1 Department of Neurosurgery, Renmin Hospital of Wuhan University, 430060 Wuhan, Hubei, China
2 Department of Neurosurgery, The Second Affiliated Hospital of Nanchang University, 330008 Nanchang, Jiangxi, China
Abstract
Intracranial space-occupying lesions (IOLs) often require precise surgical resection. Intraoperative neurophysiological monitoring (IONM), including somatosensory evoked potentials (SEPs) and motor evoked potentials (MEPs), is widely used to preserve neurological function. However, interpretation of IONM data still relies heavily on the experience of the surgeon. The aim of this study was to develop machine-learning (ML) models based on IONM data to support the assessment of lesion location relative to functional brain areas and surgical outcomes.
We initially screened 377 patients undergoing microsurgical resection of IOLs. The clinical data on these patients included demographic characteristics, quantitative IONM parameters (SEP and MEP amplitude and latency), lesion localization, and postoperative adverse events. Four ML models were developed: support vector machine (SVM), decision tree, random forest, and naïve Bayes. Model performance was evaluated using several metrics, including accuracy, sensitivity, specificity, precision, F1-score, and the area under the curve (AUC).
Significant differences in SEP and MEP parameters were observed between patient groups with lesions located in functional and non-functional brain areas (all p < 0.05). SEP and MEP parameters were both associated with lesion localization and postoperative adverse events, with differential correlation patterns observed between the two modalities. The ML models demonstrated moderate discriminative performance in predicting lesion involvement in functional areas, with the highest accuracy of 79.2% in the training set and 65.00% in the test set. The models showed good performance in predicting serious adverse events, with the best accuracy of >78% in both datasets.
ML models based on IONM data may help to assess lesion location relative to functional brain areas, as well as the prediction of postoperative outcomes. These findings suggest that ML-assisted analysis of IONM data may provide an exploratory framework for understanding lesion localization and postoperative outcomes, rather than a clinically deployable decision-support tool.
Keywords
- intraoperative neurophysiological monitoring
- evoked potentials, somatosensory
- evoked potentials, motor
- machine-learning
- postoperative complications
- brain mapping
Intracranial space-occupying lesions (IOLs) refer to a group of diseases that occupy a certain space within the cranial cavity. IOLs present with symptoms such as increased intracranial pressure, headache, and sensory and motor function impairment [1, 2]. These lesions include brain tumors, cerebrovascular diseases, and brain abscesses. The prognosis of patients is closely related to factors such as age, gender, type of lesion, and location of the lesion [3]. The scope of surgical resection is determined by whether the lesion is located in the functional area, while the experience of neurosurgeons can directly affect the prognosis of patients with hemiplegia and their quality of life [4, 5, 6]. The lesion location and risk of the surgery area are always determined by magnetic resonance imaging (MRI) prior to neurosurgery. However, because the normal anatomical structures are twisted due to the lesion, the functional area cannot be distinguished during surgery, causing postoperative functional impairment in patients. Therefore, Intraoperative neurophysiological monitoring (IONM) is needed during neurosurgery to determine the functional area and protect neurological function.
IONM is often used to evaluate the functional integrity of target neural structures in neurosurgery [7, 8]. It has been widely used in various neurosurgical procedures, including skull base lesion resection, aneurysm clipping, and catheter drainage. Somatosensory evoked potentials (SEPs) and motor evoked potentials (MEPs) are the most commonly used monitoring methods [9, 10].
Currently, the extent of surgical resection for IOLs is mainly based on the experience of neurosurgeons and electrophysiologists, with a lack of reliable methods to determine the surgical boundaries [11]. This results in a high surgical threshold and relatively random patient outcomes. Better methods are needed to control artificial errors. Machine-learning (ML) is proposed as a way to solve this problem. Therefore, based on IONM data from patients, we used ML to establish a predictive model assessing whether IOLs were in functional areas, and for the prediction of patient outcomes during the operations.
This retrospective study included patients who underwent microsurgical resection of IOLs at our institution between March 2019 and July 2022. Initially, 377 consecutive patients were identified, and these were eligible for inclusion if they met the following criteria:
(1) availability of complete IONM data, including SEP and MEP;
(2) availability of complete clinical records and follow-up information;
(3) availability of preoperative and postoperative MRI for lesion localization and evaluation of surgical extent.
Patients were excluded if electrophysiological data were incomplete or technically inadequate (n = 43) or if follow-up data were missing or insufficient for the assessment of outcome (n = 134). After applying these criteria, a final cohort of 200 patients was included in the analysis. The patient selection process and data filtering steps are shown in the study flowchart (Fig. 1).
