1 Department of Chinese Medicine and Health Care, Changchun Humanities and Sciences College, 130117 Changchun, Jilin, China
2 School of Pharmaceutical Sciences, Jilin University, 130012 Changchun, Jilin, China
3 International College, Krirk University, 10220 Bangkok, Thailand
Abstract
Medicinal plants have long been used to manage hyperuricemia (HUA) and gout, with flavonoids identified as key bioactive components. However, the therapeutic effects of flavonoids on HUA remain inadequately characterized. Thus, we conducted a systematic review and meta-analysis to evaluate the impact of flavonoids on serum uric acid (UA) and xanthine oxidase (XOD) activity in animal models. A comprehensive literature search of the PubMed and Web of Science databases (January 2007–December 2024) identified 21 eligible studies involving 550 male mice. A random-effects meta-analysis revealed that flavonoid administration significantly reduced serum UA (standardized mean difference [SMD] = –2.22, 95% confidence interval (CI): –2.80 to –1.64; p < 0.001) and XOD activity (SMD = –1.79, 95% CI: –2.50 to –1.08; p < 0.001). Subgroup analyses indicated that chalcones, flavonols, and flavones exerted the most pronounced hypouricemic effects. Molecular docking of 28 flavonoids to XOD yielded binding affinities ranging from –8.4 to –10.9 kcal/mol, with molecular dynamics simulations confirming stable ligand–enzyme complexes; flavones exhibited the highest stability. Collectively, these data provide robust preclinical evidence that flavonoids ameliorate hyperuricemia via dual mechanisms of UA reduction and XOD inhibition, and support the further clinical development of flavonoids as potential natural agents for managing HUA.
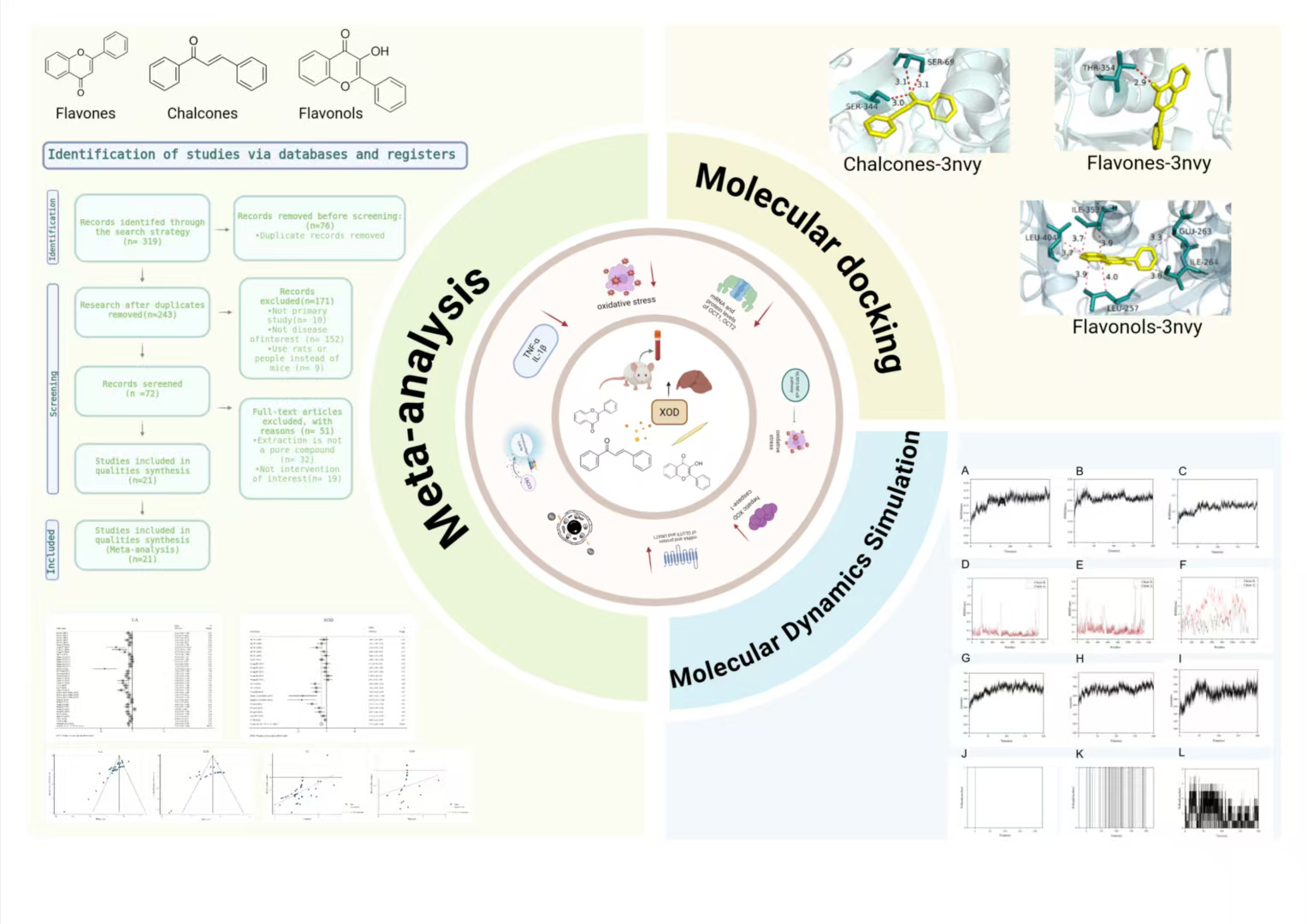
Graphical Abstract
Keywords
- medicinal plants
- natural products
- hyperuricemia
- flavonoids
- meta-analysis
- molecular dynamic simulation
Hyperuricemia (HUA), defined as a serum uric acid (UA) concentration exceeding 420 µmol/L on two separate occasions under standard dietary conditions, is now the second most prevalent metabolic disorder after diabetes [1]. Global epidemiological data reveal pronounced regional disparities, with a prevalence of 20.1% in the United States versus 13.3% in China (19.4% in males vs. 7.9% in females) [2]. Notably, more than 180 million individuals are affected in China alone, and over 50% of cases occur in adults aged 18–35 years, indicating a trend toward earlier onset [2]. Beyond gout, HUA is an independent risk factor for cardiovascular disease, chronic kidney disease, and type 2 diabetes mellitus, constituting a major public-health threat [3].
The cornerstone of contemporary HUA therapy is the inhibition of xanthine oxidase (XOD) to suppress UA synthesis and/or the use of uricosuric agents to enhance renal UA excretion, via blockade of uric acid transporter 1 (URAT1). Allopurinol and febuxostat remain first-line XOD inhibitors [4]; however, their clinical utility is constrained by serious adverse events. The HLA-B*5801 allele confers a 7- to 30-fold increase in the risk of life-threatening cutaneous reactions among Asian populations [5], whereas febuxostat has recently been associated with elevated cardiovascular mortality [6]. Consequently, safer and more cost-effective alternatives are urgently required.
Flavonoids—ubiquitous plant secondary metabolites built on a C6-C3-C6 scaffold—have been repeatedly reported to lower UA in potassium oxonate-induced rodents [7]. Mechanistic investigations indicate that these polyphenols attenuate purine catabolism by inhibiting XOD [8], modulate renal organic ion transporters (including GLUT9, URAT1, OAT1 and OAT3) [9]. and blunt NLRP3 inflammasome-driven oxidative stress [10]. Additionally, certain flavonoids can increase urinary UA excretion and fractional excretion of urate (FEUA) without suppressing hepatic XOD activity [11]. Nevertheless, the magnitude, precision and subclass-related heterogeneity of their hypouricemic action have not been quantitatively integrated, and no meta-analytic synthesis focusing exclusively on purified flavonoids in validated murine models of HUA is currently available [12].
