1 Department of STEM Education and Professional Studies, Old Dominion University, Norfolk, VA 23529, USA
Abstract
As artificial intelligence (AI) becomes increasingly embedded in healthcare applications, concerns have emerged around the trustworthiness, interpretability, and context-awareness of these systems. Knowledge Organization Systems (KOS) hold considerable potential to address these challenges by supporting semantic standardization, explainability, and domain alignment. This study presents a bibliometric analysis of scholarly publications referencing both AI and healthcare concepts to examine how KOS are positioned within this evolving discourse. The findings indicate that while early literature frequently and explicitly referenced KOS—such as ontologies, controlled vocabularies, and classification systems—their visibility has declined relative to newer paradigms such as machine learning and large language models. Nevertheless, KOS-related terms remain conceptually linked to key healthcare domains, including diagnostics, therapeutics, and administration, albeit occupying a more peripheral role in the broader AI research landscape. These terms most often co-occur with topics such as natural language processing, information extraction, and the semantic enrichment of unstructured clinical data. The findings show the continued relevance of KOS in AI-healthcare discourse while highlighting the need for more deliberate alignment between KOS and emerging AI methodologies. Future work should explore frameworks that bridge conceptual presence with technical deployment. KOS may thereby offer critical contributions to the development of transparent, context-sensitive, and ethically grounded AI systems in healthcare.
Keywords
- knowledge organization systems
- artificial intelligence
- healthcare
- bibliometric analysis
- health informatics
- semantic interoperability
The healthcare sector stands at a critical juncture where the convergence of artificial intelligence (AI) and knowledge organization systems (KOS) is no longer optional but imperative. Over the last decade, the healthcare information technology landscape has undergone transformative shifts, driven in part by federal policies such as the Health Information for Economic and Clinical Health (HITECH) Act (2009) and the 21st Century Cures Act (2016). These legislative policies incentivized the adoption of digital systems, catalyzing a transition from paper-based records to integrated digital frameworks like electronic health records (EHRs), clinical decision support systems (CDSS), and remote monitoring devices (Aradhya et al, 2023). These innovations have revolutionized clinical workflows and also unlocked opportunities to enhance patient outcomes, streamline data interoperability, and accelerate research at scale.
Building on this foundation, AI has emerged as a transformative force in
healthcare. Broadly defined, AI refers to the field of computer science focused
on developing systems that attempt to simulate aspects of human intelligence
(Secinaro et al, 2021). Applied within health informatics, AI
technologies—including machine learning (ML) and natural language processing
(NLP)—leverage vast datasets to identify patterns imperceptible to human
analysis, enabling real-time, evidence-based decision-making (Al Kuwaiti et al, 2023; Amiri, 2024; Secinaro et al, 2021). Applications range from
high-accuracy medical imaging (e.g., detecting tumors with
Despite the promising contributions of AI in healthcare, its adoption is not without challenges. AI’s potential to revolutionize healthcare may be stymied by unprecedented volumes of unstructured and semi-structured data, which may have poor data quality standards and lack semantic consistency. Ethical concerns, data privacy issues, and the need for transparent and explainable AI systems remain significant barriers (Antoniadi et al, 2021; Saraswat et al, 2022). Furthermore, the efficacy of AI-enabled systems hinges on their ability to interpret and contextualize healthcare data. However, fragmented data interoperability, algorithmic bias, and “hallucinations” in large language models (LLMS) may undermine their real-world efficacy. Such challenges underscore a pressing need for KOS, to ensure AI accuracy, trustworthiness, and scalability.
KOS are schemes designed to support the structured organization of information by modeling the underlying semantic structure of a domain and acting as knowledge representation resources (Hjørland, 2016; Mazzochi, 2018; Souza et al, 2012). KOS play a foundational role in enabling knowledge representation and interoperability as they have been embedded into diverse clinical systems. Furthermore, those designed for clinical contexts have had their use mandated by federal policies.
In healthcare settings, KOS such as the Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT) from the International Health Terminology Standards Development Organization, International Classification of Diseases (ICD) developed and maintained by the World Health Organization, Logical Observation Identifiers Names and Codes (LOINC) from LOINC.org, and Medical Subject Headings (MeSH) provided by the USA National Library of Medicine (NLM) establish explicit semantic relationships and represent both relationships and properties of concepts in knowledge models (Zeng et al, 2020). Bronnert et al. (2012) categorize these KOS based on their responsibilities in various applications as administrative (for administration functions like billing, reimbursement, information classification, or secondary data aggregation); clinical (for electronic clinical data exchange and aggregation); reference (for representation of concepts); and interface (to support interoperability) terminologies.
KOS organize and represent information in healthcare through standardized terminologies, metadata-like models, classification and categorization schemes, taxonomies, ontologies, semantic networks, and other knowledge structures. They capture the complexity and semantics of medical knowledge, including diseases and diagnoses, treatment and medications, outcomes and research, and decision support (Bodenreider et al, 2018; Fung and Bodenreider, 2019; Fung and Bodenreider, 2023). These knowledge structures provide explicit definitions that prevent semantic conflicts from occurring and enable semantic interoperability. Semantic conflict often occurs within data and information communication processes, when names carry different meanings at different times and can add confusion. One such example of semantic conflict is outlined by Zeng et al. (2020) when they describe the issues around the naming of the 2009 Influenza A virus subtype H1N1 outbreak and later the Coronavirus Disease 2019 (COVID-19) outbreak in 2020.
Semantic interoperability in healthcare systems refers to the ability of different computer systems to exchange information while preserving the intended meaning and interpretation of data (Arvanitis, 2014). Beyond simple data exchange, this ensures that the receiving system understands and processes information as intended by the transmitting system. This requires technical standards such as Health Level 7 and KOS such as those previously mentioned that define relationships between medical concepts.
These KOS are particularly crucial for decision support systems as they enable rule-based efficient reasoning and reduce redundancy in knowledge bases. For example, Ahmadian et al. (2011) demonstrate how hierarchical relationships within terminologies like SNOMED CT allow a single clinical rule to apply across multiple subtypes of a disease category (e.g., cardiovascular conditions), simplifying rule authoring and enhancing system scalability. Such integration enables decision support systems to recognize conditions like hypertension, arrhythmias, and valve disorders as related, improving alignment with clinical protocols and guidelines, simplifying maintenance, and improving the consistency and reliability of AI-driven decision support. Despite their potential, the roles or contributions of KOS in AI-driven healthcare remain underexplored, and when present, are inconsistently applied and often technologically fragmented. Challenges such as inconsistencies in granularity and semantic misalignment, and limited interoperability between terminologies like ICD-10 and SNOMED CT, hinder the effective use of structured knowledge in AI-based analytics and applications.
Gaining insight into how KOS are positioned within AI-driven healthcare research necessitates a comprehensive examination of existing literature. Systematic reviews and bibliometric analyses are commonly used to examine both the current state of a field and the evolution of its subtopics. While recent studies focusing on AI in healthcare have primarily focused on broad trends, with researchers investigating NLP methodologies in clinical settings (Shaikh et al, 2023; Wang et al, 2020), they often overlook the critical role of KOS in AI implementation. Additionally, although some studies present promising frameworks for KOS-AI integration, such as Greenberg et al.’s (2021) work on Helping Interdisciplinary Vocabulary Engineering (HIVE)-enabled KOS and Jia et al’s (2019) knowledge graph-based diagnostic systems, no comprehensive bibliometric analysis has examined how these technologies might converge in healthcare applications.
The rapid advancement of both KOS and AI technologies highlights a significant gap, especially considering applications like ontology-driven clinical decision support and semantic search frameworks. Hong and Zeng (2022) summarize efforts in the literature to automatically assign ICD codes to health-related documents, describing evolving approaches such as rule-based systems, traditional ML, and more recent neural network-based methods. These approaches have led to improved information retrieval and enhanced capabilities in identifying chronic pain and exploring causality.
However, challenges remain regarding data acquisition and effective alignment of KOS. For example, Medical Literature Analysis and Retrieval System Online (MEDLINE) indexing, which provides valuable metadata for biomedical literature, transitioned to fully automated indexing in 2022 using the Medical Text Indexer-NeXt Generation (MTIX) algorithm, with human review for quality assurance (National Library of Medicine, 2023). However, the risk remains that nuances and contextual information might be missed, especially since only titles and abstracts are included, potentially leading to misclassification or incomplete indexing. Other foundational analyses utilizing KOS resources, such as the Unified Medical Language System (UMLS) (Jing, 2021; Kim et al, 2020), also exist. However, these studies tend to concentrate on individual components rather than their integration and interoperability within AI systems. They also fail to provide a comprehensive overview of the field, including methodologies, opportunities, and remaining challenges. As AI systems become increasingly incorporated into healthcare, it is crucial to address this gap by exploring the diverse ways KOS are being implemented.
By employing bibliometric analysis, the study aims to explore the occurrence of terminology related to KOS and AI, as well as the co-occurrence of health science research topics in scholarly literature published over the past decade. Bibliometric analysis enables inferences about prevailing themes, conceptual structures, and emerging research fronts by analyzing the frequency and co-location of terms within a defined corpus. This approach provides an overall understanding of the evolving landscape, revealing patterns of scholarly attention, disciplinary convergence, and opportunities for innovation. While this study explores literature at the intersection of KOS, AI, and healthcare, it does not evaluate the technical integration of KOS into AI systems, investigating instead co-occurrence and thematic clustering of relevant terms in scholarly publications to reveal patterns of attention and areas of conceptual convergence across these domains. The analysis is intended to provide valuable insights for researchers, practitioners, and policymakers working to advance this critical field. The paper addresses the following research questions:
(1) What thematic areas and AI methodologies emerge most prominently from literature addressing KOS-related terms in healthcare applications (2014–2024)?
(2) How has scholarly attention to knowledge organization systems evolved within AI-driven healthcare literature over the past decade?
(3) What gaps, challenges, or opportunities are revealed in the scholarly literature on knowledge organizations systems within AI-driven healthcare research?
Bibliometric analysis offers a robust methodological framework for systematically examining the evolving conceptual positioning of KOS into AI-driven healthcare. To map the intellectual structure of this interdisciplinary domain, the study adopts a multi-method bibliometric approach, including co-occurrence network analysis, walktrap clustering, and multidimensional scaling (MDS). Co-occurrence network analysis was selected as the primary analytical method due to its proven effectiveness in visualizing conceptual relationships among terms within scientific literature (Kirtania, 2023; Singh and Singh, 2018; van Eck et al, 2010). Specifically, author keyword co-occurrence networks enable the identification of core research themes and their interconnections without imposing predetermined classifications on the natural structure of scholarly discourse.
The Walktrap clustering algorithm was used for community detection, based on its demonstrated effectiveness in identifying cohesive and functionally meaningful modules (thematic clusters or research areas) within bibliometric networks (Brusco et al, 2024; Jamison et al, 2021; Pons and Latapy, 2006). This hierarchical clustering algorithm identifies communities or clusters within a network based on the idea that nodes sharing frequent connections are more likely to belong to the same conceptual cluster (Hosseini et al, 2024). Walktrap’s random walk approach offers notable advantages over other community detection algorithms by detecting natural thematic clusters that reflect underlying scholarly discourse patterns without requiring predefined parameters.
To complement these methods, multidimensional scaling (MDS) was also used. MDS is a dimensionality reduction technique that provides a spatial representation of conceptual distances among key terms, i.e., representing semantic proximity. It effectively reveals the underlying intellectual structure and facilitates interpretation of multiple analytical dimensions (Hout et al, 2013; van Eck et al, 2010). Together, these approaches clarify the key research themes and patterns in the conceptual alignment of KOS and AI within healthcare literature.
