tesque, Japan) 0.05% MES (FUJIFILM Wako Pure Chemical Corp., Japan), 1% sucrose, and CIM contained Gamborg’s B5 salts (FUJIFILM Wako Pure Chemical Corp., Japan), 0.05% MES, 2% glucose, Gamborg’s B5 vitamins, 2 µM 2,4-D, and 0.45% gellan gum. The shoot inducing medium (SIM) contained MS medium, 12.5 µM 2-ip, and 0.45% gellan gum.
The sequences of all primers used in this study are shown in Table 1.
| Primer name | Sequence (5′-3′) |
| Atu0972F | CGGAATACAAGGCGCTGGAAACG |
| Atu0972R | GTCGACAGTGAGGGTATGGGAATATTGC |
| AvrII-A281virF R | AGAGAGCCTAGGCGTTGAAATCGACGGTATCCCGA |
| BamHI-L3 R | AGAGAGGATCCGTCTTCGATGAATTGCGTGATCTG |
| BamHI-repABC F | AGAGAGGATCCGCATCATAATCTCCGCATGAACAG |
| HindIII-cos R | GAGAGAAGCTTCGTTGACATGAGGTTGCCCCGTAT |
| PlacIq R | AGAAAATACCGCATCAGGCGCTCTTC |
| R3-Bg F | GCGATCATGTTGAATGCACCTGATCCTATACC |
| R3-Pst R | GCTACCACCAGGACGAATCGACATTCATC |
| repB CF | TTTCGCTCGGGTATTCGAAGCTGTCAC |
| repB CR | GCGGCCTTTTTGTCGATCATTAACGTCAGG |
| RI-Sbf-L3 F | AGAGAGAATTCCTGCAGGGACCTTCGTTGAGCAGA |
| SbfI-cos F | GAGACCTGCAGGATACTCGCACCGAAAATGTCAGC |
| SpeI-repABC R | AGAGAGACTAGTCTCTTGTCACGGTTAGGAGGCAC |
| virG qF | TAGGTCGTGAAGATGGGCTTGAGAT |
| virG qR | TTGCTCCTAGCTCGAGTGCAACAAC |
| XbaI-trfA F | GAGTTCTAGAATACCAAGTACGAGAAGGACGGCCA |
| XhoI-A281virF F | GAGAGAACTCGAGTGACCCGGGAATTTGCACCCTT |
| Y311H F | CGCTCCCAGAAATCCACGCTTCGTTCAAAAGGTCG |
| Y311H R | CTTTTGAACGAAGCGTGGATTTCTGGGAGCGCCGA |
The VL arm (121681-124216) located downstream of the virulence region in
pTiEHA101 (Genbank accession: NZ_KY000035) was amplified by PCR from pTiEHA101
using primers, RI-Sbf-L3 F and BamHI-L3 R, then digested with EcoRI and
BamHI. The EcoRI-BamHI fragment was inserted between
the EcoRI and BamHI sites in p15AGm (Genbank accession:
LC792546), resulting in p15AGm::VL. Then the cos site which had been amplified by
PCR from bacteriophage
Twenty ng of total DNA extracted from EHA101 (C58C1 carrying pTiEHA101), C58C1 carrying pMD::repABC, C58C1 carrying pMD::Y311H and C58C1 carrying pCU307D was used as a template for the PCR. PCR was performed on the CFX96™ Real-Time PCR Detection System (BIO-RAD, USA) with Premix Ex Taq™ (Probe qPCR) (RR390A, Takara, JAPAN) according to the manufacturer’s instruction. The gene-specific primers were designed to produce 126 (primers; virG qF and virG qR), 117 (primers; repB CF and repB CR) and 146 (primers: Atu0972F and Atu0972R) bp DNA fragments from the genes of virG, repB in Ti-plasmids and Agrobacterium Atu0972 gene used as an internal control, respectively. The sequences of the primers are shown in Table 1. Relative copy numbers were calculated by normalization with those of the Atu0972 gene.
Arabidopsis calli were freshly broken into fine pieces as described in 2.1. Agrobacterium cells were co-cultured with the crushed calli for 48 hours, then the calli were washed with sterilized water three times. The washed calli were incubated on CIM with 100 µg/mL of carbenicillin for a further 24 hours to allow transient expression in plant cells.
