1 Department of Microbiology, Kogi State University, 272102 Anyigba, Nigeria
2 Department of Microbiology, University of Nigeria, 410001 Nsukka, Nigeria
3 Department of Microbiology, Catholic University of Central Africa, 11628 Yaoundé, Cameroun
4 Department of Molecular Biology, Biotechnology, and Biochemistry, Wroclaw University of Science and Technology, 50-370 Wrocław, Poland
5 Discipline of Microbiology, School of Life Sciences, College of Agriculture, Engineering and Science, Westville Campus, University of KwaZulu-Natal, 4000 Durban, South Africa
Abstract
Escherichia coli (E. coli) is the most prominent bacterial pathogen that causes urinary tract infections (UTIs), and the rate of resistance to most used antibiotics is alarmingly increasing.
This study assessed the hostel gutters of two Nigerian universities, the University of Nigeria, Nsukka (UNN) and Kogi State University, Anyigba (KSU), for E. coli and its antimicrobial resistance genes (ARGs). Oxoid Chromogenic UTI agar was used to isolate uropathogenic E. coli (UPEC), identified using standard biochemical tests. The virulence and resistance genes of the isolates were further characterized using molecular techniques.
A total of 906 UPEC were isolated in this study, of which 63 isolates were selected for antimicrobial susceptibility tests. The UPEC isolates showed 100% resistance to amoxicillin/clavulanic acid, vancomycin, and penicillin G, while a complete sensitivity of the isolates to meropenem and ciprofloxacin was observed. The index of isolates showing multidrug resistance ranged from 0.33 to 0.73. The level of multiple drug resistance (MDR) exhibited by the UPEC isolates from effluent was significantly higher compared to those from influent (p < 0.05). The ARGs detected were blaOXA-1 8 (38.1%), blaCTX-M3 8 (38.1%), and ant(2)-la 20 (95.2%). Virulence genes encodings beta-glucuronidase (uidA) and hemolysin A (hlyA) were detected in 95.2% of UPEC isolates.
The current study showed that UPEC is widely distributed in the environment of two Nigerian universities. The index range of MDR and the circulation of ARGs and virulence genes in the environment suggest a potential health concern, thus warranting further investigation.
Keywords
- antibiotic-resistant genes
- antibiotic susceptibility testing
- Escherichia coli
- urinary tract infection
- wastewater environments
Globally, wastewater represents an important hotspot for the rapid dissemination of antibiotic-resistant bacteria (ARB) and antibiotic-resistance genes (ARGs) in the environment [1]. Antimicrobial resistance (AMR) can spread between people, animals, and the environment through different pathways [2]. Significantly, the environment has been identified as a bridge for different transmission pathways, from animals to compost, soil to water, and sediments to sewage. As a reservoir, the environment may simultaneously mix mobile genetic elements (MGEs) before their eventual diffusion into human and animal hosts [3]. Notably, the ARGs are not biodegradable and can be distributed among microbial communities when they find their way into natural environments, which carries additional implications for promoting the emergence of multidrug-resistant organisms. Discharging improperly treated effluent into surface waters greatly impacts the quality of these surface waters. The presence of antibiotics, ARB, and ARGs causes changes to indigenous microbial communities in surface waters and contributes to the acquisition and spread of waterborne diseases in people who use them [1].
Escherichia coli (E. coli) was recently listed among the
high-priority pathogens since their presence may threaten human health [4]. In
addition, the transmission of uropathogenic E. coli strains via
environmental water-related routes remains an emerging concern globally [4].
