1 Department of Molecular Biology and Genetics, Faculty of Science, Atatürk University, 25240 Erzurum, Turkey
2 Department of Molecular Biology and Genetics, Graduate School of Natural and Applied Sciences, Atatürk University, 25240 Erzurum, Turkey
Abstract
Komagataella phaffii (K. phaffii) (Pichia pastoris), also called biotech yeast, is a yeast species with many applications in the biotechnology and pharmaceutical industries. This methylotrophic yeast has garnered significant interest as a platform for the production of recombinant proteins. Numerous benefits include effective secretory expression that facilitates the easy purification of heterologous proteins, high cell density with rapid growth, post-translational changes, and stable gene expression with integration into the genome. In the last thirty years, K. phaffii has also been refined as an adaptable cell factory that can produce hundreds of biomolecules in a laboratory setting and on an industrial scale. Indeed, over 5000 recombinant proteins have been generated so far using the K. phaffii expression method, which makes up 30% of the total cell protein or 80% of the total released protein. K. phaffii has been used to manufacture more than 70 commercial products in addition to over 300 industrial processes that have been granted licenses. Among these are useful enzymes for industrial biotechnology, including xylanase, mannanase, lipase, and phytase. The others are biopharmaceuticals, which include human serum albumin, insulin, hepatitis B surface antigen, and epidermal growth factor. Compared to other expression systems, this yeast is also considered a special host for synthesizing subunit vaccines, which have recently been supplanted by alternative vaccination types, such as inactivated/killed and live attenuated vaccines. Moreover, efficient production of recombinant proteins is achieved through multi-level optimization methods, such as codon bias, gene dosage, promoters, signal peptides, and environmental factors. Therefore, although K. phaffii expression systems are efficient and simple with clearly established process procedures, it is still necessary to determine the ideal conditions since these vary depending on the target protein to ensure the highest recombinant protein generation. This review addresses the K. phaffii expression system, its importance in industrial and biopharmaceutical protein production, and some bioprocessing and genetic modification strategies for efficient protein production. K. phaffii will eventually continue contributing as a potent expression system in research areas and industrial applications.
Keywords
- Komagataella phaffii
- Pichia pastoris
- yeast
- expression system
- recombinant protein
Biological expression systems are used in medical and industrial fields to produce heterologous proteins such as recombinant therapeutics, vaccines, and agricultural products [1]. Various expression systems are needed to express heterologous proteins for both therapeutic and research purposes. Different hosts, such as mammalian cells [2, 3, 4], yeasts [5, 6, 7], and bacteria (mainly E. coli) [8, 9, 10], are widely used as expression platforms for the production of biopharmaceuticals and industrial enzymes [11]. These hosts should be selected with both economic and qualitative considerations in mind. Industrial production depends on the expected high economic yield of the product and the use of cheap media components [5]. This is possible through recombinant DNA (rDNA) technology, which allows a specific gene of interest to be cloned and expressed in a suitable system [12]. Since this technology has the necessary tools to produce the desired proteins in a native form, the recombinant protein can be obtained in its pure form and in large quantities. In this way, biotechnology companies producing rDNA products in human and animal health have begun to grow rapidly [1]. In 1982, Humulin™ (biosynthetic human insulin) was produced in E. coli and commercialized for human use as the first recombinant biopharmaceutical. Prokaryotic hosts can express any gene, but the proteins produced may not always have the desired stability or biological activity. Additionally, every property of the recombinant protein must be identical to the native protein. Meanwhile, the final product may be contaminated by toxic components of bacterial origin, which represents a major problem, especially if the expressed protein is required for therapeutic purposes. Therefore, higher expression systems capable of many post-translational modifications are needed, especially during the production of eukaryotic proteins [1]. In this context, selecting a suitable host system for the specified purpose is necessary, with eukaryotic expression systems being used as an alternative to the prokaryotic system for expressing heterologous proteins.
The fields of genomics and proteomics, which have an important role in identifying potentially useful proteins, have triggered the need to develop high-throughput heterologous expression systems [13]. In recent years, yeasts have become attractive eukaryotic hosts for producing heterologous proteins. Unlike prokaryotes such as E. coli, significant advantages such as rapid growth in protein-free media, the ability to perform post-translational modifications, and disulfide bond formation make yeasts popular for use as expression systems [14, 15]. Yeast expression offers several advantages compared to other eukaryotic expression systems, such as mammalian and insect cells. For example, yeasts require simple cultivation techniques for maintenance, resulting in rapid cell growth, and they are highly adaptable to large-scale production [16]. Thus, many yeasts, including Saccharomyces cerevisiae, Komagataella phaffii, Pichia stipitis, Kluyveromyces lactis, Hansenula polymorpha, Schizosaccharomyces pombe, Yarrowia lipolytica, Schwanniomyces occidentalis, and Arxula adeninivorans are used as hosts for producing recombinant products in industrial biotechnology. These act as versatile hosts for genetic modifications, which makes them an ideal yeast expression system because yeasts can express heterologous genes through chromosomal integration of an expression cassette or episomal plasmids [17].