The Bibliometrix software (Version 5.1.1, K-Synth team, Naples, Italy) was selected to conduct this analysis, as it comprehensively implements these techniques within a unified analytical framework (Aria and Cuccurullo, 2017). Biblioshiny is the interactive web interface for the Bibliometrix R-package (https://www.bibliometrix.org/home/), a well-established tool supporting science mapping through integrated data collection, analysis, and visualization capabilities (Aria and Cuccurullo, 2017). Although Biblioshiny and comparable tools like VOSviewer (Version 1.6.20, (van Eck and Waltman, 2023); Leiden, The Netherlands) and CiteSpace (Version 6.4.R2, (Chen, 2025), Philadelphia, PA, USA) have advanced science mapping capabilities (Kim et al, 2020; Biranvand et al, 2020), their specific application to examining KOS-related terms within AI-driven healthcare remains notably underexplored, underscoring the value and novelty of this methodological approach.
Finally, to identify and quantify the prevalence of KOS and ML methods across the corpus, a Python script was used to perform the following tasks:
(1) Comprehensive synonym matching: Terms are mapped to all common variations, abbreviations, and full names. For instance, the International Classification of Diseases was mapped to include ICD, ICD-9, ICD-10, ICD-11, and its full name.
(2) Text field coverage: The analysis searched across four key bibliographic fields for each article -Abstract, Title, and both keyword fields to ensure that relevant mentions in any field would be captured.
(3) Deduplication: To avoid overcounting, each term was counted only once per article, regardless of how many times it appeared in various forms.
(4) Word Boundary Detection: Using regular expressions for short abbreviations to prevent false positives from partial string matches, for example, distinguishing between the word “go” and the acronym Gene Ontology (GO).
The search query targeted scholarly articles indexed in major bibliographic databases focusing on medicine and health sciences. The query was developed through (1) generating initial terms, (2) identifying relevant concepts and associated terms via a preliminary literature review of keywords, abstracts, and titles, and (3) refining terms using subject headings from the databases. Terms were selected to represent three conceptual domains:
• KOS: Included general KOS terminology (e.g., ontology, taxonomy) and healthcare specific implementations (e.g., SNOMED, ICD). Using general terms ensured inclusion of broader research applying knowledge organization principles beyond explicitly named standardized terminologies.
• AI: Captured terms representing both general AI approaches and specific methodologies ensuring broad representation of computational intelligence applications.
• Healthcare context: This dimension included both general health terms and specific application domains. Explicit terms (e.g., electronic health records and precision medicine) ensured targeted relevance, while broader terms provided coverage of diverse healthcare applications.
To obtain the corpus, a search was conducted using the following query:
(“Knowledge Organization Systems” OR “KOS” OR “Knowledge Graph*” OR “Ontolog*” OR “Taxonom*” OR “Thesaurus” OR “Classification System*” OR “Controlled Vocabular*” OR “Semantic Network” OR “Semantic Reasoning” OR “Data Fusion” OR “Knowledge Representation” OR “Semantic Integration” OR “ICD*” OR “SNOMED*” OR “UMLS” OR “clinical terminolog*”) AND (“Artificial Intelligence” OR “AI” OR “Machine Learning” OR “ML” OR “Deep Learning” OR “Large Language Model*” OR “LLM*” OR “Neural Network*” OR “Natural Language Processing” OR “NLP” OR “xai” OR “Explainable Artificial Intelligence”) AND (“Healthcare” OR “Health Care” OR “health” OR “Medicine” OR “Biomedical” OR “Clinical” OR “Electronic Health Record*” OR “EHR” OR “Precision Medicine”)
The literature search was conducted in two major academic databases, Web of Science (WoS) and Scopus. To refine the corpus, filters were applied to include only records published between 2014 and 2024, limited to peer-reviewed journal articles written in English. For the WoS database, the search was confined to three citation indexes, the Science Citation Index Expanded (Sci-Expanded), the Social Science Citation Index (SSCI), and the Emerging Sources Citation Index (ESCI). After removing duplicate records (n = 1604), a total of 5923 unique articles remained. A final eligibility screening excluded records that were not journal articles, were retracted, or failed to meet predefined eligibility criteria. This resulted in 5836 articles retained for bibliometric analysis. The following Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) flow diagram (Fig. 1) outlines the inclusion extraction process.
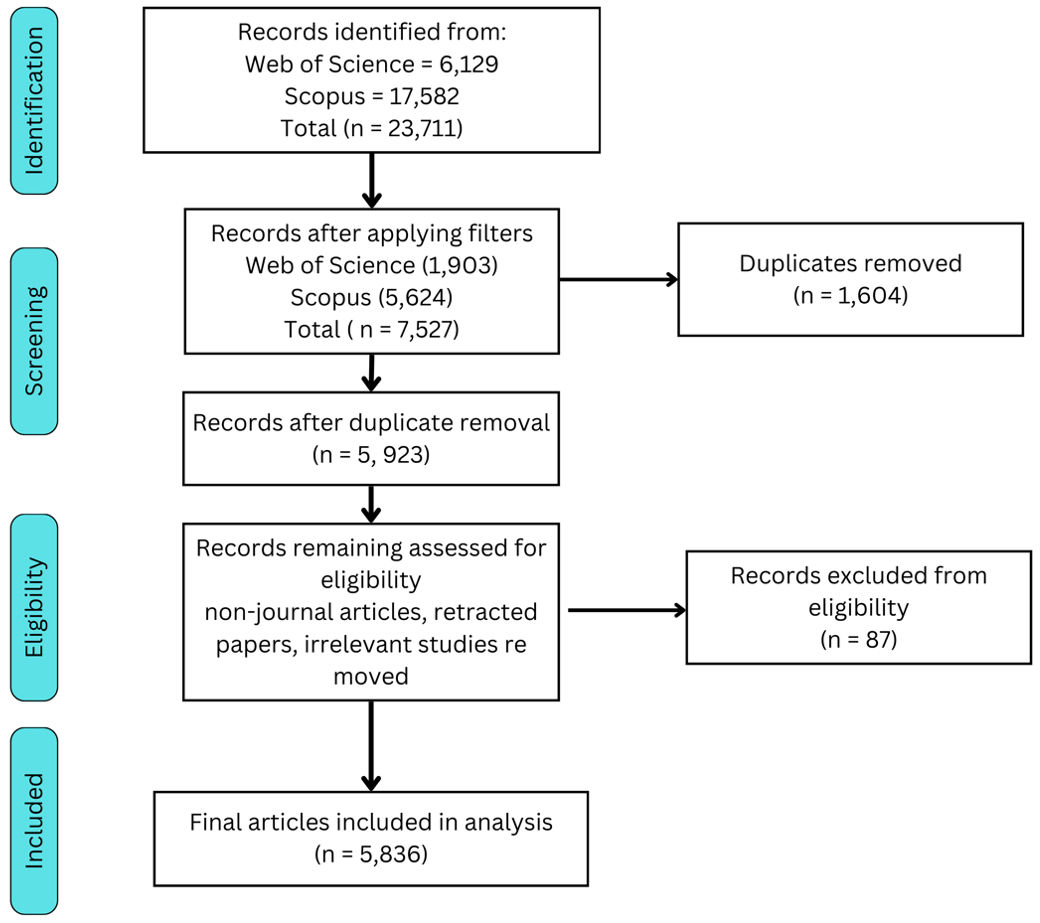 Fig. 1.
Fig. 1.
PRISMA flow diagram to show the inclusion-extraction process. Diagram created by author and adapted from Page et al. (2021). PRISMA, Preferred Reporting Items for Systematic Reviews and Meta-Analyses.
The results of the analysis are presented in the following sections beginning with an assessment of the characteristics of the dataset. Subsequently, the conceptual structure of words in the corpus of documents was assessed.
The bibliometric analysis covered a total of 5836 documents published between 2014 and 2024. These documents were sourced from 1631 distinct journals, reflecting the multidisciplinary nature of research at the intersection of KOS and AI in healthcare. The annual growth rate of publications in the domain was 27.72%, indicating a rapidly increasing interest and research activity in the application of AI and KOS in health care contexts. This growth suggests an expanding body of literature, with new advancements and applications emerging consistently over the last decade.
A total of 26,054 authors contributed to the body of literature, which signifies a large research community contributing to the field. However, with only 70 documents being single-authored and an average number of co-authors per document of 7.77, there is a clear emphasis on collaboration amongst authors, likely driven by the need for interdisciplinary expertise. There is also a modest level of international co-authorship at 8.1%, which suggests research is being conducted across borders, strengthening resources and expertise.
The corpus had an average document age of 3.6 years, indicating that the literature is relatively recent and aligned with current technological developments in AI and healthcare. On average, each document received 14.14 citations, suggesting a healthy level of academic visibility and engagement within the field. The documents reflect a rich diversity of topics with 13,320 unique author keywords across a broad range of AI methodologies (e.g., ML, deep learning, NLP) discussed further in Section 5.3.1 and healthcare applications (e.g., electronic health records, clinical decision support, precision medicine).
Further analysis revealed that of the total articles processed, 5508 (94.4%) mentioned at least one KOS, with 3675 (63%) articles mentioning general KOS types and 4786 mentioning specific KOS, indicating a strong methodological focus on knowledge organization principles within the corpus. Ontologies dominated the KOS landscape with 1911 articles (32.7%), reflecting their central role in the KOS-AI landscape, and classification schemes were mentioned in 805 articles (13.8%). Taxonomies appeared in 546 articles (9.4%), while Knowledge graphs were mentioned in 258 articles (4.4%). Other general types of KOS showed lower but still significant frequencies: Terminology (202 articles, 4.0%), Controlled Vocabulary (123 articles, 2.1%), Thesaurus (49 articles, 0.8%), and Semantic Network (28 articles, 0.5%). The relatively low frequency of traditional library science terms like thesaurus and controlled vocabulary suggests a preference for specific terminology within the bioinformatics field.
For specific KOS, the ICD emerged as the most frequently mentioned specific KOS, appearing in 1809 articles (31.0%). UMLS was mentioned in 195 articles (3.3%), and SNOMED in 145 articles (2.4%). Other medical/clinical KOS mentioned include Current Procedural Terminology (CPT), MeSH, Diagnostic and Statistical Manual of Mental Disorder (DSM), Health Level Seven International (HL7), RxNorm, Fast Healthcare Interoperability Resources (FHIR), LOINC, Healthcare Common Procedure Coding System (HCPCS), and National Drug Code (NDC). There were also mentions of Biological/Scientific KOS, including the GO in 1421 articles (24.3%). This reflects GO’s established role as the de facto standard for gene function annotation across model organisms and its integration into numerous bioinformatics workflows. The Kyoto Encyclopedia of Genes and Genomes (KEGG) was mentioned in 614 articles (10.5%), which shows its use for research activity in metabolic pathways, systems biology, and drug discovery. The Disease Ontology is mentioned in 247 articles (4.2%). Other KOS mentioned include the Protein Ontology, Human Phenotype Ontology, Cell Ontology, Reactome, Anatomy Ontology, PubChem, UniProt, Chemical Entities of Biological Interest (ChEBI), and the National Center for Biotechnology Information (NCBI) Taxonomy.
As illustrated in Fig. 2, the annual scientific production of publications showed a relatively modest and steady growth with minor fluctuations between 2014 and 2017, likely representing the field’s early exploration of foundational concepts and potential applications. A noticeable uptick occurred beginning in 2018, which may coincide with heightened policy interest and the forthcoming implementation of the 21st Century Cures ACT, enacted in 2016 and taking effect in 2020. This legislation emphasized interoperability, standardized data exchange, and expanded access to health information, potentially creating a regulatory environment that may have driven more research into the use of KOS and AI applications in healthcare.
