1 Department of Cardiology, Beijing Anzhen Hospital, Capital Medical University, National Clinical Research Center for Cardiovascular Diseases, 100029 Beijing, China
2 Department of Geriatric, Huangshi Central Hospital, Affiliated Hospital of Hubei Polytechnic University, Edong Healthcare Group, 435000 Huangshi, Hubei, China
3 Beijing Key Laboratory of Preclinical Research and Evaluation for Cardiovascular Implant Materials, Animal Experimental Centre, Fuwai Hospital, National Centre for Cardiovascular Disease, Chinese Academy of Medical Sciences and Peking Union Medical College, 100037 Beijing, China
4 Heart Failure Care Unit (HFCU), Heart Failure Center, Fuwai Hospital, National Center for Cardiovascular Disease, Chinese Academy of Medical Sciences and Peking Union Medical College, 100037 Beijing, China
5 Department of Comprehensive Internal Medicine, First Affiliated Hospital of Xinjiang Medical University, 830011 Urumqi, Xinjiang, China
6 Key Laboratory of Clinical Research for Cardiovascular Medications, National Health Committee, 100037 Beijing, China
†These authors contributed equally.
Abstract
Coronary heart disease (CHD) arises from a complex interplay of genetic and environmental factors. This study examines the influence of endothelial lipase gene polymorphisms (rs2000813 and rs3813082) and their interactions with traditional cardiovascular risk factors on CHD susceptibility.
This retrospective case–control study enrolled 900 CHD patients and 900 control subjects. We evaluated associations between conventional cardiovascular risk factors and polymorphisms at the rs2000813 and rs3813082 loci in the endothelial lipase gene. Multifactorial analysis was used to assess interactions between traditional risk factors and these polymorphisms. Additionally, we developed a predictive model integrating genetic variants and clinical variables to estimate CHD risk.
No significant differences were observed in the distribution of rs2000813 genotypes (CC, CT, TT) and alleles (C, T), or rs3813082 genotypes (AA, AC, CC) and alleles (A, C) between CHD and control groups, including among males. However, in females with CHD, the rs2000813CT genotype was significantly more frequent (49.30%) than in controls (37.80%), whereas the CC genotype was less frequent in the CHD group (45.00%) than in controls (55.20%). Multivariate logistic regression identified the rs2000813CT genotype, hypertension, ages ≥60 years, body mass index (BMI) values ≥28 kg/m2, total cholesterol (TC) ≥6.2 mmol/L, and apolipoprotein B (ApoB) ≥1.1 g/L as potential risk factors for CHD in women (p < 0.05). Gene–environment interaction analysis revealed that BMI exerted the greatest influence (12.62%). A predictive model incorporating rs2000813 genotypes estimated CHD risk in women with an area under the curve (AUC) of 0.804.
The rs2000813CT endothelial lipase genotype is potentially associated with an increased CHD risk in females, whereas the CC genotype may confer a protective effect. Integrating endothelial lipase gene variants with traditional cardiovascular risk factors enhances CHD risk prediction in women. Synergistic interaction between endothelial lipase polymorphisms and environmental factors appears to influence CHD occurrence in this population.
Keywords
- gene-environment interaction
- gene polymorphisms
- endothelial lipase
- coronary heart disease
- traditional cardiovascular risk factors
Coronary heart disease (CHD) encompasses a spectrum of cardiovascular disorders resulting from myocardial ischemia, hypoxia, or necrosis due to coronary artery stenosis or occlusion. It is characterized by high morbidity and mortality rates. According to the World Health Organization’s Global Health Estimates report from December 2020, CHD remains the leading cause of death worldwide, accounting for 16% of all fatalities [1]. In China, the prevalence of CHD is increasing. The China Health Statistics Yearbook 2019 reports that in 2018, the CHD mortality rate was 120.18 per 100,000 in urban populations and 128.24 per 100,000 in rural population [2].
As a chronic condition, CHD is manageable but incurable, requiring long-term diagnosis and treatment. Its etiology primarily stems from a combination of factors, including dyslipidemia, coronary artery endothelial dysfunction, and chronic inflammation [3, 4]. Recent studies have underscored the critical role of genetic variants in CHD and pathophysiology [5, 6]. Despite the implementation of clinical interventions, such as preventive and therapeutic medications, along with guidelines for managing lipid metabolism disorders and chronic inflammatory, the prevalence of CHD remains high [7]. Elucidating the regulatory mechanisms of gene variants in lipid metabolism, inflammatory responses, and vascular endothelial injury is essential to address the underlying causes of CHD and may pave the way for significant advances in its prevention and treatment [8].
Endothelial lipase (EL), encoded by the endothelial lipase gene, is primarily synthesized by vascular endothelial cells. Its expression is upregulated in response to acute inflammatory stimuli, endotoxin exposure, and alterations in vascular wall shear stress. Hirata et al. [9] identified the endothelial lipase gene on the long arm of chromosome 18 (18q21.1), comprising 10 exons and 9 introns. EL plays a pivotal role in lipid metabolism and is implicated in metabolic syndromes, including inflammation and atherosclerosis [10]. This study investigates the effects of endothelial lipase polymorphisms rs2000813 and rs3813082, alongside traditional cardiovascular risk factors and their interactions, on susceptibility to CHD. Furthermore, it aims to develop a risk-prediction model integrating genetic and conventional factors. This model seeks to identify high-risk individuals for early intervention, enhance the prognosis of cardiac events, and provide a theoretical foundation for novel approaches to the diagnosis, treatment, and prevention of CHD.