Initially isolated from a French chestnut tree, Komagataella phaffii (K. phaffii) is a methylotrophic yeast. French mycologist and cytologist Alexandre Guilliermond identified K. phaffii as Zygosaccharomyces pastori [18]. However, a new genus called Komagataella was suggested while redefining yeast taxonomy using information from DNA sequences. Thus, Komagataella pastoris was split into K. pastoris and K. phaffii [19]. Most industrially used strains called P. pastoris belong to the species K. phaffii [20], which is used for protein expression in most biotechnological applications. Alternatively, over 40 years ago, it was evaluated for single-cell protein (SCP) production as an animal feed additive [21, 22]. However, the remarkable increase in methanol prices due to the 1973 oil crisis made single-cell protein production processes uneconomical. When K. phaffi was initially developed as a heterologous protein expression system in the 1980s, a potent and tightly regulated AOX1 promoter was employed [21]. Subsequently, this yeast attracted the attention of researchers and became the most widely used yeast expression system in recombinant protein production research. Consequently, twenty years ago, Invitrogen licensed this system to ensure its availability to researchers worldwide, and K. phaffii was generally recognized as safe (GRAS) by the Food and Drug Administration (FDA) [23]. Indeed, K. phaffii is an excellent expression host for the production of industrial enzymes and biopharmaceuticals, as well as other heterologous proteins [24]. The technical advantages of this system are site-specific integration, a leader sequence for secretion of the heterologous protein, increased copy number, and post-translational modifications [12].
A wide range of proteins and peptides are produced from cells that are not naturally produced, thanks to recombinant DNA (rDNA) technologies. With the advent of recombinant DNA technology, the commercialization of new enzymes and drugs used in industrial applications with therapeutic properties has increased significantly [13]. Moreover, the market is expanding due to the rising demand for recombinant proteins for use in various biotechnological procedures, meaning revenues from commercialization have increased rapidly over the last decade [25, 26, 27, 28]. The first biotechnology items on the global market to be created by rDNA were industrial enzymes, which are used to treat food and in feed, paper pulp, detergents, and health care, as well as pharmaceuticals, which include insulin and the hepatitis B vaccine, interferons, and erythropoietin [29]. One of the most important branches of genetic engineering is the expression of recombinant proteins using biological expression systems. Furthermore, Komagataella phaffii is one of the eukaryotic expression systems developed over the last three decades as a flexible cell factory to produce thousands of biomolecules, both in the laboratory and on an industrial scale [30]. Additionally, it offers greater physiological advantages compared to other widely employed host cells [31, 32].
Research on using this yeast to produce recombinant proteins has continued, with some engineered wild-type, auxotrophic, and protease-deficient K. phaffii strains in the literature reported in Table 1 (Ref. [31, 33, 34, 35, 36]).
| Type | Genotype | Reference | ||
| Strain | Wild-type | X-33 | Wild-type | Life Technologies™ |
| CBS7435 (NRRL Y-11430) | Wild-type | Centraalbureau voor Schimmelcultures, the Netherlands | ||
| CBS704 (DSMZ 70382) | Wild-type | |||
| Auxotrophic | GS115 | his4 | Life Technologies™ | |
| KM71 | his4, aox1::ARG4, arg4 | Life Technologies™ | ||
| KM71H | aox1::ARG4, arg4 | Life Technologies™ | ||
| BG09 | arg4::nourseo Δlys2::hyg | BioGrammatics | ||
| GS190 | arg4 | [33] | ||
| GS200 | arg4 his4 | [34] | ||
| JC220 | ade1 | [33] | ||
| JC254 | ura3 | [33] | ||
| JC227 | ade1 arg4 | [35] | ||
| JC300-JC308 | ade1 arg4 his4 ura3 | [35] | ||
| CBS7435 his4 | his4 | [36] | ||
| CBS7435 Mut |
aox1, his4 | [36] | ||
| CBS7435 met2 arg4 | aox1, arg4 | [36] | ||
| Protease deficient | SMD1163 | his4 pep4 prb1 | [31] | |
| SMD1165 | his4 prb1 | |||
| SMD1168 | his4 pep4::URA3 ura3 | |||
| PichiaPink | ade4 |
Overall, the advantages of using the K. phaffii expression system in protein production include high cell density fermentation, higher folding efficiency, genetic stability, strong promoter, and a mature secretion system (using Kex2 as the signal peptidase) [31, 37]. Further, using this inexpensive system means there is no possibility of bacteriophage contamination, with a high level of efficiency exhibited in a virtually protein-free medium [38, 39]. Various post-translational modifications, including polypeptide folding, methylation, acylation, glycosylation, proteolytic adjustment, and targeting to subcellular compartments can be performed by these methylotrophic yeasts and cells, which can secrete proteins into the growth medium. Thus, simple processes can purify secreted proteins from the growth medium [39]. Industrial bioreactors enable this system to manufacture desired proteins on a big scale from tiny culture volumes [31]. A schematic representation of recombinant protein production using K. phaffii is given in Fig. 1 (Ref. [40]).
