1 Department of Pathology, Affiliated Hospital 2 of Nantong University, 226001 Nantong, Jiangsu, China
2 Faculty of Medicine, Macau University of Science and Technology, 999078 Macau, China
Abstract
Primary ovarian insufficiency (POI), characterized by infertility and an elevated risk of multiple comorbidities, affects approximately 1% of women <40 years of age. Although mitochondrial dysfunction has been associated with POI, the specific mitochondrial proteins involved in its pathogenesis remain largely unidentified. To address this gap, the present study investigated the causal relationship between mitochondrial proteins and POI using a bidirectional Mendelian randomization (MR) approach.
Bidirectional MR analysis was conducted using genetic data derived from the INTERVAL and FinnGen databases. Plasma data for 3622 proteins, including 66 mitochondrial proteins, were examined. Genetic instruments were selected based on a stringent genome-wide significance threshold, and causal associations were estimated using the inverse-variance weighted method.
The analysis revealed that higher levels of 39S ribosomal protein L14 (odds ratio [OR] = 0.40, p = 0.009), oligoribonuclease (OR = 0.64, p = 0.031), and mitochondrial fission regulator 1 (OR = 0.60, p = 0.044) were significantly associated with a reduced risk POI. In contrast, higher levels of coiled-coil domain-containing protein 90B were associated with an increased risk of POI (OR = 1.80, p = 0.042).
This study identified key mitochondrial proteins associated with a reduced risk of POI, highlighting the potential role of mitochondrial pathways in POI pathogenesis and offering possible targets for future diagnostic and therapeutic interventions.
Keywords
- primary ovarian insufficiency
- mitochondrial dysfunction
- mendelian randomization
- mitochondrial ribosomal protein l14
- genome-wide association studies (GWAS)
Primary ovarian insufficiency (POI), previously referred to as premature ovarian failure, is a clinical condition characterized by a broad spectrum of manifestations and affects approximately 1% of women under 40 years of age [1, 2]. The condition is characterized by amenorrhea, hypergonadotropic hypogonadism, and hypoestrogenism, which has significant implications for both reproductive and systemic health [3]. POI has profound impacts on female fertility and is a major cause of infertility in affected individuals. Additionally, its onset has been associated with an increased risk of osteoporosis, cardiovascular disease, and reduced quality of life [4, 5]. Despite its clinical significance, the underlying pathogenesis of POI remains unclear, and effective therapeutic options remain limited [6].
One promising area of investigation involves the role of mitochondrial function in POI [7]. Mitochondria, the primary site of ATP production, play a pivotal role in apoptosis regulation and intracellular signaling [8]. An increasing number of studies indicate that mitochondrial dysfunction may contribute to the development of POI. For example, oxidative stress, mitochondrial DNA (mtDNA) mutations, and impaired mitochondrial biogenesis may result in accelerated depletion of ovarian follicles [9]. Furthermore, specific mitochondrial proteins appear to be involved in folliculogenesis and oocyte survival, suggesting that mitochondrial dysfunction may be a crucial element in the etiology of POI [10].
Nevertheless, despite these insights, significant gaps remain in our understanding. Current research has not yet fully elucidated the specific mitochondrial proteins implicated in the development and progression of POI. Identifying these proteins could facilitate the development of novel biomarkers for early diagnosis and potential targets for therapeutic intervention. Most existing studies are limited by observational designs, highlighting the need for robust methodologies capable of establishing causality, such as Mendelian randomization (MR) [11, 12].
In this study, we employed a bidirectional MR approach, using genetic data from the INTERVAL study and the FinnGen database, to investigate the association between mitochondrial proteins and the risk of POI [13, 14]. The objective of this study was to identify mitochondrial proteins that influence the development of POI and are significantly affected by the condition. By applying genetic instrumental variable (IV) methods, we aimed to overcome the confounding biases that have previously limited progress in this field and to establish more robust causal inferences. These findings may contribute to the development of novel diagnostic tools and therapeutic strategies, offering potential advances in the clinical and scientific management of POI.
MR employs genetic variants as instrumental proxies to assess causal relationships between exposures and outcomes. This method is based on three fundamental assumptions: instrument relevance, instrument independence, and exclusion-restriction criterion [15]. In accordance with these principles, our study employed an MR framework to investigate the bidirectional causal effects of mitochondrial proteins on the risk of POI, as illustrated in Fig. 1. This approach allows for the examination of the complex interplay between genetic and environmental factors, providing a more nuanced understanding of how mitochondrial proteins may influence POI development and offering insights that could guide future interventions.