Notably, multidrug resistance in E. coli is becoming a growing concern
for humans worldwide. Intrinsically, E. coli is vulnerable to virtually
every important clinical antimicrobial. However, the microbe can store resistant
genes, often via horizontal gene transfer [5]. The highest challenging E.
coli mechanisms match with the ability to acquire extended-spectrum
The WHO has rated wastewater treatment plants (WTPs) as the leading cause of environmental contamination by antibiotics [4]. WTP refining has been shown to select antimicrobial-resistant isolates [8], and reports of multidrug resistance are common in hospital environments [9]. Effluents of WTPs, as well as wastewater directly dumped into the University of Nigeria, Nsukka (UNN), and Kogi State University (KSU)’s hostel drains are used to water vegetables as well as other food crops within the area, and this may add to causes of human infections after such vegetables are consumed. Reportedly, these disposal habits have been associated with antibiotic resistance spread via horizontal gene transfer [10]. In the study areas, little or no data regarding the occurrence and distribution of UPEC and associated genes in wastewater environments are available. Thus, it is important to appraise the quality of effluent and surface waters in the area and to determine if wastewater plays a role in the spread of virulence genes and antibiotic resistance. Therefore, the current study was designed to isolate UPEC from water samples obtained from UNN-WTP presumptively and hostel drains at UNN and KSU, determine the antimicrobial susceptibility patterns of the isolates, and further screen the isolates showing multiple drug resistance for resistance and virulence genes using the molecular technique.
The study area comprised two university environments: The University of Nigeria, Nsukka (UNN), and Kogi State University, Anyigba (KSU). UNN is situated at 6°52′10.07′′N and 7°24′2.46′′E in the Southeastern part of Nigeria while KSU is situated at 7°29′3.60′′N and 7°10′51.62′′E in the North-central region of Nigeria (Fig. 1). Irrigation with wastewater in the surrounding of UNN-WTP and hostel drains has been a common practice since 2000 at KSU and 1976 in UNN. One rationale behind selecting these universities is that farming activities go on around the site of the UNN-WTP, and hostel drains and wastewater are scooped to irrigate the vegetable farms during dry periods. In addition, students patronize these farmers for their farm produce, which may pose a risk of infection when contaminated. Notably, an outbreak of UTI and gastrointestinal tract infections was recorded at both campuses due to the interaction with hospital management boards.
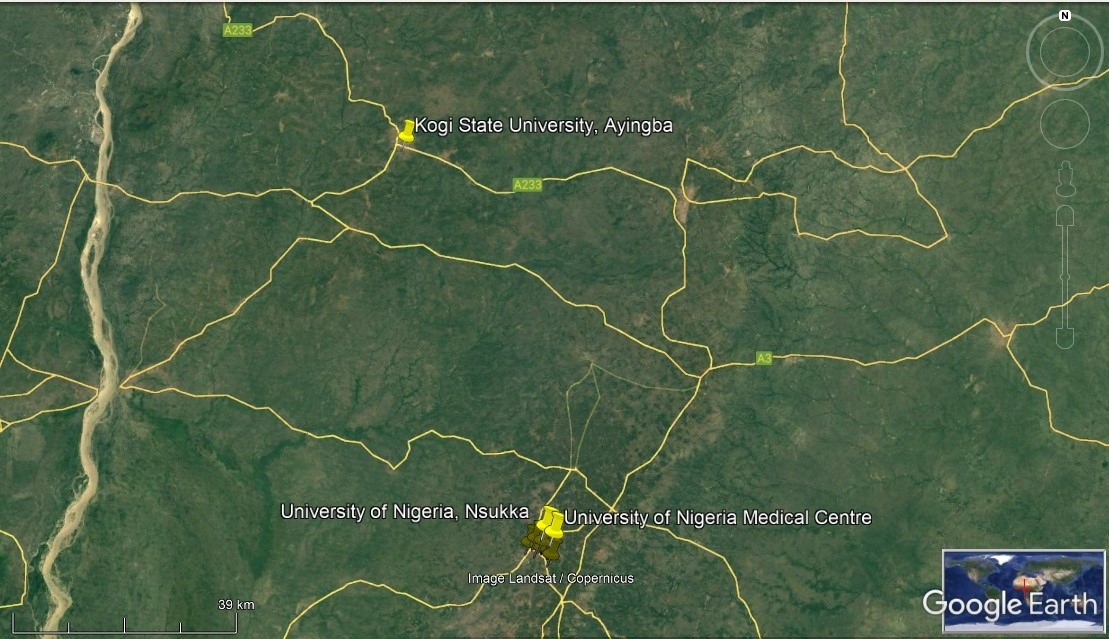 Fig. 1.