The expression assay of GUS reporter gene was done by GUS histochemical assay
and MUG assay in Arabidopsis cells. For GUS histochemical assay, the
transiently transformed Arabidopsis cells were incubated at 37
°C for 4 hours in the GUS staining solution (50 mM Na3PO4 pH
7.2, 0.5 mM K3[Fe(CN)6], 0.5 mM K4[Fe(CN)6], 0.1% Triton
X-100, 0.5 mg/mL 5-bromo-4-chloro-3-indolyl glucuronide (X-gluc), 2mM
MgCl2). After incubation, the buffer was removed and added with a stop
buffer containing 50 mM MES and 50 mM KCl. Microscopic analysis was done to
determine differences between Agrobacterium strains. For the MUG assay,
the Arabidopsis calli were added with 1x Tris Buffered Saline (TBS) and
0.1% Triton x-100. These were squeezed using the disposable homogenizer. This
was centrifuge at 15,000 rpm for 10 minutes and the supernatant was transferred
into a new tube. After, 5 µL of the extract was incubated at 37 °C
for 1 hour in 50 µL 4-methylumbelliferyl
We used the Arabidopsis ecotype Wassilewskija (WS) for our tissue culture experiments, as Czakó et al. [10] have reported that cultured WS cells have the highest efficiency of regenerating shoots among the examined ecotypes. It is known that long-term incubation of Arabidopsis calli on CIM containing 2,4-D inhibits shoot regeneration in SIM containing cytokinin [11]. Our previous study demonstrated that a high cell density of cultured cells caused by extended incubation on CIM had an inhibitory effect on shoot regeneration [12]. To confirm the shoot-regenerating ability of calli that had been maintained for a long time, calli derived from Arabidopsis WS root segments by culturing on CIM were subcultured every two weeks for 26 months. Ground calli (see 2.1) were then grown on CIM for two weeks and then transferred onto SIM (Fig. 1). A large number of shoots were formed on calli that had grown larger. More shoots formed on parts of the calli in contact with the medium (Fig. 1C) compared to those on aerial parts. These results clearly showed that calli that had been subcultured for a long period retained their ability to regenerate shoots on SIM. We chose Arabidopsis calli as plant materials for transient expression experiments since they can be able to be used for model experiments for genome editing.
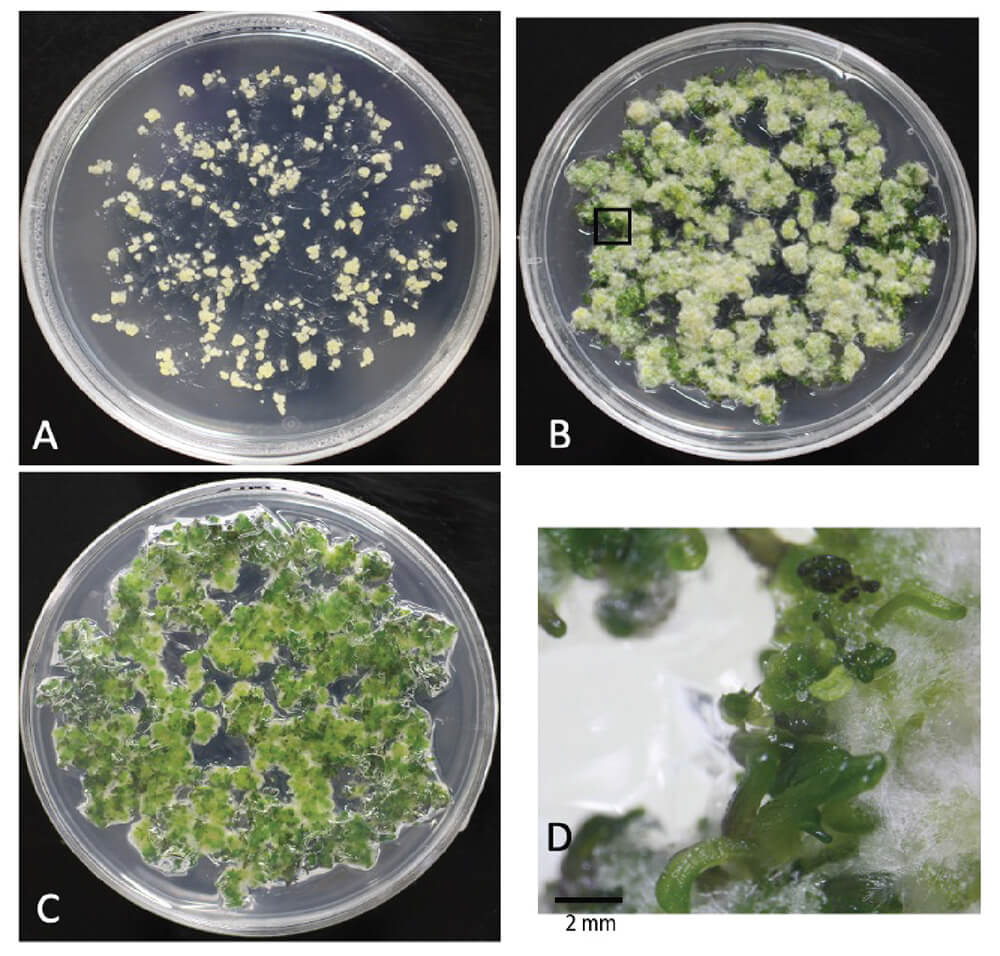 Fig. 1.