 Fig. 1.
Fig. 1.
Study design for MR analysis. The analysis is based on three
core hypotheses: (1) a strong association between IVs
and exposure factors, (2) no association between IVs and confounders, and (3) IVs
influence the outcome solely through the exposure factors and not through other
pathways. In this study, genome-wide statistical significance was as p
To investigate mitochondrial proteins, plasma protein data for 3622 proteins from 3301 healthy participants in the INTERVAL (https://www.intervalstudy.org.uk/) study were analyzed. The INTERVAL study is a prospective cohort of predominantly European-ancestry blood donors in the United Kingdom, as previously described [13]. This dataset includes 1927 genotype-protein associations (pQTLs), encompassing trans-associated loci for 1104 proteins [16]. This study provides new insights into the genetic regulation of protein expression. Subsequently, 66 datasets corresponding to mitochondrial proteins were screened. In parallel, POI was examined using the dataset labeled finn-b-E4_OVARFAIL, sourced from the Finnish FinnGen database (https://r8.risteys.finngen.fi/phenocode/E4_OVARFAIL), which includes data collected up to 2021 [17]. The analysis included 429,209 individuals of Finnish ancestry. After applying sex-specific filters (females only), the effective sample size was 239,906 individuals, comprising 542 POI cases and 239,364 controls. POI cases were identified based on nationwide health registries, including hospital discharge records, cause-of-death data, and drug reimbursement records, based on relevant International Classification of Diseases (ICD) codes (ICD-10: E28.3). The use of this dataset guarantees that our investigation focuses on the population most affected by POI, thus enhancing the relevance of the findings.
To further satisfy the MR independence and exclusion-restriction assumptions,
each single nucleotide polymorphism (SNP) and its high-linkage disequilibrium
(LD) proxies were systematically screened for prior genome-wide significant
associations with potential confounders or traits related to the outcome.
Specifically, for every IV retained after clumping and F-statistic filtering, we
queried LDlinkR (LDtrait function) using European reference populations, defining
high-LD proxies as those with r2
SNPs associated with mitochondrial protein levels were selected as IVs through
rigorous screening. To ensure an adequate number of IVs while capturing relevant
genetic variants associated with mitochondrial protein levels, IVs were required
to show a genome-wide significant association with the exposure (p
Furthermore, palindromic and proxy SNPs were removed to prevent inaccuracies in
the MR analysis. To mitigate potential weak instrument bias, an F-statistic
threshold of
Statistical analyses were conducted using R software (version 4.2.2; R
Foundation for Statistical Computing, Vienna, Austria). The following R packages
were used: ggplot2 (version 3.4.0; tidyverse), LDlinkR (version 3.1.3; National
Cancer Institute, Division of Cancer Epidemiology & Genetics, Bethesda, MD, USA),
MendelianRandomization (version 0.5.1), MRPRESSO (version 1.0; Broad Institute of
MIT and Harvard, Cambridge, MA, USA), and TwoSampleMR (version 0.6.17; MRC
Integrative Epidemiology Unit, University of Bristol, Bristol, UK). Analyses
were repeated after excluding outlier SNPs (Supplementary Tables 1,2).
