1 Department of Obstetrics and Gynecology, Zhongda Hospital, School of Medicine, Southeast University, 210009 Nanjing, Jiangsu, China
2 Department of Pharmacology, Jiangsu Provincial Key Laboratory of Critical Care Medicine, School of Medicine, Southeast University, 210009 Nanjing, Jiangsu, China
3 Department of Obstetrics and Gynecology, Nanjing Women and Children’s Healthcare Hospital, 210004 Nanjing, Jiangsu, China
Abstract
This research sought to examine alterations in plasma circular RNA (circRNA) expression in women with postpartum depression (PPD), assessing its potential utility as an auxiliary diagnostic biomarker for this condition.
Women examined at 42 days postpartum were recruited between June 2024 and December 2024, during which plasma samples and relevant data were collected. A discovery cohort was established, consisting of 3 women with PPD and 3 control participants. This was followed by a validation cohort of 50 women with PPD and 50 controls, identified using the Structured Clinical Interview. The discovery cohort underwent plasma circRNA microarray analysis, whereas the validation cohort used reverse transcription-quantitative polymerase chain reaction (RT-qPCR) to quantify candidate circRNAs and five circRNAs previously associated with major depressive disorder (MDD).
Plasma levels of circRNA derived from the Interferon Gamma Receptor 2 gene (circIFNGR2), circRNA derived from the ATP-Binding Cassette Subfamily C Member 5 gene (circABCC5), circRNA derived from the Activating Transcription Factor 7 Interacting Protein gene (circATF7IP) were significantly upregulated in the PPD group, whereas circRNA derived from the Dystrophia Myotonica Protein Kinase gene (circDYM) was significantly downregulated (p < 0.05). The area under the curve (AUC) of the four circRNAs, circIFNGR2, circABCC5, circDYM, and circATF7IP, was 0.62 (sensitivity 28%, specificity 96%), 0.64 (sensitivity 36%, specificity 96%), 0.69 (sensitivity 54%, specificity 84%), and 0.65 (sensitivity 62%, specificity 72%), respectively. The joint AUC of circIFNGR2, circABCC5, circDYM, and circATF7IP was 0.80 (sensitivity 78%, specificity 70%). After adjustment for covariates, circIFNGR2, circABCC5, and circATF7IP remained independent predictive factors.
Changes in the levels of circIFNGR2, circABCC5 identified by microarray analysis, as well as circDYM, and circATF7IP associated with MDD, were associated with the occurrence of PPD. These findings support the potential utility of circRNAs as adjunct diagnostic biomarkers for PPD and offer valuable insights into the molecular mechanisms underlying the disorder.
The study has been registered on https://zenodo.org/ (registration number: zenodo. 17906223; registration link: https://zenodo.org/records/17906223).
Keywords
- postpartum depression
- circular RNA
- plasma
- biomarker
Postpartum depression (PPD) encompasses depressive episodes of varying severity occurring within one year after childbirth. Persistent low mood, a marked reduction in interest or pleasure in activities, and feelings of fatigue or diminished energy constitute the three core symptoms [1]. As an obstetric complication, PPD represents a major contributor to maternal mortality worldwide [2, 3], with an estimated prevalence of approximately 23% [4]. PPD can severely impair maternal physical and mental health, disrupt family relationships, and exert long-term adverse effects on a child’s cognitive, emotional, and behavioral development [5, 6, 7, 8].
Recognition and diagnosis of PPD remain challenging. Postpartum changes in sleep, appetite, and energy related to infant care often overlap with depressive symptoms, making differentiation difficult. Additionally, some women are reluctant to report mood changes due to cultural stigma. Although the American College of Obstetricians and Gynecologists (ACOG) recommends routine depression screening at least once during pregnancy and the postpartum period, such screening is effective only when patients who screen positive receive proper diagnosis, treatment, or referral for appropriate medical care [9, 10]. Many obstetric clinicians are unable to provide a diagnosis of depression without psychiatric support, and one study reported a referral rate of only 8% among patients who screened positive [11]. Consequently, only a minority of patients access mental health services, and an even smaller proportion receives comprehensive treatment.
With advances in biomedical technology, an increasing number of studies are now exploring biological markers. In mammals, over 98% of gene transcripts are non-protein-coding RNAs (ncRNAs), which perform essential regulatory roles in diverse biological processes [12, 13]. Circular RNAs (circRNAs), a covalently closed-loop subtype of ncRNAs, were long regarded as splicing by-products due to their unique structure, unknown functions, and low expression levels. However, with advances in bioinformatics and high-throughput sequencing technologies, many circRNAs have been found to play roles in the pathogenesis of diverse neuropsychiatric disorders [14, 15, 16, 17]. Moreover, owing to their intrinsic resistance to exonucleases such as Ribonuclease R (RNase R), circRNAs exhibit superior stability to their linear counterparts [18], highlighting their potential as diagnostic biomarkers in clinical settings.