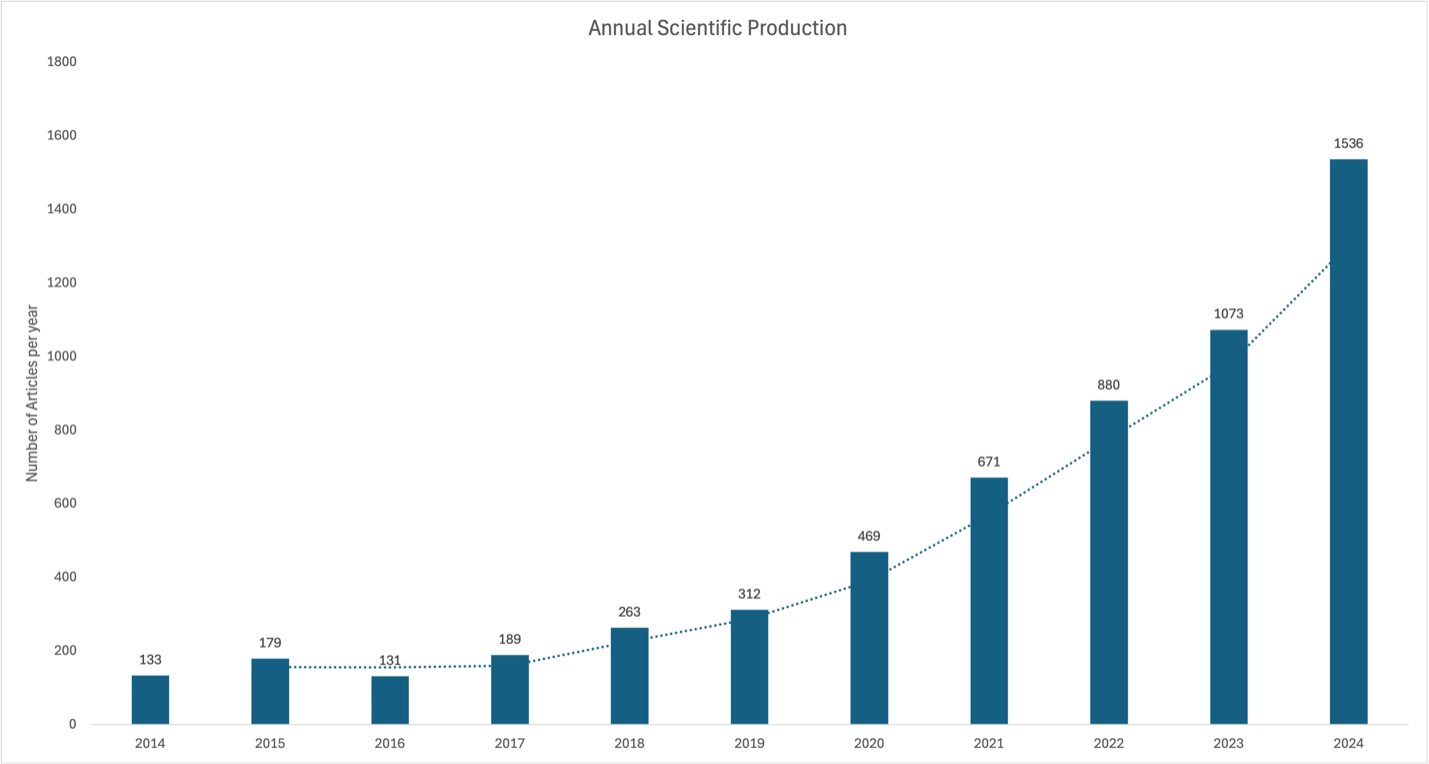 Fig. 2.
Fig. 2.
Annual scientific production.
The significant acceleration between 2020 and 2023 likely reflects a confluence of transformative developments. Most notably, the COVID-19 pandemic generated exponential growth in data volumes and highlighted an urgent need for AI-driven decision support, diagnostics, and information management. Another possibility is that the maturation of AI technologies, including the rapid advancement and diffusion of LLMs, gave rise to new research pathways in areas such as clinical text mining and automated knowledge representation.
Publication trends over time show that a select group of journals have emerged as key publication sources for research at the intersection of AI and KOS in healthcare. Fig. 3 shows that the Journal of Biomedical Informatics exhibits the most consistent and sustained growth, with 20 publications on these topics in 2014 and 228 in 2024. This underscores its central role in advancing research on data management, knowledge representation, and computational frameworks in health informatics.
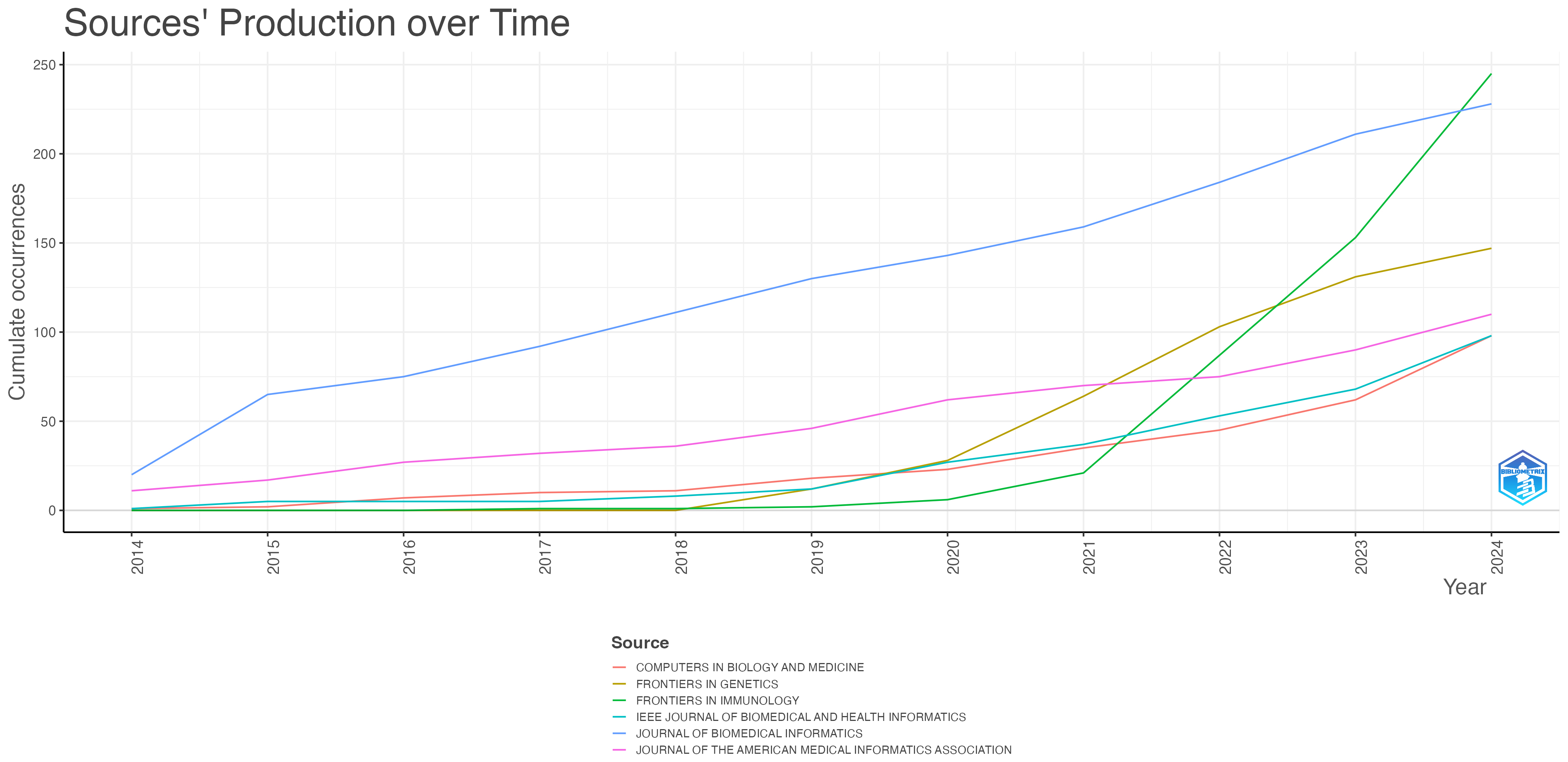 Fig. 3.
Fig. 3.
Cumulative source production over time.
In contrast, Frontiers in Immunology, which had minimal publications in the early years, demonstrates explosive growth beginning in 2020, surpassing the Journal of Biomedical Informatics by 2024 with over 245 publications. The surge (2021–2024) as observed in the Journal of Biomedical Informatics and Frontiers in Immunology, suggests a growing interest in potentially applying KOS-AI frameworks to immunological research driven by the COVID-19 pandemic, which intensified research focused on vaccine development, immune response modeling, and epidemiological surveillance.
Journal of the American Medical Informatics Association (JAMIA) shows a steady upward trajectory, reflecting its consistent role in supporting research on medical informatics. Similarly, Frontiers in Genetics also shows marked growth after 2018, accelerating after 2020, suggesting a rising emphasis on genomics, bio-ontologies, and precision medicine as emerging domains for KOS-AI integration. The Institute of Electrical and Electronics Engineers (IEEE) Journal of Biomedical and Health Informatics and Computers in Biology and Medicine both display a steady but slower rate of growth, reflecting their focus on technical aspects of healthcare, particularly in algorithmic development, and computational methods.
Collectively, these trends indicate a broadening of disciplinary engagement with KOS-AI research, extending beyond traditional biomedical informatics into immunology, genetics, and computational biology. The growing diversity of contributing journals points to converging interdisciplinary approaches where KOS-AI applications are increasingly used to tackle challenges in specialized medical fields and the increasing complexity of healthcare data.
Fig. 4 presents a horizontal bar chart depicting the top 20 most frequently occurring author-assigned keywords, derived from a broader set of 13,320 unique terms. The keyword “machine learning” dominates the dataset with 1180 occurrences (22%), reinforcing its role as the primary AI methodology employed in healthcare research. Closely following are “natural language processing” (460, 9%), “deep learning” (340, 6%), and “artificial intelligence” (328, 6%), each reflecting a growing emphasis on both foundational AI techniques and the emergence of language-based models in clinical and biomedical applications.