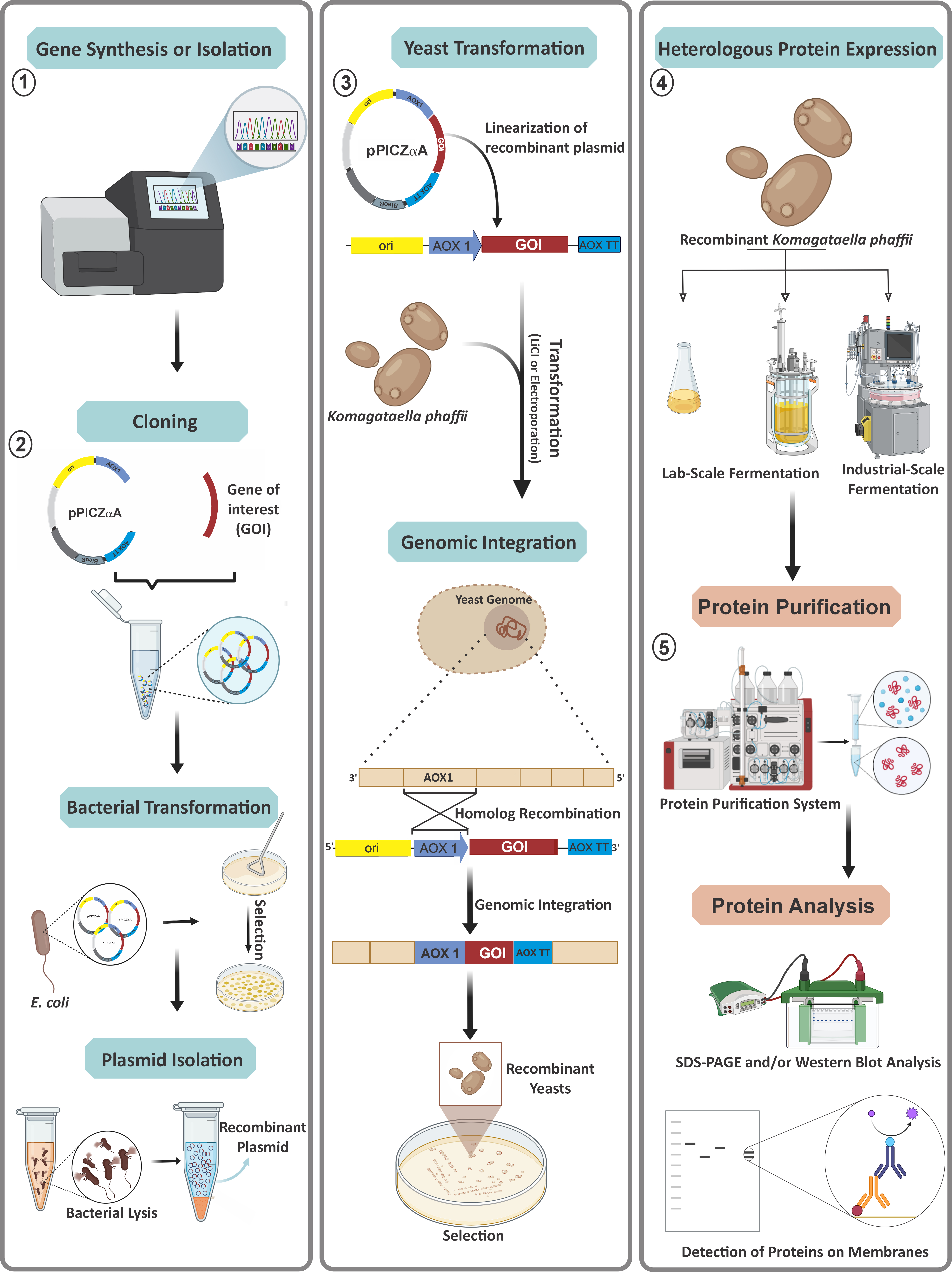 Fig. 1.
