1 School of Information Studies, Syracuse University, Syracuse, NY 13244, USA
Abstract
Knowledge Organization (KO) has historically been used to structure biological knowledge, from taxonomy to ontologies. This becomes increasingly challenging as life sciences evolve into a data-intensive domain. The advent of artificial intelligence (AI) has enabled knowledge organization systems (KOSs) to assume active roles in computational workflows rather than serve as passive repositories. This thematic review examines the evolution of KOSs in AI-augmented biological research by situating them within scientific paradigmatic and epistemological shifts. By synthesizing foundational theories from library and information science, philosophy of science, and biological systematics, we propose the Knowledge Organization Analysis Framework (KOAF) to capture bio-KOSs’ developments across functional sophistication, automation degree in system construction, and reasoning and inference capability. Representative empirical studies show that bio-KOSs enable semantic interoperability and data integration, while also contributing to hypothesis generation and reasoning. We argue that advanced bio-KOSs increasingly function as epistemic agents in scientific discovery. This transformation marks KOSs as theoretical frameworks shaping scientific inquiry through AI-KO convergence and highlights the need for future research on accountability, epistemic integrity, and scientific trustworthiness in AI-driven knowledge discovery.
Keywords
- knowledge organization systems
- biological ontologies
- AI-driven biological research
- knowledge discovery
Emerging artificial intelligence (AI) models and algorithms are accelerating scientific inquiry and knowledge discovery. This is evident at different stages of the research process, including literature search, hypothesis generation, experimental design, data collection and analysis, and scientific writing (Reddy and Shojaee, 2025; Salvagno et al., 2023; Wang et al., 2023). During the COVID-19 pandemic, AI was applied to monitor outbreaks, trace contacts of infected individuals, and develop drugs and vaccines (Vaishya et al., 2020). Evidence shows that retrieval-augmented generation (RAG) can embed external knowledge bases (KBs) or knowledge graphs (KGs) to improve machine learning (ML) model performance in ideation and innovation (Wang et al., 2024). A new term, “AI scientists”, has emerged to describe AI agents that can assist humans in research or even automate scientific inquiry (Lu et al., 2024; Pool, 2024). Studies have found that AI scientists can generate insightful research ideas, design and execute experiments, and draft a full scientific paper when trained with structured domain-specific knowledge (Gottweis et al., 2025; Kumbhar et al., 2025; Lu et al., 2024; Si et al., 2024).
Life sciences provide a compelling example of using AI to power research and innovation. Using well-established knowledge organization systems (KOSs), AI and other ML models significantly increased researchers’ abilities to collect and annotate large-scale experimental data with greater precision and proficiency. For instance, PubMedBERT, a biomedical natural language processing (NLP) model pretrained with PubMed data, achieved better performance using domain-specific vocabulary and entities compared to baseline Bidirectional Encoder Representations from Transformers (BERT) (Gu et al., 2021). Doğan and colleagues annotated the National Center for Biotechnology Information (NCBI) disease corpus, a collection of 793 PubMed abstracts in which mentions of diseases were mapped with Medical Subject Headings (MeSH) and Online Mendelian Inheritance in Man (OMIM). Their annotations provided a high-quality gold standard for ML-based applications (Doğan et al., 2014).
Synthesizing heterogeneous data from domain-specific databases and repositories can enable AI models to conduct complex scientific reasoning. The BioLunar framework developed by Wysocki et al. (2024) used Large Language Models (LLMs), a type of generative AI, to harmonize distributed experimental evidence of disease inference and discovery. They integrated multiple KBs including the Clinical Interpretation of Variants in Cancer (CIViC), Oncology Knowledge Base (OncoKB), Gene Ontology (GO), Human Kyoto Encyclopedia of Genes and Genome (KEGG), Reactome, WikiPathways, and PubMed API to continuously compile clinical, experimental, and population genetic study outcomes. Together, these KBs were implemented into LLM through RAG to construct LLM-supported scientific workflows that make way for automatic scientific discovery.
Operationally, these advancements rely on knowledge infrastructures to provide the models with training data. At a conceptual level, classification systems define the scope of concepts and relations based on empirical evidence. A crucial issue is how to effectively classify knowledge to reflect verified truth to the broadest extent possible for AI models to use (Beghtol, 2009; Hjørland, 2016). Multiple classification theories have been proposed to address this question in library and information science (LIS). Early LIS theorists such as Bliss proposed that the most effective library classification systems are those that align closely with the scientific classification they represent (Bliss, 1935). The theory of scientific and educational consensus advocates that Knowledge Organization (KO) should be based on the consensual foundations established by knowledgeable people in their own studies and mirror the ways they acquire knowledge (McDonald and Levine-Clark, 2019). However, this approach has been challenged as it only considers consensual knowledge and excludes conflicting interpretations of reality.
KOSs are not simply spectrums of ‘formality’. Instead, they are selections of properties that represent domains and the semantic relationships between concepts within (Hjørland, 2015). The knowledge about a concept reflects not only a representation of its features, but also an explicit representation of the causal mechanisms that link those features to form a coherent whole (Spiteri, 2008). The pragmatic purpose of KO incorporates the principle of maximal efficiency to theories of consensus. To achieve maximum efficiency in information retrieval, the representation and organization of knowledge ought to be domain-/situation-specific yet dispersed enough for both humans and machines to rapidly recognize what is the right knowledge for detailed circumstances and to find relevant knowledge.
However, the rise of data-intensive sciences challenges conventional KO for scientific knowledge, which Hope et al. (2023) termed the “outer world”, in contrast to a scientist’s “inner cognitive world”, bounded by information processing capacity and personal knowledge. Unprecedented volume, velocity, and heterogeneity of data, from benchmark experimentations to model predictive results, increases the scale and complexity of knowledge organization practices. The explosion of information challenges the limitations of human cognition to find, assimilate, and manipulate information in order to perform synthesis and reasoning (Hope et al., 2023). Contemporary biological discovery requires coordination of complex multi-step data analysis pipelines and interpretation of controlled experiment results by applying evidence from different curated databases and literature (Wysocki et al., 2024). These developments create a critical need for KOSs to provide accurate, relevant, and structured knowledge to interoperate with AI applications and scientific research workflow management systems.
A more pressing question is whether rule-based, symbolically logical, propositionally friendly, and situation-specific organizations of knowledge remain the best solution for AI to conduct scientific reasoning and discovery. Researchers in knowledge representation (KR) have begun to challenge the predominant logical approaches to representing semantic knowledge and propositional language of thought (Browning and LeCun, 2023). Despite the success of current statistics-based LLMs on benchmarks such as the Turing Test and the Winograd schema challenge, researchers are doubtful whether these systems truly possess semantic understanding, as they lack explicit symbolic representations (Browning and LeCun, 2023). This raises fundamental questions: Does the success of these statistical models reveal inherent limitations in propositional approaches to knowledge representation? Can semantic understanding emerge from patterns in data without formal logical structures?
These theoretical debates are not merely philosophical; they have direct implications for how AI systems engage with complex scientific domains such as biology. The dynamics in current biology research and AI-powered scientific knowledge discovery pose pressing practical challenges to existing knowledge organization theories and practices. Researchers expect AI systems to not just identify problems, but capable of interpreting them and explaining their solutions, a process that often remains opaque in current AI analysis (Han and Liu, 2022). This “black box” problem has prompted biomedical researchers to turn to explainable AI (XAI) that provides transparency in decision-making (Karim et al., 2023). Meanwhile, LLMs excel in NLP, enabling the models to streamline literature reviews, synthesize heterogeneous research findings, and generate hypotheses (Thapa and Adhikari, 2023). However, to perform any of these tasks effectively, LLMs need extensive pre-training with comprehensive, domain-specific, and up-to-date knowledge.