This study employed a case-control design. The sample size was calculated using
the following parameters: an odds ratio (OR) of 1, a significance level
(
Between January 2019 and December 2021, we enrolled 900 patients diagnosed with coronary artery disease (CAD) who underwent coronary angiography or percutaneous coronary intervention (PCI) at the First Affiliated Hospital of Xinjiang Medical University as the case group. Concurrently, 900 patients who did not meet the diagnostic criteria, as confirmed by coronary angiography or coronary computed tomography angiography (CTA) during the same period, were selected as the control group. Subsequently, 720 participants from each group were randomly allocated in an 8:2 ratio to form the modeling subset, with the remaining 180 participants from each group assigned to the validation subset.
Inclusion criteria: (1) Patients aged
Exclusion criteria: Patients were excluded if they met any of the following conditions: (1) Incomplete clinical data. (2) Presence of severe heart failure, cardiogenic shock, malignant arrhythmias, tumors, autoimmune diseases, or conditions associated with aortic stenosis. (3) Refusal to participate or noncompliance with the study protocol.
Inclusion criteria: (1) Participants aged
Exclusion criteria: Patients were excluded if they met any of the following: (1) Incomplete clinical data. (2) Presence of severe heart failure, cardiogenic shock, malignant arrhythmias, tumors, or autoimmune diseases. (3) Refusal to participate or noncompliance with the study protocol.
(1) CHD: Diagnosis required typical angina symptoms and/or non-invasive evidence
of myocardial ischemia, coupled with coronary angiography demonstrating
(2) Hypertension: Defined as systolic blood pressure
(3) Type 2 diabetes mellitus: Diagnosed based on random blood glucose
(4) Smoking calculator: A pack year is defined as twenty cigarettes smoked everyday for one year [15].
(5) Alcohol consumption: Defined as ethanol intake
Baseline data, including sex, age, body mass index (BMI), smoking history, alcohol consumption, hypertension, and type 2 diabetes mellitus, were collected from all participants.
After fasting for
Upon hospital admission, 3 mL of venous blood was collected from the antecubital vein of each participant and anticoagulated with 2% ethylenediaminetetraacetic acid (EDTA) disodium salt. Leukocytes were isolated by centrifugation, and genomic DNA was extracted using the phenol-chloroform method, then stored at –80 °C. The SNPscnTM high-throughput assay was employed to genotype the endothelial lipase gene loci rs2000813 and rs3813082. Based on nucleotide variations, the rs2000813 locus was classified into CC, CT, and TT genotypes, and rs3813082 locus into AA, AC, and CC genotypes (Supplementary Table 1).
Statistical analyses were conducted using SPSS software (version 26.0, IBM
Corp., Armonk, NY, USA). Categorical data were reported as counts and percentages
(n [%]) and compared using the chi-square (
Baseline characteristics (Table 1) revealed no significant differences between
the case and control groups in sex distribution, age, alcohol consumption, TG
levels, ApoA, SUA, Scr, or other variables. However, the case group exhibited
significantly higher prevalence rates of smoking (44.70% vs. 36.60%, p
= 0.001), hypertension (48.60% vs. 43.10%, p = 0.023) and diabetes
(26.20% vs. 11.90%, p
| Variable | Control group (n = 900) | Case group (n = 900) | p-value | |
| Male, n (%) | 589 (66.40) | 612 (68.00) | 1.324a | 0.271 |
| Age, years | 55.75 |
55.98 |
–0.511b | 0.609 |
| Smoking history, n (%) | 329 (36.60) | 402 (44.70) | 12.275a | 0.001 |
| History of drinking, n (%) | 268 (29.80) | 265 (29.40) | 0.024a | 0.918 |
| Hypertension, n (%) | 388 (43.10) | 437 (48.60) | 5.373a | 0.023 |
| Diabetes, n (%) | 107 (11.90) | 236 (26.20) | 59.937a | |
| BMI, kg/m2 | 25.40 (23.44, 29.36) | 28.03 (25.77, 31.25) | –11.307c | |
| TG, mmol/L | 1.66 (1.17, 2.26) | 1.66 (1.19, 2.40) | –0.845c | 0.398 |
| TC, mmol/L | 4.33 (3.89, 4.78) | 4.44 (4.01, 5.10) | –4.713c | |
| HDL-C, mmol/L | 1.08 |
1.00 |
5.037b | |
| LDL-C, mmol/L | 2.73 (2.51, 3.05) | 2.75 (2.60, 3.31) | –4.248c | |
| ApoA, mmol/L | 1.17 (1.06, 1.30) | 1.17 (1.01, 1.30) | –1.761c | |
| ApoB, mmol/L | 0.84 (0.68, 0.98) | 0.87 (0.73, 1.02) | –3.614c | |
| Lp(a), mmol/L | 153.91 (110.88, 215.00) | 176.65 (123.16, 265.80) | –4.775c | |
| SUA, µmol/L | 311.61 |
316.44 |
–1.150b | 0.250 |
| Scr, µmol/L | 71.00 (59.09, 82.00) | 71.01 (61.00, 83.00) | –1.195c | 0.232 |
| WBC, |
6.68 (5.53, 7.52) | 8.61 (6.97, 11.06) | –17.829c | |
| NE, |
3.82 (2.98, 4.60) | 5.66 (4.07, 8.41) | –18.466c | |
| PLT, |
216.00 (182.00, 250.00) | 223.50 (191.00, 269.75) | –5.328c | |
| CK-MB, µg/L | 13.60 (10.10, 16.30) | 18.33 (12.47, 38.37) | –14.625c | |
| cTnI, µg/L | 0.00 (0.00, 0.00) | 0.00 (0.00, 0.02) | –6.531c | |
| cTnT, µg/L | 0.00 (0.00, 0.03) | 0.02 (0.00, 0.27) | –20.866c |
BMI, body mass index; TG, triglyceride; TC, total cholesterol; HDL-C,
high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; ApoA, apolipoprotein A;
ApoB, apolipoprotein B; Lp(a), lipoprotein (a); SUA, serum uric acid; Scr, serum
creatinine; WBC, white blood cell; NE, neutrophil count; PLT, platelet; CK-MB,
creatine kinase-MB; cTnI, cardiac troponin I; cTnT, cardiac troponin T; a:
Supplementary Table 2 presents the Hardy-Weinberg equilibrium (HWE) test results. Genotype distributions for both the CHD and control groups conformed to HWE, indicating that the study population represents a Mendelian population in genetic equilibrium. This finding supports the genetic representativeness the sample, a critical factor for the validity of genetic association studies. Additionally, haplotype linkage disequilibrium (LD) analysis of the rs2000813 and rs3813082 loci in the endothelial lipase gene was performed using the SHEsis online platform. The results yielded a D’ value of 0.158 and an r2 value of 0.001, suggesting no significant LD between these loci. Thus, rs2000813 and rs3813082 are genetically independent, an important consideration for evaluating their individual and combined effects on CHD susceptibility.