Despite proven urate-lowering efficacy, the first-line XOD inhibitors
allopurinol and febuxostat carry distinct safety burdens. The HLA-B*5801 allele
reaches 7–8% in Asian populations (versus
To address this knowledge gap, we performed a Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA)-compliant systematic review and meta-analysis of 21 rodent studies coupled with computational mechanistic validation. Our objectives were (i) to estimate the pooled effect of flavonoids on serum UA and XOD activity, (ii) to dissect subclass-specific responses, and (iii) to corroborate experimental findings with molecular docking and long-timescale molecular dynamics (MD) simulations. The results provide a rigorous evidence base to guide the translational development of flavonoid-based therapeutics for hyperuricemia.
The systematic review and meta-analysis was conducted in strict accordance with the PRISMA 2020 statement [13]. The methodological quality of included animal studies was appraised with the Systematic Review Centre for Laboratory Animal Experimentations (SYRCLE) guidelines [14].
A systematic search was executed through comprehensively searching in online databases PubMed and Web of Science for studies published between January 2007 and December 2024. The searches were conducted by information specialist MJ Zhao. The following keywords were used in the search: (flavonoids) and (hypouricemic OR hyperuricemia OR hyperuricemia or gout or hyperuricemia) AND (mice). Initial studies were imported to the EndNote 20 (Clarivate Analytics, Philadelphia, PA, USA) software. After removing duplicate studies, two independent researchers (MJ Zhao and Q Xiao) screened the titles and abstracts of the remaining articles. Studies that did not meet the inclusion criteria were excluded. When necessary, the full texts of the remaining publications were evaluated. Any disagreements regarding the inclusion or exclusion of studies were resolved through discussion among the authors until consensus was reached.
The criteria for included animal studies were as follows:
(1) original research that presented original data;
(2) study performed in male mice weighing 18 to 30 g;
(3) disease of interest (hyperuricemia induced by potassium oxonate);
(4) intervention of interest (flavonoids that were pure natural products);
The criteria for excluded animal studies were as follows:
(1) in vitro or ex vivo models;
(2) complex models of HUA and other disorders;
(3) studies not using one of pure compound as an intervention group;
(4) not interested intervention;
(5) reviews, conference abstracts, editorials, clinical trials, case reports and incomplete articles;
(6) not an English-language publication.
Two authors (JJ Xu and Q Xiao) independently extracted study characteristics The extracted information included the first author’s name, year of publication, animal details (species, weight, sex, strain, control group), and reagent characteristics (flavonoid type, sample size, study duration, intervention dose, potassium oxonate dose). Standard deviations (SD) and means for intervention and control groups were also recorded. If study data were only available as graphs, Origin 2025 (OriginLab Corporation, Northampton, MA, USA) software was used for extraction. Any discrepancies in measurements were resolved through discussion between the two authors.
The quality assessment was conducted independently by two researchers (Q Xiao and MJ Zhao) to ensure inter-rater reliability and to minimize the potential bias. Any discrepancies in the risk of bias assessment were resolved through consensus discussions, ensuring a standardized and objective evaluation of study quality. The methodological rigor of the included studies was critically appraised using SYRCLE’s Risk of Bias tool, a validated instrument specifically designed for assessing the risk of bias in animal intervention studies.
XOD was selected as the core target for molecular docking analysis. The 3D structures of the active compounds were retrieved from PubChem, and the crystal structure of XOD (PDB ID: 3NVY) was downloaded from the PDB database (RCSB Protein Data Bank, New Brunswick, NJ, USA). Solvent molecules were removed using PyMOL, and the target protein was prepared for hydrogenation and charge assignment using MGLTools 1.5.7 (The Scripps Research Institute, La Jolla, CA, USA). Both the protein and the 18 compounds were saved as “pdbqt” files, and the docking box was configured. Molecular docking was performed using Autodock suite 4.2.6 (The Scripps Research Institute, La Jolla, CA, USA), with binding affinities calculated in kcal/mol. Binding affinities less than –5.00 kcal/mol were considered indicate strong binding interactions.
To assess complex stability, 200 ns atomistic MD simulations were conducted with GROMACS 2020.6 (GROMACS Development Team, Stockholm, Sweden) for the three highest-affinity representatives (chalcones, flavones, flavonols) bound to XOD. The Amber99SB-ILDN force-field was used for the protein; ligands were parameterised with GAFF2 and AM1-BCC charges. Each system was solvated in a cubic TIP3P water box (minimum 1.2 nm buffer) and neutralised with Na+/Cl– ions to 0.15 M. Following energy minimisation (steepest descent), 1-ns NVT (300 K, V-rescale) and 1-ns NPT (1 bar, Parrinello–Rahman) equilibrations were performed with position restraints on heavy atoms. Production runs used 2-fs time steps, LINCS constraints, and PME electrostatics. Trajectories were saved every 10 ps for subsequent analysis of root-mean-square deviation (RMSD), root-mean-square fluctuation (RMSF), solvent-accessible surface area (SASA), and hydrogen-bond occupancy.
Standardised mean differences (SMD, Hedges’ g) with 95% confidence intervals
(CI) were pooled under a random-effects model (DerSimonian–Laird). Heterogeneity
was quantified with I2 and
Database searching retrieved 319 records; 76 duplicates were removed. After title/abstract screening, 222 articles were excluded (171 non-primary studies, non-matching models, or rat protocols; 51 non-pure compounds or irrelevant interventions). Twenty-one studies met the inclusion criteria and were entered into quantitative synthesis (Fig. 1).
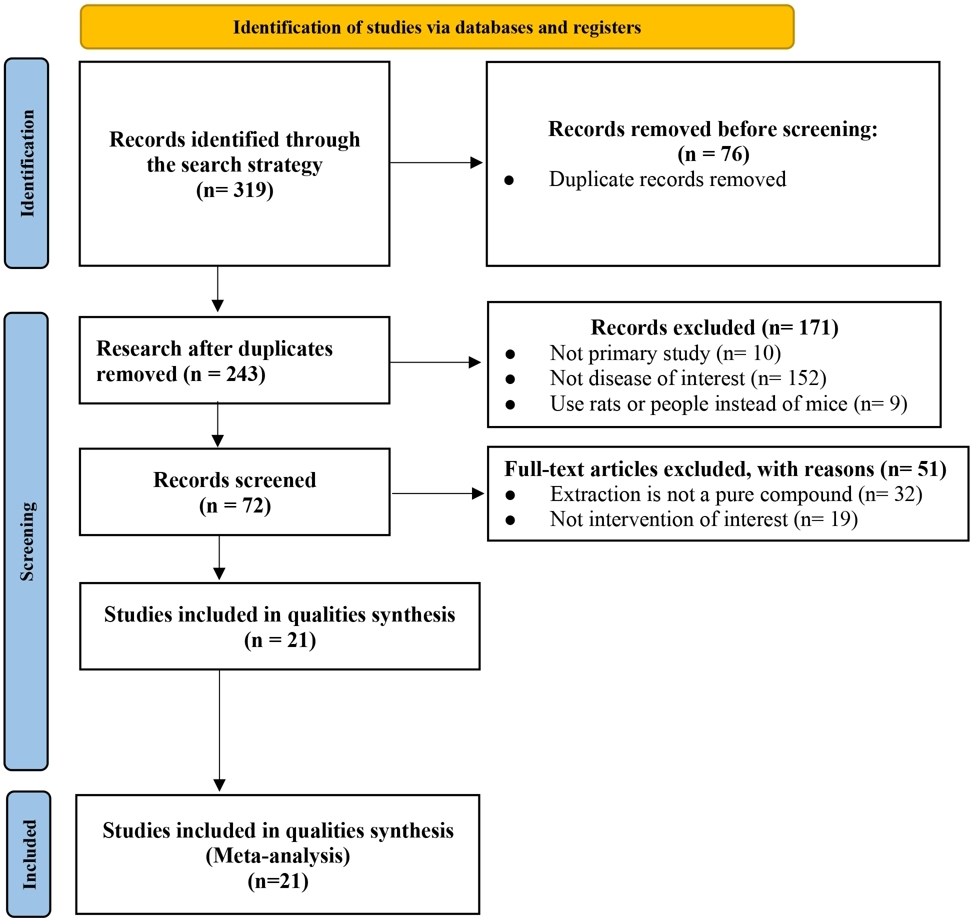 Fig. 1.
