1 Department of Computer Engineering, Faculty of Engineering, Igdir University, 76000 Igdir, Turkey
2 Department of Electronics and Information Technologies, Faculty of Architecture and Engineering, Nakhchivan State University, AZ 7012 Nakhchivan, Azerbaijan
3 Informatics and Computer Systems Department, College of Computer Science, King Khalid University, 61421 Abha, Asir, Saudi Arabia
4 Department of Neurology, University of Health Sciences, Van Education and Research Hospital, 65000 Van, Turkey
5 Department of Radiology, University of Health Sciences, Van Education and Research Hospital, 65000 Van, Turkey
Abstract
Accurate and timely segmentation of ischemic stroke lesions from diffusion-weighted imaging (DWI) is crucial for diagnosis and treatment planning. Manual segmentation is labor-intensive, time-consuming, and prone to inter-observer variability. This study aims to develop and validate a novel deep learning framework that overcomes the common trade-off between high segmentation accuracy and the computational efficiency required for practical clinical use.
We developed FasterNet and Attention-Gated UNet (FA-UNet), a hybrid U-Net-based architecture. The model’s design features two key innovations: a computationally efficient FasterNet block at the bottleneck to capture global lesion context and multi-scale attention gates (MSAGs) on the skip connections to adaptively refine features and suppress noise. The model was trained and validated on the public Ischemic Stroke Lesion Segmentation (ISLES) 2022 dataset (n = 250 patients) and its performance was assessed on an independent, private test set of 600 DWI scans from 80 patients. FA-UNet’s performance was benchmarked against several state-of-the-art U-Net variants using the Dice coefficient, Intersection over Union (IoU), sensitivity, and precision as primary outcome measures.
On the independent test set (n = 80), the proposed FA-UNet model achieved a Dice coefficient of 0.8676 and an IoU of 0.7584. This performance surpassed all benchmarked architectures, including U-Net, U-Net3plus, and CMU-Net. Compared with the next best performing model, this represents a relative improvement of approximately 1.64% in the Dice score and 1.42% in IoU.
The FA-UNet architecture establishes a new state-of-the-art performance benchmark for automated ischemic stroke segmentation. By effectively balancing high accuracy with computational efficiency, it offers a robust, reliable, and clinically viable tool.
Keywords
- health
- deep learning
- ischemic stroke
- diffusion-weighted imaging
- image segmentation
- U-Net
Ischemic stroke represents a major global health issue due to its widespread occurrence and serious consequences, ranking among the top causes of death and long-term disability in adults [1]. Survivors often experience lasting impairments in movement, thinking, and communication, which drastically lower their quality of life and impose heavy socioeconomic costs [2]. Most strokes are ischemic in nature, resulting from blocked blood flow in the brain, which causes significant neuronal injury and neurological impairments [3]. Accurate and prompt identification and characterization of stroke lesions are critical for effective treatment planning and rehabilitation [4].
Conventional medical imaging methods, especially magnetic resonance imaging (MRI) and computed tomography (CT), play a vital role in the diagnosis and management of ischemic stroke [5]. In the acute phase, MRI techniques like diffusion-weighted imaging (DWI) and perfusion-weighted imaging (PWI) are particularly valuable, offering precise information about the size, location, and blood flow characteristics of the infarct [6]. For chronic stroke evaluation, T1-weighted (T1W) MRI remains essential, thanks to its high spatial resolution, which enables the monitoring of lesion progression and long-term structural changes in the brain [7, 8].
Lesion segmentation, the accurate outlining of damaged brain tissue, has been performed manually by skilled radiologists [9, 10]. While these methods are known for their precision, they come with considerable drawbacks, such as being time-consuming and subject to variations between and within observers [11]. These challenges underscore the pressing need for automated segmentation solutions that offer greater consistency, speed, and reliability.
Recent advances in machine learning, particularly deep learning methods, have shown considerable promise in medical image analysis tasks, significantly advancing the field of lesion segmentation [12, 13]. Among these methods, convolutional neural networks (CNNs) have become the cornerstone due to their powerful capability to automatically learn and extract intricate spatial features directly from raw imaging data without explicit manual feature engineering [14, 15]. CNNs excel in capturing complex hierarchical representations of images, effectively addressing variations in scale, texture, and contextual information relevant for accurate lesion detection [16, 17].
The U-Net architecture, in particular, has revolutionized medical image segmentation tasks with its novel encoder-decoder structure complemented by skip connections [18]. This architecture effectively retains spatial information lost during down-sampling in the encoder stage and restores high-resolution features through the decoder, crucial for precise delineation of lesions [19]. By integrating skip connections, U-Net facilitates the fusion of low-level and high-level image features, thus enhancing the accuracy of multi-scale feature extraction. Its simplicity, computational efficiency, and effectiveness in handling limited datasets have contributed to its widespread adoption across various medical segmentation tasks, including stroke lesion segmentation [20]. Subsequent enhancements to the U-Net framework have further elevated its segmentation performance. Variants incorporating attention mechanisms, such as Attention U-Net, introduce selective focus on relevant spatial regions, enhancing model sensitivity to critical features. Dense U-Net variants employ densely connected layers to promote efficient gradient flow and feature reuse, thereby boosting segmentation robustness. More recent developments have integrated transformer-based architecture, capitalizing on self-attention mechanisms to capture both local and global context [17, 21]. Such approaches provide substantial improvements in handling the spatial complexity and variability inherent in stroke lesions. However, despite these advancements, significant computational demands associated with transformer-based models necessitate careful optimization and efficiency improvements for practical clinical deployment.
Despite the advances brought by U-Nets and related deep learning methods, several critical challenges persist in ischemic stroke lesion segmentation [22]. Lesions typically exhibit significant variability in size, shape, and intensity, complicating accurate model training and segmentation results [19, 23]. Furthermore, distinguishing ischemic lesions from healthy brain tissue is challenging due to low contrast and subtle differences in intensity, especially in regions affected by partial volume effects [24]. These issues result in segmentation errors, limiting the clinical applicability and generalization of current models. Efforts to overcome these limitations include developing models with enhanced boundary recognition capabilities, multi-scale feature extraction, and improved generalization techniques. Approaches utilizing attention mechanisms and transformer-based architecture have also been explored, capitalizing on their ability to integrate both global and local contextual information [22, 25]. Nevertheless, the computational complexity of these methods often restricts their practical deployment in clinical settings. Additionally, the scarcity of adequately annotated imaging datasets remains a substantial barrier to developing robust and generalizable models. Techniques such as weakly supervised learning and active learning frameworks have been proposed to mitigate the dependence on extensive manual annotations, promoting more efficient and effective model training [26]. Thus, while considerable advancements have been achieved in automated lesion segmentation for ischemic stroke, significant gaps remain, necessitating further research and innovation. Addressing these challenges through improved methodologies and efficient utilization of available data is critical for advancing stroke diagnosis, treatment, and rehabilitation outcomes.
In this study, we propose FasterNet and Attention-Gated UNet (FA-UNet), a novel deep learning framework designed to overcome the key challenges in ischemic stroke segmentation. Existing methods often struggle with the high variability of lesions, low image contrast, and the computational demands of advanced models. Our model addresses these issues by integrating a highly efficient FasterNet block at the bottleneck of a U-Net architecture and a multi-scale attention gate (MSAG) mechanism on the skip connections. This design allows for both robust feature extraction and intelligent feature refinement. The primary contributions of this research are as follows.
(1) This research proposes FA-UNet, a novel hybrid deep learning architecture that is specifically designed to enhance the accuracy and robustness of ischemic stroke segmentation by addressing key challenges such as lesion heterogeneity and low image contrast.
(2) The architecture introduces a FasterNet block at its bottleneck, which replaces standard convolutions with a more efficient operator that utilizes Partial Convolutions (PConvs) to reduce computational redundancy and memory access. This efficiency allows the model to effectively learn long-range spatial dependencies and global contextual information, critical for accurately segmenting large, heterogeneous lesions and maintaining their structural integrity.