Supplementary Table 3 summarizes the distribution of endothelial lipase
gene polymorphisms between the CHD and control groups. No significant differences
were observed in the distribution of rs2000813 genotypes (CC,
CT, TT) and alleles (C, T), or in
rs3813082 genotypes (AA, AC, CC) and alleles
(A, C) between the groups (all p
Supplementary Table 4 details the distribution of endothelial
lipase genes polymorphisms in male participants, comparing the CHD and control
groups. No significant differences were observed in the distribution of
rs2000813 genotypes (CC, CT, TT) and alleles
(C, T), or rs3813082 genotypes (AA,
AC, CC) and alleles (A, C) between the CHD
and the control groups among men (all p
| SNP | Genotype | Control group (n = 241) | Case group (n = 229) | p-value | |
| rs2000813 | CC | 133 (55.20) | 103 (45.00) | 6.417 | 0.040 |
| CT | 91 (37.80) | 113 (49.30) | |||
| TT | 17 (7.10) | 13 (5.70) | |||
| C | 357 (74.07) | 319 (69.65) | 2.267 | 0.132 | |
| T | 125 (25.93) | 139 (30.35) | |||
| Dominant model | CC | 133 (55.20) | 103 (45.00) | 4.895 | 0.027 |
| CT + TT | 108 (44.80) | 126 (55.00) | |||
| Recessive model | TT | 17 (7.10) | 13 (5.70) | 0.373 | 0.542 |
| CT + CC | 224 (92.90) | 216 (94.30) | |||
| Additive model | CC | 133 (55.20) | 103 (45.00) | 0.001 | 0.974 |
| TT | 17 (7.10) | 13 (5.70) | |||
| rs3813082 | AA | 193 (80.10) | 180 (78.60) | 0.651 | 0.722 |
| AC | 42 (17.40) | 45 (19.70) | |||
| CC | 6 (2.50) | 4 (1.70) | |||
| A | 428 (88.80) | 405 (88.43) | 0.032 | 0.859 | |
| C | 54 (11.20) | 53 (11.57) | |||
| Dominant model | AA | 193 (80.10) | 180 (78.60) | 0.157 | 0.692 |
| AC + CC | 48 (19.90) | 49 (21.40) | |||
| Recessive model | CC | 6 (2.50) | 4 (1.70) | 0.311 | 0.577* |
| AC + AA | 235 (97.50) | 225 (98.30) | |||
| Additive model | AA | 193 (80.10) | 180 (78.60) | 0.266 | 0.753* |
| CC | 6 (2.50) | 4 (1.70) |
SNP, single nucleotide polymorphism. *: There is a frequency
Supplementary Table 5 outlines the classification of blood lipid parameters, SUA, and routine hematological indices. These classifications are based on the 2022 Chinese Clinical Blood Lipid Test Guide [17], the 2019 Basic Diagnosis and Treatment Guide for Gout and Hyperuricemia [18], and the ninth edition of the Diagnostic Standard [19]. These authoritative guidelines provide a standardized framework for assigning values and interpreting the diverse biochemical and hematological data in this study, ensuring consistency and reliability.
Spearman correlation analysis revealed that positive associations between CHD in
women and multiple factors, including lifestyle factors (smoking and alcohol
consumption), clinical conditions (hypertension and type 2 diabetes mellitus),
and biochemical markers (BMI, TC, ApoB, Lp(a), WBC, NE, and PLT). The
rs2000813 CT genotype also showed a positive correlation with
CHD, whereas the rs2000813 CC genotype was inversely associated with CHD
in women. Univariate logistic regression analysis identified hypertension, type 2
diabetes mellitus, advanced age, elevated BMI, and increased levels of TC, ApoB,
Lp(a), WBC, NE, and PLT as risk factors for CHD in women, each with distinct ORs.