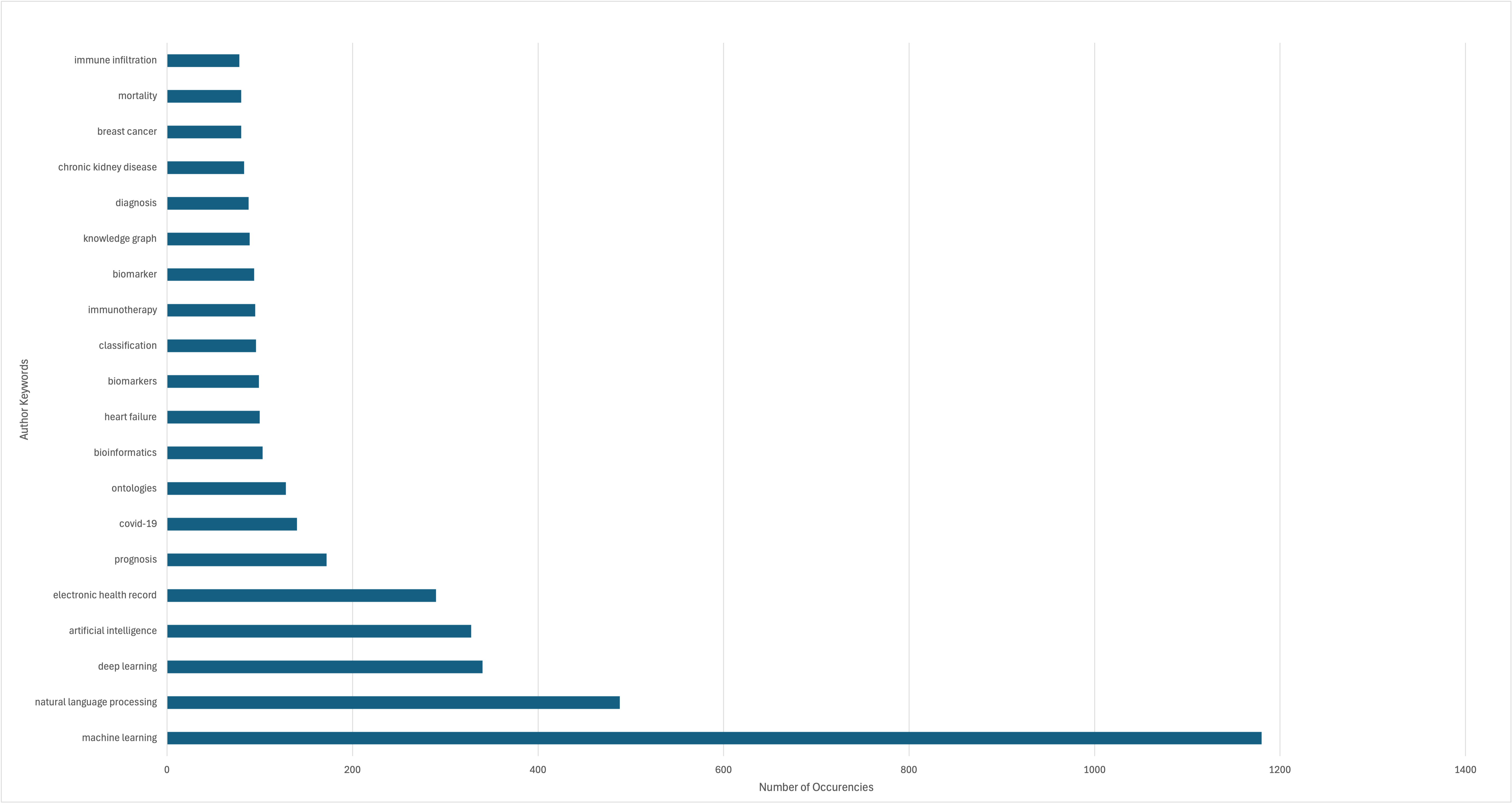 Fig. 4.
Fig. 4.
Top 10 occurring author keywords.
The keyword distribution also highlights increased attention to knowledge-centric terms such as “electronic health records”, “ontologies”, “semantic web”, and “knowledge graphs”. This may indicate sustained interest in leveraging structured domain knowledge and standards in AI workflows. Other important terms reflected include “information extraction”, “text mining”, “data fusion”, and “random forest”, perhaps demonstrating the methodological diversity of the field and an alignment towards data preprocessing, model training, and interpretability.
Fig. 5 presents a trend analysis of author-assigned keywords showing frequency and temporal evolution across the corpus. Each line represents the variability and interquartile range of a keyword’s occurrence, from the first quartile (Q1) to the third quartile (Q3), to indicate when topics peaked or emerged. The line extends from the first quartile (25th percentile) to the third quartile (75th percentile) of the term’s frequency, with wider lines suggesting a broader range of occurrences in the literature. In contrast, the circles represent the median frequency of a term in a given year. The larger the circle, the higher the median frequency of that term in that year, which indicates the relative importance of a term in the research literature during that period.
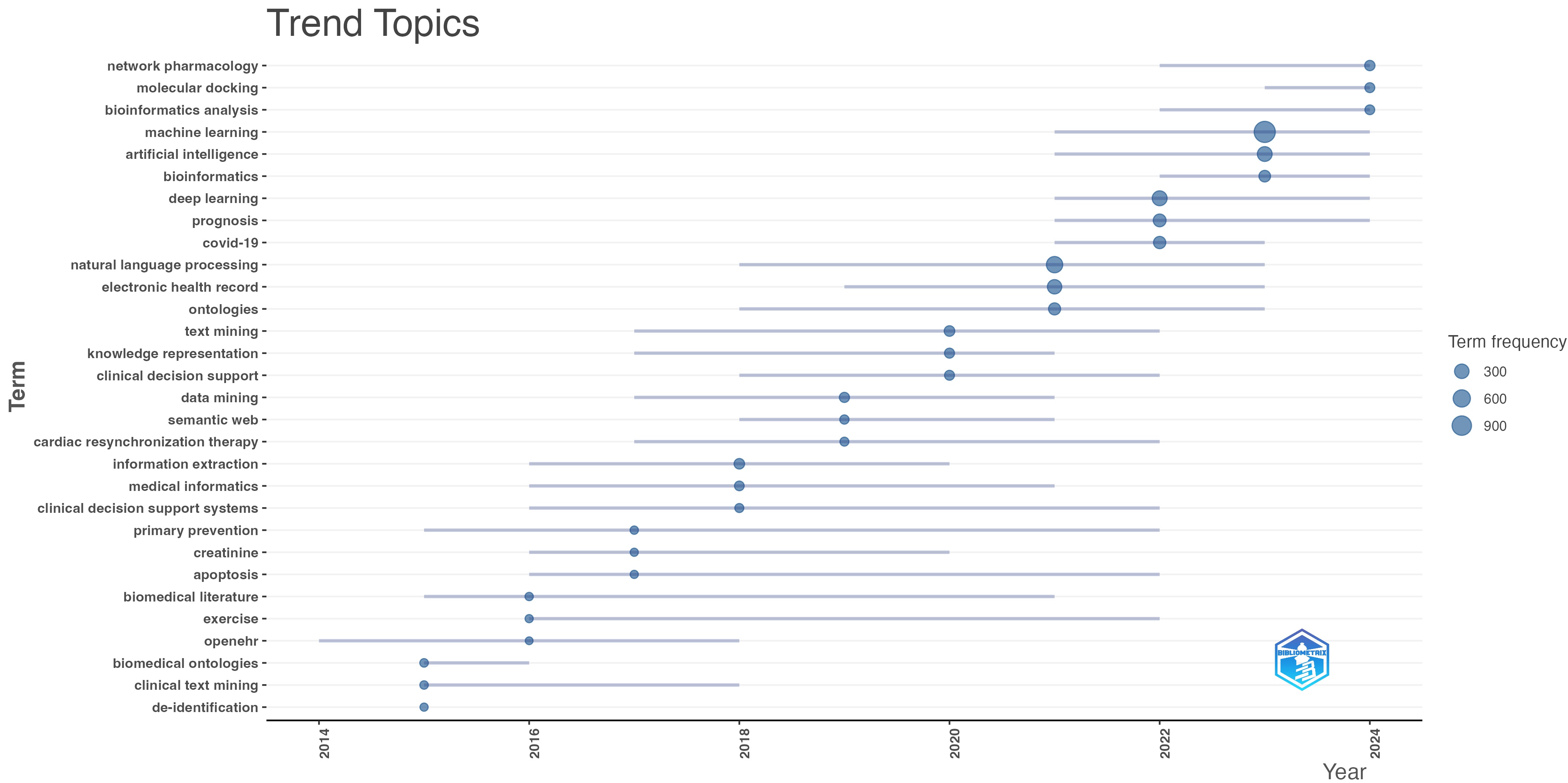 Fig. 5.
Fig. 5.
Topical trends from author keywords.
The visualization reveals that keywords such as “machine learning”, “deep learning”, and “artificial intelligence” show bursts of interest concentrated in recent years, especially from 2022 to 2024, showing a recent uptick in research activity in core AI domains. COVID-19 appears as a dominant emerging topic, with strong frequency and tight concentration in 2020–2022, reflecting the surge in pandemic-related health data analysis and AI-enabled public health surveillance. Keywords like “clinical decision support”, “medical informatics”, and “electronic health record” appear consistently and span much of the timeline with moderately sized bubbles. These represent foundational and enduring topics over which new AI techniques have been deployed.
Notably, KOS-related terms such as “knowledge representation”, “ontologies”, and “semantic web” are present across multiple years, with bubble sizes indicating sustained relevance and ongoing efforts for integration into AI models. In particular, the recent activity around the semantic web reflects the rising role of semantic technologies in making AI models more interpretable, standardized, and context-aware, particularly in biomedical data integration. Additional terms such as “text mining”, “data mining”, “case-based reasoning”, and “word-sense disambiguation” also appear prominently, reinforcing the emphasis on extracting structured insights from unstructured health data sources such as electronic health records, clinical notes, and biomedical literature perhaps for use in context-aware AI models that leverage structured knowledge frameworks.
This section describes the main themes and trends discussed in the literature. To facilitate interpretation of thematic clusters identified in the analysis, it might be helpful to understand the broad application contexts in which KOS-related and AI-driven approaches co-occur in healthcare literature, typically spanning four domains: (1) Research applications, focusing on knowledge discovery and hypothesis generation; (2) diagnostic applications, which support clinical decision making in identifying conditions; (3) therapeutic applications which guide treatment selection and personalization; and (4) administrative applications which enhance healthcare operations and information management. The relations among concepts are interpreted through this framework to identify the most important topics covered, their evolution over time, and any subfields.
The co-occurrence network provides insights into the conceptual structure of the domain by visualizing associative relationships between author keywords, revealing both the dominant themes and their interconnections in KOS-AI healthcare research. This approach is particularly valuable for identifying the semantic bridges between KOS and AI methodologies in healthcare applications. The keyword co-occurrence used the walktrap algorithm to provide a more nuanced understanding of how research themes naturally group together based on semantic proximity rather than predetermined categories. The network reveals distinct thematic communities, indicating how closely related topics cluster together based on their frequent co-occurrence. The normalization by association ensures that stronger relationships are emphasized.
In Fig. 6, it is shown that the largest and most central node is “ML” reinforcing its dominant theme in the AI-driven healthcare research. Its many direct connections to various subfields indicate that ML is applied across multiple domains including clinical decision support, genomics and bioinformatics, medical imaging, epidemiology and chronic disease research. The thick edges between ML and other nodes indicate high co-occurrence frequency.