Given the diverse pathogenesis of PPD, we also include 5 circRNAs previously validated by our group as differentially expressed in major depressive disorder (MDD): circRNA derived from the Homeodomain Interacting Protein Kinase 2 gene (circHIPK2), circRNA derived from the Zinc Finger and BTB Domain Containing 25 gene (circZBTB25), circRNA derived from the Activating Transcription Factor 7 Interacting Protein gene (circATF7IP), circRNA derived from the Dystrophia Myotonica Protein Kinase gene (circDYM), and circRNA derived from the Stromal Antigen 1 gene (circSTAG1). Functional studies in animal models indicated that therapeutic intervention targeting these circRNAs effectively mitigated depressive-like behaviors in mice [19, 20, 21, 22]. These findings suggest that circRNAs have potential as objective diagnostic biomarkers for depression. To date, no comparable studies have examined this relationship in patients with PPD. This study aims to preliminarily explore and validate the potential association between plasma circRNA levels and PPD by analyzing samples collected from women at 42 days postpartum.
A discovery cohort of three women with PPD and three control participants, along
with a validation cohort (n = 50 with PPD, n = 50 controls), was recruited from
women who attended a 42-day postpartum follow-up visit at Zhongda Hospital and Nanjing Women and Children’s Healthcare Hospital between
June 2024 and December 2024. All participants completed the psychological
questionnaire, including the Edinburgh Postnatal Depression Scale (EPDS), Social
Support Rating Scale (SSRS), Pittsburgh Sleep Quality Index (PSQI), and
sociodemographic questionnaires under the guidance and supervision of a trained
physician, following predefined inclusion and exclusion criteria. The control
group was defined by EPDS scores (
Participant selection criteria were as follows: (1) age from 18 years to 45 years, who received routine prenatal care, delivery, and postpartum follow-up at the participating hospital; (2) no serious cardiovascular, neurological, pulmonary, or renal diseases; (3) currently breastfeeding and not taking any medication; and (4) submission of a signed informed consent form. Participant exclusion criteria were: (1) twin pregnancy; (2) pre-existing diabetes before conception; (3) pre-pregnancy hypertensive disorders and preeclampsia; (4) hyperthyroidism; (5) history of miscarriage or stillbirth; (6) clinical diagnosis of schizophrenia and bipolar disorder; (7) family history of mental illness; (8) lack of clinical information; and (9) missing questionnaire information and incomplete clinical information.
The EPDS was used to assess PPD in mothers and consisted of 10 questions with a
total score of 30 points. Scores
The SSRS was used to quantify an individual’s access to practical help,
emotional support, and active support-seeking from family, friends, and social
networks. It included 10 entries divided into three dimensions: objective support
(three entries), subjective support (four entries), and utilization of social
support (three entries). Higher scores indicated better social support. The
internal Cronbach’s
The PSQI is a widely employed instrument for assessing sleep disturbances in
clinical and research settings, encompassing seven dimensions and scored on a
0–21 scale where elevated scores denote poorer sleep quality. The internal
Cronbach’s
Sociodemographic information was collected through a custom-designed questionnaire in Chinese. This covered various domains, including age, marital status, ethnicity, educational status, religious belief, mood disorders, number of pregnancies, number of births, and mode of delivery. Clinical laboratory data covering the antenatal and postpartum periods were retrieved from the institutional electronic medical records. This encompassed assessments of thyroid and liver function from mid to late pregnancy, as well as complete blood count parameters during the postpartum period.
Venous blood samples (2 mL) were collected from the antecubital vein of each
participant in the early morning following an overnight fast, using K2EDTA-coated
vacuum blood collection tubes (ST750EK, Streck, NE, USA). After gentle
inversion, samples underwent centrifugation (1000
For circRNA microarray analysis, sample preparation and microarray hybridization were performed following the Arraystar microarray protocols (AS-S-CR-M-1.0 and ASLNC-MV3.0, Arraystar, TX, USA). To enrich circRNAs, total RNA samples were digested with RNase R (RNR07250, Epicentre, WI, USA) to remove linear RNAs. The enriched circRNAs were then amplified and transcribed into fluorescent circRNA utilizing a random priming method (Arraystar Super RNA Labeling Kit; AS-MS-0250, Arraystar, TX, USA). Labeled circRNAs were hybridized onto the Arraystar Human circRNA Array V2 (8x15K, Arraystar, TX, USA). After washing, the slides were scanned using the Agilent Scanner G2505C (G2505C, Agilent Technologies, CA, USA), and array images were analyzed with Agilent Feature Extraction software (version 11.0.1.1, Agilent Technologies, CA, USA).