Multivariate logistic regression further confirmed the following as significant
predictors: hypertension (OR, 2.305; 95% confidence interval (CI), 1.438–3.696;
p
| Risk factor | One-way | Risk factor | Multi-factor | ||
| OR (95% CI) | p-value | OR (95% CI) | p-value | ||
| Hypertension | 2.301 (1.590, 3.332) | Hypertension | 2.305 (1.438, 3.696) | ||
| Diabetes | 2.304 (1.474, 3.602) | Diabetes | 1.286 (0.749, 2.209) | 0.362 | |
| Age |
2.130 (1.440, 3.150) | Age |
2.267 (1.392, 3.692) | ||
| BMI |
4.438 (2.925, 6.735) | BMI |
8.634 (4.653, 16.021) | ||
| TC |
6.768 (1.966, 23.299) | TC |
6.437 (1.074, 38.577) | 0.042 | |
| ApoB |
2.461 (1.453, 4.168) | 0.001 | ApoB |
2.504 (1.138, 5.512) | 0.023 |
| Lp(a) |
1.637 (1.045, 2.677) | 0.032 | Lp(a) |
1.422 (0.798, 2.534) | 0.233 |
| WBC |
4.409 (2.312, 8.408) | WBC |
2.148 (0.303, 15.741) | 0.438 | |
| 2.0 |
0.416 (0.258, 0.671) | 2.0 |
1.102 (0.084, 14.544) | 0.941 | |
| NE |
4.877 (2.668, 8.915) | NE |
4.015 (0.214, 75.168) | 0.352 | |
| PLT |
3.452 (2.186, 5.445) | PLT |
3.064 (0.341, 27.560) | 0.318 | |
| rs2000813CC | 0.664 (0.460, 0.955) | 0.027 | rs2000813CC | - | 0.082 |
| rs2000813CT | 1.606 (1.112, 2.318) | 0.012 | rs2000813CT | 1.614 (1.010, 2.579) | 0.045 |
BMI, body mass index; TC, total cholesterol; ApoB, apolipoprotein B; Lp(a), lipoprotein (a); WBC, white blood cell; NE, neutrophil count; PLT, platelet.
This study suggests that the rs2000813 CT genotype may increase the risk of CHD in women, while the rs3813082 locus of the endothelial lipase gene may influence lipid metabolism. To explore these genetic contributions further, we analyzed interactions between these loci and environmental factors—including smoking, alcohol consumption, hypertension, type 2 diabetes mellitus, age, BMI, and specific blood markers—using the MDR method. This approach constructed a multilevel interaction model integrating genetic and environmental variables, demonstrating strong predictive performance (Supplementary Table 6).
MDR analysis focused on interactions between the rs2000813 and rs3813082 loci of the endothelial lipase gene. Supplementary Fig. 1A illustrates risk-factor combinations associated with CHD in women. Dark-colored cells denote high-risk combinations, indicating increased CHD susceptibility, whereas light-colored cells represent low-risk combinations, suggesting reduced disease likelihood. White cells indicate no significant association with CHD. Supplementary Fig. 1B, highlights positive interactions between these loci in red, quantifying their individual main effects. Notably, rs2000813 exhibited a stronger influence on CHD risk than rs3813082.