Fig. 1.Schematic representation of recombinant protein production using K. phaffii (modified from Barone et al. [40] and created with https://www.biorender.com/). ① Gene synthesis or isolation; ② cloning of the gene, transformation of bacteria, and plasmid DNA isolation; ③ transformation of K. phaffii cells and integration of recombinant plasmid into the yeast genome; ④ heterologous protein expression by K. phaffii; ⑤ protein purification and analysis. The blue or orange color of the rectangular-shaped text field indicates the upstream (blue) or downstream (orange) process, respectively.
K. phaffii has several strong or weak promoters that can be used to
drive the expression of heterologous genes in both an inducible and constitutive
manner. The strong methanol-inducible alcohol oxidase (alcohol oxidase 1,
AOX1, and to a lesser extent alcohol oxidase 2, AOX2) promoters
(P
P
| Corresponding gene | Regulation | Reference | ||
| Promoter | Inducible | dihydroxyacetone synthase (DAS) | Inducible with methanol | [47, 48] |
| formaldehyde dehydrogenase 1 (FLD1) | Inducible with methylamine or methanol | [49] | ||
| isocitrate lyase (ICL1) | Repressed by glucose; induced by ethanol (in absence of glucose) | [50] | ||
| putative Na |
Induction upon phosphate starvation | [51] | ||
| thiamine biosynthesis gene (THI11) | Repressed by thiamin | [52] | ||
| alcohol dehydrogenase (ADH1) | Repressed by methanol and glucose; induced by ethanol and glycerol | [53] | ||
| enolase (ENO1) | Repressed by glucose, ethanol, and methanol; induced by glycerol | [53] | ||
| glycerol kinase (GUT1) | Repressed by methanol; induced by glycerol, glucose, and ethanol | [53] | ||
| Constitutive | translation elongation factor 1 (TEF1) | Constitutive expression from glucose and glycerol | [54] | |
| 3-phosphoglycerate kinase (PGK1) | Constitutive expression from glucose, and to a lesser extent from glycerol and methanol | [55] | ||
| potential glycosyl phosphatidyl inositol (GPI)-anchored protein (GCW14) | Constitutive expression from glucose, glycerol, and methanol | [56] | ||
| high-affinity glucose transporter (G1) | Repressed by glycerol; induced upon glucose limitation | [57] | ||
| putative aldehyde dehydrogenase (G6) | Repressed by glycerol; induced upon glucose limitation | [57] |
Moreover, this methylotrophic yeast is of interest as a cell factory for
producing recombinant proteins as it effectively meets laboratory and industrial
setup requirements [58]. Typically, this yeast may be grown in dynamic culture
systems, where the cell culture medium is continuously changed, and various
variable elements influence how productive the final products are. It can secrete
large amounts of protein in a bioreactor while growing quickly at high cell
densities (
Numerous models have been created at the metabolic and bioreactor levels, and K. phaffii has emerged as a significant microorganism for both basic and applied research and industrial applications [61]. Metabolic models are useful tools in biotechnology that aid in understanding and optimizing the production process [62]. Bioreactor models for K. phaffii can be used to optimize bioreactor operations and provide insights into cellular metabolism. A comprehensive macroscopic bioreactor model for K. phaffii was built by Hong et al. (2021) [63], and it includes descriptions of the substrates, biomass, total protein, other medium components, and off-gas components. To explain the absorption and evolution rates for medium components and off-gas components, the study introduces species and elemental balances. In addition, a pH model was built to describe the precipitation of medium components and the synthesis of recombinant protein utilizing an overall charge balance, acid/base equilibria, and activity coefficients. It was stated that by reducing the number of components needed to meet the cellular requirements, the model for medium components with pH and precipitation can be utilized to improve the chemically defined medium.
In another study, innovative bioprocessing strategies combining physiological
control and strain engineering were developed to improve the recombinant protein
production of K. phaffii. Initially, two Candida rugosa lipase
1 producer clones were compared in chemostat cultures with varying
oxygen-limiting circumstances concerning their gene dosage performance under
constitutive P
Since K. phaffii is widely recognized as essential to the
biotechnological processes used to produce a variety of bioproducts, successful
genetic modification of this yeast requires advanced gene editing techniques.