Validation of differentially expressed circRNAs: 22 circRNAs identified as differentially expressed in the microarray analysis were selected for validation. Among them, 5 circRNAs (circDYM, circHIPK2, circZBTB25, circSTAG1, circATF7IP) that had previously been reported to be closely associated with MDD are included for validation using reverse transcription-quantitative polymerase chain reaction (RT-qPCR). In the validation cohort, total RNA was extracted from plasma using the miRNeasy® Serum/Plasma Kit (cat. No. 217184, Qiagen, MD, USA) according to the manufacturer’s instructions. RNA concentrations were detected using the One Drop spectrophotometer (Thermo Fisher Scientific, MA, USA). Reverse transcription was performed using the HiScript III All-in-one RT SuperMix Perfect for qPCR Kit (R333-C1, Vazyme, Nanjing, Jiangsu, China). The reaction system was configured, and amplification was carried out using SYBR Green Premix Pro Taq HS qPCR Tracking Kit (AG11733, Accurate Biology, Changsha, Hunan, China) following the manufacturer’s protocol, with cDNA as the template. Relative gene expression levels were calculated using the 2-ΔΔCT method. Controls were normalized for each experiment. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as an internal control. We employed the “reverse primer approach” by designing upstream and downstream primers on either side of the circRNA-specific antisense splicing site, allowing the primers to extend outward across the junction. Simultaneously, we verified the single peak in the melting curve to ensure primer specificity. Primers were synthesized by Geneary (Shanghai, China), and the primer sequences are shown in Supplementary Table 1.
SPSS version 26.0 (International Business Machines Corporation, Armonk, NY, USA)
and GraphPad Prism version 8.0.2 (GraphPad Software, Boston, MA, USA) were used
for data analysis. The normality of continuous variables was assessed using the
Shapiro-Wilk test, while the categorical variables were analyzed using the
chi-square test. For continuous variables, an independent samples t-test
was employed when the data were normally distributed; otherwise, the Mann-Whitney
U-test was used. Spearman’s rank correlation was utilized to examine linear
relationships between levels of circRNAs and severity of PPD, degree of social
support, and sleep condition. To adjust for potential confounders, both
univariate and multivariate logistic regression models were employed, reporting
odds ratios (ORs) with corresponding 95% confidence intervals (CIs). The
diagnostic efficacy of candidate circRNAs for PPD was assessed by the area under
the curve (AUC) of the receiver operating characteristic (ROC), with the Youden
index determining the optimal sensitivity and specificity cutoff. Statistical
significance was defined as a two-tailed p-value
In the discovery cohort, the mean ages of the PPD and control groups were 30.33
| Characteristics | N (n = 50) | PPD (n = 50) | t/U/ |
p-value | |
| Demographic characteristics | |||||
| Age (years), mean (SD) | 30.54 (3.62) | 31.28 (3.81) | –0.99 | 0.32 | |
| Married, n (%) | 48.00 (96.00) | 50.00 (100.00) | - | 0.50$ | |
| Han nationality, n (%) | 47.00 (94.00) | 48.00 (96.00) | - | 1.00$ | |
| Undergraduate and junior college, n (%) | 32.00 (64.00) | 38.00 (76.00) | 2.79 | 0.25 | |
| Master’s degree or above, n (%) | 15.00 (30.00) | 8.00 (16.00) | |||
| Junior high school, vocational high school, and below, n (%) | 3.00 (6.00) | 4.00 (8.00) | |||
| Having religious belief, n (%) | 2.00 (4.00) | 5.00 (10.00) | - | 0.44$ | |