Fig. 1.
Map representation of study area/sampling sites. Source: Google Earth (Access date: 25 January 2024).
Grab samples of hostel drains, influent, and effluent of wastewater were obtained from UNN at the marked points, while hostel drain samples were obtained only from KSU. A total of 27 samples and 18 samples were obtained from hostel drains at KSU and UNN, respectively. In the latter location, 9 samples each were collected from WTP influent and effluent. At each point, 250 mL of sample volume was collected into a well-labeled sterile bottle. Samples were then transported in ice boxes to the laboratory for analysis. In total, 63 samples were collected over 9 months.
Chromogenic UTI agar (Oxoid/Thermo Fisher Scientific, Waltham, MA, USA) was employed from each sample to recover UPEC. Gram staining and regular biochemical analyses were conducted on each isolate [11]. Tryptic soy broth (Oxoid, Basingstoke, UK; Catalog number: CM0876) and 15% glycerol (Thermo Fisher Scientific, Waltham, MA, USA, Catalog number: A16205.AP) were used to preserve the presumptive isolates for additional tests and molecular confirmation of the UPEC strains. The glycerol stocks in Eppendorf bottles were stored in a biofreezer (Thermo Fisher Scientific, Model: IUE30086LA) at –80 °C. An E. coli (NCTC 13353) positive control was used. The color (pink/red) shown on UTI chromogenic agar plates was used to identify the organisms.
This test was performed on every confirmed UPEC isolate following the Kirby–Bauer disc diffusion method [12] and the guidelines of the Clinical and Laboratory Standards Institute [13]. Altogether, 15 antibacterial susceptibility testing discs (Oxoid Thermo Fisher Scientific) were used to test the susceptibilities of the isolates to ciprofloxacin (CIP 5 mcg), amoxicillin/clavulanic acid (AMC 30 mcg), gentamycin (CN 30 mcg), ampicillin (AMP 10 mcg), meropebem (MEM 10 mcg), streptomycin (S 10 mcg), erythromycin (E 15 mcg), cefotaxime (CTX 30 mcg); sulphamethoxazole/trimethoprim (SXT 25 mcg), vancomycin (VA 30 mcg), levofloxacin (LEV 1 mcg), penicillin G (P 10 IU), ceftazidime (CAZ 30 mcg), nitrofurantoin (F 300 mcg), and tetracycline (TE 30 mcg).
Genomic DNA was extracted from each isolate pure culture grown on nutrient agar
for 24 hours (37 °C) via the traditional method of boiling [14]. The
template DNA was briefly prepared by suspending a loopful of bacterial cells from
an overnight culture in 1 mL of sterile distilled water. The suspensions of
bacterial cells were then heated (100 °C for 5 min) and centrifuged
(12,000
PCR assays were performed using MyGene Gradient Thermal Cycler (Model MG96G; LongGene Scientific Instruments Co., Ltd., East Lyme, CT, USA), as described elsewhere [15, 16]. The 50 µL total reaction volume contained 0.5 µL each of forward and reverse primers (Inqaba Biotechnological Industries, Pretoria, South Africa), 25 µL of the PCR master mix (Thermo Scientific), 14 µL nuclease-free water, and 10 µL of the template DNA. The PCR cycling conditions included an initial activation at 94 °C for 3 min, followed by 30 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, and extension at 74 °C for 1 min. Final elongation was at 74 °C for 9 min following the method of Iredell et al. [15]. E. coli strain (NCTC 13353) was utilized as the positive control for the generic identification of E. coli. Table 1 (Ref. [15, 16]) contains the genes targeted, oligonucleotide sequence primers used, and the expected amplicon sizes.