Supplementary Fig. 2 presents the MDR analysis of interactions between the rs2000813 and rs3813082 loci of the endothelial lipase gene and environmental factors in in relation to CHD in women. Bar histograms distinguish the case group (left) from the control group (right), with dark cells indicating high-risk combinations, light cells representing low-risk combinations, and white cells denoting no significant association with CHD. This visualization elucidates the gene-environment interactions influencing CHD in women, offering insights into the complex interplay of genetic and environmental factors.
Supplementary Fig. 3 depicts interaction tree highlighting patterns among risk factors for CHD in women. Traditional cardiovascular risk factors-including neutrophil count, BMI, platelet count, and WBC-cluster on a single main branch, exhibiting a strong negative interaction (shown in dark blue). The introduction of the rs2000813 locus, alongside other risk factors such as alcohol consumption, age, smoking, lipoprotein(a), type 2 diabetes mellitus, hypertension, apolipoprotein B, and total cholesterol, attenuates this negative interaction, transitioning towards a positive interaction (indicated by green and yellow). Ultimately, a weak positive interaction emerges with rs3813082.
In the interaction ring network of the MDR model (Fig. 1), node values represent the information gain from individual attributes (main effects), while values between nodes reflect the information gain from attribute pairs (interaction effects). The main effects of single attributes, ranked in descending order, are as follows: BMI, NE, PLT, WBC, hypertension, age, type 2 diabetes mellitus, TC, ApoB, smoking, alcohol consumption, rs2000813, Lp(a), and rs3813082. Among the interaction effects, the strongest positive interaction occurs between rs2000813 and rs3813082, whereas the most pronounced negative interaction is observed between BMI and NE.
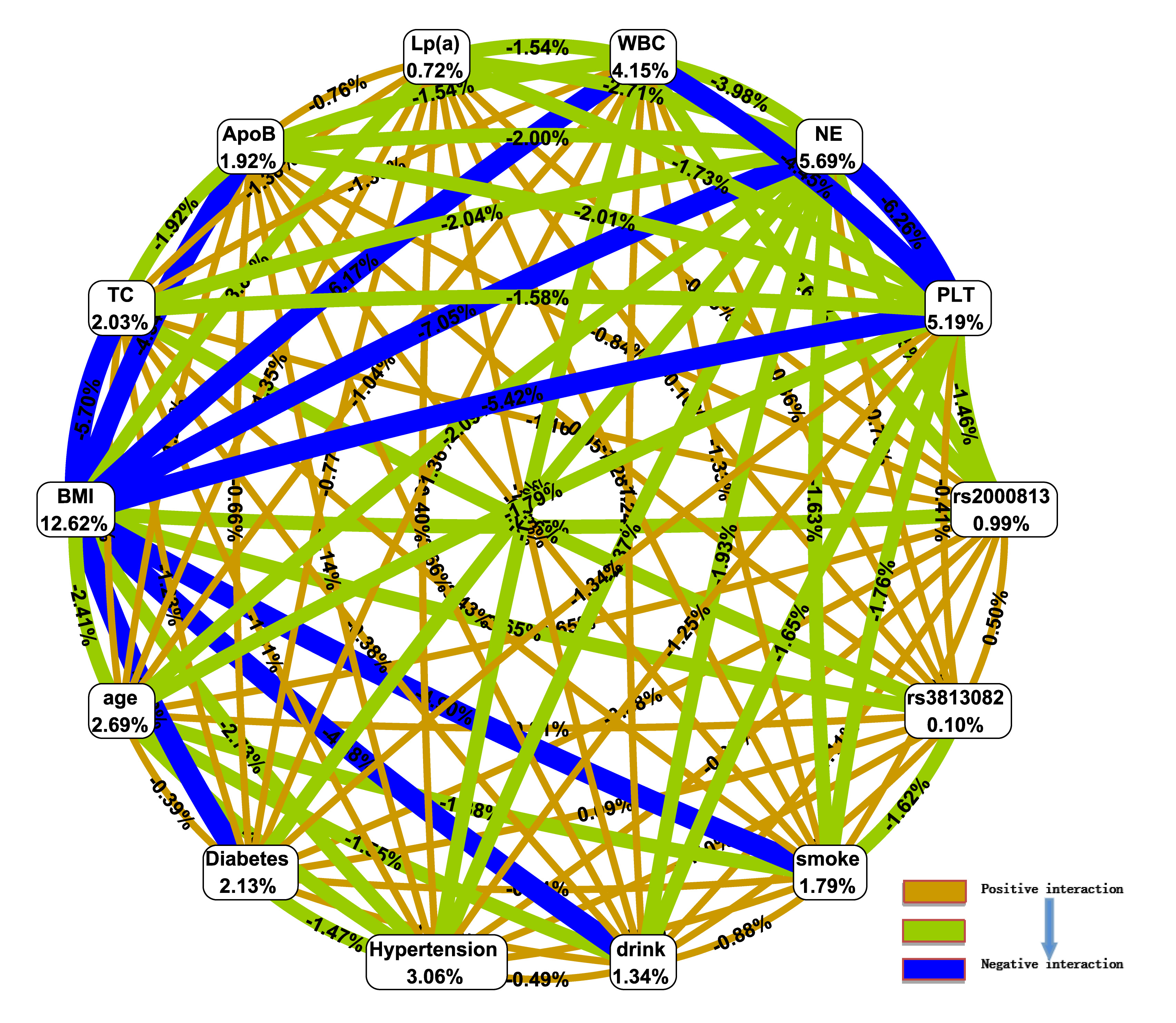 Fig. 1.
Fig. 1.
Interaction ring network of rs2000813 and rs3813082 loci and gene-environment effects in the multifactor dimensionality reduction (MDR) model. WBC, white blood cell; NE, neutrophil count; PLT, platelet; Lp(a), lipoprotein (a); ApoB, apolipoprotein B; TC, total cholesterol; BMI, body mass index.
Logistic regression analysis, conducted using SPSS software (version 26.0),
generated prediction probabilities for four models. Model Y1 was defined by the
rs2000813 CT genotype, while Model Y2 by the rs2000813 CC
genotype. Model Y3 included traditional cardiovascular risk factors (e.g.,