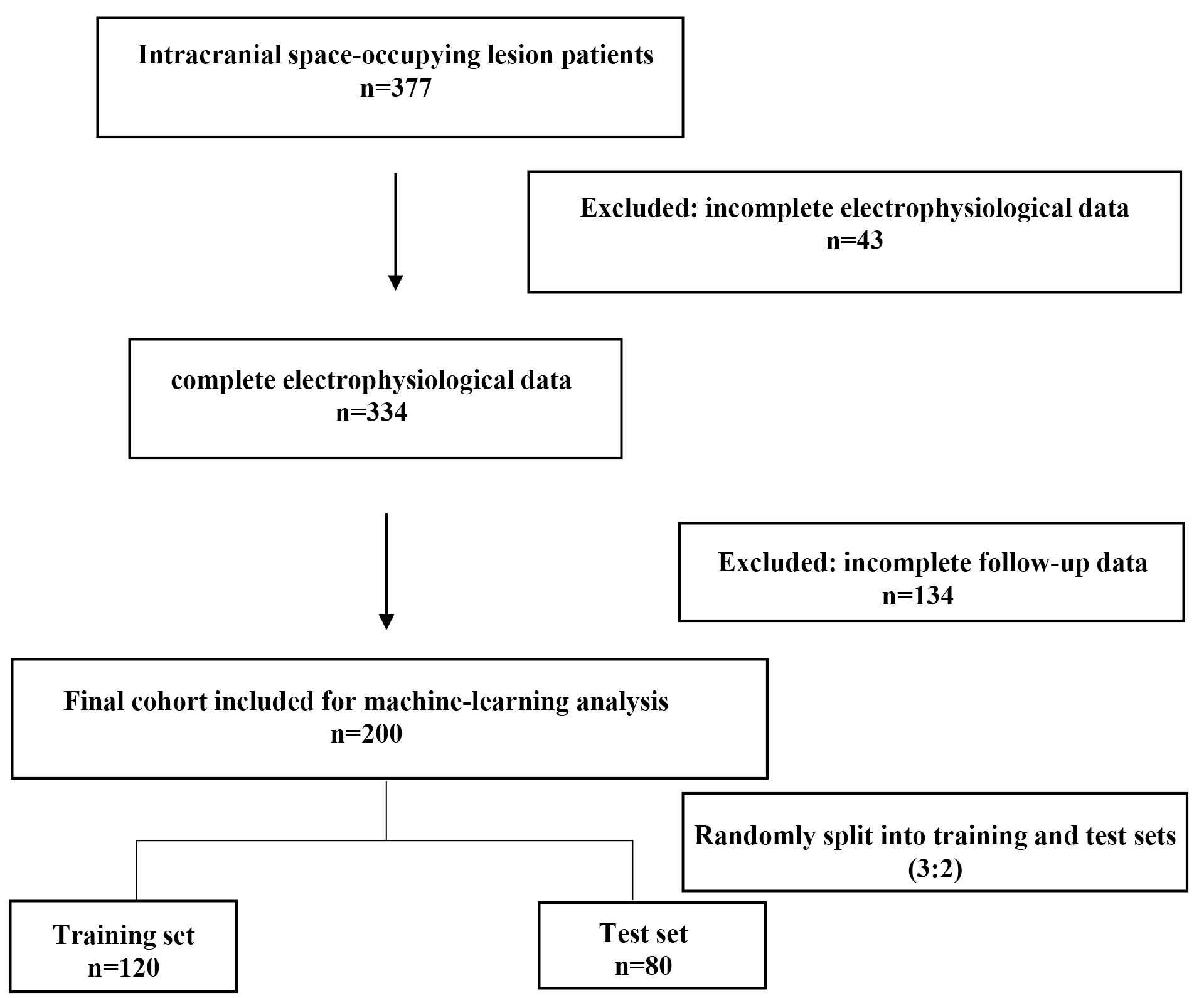 Fig. 1.
Fig. 1.
Flowchart.
The final cohort was randomly divided into a training set (n = 120) and an independent test set (n = 80) using a 3:2 ratio to ensure sufficient data for model training, while preserving an independent dataset for performance evaluation.
The original data are provided in the Supplementary Material.
Baseline demographic and clinical data were collected for all patients, including age, sex, pathological disease type, presence of preoperative limb paralysis, and lesion location. Lesions were classified as involving functional or non-functional brain areas based on preoperative imaging findings and intraoperative anatomical assessment.
All patients underwent standardized preoperative neurological examinations, including assessment of muscle strength, coordination, sensory function, and cranial nerve status. Muscle strength was graded according to criteria established by the Chinese Medical Association.
Intraoperative neurophysiological monitoring was performed in all patients using the Xltek Protektor32 monitoring system (DBA Excel-Tech Ltd., Oakville, ON, Canada) under a standardized protocol.
Transcranial electrical stimulation was delivered via disposable scalp needle electrodes placed at C3 and C4 according to the international 10–20 electroencephalography (EEG) system. During stimulation of the left limb, C4 served as the anode and C3 as the cathode, whereas during stimulation of the right limb, C3 served as the anode and C4 as the cathode.
MEP signals were recorded using disposable, twisted-pair needle electrodes placed in the abductor pollicis brevis and abductor digiti minimi muscles for the upper limbs and the abductor hallucis for the lower limbs, with the tibialis anterior muscle serving as the ground electrode. The recording bandwidth was set at 30–1000 Hz. Each stimulus consisted of a train of four pulses, with an inter-pulse interval of 2 ms. The stimulation intensity ranged from 150 to 300 V, determined individually at the beginning of each procedure. To minimize interference with surgical manipulation, MEP testing was coordinated with the surgical team and performed at approximately 10-minute intervals.
SEP monitoring was performed using peripheral nerve stimulation delivered through disposable adhesive electrodes. The median nerve was stimulated for upper-limb SEP recording, and the posterior tibial nerve was stimulated for lower-limb SEP recording. Stimulation frequencies were 4.71 Hz for upper limbs and 3.27 Hz for lower limbs, with a pulse width of 0.2 ms.
SEP signals were recorded using disposable scalp needle electrodes placed at C4′, C3′, and Cz′ (1 cm posterior to the corresponding 10–20 system landmarks), with Fz as the reference electrode. The recording bandwidth was set at 30–1000 Hz. The stimulation intensity ranged from 10 to 30 mA and was determined individually at the start of surgery. Averaged waveforms were obtained through repeated stimulation, with recordings performed at approximately 10-minute intervals to minimize electrical interference.
Although raw SEP and MEP waveforms were recorded intraoperatively, electrophysiological features used for analysis were limited to quantitative parameters, including signal amplitude and latency, rather than raw waveform-based modeling. This approach was adopted due to practical constraints and to ensure interpretability and robustness of the predictive models.