(3) The model integrates a MSAG on its skip connections to provide an intelligent feature refinement mechanism. By processing features through three parallel branches with different receptive fields, the MSAG creates a dynamic spatial attention map that adaptively amplifies salient lesion characteristics while suppressing background noise, crucial for improving detection sensitivity and boundary precision in low-contrast DWI images.
(4) The framework’s performance was confirmed through a rigorous validation on gold-standard DWI scans, using the public Ischemic Stroke Lesion Segmentation (ISLES) 2022 dataset for training and a private dataset for testing. This evaluation demonstrated the model’s superiority over several state-of-the-art U-Net variants, establishing a new benchmark with a Dice coefficient of 0.8676 and an Intersection over Union (IoU) of 0.7584.
The automated segmentation of ischemic stroke lesions from medical images has become a significant area of research, with deep learning techniques, particularly those based on CNNs, emerging as the state-of-the-art. Numerous studies have focused on developing robust models for both CT and MRI scans, with a strong emphasis on enhancing segmentation accuracy, a critical step for clinical diagnosis and treatment planning. A deep novel learning using a feature-enhanced network (FEN) model was presented to accurately segment stroke lesions in brain MRI scans [27]. It integrates multi-scale features and attention mechanisms to better distinguish stroke-affected regions from healthy tissue. Wang et al. [28], proposed a novel approach to improve the segmentation of acute ischemic stroke (AIS) lesions in non-contrast CT (NCCT) brain scans using a hybrid knowledge distillation strategy. This work combined offline knowledge distillation with self-distillation to help the segmentation model learn more effectively from limited and low-contrast NCCT data, which traditionally presents challenges due to poor lesion visibility. A feature selection using a wrapper-based approach was presented to identify the best combination of MRI sequences (modalities) for accurately segmenting ischemic stroke lesions [29]. The method evaluates various subsets of MRI sequences and selects the most effective sequences to improve segmentation accuracy, reduce computational cost, and minimize data redundancy. Medical imaging and machine learning were utilized to support clinicians in rapid, accurate stroke evaluation, potentially improving treatment decisions and patient outcomes [26]. Another advanced deep learning model used DWI for accurately segmenting cerebral vascular occlusions [30]. The novel segmentation model integrates ConvNeXtV2’s strong feature extraction capabilities and Global Response Normalization (GRN) ability to stabilize and enhance training dynamics, leading to better localization of vascular occlusions.
Segmentation–reSegmentation Net (SrSNet) is a novel deep learning model that aims to improve the accuracy of stroke lesion segmentation in brain imaging [31]. The study proposed a two-stage framework: the first stage roughly identifies potential lesion areas, and the second stage refines these predictions with greater precision. By comparing mirrored brain regions, the model enhances its ability to distinguish lesions from normal tissue. Sun et al. [25], introduced a novel deep learning framework to improve the diagnosis of acute ischemic stroke using minimal labeled data. A multi-grained contrastive learning approach combined unsupervised pretraining with contrastive learning at multiple levels and utilizes both global and local features from brain MRI scans to enhance the model’s ability to segment stroke lesions and classify the time from stroke onset and learns rich and discriminative representations. Stroke lesions in brain MRI scans were accurately identified using advanced deep learning [24]. The proposed model fuses feature from multiple deep neural network layers, combining both low-level texture and high-level semantic information to improve segmentation performance. By integrating these diverse features, the model captures more comprehensive lesion characteristics, leading to enhanced accuracy and robustness in detecting acute ischemic stroke regions. Multimodal difference aware network (MDANet), a deep learning framework, was developed to improve brain stroke lesion segmentation by effectively [21]. The model utilized multimodal MRI data to learn the differences and complementary information across multiple MRI modalities (like DWI, T2-weighted imaging (T2WI), etc.) and therefore enhance the accuracy of stroke detection. Segmenting brain lesions caused by acute ischemic stroke was analyzed using deep learning [32]. The model developed to accurately detect and outline the stroke-affected brain regions. Additionally, the study explores how specific features extracted from these lesions—such as size, location, or intensity—relate to patient outcomes measured by the modified Rankin Scale (mRS). Res2U++, a novel deep learning architecture developed to improve the segmentation of ischemic stroke lesions in brain imaging [20]. Built upon U-Net and Res2Net principles, Res2U++ incorporates multi-scale feature extraction and dense skip connections to capture both fine-grained and contextual information, which is critical for accurate lesion detection.
Advanced deep learning approaches have been widely used to analyze and detect medical images in the and particularly ischemic stroke applications, such as stroke detection, lesion segmentation, perfusion analysis, and outcome prediction using CT and MRI scans [11, 33, 34, 35, 36]. Rahman et al. [33], presented a deep learning model based on CNNs to automate the segmentation process, improving both accuracy and consistency compared to traditional manual methods. EnigmaNet is a novel deep learning framework that enhances performance by integrating attention mechanisms that help the network focus on critical lesion features while suppressing irrelevant background noise [34]. Transformer-based architectures with CNNs techniques have been used to segment stroke lesions in brain MRI scans [23, 37]. Circular feature interaction (CFI) has been used with hybrid CNN and Transformer network to improve the automatic segmentation of AIS lesions in NCCT scans [38]. The module enables more effective feature exchange between different layers and paths within the network. Pathological Asymmetry-Guided Progressive Learning (PAGPL) is a deep learning model that was proposed to improve the accuracy of AIS infarct segmentation from brain imaging [39]. It leverages the natural anatomical symmetry of the brain to better detect stroke-related abnormalities. Bal et al. [40] proposed Two-pathway 3D deep neural network to address challenges posed by stroke lesion variability in shape, size, and intensity across different patients. The model incorporates two parallel processing paths: a local path that focuses on fine-grained features for detailed lesion boundaries, and a global path that captures broader contextual information from the surrounding brain structures.
A comprehensive review of the recent literature reveals several key trends and strategies. The field is predominantly characterized by the enhancement of U-Net-based architecture through various sophisticated modules designed to improve feature representation. These include multi-scale feature fusion networks that capture both fine-grained and contextual information, and the integration of attention mechanisms to help models focus on salient lesion regions while suppressing background noise. More recently, hybrid architecture combining CNNs with Transformers have been explored to leverage the latter’s strength in modeling long-range spatial dependencies. Other notable strategies involve leveraging anatomical priorities like brain symmetry, employing two-stage refinement processes for greater precision, and using knowledge distillation to overcome the challenges of limited or low-contrast data. While these advanced methods have progressively improved segmentation accuracy, a persistent challenge lies in balancing high performance with computational efficiency, a crucial factor for practical deployment in time-sensitive clinical settings. Many powerful models, particularly those incorporating Transformers or complex multi-pathway systems, incur significant computational overhead, which can limit their applicability. The present study addresses this critical trade-off. Our proposed model, FA-UNet, is designed to achieve state-of-the-art accuracy while maintaining high computational efficiency. By integrating a novel FasterNet block at the network’s bottleneck, we effectively capture global context with significantly lower latency than conventional methods. This is synergistically combined with a MSAG that intelligently refines features with different resolutions. By doing so, our work aims to bridge the gap between high-performance, complex models and the need for fast, reliable, and clinically viable segmentation tools.
This section presents the comprehensive technical framework of our proposed network, FA-UNet, designed for the precise and efficient segmentation of ischemic stroke lesions in DWI. We provide a detailed exposition of the network’s architecture, the dataset and preprocessing protocols, the implementation and training strategy, and the quantitative metrics employed for performance evaluation. The architectural design of FA-UNet, illustrated in Fig. 1, is founded on a hybrid U-Net structure that synergizes a computationally efficient backbone with an advanced feature fusion mechanism to address the key challenges of lesion heterogeneity and low image contrast inherent in stroke imaging.