| Organism/pathogen | DNA target | Virulence factor/gene product | Primer sequence (5′ to 3′) | Product size (bp) | PCR cycling conditions | References | |
| E. coli | uidA | Beta-D-glucuronidase | F | AAAACGGCAAGAAAAAGCAG | 147 | Initial activation at 94 °C for 3 min, 30 cycles comprising denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, extension at 74 °C for 1 min and final elongation at 72 °C for 9 min | [15, 16] |
| R | ACGCGTGGTTACAGTCTTGCG | ||||||
| UPEC | hlyA | Hemolysin A | F | TGTTGAAAGATCAGTCCTCA | 500 | Same as above | |
| R | CTGCGTAGATATTGGCTGAG | ||||||
Note: Gel electrophoresis consisted of template DNA (10 µL), 6
µL positive control as well as 3 µL DNA ladder (ladder of
DNA+1 kb; Invitrogen, Waltham, MA, USA), 2.5% (w/v) agarose gels at 100 V (30
min) in 1
As described elsewhere, a hot start PCR technique was used to screen the
isolates for bla OXA, bla CTM1, and ant(2)-Ia resistance genes [14].
Table 2 (Ref. [16, 17]) shows the expected primers and amplicon sizes. The 20
µL total PCR reaction volume contained 1.5 µL template,
10 µL 1
| Primers name | Primers sequence (5ʹ–3ʹ) | Amplicon size (bp) | PCR conditions | References |
| ant(2)-laF | CATCATGAGGGAAGCGGTG | 787 | Initial activation step (94 °C for 5 min); 30 cycles: denaturation (94 °C for 1 min), annealing (60 °C for 1 min), extension (74 °C for 2 min); final elongation (74 °C for 15 min) | [16, 17] |
| ant(2)-laR | GAGTACCTTGGTGATCTCG | |||
| blaCTX-M1F | AAAAATCACTGCGCCAGTTC | 585 | Same as above | |
| blaCTX-M1R | AGCTTATTCATCGCCACGTT | |||
| blaOXA-F | TCAACAAATCCCCAGAGAAG | 276 | Same as above | |
| blaOXA-R | TCCCACACCAGAAGAACCAG |
Data were analyzed using SPSS version 20 (IBM Corp., Armonk, NY, USA). The
chi-square test or one-way analysis of variance (ANOVA) was used where
appropriate to compute p-values. p values of
The influent from UNNWTP had a higher viable UPEC count (mean count:
3.3
| Month/year | Inf | Eff | Nkr | Alv | Dan | Och | Ink | Mean |
| July 2017 | 3.2 |
3.5 |
3.1 |
1.5 |
2.3 |
4.2 |
4.0 |
3.5 |
| August 2017 | 4.0 |
5.3 |
1.0 |
2.3 |
4.0 |
3.8 |
3.9 |
4.5 |
| September 2017 | 4.9 |
3.3 |
1.2 |
9.0 |
3.6 |
3.2 |
3.5 |
5.4 |
| October 2017 | 5.4 |
4.5 |
2.5 |
5.2 |
3.2 |
4.5 |
3.7 |
5.9 |
| November 2017 | 5.6 |
5.8 |
2.2 |
1.0 |
2.9 |
4.2 |
2.0 |
5.7 |
| December 2017 | 5.2 |
4.8 |
3.0 |
3.0 |
3.8 |
3.6 |
3.6 |
5.7 |
| January 2018 | 6.3 |
3.5 |
2.8 |
4.5 |
3.6 |
3.2 |
3.5 |
9.9 |
| February 2018 | 5.8 |
5.2 |
1.6 |
1.0 |
2.9 |
4.2 |
2.0 |
6.3 |
| March 2018 | 4.5 |
4.8 |
1.8 |
3.0 |
3.8 |
3.6 |
3.6 |
9.3 |
Note: Inf, influent; Eff, effluent; Nkr, Nkruma female hostel; Alv, Alvan Ikoku male hostel; Dan, Dangana male hostel; Och, Ocheja male hostel; Ink, Inikpi female hostel.