The original pathological disease types were recorded for all patients. However, due to the limited sample size, further stratification into multiple disease subtypes would have resulted in small subgroup sizes and reduced statistical power. Therefore, disease types were not further subdivided during model development.
Postoperative functional outcomes were assessed primarily through structured
telephone follow-up. Limb function was evaluated using patient-reported scores on
a scale of 0–100, with scores
To integrate dynamic electrophysiological signals with static demographic and clinical variables, four ML models were developed: support vector machine (SVM), decision tree, random forest, and Bayesian models. Model development and statistical analyses were conducted using Python (version 3.13.12, Python Software Foundation, Wilmington, DE, USA) in a Jupyter Notebook environment, selected for compatibility with the analysis workflow used in this study.
Descriptive statistics were summarized as means with 95% confidence intervals. Group comparisons were performed using Student’s t-test. Correlation analyses among clinical and electrophysiological variables were conducted and visualized using heatmaps.
Model performance was evaluated using confusion matrices and receiver operating characteristic (ROC) analysis. Predictive performance was assessed in both the training and independent test datasets.
Given the substantial class imbalance and distributional differences between the serious adverse event (SAE) and functional-area prediction tasks, separate model tuning strategies were applied for SAE prediction. Hyperparameters were optimized to improve minority-class sensitivity, and decision thresholds were adjusted to account for outcome imbalance.
All patients in this study received combined intravenous and inhalation
anesthesia with a minimum alveolar concentration (MAC)
A total of 200 patients were included in the final analysis. The baseline demographic and clinical characteristics of the study population are summarized in Table 1. The mean age was 52.2 years, and 42.5% of patients were male. Malignant tumors accounted for 73.5% of cases. Lesions involving functional brain areas were identified in 53.0% of patients. Preoperative limb paralysis was present in 18.5% of patients, and serious adverse events occurred in 11.5% of patients during follow-up.
| Characteristics | Overall | SAE (n = 23) | Functional area (n = 106) |
| Age, years (mean |
52.2 |
51.8 |
53.7 |
| Male, n (%) | 85 (42.5%) | 10 (43.5%) | 45 (42.5%) |
| Malignant tumor n (%) | 147 (73.5%) | 11 (47.8%) | 72 (67.9%) |
| Lesion in functional area, n (%) | 106 (53.0%) | 18 (78.3%) | 106 (100.0%) |
| Serious adverse event (SAE), n (%) | 23 (11.5%) | 23 (100.0%) | 18 (17.0%) |
| Any preoperative limb paralysis, n (%) | 37 (18.5%) | 6 (26.1%) | 29 (27.4%) |
SEP amplitude and latency differed significantly between patients with lesions
located in functional and non-functional brain areas. As shown in Fig. 2A,B, SEP
amplitude of both the left and right upper limbs was significantly different
between the two groups (left: p = 6.07
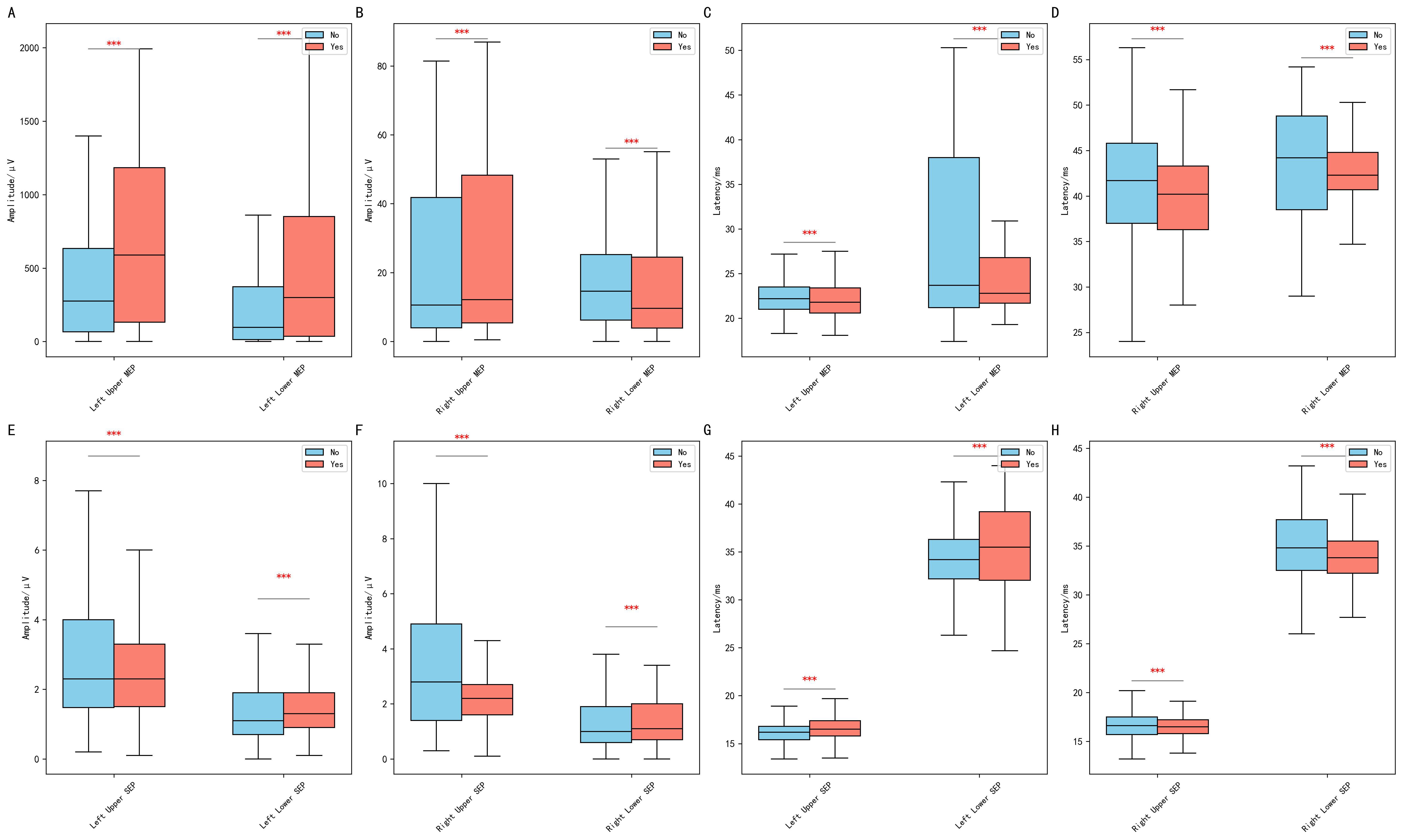 Fig. 2.