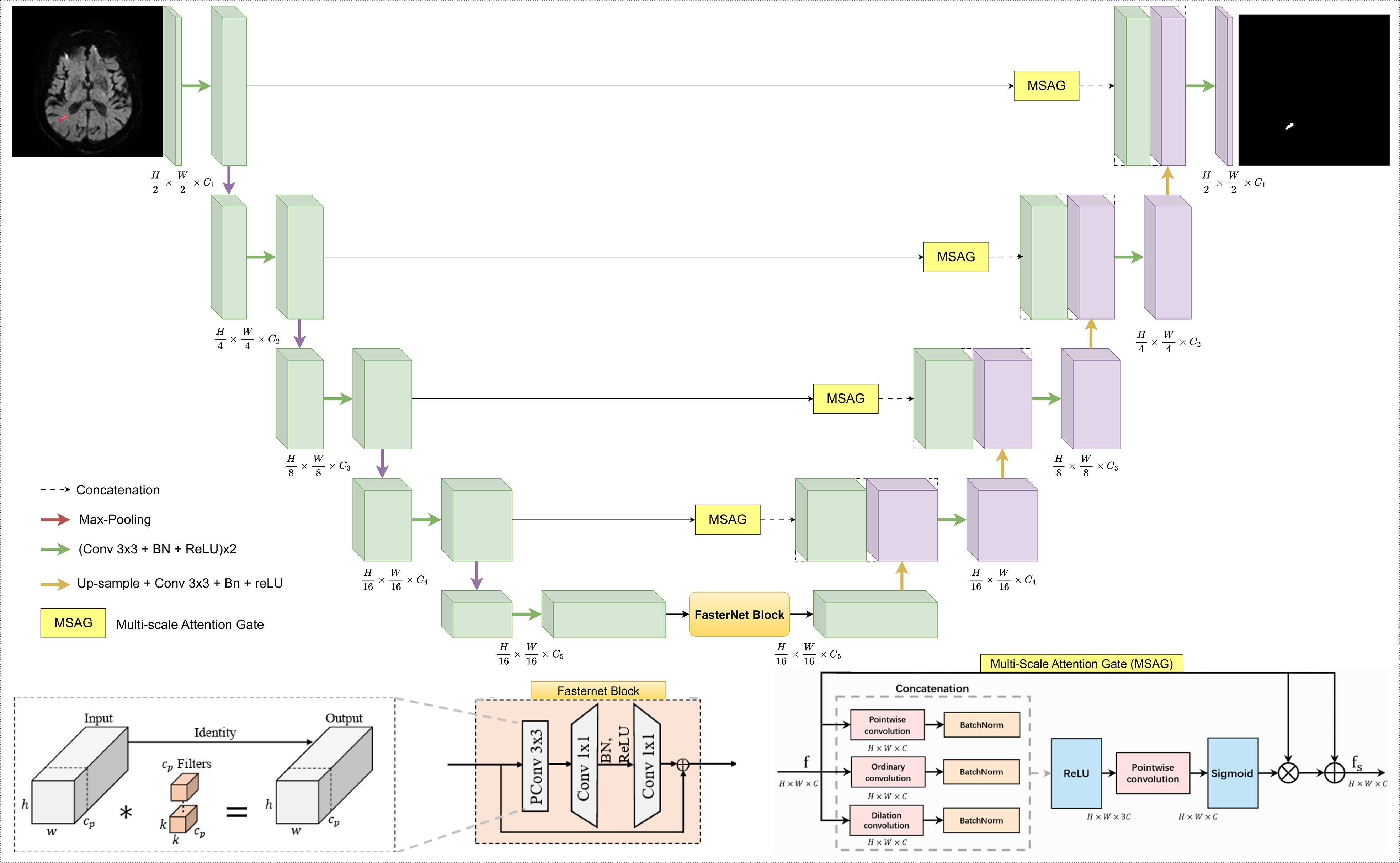 Fig. 1.
Fig. 1.
Detailed architecture of the proposed FA-UNet model. The network employs a symmetric encoder-decoder structure enhanced with two key components: a FasterNet block at the bottleneck to efficiently capture global features, and MSAGs on the skip connections. The MSAGs function to suppress irrelevant information and emphasize salient lesion characteristics from the encoder path before fusion, improving the accuracy of the final segmentation. FA-UNet, FasterNet and Attention-Gated UNet; Conv, convolution; BN, batch normalization; ReLU, rectified linear unit; MSAGs, multi-scale attention gates; PConv, partial convolution.
The proposed FA-UNet adopts the symmetric encoder-decoder structure of the U-Net architecture, a proven foundation for medical image segmentation. This foundational structure is enhanced with two principal architectural innovations: a computationally efficient FasterNet block positioned at the network’s deepest point (the bottleneck), and multi-scale attention gates (MSAGs) integrated into the skip-connections that link the encoder to the decoder. The design philosophy of FA-UNet is to resolve a fundamental dilemma faced by modern deep learning architectures. While standard CNNs struggle to effectively capture global contextual information due to their inherently limited receptive fields, Transformer-based models, which excel at this task, often incur significant computational costs that can restrict their practical deployment. FA-UNet aims to establish an efficient and powerful balance between these two approaches. The core strategy of architecture is to efficiently capture global context using the FasterNet block and to intelligently fuse this information with local, high-resolution details via MSAG. This hybrid design, the schematic of which is illustrated in Fig. 1, is specifically engineered to address the core challenges of ischemic stroke segmentation, namely the high variability in lesion size, shape, and intensity, as well as low image contrast.
The encoder path of FA-UNet functions as a hierarchical feature extractor, progressively down-sampling the input DWI image to capture increasingly complex and abstract semantic features. This path culminates in a highly efficient bottleneck that processes the most abstract feature representations.
The main body of the encoder consists of a series of standard encoding blocks.
Each block comprises two consecutive 3
At the deepest point of the encoder, where the feature maps contain the most abstract semantic information, the standard convolutional blocks are replaced by a FasterNet Block. FasterNet was proposed by Chen et al. [41] in 2023 to show that lowering the arithmetic count of a network is not enough to shorten inference time unless the design also permits a high rate of executed operations per second on real devices. Depthwise convolution, the usual choice in mobile architectures, reduces theoretical floating-point operations but spends so much time on memory transfers that its realised throughput floating-point operations per second (FLOPs) stays low. To break that bottleneck the authors replaced depthwise layers with a partial convolution followed by two pointwise convolutions, creating the macro-unit now called the FasterNet Block, as depicted in Fig. 1. This architectural choice is a direct response to a critical challenge in designing efficient neural networks: simply reducing the theoretical number of FLOPs does not guarantee a proportional decrease in actual inference latency. Many efficient operators, such as the commonly used depthwise convolution, suffer from low computational speed (measured in FLOPS) due to frequent and inefficient memory access, which becomes the primary performance bottleneck. To break the memory access bottleneck, the FasterNet architecture is built around a novel partial convolution (PConv). PConv is designed to reduce both computational redundancy and memory access simultaneously by exploiting the high similarity often found across different channels of a feature map. Instead of applying filters to all input channels, PConv applies a standard convolution to only a partial set of the input channels (cp), while the remaining channels are left untouched through an identity mapping. This design drastically reduces the computational load and memory access. The FLOPs of a PConv and its memory access are given by Eqn. 1 and Eqn. 2.
To ensure that information from all channels is effectively aggregated, the
PConv layer is followed by a standard pointwise convolution (PWConv), which
performs a 1
The complete FasterNet Block, as implemented in FA-UNet, consists of a PConv layer followed by two PWConv layers, forming an inverted residual block structure where the channel count is expanded and then projected back down. The total floating-point operations for this entire block can be calculated in Eqn. 4.
This design not only achieves high practical speed (high FLOPS) but is also highly effective at feature transformation, having been shown to approximate the representational power of a standard convolution with much greater efficiency. The performance enhancement in ischemic stroke segmentation stems from two primary advantages. First, the block’s high computational efficiency reconciles speed and power, allowing FA-UNet to incorporate a deep and powerful feature extractor at the bottleneck without incurring prohibitive latency. This is crucial for developing models that are both accurate and practical for time-sensitive clinical applications like acute stroke assessment. Second, and equally important, is the block’s superior contextual modeling for lesion delineation. Its ability to efficiently model long-range spatial dependencies is vital for segmenting stroke lesions, which are often highly heterogeneous in size and shape. By perceiving the entire context of a large or irregularly shaped lesion, the FasterNet bottleneck enables FA-UNet to generate more coherent and complete segmentation masks. This global understanding helps the model to distinguish true lesion areas from mimics and reduces the risk of fragmenting a single large infarct into multiple, disconnected parts, thereby improving the overall accuracy of lesion volume assessment.