This thematic review surveys generative AI applications in scientific research to examine the evolving roles of KOSs in AI development implementation. We focus specifically on biological and biomedical research, where the heterogeneous and large-scale data present significant challenges in AI model precision and interoperability. While collaboration between human scientists and generative AI has become more prevalent, it is grounded in preexisting bodies of biological knowledge rather than originating independently. Biological KBs accumulated from years of work can and should play an important role for high-quality AI modelling and applications. Throughout this review, “AI” primarily refers to LLMs. Other relevant ML models are discussed but not prioritized.
Applying a thematic analysis approach allows us to synthesize relevant literature and sift out patterns and trends. We aim to answer the following questions:
RQ1: How do different types of KOSs in biological sciences vary across functional sophistication, degree of automation in system construction, and reasoning capabilities in supporting AI-augmented research?
RQ2: How does the functional evolution of KOSs empowered by AI technologies propelled the paradigmatic transitions in biological research?
The structure of this thematic review will be as follows. First, KO and classification theories are introduced to guide this study. Next, we define biological science paradigms through philosophy of science theories and review major developments in AI-assisted biological research. We combine KO and KR theories to address our research questions. Finally, we conclude with future directions of KO for AI-assisted biological research.
This study adopts a thematic review approach to provide a general understanding of theories of KO and their empirical applications in biological science research, ranging from traditional KOSs to contemporary infrastructures embedded within AI models. This study does not aim to compile a comprehensive body of literature for an exhaustive review but instead prioritizes publications of theoretical significance and intellectual diversity that are representative in this domain. We analyze these articles by synthesizing major themes, paradigmatic examples, and critical perspectives that illuminate the evolving role of KO in AI-assisted biological research.
We manually identified and selected relevant literature through iterative keyword searches in scholarly databases, including Google Scholar, Semantic Scholar, and PubMed. Keywords and search terms were adapted and refined throughout the process to reflect emerging patterns and conceptual intersections across biology, KO, and AI. To broaden the research, alternative terms such as knowledge graph and knowledge representation were also used in place of KO. In addition to database searches, snowball sampling techniques (e.g., forward and backward citation tracking) were employed to capture high-impact studies. ArXiv and BioRXiv were further consulted to include preprints in AI and biological research.
The literature reviewed in this study is organized into two tiers. The first tier includes papers that describe and discuss conceptual frameworks for constructing and applying KO with AI technologies. We selected literature based on its theoretical significance and empirical contributions to understanding the evolving relationship between KO and AI-assisted biological research. This foundational set of work sets up the theoretical groundwork for examining science paradigms, classification theories, and the epistemological foundations of biological systematics, drawing from LIS, philosophy of science, and biological classification theory. Priority was given to influential publications that have shaped past and contemporary understanding of KOSs, the formation and transformation of scientific paradigms, and the evolution of biological classifications. Studies at the intersection of KR and AI research were also included to bridge traditional classification approaches with emerging computational methodologies.
The second tier focuses on empirical studies that demonstrate the active implementation of KOSs in contemporary biological research, particularly those incorporating AI technologies. Literature was excluded if it focuses solely on technical implementation details without addressing broader implications for KO theory or biological research paradigms. These studies include, for instance, efforts to construct KGs as advanced mapping approaches for organizing biological knowledge. Studies addressing KO in non-biological domains were excluded unless they offered theoretical insights directly applicable to biological classification systems. Papers presenting only preliminary results, purely speculative frameworks lacking empirical validation, or insufficient methodological detail for dimensional assessment were also excluded from the target paper pool, although some were kept as background literature when they contributed valuable theoretical insights.
Literature in the second tier, referred to as target papers, was evaluated using the proposed Knowledge Organization Analysis Framework (KOAF). This framework is built on three dimensions: functional sophistication (ranging from simple tasks such as terminology disambiguation to more advanced operations such as AI-driven knowledge discovery), automation degree in system construction (from manual curation only to fully automated systems), and reasoning and inference capability (from basic lookup functions to hypothesis generation). These three dimensions were designed to better structure the review of selected papers as well as serve as potential evaluation criteria for AI-powered bio-knowledge systems. The framework also helped us decide the target paper selection criteria: papers for the second tier should present concrete applications of KOSs in biological research contexts, demonstrate measurable impacts on research workflows or discovery processes, and provide sufficient technical detail to enable dimensional analysis within the KO analysis framework.
The final target paper pool includes 27 scholarly publications, spanning from 2003 to 2025 (see Appendix Table 4 for the full list). The analysis result on the target papers is visualized using the plotly R package (Sievert, 2020). We acknowledge that these 27 papers in the list may not be sufficiently inclusive of all existing literature that would qualify under the inclusion criteria. Given the rapidly expanding intersections of KO, AI, and biological research, a comprehensive coverage of all relevant studies is neither feasible nor necessary for the thematic analysis approach employed in this review. Instead, the papers included in this review were chosen to give representative examples across the three dimensions of the KO analysis framework, ensuring adequate coverage of different functional sophistication levels, degrees of automation in system construction, and reasoning capabilities. The temporal span reflects both the historical development of computational approaches to biological knowledge organization and the recent acceleration while keeping a focus on contemporary applications and emerging paradigms. The iterative selection process allowed refinement of inclusion parameters as emerging themes and conceptual intersections became clearer, ensuring that the final literature corpus represented both the theoretical foundations and empirical manifestations of knowledge organization systems in AI-augmented biological research.
To understand the evolving role of KO in AI-powered scientific discovery, we begin by situating KO within the context of scientific paradigms. Scientific paradigms define the intellectual boundaries and methodological norms within which research is conducted, thus shaping how knowledge is classified, interpreted, and disseminated (Kuhn, 1962). As scientific inquiry becomes increasingly interdisciplinary and data-intensive, traditional KO models and theories are being challenged and redefined. Beginning with paradigms allows us to critically examine the conceptual foundations upon which KO systems are built, and to assess their adaptability to the current scientific realm.
In LIS, KO refers to activities such as describing, representing, and organizing documents, works, and concepts. In broader sense, KO can be understood as an institutional practice involved in the production and dissemination of knowledge. The narrower definition applies primarily within the LIS domain and encompasses document description, indexing and classification activities carried out in libraries, bibliographical databases, archives, and other forms of “memory institutions” (Hjørland, 2008).
Conventionally, librarians, archivists, and information specialists performed these practices. Today, technological advancements enable the embedding of well-structured KOSs into various tools and systems to perform the tasks that once required extensive human intervention. These developments have, in turn, driven advances in KOS design. With varying degrees of complexity and purpose, embedded KOSs help reduce ambiguities, control synonyms, and establish explicit semantic relationships between concepts based the types of information being handled (Zeng, 2008).
KO practices are not static. Their evolution reflects perceptual changes of similarity and difference among the objects we see. These categorization and classification abilities, fueled by cognition, in turn reinforce how we see things and the relations between them (Atran and Medin, 2008). In the field of biology, the science of classification formed its separate paradigm—systematics. Once known as “the Queen of Sciences”, systematics contributes to taxonomic theories that manage and interpret vast amounts of potentially conflicting information about the origins of life and organismal features, guided by diverse philosophical ontologies such as foundationalism and constructive empiricism (Brower and Schuh, 2021).
Linnaeus’s Systema Naturae endured for centuries because of its ability to convey, through language, the observed hierarchical structure of the living and inanimate world (Padial and De la Riva, 2021). Over time, several schools of philosophical perspectives emerged, from early taxonomy theory to Darwinism phylogenetic inference. Darwin paved the path towards a paradigmatic shift that gradually replaced both traditional classifying methods of species (e.g., by similarities or differences, degrees of divergence, or diagnosed populations) and the abstract notion of hierarchical levels attributed to natural or divine order during Linnaeus’s era. In contrast, phylogenetic trees offered a newer and more complete synthesis, deciding species classification by their phylogenetic lineage. Relationships cannot be directly observed but must be inferred through empirical evidence and patterns of variation (De Queiroz, 2007; Sites Jr and Marshall, 2004; Wiley, 1978). This framework shifted the focus of evolutionary biology from intrinsic reproductive isolation as the sole driver of speciation towards a more integrated genetic understanding of the origin of species (Padial and De la Riva, 2021).