Fig. 2.
MEP and SEP according to lesion location in eloquent
areas. Comparative analysis of motor evoked potentials (MEPs) and somatosensory
evoked potentials (SEPs). (A,B) Comparison of SEP amplitude between patients with
intracranial lesions located in functional and non-functional brain areas for the
left and right limbs, respectively. (C,D) Comparison of SEP latency between the
two groups for the left and right limbs, respectively. (E,F) Comparison of MEP
amplitude between patients with lesions located in functional and non-functional
brain areas for the left and right limbs, respectively. (G,H) Comparison of MEP
latency between the two groups for the left and right limbs, respectively. (Statistical significance is indicated as follows: ***p
SEP latency also differed significantly between the two groups. As shown in Fig. 2C,D, patients with functional-area lesions exhibited shorter SEP latencies in
both the upper limbs (left: p = 8.46
MEP parameters also showed significant differences between patients with lesions
located in functional and non-functional brain areas. As shown in Fig. 2E,F, MEP
amplitude for the upper limbs differed significantly between the two groups
(left: p = 3.14
MEP latency also differed significantly between the two groups. As shown in Fig. 2G,H, patients with functional-area lesions exhibited greater deviations in MEP
latency compared to those with non-functional-area lesions (upper limbs:
p = 5.67
Correlation analysis revealed associations between electrophysiological parameters, lesion localization, and clinical outcomes. As shown in Fig. 3, SEP and MEP parameters were positively correlated with lesion involvement in functional brain areas, and negatively correlated with the occurrence of serious adverse events.
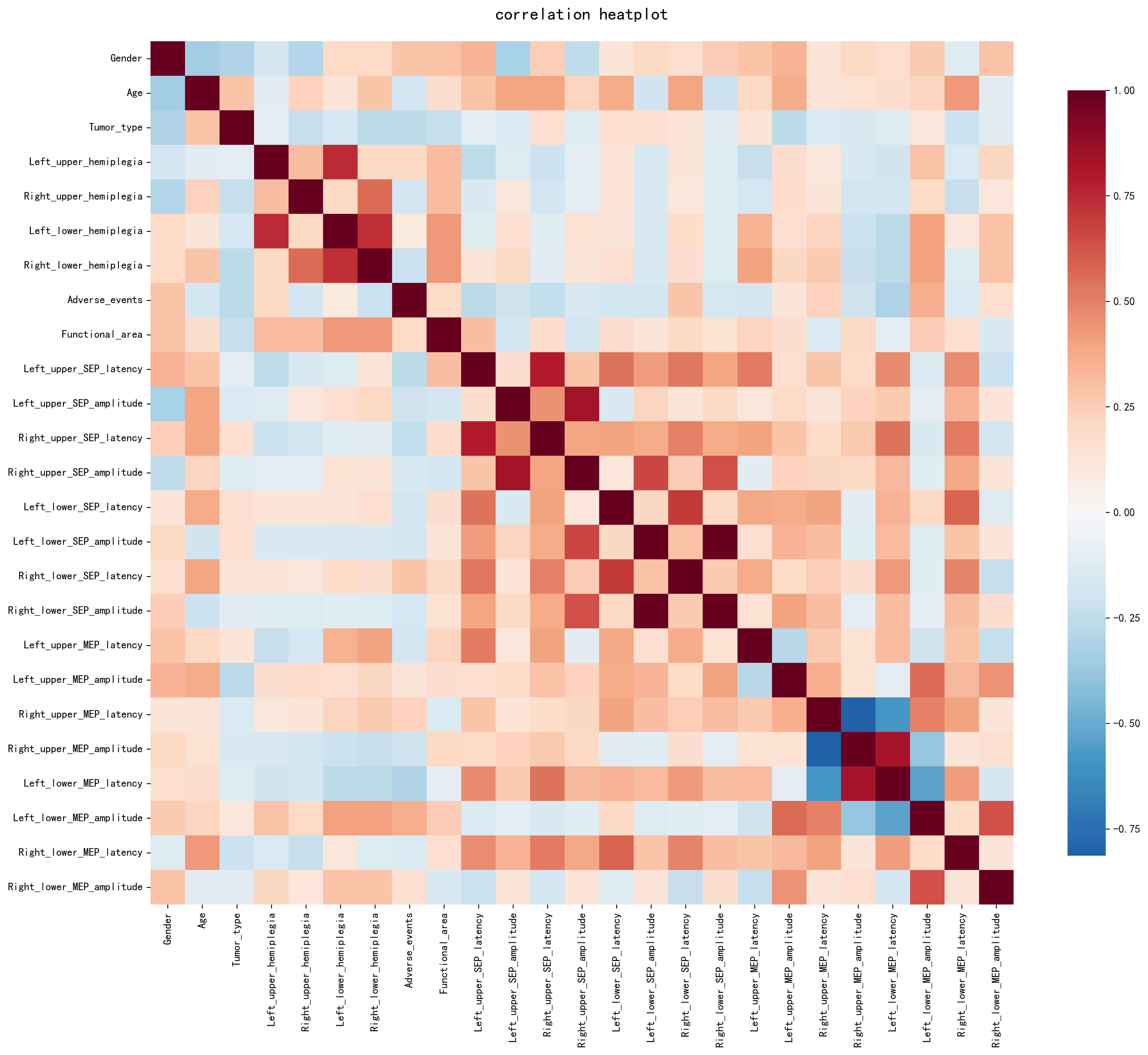 Fig. 3.