smoking, hypertension, BMI), while Model Y4 combined all Model Y3 factors with
both rs2000813 genotypes. ROC curve analysis revealed that Model Y3
(risk ratio index [RRI]
| Predictors | Youden’s index | Cut off point | Sensitivity (%) | Specificity (%) | AUC (95% CI) | p-value |
| Model Y1 | 0.115 | 0.495 | 0.493 | 0.622 | 0.558 (0.506, 0.610) | 0.030 |
| Model Y2 | 0.102 | 0.487 | 0.550 | 0.552 | 0.551 (0.499, 0.603) | 0.056 |
| Model Y3 | 0.452 | 0.344 | 0.825 | 0.627 | 0.803 (0.765, 0.842) | |
| Model Y4 | 0.458 | 0.515 | 0.686 | 0.772 | 0.804 (0.766, 0.843) |
AUC, area under the curve.
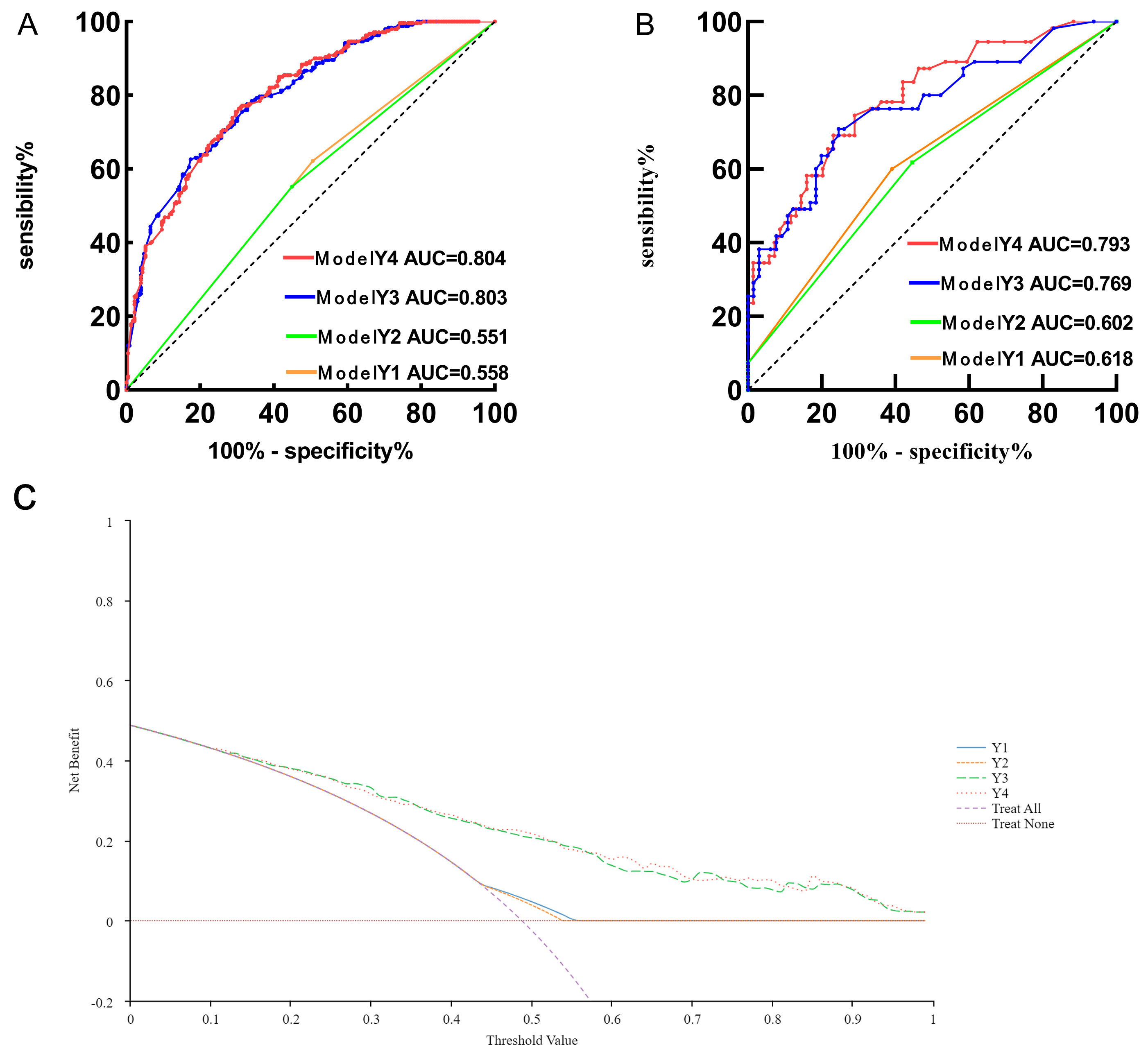 Fig. 2.
Fig. 2.
Performance evaluation of coronary heart disease risk models in female patients — ROC curves for training (A) and validation (B) sets, and decision curves for training set (C). AUC, area under the curve; ROC, receiver operating characteristic.
In this study, 1800 participants were enrolled and divided into a modeling
subset (1440 participants) and a validation subset (360 participants) in an 8:2
ratio. These models were reconstructed and validated in female patients. Of
these, 590 were women, with 470 (229 CHD, 241 Controls) in the modeling subset
and 120 (51 CHD, 69 Controls) in the validation subset. Statistical analysis
confirmed no significant differences in the distribution of factors such as
smoking status, hypertension prevalence, or rs2000813 genotypes between
the two subsets (p
| Items | All (n = 590) | Modeling subset (n = 470) | Validate subset (n = 120) | p-value |
| CHD | 288 (48.1) | 229 (48.7) | 51 (42.5) | 0.476 |
| Smoking history | 10 (1.7) | 8 (1.7) | 2 (1.7) | 1.000 |
| History of drinking | 6 (1.0) | 6 (1.3) | 1 (0.8) | 0.882 |
| Hypertension | 310 (52.5) | 242 (51.5) | 68 (56.7) | 0.598 |
| Diabetes | 128 (21.7) | 107 (22.8) | 21 (17.5) | 0.458 |
| Age |
395 (66.9) | 313 (66.6) | 82 (68.3) | 0.937 |
| BMI |
193 (32.7) | 158 (33.6) | 35 (29.2) | 0.650 |
| TC |
22 (3.7) | 21 (4.5) | 1 (0.8) | 0.172 |
| ApoB |
95 (16.1) | 73 (15.5) | 22 (18.3) | 0.758 |
| Lp(a) |
110 (18.6) | 88 (18.7) | 22 (18.3) | 0.995 |
| WBC |
73 (12.4) | 59 (12.6) | 14 (11.7) | 0.966 |
| 2.0 |
478 (81.0) | 379 (80.6) | 99 (82.5) | 0.898 |
| NE |
88 (14.9) | 71 (15.1) | 17 (14.2) | 0.967 |
| PLT |
131 (22.2) | 114 (24.3) | 17 (14.2) | 0.060 |
| rs2000813CC | 295 (50.0) | 236 (50.2) | 59 (49.2) | 0.979 |
| rs2000813CT | 260 (44.1) | 204 (43.4) | 56 (46.7) | 0.814 |
CHD, coronary heart disease; BMI, body mass index; TC, total cholesterol; ApoB, apolipoprotein B; Lp(a), lipoprotein (a); WBC, white blood cell; NE, neutrophil count; PLT, platelet.