Fig. 1.
Shoot regeneration from Arabidopsis calli. Routinely subcultured calli on day 14 after the last subculture were transferred to shoot inducing medium (SIM). Images show identical calli on the SIM. (A) Calli on day 0 after transfer to SIM. (B,C) Calli on day 14 after transfer to SIM. Plate with the calli on the indicated day after transfer to the SIM. Each picture was taken from the table side (B) or the other side (C). (D) Area with black frame in (B). Bar = 2 mm.
In order to improve the efficiencies of T-DNA transfer by Agrobacterium, we attempted to increase the copy numbers of Ti-plasmids in Agrobacterium cells. Ti-plasmids are megaplasmids with sizes of 200–250 kbp and have low copy numbers. The replication origins of Ti-plasmids are the repABC operons which are commonly found in alfaproteobacteria plasmids [13, 14]. EHA101 is called the hypervirulence strain [15] and we chose this strain as the initial material to manipulate. It had been reported that a mutation, Y299H in the repB gene in pRiA4b increased the copy numbers of the plasmids and the alignment of amino acid sequences from several repABC revealed that the tyrosine residue was conserved among a variety of repB proteins [16]. We introduced the same amino acid change, tyrosine to histidine in the repB gene in pTiEHA101 (originated from pTiBo542) (Fig. 2A). Fig. 2B,C shows the comparison of copy numbers in pTiEHA101/C58C1 (common strain name, EHA101), pMD::repABC/C58C1 (wild type repABC gene) and pMD::repH (repH: repABC with the mutation Y311H), determined by qPCR. The copy numbers of pMD::repABC were approximately 7.2-fold higher than those of pTiEHA101, probably owing to the difference of their sizes. The size of pMD::repABC is 6.8 kbp, while that of pTiEHA101 is 189 kbp, suggesting plasmids with smaller sizes give more copy numbers compared to those with larger sizes and the identical replication origin. In addition, introduction of the mutation, Y311H gave approximately twice copy numbers, compared with wild-type repABC and 14.2-fold, and 7.2-fold, compared with pTiEHA101 and pMD::repABC, respectively. The increased ratios caused by the Y311H mutation were much lower (approximately 2.0-fold) than the Y299H mutation (approximately 10-fold) in pRi repABC [16] when plasmids with the same sizes were examined. The effects on copy numbers by reduced sizes were much greater than those by the Y311H mutation.