The water samples from female hostel drains of KSU had a higher number of viable
UPEC count than those from the UNN female hostel (mean colony count: 3.3
Fluctuations were observed in the UPEC mean colony counts across the sampling
months. Microbial counts of over 108 were recorded in all nine months for
influents isolates, and in the effluents, more than 107 counts were recorded
in August (5.3
A total of 906 UPEC were isolated in this study, of which 147 isolates were from
samples drawn from UNN hostel drains (55 from Alvan Ikoku, 92 from Nkrumah), 167
were from UNN-WTP influent, and 192 were from UNN-WTP effluent. The remaining 400
isolates were from KSU hostel drains, with the most isolates (150) recorded from
Inikpi, followed by Ocheja (131) and Dangana (119). A total of 63 isolates were
selected for antibiotics susceptibility tests from 906 isolated E.coli,
and 21 were selected for molecular analyses. The 63 UPEC isolates were chosen to
represent a number of samples collected from seven sites over 9 months (7
| Treatments | Influent | Effluent | Statistical significance | ||||
| Antibiotics | Resistant | Sensitive | Intermediate | Resistant | Sensitive | Intermediate | Chi-square |
| AMC | 9 | 0 | 0 | 9 | 0 | 0 | NA |
| CIP | 0 | 9 | 0 | 0 | 9 | 0 | 0.331 |
| CN | 7 | 1 | 1 | 7 | 1 | 1 | 0.929 |
| AMP | 8 | 0 | 1 | 8 | 0 | 1 | 0.131 |
| MEM | 0 | 9 | 0 | 0 | 9 | 0 | NA |
| S | 8 | 0 | 1 | 8 | 0 | 1 | NA |
| E | 8 | 1 | 0 | 8 | 1 | 0 | NA |
| CTX | 0 | 9 | 0 | 0 | 9 | 0 | NA |
| SXT | 0 | 0 | 0 | 0 | 0 | 0 | NA |
| VA | 9 | 0 | 0 | 9 | 0 | 0 | NA |
| LEV | 2 | 7 | 0 | 2 | 7 | 0 | NA |
| P | 9 | 0 | 0 | 9 | 0 | 0 | NA |
| CAZ | 0 | 9 | 0 | 0 | 9 | 0 | NA |
| F | 2 | 8 | 0 | 2 | 8 | 0 | NA |
| TE | 8 | 0 | 1 | 8 | 0 | 1 | 0.303 |
Note: WTP, Waste water treatment plant; UNN, University of Nigeria Nsukka; NA, not applicable; AMC, amoxicillin/clavulanic acid; CIP, ciprofloxacin; CN, gentamycin; AMP, ampicillin; MEM, meropebem; S, streptomycin; E, erythromycin; CTX, cefotaxime; SXT, sulphamethoxazole/trimethoprim; VA, vancomycin; LEV, levofloxacin; P, penicillin G; CAZ, ceftazidime; F, nitrofurantoin; TE, tetracycline.
The antibiotic susceptibility test results showed that the UPEC isolates were 100% resistant to AMC, VA, and P. In decreasing order, the antibiotic resistance of the E. coli isolates was 95.2%, 88.9%, 69.8%, 66.7%, and 65.1% against E, S, AMP, TE, and CN, respectively. Conversely, the E. coli isolates were most sensitive to MEM (100%), CTX (87.3%), CIP (81.0%), and SXT (85.7%) (Supplementary Fig. 1).
The E. coli isolates from both campuses demonstrated a 100% resistance to AMC, S, and P. In general, the isolates from the KSU water drains showed more resistance to the antibiotics compared to those from UNN (Supplementary Fig. 2). However, no significant difference was observed in the antibiotic resistance profiles between UPEC isolates from both universities (p = 0.09) (Table 5).