Since systematics studies the observable morphological similarity and also fosters ontological discussions of nature and origins of biological diversity, biological classifications have significant scientific value as indices of accumulated knowledge (Brower and Schuh, 2021). As Foucault (1970) described in The Order of Things, the naturalists “distinguishes the parts of the natural bodies with his eyes, describes them appropriately according to their number, form, position, and proportion, and he names them”, emphasizing that the classification depends on what can be perceived and conceptualized within a given historical framework. Under this view, every classificatory system reflects not a timeless natural order but the epistemic conditions of its era. Brower and Schuh (2021) similarly argue that biological classifications embody the scientific perspectives and methodological constraints within which they are produced. Yet, past errors and the abandonment of outdated beliefs retain value, as they represent the ongoing confrontation with complex, indirect, fragmented evidence analyzed through assumptions that are themselves subject to refutation (Padial and De la Riva, 2021). Collectively, these perspectives reflect the underlying values of historically evolving systems of KO. Each system indexes how biological species were understood at a particular moment in scientific history.
When Kuhn first introduced the term paradigm, he argued that scientific paradigms emerge through the continual identification of anomalies and repeated experimentation that generate new narratives within existing domain. Observations evolve into rules, rules develop into theories, and theories show the invisible boundaries of scientific consensus that constitute paradigms. The discovery of knowledge, therefore, is not a linear process but one driven by recurring cycles of anomaly detection and conceptual reconstruction (Kuhn, 1962). Scientists owe their success to their ability to identify and solve problems using conceptual and instrumental techniques that already exist within the prevailing paradigm.
The contemporary scientific process, however, is different from the one Kuhn described. Unlike in Kuhn’s time, most researchers today are trained and socialized within paradigm-centered scientific communities, where their work is shaped by daily engagement with experiments and collaborative infrastructures. Modern science advances increasingly through networked accumulation and tools-mediated integration. The exponential growth of scientific knowledge gives rise to competition between scientists who pursue similar goals with comparable training, ambition, and resources (Wang and Barabási, 2021). To reach the frontiers of discovery more efficiently, scientists have narrowed their specialization and collaborate extensively to integrate diverse knowledge backgrounds essential for modern scientific research (Wang and Barabási, 2021).
Jones described this phenomenon as the “burden of knowledge”, arguing that innovators must undertake extensive education and training to reach the frontier of discovery (Jones, 2009). As scientific knowledge continues to expand exponentially, the structure of research itself has evolved. The growing complexity of contemporary science challenges the solitary model of inquiry that characterized earlier paradigms. One major response to this challenge is the rise of multidisciplinary collaboration. This increasing collaboration reflects the difficulty of conducting complex studies individually. This is not because scientists today are less productive (in fact, individual productivity is stably rising from 1900 to 2000), but because the amount of knowledge has been accumulating from the exponential growth of science and fewer new paradigm shifts happen within the existing fields (Gingras and Wallace, 2010).
Therefore, Kuhn’s view that emerging scientific paradigms must derive from new theories may not be entirely suitable for describing contemporary scientific discovery. Ankeny and Leonelli (2016) argued that paradigms are no longer useful as a framing concept for three reasons. First, paradigms tend to be highly static and inflexible in which changes only occur in dramatic fashion. Second, Kuhn considers conflicting paradigms to be incommensurable, therefore unable to coexist except in extreme moments of crisis. This does not align well with current interdisciplinary research, where different epistemic approaches often appear in parallel or at the intersection between paradigms to uncover questions. Third, Kuhn prioritizes theoretical knowledge as the primary output of science, while marginalizing applied science and technological innovations. Kuhn also limited scientific communication to textual documents, excluding other forms such as datasets, graphs, and charts from his definition of legitimate scientific outputs (Ankeny and Leonelli, 2016). A fourth point can be added to this argument, that is, the melding and interconnections of multidisciplinary scientific research do not follow the dramatic rupture with which Kuhn associated the emergence of new paradigms.
Instead, the new paradigms reflect a gradual convergence of existing domains, blurring disciplinary boundaries through the alignment of key concepts. These concepts, while semantically and distinctively defined, imply overlapping conceptual meanings. Much of today’s scientific progress, particularly in fields such as bioinformatics and computational biology, is driven less by theoretical contributions but more by revealing hidden patterns and relationships among large and complex data entities (Leonelli, 2016). In biology, Noble (2002) characterized this integrative approach through a “middle-out” framework for biological modeling, in which researchers work simultaneously upward from molecular mechanisms to physiological systems and downward from system functions to cellular components, with continuous integration occurring at intermediate levels. This approach transcends the traditional dichotomy between reductionist “bottom up” and holistic “top down” strategies. Data as empirical evidence have theoretical value as it embodies knowledge through classification (Leonelli, 2016). This coincides to Hjørland’s point that knowledge is embedded in larger frameworks of human activities (Hjørland, 2015). In data-intensive biology, research often starts at whichever level with sufficient data (e.g., pathways, cells, tissues, organs) and extends both vertically and horizontally to connect across multiple biological scales.
The application of NLP in biomedical research enables information extraction from electronic health records for hidden clinical narrative discovery (Houssein et al., 2021). Similarly, the development of GenBank, a public repository of annotated nucleotide sequences maintained by the NCBI, shows how scientific progress can be driven by the creation of shared data ecosystems that support reproducibility, retrieval, and the synthesis of biological knowledge (Benson et al., 2014). These examples of interdisciplinary fusion represent forms of convergence, not “paradigm shifts” in Kuhn’s conventional sense, but rather, gradual adaptations across domains facilitated by interoperable terminology, method, and data infrastructures.
This emerging paradigm of science combines individual efforts into a form of collective intelligence, defined as the general ability of a group to perform a wide variety of tasks and achieve deeper integration of scientific knowledge (Woolley et al., 2010). This shift raises questions about how organization of scientific knowledge within interdisciplinary paradigms are conceptualized evolve in a consistent pace with frontier discoveries. Effective classifications of knowledge enhance researchers’ capacity to identify connections across diverse literature or databases, thereby generating novel combinations of previously distinct insights (Szostak et al., 2016). As interdisciplinary research increasingly involves larger teams and more heterogeneous datasets, classification systems and KOSs at large are being adopted as foundational knowledge in science processes, even though such a role of KOSs is not always fully recognized by the researchers who adopted them.
Unlike paradigms that guide research through agreed identification and interpretation of concepts and epistemology (Kuhn, 1962), KOSs can support research by offering semantic alignment, enabling diverse teams to describe, use, and interpret data in consistent ways. Researchers who use KOSs implicitly accept, even if they might not be aware of it, the definition of entities and processes contained within KOSs at the moment they consulted it (Leonelli, 2016). For example, GO is one of the most successful bio-ontologies that offers comprehensive, structured, computer-accessible representation of genes and genetic functions (Ashburner et al., 2000; Pesquita et al., 2009). It provides a framework and a set of concepts for describing the functions of gene products by assigning GO annotations to the association between a specific gene product and a GO concept. Together, they make a statement pertinent to the function of that gene. There are three types of GO concepts in GO to represent the distinct aspects of how gene functions can be described: molecular function, cellular component, and biological process (Dessimoz and Škunca, 2017). The logic is that a gene encodes a gene product, and that gene product carries out a molecular-level process or activity (molecular function) in a specific location relative to the cell (cellular component), and this molecular process contributes to a larger biological goal (biological process).
A biologist who uses GO data for their research underlyingly means they accept the descriptive predicates that are in the GO framework, including definitions and conceptual boundaries. Descriptive metadata captured in KOSs thus have the same epistemic value as testable hypotheses: they are theoretical statements whose validity and meaning can only be interpreted and assessed with empirical evidence and the conditions under which that evidence has been produced. This crucial role in expressing disciplinary embodied knowledge qualifies KOS as a form of theory (Hjørland, 2015; Leonelli, 2016).