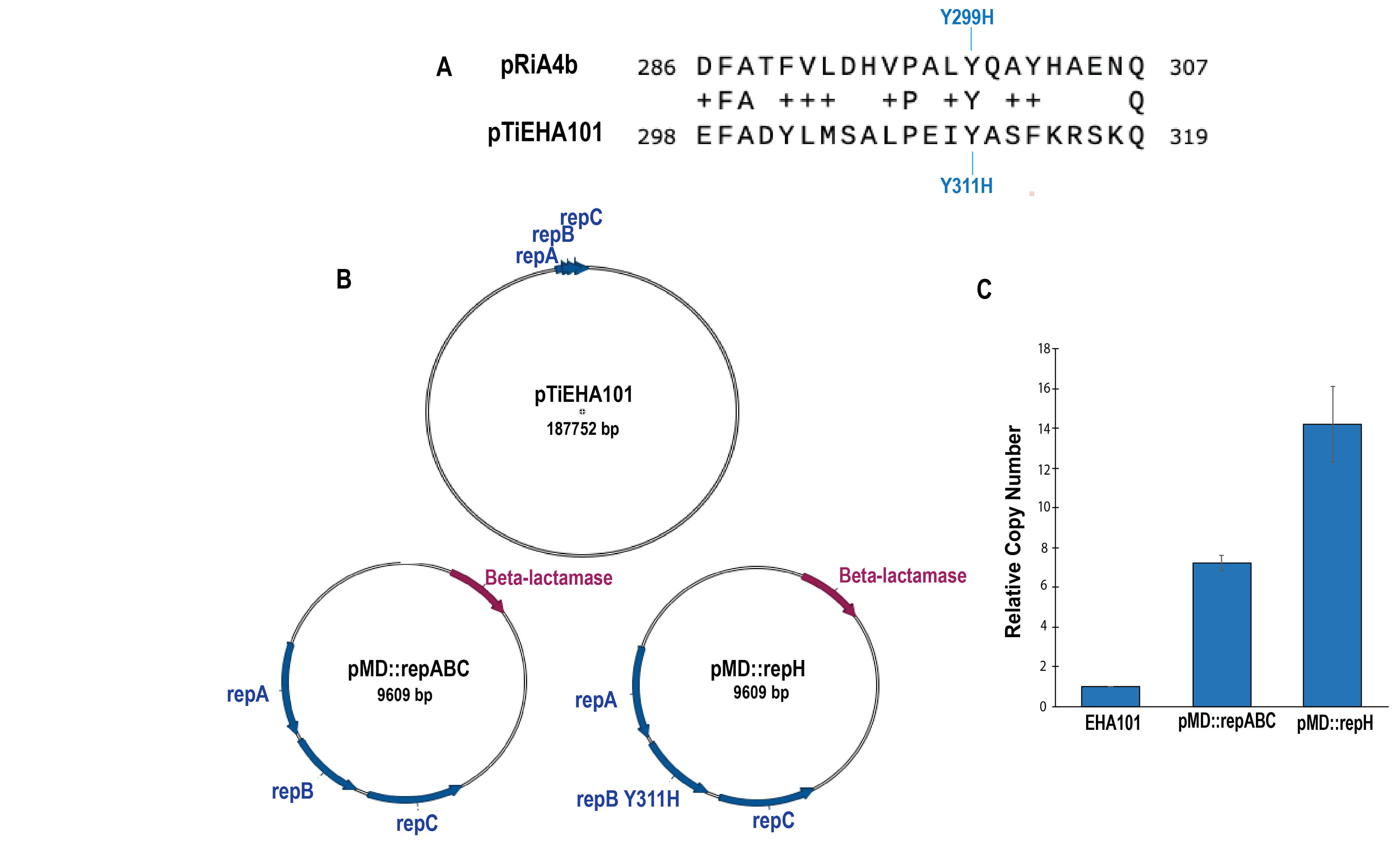 Fig. 2.
Fig. 2.
Relative copy numbers of pTiEHA101-derivatives in Agrobacterium cells. (A) Alignment of wildtype RepB protein sequences in pRiA4b and pTiEHA101 corresponding to mutation sites Y299H and Y311H. (B) Schematic structures of plasmids used in the experiments. EHA101; pTiEHA101, pMD::repABC; a small-sized plasmid with wild type repABC, pMD::repH; a small-sized plasmid with repH. (C) The relative copy numbers were determined by qPCR. The vertical axis shows the plasmid copy number. Relative copy numbers are shown as normalized values to those of EHA101. Experiments were individually performed three times. Error bars represent standard error of the mean.