The primary method used was the Inverse Variance Weighted (IVW) approach, which
is known for its robustness in the absence of pleiotropy [22]. To address
potential biases from horizontal pleiotropy, several sensitivity analyses were
performed, including MR-PRESSO, MR-Egger, and Weighted Median regression. The
MR-Egger intercept p-value
MR analyses using the IVW method revealed significant associations between
specific mitochondrial proteins and POI risk (Figs. 2,3). Among the 66
mitochondrial proteins tested, 4 demonstrated significant causal associations
with POI risk. Coiled-coil domain-containing protein 90B was positively
associated with POI risk (odds ratio [OR] = 1.80, 95% confidence interval [CI]:
1.02–3.19, p = 0.042), indicating increased risk. In contrast, 3
proteins demonstrated protective associations. For 39S ribosomal protein L14
(mitochondrial), the IVW method revealed a significant inverse association with
POI (OR = 0.40, 95% CI: 0.20–0.80, p = 0.009), suggesting a protective
effect. This association, with a beta (
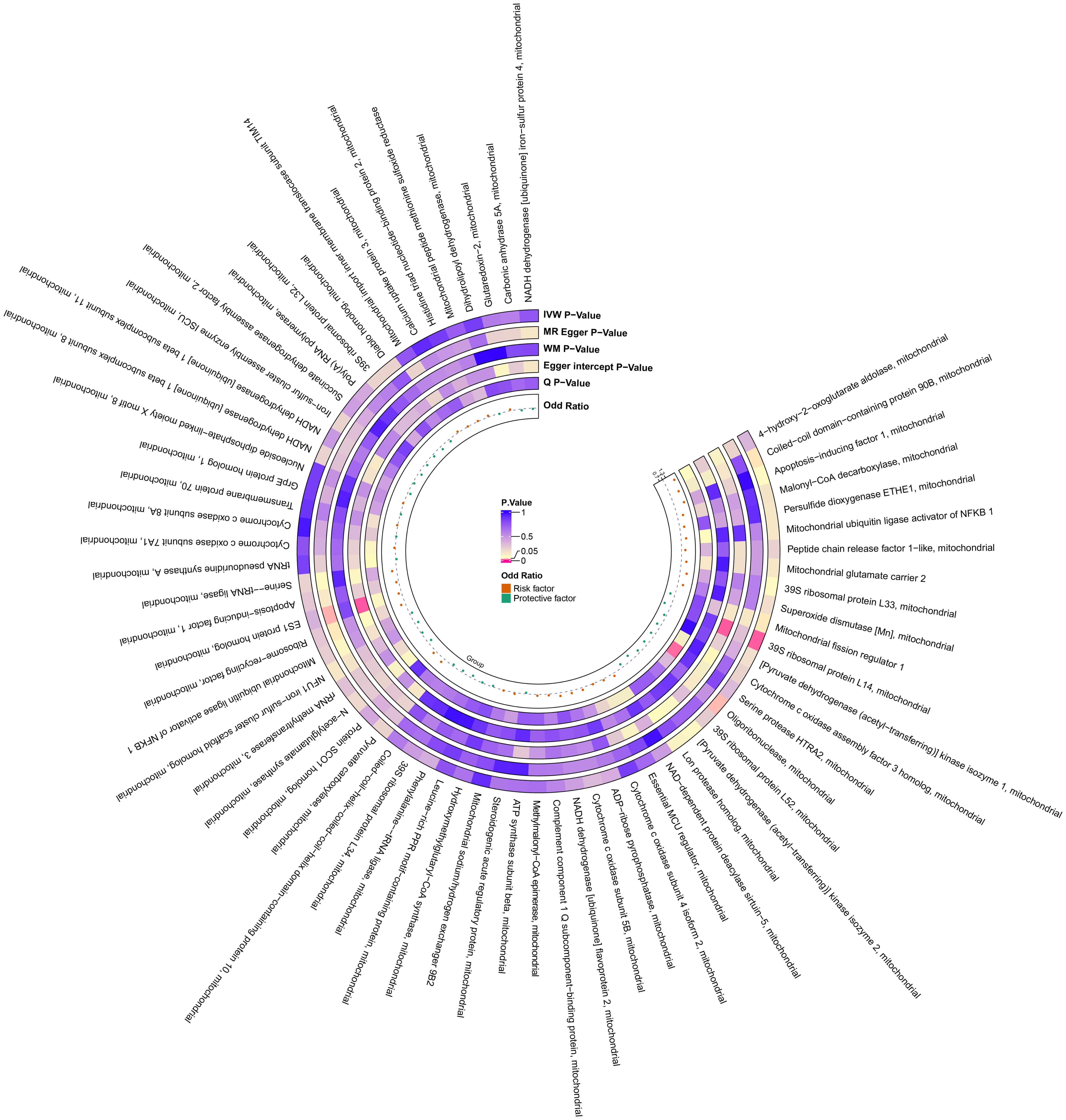 Fig. 2.
Fig. 2.
Preliminary MR estimates of the effects of mitochondrial proteins on POI. From the inner to outer circles, the estimates represent: Beta-value, Q p-value, MRPRESSO global test p-value, Egger intercept p-value, WM p-value, MR-Egger p-value, and IVW p-value, respectively. The color intensity reflects the magnitude of the p-value. POI, primary ovarian insufficiency; WM, weighted median.