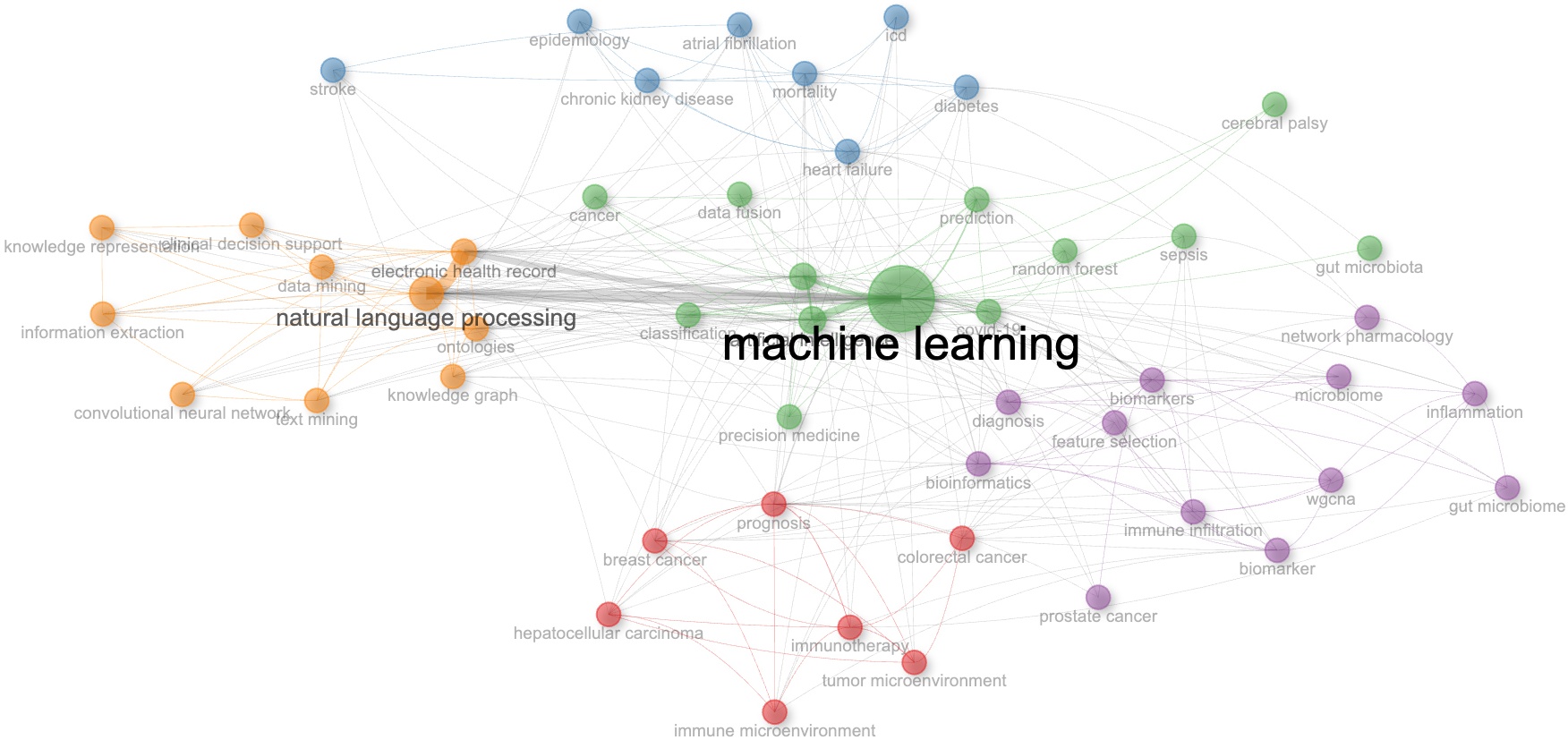 Fig. 6.
Fig. 6.
Co-occurrence network of author keywords.
The walktrap algorithm was used to identify thematic communities, each represented by a distinct color, providing insights into how research topics are conceptualized and grouped. The green cluster, for instance, encompasses keywords related to structured knowledge representation and AI-based text processing. These keywords include NLP, knowledge representation, information extraction, SNOMED CT, ontology, knowledge graphs, clinical decision support, and electronic health records. The presence of these keywords suggests a focus on the active use of KOS for knowledge extraction, semantic reasoning, and semantic interoperability in healthcare AI applications.
The red cluster focuses on keywords such as biomarkers, microbiome, gene expression, proteomics, prediction models, and artificial neural networks. This represents AI applications in genomics and precision medicine (primarily research and diagnostic applications), with the connection to ML, indicating that AI is widely used for biomarker discovery, disease classification, and molecular data analysis. The presence of omics terms suggests that AI is also being used for multi-omics integrations, which supports personalized medicine (a therapeutic application focused on tailoring treatments to individual patient characteristics). The orange cluster focuses on population health, chronic disease risk prediction and healthcare outcomes analysis, representing a multi-domain application that spans research, diagnostic and administrative functions. It involves pattern identification in chronic disease progression. The strong link to ML may imply that AI plays a role in predictive modeling for both individual patient interventions (therapeutic applications) and population-level healthcare planning (administrative applications).
The blue clusters represent the use of AI to support drug discovery and pharmaceutical research, primarily functioning as research applications focused on development of various therapies. The purple cluster indicates AI’s increasing role in oncology decision support, spanning both diagnostic and therapeutic applications. For diagnostics, this may include cancer detection, and prognostic modeling, involving medical image analysis and pathology interpretation. The therapeutic dimension involves treatment selection, response prediction, and personalized cancer care protocols.
A factorial analysis of keywords was conducted to explore the conceptual structure of KOS-AI healthcare research. This analysis employs multidimensional scaling (MDS) to reduce high-dimensional co-occurrence data into a two-dimensional space, where the spatial proximity of terms reflects their semantic and topic-relatedness across the literature corpus. The map presented in Fig. 7 includes only high-frequency keywords, those appearing in at least 407 publications (minDegree = 407). This threshold ensures that the analysis reflects the most influential terms in the field. Generic clinical descriptors (e.g., male, adult, cohort study) were excluded to focus the analysis on intellectually significant terms. Three distinct thematic clusters represented in red, green, and blue, were identified, each representing a major domain of scholarly activity within the corpus.
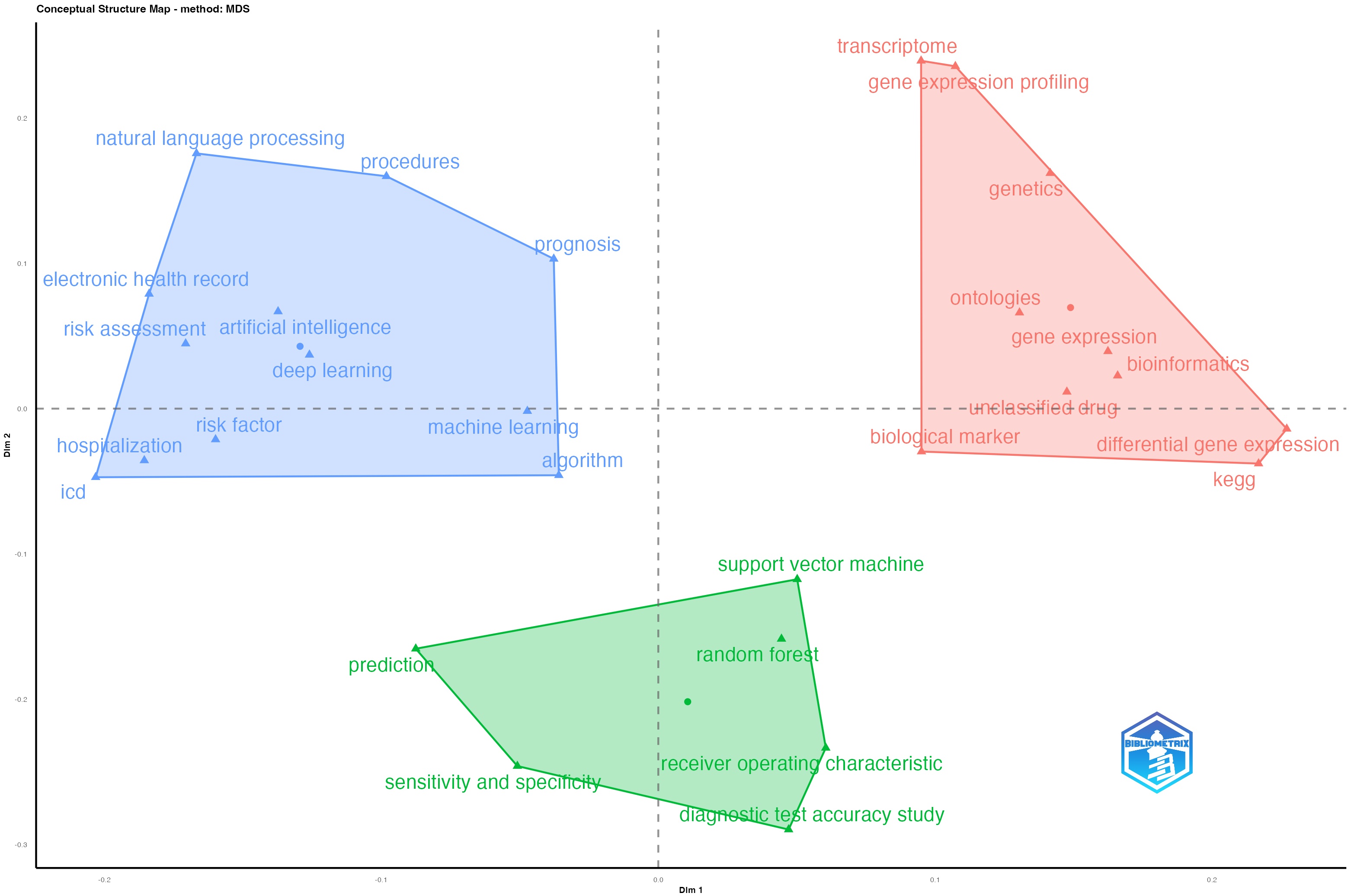 Fig. 7.
Fig. 7.
Factorial Analysis of Keywords using multidimensional scaling.
The blue cluster represents research focused on data-driven clinical decision-making and risk modeling, particularly related to diagnostic and administrative tasks. Frequently occurring terms include “natural language processing”, “electronic health record”, “risk factor”, “prognosis”, and “ICD”. This cluster reflects the conceptual role of KOS in tasks such as disease classification, predictive diagnostics, and population health monitoring, where terminologies like ICD and SNOMED CT support semantic standardization and interoperability. The presence of both NLP and EHR suggests a strong emphasis on semantic enrichment of narrative clinical data to support AI applications in clinical documentation, decision support, and billing workflows.
The green cluster reflects work on AI and model evaluation and performance assessments, which is important to clinical applicability and trustworthiness, especially for diagnostic accuracy. It includes terms like “support vector machines”, “random forest”, “receiver operating characteristic”, “sensitivity and specificity”, and “diagnostic test accuracy study”. This cluster involves focus on model development, validation and performance benchmarking, activities that are critical to ensuring reliable diagnostic tools, particularly those involving image analysis, lab interpretation, and risk stratification.
The red cluster focuses on therapeutic and precision medicine applications. This cluster primarily encompasses research applications aimed at genomics, molecular biology, drug discovery and development. The presence of key terms such as “gene expression”, “genetics”, “biological marker”, and “KEGG”, a pathway database (Kanehisa and Goto, 2000), points to AI methods being used for biomarker identification, pharmacogenomics, and pathway analysis. The presence of “ontologies” indicates the reliance on structured knowledge in AI-driven bioinformatics and biomarker discovery and facilitates semantic integration across disparate sources in therapeutic research. These findings highlight the importance of semantic integration, and structured knowledge representation for target identification, therapy optimization, and personalized medicine.
To explore research themes across the corpus, a bivariate thematic mapping (Fig. 8) was performed based on keyword co-occurrence analysis. This technique uses two key metrics: centrality and density. Centrality measures how connected a theme is to other themes in the network. It indicates the theme’s overall importance to the field and plays a bridging role between subfields. Density reflects the internal cohesion of a theme. High density signals a well-developed and conceptually mature area with strong internal connectivity among keywords, while low density suggests an emerging, fragmented, or underdeveloped theme. The positions of the nodes in the thematic map corresponds with its strategic role within the broader research landscape, while the size of the nodes represents the volume of research activity associated with that theme. The thematic map is divided into four quadrants, each representing a distinct type of research theme.