Fig. 1.
PRISMA flowchart of literature search and study selection (21 studies included). PRISMA, Preferred Reporting Items for Systematic Reviews and Meta-Analyses.
The 21 articles comprised 550 male mice. Detailed intervention parameters and animal characteristics are summarised in Table 1 (Ref. [15, 16, 17, 18, 19, 20, 21, 22, 23, 24]). Nine strains were represented: Kun-Ming (n = 9 studies), ICR (6), C57BL/6 (3), Albino Swiss (2) and BALB/c (1). Hyperuricemia was induced by potassium oxonate (PO) at 250–300 mg kg-1 day-1 for 1–28 days. Flavonoid interventions included chalcones (6 datasets), flavonols (18), flavones (7) and others (8); doses ranged 0.5–700 mg kg-1 day-1. All studies employed allopurinol (5–10 mg kg-1) as positive control. Chemical structures and subclass assignments of the investigated flavonoids are provided in Table 2.
| Flavonoid | Mouse Weight (g) | Sample Size (C/T) | Mice Species | Control Allopurinol (mg/kg) | Study length (Days) | Intervention Dose (mg/kg/d) | PO Dose (mg/kg/d) | Study |
| Morin | 25–28 | 2/10 | ICR | 10 | 3 | 50,100 | 250 | [15] |
| Puerarin | ||||||||
| Myricetin | ||||||||
| Apigenin | ||||||||
| Quercetin | ||||||||
| Kaempferol | ||||||||
| Okanin | 30 | 2/6 | ICR | 13.6 | 1 | 28.8 | 250 | [16] |
| (-)-2,3-cis-3,4-cis-3,3′4,4′,7,8-hexahydroxyflavan | 30.6 | |||||||
| (-)-2,3-cis3,4-cis-4′-methoxy-3,3′,4,7,8-pentahydroxyflavan | 32.0 | |||||||
| Morin | 18–22 | 8/8 | Kunming | 2.5/5 | 7 | 10–80 | 250 | [17] |
| Rutin | 23–27 | 2/10 | BALB/c | 10 | 7 | 75–300 | 250 | [18] |
| Astilbin | 1, 9 | |||||||
| Genistein | 4.5, 18 | |||||||
| Apigenin | 175, 700 | |||||||
| Quercetin | 100, 400 | |||||||
| 3,5,2′,4′-tetrahydroxychalcone | 18–22 | 10/10 | Kunming | 1 | 7 | 0.5, 2.0 | 500 | [19] |
| Apigenin | 25–30 | 3/6 | Albino Swiss | 10 | 3 | 25 | 250 | [20] |
| Luteolin | ||||||||
| Quercetin | 18–22 | 6/6 | Kunming | 5 | 7 | 25, 100 | 250 | [21] |
| Rutin | 18–22 | 8/8 | Kunming | 5 | 7 | 25, 50, 100 | 250 | [22] |
| Astilbin | 20 |
15/15 | Kunming | / | 7 | 5, 10, 20 | - | [23] |
| Baicalein | 20–22 | 6/6 | ICR | / | 21 | 50 | 300 | [24] |
| Subgroup | No. | Name | R1 | R2 | R3 | R4 | R5 | R6 | |
| Chalcones | 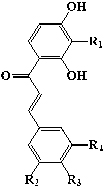 |
1 | 3,5,2′,4′-tetrahydroxychalcone | -H | -OH | -H | -OH | ||
| 2 | Okanin | -OH | -OH | -OH | -H | ||||
| 3 | Phloretin | 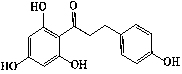 |
|||||||
| Flavones | 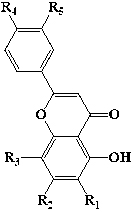 |
4 | Apigenin | -H | -OH | -H | -H | -OH | |
| 5 | Baicalein | -OH | -OH | -H | -H | -H | |||
| 6 | Luteolin | -H | -OH | -H | -OH | -OH | |||
| 7 | Luteolin-4′-O-glucoside | -H | -OH | -H | -OH | -OGlc | |||
| 8 | Scutellarin | -OH | -OGlu A | -H | -OH | -H | |||
| 9 | Vitexin | -H | -OH | -OGlc | -OH | -H | |||
| Flavonols | 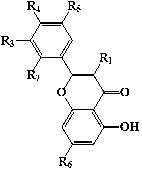 |
10 | Astilbin | -ORha | -H | -OH | -OH | -H | -OH |
| 11 | Hesperidin | -H | -H | -OH | -OCH3 | -H | -OGlc-(6 | ||
| 12 | Kaempferol | -OH | -H | -H | -OH | -H | -OH | ||
| 13 | Kaempferol-3-O-sophoroside | -Osophoroside | -H | -H | -OH | -H | -OH | ||
| 14 | Sodium kaempferol-3′-sulfonate | -OH | -H | -H | -OH | -SO3Na | -OH | ||
| 15 | Morin | -OH | -OH | -H | -OH | -H | -OH | ||
| 16 | Quercetin | -OGlc | -H | -OH | -OH | -H | -OH | ||
| 17 | Myricetin | -OH | -H | -OH | -OH | -OH | -OH | ||
| 18 | Rutin | -OGlc-(6 |
-H | -OH | -OH | -H | -OH | ||
| 19 | 6,8,3′,4′-tetrahydroxyflavanone-7-C- |
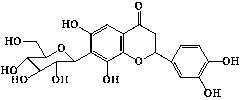 |
|||||||
| 20 | 6,8,4′-trihydroxyflavanone-7-C- |
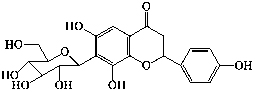 |
|||||||
| Others | Flavan-3-ol | 21 | (-)-2,3-cis-3,4-cis-3,3′4,4′,7,8-hexahydroxyflavan | -OH | 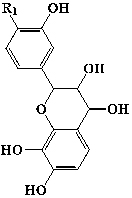 |
||||
| 22 | (-)-2,3-cis-3,4-cis-4′-methoxy-3,3′,4,7,8-pentahydroxyflavan | -OMe | |||||||
| Isoflavones | 23 | Genistein | 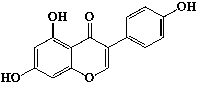 |
||||||
| Anthocyanidins | 24 | Anthocyanin | 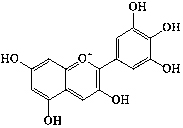 |
||||||
| 25 | Epigallocatechin-3-gallate | 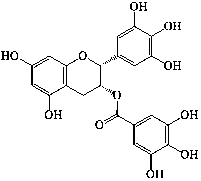 |
|||||||
| 26 | Epiphyllocoumarin-3-O- |
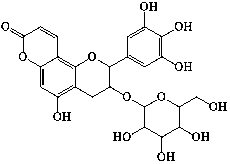 |
|||||||
| 27 | 2′,4′-dihydroxychalcone-(4-O-7′′)-8′′,4′′′-dihydroxyflavanone | 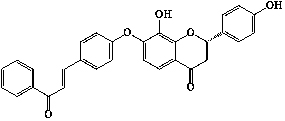 |
|||||||
| 28 | Puerarin | 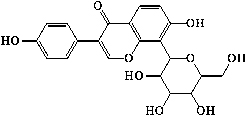 |
|||||||
As the final product of purine metabolism, UA is crucial for diagnosing HUA and
understanding the pathogenesis of gout and rheumatoid arthritis [25].