| A history of pre-pregnancy anxiety or depression, n (%) | 0.00 (0.00) | 6.00 (12.00) | - | 0.03$ | |
| Gravidity, median (interquartile range) | 1.00 (1.00, 2.00) | 1.00 (1.00, 2.00) | 1249.50 | 1.00 | |
| Parity, median (interquartile range) | 1.00 (1.00, 1.00) | 1.00 (1.00, 1.00) | 1225.50 | 0.79 | |
| Planning pregnancy, n (%) | 39.00 (78.00) | 36.00 (72.00) | 0.48 | 0.49 | |
| Natural conception, n (%) | 47.00 (94.00) | 40.00 (80.00) | 4.33 | 0.04 | |
| Exercise during pregnancy, n (%) | 46.00 (92.00) | 36.00 (72.00) | 6.78 | 0.01 | |
| Spontaneous labor, n (%) | 20.00 (40.00) | 30.00 (60.00) | 4.00 | ||
| Gestational weeks at delivery, median (interquartile range) | 39.79 (38.71, 40.18) | 39.14 (38.71, 40.00) | 1070.50 | 0.22 | |
| Postpartum BMI (kg/m²), mean (SD) | 23.10 (2.87) | 23.37 (3.08) | –0.45 | 0.66 | |
| Newborn is a boy, n (%) | 23.00 (46.00) | 27.00 (54.00) | 0.64 | 0.42 | |
| Laboratory parameters, mean (sd)/median (interquartile range) | |||||
| WBC, (109/L) | 6.26 (5.71, 7.30) | 6.62 (6.03, 7.93) | 964.50 | ||
| RBC, (1012/L) | 4.48 (4.33, 4.67) | 4.58 (4.42, 4.75) | 1040.00 | 0.15 | |
| HGB, (g/L) | 132.50 (127.00, 138.00) | 134.50 (128.00, 139.00) | 1147.50 | 0.48 | |
| Platelets, (109/L) | 240.50 (207.50, 275.00) | 243.00 (205.00, 278.25) | 1223.00 | 0.85 | |
| NEUT count, (109/L) | 3.48 (3.08, 4.23) | 3.90 (3.21, 4.97) | 991.00 | 0.07 | |
| LYMPH count, (109/L) | 2.17 (1.89, 2.55) | 2.32 (2.08, 2.63) | 1064.50 | 0.20 | |
| MONO count, (109/L) | 0.36 (0.31, 0.42) | 0.37 (0.31, 0.45) | 1184.00 | 0.65 | |
| ALB, (g/L) | 35.65 (34.15, 37.30) | 36.20 (34.60, 37.65) | 1092.00 | 0.28 | |
| TC, (mmol/L) | 6.85 (5.96, 7.56) | 7.15 (6.43, 7.96) | 959.50 | 0.06 | |
| ALP, (U/L) | 148.85 (115.63, 186.80) | 139.55 (109.75, 183.33) | 1179.00 | 0.63 | |
| LDH-L, (U/L) | 179.45 (161.40, 199.43) | 181.65 (166.00, 196.03) | 1194.50 | 0.70 | |
| TSH, (µIU/mL) | 1.77 (1.29, 2.38) | 2.25 (1.48, 3.00) | 934.00 | 0.04 | |
| FT4, (pmol/L) | 13.05 (11.80, 14.40) | 12.70 (11.95, 13.60) | 1082.00 | 0.32 | |
| Anti-TPO, (IU/mL) | 13.25 (10.68, 18.88) | 12.90 (9.00, 20.90) | 1158.00 | 0.63 | |
| Questionnaire | |||||
| PSQI score, median (interquartile range) | 4.00 (1.00, 5.25) | 9.00 (7.00, 11.00) | 216.00 | ||
| SSRS score, mean (sd) | 42.12 (4.72) | 33.78 (5.63) | 8.03 | ||
| Positive self-harm intention, n (%) | 0.00 (0.00) | 30.00 (60.00) | - | ||
Note: N, control group; t, Student’s t-test; U, Mann-Whitney U test;
Abbreviations: PPD, postpartum depression; SD, standard deviation; BMI, body mass index; WBC, white blood cell; RBC, red blood cell; HGB, hemoglobin; NEUT, neutrophil; LYMPH, lymphocyte; MONO, monocyte; ALB, albumin; TC, total cholesterol; ALP, alkaline phosphatase; LDH-L, lactate dehydrogenase-L form; TSH, thyroid-stimulating hormone; FT4, free thyroxine; Anti-TPO, anti-thyroid peroxidase antibody; SSRS, Social Support Rating Scale; PSQI, Pittsburgh Sleep Quality Index.
| Variable | B | SE | OR (95% CI) | p-value |
| A history of pre-pregnancy anxiety or depression | 2.69 | 1.48 | 14.75 (1.67, 1940.94) | 0.01# |
| Assisted reproduction | 1.37 | 0.69 | 3.92 (1.01, 15.22) | |
| Cesarean section delivery | –0.81 | 0.41 | 0.44 (0.20, 0.99) | |
| Exercise during pregnancy | –1.50 | 0.61 | 0.22 (0.07, 0.74) | 0.01 |
| WBC, (109/L) | 0.36 | 0.16 | 1.44 (1.05, 1.98) | 0.03 |
| TSH, (µIU/mL) | 0.41 | 0.21 | 1.51 (1.00, 2.27) | |
| SSRS scores | –3.00 | 0.06 | 0.74 (0.66, 0.83) | |
| PSQI scores | 0.79 | 0.16 | 2.21 (1.62, 3.03) |
Abbreviations: OR, odds ratio; CI, confidence interval; #, perform a firth regression test.