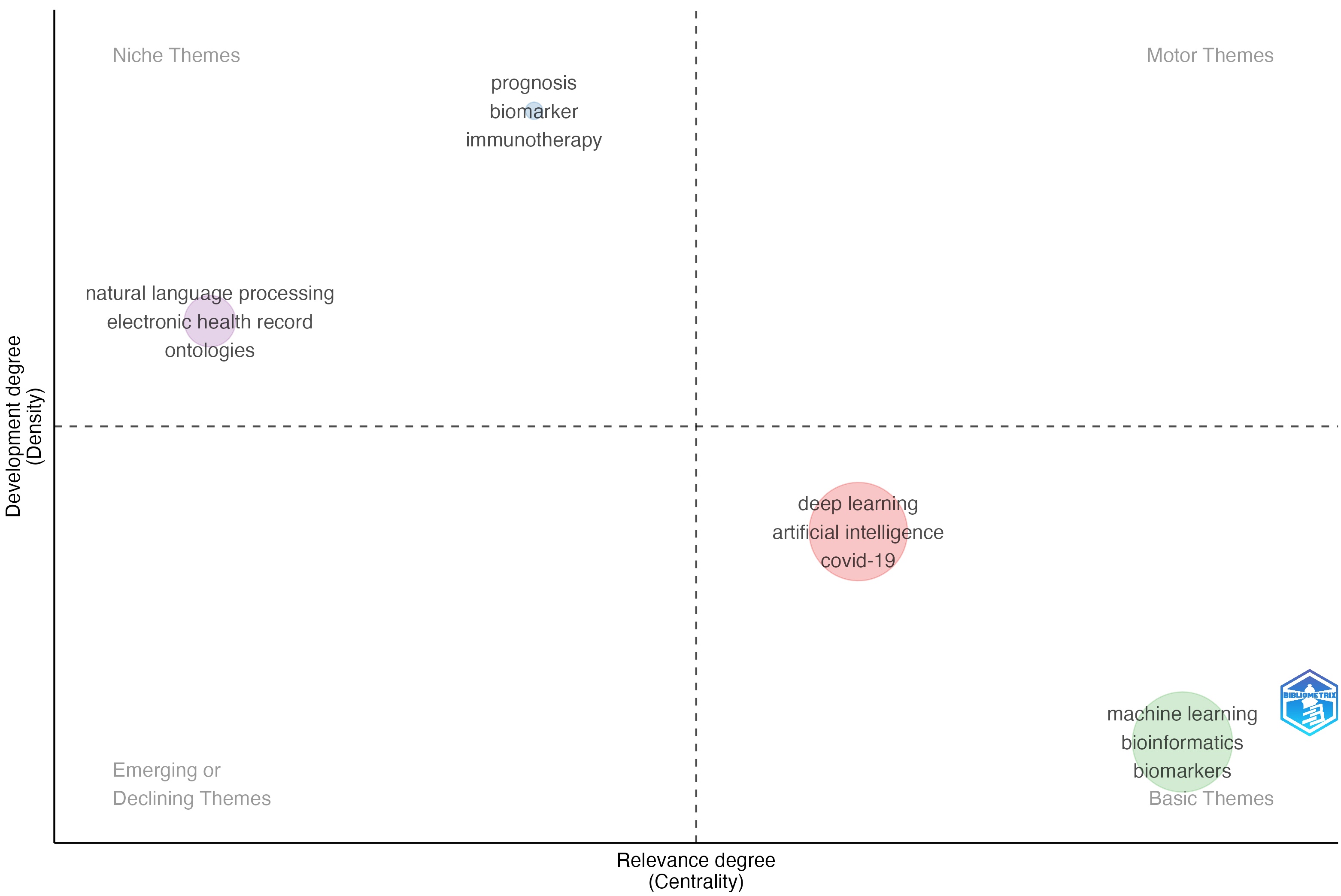 Fig. 8.
Fig. 8.
Thematic bivariate map of keywords.
Motor themes are highly central and well-developed, referring to methods that are influential and anchoring the field. Although no cluster is located centrally in this quadrant, the deep learning cluster is moving towards this area. This cluster (deep learning, artificial intelligence, COVID-19) indicates research in this area is dominant and established. The proximity of deep learning to artificial intelligence and deep learning reflects the surge in research during the pandemic era (2020–2024), particularly focused on applying deep learning to medical imaging, diagnostics, and predictive modeling.
Basic themes are central but less developed, foundational topics that are still evolving conceptually. The machine learning cluster appears in this lower-right quadrant, suggesting it is a core but still-developing area within the research landscape. While highly relevant, its internal cohesion is lower than deep learning, perhaps because of its broad applicability across diverse subfields. Keywords like bioinformatics and biomarkers indicate that machine learning serves foundational roles in omics data analysis, molecular diagnostics, and personalized medicine.
Niche themes, in the upper left quadrant, are well developed but peripheral, representing specialized domains that are internally cohesive but less connected to the broader research structure. Terms such as “natural language processing”, “ontologies”, and “electronic health records” are present here. This indicates that there is significant work on semantic processing, knowledge representation, and clinical text mining. Still, these themes remain less embedded in the dominant thematic core of KOS-AI research network. Also located in this quadrant is a cluster focused on prognostic modeling and immune response prediction. The co-occurrence of the terms “prognosis”, “biomarker”, and “immunotherapy” indicates work on determining therapeutic outcomes based on genomic and immunologic markers.
The placement of these clusters is additionally supported by article-level data. For example, articles related to deep learning span the years 2020–2024 and appear in journals focused on medical informatics and heart failure. Articles related to machine learning span the years 2022–2024 and appear in journals focused on medical informatics and cellular/molecular biology. Articles related to NLP appear in journals that deal with emergency care, and the study of toxins. Finally, the articles for prognosis appear in genetics and immunology articles. PageRank scores reinforce the influence of these articles, where higher PageRank scores suggest a more influential article. For example, Almog et al.’s (2020) article on deep learning with electronic health records has the highest PageRank (0.318), underscoring deep learning’s centrality within the citation network. Overall, these results point to a need for advancing the interoperability and interpretability of AI models through tighter integration with KOS, such as ontologies, terminologies, and classification schemes, which may enhance predictive accuracy and clinical impact and lead to field-defining innovations.
To trace how research priorities and conceptual focus areas have shifted, a longitudinal thematic evolution map was generated (Fig. 9). This visualization spans five distinct periods (2014–2016, 2017–2018, 2019–2020, 2021–2022, and 2023–2024), capturing the progression and thematic transitions of high-frequency terms within KOS-AI healthcare research over time. Each rectangle represents a thematic cluster, while connecting lines represent conceptual continuity, mergers or divergence. Through analysis of the emergence, persistence, and transformation of key themes, the map shows how early emphasis on semantic tools and text analytics, evolved into a mature, AI-centered ecosystem characterized by predictive modeling, precision medicine, and structured clinical data integration.
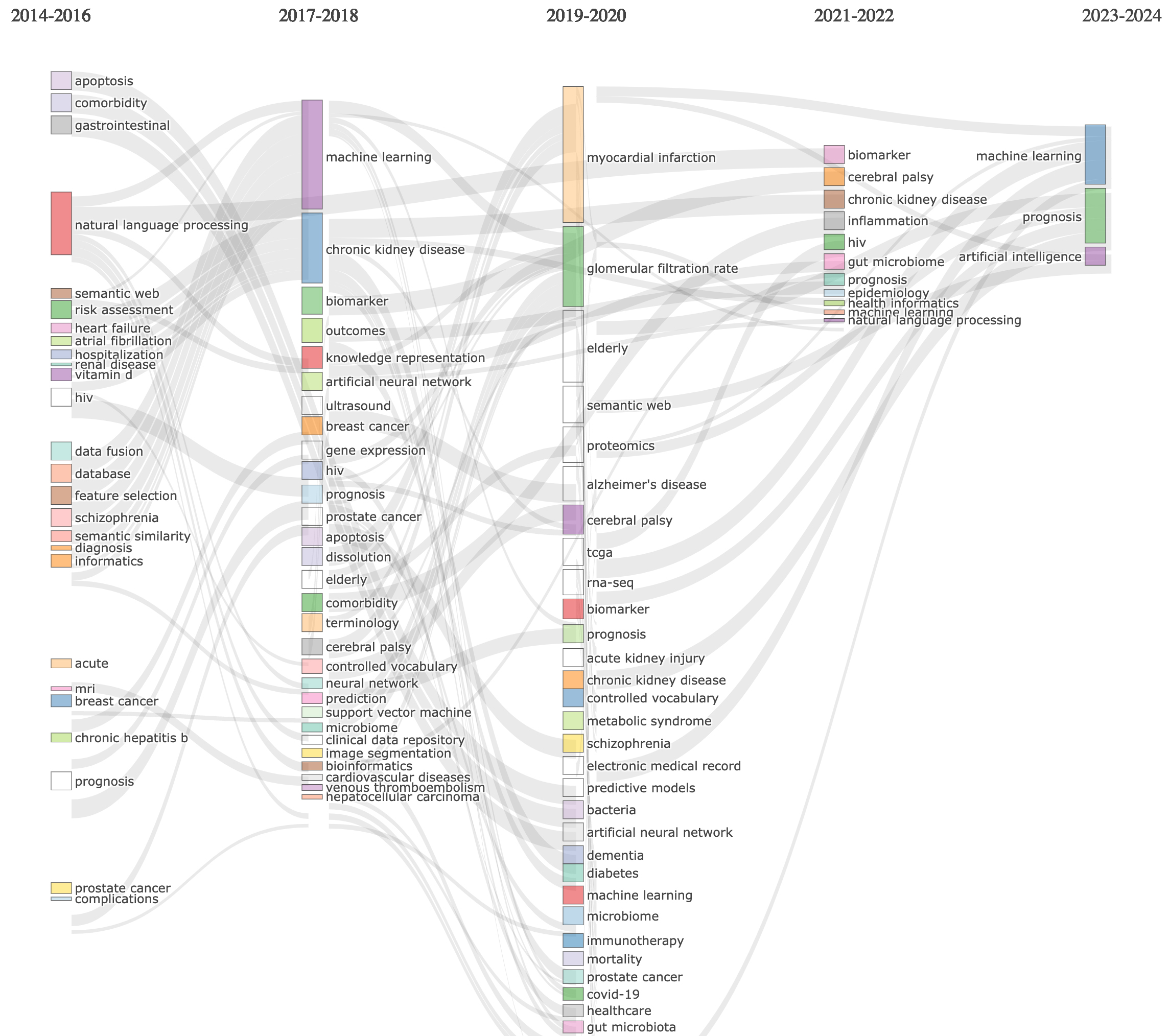 Fig. 9.
Fig. 9.
Longitudinal thematic map analysis of concepts.
Early research (2014–2016) shows a strong focus on NLP, semantic web technologies, and data fusion. These themes highlight the field’s initial efforts to enable semantic interoperability, text mining and knowledge extraction from unstructured clinical records. The prominence of NLP in this period suggests a push toward text mining and automated extraction from clinical records. Chronic disease topics like heart failure and prostate cancer were present, suggesting an early application of KOS-AI methods to research in these areas. At this stage, ML begins to appear as a basic theme, suggesting that it is being used but not driving new areas of research.
By 2017–2018, ML emerged as a dominant theme, accompanied by terms such as biomarkers, artificial neural networks, and electronic health records (EHRs). This indicates a methodological shift toward AI-driven predictive models for cancer and cardiovascular disease, and chronic kidney disease. Simultaneously, the presence of terms like terminology, controlled vocabulary, and knowledge representation may highlight a reliance on structured data and clinical terminologies to support AI methods. The period from 2019–2020 marks a major inflection point with the emergence of COVID-19 as a focal topic. Natural language and deep learning start to become more prominent, suggesting a movement of AI-related topics towards greater centrality and influence, as well as the use of deep learning, predictive models, and prognostic analytics for risk stratification, patient monitoring, and outbreak modeling. New themes such as myocardial infarction, Alzheimer’s disease, and chronic kidney disease indicate the increasing use of AI applications across clinical domains. The visibility of electronic health records, along with concepts such as proteomics, genomic data, and EHRs, also suggests integration of structured biomedical and clinical data into predictive frameworks.