These results led us to make the size of the helper Ti-plasmid reduce by cloning
only the virulence (vir) region. According to information reported previously
[15], we designed the strategy to clone the vir region by introducing two cos
sites at the ends of the vir region, a selectable marker gene and the p15A
replication origin for E. coli, then packaging as
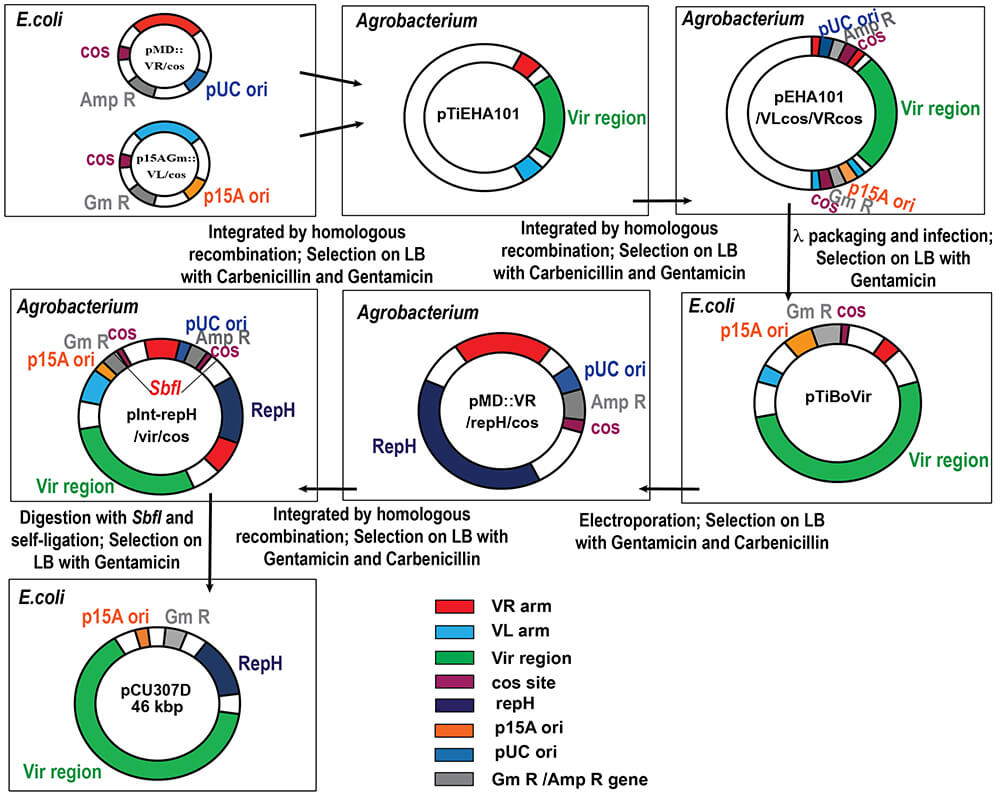 Fig. 3.
Fig. 3.
Schematic diagram of pCU307D construction. Abbreviations indicate the followings: cos, the recognition sequence by lambda terminase; AmpR, the ampicillin/carbenicillin resistance gene; VR, homologous sequence to pTiEHA101 