CircRNA microarrays were used to compare plasma circRNA levels between PPD
patients and matched controls in the discovery cohort. Among the 7683 circRNAs
detected by microarray profiling (Fig. 1A–C), 22 were found to be differentially
expressed between the PPD group and the control group (
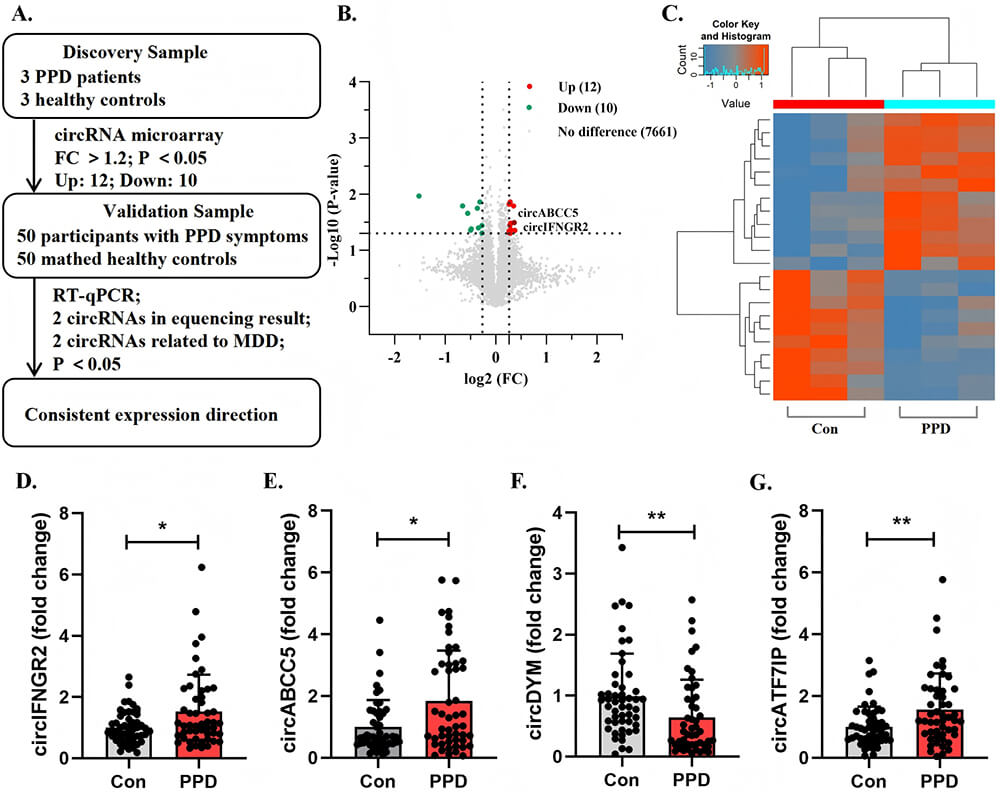 Fig. 1.
Fig. 1.
Screening and validation of differentially expressed
circular RNAs. (A) Experimental flowchart. (B,C) Volcano plot and heatmap of
microarray analysis results. (D–G) Differentially expressed circRNAs verified by
RT-qPCR in the validation cohort. (D) Relative expression levels of circIFNGR2.
(E) Relative expression levels of circABCC5. (F) Relative expression levels of
circDYM. (G) Relative expression levels of circATF7IP. Note: * p
Spearman correlation analysis demonstrated that circIFNGR2 was not correlated with EPDS scores (r = 0.123, p = 0.224), SSRS scores (r = –0.158, p = 0.117), or PSQI scores (r = 0.152, p = 0.132). CircABCC5 was not correlated with EPDS scores (r = 0.192, p = 0.056) or SSRS scores (r = –0.095, p = 0.350) but was positively associated with PSQI scores (r = 0.199, p = 0.047). CircDYM was not significantly correlated with EPDS scores (r = –0.155, p = 0.123) but was positively associated with SSRS scores (r = 0.219, p = 0.029) and negatively associated with PSQI scores (r = –0.248, p = 0.013). CircATF7IP was positively correlated with EPDS scores (r = 0.236, p = 0.018) and PSQI scores (r = 0.337, p = 0.0006) and negatively associated with SSRS scores (r = –0.306, p = 0.002) (Fig. 2). Further studies indicate that only circABCC5 level shows a positive correlation with WBC count (r = 0.205, p = 0.041) (Supplementary Fig. 1).