Genetic modification of K. phaffii has traditionally been based on
homologous recombination mechanisms, although this method results in low
efficiency even when long homologous arms are required. The main reason for this
low efficiency is the predominance of non-homologous end joining (NHEJ), a
natural gene repair mechanism that suppresses homologous recombination in this
yeast species [65]. The CRISPR–Cas9 system, which is currently revolutionizing
the fields of biotechnology, genetic engineering, and protein production,
provides significant advances in these fields by editing the genomes of
microorganisms such as K. phaffii. Recent studies have focused on
increasing the efficiency of gene manipulations in K. phaffii using
CRISPR–Cas9. In one of these studies, a strain of K. phaffii with a
non-homologous end-junction defect (
Methanol induction has many drawbacks while also being a powerful tool for
promoting and controlling the expression of recombinant proteins. One of the
disadvantages is that methanol is flammable and toxic. Methanol is highly used in
recombinant protein production, although its storage causes technical
difficulties. Based on the EasySelect™ Pichia Expression kit
(Thermo Scientific, Waltham, MA, USA), at least 0.5% (wt/vol) methanol use is
needed during recombinant protein production [68]. Maximum production was
achieved with a methanol level of 2–2.5% (wt/vol) [69]. Yeasts can normally
withstand up to 5% of methanol, but concentrations above 5% are extremely
harmful to cell viability, halting the production process [70]. In addition, a
high oxygen supply is needed for catabolism, which means increased production
costs as this promotes high heat release. Moreover, since methanol is a petroleum
product, its cost invariably increases during petroleum crises. Thus, the
preference is to avoid using methanol for producing pharmaceuticals or edible
products [23]. On the other hand, hydrogen peroxide (H
While protein production via methanol induction remains popular, efforts are underway to eliminate its disadvantages. Indeed, there have been efforts to replace methanol as a single carbon source in K. phaffii, mainly due to strict process control for large-scale methanol induction alongside safety concerns. Some methanol-free expression systems were created to remove the drawbacks of using methanol in the production of recombinant proteins, and the number of studies on methanol-free K. phaffii expression systems in the literature has increased. These studies aimed to investigate new alternative promoters to the AOX promoters, inhibit and/or activate current transcription factors, and obtain mutant strains capable of expressing methanol independently.
Indeed, a study by Shen et al. (2024) [71] identified a novel promoter
that can achieve biphasic expression without the need for an inducer and is
closely related to the growth status of cells. In K. phaffii cells
carrying a single-copy green fluorescent protein (GFPuv) gene, P
Garrigós-Martínez et al. (2021) [72] first performed a
comparative kinetic characterization of two expression systems, including
commercial PDF and UPP promoters (P
Mombeni et al. (2020) [73] replaced the promoter region of the methanol
oxidase gene (P
Shirvani et al. (2019) [74] developed a new expression vector (pEP
(
Takagi et al. (2019) [75] developed a methanol-free K. phaffii
expression system by arranging a strong methanol-inducible
P
Another example in the literature is the study by Wang et al. (2017)
[76], where to obtain a methanol-free K. phaffii expression system, the
authors tested the functions of the trans-acting factors of the AOX1
promoter and found that the factors were arranged combinatorially. Furthermore,
in the Wang et al. [76] study, three transcription factors (Mig1,
Mig2, and Nrg1) were defined and deleted, and the Mit1 transcription activator
was overexpressed. Thus, a methanol-free P
In another study, Shen et al. (2016) [77] focused on constructing a new
methanol-free expression system targeting kinases by activating or inhibiting the
AOX1 promoter using different carbon sources. For this purpose, two
kinase mutants (
Protein production for industrial use started at the end of the nineteenth century. Meanwhile, the industry has been investigating new components to improve the efficiency of the process and analyzing alternatives to make the process most cost-effective. The development of recombinant DNA technology has made significant advances in bioprocess optimization owing to the possibility of using recombinant organisms. This technology offers several alternatives for the production of proteins, offering new and/or better properties for industrial processes [78].
Yeast expression systems are one of the most frequently used alternatives for high-scale production of various proteins. In fact, they have become an economical and efficient source of industrially important proteins. Recently, yeast expression systems have been used in industrial biotechnology to produce different product types. These mainly include the production of organic acids, pigments, fatty acids, vitamins, other food components, pharmaceuticals, and therapeutic molecules, as well as enzymes and biofuels [17].