The decoder path is responsible for reconstructing a high-resolution segmentation map from the low-resolution, abstract features generated by the encoder. It progressively up-samples the feature maps while intelligently fusing them with high-resolution information from the encoder via skip connections.
The decoder mirrors the encoder’s structure, consisting of a series of
upsampling blocks. At each stage, the feature map from the previous, deeper layer
is up-sampled using a 2
Before the feature maps from the skip connections are fused with the decoder’s up-sampled features, they are passed through a MSAG [42]. The purpose of this module is to address a fundamental limitation of standard U-Net architecture: naive skip connections pass unrefined information that contains both valuable, high-resolution boundary details and irrelevant background noise or imaging artifacts. In the context of ischemic stroke segmentation, this can be particularly problematic, as the subtle signal from a low-contrast lesion can be obscured by noise or similarly intense healthy tissue, leading to inaccurate or incomplete segmentation. The MSAG functions as an adaptive feature refinement module that intelligently filters this information to suppress unimportant features and enhance the valuable ones, thereby improving the overall precision and robustness of the model, as illustrated in Fig. 1.
The core strength of the MSAG lies in its ability to analyze features at
multiple scales simultaneously, which is an essential capability for handling the
high heterogeneity of ischemic stroke lesions. To achieve this, the incoming
feature map is processed through three parallel convolutional branches, each
designed to extract a distinct type of feature. The first is a pointwise
convolution, which has a 1
The outputs from these three branches, each followed by a Batch Normalization layer, are concatenated and passed through ReLU activation. A final pointwise convolution followed by a Sigmoid activation function then generates a dynamic spatial attention map. This map acts as a learned, data-driven filter, assigning an important score between 0 and 1 to each pixel in the feature map. This recalibration process is crucial for improving performance in low-contrast DWI images, as the attention map can learn to amplify the faint signals of an ischemic lesion while suppressing confounding signals from background noise or healthy tissue, such as the periventricular white matter. This selective amplification leads to both higher detection sensitivity and fewer false positives, resulting in more precise boundary delineation. This entire process is formally defined in Eqn. 5.
Here,
The primary dataset used for training and validation in this study is the ISLES 2022 dataset, which was developed in collaboration with Medical Image Computing and Computer Assisted Interventions (MICCAI) and other institutions and serves as a crucial resource for research in brain stroke segmentation [43]. The main goal of this dataset is to facilitate the accurate segmentation of acute and subacute stroke lesions from MRI scans collected from various clinical centers. It consists of 400 MRI cases from a range of imaging sources, covering a wide spectrum of stroke lesion characteristics. Of these, 250 cases are publicly accessible for training purposes, while the remaining 150 are reserved for independent evaluation. The inclusion of fluid-attenuated inversion recovery (FLAIR), DWI, and apparent diffusion coefficient (ADC) sequences in ISLES 2022 offers complementary diagnostic information that significantly improves the accuracy of lesion detection.
For the purpose of this study, only DWI scans, which are of paramount importance in stroke assessment, were utilized. A selection of 2600 images from 250 patients in the ISLES 2022 dataset, chosen by radiologists (excluding images with very small pixels and some unclear details), was used, with 80% of this data allocated for training and the remaining 20% for validation. The test data was obtained from Van Education and Research Hospital between 2024 and 2025, with the ethical approval number 60KAEK/2025-02-23. This test set comprises a total of 600 images from 80 patients. Fig. 2 displays several randomly selected DWI images from the ISLES 2022 dataset and our private dataset with their corresponding ground truth masks, providing a clear view of the raw data and its annotated counterparts. As illustrated in Fig. 2, the DWI images and their ground truth masks not only show the typical appearance of ischemic stroke lesions but also underscore the inherent variability and complexity in how these lesions present.
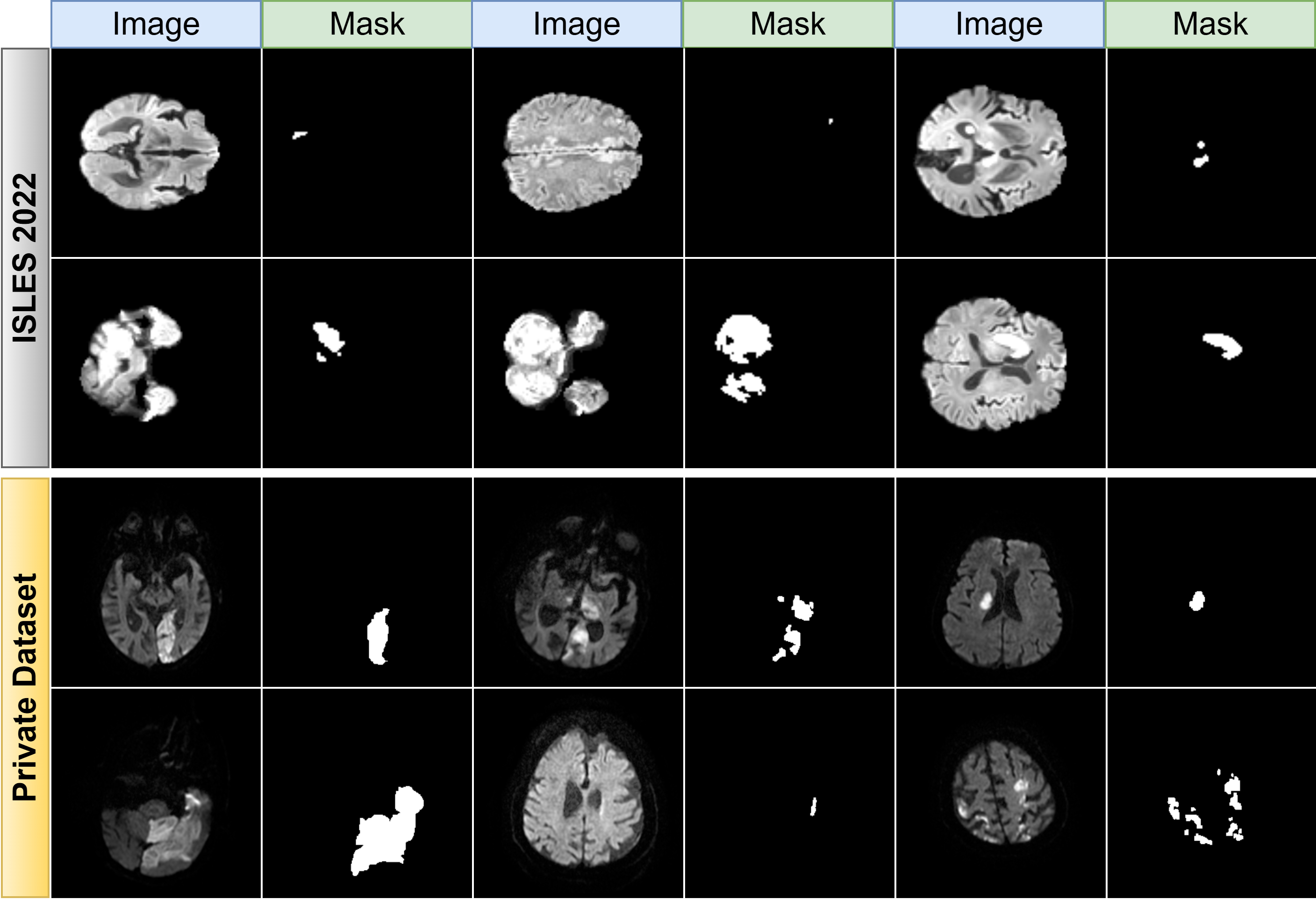 Fig. 2.
Fig. 2.
Representative DWI samples from ISLES 2022 and private datasets. Examples of diffusion-weighted imaging (DWI) scans and corresponding ground truth masks. The top panel shows samples from the public Ischemic Stroke Lesion Segmentation (ISLES) 2022 dataset (training/validation), and the bottom panel shows samples from the private test dataset, illustrating the data’s heterogeneity.