The convergence of scientific paradigms and the increasing reliance on structured relational KBs in interdisciplinary research reflect a growing need for shared understanding, often referred to as common knowledge (Neumann, 2010). This includes a collective foundation of terms, concepts, assumptions, and frameworks that enable coordination and collaboration across diverse domains. Although conceptual meanings may vary across disciplines, such ambiguity can be minimized using precise and formal structures of language. Conceptual clarity depends not only on empirical accuracy but also on the formal structure of the language used to express it (Reichenbach, 2012). In contemporary science, this shared structure is frequently established through human-mediated KR (Qin, 2020), where ontologies and classificatory systems mediate raw data and conceptual reasoning.
A related concept in linguistics and philosophy of AI is common sense, which serves as a measurement of whether an AI system is sufficiently intelligent to process and understand human language and its underlying meaning, thereby performing referring and reasoning (Browning and LeCun, 2023). For humans, common sense runs subconsciously in almost every decision in daily life. For instance, in the Winogard Schema Challenge, the following pair of sentences is used to test a machine’s capacity for common-sense reasoning:
The trophy doesn’t fit inside the suitcase because it is too large.
The trophy doesn’t fit inside the suitcase because it is too small.
Common sense allows us to effortlessly determine what “it” refers to in each sentence, based on the implied size difference. Although LLMs can pass such challenges, it does not necessarily imply that LLMs can truly understand the meaning of semantic knowledge and common sense (Browning and LeCun, 2023). Studies also show that LLMs can exhibit hallucinations by fabricating evidence that contradicts real-world facts (Huang et al., 2025b; Lu et al., 2024). While deploying RAG into LLMs can reduce hallucination, the performance can vary depending on how well models architecture uses external KBs, where in these situations they function as a form of semantic modeling.
KOSs map between model-theoretic representations of scientific domains and the natural language expressions researchers use to describe them. By codifying knowledge into structured forms, it enables conceptual regularity to support interpretation and semantic interoperability. Establishing connections between structured representations of knowledge and natural language is essential to enabling AI to not only process data, but to reason with it, thus truly achieving trustworthy AI-driven science inquiry.
KOSs can be categorized into four types by structural sophistication. The simplest are term lists, such as glossaries or dictionaries of biological terms. Term lists are sets of terms often arranged in alphabetical order. The second type includes structured representation of entities such as places, organizations, and persons, typically taking the forms of gazetteers, directories, or name authority files. The third type makes up classification and categorization systems, including subject headings, classification and categorization schemes, and taxonomies. This category of KOSs aims to reduce ambiguity, control synonyms, and establish relationships between entities. The Linnaean taxonomy and contemporary phylogenetic classification systems are representative examples of KOSs.
The fourth and most complex type consists of relationship models, such as thesauri, semantic networks, and ontologies. These systems explicitly represent concepts with unambiguous terms and relationships between entities not limited to hierarchical structure. For example, the Unified Medical Language System (UMLS) semantic network contains 127 semantic types and 54 relationships of concepts represented in the UMLS Metathesaurus for biomedicine.
Ontologies are sometimes referred to as KBs because they consist of symbolic structures that represent the world as shaped by human cognition and perception (Levesque and Lakemeyer, 2022). Such representations are semantically rich, featuring unambiguous vocabularies for entities and explicit relations grounded in domain-specific classification theories.
KGs have appeared as a new form of KOS, widely used for data extraction and LLM training. Recent studies employing LLM rely on structured database and RAG pipelines to store and distillate factual knowledge. KGs have emerged as a promising solution, offering an intuitive medium for representing and synthesizing knowledge that can be interpreted by both humans and machines (Xiao et al., 2025). Conversely, LLM-augmented KGs address challenges in generalizing large-scale textual information and handling incomplete or dynamic data (Pan et al., 2024). Synergizing LLMs and KGs iteratively compensates for general and domain-specific knowledge to achieve better performance for advanced tasks.
In the context of KO, classification theories and systems are socially and culturally dependent artifacts that directly reflect the contextual realities in which they are developed (Beghtol, 2009). Classification theories serve as intellectual foundations for the practice of KO. Regardless of their specific names or forms, all KOSs are based on a set of principles that guide KO practices. The conceptual basis for a KOS defines its purpose and warrant, shaping both its content and structure. In this sense, classification theories rationalize the need for and functions of KOSs while defining the semantic relations of concepts they represent.
Some scholars have further argued that KOS itself can be regarded as a classification theory. The case of KBs well illustrates this perspective. As modern instantiations of KOSs, KBs have been viewed by logicist AI as theories because they comprise collections of terms with associated axioms designed to constrain unintended interpretations and to enable the derivation of new information from ground facts. KBs are often crafted in ways that both reflect common-sense knowledge in a declarative form while taking advantage of the reasoning capabilities used by a given automated system (Smith and Welty, 2001).
In systematics, the groundbreaking Linnaean taxonomy functioned as a conceptual theory in eighteenth-century botany, as stated in Data-Centric Biology: A Philosophical Study:
“[Bio-ontologies] constitute a form of scientific theorizing that has the potential to affect the direction and practice of experimental biology in the long term, and yet they do not aim to introduce new language into biological discourse but rather attempt to gather and express consensus on what constitutes established knowledge” (Leonelli, 2016, pp. 121–122).
This interpretation of bio-ontologies derives from pragmatic epistemology and a structuralist ontological stance, which view theories as tools that scientists employ to define, refine, search for, and evaluate answers to empirical and theoretical questions (Balzer and Moulines, 1996). Treating ontologies as classificatory theories emphasizes their role in enabling the dissemination and retrieval of research materials while providing systematic scrutiny and interpretation of empirical evidence. The theoretical value of descriptive predicates lies not in their capacity to explain specific phenomena, but rather in their ability to classify entities that sustain both stability and dynamism in the evolving understandings of life science.
Recent studies have proposed a paradigm shift from KOSs to knowledge organization ecosystems (KOEs), highlighting the collaboration of interdependent components such as individuals, knowledge organization models and organizations, within platform-based socio-technical systems (STSs) (Bagchi, 2021). The KOE framework encompasses three interrelated dimensions: individuals (designers, domain experts, transdisciplinary ethics experts); technology (representational appropriateness of knowledge models, inter and intra domain knowledge artifacts’ interoperability); and organizations (technology consortiums) (Bagchi, 2021). Informed by foundational works in socio-technical ethics and the philosophical ramifications of KO, this framework establishes a conceptual bridge between STS and KO theories, thereby illuminating their influences on classical AI problems (e.g., knowledge-based AI systems).
Biology is among the most classification-oriented natural sciences, so devoted to classifications such that the special division, biological systematics, emerged to exclusively develop classifications under the framework of “theory + philosophy + conceptual history” (Pavlinov, 2021). Since the 18th century, systematics and its effort to create natural systems of living beings was considered a branch of natural philosophy and the product of human cognitive activity. There are two main components in the conceptual history of systematics—anagenetic and cladogenetic. Anagenetic means a sequential progression of taxonomic groups from less to more, while cladogenetic means fragmentation and multiplication (Pavlinov, 2021). Two components correspond to the unification and diversification trends separately in systematics, thus developing into an open information system that allows the third component—network component, to play its role in exchange and combination.
Influenced by important scholars and their philosophical contributions, e.g.,
Charles Darwin and Ernst Haeckel, systematics embarks on different periods of
theories. Classification systems were proposed suitable for direct interpretation
of systematics. The structuralist approach to classification systems (
where
The new systematics era encountered fallacies by confusing the mission of taxonomy with population genetics. Population genetics is experimental and conducted in real time focusing on genetic phenomena within and between populations of living species, while taxonomy is comparative, historical, and non-experimental, and emphasizes at and above the species level. Some scholars predict the decay of systematics because of advancements in technology. Systematics seems vulnerable against trendy, modern, and easily fundable sciences like genetics or molecular science. However, narrowly focusing on one group or genus through molecular measures and an over-obsession in “DNA barcoding” of new species’ identification overlook the importance of conceptual understanding of species and the intrinsic relations leading to knowledge of natural life. Studies argue for new knowledge organization and management tools designed for taxonomists to return their focus to species, their characters and relationships. It is imperative that technology should be secondary to the pursuit of knowledge in science (Wheeler, 2023).