| Treatment | KSU | UNN | Statistical significance | ||||
| Antibiotics | Resistant | Sensitive | Intermediate | Resistant | Sensitive | Intermediate | Pearson chi-square |
| AMC | 27 | 0 | 0 | 18 | 0 | 0 | 0.3407 |
| CIP | 0 | 22 | 5 | 0 | 18 | 0 | NA |
| CN | 17 | 9 | 1 | 9 | 0 | 9 | 0.0003 |
| AMP | 20 | 7 | 0 | 16 | 2 | 0 | 0.6089 |
| MEM | 0 | 27 | 1 | 18 | 0 | 0 | NA |
| S | 27 | 0 | 0 | 18 | 0 | 0 | NA |
| E | 27 | 0 | 0 | 17 | 1 | 0 | NA |
| CTX | 0 | 27 | 0 | 6 | 12 | 0 | NA |
| SXT | 2 | 24 | 1 | 0 | 18 | 0 | 0.4960 |
| VA | 27 | 0 | 0 | 18 | 0 | 0 | NA |
| LEV | 3 | 24 | 0 | 4 | 12 | 2 | NA |
| P | 27 | 0 | 0 | 18 | 0 | 0 | NA |
| CAZ | 5 | 22 | 0 | 18 | 0 | 0 | NA |
| F | 8 | 19 | 0 | 0 | 18 | 0 | 0.0358 |
| TE | 19 | 6 | 2 | 9 | 5 | 4 | 0.0580 |
Note: NA, not applicable; KSU, Kogi State University; UNN, University of Nigeria, Nsukka.
In this study, antimicrobial resistance to AMC, VA, and P was generally observed. The index of isolates showing multidrug resistance ranged from 0.33 to 0.73, with a broad range discovered in isolates from both KSU female (Inikpi) and male (Ocheja) hostel drains (Table 6). The multiple antimicrobial drug resistance (MAR) index was calculated as the ratio of the number of antibiotics to which an organism is resistant against the total number of antibiotics to which the organism is exposed.
| Sampling Area | Campus | Sampling point | Resistant isolates | Antibiotic resistance | Number of antibiotics resisted (%) | MAR index range | |
| Female hostels drains | UNN | Nkrumah | UNKEC3 | UNKEC10 | MEM, CAZ, P, CTX, E | 5 (33.3) | 0.33–0.67 |
| UNKEC12 | UNKEC9 | ||||||
| UNKEC11 | |||||||
| KSU | Inikpi | KIKEC9 | KIKEC12 | MEM, CAZ, P, CTX, E, CN | 6 (40.0) | 0.33–0.73 | |
| KIKEC8 | KIKEC3 | ||||||
| KIKEC1 | KIKEC10 | ||||||
| Male hostels drains | UNN | Alvan Ikoku | UAVEC2 | UAVEC9 | MEM, CAZ, P, CTX, E | 5 (33.3) | 0.33–0.67 |
| UAVEC10 | UAVEC1 | ||||||
| UAVEC3 | |||||||
| KSU | Ocheja | KOCEC3 | KOCEC1 | MEM, CAZ, P, CTX, E | 5 (33.3) | 0.33–0.73 | |
| KOCEC2 | KOCEC8 | ||||||
| KOCEC11 | |||||||
| KSU | Dangana | KDGEC8 | KDGEC7 | MEM, CAZ, P, CTX, E | 5 (33.3) | 0.33–0.67 | |
| KDGEC10 | KDGEC2 | ||||||
| KDGEC12 | |||||||
| WTP | UNN | Influent | UIFEC8 | UIFEC3 | MEM, CAZ, P, CTX, E, | 5 (33.3) | 0.33–0.5 |
| UIFEC9 | UIFEC11 | ||||||
| UIFEC10 | UIFEC1 | ||||||
| UNN | Effluent | UEFEC3 | UEFEC1 | MEM, CAZ, P, CTX, E, | 5 (33.3) | 0.33–0.53 | |
| UEFEC9 | UEFEC8 | ||||||
| UEFEC2 | UEFEC7 | ||||||
| UEFEC10 | |||||||
Note: UIF, UNN-WTP influent; UEF, UNN-WTP effluent; UAV, UNN Alvan Ikoku male hostel; UNK, UNN Nkruma female hostel; KDG, KSU Dangana male hostel; KOC, KSU Ocheja male hostel; KIK, KSU Inipki female hostel; EC, E.coli; 1, January 2018; 2, February 2018; 3, March 2018; 7, July 2017; 8, August 2017; 9, September 2017; 10, October 2017; 11, November 2017; 12, December 2017; MAR, multiple antimicrobial drug resistance.