This study exclusively utilized DWI scans, selected for their paramount
importance in acute stroke assessment. The primary data for model training and
validation were sourced from the public ISLES 2022 challenge dataset. To optimize
this data for robust model training, a meticulous curation process was conducted
collaboratively by a senior radiologist, with over seven years of experience in
neuroradiology, and a specialist neurologist. Based on their joint clinical
assessment, a final subset of 2600 images from 250 patients was established. This
curation focused on excluding scans with ambiguous details or certain
micro-lesions, particularly in cases presenting multiple infarcts, to create a
more distinct and reliable dataset. The primary objective of this refinement was
to facilitate a more effective and focused training process for the segmentation
models. This data was then partitioned, with 80% allocated for training and 20%
for validation. To enhance model robustness and mitigate overfitting, a
sophisticated data augmentation strategy was systematically applied to the
training set. This regimen included geometric transformations (random rotations,
flips, elastic and grid distortions) and photometric augmentations (random
brightness and contrast adjustments). For independent testing, a separate cohort
of 600 images from 80 patients was prospectively collected at Van Education and
Research Hospital. All images were resized
to a uniform 256
The training protocol was executed on a high-performance system equipped with an NVIDIA RTX 4090 GPU (NVIDIA Corporation, Santa Clara, CA, USA) and an Intel i9 14900K processor (Intel Corporation, Santa Clara, CA, USA), using a batch size of 16. We employed the AdamW optimizer for its robust performance. The loss function chosen for this segmentation task was BCEDiceLoss, which combines Binary Cross-Entropy (BCE) and Dice Loss to leverage the strengths of both pixel-level accuracy and spatial overlap assessment. Its mathematical formulation is depicted in Eqn. 6.
In this composite function,
The performance of the proposed segmentation framework was quantitatively benchmarked against established standards using a comprehensive suite of evaluation metrics. Spatial segmentation fidelity was primarily assessed using the Dice Coefficient, which measures the normalized overlap between the predicted and ground-truth masks, and the IoU, which quantifies their degree of concordance. For a more granular analysis of pixel-level classification performance, we employed a set of complementary metrics. Sensitivity (Recall) was calculated to determine the model’s effectiveness in identifying all actual lesion pixels, while Precision was used to measure the reliability of the positive predictions made by the model. To assess the crucial ability to correctly identify healthy, non-pathological tissue and thus FP, Specificity was also computed. Finally, the F1-Score, as the harmonic mean of Precision and Sensitivity, provided a single, balanced score reflecting overall segmentation accuracy, particularly valuable in the context of the inherent class imbalance of stroke lesion datasets. All metrics were derived from the cardinal counts of true positives (TP), true negatives (TN), FP, and false negatives (FN) established through a pixel-wise comparison of the predicted outputs against the ground-truth annotations. These metrics are defined in Eqns. 7,8,9,10,11,12.
This section details the quantitative evaluation of our proposed FA-UNet model. The model was initially trained and validated on a curated set of 2600 DWI MR images from the public ISLES 2022 benchmark dataset. To test its generalization capabilities and performance on unseen clinical data, we subsequently evaluated the model on a distinct, independent dataset. This private test set, collected at the Van Education and Research Hospital, consists of 600 DWI scans from 80 patients. On this independent dataset, FA-UNet’s performance was rigorously benchmarked against several influential U-Net variants, including U-Net [44], U-Net3plus [45], CMU-Net [42], CMU-NeXt [46], AttentionU-Net [47], and U-NeXt [48]. To ensure a fair and direct comparison, all of these comparative models were retrained from scratch using the identical training protocol, data splits, and experimental setup as our proposed FA-UNet. The comparison was based on a suite of key metrics assessing spatial accuracy (Dice coefficient and IoU) and diagnostic reliability (Sensitivity, Precision, and F1-score). The comprehensive results of this comparative analysis are summarized in Table 1, providing a clear overview of each model’s segmentation performance.
| Model names | IoU | Dice | Sensitivity | Precision | F1-score |
| U-Net | 0.7323 | 0.8432 | 0.8348 | 0.8547 | 0.8432 |
| U-Net3plus | 0.7457 | 0.8526 | 0.8439 | 0.8649 | 0.8526 |
| CMU-Net | 0.7478 | 0.8536 | 0.8502 | 0.8604 | 0.8536 |
| CMU-NeXt | 0.7360 | 0.8460 | 0.8536 | 0.8416 | 0.8460 |
| AttentionU-Net | 0.7429 | 0.8507 | 0.8413 | 0.8633 | 0.8507 |
| U-NeXt | 0.7275 | 0.8403 | 0.8391 | 0.8462 | 0.8403 |
| FA-UNet (Proposed) | 0.7584 | 0.8676 | 0.8642 | 0.8723 | 0.8682 |
FA-UNet, FasterNet and Attention-Gated UNet; IoU, Intersection over Union.
The results presented in Table 1 provide a clear and consistent demonstration of the proposed FA-UNet model’s superior performance across every evaluation metric when compared against a range of influential state-of-the-art architectures. From the perspective of spatial accuracy, the Dice coefficient and IoU are the most critical metrics. Our FA-UNet model achieves the highest scores with a Dice of 0.8676 and an IoU of 0.7584. This represents a significant margin over the next-best performer, CMU-Net (Dice: 0.8536), and foundational models like U-Net (Dice: 0.8432). This marked improvement indicates a more precise delineation of lesion boundaries and a more accurate estimation of lesion volume, which are vital for clinical assessment. The lowest performance in these metrics belongs to U-NeXt, suggesting that its multilayer perceptron (MLP)-based approach may be less effective for this specific segmentation task compared to advanced convolutional methods.
Analyzing the trade-off between Sensitivity and Precision reveals deeper insights into model behavior. FA-UNet leads in both metrics, with a Sensitivity of 0.8642 and a Precision of 0.8723. This is a crucial finding, as it shows our model not only excels at identifying true lesion areas (high sensitivity, minimizing missed strokes) but also maintains high reliability in its predictions (high precision, minimizing false alarms). Other models exhibit a more pronounced trade-off; for example, CMU-NeXt has high sensitivity (0.8536) but relatively low precision (0.8416), suggesting a tendency for over-segmentation. Conversely, U-Net3plus shows high precision (0.8649) but a comparatively lower sensitivity (0.8439). FA-UNet’s ability to lead in both demonstrates a more robust and well-balanced diagnostic capability.
Considering the chronological evolution of these architectures, a clear performance trend is visible. The original U-Net sets a strong baseline. Subsequent models like AttentionU-Net (2018) and U-Net3plus (2020) show incremental gains by integrating attention mechanisms and denser connectivity. Interestingly, while CMU-Net (2023) represents a peak among the compared State-of-the-Art (SOTA) models, the even more recent CMU-NeXt (2024) shows a slight dip in overall performance, indicating that architectural novelty does not always guarantee superiority. Our proposed FA-UNet surpasses even the most recent and highest-performing models in this list, demonstrating a significant leap forward rather than a mere incremental improvement. A graphical comparison presented in Fig. 3 provides clear visual corroboration of these quantitative results, highlighting the significant performance gap between the proposed FA-UNet and the other architectures.