Apart from technology, informatics has also dramatically altered taxonomic working practices and workflows. One notable change is the discoverability and reusability of taxonomic data, both through publication venues, e.g., European Journal of Taxonomy, Pensoft Journals, databases, repositories, and KBs, e.g., DNA Data Bank of Japan, the European Molecular Biology Laboratory, GenBank, the International Barcode of Life, MorphoBank, TraitBank (Secretariat of the Convention on Biological Diversity, 2021). The DELTA (DEscription Language for TAxonomy) was developed in 2000 as an information language system specifically for biological taxonomy, providing standardization for encoding taxonomic information on biological species (Coleman et al., 2010). Domain-specific NLP algorithms are also designed for biological information extraction including cellular processes, taxonomic names, and morphological characters (Thessen et al., 2012). Advanced AI techniques such as LLMs offer even more diverse and powerful support to biological classification and KO.
Within biomedical discovery, the language interpretation capabilities of LLMs can provide a framework for integrating and synthesizing the complex evidence space and tools, systematizing and lowering the barriers to access and reason over multiple structured databases, repositories and KOSs, enriching the background knowledge through specialized ontologies and serving as interfaces to external analytical tools (Wysocki et al., 2024).
Even before the emergence of LLMs, earlier ML models employed KOSs for computational analysis in biomedicine. Bio-ontologies were used as reliable structured KBs with semantically rich relationships between properties for gene-disease association discovery and identification. Advanced genome sequencing technologies accelerated the process of exploring genomic variations (Piñero et al., 2020) and genetic markers detection (Chang et al., 2024). ML models accelerate the discovery in searching for primary causes of genetic diseases by predicting functional similarities of genes with their semantic similarity within KOSs. The backbone of this approach is the schema-guided, computer-accessible bio-ontologies such as GO and Disease Ontology (DO). By calculating the semantic distance between entities in GO, models such as Support-Vector Machines (SVM) can predict the functional similarity between gene products, therefore predicting genes that potentially share the same association with disease-triggering proteins (Kulmanov et al., 2021; Liu and Qin, 2025).
Such design has been applied in COVID-19 research and proved to be powerful. One of the key challenges during the pandemic was the diversity of concepts and the breadth of background knowledge needed to understand the disease and develop diagnostic and therapeutic solutions. Even a single research article could span multiple biomedical subfields as well as physics, chemistry, engineering, computer science, and social science. To address this, Hope et al. (2020) introduced SciSight, an exploratory search system designed to help researchers navigate the rapidly evolving COVID-19 literature. SciSight combined faceted navigation, entity extraction, and network visualizations to surface relationships between research topics, biomedical entities, and scientific groups. While not as formalized as a logic-based ontology, SciSight functioned as a lightweight, dynamic KOS that supported sense-making and collaborative discovery at scale. These approaches prove how KOSs can act as integrative infrastructures in highly dynamic, interdisciplinary problem spaces, setting the stage for their role in AI-assisted scientific reasoning.
Hope et al. (2021) also developed a method for extracting mechanistic knowledge from the COVID-19 Open Research Dataset (CORD-19) by combining automated machine reading with targeted expert curation. Their framework aggregated heterogeneous mechanistic assertions into a unified KB. It covers entities such as viral proteins, host factors, pathways, and disease processes. This enabled queries to trace causal chains and uncover cross-disciplinary connections, exemplifying that, once formalized into a structured, computable representation, diverse evidence can facilitate systematic exploration of emerging literature and support rapid hypothesis generation in crisis-driven research. Adding another layer to this ecosystem, Wang et al. (2021) demonstrated the value of domain-specific pretraining for vertical search in biomedical literature. Using large-scale corpora to adapt transformer models to biomedical language, they achieved substantial gains in retrieving relevant documents from specialized repositories. While their work focused on retrieval rather than explicit ontology construction, the improved access to precise, contextually relevant literature directly supported downstream KOS applications particularly in crisis contexts such as COVID-19, where prompt identification of relevant evidence was essential.
Supporting knowledge discovery in AI-assisted science inquiry and reasoning is another key function of Bio-KOSs. While LLMs have already shown exceptional capabilities in medical knowledge competency (Nori et al., 2023), baseline models such as OpenAI’s GPT-4 (Achiam et al., 2023) tend to hallucinate or respond with overly general answers due to the lack of domain-specific external KBs. For LLMs to achieve scientific ideation and fully automated reasoning, the framework for deep learning or adaptive inference must provide high-quality scientific artifacts such as scholarly publications, metadata as signals of quality and interest, and structured data as support (Hope et al., 2023; Zhou et al., 2024).
Complementing LLMs with RAG mechanisms enables contextual control over relevant background knowledge and facts, thereby enhancing analytical precision (Wysocki et al., 2024). The theoretical foundation of such an approach mirrors human scientific reasoning: new ideas are generated by building upon existing ones. Scholars synthesize accumulated knowledge and external evidence to form insights, answers and research directions (Hope et al., 2023). Similarly, KBs supply AI models with continuously updated knowledge derived from clinical, experimental, and population-level genetic research. Integrating multiple KOSs significantly improves the competence of AI-driven scientific ideation. Consequently, KR in biology must support not only classification but also reasoning capabilities.
Neumann (2010) introduced the modal logic as a means for computational systems to encode varying degrees of epistemic commitment, such as “believes”, “knows”, and “possibly”. In the context of biological discovery, such logic-based representations enable reasoning across incomplete data and evolving hypotheses, which is an essential capacity in a dynamic field of science characterized by ambiguity and exception. Biologists frequently work with layered hypotheses and partially overlapping models that cannot be verified fully in isolation but can gain epistemic strength when integrated through structured representation and logical rules. Bio-ontologies, with their formal semantics and intrinsic ability to manage unverified concepts and relations, have the potential to support both deductive reasoning and exploratory inference across relational structures. These systems do more than define what is known; they establish the infrastructure for discovering what is knowable. As Neumann (2010) emphasized, recognizing where knowledge comes from is just as important as knowing it in biomedical research.
In AI-driven scientific research, KOSs do more than serve as simple reflections of scientific understanding. Their value in providing quality data and knowledge infrastructures in AI for science is increasingly recognized by AI model and application developers. Meanwhile, rapid advancements in AI are introducing new methods and perspectives into the design and use of KOSs. The classification and entification of concepts in KOSs, as in the case of GO, are essential not only for data management and knowledge representation, but also for critically shaping the directions of scientific inquiry they enable (or constrain). When embedded into LLM pipelines, structured networks of entities and relations can affect the performance of AI models in executing complex research tasks. Since the release of GPT-4, numerous NLP studies have incorporated KOSs as external KBs within RAG frameworks to enhance LLM performance on advanced scientific tasks, including project workflow construction (Ramirez-Medina et al., 2025) and hypothesis or idea generation (Abdel-Rehim et al., 2025; Guo et al., 2024).
The growth of computational tools and technology changed the classifications of KOSs, resulting in increased mapping and cross-walking between different types of KOSs and disciplines (Souza et al., 2012). Different KO scholars’ opinions varied on how to classify KOSs and what parameters/dimensions should be prioritized. Smith and Welty (2001) proposed a classification based on the capability for automated logical reasoning. Their framework arranged KOSs along a complexity spectrum: simple catalogs, glossaries, thesauri, taxonomies and ontologies with logical constraints. Bergman (2007) introduced a two-dimensional framework that correlates semantic clarity with investment in construction. His model positions simple glossaries and dictionaries at the lower end because they require minimal investment and offer limited semantic expressiveness. Formal ontologies at the opposite end deliver maximum semantic clarity but demand substantial investment in time, expertise, and computational resources. Zeng (2008) expanded this understanding by mapping KOSs across functional dimensions and semantic relationships, which resulted in four categories: term lists (glossaries, dictionaries), metadata-like models (directories, authority files), classification and categorization schemes (subject headings, taxonomies, categorization schemes), and relationship models (thesauri, semantic networks, ontologies). As the sophistication level increases from term lists to next category, their abilities to reduce ambiguity, control synonyms, establish relationships, and present properties also increase.