Fig. 3.
Correlation analysis of MEP and SEP. The heatmap illustrates pairwise correlations between baseline clinical variables, lesion involvement in functional brain areas, serious adverse events, and quantitative SEP and MEP parameters. The color intensity reflects the strength and direction of correlations, with red indicating positive correlations and blue indicating negative correlations. SEP and MEP parameters show differential correlation patterns with lesion localization and postoperative adverse outcomes.
ML models integrating baseline clinical variables and electrophysiological parameters were developed to predict whether lesions were located in functional brain areas. In the training set, the prediction accuracy ranged from 60.0% to 79.2%, with the highest accuracy observed for the decision tree and random forest models (both 79.2%).
In the independent test set, the prediction accuracy ranged from 51.3% to 65.0%. Detailed performance metrics, including accuracy, sensitivity, specificity, precision, F1-score, and AUC, are summarized in Table 2. Confusion matrices and ROC curves for each model are shown in Figs. 4,5, respectively.
| Model | Accuracy | Sensitivity | Specificity | Precision | F1-score | AUC |
| SVM | 0.563 | 0.488 | 0.649 | 0.618 | 0.545 | 0.590 |
| Decision Tree | 0.513 | 0.581 | 0.432 | 0.543 | 0.562 | 0.560 |
| Random Forest | 0.538 | 0.558 | 0.514 | 0.571 | 0.565 | 0.530 |
| Naïve Bayes | 0.650 | 0.419 | 0.919 | 0.857 | 0.563 | 0.640 |
SVM, support vector machine; AUC, area under the curve; ML, machine-learning.
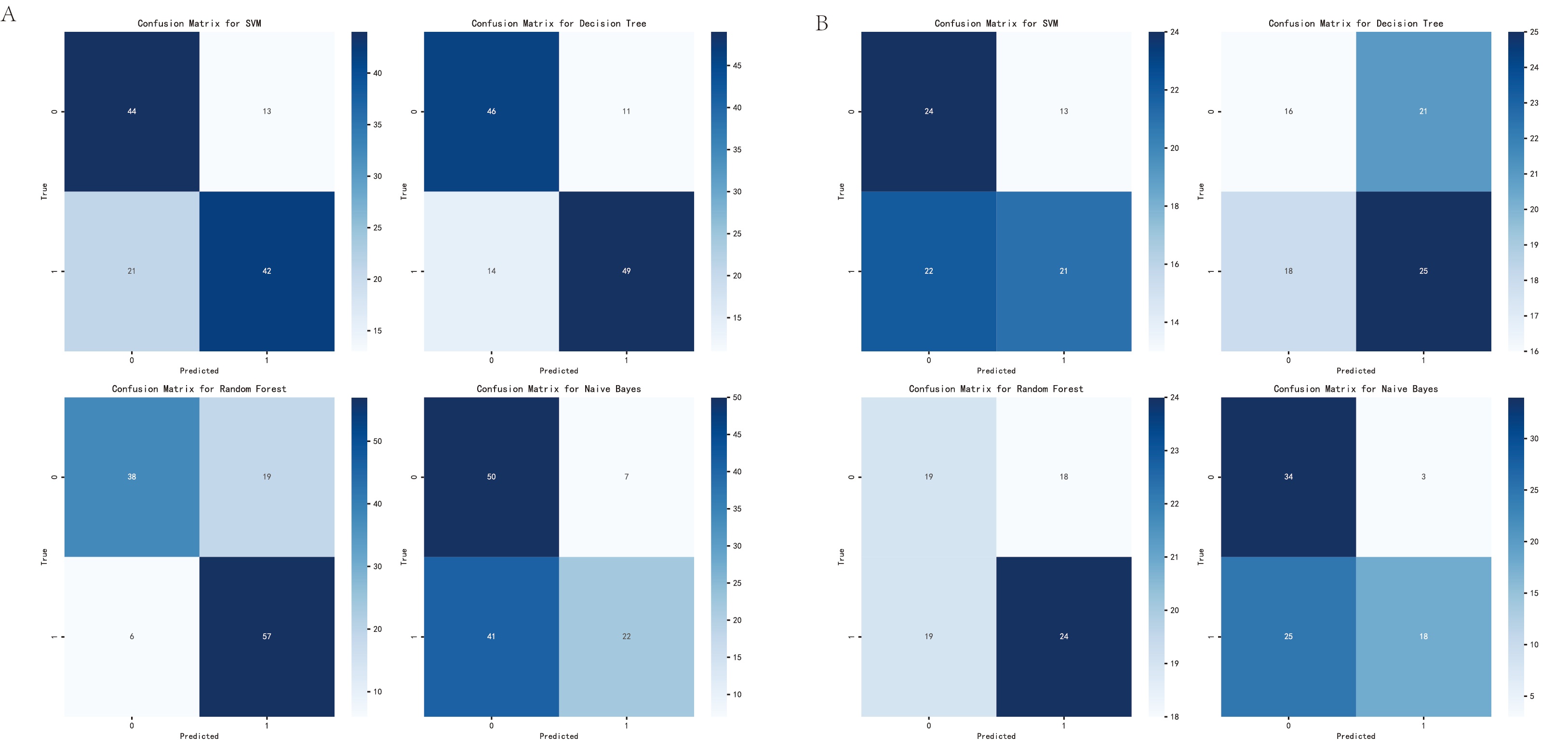 Fig. 4.