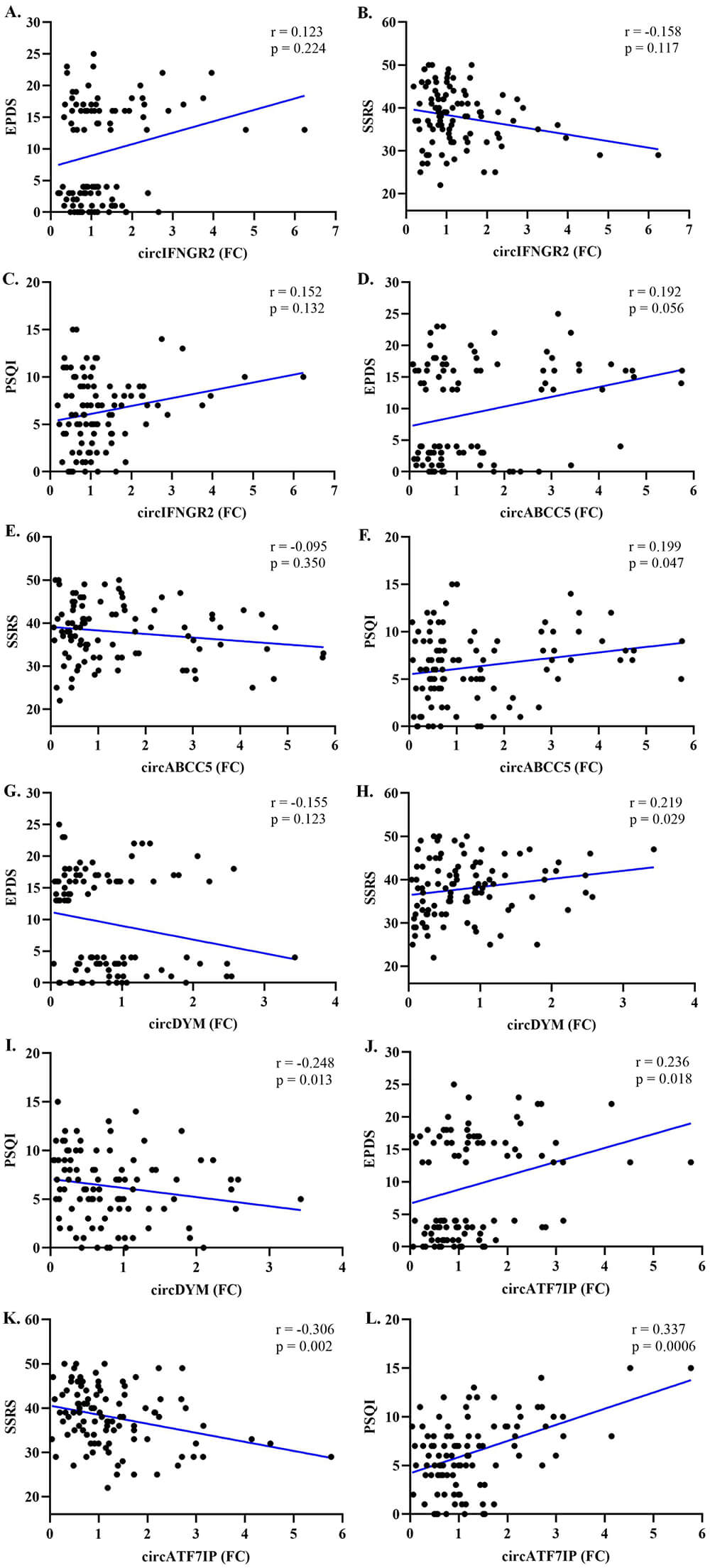 Fig. 2.
Fig. 2.
Correlation between plasma circIFNGR2, circABCC5, circDYM and circATF7IP levels and questionnaire assessments in participants. (A,D,G,J) Correlation between four differentially expressed circular RNAs and EPDS scores. (B,E,H,K) Correlation between four differentially expressed circular RNAs and SSRS scores. (C,F,I,L) Correlation between four differentially expressed circular RNAs and PSQI scores. Abbreviations: EPDS, Edinburgh Postnatal Depression Scale.