The period 2021–2022 shows the growing role of AI/ML methodologies as foundational tools, with ML, biomarkers, and diabetes emerging as dominant themes. AI is now embedded not only in diagnostic applications but also in population health management, inflammatory disease modeling, and immune system research. In the most recent period, 2023–2024, AI, ML, and prognosis dominate the thematic landscape, now motor themes, due to their high centrality and maturity. These themes reflect a mature AI ecosystem focused on clinical outcome prediction, chronic disease management, and precision medicine. NLP has re-emerged as a key theme, suggesting renewed focus on extracting and structuring clinical data. This suggests a continued demand for KOS-enabled tools to support semantic alignment and integration with AI systems.
Overall, the thematic trends shift from semantically structured knowledge extraction to ML and AI-driven modeling. The early focus on ontologies, terminologies, and semantic web frameworks, perhaps laid a foundation for later advances in predictive analytics and precision medicine. However, the declining visibility of explicit KOS-related keywords in recent periods indicates that AI adoption in healthcare still faces challenges in structured data integration. Furthermore, the resurgence of NLP as a key method in later years, paired with high-frequency terms like biomarkers, prognosis, and chronic disease, suggests that semantic structuring remains critical to unlocking AI’s full potential. As the field matures, alignment between KOS and AI may be essential for supporting interoperability, explainability, and scalable clinical implementation.
This study explores how KOS-related terms appear in the AI and healthcare literature, identifying trends in the co-presence of KOS, AI, and healthcare concepts in scholarly discourse. It expands the existing literature by providing an evidence-based mapping of research trends, thematic structures, and conceptual evolution in this domain. The findings reveal a growing interdisciplinary convergence between KOS and AI methodologies. Consistent with prior studies (e.g., Wang et al, 2020), ML remains the most dominant AI method, particularly in biomarker discovery, precision medicine, and predictive analytics. Techniques such as random forest and support vector machines frequently appear in connection with KOS-enabled data standardization and classification tasks. However, the longitudinal analysis highlights the increasing importance of NLP for clinical text analysis and information extraction from electronic health records, as well as the emergence of deep learning in areas such as medical image analysis and diagnostics.
These trends reflect a growing scholarly emphasis on unstructured data and more complex knowledge representations in AI workflows. While early research prominently featured various KOS, their relative visibility has declined in recent years, suggesting either a waning emphasis or ongoing challenges with structured knowledge resources in AI-focused healthcare literature. The continued prominence of NLP in the literature may reflect ongoing efforts to extract structured meaning from unstructured clinical narratives. In later years, there is a significant trend observed where semantic web technologies and knowledge graphs are used as mechanisms to enhance the interpretability and context-awareness of AI models, particularly in biomedical data integration. These insights underscore the ongoing importance of aligning KOS-related approaches with AI development to support goals such as trustworthiness, accuracy, and interoperability in healthcare settings (Antoniadi et al, 2021; Kim et al, 2020).
Surprisingly, while LLMs such as Generative Pre-trained Transformer (GPT-3), BioGPT, and Medical Pathways Language Model (Med-PaLM), have become more popular in AI-discourse, their presence in this study was quite muted. Further analysis (see Table 1) shows that the term LLM appeared 133 times across the corpus yet did not surface prominently in co-occurrence networks or author keyword metadata. This is significantly less than the occurrence of other ML methods, such as Support Vector Machine (SVM). Reasons such as low occurrence, a combination of publishing lag, the recent rise of LLMs (post-2022), and inconsistent metadata usage, where authors may have referred to LLMs by model names (e.g., GPT, Bidirectional Encoder Representations from Transformers [BERT]) or under broader categories like “transformers” or “NLP”, may explain this. This finding highlights a limitation of bibliometric methods, where rapidly emerging trends may be underrepresented due to delays in indexing and keyword standardization. Despite this, LLMs will likely play a critical role in the next phase of KOS-AI integration, particularly for question answering, automated summarization, and semantic inference over large-scale health data.
| ML method | Frequency | Category | Example tasks |
| Vision Transformer (ViT) | 2142 | Transformer-Based | Retinal disease classification, skin cancer detection |
| Artificial Neural Networks (ANN) | 1370 | Deep Learning/Neural Networks | Predicting disease risk, medical diagnosis support |
| Generative Adversarial Networks (GAN) | 850 | Deep Learning/Neural Networks | Medical image synthesis, patient data de-identification |
| Neural Networks (generic) | 822 | Deep Learning/Neural Networks | Cancer classification, diabetes prediction |
| Random Forest | 685 | Supervised Learning | Identifying disease risk factors, classifying gene expression data |
| Support Vector Machines (SVM) | 680 | Supervised Learning | Cancer detection from histopathology, classifying EEG signals |
| Deep Learning | 653 | Deep Learning/Neural Networks | Image based diagnostics, drug response prediction, genomics analysis |
| Logistic Regression | 520 | Supervised Learning | Risk stratification, patient adherence modeling |
| Clustering | 408 | Unsupervised Learning | Patient segmentation, genetic data grouping, discovering disease subtypes |
| Convolutional Neural Networks (CNN) | 291 | Deep Learning/Neural Networks | Tumor detection in MRIs, diabetic retinopathy screening, dermatology imaging |
| Decision Trees | 234 | Supervised Learning | Symptom triage, treatment decision modeling |
| Gradient Boosting | 233 | Ensemble Learning | Predicting hospital stay duration or readmissions, EHR analysis |
| Boosting | 211 | Ensemble Learning | Symptom classification, therapy optimization |
| XG Boost | 172 | Ensemble Learning | Prediction of readmission, sepsis, ICU patient deterioration |
| BERT | 166 | Transformer-Based | Named entity recognition in health records, question answering on EMRs |
| Transformer | 140 | Transformer-Based | Multimodal patient record summarization |
| LLMs | 133 | Transformer-Based | Clinical text summarization, radiology report generation, question answering for EMRs |
| Deep Neural Networks (DNN) | 125 | Deep Learning/Neural Networks | Lab pattern detection, personalized treatment matching |
| Embedding Techniques | 120 | Other | Mapping clinical concepts to vector space, concept similarity computation, cross-code harmonization |
| Long Short-Term Memory (LTSM) | 100 | Deep Learning/Neural Networks | Time-series EHR prediction, adverse event detection from temporal data |
ML, machine learning; BERT, bidirectional encoder representations from transformers; LLMs, large language models; EEG, electroencephalogram; MRI, magnetic resonance imaging; EHR, electronic health record; ICU, intensive care unit; EMRs, electronic medical records.
RQ1: What thematic areas and AI methodologies emerge most prominently from literature addressing KOS-related terms in healthcare applications (2014–2024)?
The analysis reveals that KOS play a critical role in standardizing and structuring knowledge across a variety of AI implementations in healthcare. These standardized frameworks serve as foundational infrastructures that support AI development in health systems by enhancing data interoperability, contextualization, and reasoning. Among AI methodologies, ML emerged as the most frequently applied technique across the literature, particularly for application in areas such as clinical decision support, genomics, and medical imaging. Its dominance reflects the adaptability of ML to both structured and unstructured healthcare data, and its centrality to predictive and diagnostic applications. To show these thematic patterns, a co-occurrence analysis of author keywords was employed. This technique maps the frequency with which keywords co-appear across the corpus and reveals the central methodological approaches and thematic areas without the need for predetermined classification. The application of the Walktrap algorithm further enhanced this analysis by detecting natural communities within the co-occurrence network, effectively delineating distinct methodological domains and their conceptual boundaries.
The results suggest that KOS are most frequently associated with NLP and information extraction techniques. This association might indicate that KOS are positioned as resources that support AI systems to process and interpret unstructured clinical text from sources such as electronic health records (EHRs), clinical notes, clinical guidelines, handbooks, protocols, biomedical literature, and other narrative sources. This alignment is key to enhancing the interpretability and semantic precision of AI applications. Finally, the consistent frequency of the “semantic web” keyword suggests a sustained emphasis on using semantic technologies to make AI models more interpretable, explainable, and context-aware, particularly in biomedical data integration. Thus, KOS are not only viewed as auxiliary tools but as potential enablers of intelligent, semantically grounded healthcare systems.
RQ2: How has scholarly attention to knowledge organization systems evolved within AI-driven healthcare literature over the past decade?
The analysis of the temporal dimensions of both co-occurrence analysis and multidimensional scaling enables the identification of evolving research priorities. The longitudinal thematic mapping highlights shifts in scholarly attention over time, revealing how concerns about accuracy, interpretability, and interoperability have gained prominence in tandem with the development of AI methodologies. This temporal perspective is crucial for understanding the evolving discourse at the intersection of KOS and AI healthcare research and for identifying emerging paradigms. It shows how early emphasis on semantic interoperability has expanded to include concerns around model transparency, evaluation, and clinical trustworthiness.
The thematic mapping factorial analysis of keywords points to concepts such as “electronic health records” and “clinical decision support systems”, where KOS-related terms co-occur in literature, emphasizing the need for semantic consistency and interoperability in AI-driven health informatics applications. Standardized vocabularies such as SNOMED CT, ICD, and UMLS enable consistent encoding of clinical concepts, enabling AI models to interpret and extract meaningful insights from large-scale, heterogeneous datasets. Their presence in the network signals recognition of their importance in supporting machine-readable logic, semantic precision, and data harmonization.
Furthermore, terms like “model evaluation” and “performance assessments” co-occur with KOS-related concepts, suggesting a growing association between knowledge representation and efforts to enhance AI interpretability and trustworthiness. KOS are often positioned in the literature as contributing to explainability by providing explicit definitions, hierarchical relationships, and domain-specific constraints that help reduce ambiguity and improve the accuracy and interpretability of AI-driven predictions and decisions. The increasing use of knowledge graphs and ontologies in genomics, precision medicine, and molecular biology also reflects a shift toward knowledge-enriched AI model development. Rather than passive data sources, KOS are framed as conceptual infrastructures that support the development of transparent, interoperable, and domain-aware AI applications.
RQ3: What conceptual gaps, challenges, or opportunities are revealed in the scholarly literature on knowledge organization systems within AI-driven healthcare research?
Although KOS are foundational to structured data representation, the bibliometric analysis reveals that they are conceptually underemphasized within the broader discourse on AI-driven healthcare. The multidimensional scaling visualization highlights fragmentation within the field. Certain terms appear spatially isolated or weakly connected, suggesting that although methodologically mature, some areas remain less connected to dominant research trajectories. Most notably, the niche themes quadrant, which included terms like NLP, EHRs, terminologies, and ontologies, indicates that while these technologies are well developed, they remain peripheral to dominant AI paradigms.
These patterns suggest a conceptual gap in the literature. While KOS are frequently framed as enabling semantic interoperability and explainability, they are not consistently foregrounded in scholarly discussions of AI development or implementation. The longitudinal thematic map further supports this finding, showing that while KOS-related terms such as “controlled vocabularies”, “semantic networks”, and “ontologies” were more prominent in early research (2014–2018), their relative visibility has declined in recent years, especially when compared to AI-centric or healthcare specific keywords, which underscores their continued underrepresentation in the evolving literature. The decline might also reflect a shift in scholarly focus toward more flexible, black-box AI models that lack built-in semantic interpretability.