Meta-analysis of 21 comparisons revealed that revealed that flavonoid
interventions significantly reduced serum UA levels compared to control groups
(SMD = –2.22, 95% CI [–2.80, –1.64], I2 = 79.7%, p
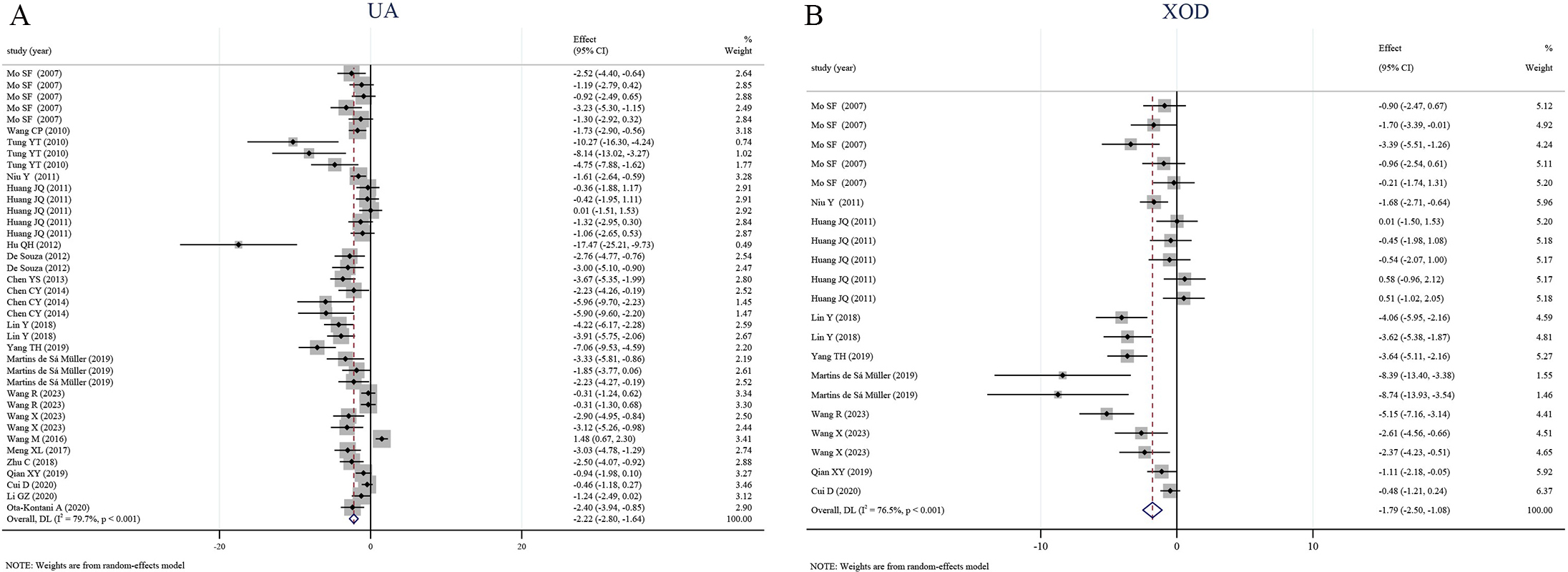 Fig. 2.
Fig. 2.
Forest plots of flavonoids effects on (A) serum uric
acid and (B) xanthine oxidase activity versus control. Pooled weighted mean
differences (random-effects) are shown with 95% CIs. flavonoid intervention
significantly reduced both outcomes (A: WMD = –2.22, 95% CI –2.80 to –1.64,
I2 = 79.7%, p
XOD is the rate-limiting enzyme in UA production; its inhibition offers a direct
index of hypouricemic efficacy [26]. Pooled data from 21 studies demonstrated
that flavonoids interventions significantly reduced serum XOD activity compared
to control groups (SMD = –1.79, 95% CI [–2.50, –1.08], I2 = 76.5%,
p
Subgroup analyses classified flavonoids into five categories—flavonols,
flavanones, flavones, chalcones, and others—and consistently revealed
significant serum-UA reductions relative to controls (random-effects; 21
studies). Flavonols: SMD = –1.99, 95% CI [–2.77, –1.22], p
When data were stratified by mouse strain, serum UA was significantly lowered in
ICR (SMD = –2.32, 95% CI [–3.23, –1.42], p
| Subgroups | Quantity | I2 | Heterogeneity (p) | SMD | 95% CI | Z test (p) | ||
| Serum UA | 21 | |||||||
| Flavonoids | ||||||||
| Flavonols | 12 | 72.0% | p |
–1.99 | [–2.77, –1.22] | p | ||
| Flavones | 3 | 11.7% | p |
–2.99 | [–3.76, –2.22] | p | ||
| Chalcones | 3 | 88.0% | p |
–3.63 | [–5.68, –1.58] | p = 0.001 | ||
| Others | 3 | 84.6% | p |
–1.27 | [–2.52, –0.03] | p = 0.046 | ||
| Animal species | ||||||||
| ICR | 7 | 72.6% | p |
–2.32 | [–3.23, –1.42] | p | ||
| Kun-Ming | 6 | 91.2% | p |
–3.27 | [–4.88, –1.65] | p | ||
| BALB/c | 5 | 0.0% | p = 0.762 | –0.61 | [–1.30, 0.09] | p = 0.088 | ||
| Albino Swiss | 2 | 0.0% | p = 0.877 | –2.56 | [–3.50, –1.63] | p | ||
| C57BL/6 | 1 | 62.5% | p = 0.070 | –1.19 | [–2.28, –0.12] | p = 0.030 | ||
| Serum XOD | 21 | |||||||
| Flavonoids | ||||||||
| Flavonols | 12 | 74.9% | p |
–2.14 | [–3.26, –1.03] | p | ||
| Flavones | 3 | 84.3% | p = 0.002 | –2.59 | [–5.09, –0.08] | p = 0.043 | ||
| Chalcones | 3 | 86.6% | p = 0.001 | –1.83 | [–3.48, –0.18] | p = 0.030 | ||
| Others | 3 | 40.6% | p = 0.186 | –0.32 | [–1.34, 0.68] | p = 0.525 | ||
| Animal species | ||||||||
| ICR | 7 | 70.0% | p = 0.003 | –1.77 | [–2.87, –0.67] | p = 0.002 | ||
| Kun-Ming | 6 | 40.8% | p = 0.133 | –2.87 | [–3.72, –2.02] | p | ||
| BALB/c | 5 | 0.0% | p = 0.780 | 0.02 | [–0.66, 0.71] | p = 0.947 | ||
| Albino Swiss | 2 | 0.0% | p = 0.925 | –8.56 | [–12.16, –4.95] | p | ||
| C57BL/6 | 1 | / | / | –0.48 | [–1.21, 0.24] | p = 0.192 | ||
SMD, standardized mean difference; UA, uric acid; XOD, xanthine oxidase.
As illustrated in Table 4 (Ref. [15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37]), the baseline characteristics that might influence outcome indicators were inadequately described in the majority of studies (19/21). Specific methodological facets—including sequence generation, allocation concealment, random outcome assessment, and blinding of caregivers or investigators—were either sparsely reported or not documented at all, leading to an “unclear” risk-of-bias classification for these domains. Nevertheless, 17 investigations (80.9%) explicitly stated that experimental animals were housed in identical macro-environmental conditions (temperature, humidity, light–dark cycle, and bedding), allowing us to assign a “low risk” rating to the item of random housing. Furthermore, all included publications provided evidence that baseline body weight, serum uric acid, and xanthine oxidase activity did not differ between treatment and control arms; consequently, the risk of confounding attrition was considered “low” across the entire data set. Incomplete outcome data were adequately addressed in every study (low risk), and neither selective reporting nor other sources of bias (e.g., early termination, unit-of-analysis errors) were detected (low risk for all trials).