An estimated USD 6.3 billion and a 4.7% compound annual growth rate (CAGR) were projected for enzymes employed in industrial applications in 2021 [25, 28]. Their global market is expected to have a CAGR of 6.3% between 2021 and 2026 and to reach USD 8.7 billion by 2026 [79]. Industrial enzymes, which are considered more environmentally friendly than chemical alternatives, are frequently preferred in various industries such as food and beverage, textile, chemical production, pharmaceuticals, and cosmetics [80, 81]. The production of more than 20 mg/L of hydroxy nitrile lyase from the tropical rubber tree Hevea brasiliensis is known as one of the first industrial production processes established in the 1990s [82]. Among the commercial K. phaffii products is Superior Stock, which is made in Michigan, USA, by Nitrate Elimination Co. Nitrates in soil, water, and animal feed can be tested using the recombinant nitrate reductase included in this kit.
Recombinant phospholipase C, which is frequently used for degumming vegetable oils, is present in K. phaffii yeast in food technology [83]. Yeasts are the most effective way to obtain Butiauxella sp. phytase, achieving yields of 22 g/L through methanol induction and 20 g/L in methanol-free conditions [84]. Thanks to improved fermentation strategies and bioreactor studies at various scales, the glycoside hydrolase yield from Lentinula edodes increased to 898 mg/L [85]. Several recombinant proteins produced by the K. phaffi expression system, mostly using the AOX1 promoter for industrial use, are provided in Table 3 (Ref. [6, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105]).
| Product name | Used strain | Used plasmid | Usage | Reference |
| Human paraoxonase 1 | X-33 | pPICZ |
Hydrolysis of organophosphates | [86] |
| Laccase | SMD1168H | pPICZ |
Biodegradation of lignin | [87] |
| Phytase | GS115 | pPIC9 | Hydrolysis of phytic acid | [88] |
| X‐33 | pPICZ |
Degradation of mannan | [89] | |
| GS115 | pGAPZ | Hydrolysis of glycoside | [90] | |
| Leukocyte protease inhibitor | JC100 | pBLHIS-SX | Immunity-associated protein | [91] |
| Human sialyltransferase | KM71H | pPICZ |
Pharmacological uses | [92] |
| Cyanate hydratase | GS115 | pPICZ |
Detoxification of cyanate and cyanide | [93] |
| X‐33 | pPICZ |
Starch saccharification | [94] | |
| GS115 | pPICZ |
Hydration of carbon dioxide to form carbonic acid | [95] | |
| Carrot antifreeze protein | GS115 | pPIC9K | Inhibition of gluten deterioration | [96] |
| Legumain | X‐33 | pPICZ |
Lysosomal protease | [97] |
| Trypsin | GS115 | pPIC9K | Hydrolysis of proteins | [98] |
| Transglutaminase | GS115 | pPIC9K | Restructured meat products | [99] |
| Lignin peroxidase | CBS704 | pJ901 | Lignin degradation | [100] |
| Myrosinase | KM71H | pPICZ |
Hydrolysis of glucosinolates | [101] |
| Xylanase | SMD1168 | pHIL-D2 | Xylan depolymerisation | [102] |
| GS115 | ||||
| Pectin methylesterase | X-33 | pGKB | Pectin degradation | [6] |
| Cellulase | GS115 | pPICZ |
Cellulose degradation | [103] |
| L-asparaginase | KM71H | pPICZ |
Chemotherapeutic agent | [104] |
| Lipase B | GS115 | pPICZ |
Hydrolysis of fats | [105] |
One of the fastest-growing areas of molecular medicine is the production of recombinant therapeutic proteins. These proteins play significant roles in the treatment of various diseases. Generally, biopharmaceuticals compensate for deficiency or damage to body proteins that are important for normal organism functions. These can be classified into enzymes, hormones, growth factors, blood factors, thrombolytics and anticoagulants, monoclonal antibodies and vaccines, interferons, and interleukins. The biopharmaceuticals market is projected to develop at a compound annual growth rate (CAGR) of 7.1% from 2022 to 2027, when it is expected to reach USD 532.2 billion [106]. The term “biopharmaceutical” refers to nucleic acid-based products, recombinant therapeutic proteins, and, more broadly, engineered cell or tissue-based products [107]. Most biopharmaceuticals approved by regulatory authorities for therapeutic applications are proteins produced by recombinant DNA technology in various expression systems [108]. To date, more than 5000 recombinant proteins have been produced in the K. phaffii expression system, accounting for 80% of the total secreted protein or 30% of the total cell protein [30]. In recent years, studies on the production of biopharmaceutical proteins with the K. phaffii expression system have increased in the literature. Some examples are given in Table 4 (Ref. [69, 109, 110, 111, 112, 113, 114, 115, 116, 117, 118, 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132, 133, 134, 135, 136, 137]).