CircIFNGR2 showed an AUC of 0.62 (95% CI: 0.51–0.73, p
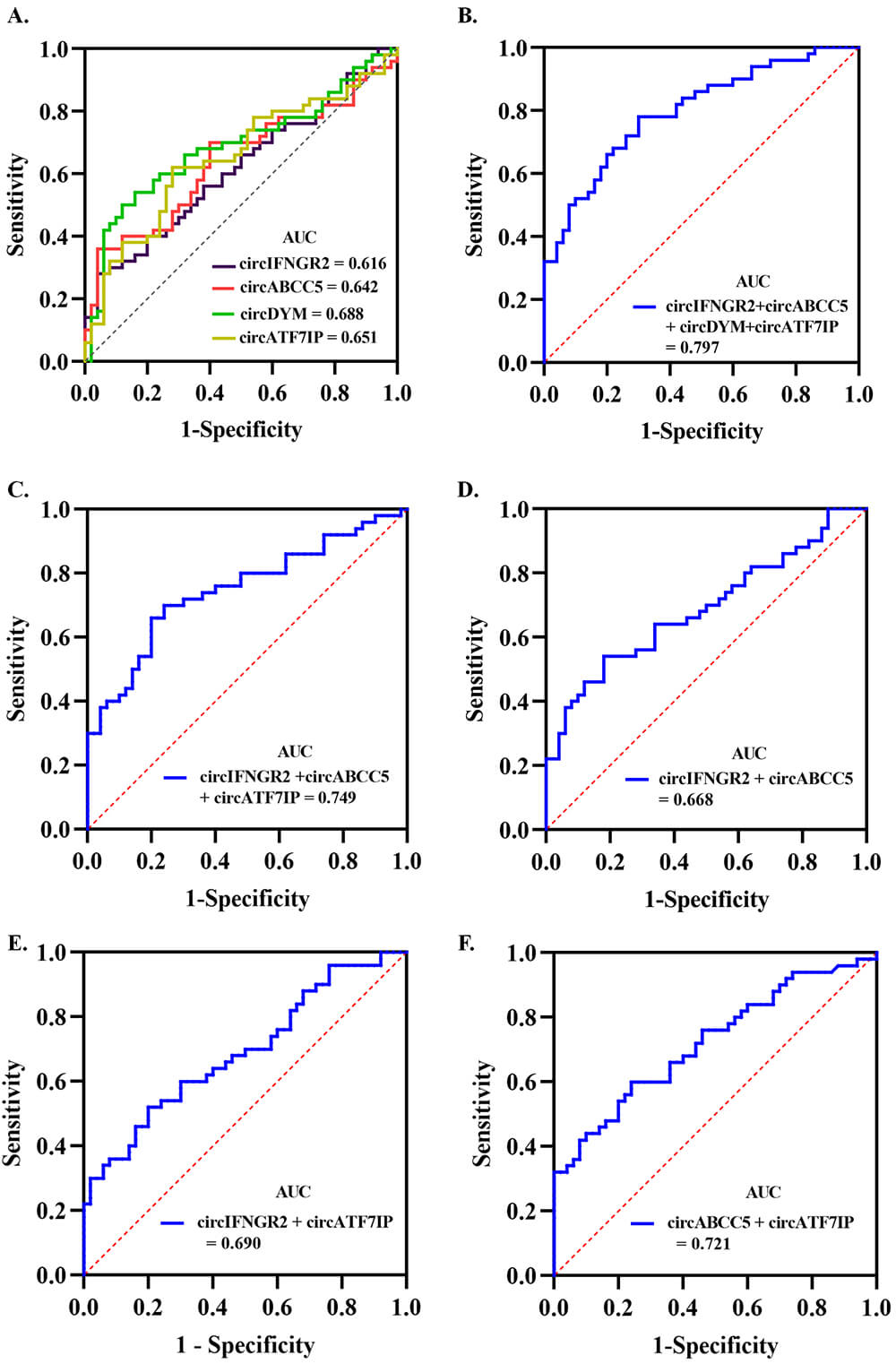 Fig. 3.
Fig. 3.
AUC for ROC analysis of circRNAs that distinguish individuals with PPD symptoms from a normal control population. (A) AUC for the independent prediction of PPD by 4 circRNAs. (B) AUC for the combined prediction of PPD by 4 circRNAs. (C) AUC for the combined prediction of PPD by 3 circRNAs identified as relatively independent predictive factors. (D–F) AUC for the combined prediction of PPD by 2 circRNAs in each combination. Abbreviations: AUC, area under the curve; ROC, receiver operating characteristic.