| Study | A | B | C | D | E | F | G | H | I | J |
| Mo SF (2007) [15] | U | U | U | Y | U | N | U | Y | Y | Y |
| Tung YT (2010) [16] | U | U | U | Y | U | N | U | Y | Y | Y |
| Wang CP (2010) [17] | U | U | U | Y | U | U | U | Y | Y | Y |
| Huang JQ (2011) [18] | U | U | U | Y | U | N | Y | Y | Y | Y |
| Niu Y (2011) [19] | U | U | U | Y | U | N | Y | Y | Y | Y |
| De Souza (2012) [20] | U | U | U | Y | U | N | Y | Y | Y | Y |
| Hu QH (2012) [21] | U | U | U | U | U | N | Y | Y | Y | Y |
| Chen Y (2013) [22] | U | U | U | Y | U | N | U | Y | Y | Y |
| Adachi SI (2017) [32] | U | U | U | Y | U | N | Y | Y | Y | Y |
| Wang M (2016) [23] | U | U | U | Y | U | U | U | Y | Y | Y |
| Meng XL (2017) [24] | U | U | U | U | U | U | Y | Y | Y | Y |
| Lin Y (2018) [35] | U | U | U | U | U | N | Y | Y | Y | Y |
| Zhu C (2018) [34] | U | U | U | Y | U | N | U | Y | Y | Y |
| Martins de Sá Müller (2019) [29] | U | U | U | Y | U | N | Y | Y | Y | Y |
| Qian XY (2019) [33] | U | U | U | Y | U | U | U | Y | Y | Y |
| Yang TH (2019) [30] | U | U | U | Y | U | U | U | Y | Y | Y |
| Cui DL (2020) [36] | U | U | U | Y | U | U | Y | Y | Y | Y |
| Li GZ (2020) [37] | U | U | U | Y | U | U | U | Y | Y | Y |
| Ota-Kontani A (2020) [31] | U | Y | U | Y | U | U | Y | Y | Y | Y |
| Wang R (2023) [27] | Y | U | U | Y | U | U | U | Y | Y | Y |
| Wang X (2023) [28] | U | U | U | U | U | U | U | Y | Y | Y |
Annotation: U, unclear; Y, Yes; L, low risk; N, No; H, high risk; A, Sequence generation; B, Baseline characteristics; C, Allocation concealment; D, Random housing; E, Performance blinding; F, Random outcome assessment; G, Blinding of outcome assessors; H, Incomplete data reporting; I, Selective outcome reporting; J, Other sources of bias.
As shown in Fig. 3, funnel-plot appraisal of serum UA and XOD outcomes revealed
marked asymmetry, with numerous small-effect studies lying outside the
pseudo-confidence limits, strongly suggestive of a small-study effect. Egger’s
linear regression corroborated this impression, yielding statistically
significant intercepts for both UA (t = 3.11, p = 0.007) and XOD (t =
2.89, p = 0.012), thereby confirming the presence of publication bias.
Such bias could, in principle, inflate or deflate the pooled effect estimates and
compromise the validity of any therapeutic inference. To gauge the robustness of
the summary statistics we therefore implemented the trim-and-fill algorithm
(Lönnér estimator) under a random-effects model. After two iterative
cycles no additional trial was imputed and the adjusted pooled SMD for UA
remained –2.20 (95% CI –2.79 to –1.63), virtually identical to the original
–2.22, indicating that the asymmetry had negligible influence on the overall
estimate. Complementary Harbord and Peters tests likewise returned
non-significant bias coefficients (both p
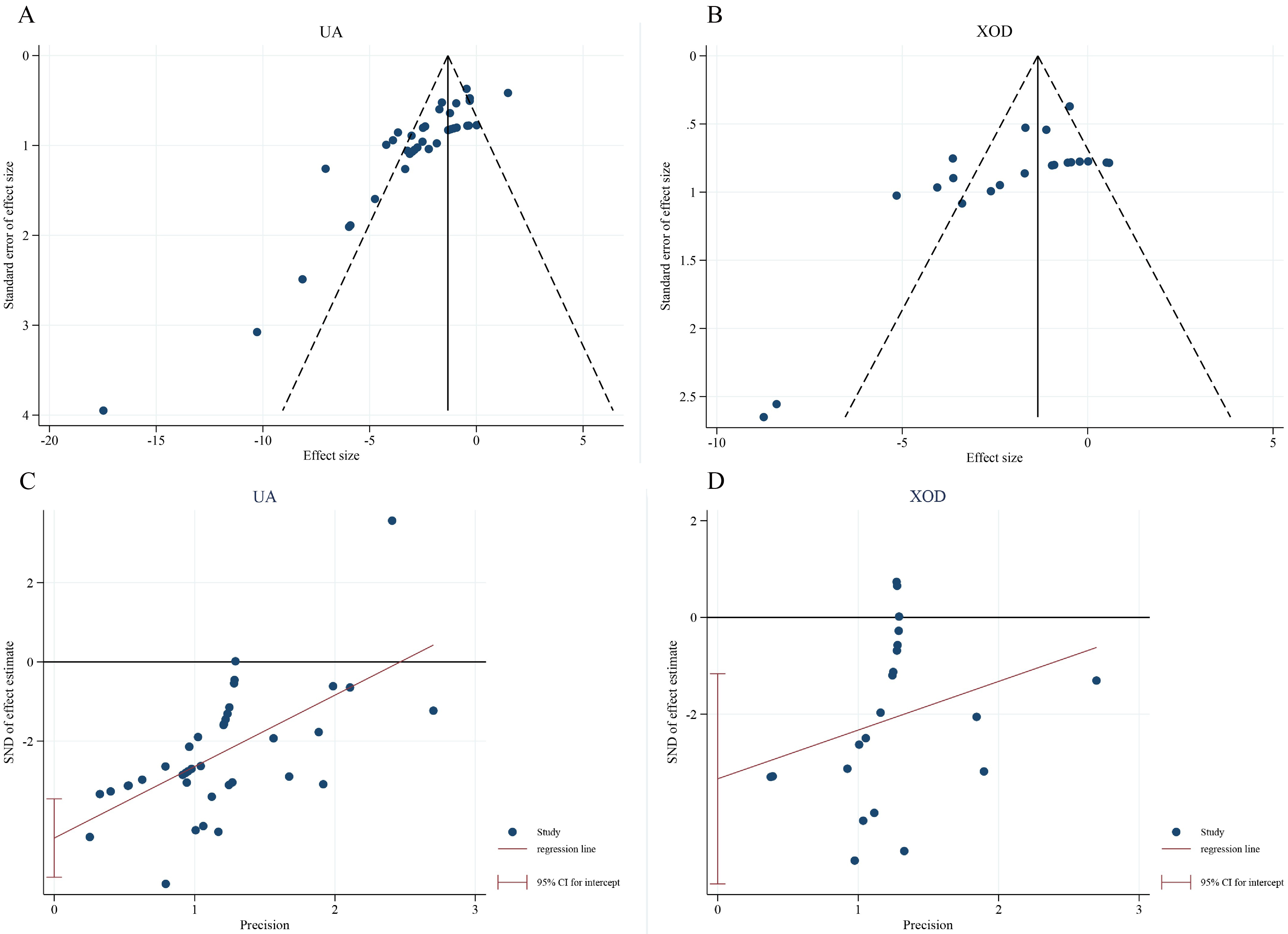 Fig. 3.
Fig. 3.
Funnel plots and Egger’s regression-based publication bias tests for the 21 included studies. (A) Funnel plot for serum UA; (B) funnel plot for XOD activity. Open circles represent individual comparisons; diagonal dashed lines indicate the 95% pseudo-confidence limits. Egger’s linear regression intercept is displayed in the lower left corner of each panel, confirming small-study effects for both outcomes. (C) Egger’s linear regression plot for serum uric acid (UA); (D) Egger’s linear regression plot for xanthine oxidase (XOD). Blue scatter points in the plots represent individual studies included in the meta-analysis; pink diagonal lines denote the Egger’s linear regression fitting lines, and black horizontal lines indicate the reference lines for pooled effect sizes. The red vertical lines with error bars correspond to the 95% confidence intervals (95% CI for intercept) of the intercepts.