Fig. 4.
Prediction of functional area. (A) Confusion matrices of the support vector machine, decision tree, random forest, and naïve Bayes models in the training set. (B) Confusion matrices of the same models in the independent test set. These matrices illustrate the distribution of true positive, true negative, false positive, and false negative predictions for each model when classifying whether intracranial lesions involve functional brain areas.
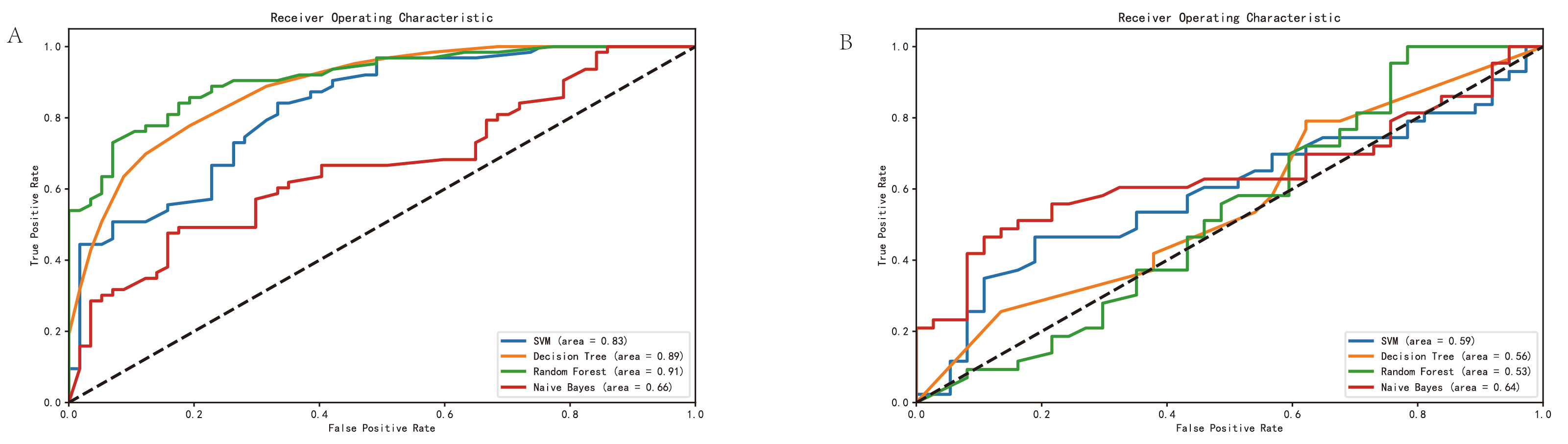 Fig. 5.
Fig. 5.
ROC curves of functional area. (A) ROC curves of the support vector machine, decision tree, random forest, and naïve Bayes models in the training set. (B) ROC curves of the same models in the independent test set. The ROC curves illustrate the trade-off between sensitivity and specificity across different classification thresholds. The AUC for each model is indicated in the legend. ROC, receiver operating characteristic.
The same ML framework was applied to predict serious adverse events. Given the low positive rate and the imbalanced distribution of SAEs in the original dataset, task-specific parameter tuning was performed before model prediction. In the training set, the prediction accuracy ranged from 87.5% to 91.7%, while the AUC values ranged from 0.92 to 0.98.