During the COVID-19 pandemic period (2019–2021), there was a noticeable shift toward AI-driven solutions in response to urgent healthcare challenges. Some publications during this period highlight the potential scalability and adaptability of KOS-informed approaches when confronted with large, rapidly changing data environments. This trend not only highlights the potential of KOS in supporting real-time clinical decision making and epidemiological forecasting but also exposes current limitations in how they are positioned or operationalized. Rather than indicating the absence of KOS altogether, the findings suggest it reveals an underexplored area of research where structured knowledge resources could more fully contribute to the development of transparent, context-aware, and interoperable AI systems in healthcare. Rather than indicating the absence of KOS altogether, the findings suggest that their role is often implicit or fragmented, lacking the conceptual integration needed to support large-scale AI adoption. Addressing this may require future research to focus on the development of AI systems that are explicitly informed by ontologies and knowledge graphs for reasoning and interpretation, standards for integrating unstructured narrative data (e.g., clinical notes) with KOS, and mechanisms that support ethical alignment, such as transparent labeling, traceability, and bias reduction enabled by KOS.
The intersection of KOS and AI within healthcare research presents both opportunities and challenges that have significant implications for research methodologies, clinical practice, and policy development. As AI systems become increasingly embedded in healthcare applications, the literature reveals growing attention to the need for structured, standardized, and interoperable knowledge representations. The findings highlighted how KOS are positioned in the literature as contributing to AI interpretability, while also pointing to conceptual and structural barriers that may hinder their broader integration. These insights contribute to ongoing discussions about how AI-driven healthcare systems could be ethically governed by integrating values of fairness, accuracy, and transparency, areas where KOS are frequently mentioned as supporting structures. This section explores implications for future research directions and policy frameworks. The following are some suggestions for consideration by researchers and policymakers in this space.
Encourage developing and adopting standardized KOS across healthcare systems to facilitate seamless data exchange and semantic interoperability. A review of articles in the dataset explicitly mentioning ML methods provides insights into some best practices for integrating KOS into AI systems. Of the total articles in the corpus (n = 5836), 78.4% of articles mentioned at least one ML method (a Python script was used to generate these numbers). Table 1 shows the top twenty ML methods across various categories, including Supervised and Unsupervised Learning Models, Transformer-Based Models, Deep Learning and Neural Networks, and Ensemble Learning.
The integration of KOS into ML models can be leveraged to enhance model interpretability and semantic reasoning. Several approaches could be adopted to seamlessly incorporate KOS. For example:
• Deep Learning and Neural Networks: Encode domain ontologies such as SNOMED CT either as categorical input features or by leveraging their hierarchical structure to inform the feature space. This can guide representation learning so that the model’s internal representations reflect clinically meaningful relationships and classifications (Lee et al, 2021; Oudah and Henschel, 2018; Sahoo et al, 2022). Alternatively, embeddings derived from knowledge graphs constructed from these ontologies can be integrated into the model input layer to enhance semantic understanding and improve interpretability.
• Transformer-Based Models: Enhance language models by linking text entities to structured KOS, e.g., SNOMED, Diabetes Mellitus Diagnosis Ontology (DDO), enabling the model to learn from contextualized semantic relationships during training and inference (Furqon et al, 2022; Jia et al, 2019; Wang et al, 2022). Another option would be to insert ontology-derived triples into the model’s architecture during pretraining, enriching the model with domain-specific semantics (He et al, 2019; Pan et al, 2024). Alternatively, domain ontologies could be used to generate concept embeddings, which are then used as model inputs to enhance semantic representation (Alawad et al, 2018; Chandrashekar et al, 2018; Karadeniz and Özgür, 2019; Topaz et al, 2019).
• Support Vector Machines & Random Forests: Utilize ontological hierarchies to create structured feature representations that encode semantic proximity or taxonomic depth. Additionally, association rules or decision paths derived from knowledge graphs can be integrated to inform feature selection or post hoc explanation, to improve transparency and trustworthiness in model outputs (Kulmanov et al, 2021; Paulheim, 2016).
Interdisciplinary collaboration between KOS experts and AI researchers could focus on generating additional ideas for leveraging the strengths of both knowledge structures and AI/ML techniques.
The introduction outlined some challenges in applying AI to healthcare, including algorithmic bias in ML models and hallucinations from generative language models. To mitigate these issues, it is advisable to ensure that datasets are diverse and representative. This can be done by implementing fairness-aware techniques such as understanding the processes by which clinicians and algorithms jointly contribute to final decisions, developing affirmative ML systems that considers the views of minority groups, using diverse and representative datasets, regularly auditing and validating AI algorithms, and educating clinicians and patients on AI bias (Grote and Keeling, 2022; Ueda et al, 2024).
In addition to this measure, supporting research into explainable AI (XAI) techniques that can provide transparency into the decision-making process of AI algorithms should be encouraged. The findings of the study revealed ML as a dominant theme, with the presence of terms like “model evaluation and “performance assessments” underscoring the importance of AI transparency and trust. Moreover, increasing occurrences of keywords such as “semantic web” and “knowledge graph” suggest a growing trend towards semantic technologies to make AI models more interpretable and context-aware in healthcare data integration. As previously discussed, KOS provide explicit definitions and semantic relationships that help to reduce ambiguity and improve the accuracy and interpretability of AI-driven predictions and decisions (Bodenreider et al, 2018; Bronnert et al, 2012; Fung and Bodenreider, 2023; Zeng et al, 2020). Yet despite this, the longitudinal thematic map suggests that structured data remains a persistent challenge in the discourse surrounding AI adoption in healthcare. This highlights the continued importance of research focused on leveraging ontologies, knowledge graphs, electronic health records, and NLP to support the development of AI systems that are more interpretable and trustworthy for healthcare professionals and patients.
The intersection of KOS and AI-driven healthcare is an area of growing scholarly attention, as evidenced by the 27.72% annual growth rate in publications over the past decade. To ensure the continued and responsible development of research in this evolving field, additional strategic actions should be taken to foster collaboration and strengthen governance frameworks.
(1) Fostering Interdisciplinary Collaboration: The high number of co-authors per document (7.77) in this research area suggests that meaningful progress relies on interdisciplinary collaboration between AI researchers, healthcare professionals, and KOS experts. Future research could benefit from fostering such collaborations to address the challenges identified, particularly concerning standardization and interoperability of healthcare data. To strengthen this collaboration, institutions should establish joint research programs and knowledge-sharing opportunities to bring together experts from different domains. Cross-sector partnerships between academia, industry, and healthcare organizations may also be encouraged.
(2) Developing Ethical Guidelines and Governance Frameworks: As AI becomes more embedded in healthcare decision-making, ethical concerns such as data privacy, algorithmic bias, and accountability become more critical and should be proactively addressed. Ethical guidelines should emphasize FAIR data principles (Findable, Accessible, Interoperable, Reusable) (see https://www.go-fair.org/fair-principles/), ensuring that the KOS and other structured knowledge resources used are widely available and responsibly managed in AI-driven healthcare applications. Finally, establishing robust governance frameworks to ensure that these systems are incorporating KOS in ways that enhance their explainability and align with regulatory standards is crucial.
This bibliometric analysis provides a comprehensive exploration of how KOS-related terms appear in the evolving landscape of AI and healthcare literature over the past decade. These findings suggest that while AI is becoming increasingly embedded in clinical decision making, KOS remain conceptually important for achieving semantic interoperability, accuracy, and interpretability in AI-driven systems. The study highlights both the transformative potential and the persistent challenges of KOS-AI synergies. While machine learning remains the dominant methodology, the resurgence of NLP and the growing role of semantic technologies such as knowledge graphs underscore a continuing need to bridge unstructured clinical data with structured knowledge models.
However, the relative decline in visibility of KOS-specific terms, particularly in more recent AI literature, suggests that discourse around the emerging paradigms, such as LLMs, has not yet consistently engaged with the structured knowledge principles that KOS provide. This points to a conceptual and developmental gap, indicating an urgent need for frameworks that explicitly embed KOS into these systems. A key insight from the analysis illustrated the importance of designing robust, scalable, and ethically aligned methodologies that integrate KOS into diverse AI applications. Future progress in this area will require collaborative engagement among researchers, policymakers, and industry leaders, which should prioritize ethical oversight, domain alignment, and methodological innovations to unlock the full potential of knowledge organization approaches with AI methodologies in healthcare delivery and clinical decision-making.
The study offers important insights but has a few limitations that may influence the interpretation and scope of the findings. First, the analysis was restricted to publications indexed in Web of Science (WoS) and Scopus, potentially excluding relevant work from databases such as PubMed, IEEE Xplore, or domain-specific repositories. Second, although carefully constructed, the search query may have missed studies that addressed KOS or AI-related concepts without using standardized terminology or explicit references to the specific terms included. Third, the analysis relied heavily on metadata fields (e.g., author keywords), which may have underrepresented emerging concepts or interdisciplinary concepts like “LLM” due to inconsistent indexing or variation in terminology across disciplines. Further, bibliometric methods are subject to a lag in capturing rapidly evolving research trends, which likely contributed to the limited visibility of recent developments, including the rise of generative AI.
In addition, the bibliometric methods employed in this study, while well suited for mapping large-scale patterns, impose limitations in distinguishing between interrelated but conceptually distinct categories. Although the document retrieval prioritized records containing KOS-related terms, the subsequent co-occurrence, clustering, and thematic mapping analyses were conducted on the full set of keywords without categorizing them by functional domain (e.g., KOS, AI method, healthcare application). As a result, the analysis does not explicitly model the relationships among these categories or differentiate how KOS terms function—as reference terminologies, ontologies, or interface tools—within AI-driven healthcare contexts. This might have contributed to the relatively peripheral position of KOS-related terms in the resulting thematic structures, despite their central importance in health information systems. Moreover, the method is unable to disambiguate polysemous terms such as “standardization” or “interoperability” which may be used differently across disciplines.
Despite these limitations, this study provides a high-level, exploratory view of how KOS-related concepts are represented in scholarly discourse, rather than a fine-grained evaluation of technical integration or implementation mechanisms. Future research should address these constraints by expanding bibliometric coverage to include a broader range of sources and employ more nuanced search strategies, including full-text analysis, or citation analysis. Mixed-method approaches that combine bibliometric mapping with qualitative content analysis or expert coding could enable better differentiation of concept functions and more precise modeling of KOS-AI interactions.
Further, empirical case studies of KOS-AI integration in real-world clinical environments could provide richer insight into operational challenges, success factors, and implementation pathways that are not visible through bibliometric analysis alone. Research should also prioritize the development of toolkits that facilitate the incorporation of KOS into AI model training, evaluation, and deployment pipelines. This includes attention to semantic alignment, explainability, and interoperability. In sum, advancing from conceptual co-presence to meaningful integration of KOS into AI will be essential to building trustworthy, transparent, and context-aware healthcare systems capable of supporting precision medicine, clinical analytics, and the ethical stewardship of healthcare knowledge and decision support.
All data reported in this paper will be shared by the corresponding author upon reasonable request.
JC conceptualized the research idea, analyzed the data, interpreted the data, and wrote the manuscript. EA helped with data analysis and manuscript writing. Both authors contributed to editorial changes in the manuscript. Both authors read and approved the final manuscript. Both authors read, approved the final manuscript, and agreed to be accountable for all aspects of the work.
We gratefully acknowledge the feedback of the reviewers for their thoughtful and constructive feedback which helped to strengthen the clarity and framing of this manuscript. We additionally acknowledge the assistance and support of the Department of STEM Education and Professional Studies at Old Dominion University.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.