| Product name | Used strain | Used plasmid | Usage | Reference |
| Bovine lactoferrin | KM71H | pJ902 | Transferrin and antibacterial protein | [109] |
| Snakin‐1 | GS115 | pPIC9 | Antimicrobial peptide | [110] |
| Fowlicidin‐2 | X‐33 | pPICZ |
Antimicrobial peptide | [111] |
| Bovine IFN‐ |
GS115 | pPIC9K | Prevention and therapy of viral diseases | [112] |
| CHIKV‐C‐E3‐E2‐6K‐E1 | GS115 | pPIC9K | Recombinant subunit vaccine | [113] |
| Gp350 | GS115 | pPICZ |
Recombinant subunit vaccine | [69] |
| Streptokinase | X‐33 | pPICZ |
Thrombolytic drug | [114] |
| TFPR1 | X‐33 | pPICZ |
Adjuvant | [115] |
| IL‐1 |
GS115, SMD1168, X‐33 | pPICZA | Proinflammatory cytokine | [116] |
| IL‐15 | X‐33 | pPICZ |
Differentiation and proliferation of T, B, and NK cells | [117] |
| BoNT Hc | X-33 | pPICZ |
Recombinant subunit vaccine | [118] |
| CFP10-Fc |
GS115 | pPICZ |
Recombinant subunit vaccine | [119] |
| Glycoprotein D | GS115 | pPIC9K | Recombinant subunit vaccine | [120] |
| IL‐11 | GS115 | pPINK |
Thrombopoietic growth factor | [121] |
| Staphylokinase | GS115, KM71H | pPICZ |
Thrombolytic drug | [122] |
| Apidaecin | SMD1168 | pPIC9K | Antibacterial peptide | [123] |
| Class I chitinase | KM71H | pPICZ |
Antifungal peptide | [124] |
| DM64 | X‐33 | pPICZ |
Anti‐myotoxic | [125] |
| hIFN‐ |
X‐33, GS115, KM71H, CBS7435 | pPICZ |
Critical cytokine for innate and adaptive immunity | [126] |
| CecropinA‐thanatin | X‐33 | pPICZ |
Antimicrobial peptide | [127] |
| Tachyplesin I | GS115 | pGAPZ |
Antibacterial peptide | [128] |
| PAF102 | X‐33 | pGAPZA | Antifungal peptide | [129] |
| Hispidalin | GS115 | pPICZ |
Antimicrobial peptide | [130] |
| CGA-N12 | GS115 | pPIC9 | Antifungal peptide | [131] |
| Azurin | X-33 | pPICZ |
Anticancer protein | [132] |
| Human telomerase inhibitor 1 (hPinX1) | X-33 | pPICZ |
Human telomerase inhibitor protein | [133] |
| Chitinase | GS115 | pPICZA | Antifungal peptide | [134] |
| Enterocin E-760 | X-33 | pPICZ |
Antimicrobial peptide | [135] |
| Human serum albumin | X-33 | pPICZ |
Therapeutic protein | [136] |
| Turgencin A | GS115 | pPICZ |
Antimicrobial peptide | [137] |
Recombinant protein vaccines are important in biopharmaceutical proteins and have attracted considerable attention recently. These vaccines are considered a safer option than those made from live viruses since they are non-replicating and do not contain any infectious elements of an attenuated viral particle. Moreover, the technology has undergone extensive testing, and these vaccinations often only cause minimal adverse effects [138, 139]. As a result, numerous recombinant protein vaccines are currently being used in clinical settings across the globe [140]. Using recombinant yeasts as vaccines may represent an interesting substitute because of their immunogenicity, versatility, and production cost benefits. These yeasts can be used in vaccination preparations without requiring the antigens to be purified due to their safety during administration. Furthermore, thousands of dosages can be obtained per liter of culture owing to the high cell densities used in yeast cultivation [141]. Clinical experiments have previously assessed the viability of administering these vaccinations to humans, yielding encouraging outcomes in terms of immunogenicity, safety, and tolerance [142, 143]. The vaccine candidates against viral diseases produced by the K. phaffii expression system, which has been used to produce several established vaccines and therapeutics, are given in Table 5 (Ref. [143, 144, 145, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155]).