located at 5′ of the vir region; VL, homologous sequence to pTiEHA101 located at 3′ of the vir region; RepH, the replication origin for Agrobacterium cell; SbfI, the recognition sequence by SbfI; p15 ori and pUC ori, the replication origins for E. coli cells.
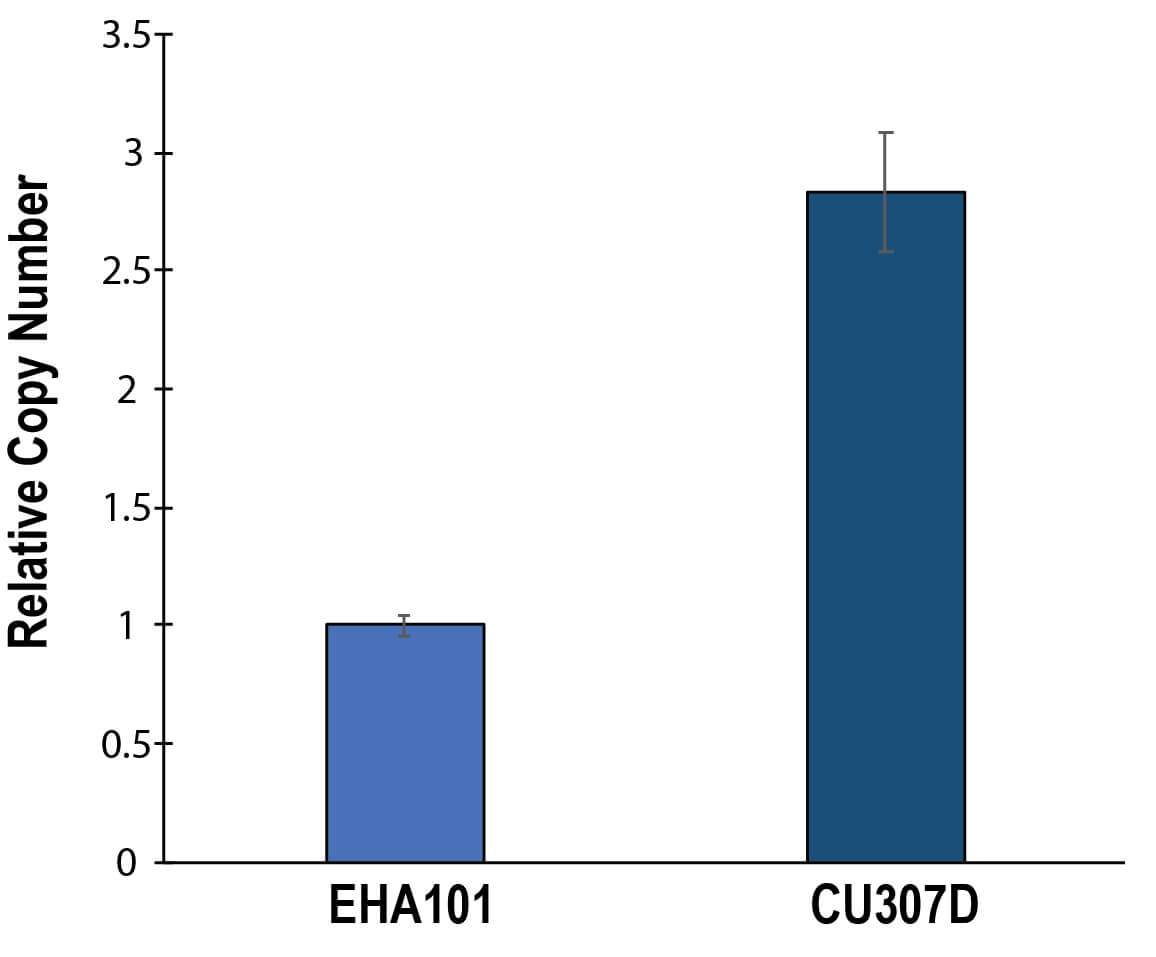 Fig. 4.
Fig. 4.
Determination of relative copy numbers of pCU307D. The relative copy numbers were determined by qPCR. Relative copy numbers were calculated as normalized values to those of EHA101. Experiments were individually performed three times. Error bars represent standard error of the mean.
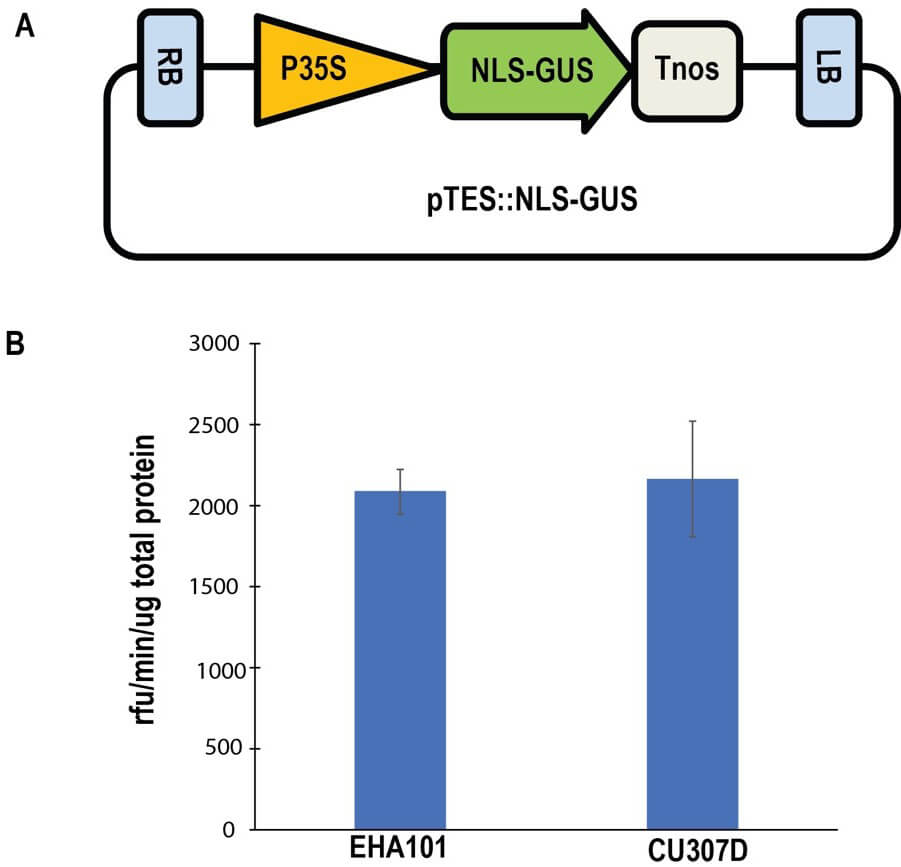 Fig. 5.