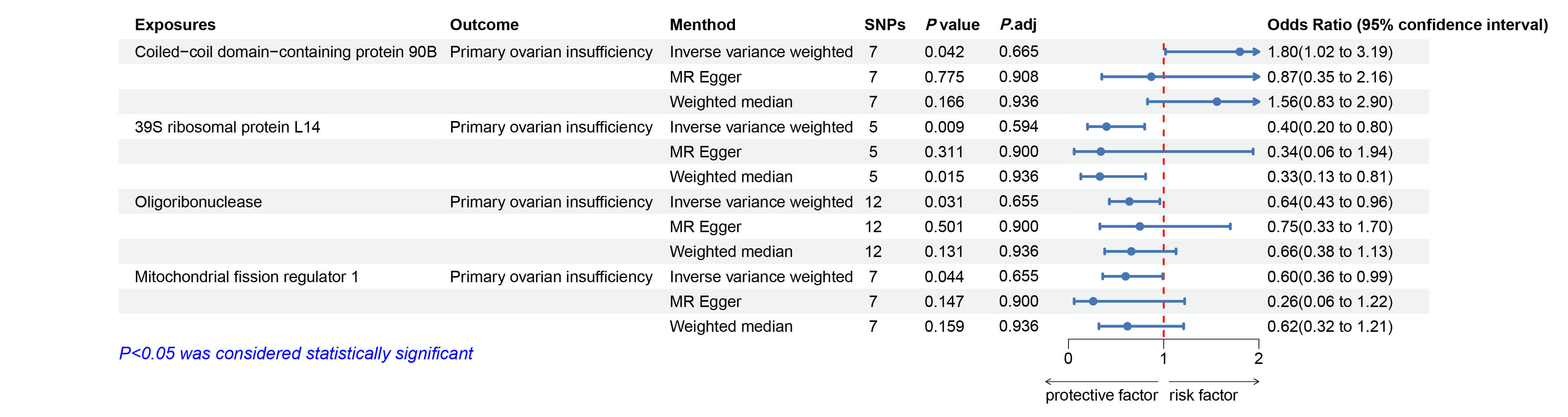 Fig. 3.
Fig. 3.
Forest plots illustrating the causal relationship between
mitochondrial proteins and POI. Significant results (p
Similarly, oligoribonuclease (mitochondrial) was significantly associated with POI, with the IVW analysis yielding an OR of 0.64 (95% CI: 0.43–0.96, p = 0.031), suggesting a protective effect. Finally, mitochondrial fission regulator 1 showed a significant inverse association with POI using the IVW method (OR = 0.60, 95% CI: 0.36–0.99, p = 0.044), indicating that this mitochondrial protein may also reduce the risk of POI.
In summary, the IVW analysis consistently suggests that higher levels of 39S ribosomal protein L14, oligoribonuclease, and mitochondrial fission regulator 1 are associated with a reduced risk of POI, whereas higher levels of coiled-coil domain-containing protein 90B may increase the risk. However, after applying the BH-FDR correction, the strength of the associations was attenuated. Therefore, these findings shouldbe interpreted with caution. No significant heterogeneity, pleiotropy, reverse causality, or outlier IVs were detected for the effect of mitochondrial proteins on POI (Supplementary Table 1). Scatter, funnel, and forest plots illustrating these associations and sensitivity analyses are provided in Supplementary Fig. 1.