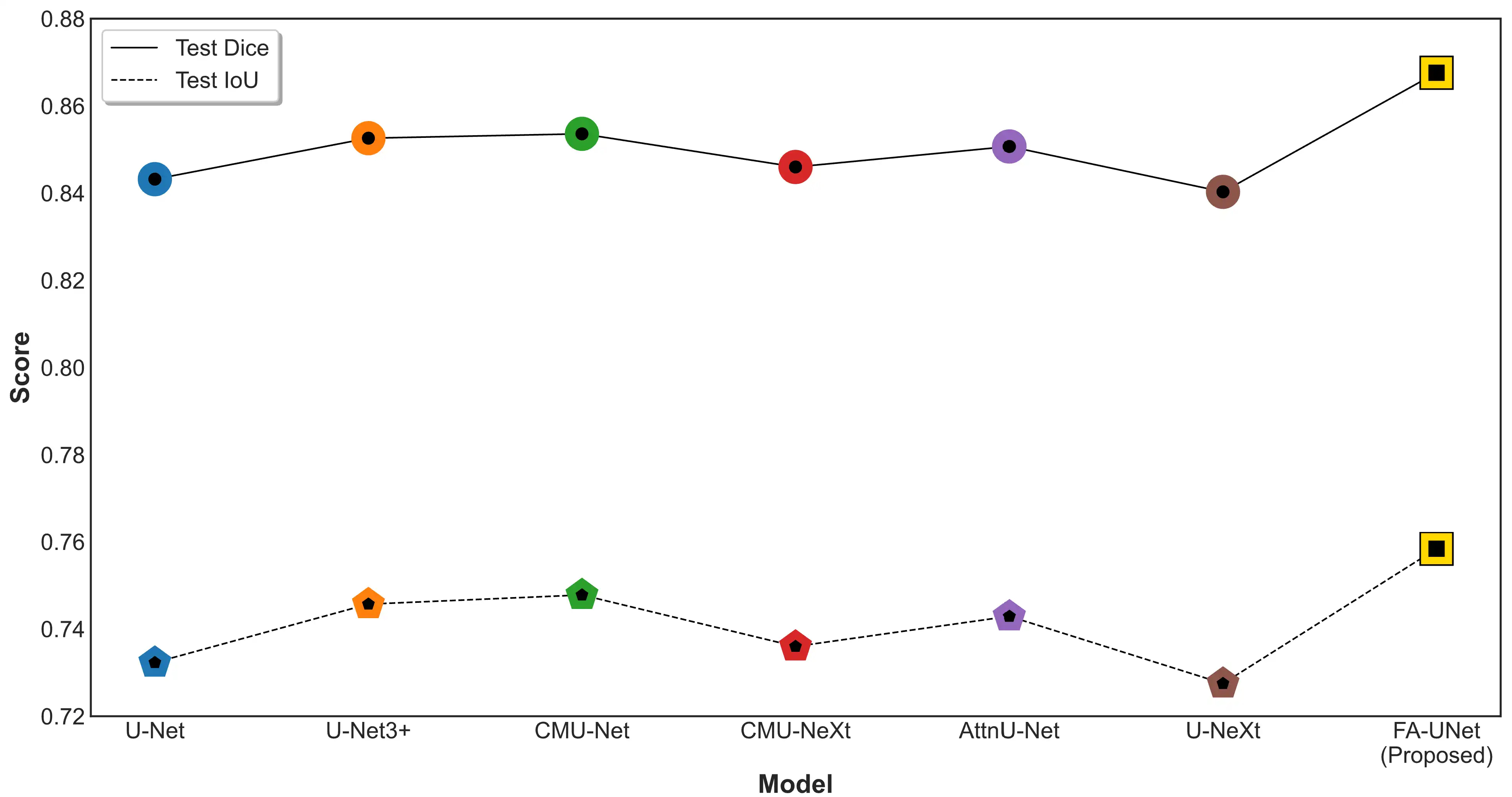 Fig. 3.
Fig. 3.
Comparative analysis of segmentation performance on the independent test set. The plot compares the Test Dice (solid line) and Test IoU (dashed line) scores for the proposed FA-UNet against several U-Net-based variants. AttnU-Net, Attention U-Net.
As depicted in Fig. 3, the proposed FA-UNet model demonstrates a clear and consistent performance advantage over all benchmarked U-Net variants. In terms of the Dice coefficient, which measures spatial overlap, FA-UNet not only surpasses the other models but also shows a significant leap in performance compared to the next-highest competitor, CMU-Net. This trend is mirrored in the IoU scores, where FA-UNet again occupies the highest position, visually confirming the quantitative results and highlighting a consistent and meaningful improvement in segmentation accuracy. This graphical evidence substantiates that the architectural enhancements in FA-UNet translate directly to a more robust and precise segmentation capability. Fig. 4 displays segmentation results on two representative cases from the test set.
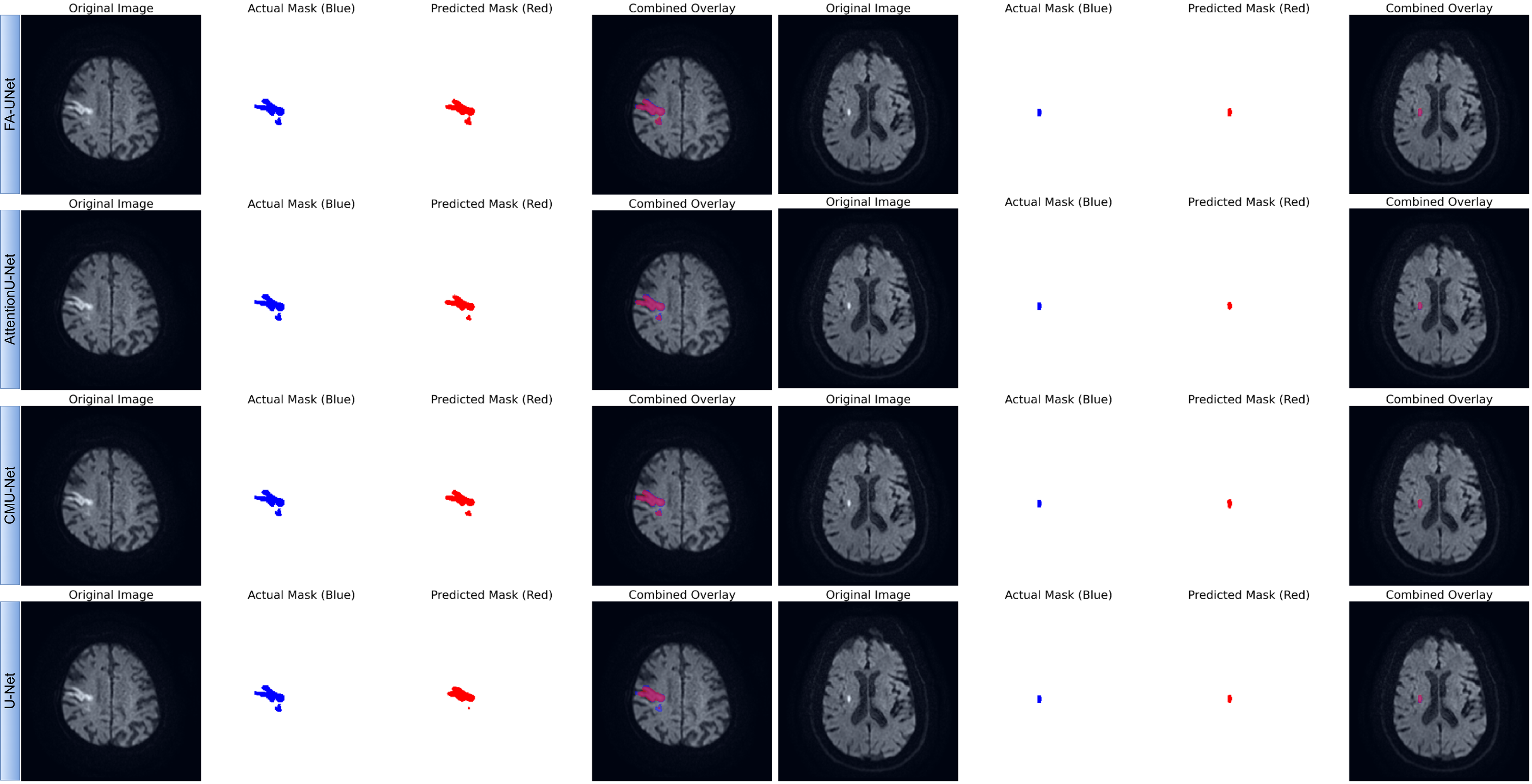 Fig. 4.
Fig. 4.
Qualitative comparison of segmentation performance for FA-UNet and benchmark models. The figure presents segmentation results on two representative cases from the test set: a large, morphologically complex lesion (left panel) and a small, well-defined lesion (right panel). For each model, the predicted mask (red) is compared against the ground truth mask (blue). The ‘Combined Overlay’ visualizes their spatial agreement, where the overlapping region appears pink or purple.