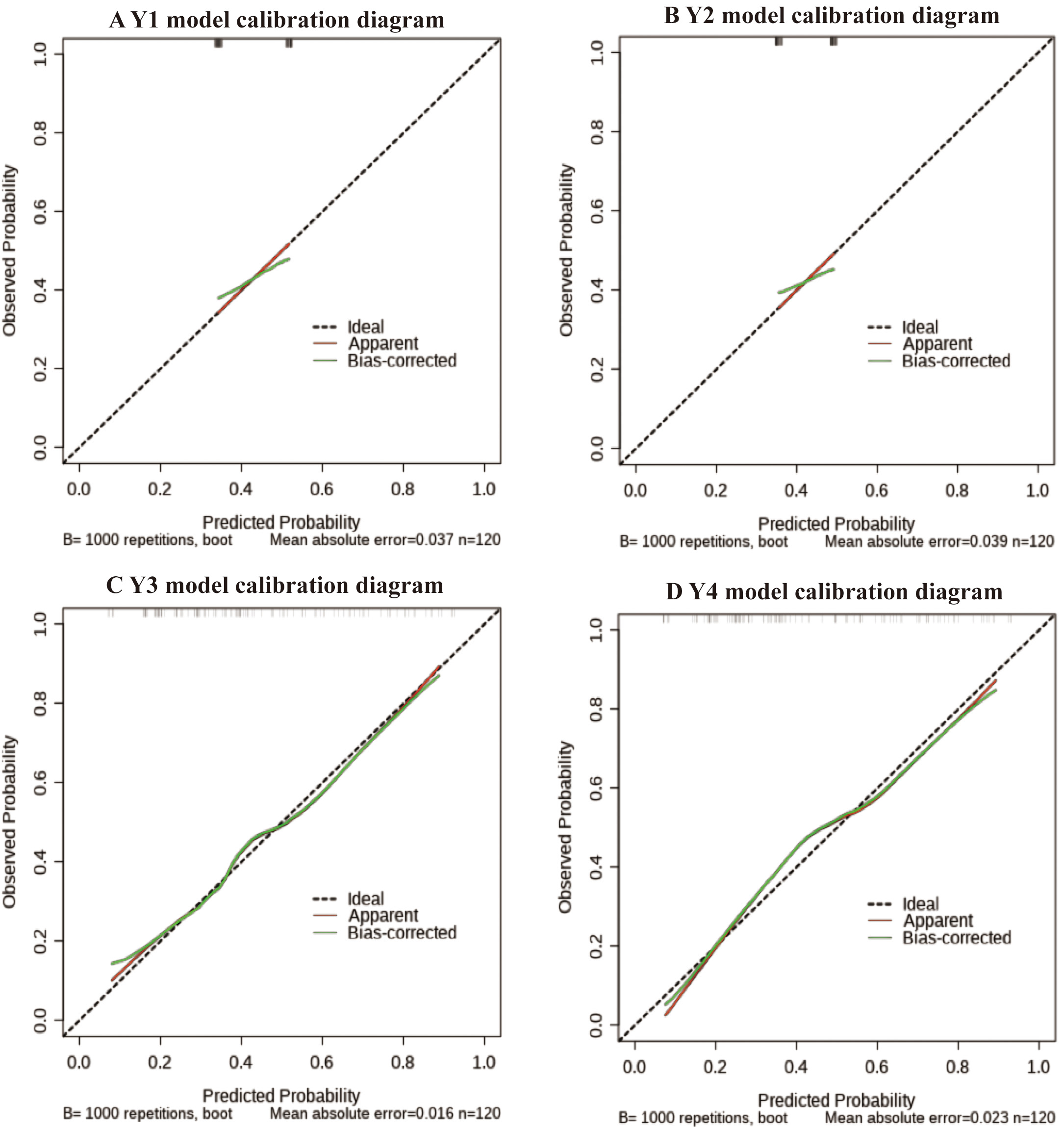 Fig. 3.
Fig. 3.
Calibration diagram of female coronary heart disease risk model. (A) Calibration curve of predicting CHD risk in women in the Model Y1. (B) Calibration curve of predicting CHD risk in women in the Model Y2. (C) Calibration curve of predicting CHD risk in women in the Model Y3. (D) Calibration curve of predicting CHD risk in women in the Model Y4. CHD, coronary heart disease. Calibration curve likely adhered closest to the 45-degree ideal line, indicating minimal deviation between predictions and actual outcomes.
CAD is widely recognized for its genetic basis, with cardiovascular diseases (CVD) development driven by interactions between genetic and traditional risk factors. The association between gene polymorphisms and CHD is a prominent focus of global research due to its implications for understanding and managing CAD [20]. Investigating gene-environment interactions is essential not only for addressing the “missing heritability” of complex traits but also for elucidating the biological mechanisms underlying multifactorial diseases. This approach is central to genetics, providing deeper insights into how genetic and environmental factors jointly shape complex health conditions [21]. Genetic variants may not directly cause disease but can influence its onset and progression through subtle interactions with other genes or environmental exposures. This complexity underscores that genetic contributions to disease often manifest significantly only in combination with additional genetic or environmental factors [22]. Studying these interactions is critical for understanding disease etiology and the impact of environmental exposures on health outcomes. Focusing on clinical data and EL gene polymorphisms in CHD patients, this research aims to uncover associations between gene interactions and disease susceptibility. It also seeks to develop a risk prediction model to identify high-risk populations, offering valuable insights for targeted public health strategies and interventions.
Endothelial lipase, encoded by the endothelial lipase gene, is predominantly synthesized by vascular endothelial cells and exhibits robust phospholipase activity with minimal triglyceride lipase activity. This enzyme critically regulates serum HDL-C metabolism, influencing cholesterol transport and cardiovascular health [23]. HDL-C is widely acknowledged for its protective role against atherosclerosis, mediated by anti-inflammatory and antioxidant effects, inhibition of platelet aggregation, and promotion of re-endothelialization. Its association with the incidence and prognosis of CHD underscores its cardiovascular significance [24]. Initially, endothelial lipase was thought to contribute to CHD pathogenesis primarily through HDL-C regulation. Subsequent research, however, demonstrated additional roles, including facilitating macrophage adhesion to the vascular endothelium, promoting platelet aggregation and endothelial cell proliferation, and contributing to atherosclerosis. Notably, suppressing endothelial lipase expression during inflammation may reduce the severity of coronary atherosclerosis [25]. Among the triglyceride lipase family, endothelial lipase is considered the most influential in atherosclerosis, emphasizing its relevance to cardiovascular diseases research [26].