| Variables | OR (95% CI) | p-value | Adjusted OR (95% CI) | p-value |
| circIFNGR2 | 2.10 (1.17, 3.76) | 0.01 | 2.26 (1.21, 4.23) | 0.01 |
| circABCC5 | 1.71 (1.19, 2.45) | 0.01 | 2.09 (1.35, 3.23) | |
| circDYM | 0.43 (0.22, 0.84) | 0.01 | 0.50 (0.23, 1.10) | 0.84 |
| circATF7IP | 2.05 (1.21, 3.46) | 0.01 | 1.74 (1.01, 3.02) |
PPD is widely recognized as a multifactorial disorder involving biological, psychological, and social factors [23, 24]; however, its precise pathogenesis remains unclear. Investigating circRNAs as auxiliary diagnostic markers for PPD may help overcome the subjectivity and limitations inherent to screening questionnaires, offering an objective molecular basis for diagnosis. In this study, individual circRNAs showed AUC values ranging from 0.62 to 0.69, with low sensitivity (28%–62%) but high specificity (72%–96%). When the 4 circRNAs were combined into a diagnostic panel, the AUC increased to 0.80, with a sensitivity of 78% and specificity of 70%. Furthermore, comparing diagnostic models with 2 versus 3 circRNAs revealed that increasing the number of circRNAs in the model gradually improved both the AUC and sensitivity. These findings indicate that although individual circRNAs possess diagnostic potential, their discriminative power alone is limited, likely reflecting the complex pathogenic mechanisms underlying PPD. The combined use of 4 circRNAs significantly enhanced the diagnostic performance, as reflected by the increased AUC.
The EPDS is the most widely employed tool for PPD screening in both clinical and research settings. Although previous study has reported that an EPDS cutoff score of 10 can achieve sensitivity and specificity exceeding 80% [25], it remains a symptom-based self-report measure. Cultural stigma may lead some individuals to underreport or conceal their symptoms, resulting in the underestimation of true prevalence. In contrast, the predictive model developed in this study, based on multiple circRNAs, allows more objective identification of individuals at risk for PPD symptoms.
Beyond improving diagnostic objectivity, understanding the molecular basis of PPD is essential for elucidating its underlying mechanisms. The neuroimmune-inflammatory hypothesis has garnered increasing attention in depression research. As the resident immune cells of the brain, microglia play a pivotal role in driving neuroinflammation through phenotypic shifts and the release of inflammatory factors. Observations of activated microglia in specific brain regions, ranging from animal models of depression [26] to human clinical studies [27, 28], support the notion that microglia-mediated neuroinflammation is a key mechanism in MDD [29]. Importantly, circRNAs have emerged as key regulators within this pathway. For instance, circDYM acts as a miR-9 sponge, suppressing microglial activation through heat shock protein 90 (HSP90) ubiquitination, ultimately ameliorating depression-like behaviors [30]. Similarly, downregulation of circATF7IP has been shown to attenuate microglia-induced neuroinflammation and alleviate depression-like behavior in mice [20].
In our study, correlation analysis revealed that circABCC5 levels positively
correlated with WBC count (p
Several limitations should be considered when interpreting our findings. First, our study included a small sample size. Due to the low incidence of PPD and challenges in sample collection, a 3-to-3 sample ratio was adopted for microarray analysis in the discovery cohort. Although the included samples were sufficiently representative, including individuals with severe depressive symptoms and self-harm tendencies who subsequently required hospitalization, the small sample size still poses a risk of false positives. Given the exploratory nature of this study, we validated as many relevant circRNAs as possible in the validation cohort, including those associated with MDD. The generalizability and reliability of these findings still require validation in larger, multicenter, multiethnic cohorts. Second, the dynamic changes of circRNA levels during the course of the disease cannot be fully captured by the single-timepoint sampling approach used in this study. Future research should adopt longitudinal sampling strategies to monitor circRNA expression profiles at multiple stages of pregnancy and the postpartum period. Lastly, the specific pathophysiologic mechanisms through which different circRNAs contribute to PPD require additional exploration.
In this study, we identified significant associations between circulating circRNA levels and PPD. Overall, our findings support the potential use of circRNAs as adjunct diagnostic biomarkers for PPD and offer valuable insights into the molecular mechanisms underlying the disorder.
The data that support the findings of this study are available from the corresponding author upon reasonable request and have been publicly deposited in Zenodo (https://zenodo.org/records/17906223).
FZ was responsible for the experimental operations, data organization and article writing. YZ was responsible for the experimental design and guidance. DQ was responsible for the collection and processing of clinical samples. HY was responsible for organizing the experimental results, making tables and pictures. XW was responsibe for data collection and article revision. HY was responsible for research design, revision, paper review and guidance. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
This study performed in accordance with the “Declaration of Helsinki” and approved by the Ethics Committee of Zhongda Hospital (2023ZDSYLL438-p01) and the Ethics Committee of Nanjing Women and Children’s Healthcare Hospital (2024KY-080-01). Informed consent was obtained from all participants prior to data collection.
We are deeply grateful to Dr. Zhengcheng Tu for language polishing and revision of the article.
This study was supported by the Natural Science Foundation of Jiangsu Province (Project No. BK20231423).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/CEOG47150.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.