We investigated the interactions between drugs and proteins via molecular
docking, and in this study, 28 flavonoid compounds were subjected to molecular
docking with xanthine oxidase (XOD), as shown in Fig. 4 among chalcones-type
flavonoids, 3,5,2′,4′-tetrahydroxychalcone, Okanin, and Phloretin all
exhibited a binding affinity of –8.9 kcal/mol; for flavones-type flavonoids, the
binding affinities of Apigenin, Baicalein, Luteolin, Luteolin-4′-O-glucoside,
Scutellarin, and Vitexin were –9.2, –9.8, –9.5, –10.3, –10.2, and –8.8
kcal/mol, respectively; regarding flavonols-type flavonoids, Astilbin,
Hesperidin, Sodium kaempferol-3′-sulfonate, Morin, Quercetin, Myricetin,
Rutin, 6,8,3′,4′-tetrahydroxyflavanone-7-C-
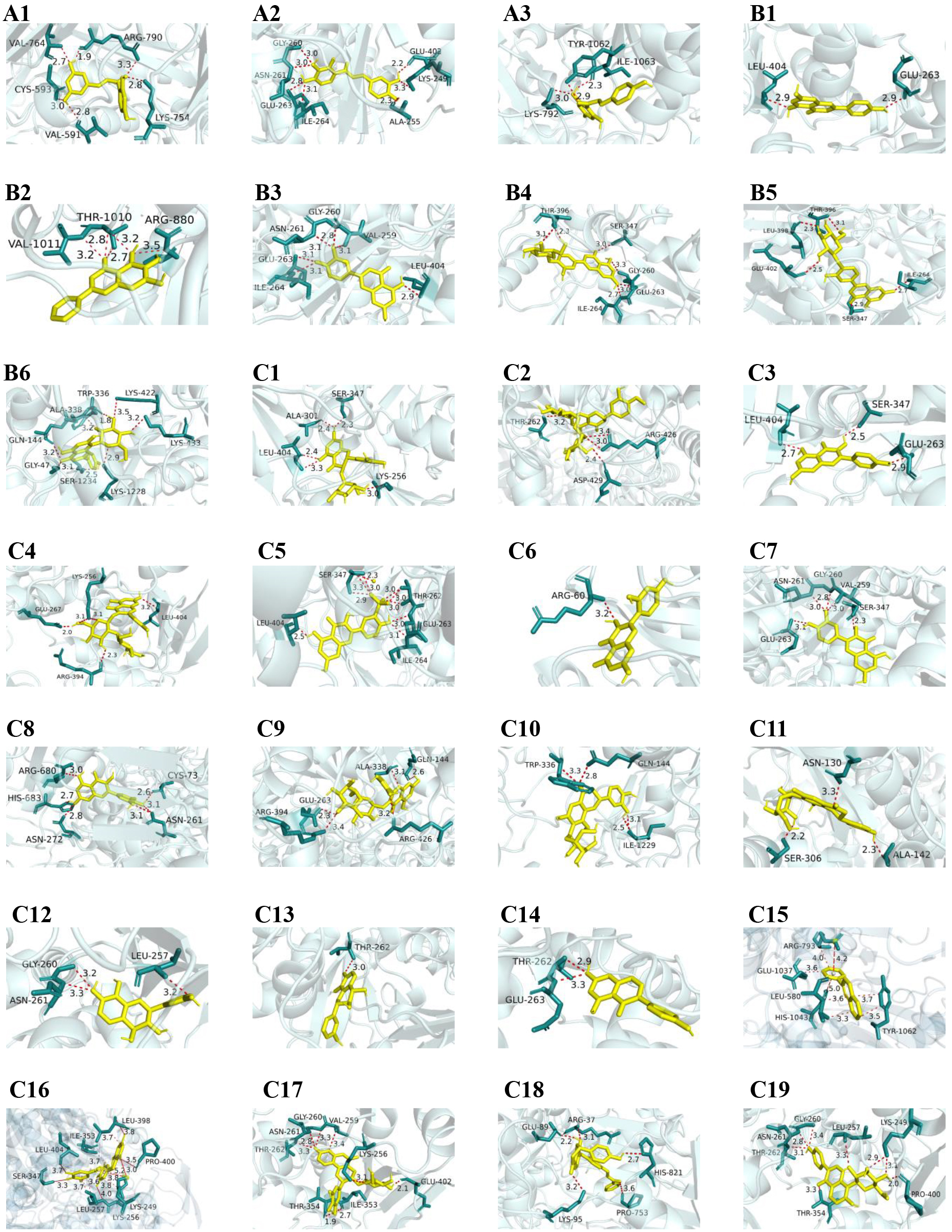 Fig. 4.
Fig. 4.
Visualization of molecular docking between active ingredients of flavonoids and XOD. (A1–A3) show the visualization of molecular docking between chalcones-type flavonoids and XOD; (B1–B6) represent the visualization of molecular docking between flavone-type flavonoids and XOD; (C1–C11) display the visualization of molecular docking between flavonols-type flavonoids and XOD; and (C12–C19) present the visualization of molecular docking between other types of flavonoids and XOD.
| Code | Mode | Affinity (kcal/mol) | Hydrogen bonds interacting residues |
| A1 | 3,5,2′,4′-tetrahydroxychalcone | –8.9 | VAL 764,ARG 790,CYS 593,LYS 754,VAL 591 |
| A2 | Okanin | –8.9 | GLY 260,ASN 261,GLU 263,ILE 264,GLU 402,LYS 249,ALA 255 |
| A3 | phloretin | –8.9 | TYR 1062,ILE 1063,LYS 792 |
| B1 | Apigenin | –9.2 | LEU 404,GLU 263 |
| B2 | Baicalein | –9.8 | VAL 1001,THR 1010,ARG 880 |
| B3 | Luteolin | –9.5 | GLY 260,ASN 261,GLU 263,ILE 264,VAL 259,LEU 404 |
| B4 | Luteolin-4′-O-glucoside | –10.3 | THR 396,SER 347,GLY 260,GLU 263,ILE 264 |
| B5 | Scutellarin | –10.2 | THR 396,LEU 398,GLU 402,ILE 264,SER 347 |
| B6 | Vitexin | –8.8 | TRP 336,ALA 338,GLN 144,GLY 47,SER 1234,LYS 1228,LYS 422,LYS 433 |
| C1 | Astilbin | –9.4 | SER 347,ALA 301,LEU 404,LYS 256 |
| C2 | Hesperidin | –10.6 | THR 262,ASP 429,ARG 426 |
| C3 | Sodium kaempferol-3′-sulfonate | –10.4 | LUE 404,GLU 263,SER 347,THR 262,ILE 264 |
| C4 | Morin | –8.6 | ARG 60 |
| C5 | Quercetin | –9.9 | GLY 260,ASN 261,VAL 259,SER 347,GLU 263 |
| C6 | Myricetin | –9.5 | ARG 680,HIS 683,ASN 272,CYS 73,ASN 261 |
| C7 | Rutin | –9.8 | ARG 394,GLU 263,ALA 338,GLN 144,ARG 426 |
| C8 | 6,8,3′,4′-tetrahydroxyflavanone-7-C- |
–8.4 | TRP 336,GLN 144,ILE 1229 |