In this study, the rs2000813 CT genotype of the endothelial lipase gene was identified as a potential risk factor for CHD in women, whereas the CC genotype exhibited a protective effect. These findings align with prior study linking endothelial lipase gene single-nucleotide polymorphisms (SNPs) to increased CHD risk, reinforcing the role of genetic factors in this disease [27]. However, Elnaggar et al. [28] suggested that the T allele of the rs2000813 (584C/T) variant may confer protection against CHD, implying a reduced risk for carriers. In contrast, Zhao et al. [29] reported that the CC genotype and C allele may predispose individuals to higher CHD risk, highlighting potential genetic contributions to disease susceptibility. The rs2000813 genotype also appears linked to sex-specific differences in CHD incidence, potentially influenced by variations in sex hormone levels. Mehilli and Presbitero [30] underscored significant disparities in clinical presentation, complications, and cardiovascular risk profiles between men and women with CHD, emphasizing the need to consider sex-specific factors. Women’s susceptibility to endothelial dysfunction and occult CHD may stem from unique factors such as inflammation, mental stress, and autonomic or neuroendocrine dysregulation [31], contributing to sex-specific cardiovascular risks. The increased CHD risk in women with the rs2000813 CT genotype may reflect its specific effects on lipid metabolism and related pathways, though further research is required to elucidate the underlying mechanisms.
In this study, no significant association was observed between
rs3813082 polymorphism of the endothelial lipase gene and CHD
(p
This study employed multifactor dimensionality reduction analysis to examine gene-gene and gene-environment interactions in CHD. A positive interaction was identified between the rs2000813 and rs3813082 loci of the endothelial lipase gene. Among individual attributes, the main effects ranked in descending order of influence were BMI, NE, PLT, WBC, hypertension, age, type 2 diabetes mellitus, TC, ApoB, smoking, alcohol consumption, rs2000813, Lp(a), and rs3813082. The strongest interaction occurred between rs2000813 and rs3813082.
Hartiala et al. [32] found that smoking attenuates the protective effect of the ADAMTS7 gene on cardiovascular health, while increased physical activity amplifies the influence of three genetic loci on serum HDL-C levels and diminishes the effect of another locus. Air pollution, meanwhile, elevates cardiovascular disease risk by altering CXCL12 expression, and X chromosome variants significantly regulate sex differences in cardiovascular disease incidence [32]. In this study, significant interactions were identified between the rs2000813 and rs3813082 loci of the endothelial lipase gene and environmental factors-including BMI, NE, PLT, WBC, hypertension, age, type 2 diabetes mellitus, TC, ApoB, smoking, alcohol consumption, and Lp(a)-collectively influencing CHD risk in women. These gene-environment interactions deepen our understanding of CHD pathogenesis and genetic susceptibility, offering critical insights for developing tailored lifestyle recommendations and therapeutic strategies.
Complex interactions between SNPs and traditional cardiovascular risk factors
are pivotal in CHD pathogenesis. Integrating these factors with SNPs in
CHD-related genes enhances our understanding of disease mechanisms. Notably,
severe coronary atherosclerosis may develop asymptomatically for years before
angina manifests [33]. Proactively identifying high-risk individuals and managing
modifiable risk factors can substantially reduce adverse cardiac events [34].
Developing an effective risk prediction model based on clinical and laboratory
data is essential for pinpointing high-risk populations, enabling targeted risk
factor management, and optimizing healthcare resource allocation. For instance,
Pattarabanjird et al. [35] combined the TID3 rs11574
polymorphism with traditional risk factors, achieving 87.0% accuracy and an AUC
of 0.840 in predicting CAD severity, markedly improving prediction efficiency. In
this study, we integrated endothelial lipase gene polymorphisms
(rs2000813, rs3813082) and traditional CHD risk factors
(smoking, alcohol consumption, hypertension, type 2 diabetes mellitus, age
This study has several limitations. First, as a single-center study with a relatively small sample size, it may be subject to selection bias, potentially limiting the generalizability of the findings. Second, the prediction model underwent only internal validation, without external validation, necessitating further assessment of its clinical applicability and broader relevance. Consequently, future research should prioritize large-scale, multicenter, prospective studies to validate the predictive utility of endothelial lipase polymorphisms (rs2000813 and rs3813082) combined with traditional cardiovascular risk factors for CHD incidence in women.
This study identified the rs2000813 CT genotype of the endothelial lipase gene as a potential risk factor for CHD in women. Furthermore, a synergistic interaction between endothelial lipase gene polymorphisms and environmental factors appears to influence CHD susceptibility in women. When integrated with traditional cardiovascular risk factors, this model exhibits robust predictive performance for CHD in women.
Data supporting the findings of this study are available from the corresponding author upon reasonable request within 1 year of publication of this article.
CH and XS have contributed equally to this work. CH, XS and YG designed the research study. XS, YG formal analyzed. XS, HA, YZ collected data. HQ, DA, MZ, JY calibrated data. CH, XS wrote the manuscript. TH, QT, YG, JZ reviewed & edited the manuscript. All authors contributed to the conception and editorial changes in the manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work. All authors read and approved the final manuscript.
This study was approved by the Ethics Committee of First Affiliated Hospital of Xinjiang Medical University (registration number K202309-08) and was conducted in accordance with the Declaration of Helsinki and Good Clinical Practice Guidelines. Also, written informed consent was obtained from all subjects included in the study.
The authors would like to thank Dr. Deyang Li for his valuable help with the statistical advice on the web interface.
This study was supported by Tianshan Elite Science and Technology Innovation Leading Talents Program of Xinjiang Uyghur Autonomous Region (High level Leading Talents; grant No. 2022TSYCLJ0023); National High Level Hospital Clinical Research Funding (2023-GSP-QN-36); Key Project of Natural Science Foundation of Xinjiang Uygur Autonomous Region (Grant No. 2023D01D13); Beijing Natural Science Foundation (grant number 7222143); National High Level Hospital Clinical Research Funding (grant number 2022-GSP-GG-9).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/RCM37356.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.