Twenty-one isolates showing resistance to multiple antibiotics were screened for antibiotic-resistance genes using a molecular approach. In 100% of the isolates, uidA and hlyA genes were detected, confirming they are from E. coli and UPEC. ARGs detected were blaOXA-1 8 (38.1%), blaCTX-M3 8 (38.1%), and ant(2)-la 20 (95.2%) (Table 7). The gel results of the virulence and antibiotic-resistance genes of UPEC isolates identified in the study are shown in Supplementary Figs. 3,4,5,6,7.
| Genes | Present no. (%) | Absent no. (%) |
| ant(2)-la | 20 (95.2) | 1 (4.8) |
| blaOXA-1 | 8 (38.1) | 13 (61.9) |
| blaCTXM3 | 8 (38.1) | 13 (61.9) |
The presence of pathogens in the environment is a great challenge because of their negative impact on human health. The presence of E. coli in the environment is frequently used to predict pathogen presence in the environment, most importantly, surface water. Evidently, WTP has been identified as a common source of E. coli contamination for most surface water worldwide [18]. Nevertheless, there is currently no documented information regarding the presence of this pathogen and the occurrence of its antimicrobial resistance genes in the study area.
In the present study, various degrees of resistance to antibiotics (100%: AMC, VA, and P; 95.2% against E; 88.9% against S; 69.8% against AMP; 66.7% against TE; 65.1% against CN) were observed, which suggest that the circulation of this environmental strains may be contributing to antimicrobial resistance spreads in the area. The high resistance levels to AMC, VA, and P observed with UPEC strains in the current study have been previously documented [7]. Previously, 100% resistance to tetracycline, amoxicillin/clavulanic acid, vancomycin, ciprofloxacin, penicillin, erythromycin, and gentamycin have been reported [19], corroborating the observations in our current study except for ciprofloxacin, which indicated a complete sensitivity. Various levels of resistance to some classes of antibiotics have been observed elsewhere. For instance, a study by Pokharel et al. [18] showed that tetracycline was the least potent, followed by ampicillin. A similar antibiotic resistance pattern was reported in more recent studies by Karam et al. [19] and Whelan et al. [20].
MDR has been regularly recorded among UPEC strains in Nigeria [17, 21, 22]. In the current study, the UPEC isolates from the KSU hostel drains had a higher multiple antimicrobial resistance range (0.33–0.73) than those from UNN (0.33–0.67). The maximum level (66.7%–73%) of multidrug resistance demonstrated by the isolates in our study is comparable with the 77% MAR level in E. coli strains previously isolated from a wastewater treatment facility [23]. According to Osundiya et al. [23], the level of non-susceptibility to amoxicillin/clavulanic acid demonstrated by the isolates in this study remains a regular occurrence in E. coli. Previously, Tang et al. [24] determined antibiotic resistance profiles of E. coli strains from a river watershed in France, and the study demonstrated antimicrobial resistance in 42% of the E. coli isolates, of which 35% exhibited MAR—the MAR index range among UPEC isolates in this study is considerably higher than those published by Tang et al. [24].
The level of potential UPEC strains observed in association with UNN-WTP influents and effluents is above the acceptable threshold. This observation may relate to the influx rates differential of domestic and industrial effluent into the plant, inefficient wastewater treatment processes, and non-adherence to standard treatment protocol. In our study, the findings of a significantly higher MAR in connection with UPEC strain from UNN effluent discharges support the assertion of Fouz et al. [2], who posited that high levels of ARB, MDR strains, and diverse ARGs are common in wastewater worldwide. Unsurprisingly, many studies have shown that effluent samples collected from urban, environmental, hospital, and pharmaceutical-treated wastewater still contain elevated levels of diverse ARGs, ARB, and antimicrobial drugs. For instance, a recent study demonstrated that the abundance of ARGs was significantly higher in effluent wastewater samples than in influents collected from low-income countries compared to high-income countries [25]. The spotting of potentially pathogenic E. coli in effluent from WTP could be associated with the inefficient removal of this bacteria during the treatment process. In another study, potentially pathogenic E. coli bacteria were reported in samples from a wastewater treatment facility at Hawassa University Referral Hospital [26]. According to the authors, the treatment could only remove larger dirt, leaving more organic matter for the remaining pathogens to use as substrates, further increasing the virulence and pathogenicity of pathogens [26].