| C9 | 6,8,4′-trihydroxyflavanone-7-C- |
–8.8 | ASN 130,SER 306,ALA 142 |
| D1 | (-)-2,3-cis-3,4-cis-3,3′4,4′,7,8-hexahydroxyflavan | –8.7 | GLY 260,LEU 257,ASN 261 |
| D2 | (-)-2,3-cis-3,4-cis-4′-methoxy-3,3′,4,7,8-pentahydroxyflavan | –8.9 | THR 262 |
| D3 | Genistein | –9.1 | THR 262,GLU 263 |
| D4 | Anthocyanin | –8.9 | ARG 793,GLU 1037,LEU 580,HIS 1043,TYR 1062 |
| D5 | Epigallocatechin-3-gallate | –9.9 | LEU 398,ILE 353,LEU 404,SER 347,LEU 257.LYS 256,LYS 249,PRO 400 |
| D6 | Epiphyllocoumarin-3-O- |
–10.8 | GLY 260,ASN 261,THR 262,THR 354,ILE 353,GLU 402,LYS 256,VAL 259 |
| D7 | 2′,4′-dihydroxychalcone-(4-O-7′′)-8′′,4′′′-dihydroxyflavanone | –10.9 | GLU 89,ARG 37,HIS 821,PRO 753,LYS 95 |
| D8 | Puerarin | –10.1 | GLY 260,ASN 261,THR 262,THR 354,PRO 400,LYS 249,LEU 257 |
To quantitatively compare the stability of XOD-flavones, XOD-flavonols and
XOD-chalcones complexes, 200-ns GROMACS trajectories were analysed by RMSD, RMSF,
SASA and hydrogen-bond metrics. RMSD records (Fig. 5A–C) show that the flavones
adduct equilibrated within 20 ns and maintained a mean value of 0.22
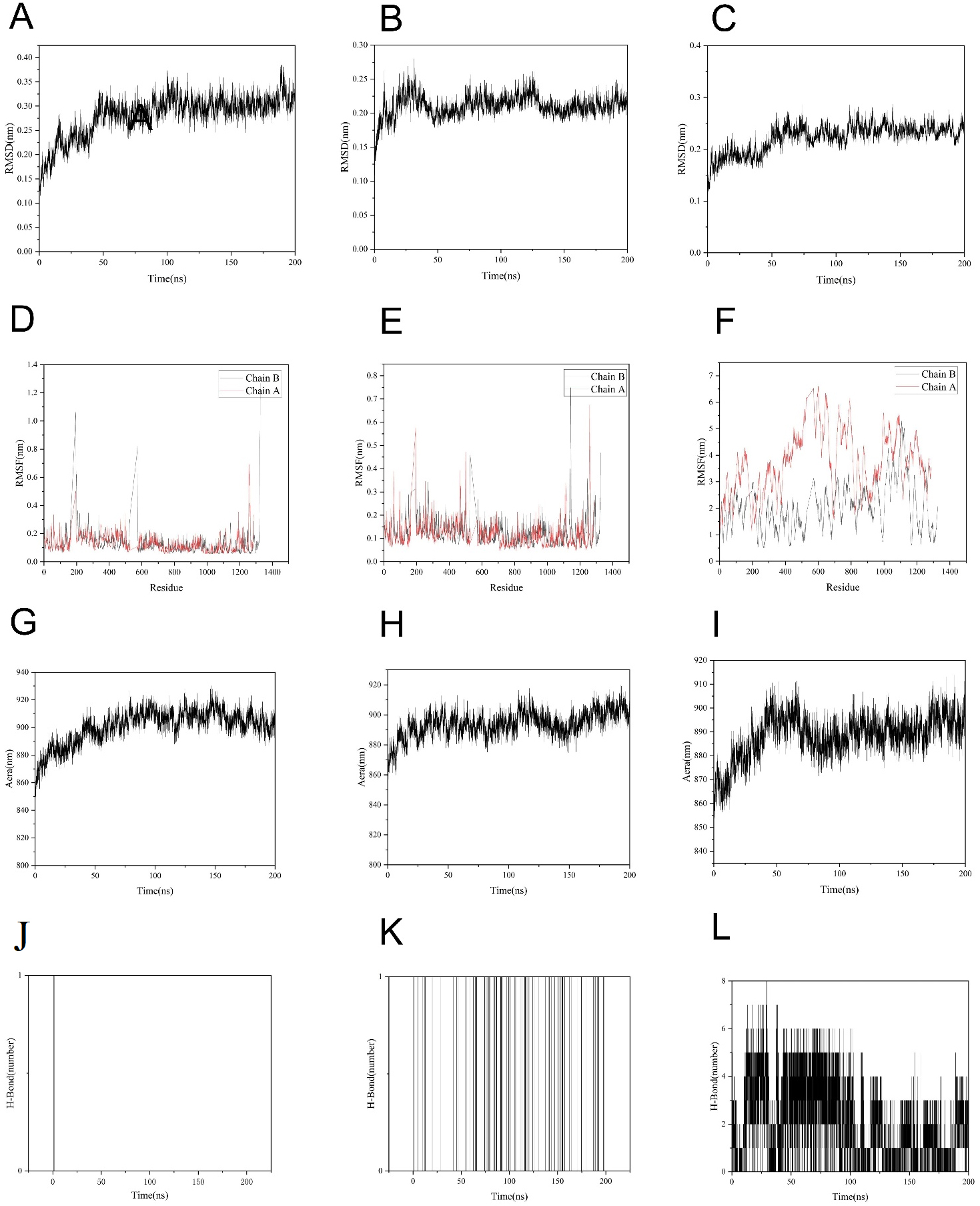 Fig. 5.
Fig. 5.
Molecular dynamics simulation between active ingredients of flavonoids and XOD. The figure presents a comprehensive analysis of the molecular dynamics simulation trajectory over 200 ns. (A–C) RMSD of the protein backbone and ligand atoms over time. (D–F) RMSF per residue for chain A and chain B of the XOD dimer. (G–I) Evolution of key interaction parameters (hydrogen bonds, hydrophobic contacts) throughout the simulation. (J–L) Analysis of binding free energy contributions and the interaction frequency of key residues. The simulation results demonstrate the stability and binding mode of flavonoids within the active site of XOD. RMSD, root-mean-square deviation; RMSF, root-mean-square fluctuation.
In the included studies, most flavonoid subclasses reduced xanthine production by inhibiting XOD activity, resulting in decreasing UA synthesis [16, 20, 27, 28, 29, 30, 31, 32]. Chalcones exhibited a dose-dependent effect, with higher concentrations yielding better UA-lowering effect [38]. XOD is present in both serum and liver, but changes in UA levels were not parallel to changes in hepatic XOD levels [15]. Another study showed that serum XOD was correlated with uric acid levels, but not with hepatic XOD levels. Conversely, serum XOD levels were correlated with UA levels but not with hepatic XOD levels [18].