| Construct name | Expression condition | Targeted disease/organism | Status | Reference |
| RBD-beta | Lab scale | SARS-CoV-2 | Pre-clinic (mouse) | [144] |
| RBD | Lab scale | SARS-CoV-2 | Pre-clinic (mouse) | [145] |
| RBD-DP | Lab scale | SARS-CoV-2 delta plus | Pre-clinic (mouse) | [146] |
| RBD | Lab scale | SARS-CoV-2 | Clinic (phase 1 and 2) | [147] |
| RBD219-N1C1 | Lab scale | SARS-CoV-2 | Pre-clinic (mouse) | [148] |
| MCPD | Lab scale | Largemouth bass iridovirus (LMBV) | Pre-clinic (Micropterus salmoides) | [149] |
| HEV-VLP | Lab scale | Hepatitis E virus | Pre-clinic (mouse) | [150] |
| ZIKV Env-NS1 | Lab scale | Zika virus (ZIKV) | Pre-clinic (mouse) | [143] |
| CRM197 | Lab scale | Typhoid fever | No clinical trial | [151] |
| HPV 52 L1 | Lab scale | Human papillomavirus (HPV) | No clinical trial | [152] |
| H7 | Lab scale | H7N9 influenza virus | Pre-clinic (mouse) | [153] |
| EVA71 | Lab scale | Enterovirus | Pre-clinic (mouse) | [154, 155] |
RBD-beta; beta variant receptor binding domain (RBD) protein, RBD-DP; RBD of the SARSCoV2 Delta Plus strain, RBD219-N1C1; mutant RBD219 subunit protein, SARS-CoV-2; severe acute respiratory syndrome coronavirus 2, MCPD; LMBV major capsid protein, HEV-VLP; Hepatitis E virus-virus-like particles, ZIKV Env-NS1; ZIKV envelope and non-structural NS1 protein, CRM197; cross-reactive-material 197, HPV 52 L1; HPV 52 L1 capsid protein, H7; hemagglutinin protein of H7N9 influenza virus, EVA71; Enterovirus A71.
In addition, more than 70 commercial products are produced by K. phaffii, and more than 300 licensed industrial processes are carried out using this expression system (http://www.pichia.com). One of the biopharmaceutical products is Kalbitor® (DX-88 ecallantide), a recombinant kallikrein inhibitor protein (Dyax, Cambridge, MA, USA). This protein, used in treating hereditary angioedema, is the first FDA-approved biopharmaceutical synthesized in K. phaffii (Thompson, 2010). Other biopharmaceutical products developed using the K. phaffii expression system include Insugen®, a recombinant human insulin manufactured by Biocon (Bengaluru, India) for treating diabetes; Medway®, recombinant serum albumin manufactured by Mitsubishi Tanabe Pharma (Osaka, Japan) for blood vessel dilation; Shanvac®, a recombinant hepatitis B vaccine, and Shanferon®, a recombinant interferon alpha 2b, both manufactured by Shantha/Sanofi (Medchal, India); Ocriplasmin®, a recombinant microplasmin manufactured by ThromboGenics (Leuven, Belgium) for the treatment of vitreomacular adhesion; the anti-IL-6 receptor single domain antibody fragment, Nanobody® ALX-0061®, for the treatment of rheumatoid arthritis; the anti-RSV single domain antibody fragment, Nanobody® ALX00171, for the treatment of respiratory syncytial virus (RSV) infection, both manufactured by Ablynx (Gent, Belgium); heparin-binding EGF-like growth factor (HB-EGF), manufactured by Trillium (Brockville, Canada) to treat interstitial cystitis (http://www.pichia.com).
Currently, low-cost and high-throughput production of many enzymes used in various industries (biopolymers, cosmetics, food processing, detergents, etc.) and medical applications are being performed on different scales by K. phaffii. Significant progress has been made to improve recombinant protein expression using this microorganism, and there are impressive advantages to using this methylotrophic yeast. However, the expression level of some proteins used in industrial applications may still need to be increased to meet commercialization demands. This is particularly important in expressing hetero-oligomers, membrane-bound proteins, or those prone to proteolytic degradation. To overcome such situations, general strategies for K. phaffii strain engineering for heterologous expression such as gene optimization (codon optimization) (i), plasmid engineering (promoter selection or gene dosage) (ii), cultivating tactics including co-expression of chaperones (iii), secretion pathway engineering (iv) and directed evolution or genome editing (v) have been preferred. Combining these approaches will make K. phaffii a promising platform for synthesizing valuable industrial and biopharmaceutical proteins.
YU—conceptualization, bibliography review and manuscript writing and editing; ID—bibliography review, manuscript, figure and table editing. Both authors contributed to editorial changes in the manuscript. Both authors read and approved the final manuscript. Both authors have participated sufficiently in the work to take public responsibility for appropriate portions of the content and agreed to be accountable for all aspects of the work in ensuring that questions related to its accuracy or integrity.
Not applicable.
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.