These KOS classification frameworks offer several insights. First, structural complexity and semantic richness directly correlate with advanced functionality in organizing and representing concepts. Second, the evolution from simple to complex KOSs parallels the transition from human-curated to machine-processable knowledge. Third, KOSs have evolved from monolithic to multi-dimensional representation of knowledge. Most studies in KO agree that the structural complexity and semantic richness lead to more advanced functionality in organizing and representing concepts and relations and fostering reasoning.
In the AI era, AI technology is used for KOSs to ingest digital “traces” of scientific thoughts and advances through scholarly publications, digital research artifacts, and even social media discussions, which in turn will help human researchers to discover and track these “traces” more easily and accurately without crossing the human cognitive threshold (Hope et al., 2023). A framework proposed by Almeida et al. (2011) considered the technological aspect of KOSs separately for humans and machines, that is, humans construct KOSs in natural language while machines use formal semantics language such as XML, RDF, OWL to encode KOSs. However, this dichotomic framework becomes problematic when describing contemporary KOSs that have integrated human curation with AI-supported processing.
The flaws and limitations of earlier approaches for analyzing KO research prompted us to develop a new one for this review: the Knowledge Organization Analysis Framework (KOAF). This framework includes three dimensions: level of functional sophistication, automation degree in system construction, and reasoning and inference capability. These dimensions align not only with what we have seen in the literature reviewed so far, but also with our goal of theoretically summarizing how current biological science research applies KOSs and how the deployment of KOSs has influenced the paradigmatic shift in AI-augmented biological science. Rather than coding the literature by topical themes alone, we used KOAF, a three-dimensional analytical framework, to map each article in a holistic view. The nuances and differences between types of KOSs cannot be captured by a simple, linear scale of “low” to “high”, although we do emphasize the progression of each level in each of the dimensions.
The first dimension is the level of functional sophistication of KOSs. We propose five distinct levels incrementally ranked from the most foundational to advanced (Table 1).
| Levels | Name | Description |
| 1 | Terminology Disambiguation | Systems at this foundational level address the most fundamental challenge in scientific communication and data integration—ambiguity. They provide controlled terms/vocabularies to ensure that a specific concept is represented consistently across different datasets, experimental mechanisms, and publications. These systems have minimal structural complexity, often consisting of simple term lists or basic hierarchies with a single, implicit relationship type. Their primary function is to build a common lexicon, thereby enabling semantic consistency in data annotation. |
| 2 | Data Structure Standardization | Building upon the foundation of consistent terminology, systems at this level adopt standardized schemas for data exchange and interoperability. They move beyond simple term lists to define a common set of data elements and their basic relationships, creating a shared “grammar” that allows effective exchanges of heterogeneous data sources. |
| 3 | Logic-Based Knowledge Modeling | Formal, logic-based KOSs represent a significant leap in structural and semantic complexity. Ontologies are the typical KOS at this level with expressive frameworks designed for computational knowledge representation. Classes (concepts) are organized in hierarchies, each of which has a set of properties (relations). Logical axioms (rules) are derived based on class relations. Logic-based KOSs model the domain semantics in a way that is not only processable but more importantly, interpretable by machines, thus enabling sophisticated computational tasks such as advanced query answering and information seeking. |
| 4 | Semantic Integration | Based on the logical blueprint provided by the previous level, systems at Level 4 integrate heterogeneous sources into cohesive networks or KGs. They primarily serve as central, query-able hubs of domain knowledge, overcoming the pervasive challenge of data fragmentation, especially in biomedical sciences. Some are equipped with tools for exploration, analysis, and application. |
| 5 | AI-Driven Knowledge Discovery | Systems at this highest level of functional sophistication represent a paradigm shift, moving beyond representation and integration of existing knowledge towards the active synthesis of new knowledge structures and generation of novel scientific insights. KOSs or frameworks at this level serve as the backbone for AI-powered knowledge discovery. AI models use KOSs as a scaffold to perform complex tasks such as extracting latent relationships from unstructured text, automating scientific reasoning, and generating novel hypotheses or research ideas. |
KOAF, Knowledge Organization Analysis Framework; KOSs, knowledge organization systems; KGs, knowledge graphs; AI, artificial intelligence.
The second dimension, degrees of automation in KOS construction, describes the extent to which computational techniques such as ML, NLP, or rule-based extraction are involved during the building, integration, and maintenance of KOSs. It captures the shift from human expert-driven curation toward data-fueled automation pipelines in constructing KOSs. To compare the extent to which human ability and computational methods contribute to these processes, we assigned a score from 1 to 4 for each paper as an indicator for the degree of automation. This dimension captures both the technical mechanisms employed and the degree of human intervention needed throughout the processes of knowledge acquisition, integration, and maintenance (Table 2).
| Degrees | Name | Description |
| 1 | Manual Curation | All knowledge elements, including terminologies, relationships, and mappings, are generated, validated, and updated exclusively by domain experts and/or trained personnel. Computational support is minimal and limited to basic data entry or indexing functions. While these systems maximize interpretability and conceptual rigor, their update cycles are slow, and scalability is constrained by expert availability. |
| 2 | Manual with Basic Automation | Human curation stays the primary driver, but the process is supported by limited computational tools such as keyword search, term suggestion, or basic text-mining functions. Automation aids candidate term identification but not in the execution of integration or validation, which is still expert led. |
| 3 | Hybrid Curation | Automated extraction, alignment, or reasoning methods—often employing NLP, ML, or rule-based pattern matching—are integrated with systematic expert oversight. Machines propose candidate knowledge additions or mappings, which are reviewed and confirmed by human curators before incorporation. This approach improves scalability while preserving quality control. |
| 4 | Fully Automated Systems | Level 4 denotes fully automated approaches to knowledge acquisition, integration, and maintenance. Most of the processing is performed by computational workflows with minimal human interventions. In both cases, the KOS curation tasks can run end-to-end without continuous human expert intervention. These are the most advanced KOSs so far and are suited for large-scale, rapid updates and integration with AI models. They typically employ ML, NLP, and KG technologies to extract and synthesize knowledge. The primary goal of these KOSs is to support new patterns identification and conceptual discovery across multiple domains. The highest scoring KOSs prove to be the most illustrative for paradigmatic shift from autonomously processing existing knowledge towards new biological science knowledge expansion. |
NLP, natural language processing; ML, machine learning.
While the second dimension assesses automation during the construction phase, the third dimension evaluates the epistemic capability of the completed KOS on how effectively it can perform reasoning, inference, and discovery beyond its explicitly encoded content. It moves from systems that merely retrieve or represent information toward those that can generate new association, infer relationships, or support hypothesis formulation. Conceptually, this dimension parallels long-standing challenges in AI research. We use five tiers to rank the extent to which a KOS works in a spectrum of knowledge organization systems’ tasks, ranging from static information storage and retrieval to structured query answering, logical reasoning, and scientific discovery (Table 3). Each tier bears all the properties from its lower tier.
| Tiers | Capability | Description |
| 1 | Simple lookup and retrieval | These systems return information verbatim from stored records without applying any computational inference. |
| 2 | Exploration along the hierarchical tree | Users of these systems can navigate up and down the structures and query the system based on kind-of or part-of relationships within taxonomies or ontologies. |
| 3 | Rule-based reasoning | KOSs at this tier are facilitated by predefined logical rules, constraints, or inference engines that can use over domain-specific ontologies or datasets, capable of deriving new facts or relationships within a domain. |
| 4 | Multi-domain reasoning | KOSs at this tier can integrate concepts and relationships across distinct ontologies or datasets, often applying complex logic rules or constraint-based reasoning to achieve interoperability and perform cross-domain inferences. |
| 5 | AI scientist | These systems not only integrate heterogeneous knowledge sources but also apply adaptive, ML-driven inference to generate novel predictions, suggest explanations, and guide further inquiry with minimal human intervention. As the highest tier of KOSs, these systems represent a paradigmatic shift from knowledge representation towards knowledge discovery. |
The results of thematic analysis on the target papers are shown in Fig. 1. The evolution of functional sophistication in bio-KOSs demonstrates a clear developmental trajectory, from foundational terminology systems to AI-driven knowledge discovery platforms. Earlier systems focused on establishing controlled vocabularies and data standards, examples including eVOC’s simple gene expression terminology (Kelso et al., 2003) and Darwin Core’s biodiversity data standards (Wieczorek et al., 2012). From literature in the middle period, we saw the emergence of sophisticated logic-based knowledge modeling through formal ontologies like GO, which introduced rich semantic relationships and cross-domain integration capabilities (Dumontier et al., 2014; Smith et al., 2007; The Gene Ontology Consortium et al., 2023; Walls et al., 2014).