A relatively high prevalence (96%) of E. coli that produces ESBL in wastewater of a rural and urban hospital in central India has been reported [10, 26]. In our study, the presumptive UPEC isolates showing multidrug resistance were all found to contain beta-beta-D-glucuronidase (uidA) and hemolysin A (hlyA). The presence of the uidA marker genetically confirmed that the isolates were of the genus E. coli [19], which, in this study, were phenotypically typed as UPEC strains. The hlyA detected in the study is commonly found in most uropathogenic E. coli (UPEC) strains reported worldwide [27]; thus, it raises serious concerns about the E. coli strains circulating in the study environment. In several case-based surveillance studies, a significant percentage of UPEC isolates have been reported to secrete hemolysin, which is directly associated with the onset of urinary tract infections (UTIs) [27]. This strain has been reported to cause acute renal failure in renal transplant patients and healthy individuals [28]. The UPEC-bearing hlyA gene was documented to increase cellular toxicity and urothelial damage [27, 28]. The gene (uidA) detected in the isolates codes for beta-D-glucuronidase, an enzyme that catalyzes the cleavage of numerous varieties of 3-glucuronides [29]. This enzyme is why the uidA gene is often employed as a marker for E. coli in water quality determination. Hence, the uidA PCR method is recommended for use because of its sensitivity for specific detection in substrate tests of E. coli by the US Environmental Protection for water quality monitoring [29].
The findings of ARGs comprising the blaOXA-1, blaCTX-M3, and ant(2)-la genes in the isolates suggest that their circulation in the environment may contribute to drug resistance in the area. Unlike the equal circulation of blaOXA genes in the study, Öztürk and Murt [28] and Bush and Jacoby [29] have reported the occurrence of blaOXA-and blaCTX-M3, respectively, as predominant genes in the environmental reservoirs. However, This result differs from the report of Chandran et al. [30], who reported blaCTX-M more frequently compared to blaTEM and ant(2)-la in isolates of E. coli from hospital-associated wastewater in India. In Enterobacteriaceae that produce ESBL, genes blaTEM, blaCTXM group1, bla SHV, and blaCTXM9 have been detected in effluents of hospitals [31]. The ESBLs detected in the study, especially CTX-M enzymes, represent a significant health concern in clinical practice [32]. As previously reported [33], the ESBL-producing UPEC strains currently account for most community and healthcare-associated infections. However, our study has a limitation, as the lack of a separate set of virulence genes in UPEC makes it difficult to ascertain the true pathogenic potential of the presumptive UPEC strains from environmental reservoirs [33]. Consequently, uncertainty may exist regarding determining the actual public health risks associated with the presence of presumptive UPEC isolates in wastewater environments.
The current study confirms the circulation of potential uropathogenic E. coli in the various wastewater environments of two Nigerian tertiary institutions. Generally, more UPEC was isolated from KSU than the UNN campus. The UPEC strains were completely susceptible to meropenem and 100% non-susceptible to amoxicillin/clavulanic acid, vancomycin, and penicillin G. The high rate of multidrug resistance observed in the UPEC isolates coupled with the detection of virulence genes in the current study suggest a public health risk in association with wastewater reuse in the area. Thus, there is a need for continuous wastewater-based surveillance programs to monitor the spread and any potential effects associated with the distribution of antimicrobial resistance and virulence genes through wastewater treatment plants and sewage-receiving hostel gutters in the area.
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
OME and CA designed the research study. OME performed the research. CA supervised the work. OME, CAO, RFA, JOA, GKN, SOS and STG analyzed the data. OME, CA, CAO, JOA, RFA, GKN, SOS, and STG wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work to take public responsibility for appropriate portions of the content and agreed to be accountable for all aspects of the work in ensuring that questions related to its accuracy or integrity.
This study did not involve human participants or animal subjects and was conducted in compliance with regional regulations, which do not require ethical approval for environmentally based research.
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/j.fbe1604038.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.