Rutin, Morin, Astilbin, anthocyanins significantly downregulated the mRNA and
protein levels of GLUT9 and URAT1, and Morin, Astilbin downregulated mRNA and
protein levels of OCT1, OCT2 in HUA mice. Anthocyanins, Phloretin downregulated
the expression of hepatic XOD, caspase-1, TNF-
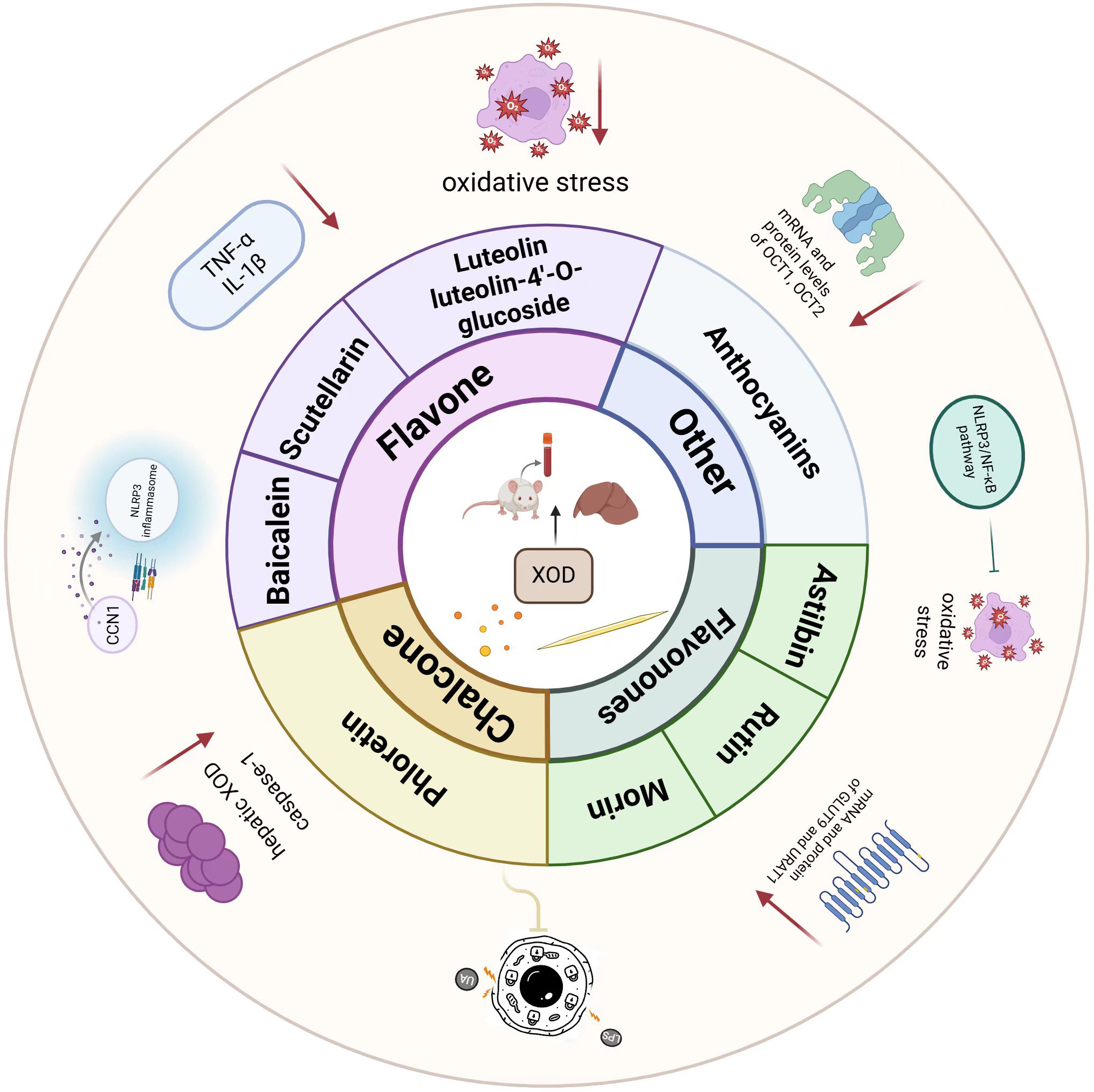 Fig. 6.
Fig. 6.
An overview of the molecular mechanisms underlying the
regulation of hyperuricemia by flavonoids. Created in BioRender. Han, Y. (2026) https://BioRender.com/cpzo8dj. The schematic
diagram illustrates the multi-targeted mechanisms by which various flavonoids
(e.g., Luteolin, Morin) ameliorate hyperuricemia. Key actions include the direct
inhibition of XOD activity, reduction of oxidative stress, and suppression of
inflammatory signaling pathways (e.g., NLRP3/NF-
Previous studies has demonstrated urate-lowering activity of dietary flavonoids, yet a quantitative, mechanism-based synthesis in validated murine hyperuricaemia (HUA) models was lacking. Our systematic review of 21 studies (550 male mice) shows that purified chalcones, flavonols and flavones consistently reduce serum uric acid (UA) and inhibit xanthine oxidase (XOD), with pooled effect sizes (SMD –2.22 and –1.79, respectively) that exceed the minimal important difference defined for animal models. Atomistic simulations corroborate stable binding of these subclasses to the molybdenum-pterin domain of XOD, providing an orthogonal line of evidence for competitive inhibition. Collectively, the data strengthen the mechanistic foundation for flavonoid-based XOD inhibitors as safer, cost-effective alternatives to allopurinol or febuxostat.
Included literatures mentioned that flavonoids exerted anti-HUA effect by
modulating hepatic production and renal excretion [39]. Specifically, flavonoids
acted via the Nrf2/HO-1/NQO1 and PI3K/AKT/NF-
Flavonoids are abundant in various foods, including vegetables, fruits, and tea, and are commonly found in traditional Chinese medicine (TCM) formulas for managing HUA [46]. Table 4 provided examples of TCM formulas that incorporate flavonoids for HUA prevention. Compared to conventional drugs such as allopurinol and febuxostat, flavonoids offer better tolerability and fewer adverse reactions [47]. Furthermore, their affordability and accessibility are also significant advantages, as they are widely available in daily diets at a low cost, facilitating patient acceptance and utilization [48]. Despite these benefits, several challenges remain in using flavonoids to treat HUA. The primary concern was their low bioavailability, which was mainly due to poor stability and limited membrane permeability [49]. The properties of flavonoids affected the pharmacokinetic process and lead to poor bioavailability [50]. Kaempferol was poorly absorbed into the bloodstream and rapidly metabolized into less active forms. Morin’s poor water solubility resulted in bioavailability of less than 1% [51]. Quercetin’s conjugation with plasma proteins also reduced its bioavailability [52]. Studies also had shown that the bioavailability of flavonoids in mice is relatively low, which may restrict their pharmacological effects and impact the accuracy and reproducibility of experimental results [53]. Furthermore, the optimal formulation of flavonoids for maximizing their pharmacological activity remained to be determined [54]. This necessitated further research and experimentation. Moreover, human clinical trials were essential to validate the efficacy and safety of flavonoids in treating HUA [55].
There are several strength points in the present study, including a comprehensive search strategy to find animal experiments on the effect of flavonoids on HUA, as well as subgroup analyses based on the level of UA and the activity of XOD. Despite statistical significance, the certainty of evidence is undermined by methodological limitations. None of the 21 included experiments employed allocation concealment or blinding, leading to an unclear risk of performance and detection bias. Empirical SYRCLE analyses indicate that lack of blinding inflates animal-study effect sizes by 8–13% [14]; consequently, our headline SMD of –2.22 may represent an upper-bound estimate. Future investigations should adopt sealed-envelope randomisation, blinded outcome assessment and pre-registration of protocols to minimise both performance bias and selective reporting. Moreover, many of the included studies also lack randomization and blinding, which are crucial for reducing bias and enhancing the rigor of the research. Furthermore, potential issues related to drug absorption and bioavailability are not addressed. Given these limitations, the results of this study should be interpreted with caution.
Species-specific differences in purine metabolism may limit the practicability
of these HUA models [8]. BALB/c mice were the only strain without a significant
UA reduction (SMD –0.61, 95% CI –1.30 to 0.09, p = 0.088). This
subgroup comprised only two comparisons from a single study, yielding extremely
wide confidence intervals and power
Despite pre-specified subgroup analyses, chalcones and “other flavonoids”
comparisons still exhibited I2
In summary, this systematic review and meta-analysis demonstrates that flavonoids derived from medicinal plants significantly reduce serum uric acid levels and inhibit xanthine oxidase activity in potassium oxonate-induced hyperuricemic mice. The therapeutic effects are particularly notable across flavonoid subclasses including chalcones, flavonols, and flavones. Molecular docking and dynamics simulations further support the stability and binding affinity of these compounds to XOD, reinforcing their potential as XOD inhibitors. These findings provide a robust preclinical evidence base supporting the development of flavonoid-based interventions for hyperuricemia. However, due to limitations such as study heterogeneity, potential publication bias, and lack of clinical validation, further well-designed clinical trials are warranted to confirm the efficacy, safety, and bioavailability of flavonoids in human populations.
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
MZ and YY conceived the study and designed the framework; MZ and QX developed the methodology; MZ and QX conducted literature retrieval, screening, data extraction and drafted the original manuscript; YH curated the data and performed formal analysis and visualization, with JX validating the results; CY performed the molecular dynamics simulations. YY reviewed and edited the manuscript. YY funded the project. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
The authors acknowledge Prof. Yi Wu and Prof. Dan Lu at Jilin University for the discussion of manuscript.
This work was supported by the Department of Science and Technology of Jilin Province, China (20220204118YY).
The authors declare no conflict of interest.
Statement: During the preparation of this work, the authors used Kimi in order to improve the readability and language of the manuscript. After using this service, the authors reviewed and edited the content as needed and take full responsibility for the content of the published article.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/IJP47870.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