The impact of POI on several mitochondrial proteins was evaluated using the IVW
method (Figs. 4,5). While associations were observed, the effect sizes were
consistently small, suggesting a limited biological impact. For
apoptosis-inducing factor 1 (mitochondrial), a significant negative association
was found (
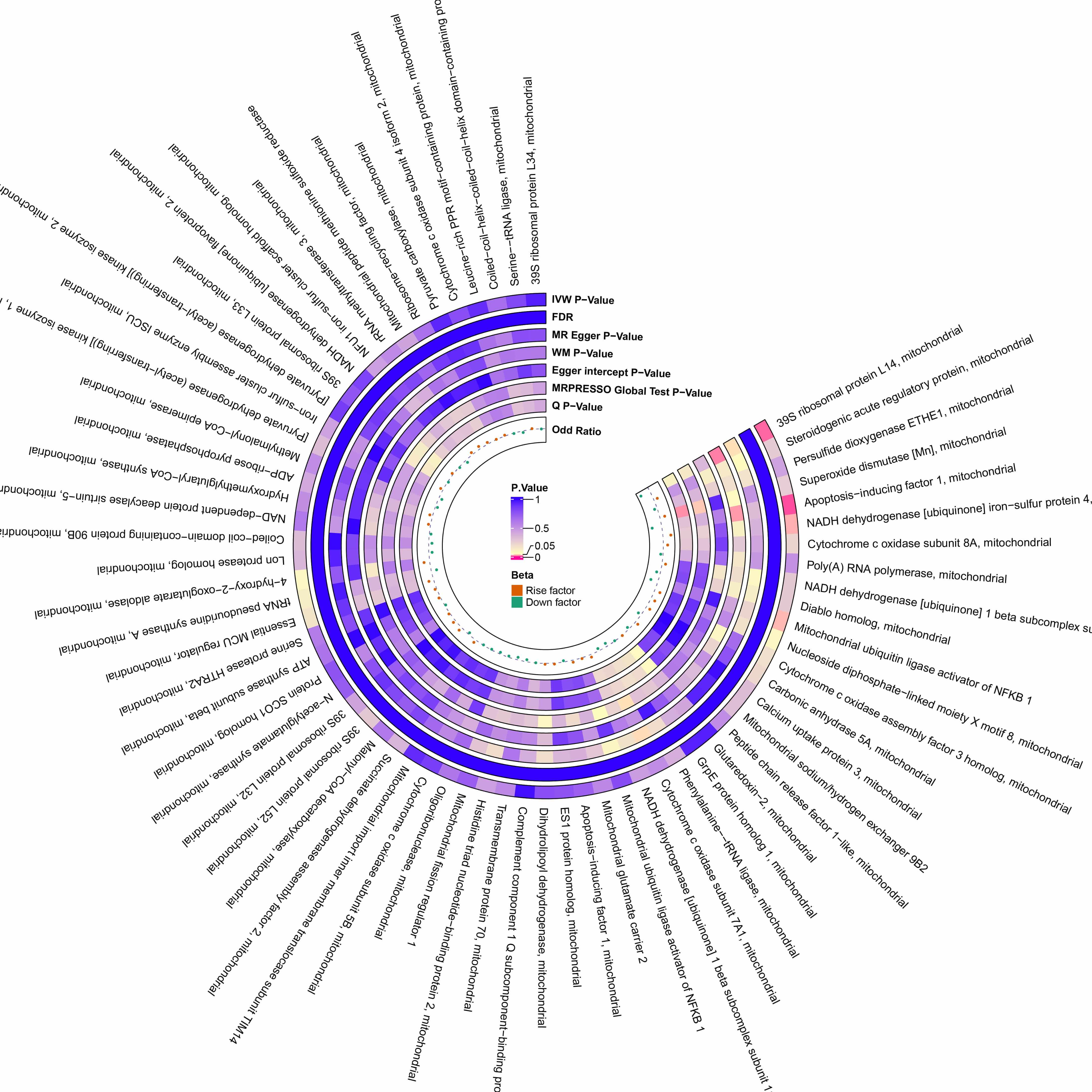 Fig. 4.
Fig. 4.
Preliminary MR estimates of the effect of POI on mitochondrial proteins. From the inner to outer circles, the estimates represent: Beta-value, Q p-value, MRPRESSO global test p-value, Egger intercept p-value, WM p-value, MR-Egger p-value, and IVW p-value, respectively. The color intensity reflects the magnitude of the p-value.
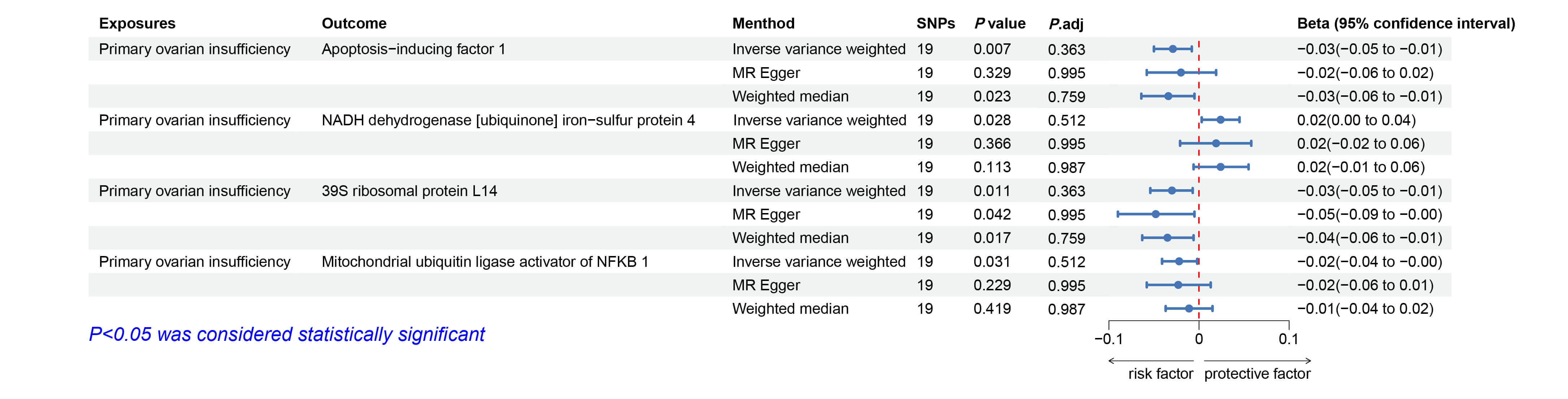 Fig. 5.