In the independent test set, the prediction accuracy ranged from 38.8% to 78.8%, and the AUC values ranged from 0.58 to 0.68. Comprehensive performance metrics are summarized in Table 3. Confusion matrices and ROC curves are presented in Figs. 6,7, respectively. Given the relatively small sample size, the number of patients experiencing serious adverse events was limited, which may have adversely affected the stability and reliability of certain model performance metrics [12].
| Model | Accuracy | Sensitivity | Specificity | Precision | F1-score | AUC |
| SVM | 0.638 | 0.556 | 0.648 | 0.167 | 0.256 | 0.580 |
| Decision Tree | 0.725 | 0.444 | 0.761 | 0.190 | 0.266 | 0.670 |
| Random Forest | 0.788 | 0.333 | 0.845 | 0.214 | 0.261 | 0.610 |
| Naïve Bayes | 0.388 | 0.889 | 0.324 | 0.143 | 0.246 | 0.680 |
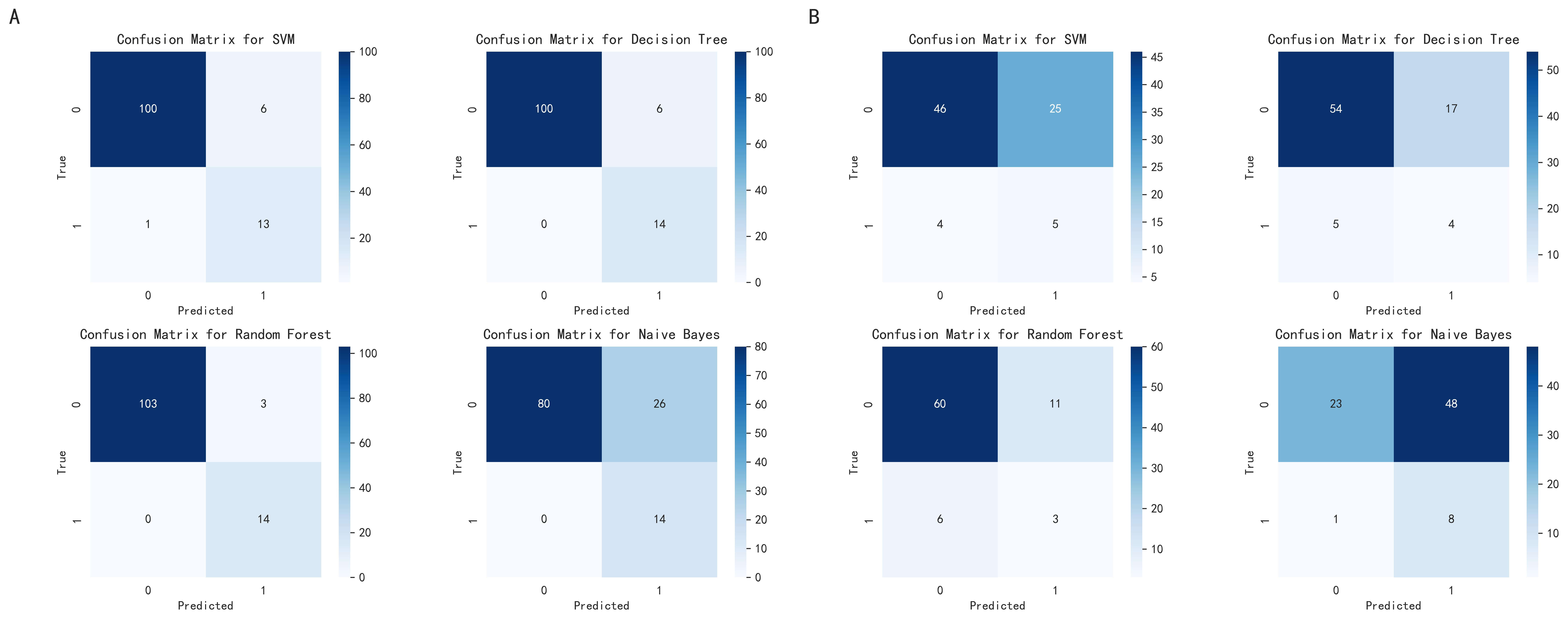 Fig. 6.
Fig. 6.
Prediction of serious adverse events. (A) Confusion matrices of the support vector machine, decision tree, random forest, and naïve Bayes models in the training set for predicting serious adverse events. (B) Confusion matrices of the same models in the independent test set. The matrices show the distribution of true-positive, true-negative, false-positive, and false-negative predictions for each model in classifying serious adverse events.
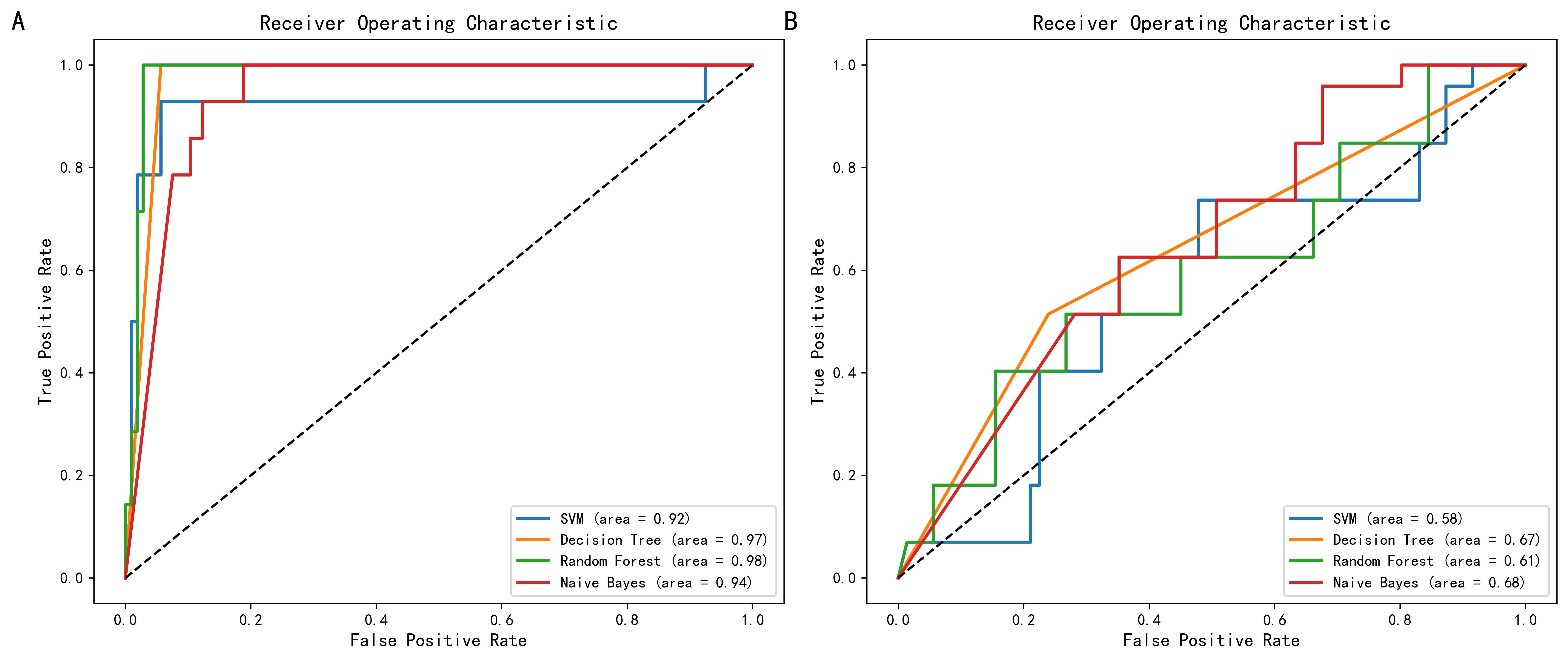 Fig. 7.
Fig. 7.