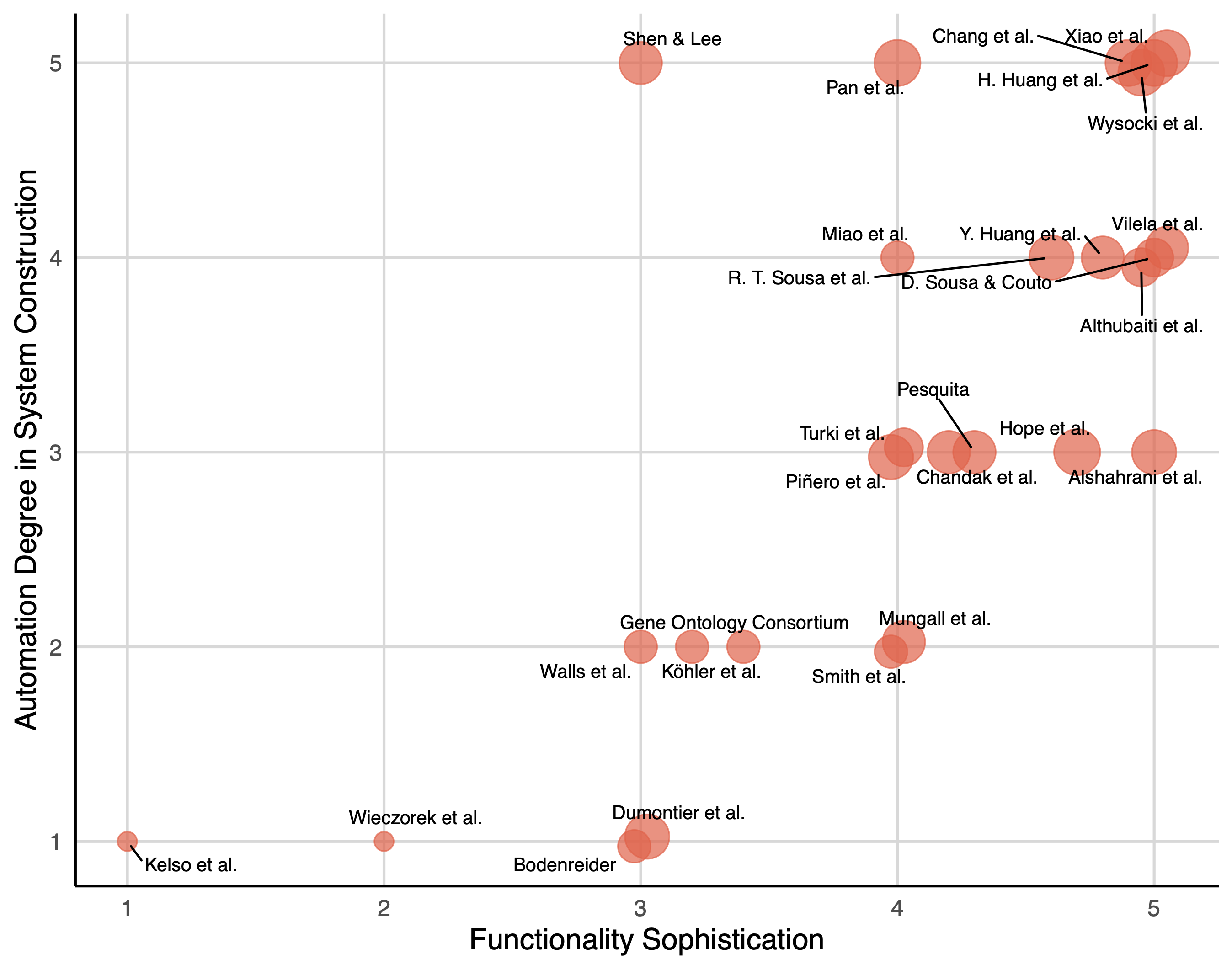 Fig. 1.
Fig. 1.
A two-dimensional mapping of target papers. The size of each point stands for the third dimension—reasoning and inference capability. See https://qiaoyiliu.github.io/KOAF/targetpaper3D.html.
The most dramatic transformation occurred from 2020 onward, with systems achieving Level 4-5 functional sophistication. For instance, TaxonGPT represents a paradigmatic example of this leap, demonstrating zero-shot taxonomic classification capabilities that generate human-readable rationales along with predictions (Huang et al., 2024). It adapts to taxonomic novelties without retraining, effectively mimicking expert taxonomic reasoning while surpassing human scalability limitations. This progression reflects the KO field’s evolution from static knowledge representation to dynamic, generative systems capable of supporting scientific reasoning and discovery.
The shift in functionality is mirrored in the automation degree in KOSs construction, as more sophisticated systems apply a hybrid approach that combined automated extraction with systematic expert validation (e.g., DisGeNET) (Piñero et al., 2020). Earlier systems were exclusively manually curated, where domain experts hand-crafted terminologies and kept ontological structures with minimal computational support. Maturation of LLMs and advanced NLP techniques were equipped to systems from 2021, signaling full automation capabilities as appears in NEKO and gene-disease discovery platform (Chang et al., 2024; Xiao et al., 2025).
Finally, the reasoning dimension in Fig. 1 shows the most dramatic change, progressing from simple lookup functions to sophisticated hypothesis generation capabilities. More fundamental KOSs merely offer basic retrieval and hierarchical navigation within predefined taxonomic or ontological structures, while Entity-relation structures and logic-based architecture allow rule-based reasoning within single domains. These KOSs began to perform logical inference to derive new facts from existing knowledge. Cross-mapping between multiple domains helped cross-ontology inference and semantic interoperability. AI-augmented systems have become more powerful in biological science in supporting hypothesis-generation and discovery.
However, the relationships among the three dimensions are not strictly linear. Some highly sophisticated systems maintain hybrid curation approaches for quality assurance (Chandak et al., 2023; Turki et al., 2024), reflecting the continued importance of expert validation in critical biomedical applications. The convergence toward higher scores across all dimensions in recent papers (2024–2025) indicates the emergence of a new class of bio-KOS appears (Chang et al., 2024; Wysocki et al., 2024; Xiao et al., 2025). These systems fundamentally transform the relationship between KO and biological scientific discovery, turning KOSs from passive knowledge tools into active players in knowledge generation.
The three dimensions in our KOAF situate KOSs both as descriptive tools and epistemic infrastructures that co-evolve with AI technologies. These dimensions may be considered as actionable criteria for evaluating and designing future KOSs for scientific knowledge. For KOS developers, this framework provides guidance on balancing human curation with computational automation to improve reliability and scalability. For biological researchers, it stresses how embedding KOSs in AI pipelines can enhance data interoperability, reproducibility, and innovation. We expect that an integration of KOS principles and AI systems will improve model transparency, reduce hallucination, and ensure that systems remain accountable to established scientific knowledge.