Fig. 5.
Forest plots illustrating the causal relationship between POI
and mitochondrial proteins. Significant results (p
Our analyses identified several drugs potentially involved in the regulation of mitochondrial proteins, focusing on their interactions with three protected genes: RNASEH1, MTFR1 and MRPL14. The principal findings of this prediction are summarized in Table 1, which lists the drugs along with their statistical significance, effect size, combined score, and associated cytokine targets [24]. This table highlights drugs that may modulate mitochondrial protein regulation, suggesting potential therapeutic applications in POI.
| Term | p-value | OR | Combined Score | Target |
| ETHYLMETHANESULFONATECTD00005938 | 0.002 | 53,055.00 | 343,272.96 | RNASEH1;MTFR1;MRPL14 |
| hypericinCTD00000183 | 0.003 | 499.43 | 2877.37 | RNASEH1 |
| jugloneTTD00008752 | 0.004 | 416.10 | 2324.86 | RNASEH1 |
| jugloneCTD00000198 | 0.004 | 369.82 | 2024.38 | RNASEH1 |
| CHEMBL332826TTD00004878 | 0.006 | 269.73 | 1394.27 | RNASEH1 |
| naringinHL60DOWN | 0.006 | 262.62 | 1350.71 | MTFR1 |
| PURPURINCTD00003820 | 0.006 | 243.37 | 1233.69 | RNASEH1 |
| quercetinTTD00010442 | 0.009 | 178.05 | 848.32 | RNASEH1 |
| protoporphyrinIXCTD00001323 | 0.009 | 171.89 | 813.07 | RNASEH1 |
| myricetinTTD00009430 | 0.009 | 160.77 | 749.95 | RNASEH1 |
| CylindrospermopsinCTD00003136 | 39,034.00 | 290,763.91 | SDHAF2;PUS1 | |
| tanespimycinHL60DOWN | 0.003 | 713.21 | 4167.85 | PUS1 |
| geldanamycinMCF7DOWN | 0.003 | 605.00 | 3439.32 | PUS1 |
| DisodiumseleniteCTD00007229 | 0.005 | 37,260.00 | 199,808.13 | SDHAF2;PUS1 |
| mepacrineHL60DOWN | 0.005 | 424.49 | 2266.91 | PUS1 |
OR, odds ratio.
The present study employed a bidirectional MR approach and identified several suggestive inverse associations between mitochondrial proteins, including 39S ribosomal protein L14, oligoribonuclease, and mitochondrial fission regulator 1, and the risk of POI. These findings suggest that mitochondrial dysfunction may play a potential crucial role in the pathogenesis and progression of POI, reinforcing the growing evidence for the essential role of mitochondria in ovarian function and female reproductive health [25].
Mitochondria, often referred to as the “powerhouses” of cells, play a critical role in energy production, cellular signal transduction, and the regulation of apoptosis [26]. Increasing evidence indicates that mitochondrial dysfunction is significantly associated with ovarian aging, a pivotal factor in POI [27, 28]. Our findings further support this association, demonstrating that mitochondrial proteins may serve as protective factors against POI and, consequently, could serve as potential therapeutic targets. For instance, the inverse association between mitochondrial fission regulator 1 and the risk of POI indicates its potential role in preserving mitochondrial integrity and sustaining energy production in ovarian cells—processes essential for folliculogenesis and oocyte viability [29, 30].