ROC of serious adverse events. (A) ROC curves of the support vector machine, decision tree, random forest, and naïve Bayes models in the training set for predicting serious adverse events. (B) ROC curves of the same models in the independent test set. The areas under the ROC curves indicate the ability of each model to discriminate patients with and without serious adverse events.
IONM has been widely adopted in neurosurgical procedures to preserve neurological function, particularly in surgeries involving lesions close to eloquent brain regions [13, 14, 15, 16, 17, 18]. However, the interpretation of SEP and MEP signals during surgery still relies heavily on the experience of neurosurgeons and electrophysiologists [19]. Individual variability in neural excitability, as well as lesion-induced displacement and deformation of functional areas, further complicates intraoperative decision-making [20, 21]. These challenges highlight the need for more objective and quantitative approaches to assist real-time surgical assessment.
In recent years, ML has been increasingly applied across multiple medical disciplines, including cardiology, neuroscience, critical care, and neurosurgery, to support clinical decision-making and reduce subjectivity [22, 23, 24, 25, 26, 27]. Rather than replacing clinician judgment, predictive models are primarily intended to serve as decision-support tools by integrating complex multidimensional data. In the context of IONM, ML approaches offer a potential framework to quantitatively interpret electrophysiological signals that are otherwise evaluated qualitatively [28, 29].
In the present study, we demonstrated that quantitative SEP and MEP parameters differ systematically according to whether IOLs involve functional brain areas. Previous studies have reported differences in the sensitivity of intraoperative MEP and SEP, during neurosurgical procedures. In certain surgical contexts, MEPs have been suggested to be more sensitive than SEPs for detecting motor pathway compromise and predicting immediate postoperative neurological dysfunction, particularly in surgeries involving eloquent motor areas [30, 31]. However, SEPs provide complementary information regarding sensory pathway integrity, and their relative sensitivity may vary depending on lesion location, surgical manipulation, and anesthetic conditions. In the present study, both MEP and SEP parameters demonstrated distinct correlation patterns with lesion localization, supporting their combined use in machine-learning–based predictive modeling. Correlation analysis suggested that SEP parameters showed a stronger association with lesion localization than MEP parameters, and that both modalities provided complementary information. Importantly, ML models that incorporated these electrophysiological features exhibited moderate discriminative ability in predicting functional-area involvement. Notably, although the predictive performance for postoperative serious adverse events appeared higher than that for lesion localization, the localization task represents a more challenging and clinically demanding objective [32]. These findings support the potential role of electrophysiological features as quantitative indicators for functional area localization, while highlighting the need for further model refinement to improve spatial precision.
Previous research has emphasized the importance of maximizing the extent of resection while preserving neurological function to improve patient outcomes. In this context, the predictive framework proposed in our study is not intended to determine surgical boundaries autonomously. Instead, it may provide supplementary information to inform intraoperative considerations, particularly in cases where functional boundaries are unclear or distorted by mass effect [33]. By integrating electrophysiological signals with clinical variables, such models may help contextualize intraoperative neurophysiological changes within a broader surgical decision-making process. Several limitations of the present study warrant consideration. First, although raw SEP and MEP waveforms were recorded, the electrophysiological features used for model development were limited to quantitative parameters such as amplitude and latency. While this approach improves feasibility and interpretability, it may not fully capture complex temporal or morphological signal characteristics. Second, SAEs represented a relatively rare outcome in the study cohort, resulting in a highly imbalanced class distribution and a limited number of positive cases. Although task-specific parameter tuning was performed, the predictive performance for SAE remained limited. This is likely attributable to the small sample size and the low prevalence of positive SAE cases, rather than to suboptimal model selection alone. Third, postoperative functional outcomes were primarily assessed through telephone follow-up rather than in-person neurological examinations. Consequently, limb function was evaluated using patient-reported scores rather than objective muscle strength testing, which may introduce subjective bias. Despite the limited predictive performance, explicitly reporting these constrained results is valuable for informing future study design, particularly with respect to outcome imbalance, feature selection, and model evaluation in IONM-based research. In summary, this study provides a proof-of-concept exploration of the feasibility and limitations of integrating SEP and MEP parameters into machine-learning–based analyses for investigating associations between functional-area involvement and postoperative outcomes in patients with intracranial space-occupying lesions. With further methodological refinement and external validation, such approaches may contribute to improving the objectivity and consistency of intraoperative neurophysiological interpretation.
In this study, we developed machine-learning models integrating intraoperative SEP and MEP parameters to explore their associations with functional-area involvement and postoperative outcomes in patients with intracranial space-occupying lesions. The results indicate that quantitative neurophysiological features derived from intraoperative monitoring are associated with lesion localization and serious adverse events. Although the predictive performance for functional-area localization was moderate, and the models showed limited ability to identify rare adverse outcomes, these findings highlight both the potential and the constraints of predictive modeling based on IONM data. Collectively, this work provides a proof-of-concept framework for future investigations, rather than a clinically deployable decision-support tool, and underscores the need for further validation in larger, well-balanced cohorts.
All primary data have been submitted as supplementary material. All data reported in this paper will also be shared by the lead contact upon request.
YZQ and XY designed the research study. ZHL performed the research. ZHL and PH analyzed the data. ZHL, XY and PH wrote the manuscript. QXC contributed to study design and the research methodology, and critically revised the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
The studies involving humans were approved by the Ethics Committee of the Renmin Hospital of Wuhan University. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study. The ethics approval number for this study is WDRY2025-K147. This study was conducted in accordance with the Declaration of Helsinki.
Not applicable.
This work was supported by the National Natural Science Foundation of China (No.82072764).
The authors declare no conflict of interest.
During the preparation of this work, the authors used ChatGPT-3.5 in order to check spelling and grammar. After using this tool, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/JIN47455.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.