This thematic analysis was intended as an exploratory study to find phenomena and trends in bio-KOSs in the context of AI, rather than a comprehensive, exhaustive study. As such, it has some inherent limitations. The corpus of 27 target papers, while representative, cannot capture the full scope of literature across bioinformatics and AI-augmented KO. The reliance on English-language publications and accessible datasets may also introduce selection bias. Moreover, the focus on conceptual and empirical exemplars, rather than exhaustive quantitative evaluation, limits the generalizability of findings. Finally, the rapidly evolving nature of AI and KO technologies means that any classification framework risks obsolescence as new models and paradigms appear. These limitations, however, reflect the exploratory nature of this thematic review and open opportunities for further refinement and validation of the proposed framework.
This study reveals that KO in AI-augmented biological research has evolved far beyond traditional classification paradigms. New theoretical frameworks are needed to explain how scientific knowledge is structured, represented, and discovered. Our analysis shows that contemporary biological sciences operate within a complex ecosystem of KOSs, ranging from basic terminology disambiguation to fully automated, AI-driven discovery platforms. The proposed KOAF offers a unique lens for understanding how bio-KOSs vary across the three dimensions of functional sophistication, automation degree in system construction, and inference and reasoning capability. Bio-KOSs have shifted from manually curated controlled vocabularies to hybrid and fully automated ontological systems that not only standardize biological data but also contribute directly to hypothesis generation and discovery. This evolution underscores the growing interdependence between human ability and ML in shaping the new landscape of KO in the AI era.
The most advanced bio-KOS applications, particularly those achieving Level 5 functional sophistication, manifest emergent properties that transcend their original design purposes. These systems support complex scientific reasoning, cross-domain integration, and novel hypothesis generation, effectively functioning as collaborative agents in scientific discovery rather than passive curators of knowledge. From these patterns, we expect a trajectory toward fully automated, reasoning-capable KOSs. As AI models continue to advance in generating scientific insights, the boundary between human and machine contributions to KO becomes increasingly blurred. This convergence calls for new epistemic frameworks that can accommodate hybrid, multidisciplinary, and collaborative modes of scientific inquiry.
Instead of dramatic theoretical and epistemological leaps as envisioned in Kuhn’s paradigm theory, current developments reflect a gradual integration of interdisciplinary tools and methodologies, helped by sophisticated KO infrastructures that enable semantic interoperability across domains. The integration of AI technologies with traditional bio-ontologies is a fundamental shift in organizing and generating scientific knowledge. Traditional biological classification systems, such as Linnaean taxonomy and the GO, are no longer mere inventories of existing knowledge. Rather, KOSs, bio-KOSs, play a constitutive role in knowledge creation through their embedding in AI pipelines. This transformation positions KOSs as theoretical frameworks, shaping research directions and constraining inquiry paths through their structural and classificatory designs. KOSs’ gradual convergence and interoperability across disciplines are reshaping the boundaries between classification, representation, and knowledge production.
As representational tools, bio-ontologies provide structured vocabularies and semantic relationships essential for data integration and retrieval. As theoretical frameworks, they decide which questions can be asked, which connections can be made, and how biological knowledge is conceptualized. Their integration into AI pipelines shows how these systems actively guide research trajectories. The findings of this thematic review extend beyond the life sciences discipline in that all data-intensive sciences are confronting similar challenges of scale, interdisciplinary, and computational integration.
Taken together, these findings suggest that bio-KOSs are evolving into active epistemic agents within AI-supported science. They not only organize and represent knowledge but increasingly shape the process of scientific discovery. As AI models become more deeply embedded in biological research, future work should investigate the long-term broader implications of automated knowledge production, including how insights are validated and how evolving AI-KOS interactions reshape scientific reasoning and research practice.
Building on these implications, several directions emerge for advancing research on AI-integrated KOSs. Future research should focus on empirically evaluating the KOAF through systematic comparisons of across scientific domains. Quantitative indicators could be developed to track how systems progress along the three dimensions, enabling longitudinal analyses of automation and reasoning capabilities. Further work is needed to establish governance and evaluation frameworks to ensure transparency, accountability, and reproducibility in AI-driven knowledge discovery. Interdisciplinary collaboration among KO scholars, computer scientists, and domain experts will be essential to translate these conceptual insights into practical design principles for future scientific infrastructures.
All data, plots, and code reported in this paper are shared via GitHub: https://qiaoyiliu.github.io/KOAF/targetpaper3D.html.
QL and JQ conceptualized the research questions and developed the KOAF analytical framework. QL identified and analyzed the target papers, conducted the comprehensive literature review, performed the thematic analysis across the three dimensions, and drafted the original manuscript. JQ provided supervision throughout the research process, guided the theoretical framework development, and contributed to refining the research direction and scope. JQ critically reviewed the analytical framework and provided substantial revisions to strengthen the theoretical contributions. Both authors collaborated on interpreting the results and their implications for knowledge organization theory. Both authors contributed to editorial revisions of the manuscript. Both authors read and approved the final manuscript. Both authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The authors thank the anonymous reviewers for their constructive feedback on earlier versions of this manuscript.
This research received no external funding.
The authors declare no conflict of interest.
See Table 4.
| Title | Author(s) | Year |
| eVOC: A Controlled Vocabulary for Unifying Gene Expression Data | Kelso et al. | 2003 |
| The Unified Medical Language System (UMLS): integrating biomedical terminology | Bodenreider | 2004 |
| The OBO Foundry: coordinated evolution of ontologies to support biomedical data integration | Smith et al. | 2007 |
| Integrating phenotype ontologies across multiple species | Mungall et al. | 2010 |
| Darwin Core: An Evolving Community-Developed Biodiversity Data Standard | Wieczorek et al. | 2012 |
| The Semanticscience Integrated Ontology (SIO) for biomedical research and knowledge discovery | Dumontier et al. | 2014 |
| Semantics in Support of Biodiversity Knowledge Discovery: An Introduction to the Biological Collections Ontology and Related Ontologies | Walls et al. | 2014 |
| Knowledge Discovery from Biomedical Ontologies in Cross Domains | Shen and Lee | 2016 |
| Neuro-symbolic representation learning on biological knowledge graphs | Alshahrani et al. | 2017 |
| The DisGeNET knowledge platform for disease genomics: 2019 update | Piñero et al. | 2020 |
| Combining lexical and context features for automatic ontology extension | Althubaiti et al. | 2020 |
| BiOnt: Deep Learning using Multiple Biomedical Ontologies for Relation Extraction | Sousa and Couto | 2020 |
| Evolving knowledge graph similarity for supervised learning in complex biomedical domains | Sousa et al. | 2020 |
| Towards Semantic Integration for Explainable Artificial Intelligence in the Biomedical Domain: | Pesquita | 2021 |
| Extracting a Knowledge Base of Mechanisms from COVID-19 Papers | Hope et al. | 2021 |
| The Human Phenotype Ontology in 2021 | Köhler et al. | 2021 |
| Biomedical knowledge graph embeddings for personalized medicine: Predicting disease-gene associations | Vilela et al. | 2023 |
| The Gene Ontology knowledgebase in 2023 | The Gene Ontology Consortium et al. | 2023 |
| Building a knowledge graph to enable precision medicine | Chandak et al. | 2023 |
| Gene-associated Disease Discovery Powered by Large Language Models | Chang et al. | 2024 |
| A framework for integrating biomedical knowledge in Wikidata with open biological and biomedical ontologies and MeSH keywords | Turki et al. | 2024 |
| An LLM-based Knowledge Synthesis and Scientific Reasoning Framework for Biomedical Discovery | Wysocki et al. | 2024 |
| UniEntrezDB: Large-scale Gene Ontology Annotation Dataset and Evaluation Benchmarks with Unified Entrez Gene Identifiers | Miao et al. | 2024 |
| Unifying Large Language Models and Knowledge Graphs: A Roadmap | Pan et al. | 2024 |
| TaxonGPT: Taxonomic Classification Using Generative Artificial Intelligence | Huang et al. | 2024 |
| Building a literature knowledge base towards transparent biomedical AI | Huang et al. | 2025a |
| Network for knowledge Organization (NEKO): An AI knowledge mining workflow for synthetic biology research | Xiao et al. | 2025 |
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.