The findings of our study indicate that mitochondrial proteins exhibiting significant associations are all capable of reducing the probability of POI, suggesting a protective effect. This highlights the intrinsic link between mitochondrial health and oocyte quality. Furthermore, these results suggest that impaired mitochondrial dynamics may contribute to fertility decline and accelerated ovarian ageing [31, 32]. Accordingly, our study provides further support for the hypothesis that mitochondrial dysfunction constitutes a fundamental mechanism underlying the pathogenesis of POI.
Interestingly, not all mitochondrial proteins exhibited a protective association with POI. Specifically, coiled-coil domain-containing protein 90B showed a positive causal relationship with POI risk, suggesting that elevated levels of this protein may increase susceptibility to the condition. Also known as MCUR1, this protein has been implicated in the regulation of mitochondrial calcium uptake and respiratory chain activity [33]. Dysregulation of mitochondrial calcium homeostasis can impair energy metabolism and increase oxidative stress, both of which are associated with ovarian aging and oocyte apoptosis [34, 35]. Therefore, the observed association between elevated coiled-coil domain-containing protein 90B levels and increased POI risk may reflect mitochondrial calcium overload-induced dysfunction, providing further insight into the complex mitochondrial mechanisms underlying POI pathogenesis.
Notably, our research also demonstrated a modest effect of POI on specific mitochondrial proteins, suggesting a potential feedback loop in which POI exacerbates mitochondrial dysfunction, thereby accelerating ovarian decline. This bidirectional relationship implies that as ovarian function deteriorates, mitochondrial health is further compromised, creating a vicious cycle that exacerbates POI progression [36]. Given the maternal inheritance of mtDNA and the dependence of oocyte quality on mitochondrial function, interventions targeting mitochondrial health may offer a promising strategy for the management and treatment of POI [37]. Available evidence suggests that antioxidants, such as coenzyme Q10 and melatonin, may enhance mitochondrial function and mitigate oxidative stress in aged oocytes [38]. These findings are particularly relevant in the context of assisted reproductive technologies, where enhancing mitochondrial function could improve oocyte quality and fertility outcomes in women with POI [6]. Moreover, recent research has identified specific mtDNA mutations and mitochondrial gene polymorphisms associated with POI, which may serve as potential diagnostic biomarkers [39]. The identification of these genetic markers in women at risk of POI could facilitate the implementation of early intervention strategies and the development of personalized treatment plans [40]. Furthermore, investigating the role of mitochondrial heteroplasmy may enhance our understanding of the complex mitochondrial mechanisms underlying not only POI but also other ovarian disorders, including polycystic ovary syndrome (PCOS) and endometriosis. Given that heteroplasmic variations can affect mitochondrial function and cellular energy metabolism, elucidating their contribution may uncover novel pathogenic pathways and shared therapeutic targets across multiple ovarian diseases [41].
Although this study has several strengths, it is important to acknowledge the limitations inherent in its research design. First, the use of genetic data from European populations may limit the generalizability of our findings to other ethnic groups. It would be beneficial for future studies to include more diverse populations to enhance the applicability of these results. Second, although MR is a robust method for inferring causality, it relies on the assumption that the selected genetic variants are valid IVs. Despite efforts to mitigate potential pleiotropy, it is not possible to rule out the possibility of residual confounding. Third, although none of the associations remained statistically significant after FDR correction, the overall pattern of nominal associations suggests that mitochondrial health is closely linked to POI and may still reflect biologically relevant relationships. Finally, while our study identified an association between mitochondrial proteins and POI, it did not elucidate the underlying molecular mechanisms. Further functional studies are required to determine how these mitochondrial proteins influence ovarian biology.
This study advances the understanding of the underlying role of mitochondria in POI by identifying specific mitochondrial proteins as potential biomarkers and therapeutic targets. By elucidating aspects of the mitochondrial-ovarian axis, it provides a foundation for developing targeted interventions aimed at delaying ovarian ageing and improving fertility outcomes in women affected by POI.
The data supporting the findings of this study are available from the corresponding authors upon reasonable request.
ZQ and YS conceived of the study and supervised the research. YK was involved in formal analysis, contributed to the software, and assisted in methodology modification. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
All the authors are very grateful for the data support provided by the IEU Open GWAS project.
This research received no external funding.
The authors declare no conflict of interest.
During the preparation of this work the authors used ChatGpt-3.5 in order to check spell and grammar. After using this tool, the authors reviewed and edited the content as needed. The author takes full responsibility for the content of the publication.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/CEOG46006.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.