To qualitatively evaluate the segmentation performance, Fig. 4 presents a visual comparison between the proposed FA-UNet and other benchmark models on two representative cases from the test set. The results for the large, morphologically complex lesion (left panel) clearly demonstrate the superiority of our proposed model. FA-UNet successfully delineates the entire lesion, achieving high spatial overlap with the ground truth mask and accurately capturing its intricate boundaries. In contrast, the other architectures, including AttentionU-Net and CMU-Net, exhibit significant under-segmentation, failing to encompass the full extent of the pathological tissue. The baseline U-Net shows the most pronounced limitation, identifying only the core of the lesion. While all models perform adequately on the smaller, well-defined lesion (right panel), the more challenging case underscores the enhanced robustness of our approach. These visual findings provide strong evidence for the architectural advantages of FA-UNet. The model’s ability to generate a complete and coherent segmentation for the complex infarct visually validates the effectiveness of the FasterNet block in capturing global contextual information and the role of the MSAG in refining boundary details. This qualitative superiority, which corroborates our quantitative metrics, underscores the model’s potential for clinical application. By preventing under-segmentation and ensuring high boundary fidelity, FA-UNet offers a more reliable tool for critical tasks such as accurate lesion volume assessment, which is essential for diagnosis and treatment planning in clinical practice.
To systematically validate the effectiveness of our proposed FA-UNet and to understand the specific contribution of its core architectural innovations, a comprehensive ablation study is conducted. This study deconstructs the model to quantify the impact of each primary component, the FasterNet block and the MSAG, on both segmentation accuracy and computational efficiency. The analysis involves a quantitative comparison of several model variants using key performance metrics (Dice, IoU) and efficiency indicators (Parameters, FLOPs, Inference Time). Furthermore, Grad-CAM is employed to qualitatively analyze the model’s decision-making process, providing visual evidence of how our proposed modules guide the network’s focus.
In this section, we quantitatively assess the individual and combined contributions of our primary architectural components through a systematic ablation study, with the results detailed in Table 2 on independent test dataset (private dataset). The evaluation begins with a Baseline Model (U-Net) and demonstrates the value added at each stage by progressively integrating our proposed modules: first, the MSAGs, followed by the FasterNet block to form the complete FA-UNet. This incremental analysis allows us to precisely measure how each component influences segmentation accuracy (Dice, IoU) while also evaluating its effect on model complexity and computational efficiency (Parameters, giga floating-point operations per second (GFLOPs), Inference Time).
| Model names | IoU | Dice | Params (M) | GFLOPs | Avg. time per image (ms) |
| Base | 0.7337 | 0.8455 | 35.33 | 136 | 2.89 |
| Base + MSAG | 0.7428 | 0.8521 | 48.75 | 180 | 3.36 |
| Base + MSAG + FasterNet Block (FA-UNet) | 0.7584 | 0.8679 | 46.42 | 180 | 3.16 |
| U-Net | 0.7323 | 0.8432 | 34.53 | 131 | 2.32 |
| U-Net3plus | 0.7457 | 0.8526 | 26.97 | 399 | 8.07 |
| CMU-Net | 0.7478 | 0.8536 | 49.93 | 182 | 3.33 |
Performance and efficiency metrics for the ablation study. All models were evaluated on an NVIDIA RTX 4090 GPU using a batch size of 16. M, Millions; ms, milliseconds; MSAG, multi-scale attention gate; GFLOPs, giga floating-point operations per second; Avg, average.
Based on the ablation study results shown in Table 2, the integration of MSAG modules into the baseline architecture results in a significant improvement in segmentation accuracy. The Dice score increases from 0.8455 to 0.8521, and the IoU score rises from 0.7337 to 0.7428. This improvement confirms the effectiveness of the MSAG in refining features, suppressing non-essential information, and emphasizing key lesion characteristics. This gain in accuracy, however, comes at a computational cost. The model’s parameter count increases from 35.33M to 48.75M, and the average inference time per image rises from 2.89 ms to 3.36 ms. This outcome illustrates a classic trade-off between performance and efficiency, where higher accuracy necessitates a more complex model.
Subsequently, the introduction of the FasterNet block at the bottleneck not only
further elevates the model’s performance but also significantly improves its
efficiency. The final FA-UNet model achieves the highest segmentation accuracy,
with a Dice score of 0.8679 and an IoU of 0.7584. Crucially, this is accomplished
while reducing the model’s complexity compared to the intermediate “Base +
MSAG” version. The number of parameters decreases from 48.75M to 46.42M, and the
inference time is reduced to 3.16 ms. This parameter reduction is achieved
because the single, highly efficient FasterNet block contains substantially fewer
weights than the two standard 3
When FA-UNet is benchmarked against other established architectures, its superior balance of performance and efficiency becomes even more evident. It achieves significantly higher segmentation accuracy than both the standard U-Net and the next-best competitor, CMU-Net, which recorded a Dice score of 0.8536. Although U-Net3plus has fewer parameters, its computational demand is more than double that of FA-UNet (399 vs. 180 GFLOPs), leading to a considerably slower inference time of 8.07 ms.
To move beyond quantitative metrics and gain deeper insight into the model’s interpretability, we employ Gradient-weighted Class Activation Mapping (Grad-CAM). This visualization technique produces heatmaps that highlight the image regions most influential in the model’s segmentation decision. Fig. 5 depicts the Grad-CAM heatmaps for representative cases from the test set. By analyzing the heatmaps from the FA-UNet, we qualitatively assess whether our architectural enhancements effectively guide the model’s focus toward the true pathological areas.
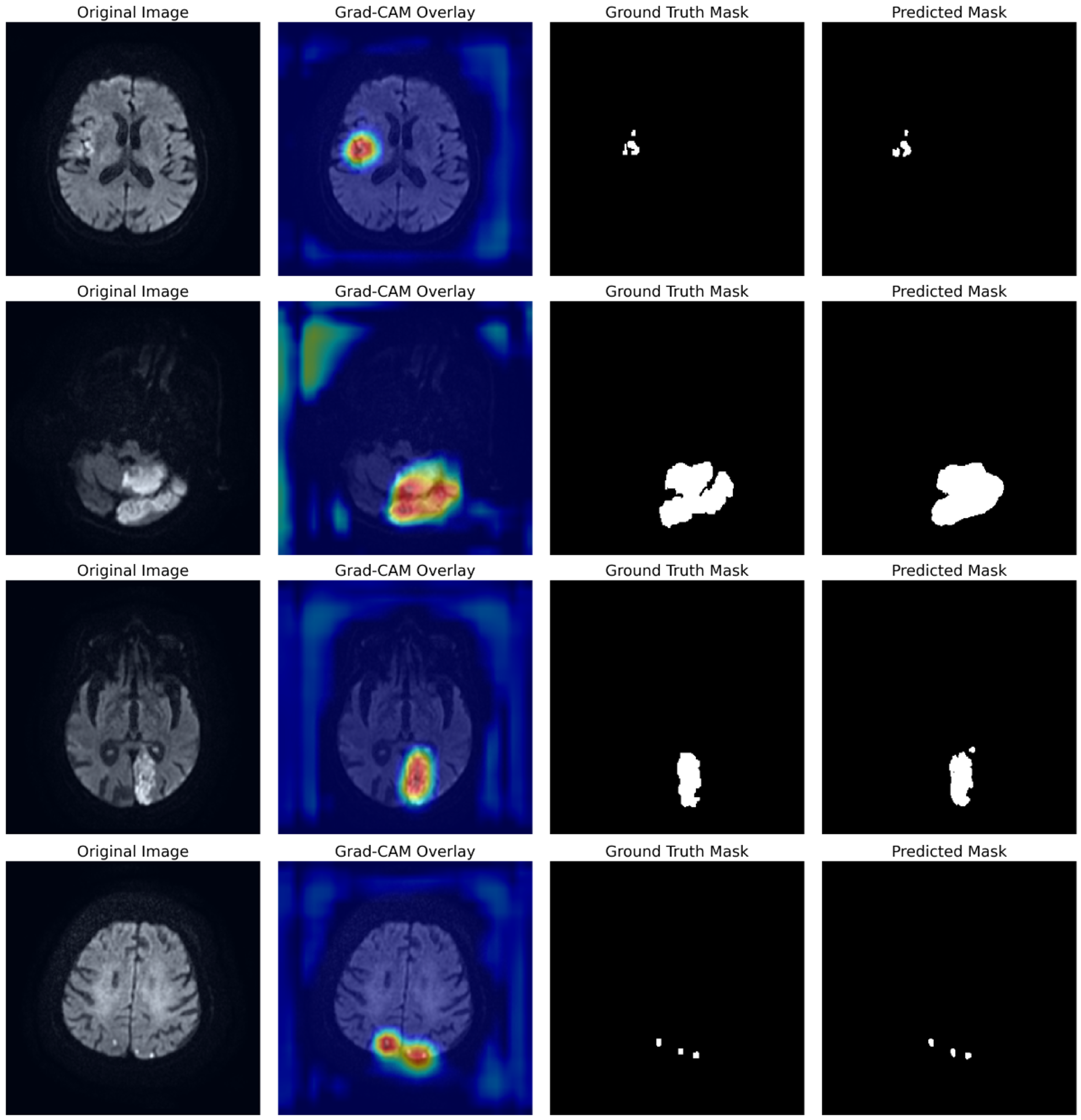 Fig. 5.