Fig. 5.
T-DNA transfer efficiencies by EHA101 and CU307D. (A) A
schematic diagram of the GUS expression construct where the T-DNA region is
flanked by Right Border (RB) and Left Border (LB). The T-DNA transfer
efficiencies were examined by transient expression of
F-box protein encoded by the virF gene in pTiC58 has been reported to enhance transformation efficiencies by Agrobacterium strain, C58 [17]. The virF gene is located in the vir region of pTiC58, while a similar gene encoding an F-box protein (165435-166301 in Genbank accession: NZ_KY000035) in agropine-type Ti-plasmids is located outside of the vir region [9]. In order to examine the effects of the A281virF gene encoding the F-box protein in pTiEHA101 (derived from pTiBo542 in Agrobacterium strain, A281) on T-DNA transfer, a plasmid carrying the gene, pRK2-A281virF was introduced into C58C1 with pCU307D, since it had been eliminated during construction of pCU307D from pTiEHA101. Introduction of pRK2-A281virF into C58C1 carrying pCU307D clearly enhanced transient expression of GUS in plant cells to 3.2 higher (Fig. 6) and the rates are similar to those of their copy numbers. We were not able to judge that the enhancement was caused by more efficient infection by Agrobacterium cells or larger amounts of T-strand transportation from the pictures of GUS staining. We named C58C1 carrying pCU307D and pRK2-A281virF as CU307DF.
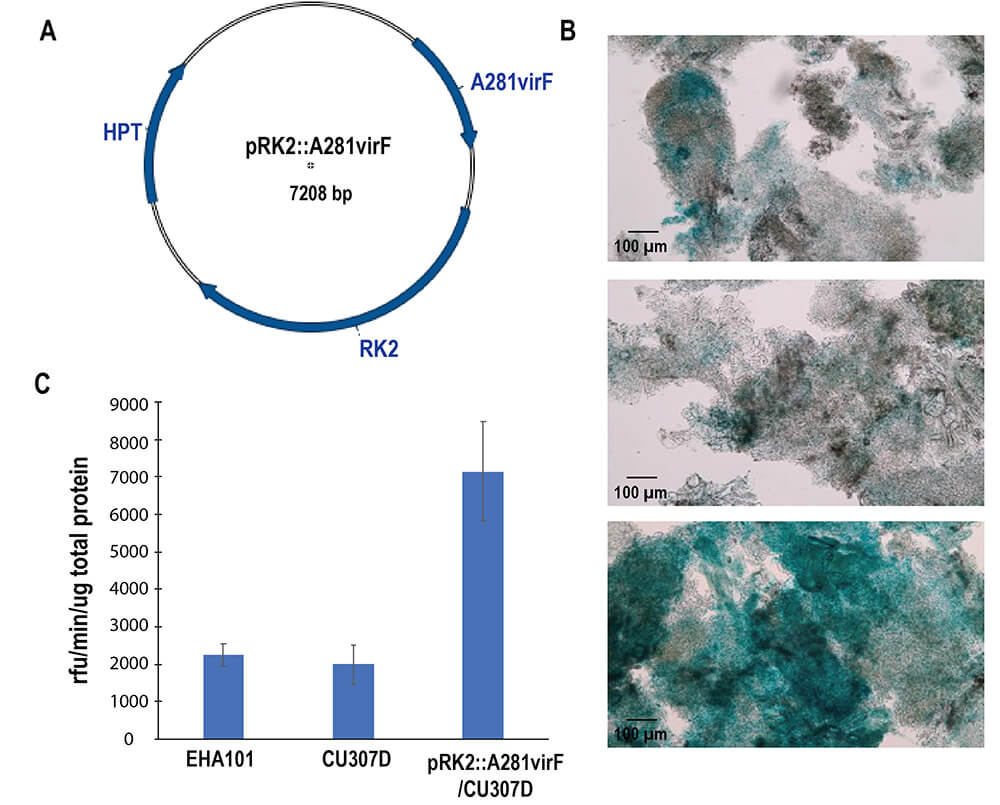 Fig. 6.
Fig. 6.