Fig. 5.
Visualization of FA-UNet’s learned attentional focus using Grad-CAM. Each row displays a different test case, showing the Original Image, the Grad-CAM heatmap overlay, the Ground Truth, and the final Predicted Mask. The heatmaps (red/yellow) highlight the model’s focus, which is consistently localized on the true lesion areas. Grad-CAM, Gradient-weighted Class Activation Mapping.
The analysis in Fig. 5 provides compelling visual evidence of FA-UNet’s robust localization capabilities. Across all presented cases, which include lesions of varying size and morphology, the model’s attentional hotspots (red/yellow) show a remarkable correspondence with the ground truth masks. For instance, the model accurately focuses on the entirety of larger infarcts (rows 1–3) as well as smaller, distinct lesions (row 4), confirming that it has learned to identify relevant pathological features rather than background noise. This direct correlation between the model’s learned focus and the final segmentation output underscores its reliability and provides a basis for clinical trust.
In this study, we introduced FA-UNet, a novel deep learning architecture designed to resolve the common trade-off between accuracy and computational efficiency in ischemic stroke lesion segmentation. The results demonstrate that the proposed model significantly outperforms existing state-of-the-art U-Net variants in terms of both spatial accuracy and diagnostic reliability. The success of FA-UNet is rooted in the synergistic interplay of two key architectural innovations: the FasterNet block at the bottleneck and the MSAGs on the skip connections. The superiority of our model stems from the ability of the FasterNet block to efficiently model global context. While conventional CNNs struggle to capture global features due to their limited receptive fields, and Transformer-based models do so at a high computational cost, FasterNet resolves this dilemma. It learns long-range spatial dependencies with lower latency by using PConv. This capability is critical for preserving the integrity of large, heterogeneous, and irregularly shaped infarcts, as visually corroborated by the case of the large, morphologically complex lesion in our results. The ablation study confirmed that the FasterNet block not only improves accuracy but also breaks the conventional performance-efficiency trade-off by reducing model complexity and inference time. The second critical component, the MSAGs, leverages a mechanism that has proven effective in its original application for addressing the challenges of low-contrast medical images, such as ultrasound [42]. While our current study did not conduct a specific experiment to isolate and measure performance based on image contrast, the high sensitivity and precision achieved by our model suggest that this adaptive feature refinement capability is highly beneficial for DWI-based stroke segmentation, where lesions often present with subtle signal differences. By processing features at three different scales and generating a dynamic attention map, this mechanism amplifies faint lesion signals while suppressing confounding signals from surrounding healthy tissue or imaging artifacts. The evidence for this is FA-UNet’s leading performance in both sensitivity and precision, demonstrating its robustness in not only detecting true lesions but also minimizing false positives. The combination of these two components endows FA-UNet with a masterful command of both local details and global context. This is qualitatively validated by Grad-CAM visualizations, which show the model’s focus consistently concentrated on true lesion areas, providing a basis for clinical trust through interpretability. Ultimately, the high Dice and IoU scores achieved by FA-UNet indicate that it offers a reliable tool for critical clinical tasks such as accurate lesion volume assessment, setting a new benchmark in the field.
Despite the promising results of the FA-UNet model presented in this study, certain limitations exist and open new avenues for future research. A primary limitation is the scope of the dataset; our model was trained on the public ISLES 2022 dataset and tested on a private dataset from a single institution. Its generalizability across different scanners, imaging protocols, and patient populations needs to be validated on more diverse, multi-center international datasets. This study also maintained a single modality focus on DWI scans. While crucial for stroke assessment, other magnetic resonance (MR) sequences like FLAIR and ADC offer complementary information that could potentially enhance performance. Furthermore, the study does not provide a detailed stratification of performance across different lesion subtypes or ages, and the exclusion of images with micro-lesions from training may limit its effectiveness on such cases. Looking forward, future work could focus on multi-modal integration by extending the FA-UNet architecture to fuse information from DWI, FLAIR, and ADC scans. Another valuable direction is using the model for longitudinal analysis to track lesion evolution over time, which could offer significant insights for predicting patient outcomes and response to therapy. The architecture could also be adapted for tasks beyond segmentation, such as predicting clinical outcome scores or classifying stroke subtypes from the initial scan. Finally, further efficiency optimization through techniques like model quantization and pruning could produce an even more lightweight version of FA-UNet for deployment in low-resource clinical settings, while expanding on our Explainable AI (XAI) analysis would further enhance clinical trust and facilitate its integration into diagnostic workflows.
In this work, we successfully developed and validated FA-UNet, a novel hybrid deep learning architecture for the accurate and efficient segmentation of ischemic stroke lesions. Our model innovatively combines a U-Net backbone with a computationally efficient FasterNet block at its bottleneck and adaptive MSAGs on its skip connections. This design effectively addresses the core challenges in ischemic stroke segmentation, including lesion heterogeneity, low image contrast, and computational demands. Comprehensive evaluations on an independent, private test set demonstrated that FA-UNet outperforms several state-of-the-art architectures, achieving a Dice coefficient of 0.8676 and an IoU of 0.7584. Critically, our model achieves this superior accuracy while maintaining the high computational efficiency necessary for practical deployment in clinical settings. Ablation studies and Grad-CAM visualizations further validated the positive contribution and interpretability of our proposed architectural components. In conclusion, FA-UNet establishes a new benchmark by successfully resolving the critical trade-off between accuracy and efficiency, offering a robust, reliable, and clinically viable tool that can enhance the diagnostic workflow for acute ischemic stroke and ultimately facilitate more timely and effective patient care.
The ISLES 2022 dataset used for training and validation in this study is publicly available through the official challenge website (https://www.isles-challenge.org/). The private test dataset generated and analyzed during the current study is not publicly available due to patient privacy and ethical restrictions but may be made available from the corresponding author upon reasonable request.
IP: Conceptualization, Methodology, Software, Formal Analysis, Investigation, Validation, Visualization, Project Administration, Funding Acquisition, Supervision, Writing—Original Draft, Writing—Review & Editing. AA: Conceptualization, Supervision, Writing—Original Draft, Writing—Review & Editing. BB: Data Curation, Investigation, Resources, Writing—Review & Editing. SI: Data Curation, Investigation, Resources, Writing—Review & Editing. All authors have read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
Experimental computations were carried out on the computing units at Igdir University’s Artificial Intelligence and Big Data Application and Research Center.
This work was supported by the grant provided by Türkiye Sağlık Enstitüleri Başkanlığı (TÜSEB) under the ‘2023-C1-YZ’ call and Project No: 33934. We would like to thank TUSEB for their financial support and scientific contributions.
The authors declare no conflict of interest.
During the preparation of this work, the authors used AI-assisted technologies for language enhancement and proofreading. After using these tools, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.