Effects of the A281virF gene on T-DNA transfer efficiencies. Transient expression of GUS was examined after co-culture of Arabidopsis cells and Agrobacterium strains indicated. All strains carry pTES::NLS-GUS shown Fig. 5A. (A) The A281virF gene was cloned into a vector with RK2 (the replication origin for Agrobacterium) and HPT (the hygromycin resistance gene). (B) Transient expressions of GUS in Arabidopsis cells were visualized by X-gluc staining. Scale bars indicates 100 µm. (C) Quantitative GUS activities in Arabidopsis calli. Experiments were individually performed three times. Error bars represent standard error of the mean.
We investigated several factors which potentially affected on T-DNA transfer by Agrobacterium cells. Smaller sizes of helper Ti-plasmids gave positive effects on copy numbers of Ti-plasmids (Fig. 2). A mutation in the repB gene also had positive effects on T-DNA transfer (Figs. 2,4). Copy numbers of pCU307D were 2.8-fold higher than those of its parental plasmid, pTiEHA101 (Fig. 4). In general higher copy numbers of plasmids enable higher expression of genes carried by the plasmids. However, efficiencies of T-DNA transfer examined by expression of the reporter gene in Arabidopsis cells were similar (Fig. 5). The positive effects of higher copy numbers on T-DNA transfer might be offset by deletion of the A281virF gene. In fact, the A281virF enhanced T-DNA transfer 2.8-fold in GUS activities when the 35S::NLS-GUS was transiently expressed in plant cells, although molecular mechanisms of enhanced T-DNA transfer by A281virF have not been investigated. This enhancement may reflect net effects of copy numbers. Since most genes in pTiBo542 except the vir genes and the repABC gene had been eliminated in pCU307D, the enhancement of T-DNA transfer may suggest expression levels of the vir genes caused by the higher copy numbers. The virF gene in pTiC58 encodes an F-box protein which is thought to be involved in degradation of plant defence proteins (reviewed by Magori and Citovsky [17, 18]). Although functional analyses of A281virF have not reported, structural similarities of these virF products may suggest their common mechanisms in T-DNA transfer by Agrobacterium.
Most binary vectors have the kanamycin resistance gene or the spectinomycin/streptomycin resistance gene as a selective marker. We confirmed CU307DF is sensitive to kanamycin and spectinomycin/streptomycin, therefore, CU307DF is compatible with a variety of binary vectors. We merely developed our transformation system for transient transformation, although it may have disadvantages for stable transformation of plant cells, owing to its high capacity to transfer T-DNA which may cause multiple integration of transgenes. Recently, gene knock-in by transient transfer of the CRISPR/Cas9 system and the donor DNA in protoplast of Nicotiana tobacum has been reported [19]. However, production of intact protoplasts requires special skills and are not necessarily applicable to many plant species. If abilities to transfer T-DNAs by Agrobacterium cells can be drastically improved, it might be going to be possible to transfer T-DNAs including a sequence encoding a sequence-specific nuclease and a donor DNA with homology surrounding the target sequence by the nuclease, leading to establish strategies for knock-ins in plant cells. Our results may be able to the initial step for the strategies mediated by Agrobacterium.
At the initial stage of construction of new helper Ti-plasmids, we attempted the
strategy, SbfI-digestion followed by self-ligation, since there was no
SbfI site in the vir region in pTiEHA101. However, the attempt was not
successful, probably owing to a large amount of non-related DNAs including other
parts of pTiEHA101 than the vir region and a megaplasmid, pAtC58 carried by
EHA101. Then we realized that the size of the vir region was close to the size
which can be accepted by
A newly produced Agrobacterium strain, CU307DF (C58C1 carrying pCU307D and pRK2::A281virF) had enhanced abilities to transfer T-DNA, compared with a conventional hypervirulent strain, EHA101. We consider that CU307DF is a good option as the host strain to deliver constructs for transient expression in plant cells.
The data and materials used in this study are available from the corresponding author.
HB conducted the whole research and wrote the manuscript. NFHA performed all experiments except qPCR and made the figures. MA designed and conducted determination of plasmid copy numbers. NT performed qPCR experiments. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work. All authors contributed to editorial changes in the manuscript.
Seeds of Arabidopsis thaliana ecotype Wassilewskija was obtained from Arabidopsis Biological Resource Center, Ohio State University 24 years ago.
We thank Dr. Ktsunori Suzuki for useful informations about Agrobacterium plasmids.
This research is supported by the Sengoku Private Fund (CU2000).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






