1 Shenzhen Maternity and Child Healthcare Hospital, Women and Children’s Medical Center, Southern Medical University, 518028 Shenzhen, Guangdong, China
2 Obstetrics and Gynecology Hospital, Fudan University, 200011 Shanghai, China
3 The Shanghai Key Laboratory of Female Reproductive Endocrine-Related Diseases, 200011 Shanghai, China
†These authors contributed equally.
Abstract
Observational studies have demonstrated a potential association between hypertensive disorders of pregnancy (HDPs) and an elevated risk of subsequent kidney impairment. The present study aimed to evaluate the possible causal relationship between HDPs and future renal dysfunction using a genetic approach.
This two-sample Mendelian randomization (MR) analysis, conducted between October 2023 and January 2024, examined the effects of HDPs—both overall and by subtype (gestational hypertension and preeclampsia/eclampsia)—on chronic kidney disease (CKD), albuminuria, and estimated glomerular filtration rate (eGFR). Genetic data were sourced from publicly available genome-wide association studies (GWAS) provided by the FinnGen, CKDGen, UK Biobank, GIANT (Genetic Investigation of ANthropometric Traits), deCODE, and DIAMANTE (DIAbetes Meta-ANalysis of Trans-Ethnic association studies) consortia. The primary method employed was inverse-variance weighting, supplemented by sensitivity analyses and instrument-strength validation to ensure causal robustness.
This study included approximately 300,000 to 600,000 individuals per kidney function trait, with around 15,000 HDP cases. Genetically predicted HDPs showed a weak but statistically significant association with albuminuria [odds ratio (OR): 1.02; 95% confidence interval (CI): 1.00–1.03; p = 0.01]. This association remained consistent after adjustments for body mass index (BMI), smoking, and type 2 diabetes (T2D) in the multivariable MR analysis (OR: 1.03; 95% CI: 1.01–1.04; p < 0.001). No significant associations were observed between HDPs, including for the subtypes, and the presence of CKD or eGFR. These results demonstrated robustness across diverse sensitivity analyses.
The genetic proxy for HDPs was identified as causally associated with maternal long-term albuminuria, independent of BMI, smoking, and T2D. These results offer novel insights bolstering the causal relationship between HDPs and long-term maternal kidney dysfunction.
Keywords
- hypertensive disorders of pregnancy
- preeclampsia
- long-term kidney function
- albuminuria
- Mendelian randomization
Hypertensive disorders of pregnancy (HDPs) comprising conditions such as preeclampsia and gestational hypertension affect approximately 10% of pregnancies worldwide. It is associated with substantial immediate maternal morbidity and mortality [1], as well as susceptibility to long-term chronic diseases, such as chronic kidney damage, which has been a common and well-documented concern [2, 3].
Acute renal damage represents a frequent manifestation of HDPs and often resolves within months postpartum. Nevertheless, various population-based observational studies have established a correlation between HDPs and long-term kidney diseases [4, 5, 6, 7, 8]. A secondary analysis of data from the Family Blood Pressure Program indicated that a history of hypertension during pregnancy was associated with an increased risk of microalbuminuria decades later [9]. Furthermore, analyses of data from the Medical Birth Registry of Norway and Swedish Medical Birth Register (MBR) demonstrated a link between preeclampsia and an increased risk of end-stage renal disease (ESRD), although the absolute risk remains low [10, 11]. In addition, meta-analysis of 18 observational studies revealed a significant positive correlation between two subtypes of HDPs, namely preeclampsia and gestational hypertension, and an increased risk of future chronic kidney disease (CKD) and ESRD [6]. However, it remains unclear whether these associations express causal relationships deriving from the adverse effects of preeclampsia itself, or if they can be attributed to underlying risk factors that predisposed women to both preeclampsia and subsequent renal diseases.
Mendelian randomization (MR) utilizes genetic variants as instrumental variables to evaluate potential causal effects of exposures on outcomes. This method leverages the inherent causal relationship between genetic variants and the exposure of interest. Because genetic alleles are randomly assigned at conception and remain largely unaffected by confounding factors that commonly plague observational studies, MR simulates the design of a randomized controlled trial. This unique feature effectively minimizes confounding and reverse causation [12]. Consequently, MR yields estimates that represent the lifelong effect of genetic predisposition to an exposure—such as HDPs—on outcomes like kidney impairment. It is essential to note that these effect estimates are not directly comparable to hazard ratios from conventional observational studies, which generally assess short-term associations over limited follow-up durations and are susceptible to unmeasured confounding. Nonetheless, MR’s capacity to approximate a natural randomized experiment underscores its strength in reinforcing causal inference.
Using a two-sample MR framework, this study examined the causal effects of HDPs, overall and by subtype, on CKD, albuminuria, and reduced estimated glomerular filtration rate (eGFR). To control for confounding, multivariable MR included adjustments for body mass index (BMI), smoking, and type 2 diabetes (T2D).
This study utilized de-identified, publicly available genome-wide association studies (GWAS) summary data. All original studies had obtained ethical approval. The reporting of this study adheres to the Strengthening the Reporting of Observational Studies in Epidemiology Using Mendelian Randomization (STROBE-MR) guidelines. Data were sourced from studies that individually analyzed populations of European ancestry to mitigate bias stemming from demographic stratification.
Fig. 1 illustrates the overall study design in a flowchart. This analysis focused on three primary exposures: HDPs overall, along with two subtypes—gestational hypertension and preeclampsia/eclampsia. Summary-level genetic association data for these phenotypes were sourced from the FinnGen (https://www.finngen.fi/en) consortium’s ninth release, which was published in late 2023 [13]. The sample sizes included 14,727 cases and 196,143 controls for any HDPs, 8502 cases and 194,266 controls for gestational hypertension, and 7212 cases and 194,266 controls for preeclampsia/eclampsia. The case definitions for HDPs, gestational hypertension, and preeclampsia/eclampsia were determined using specific phenotype codes within the FinnGen consortium: O15_HYPTENSPREG, O15_GESTAT_HYPERT, and O15_PRE_OR_ECLAMPSIA, respectively. These codes are derived from the International Classification of Diseases (ICD) system, providing a standardized and reliable framework for phenotype classification, thereby enhancing the validity of the analysis (Table 1, Ref. [13, 14, 15, 16, 17, 18]).
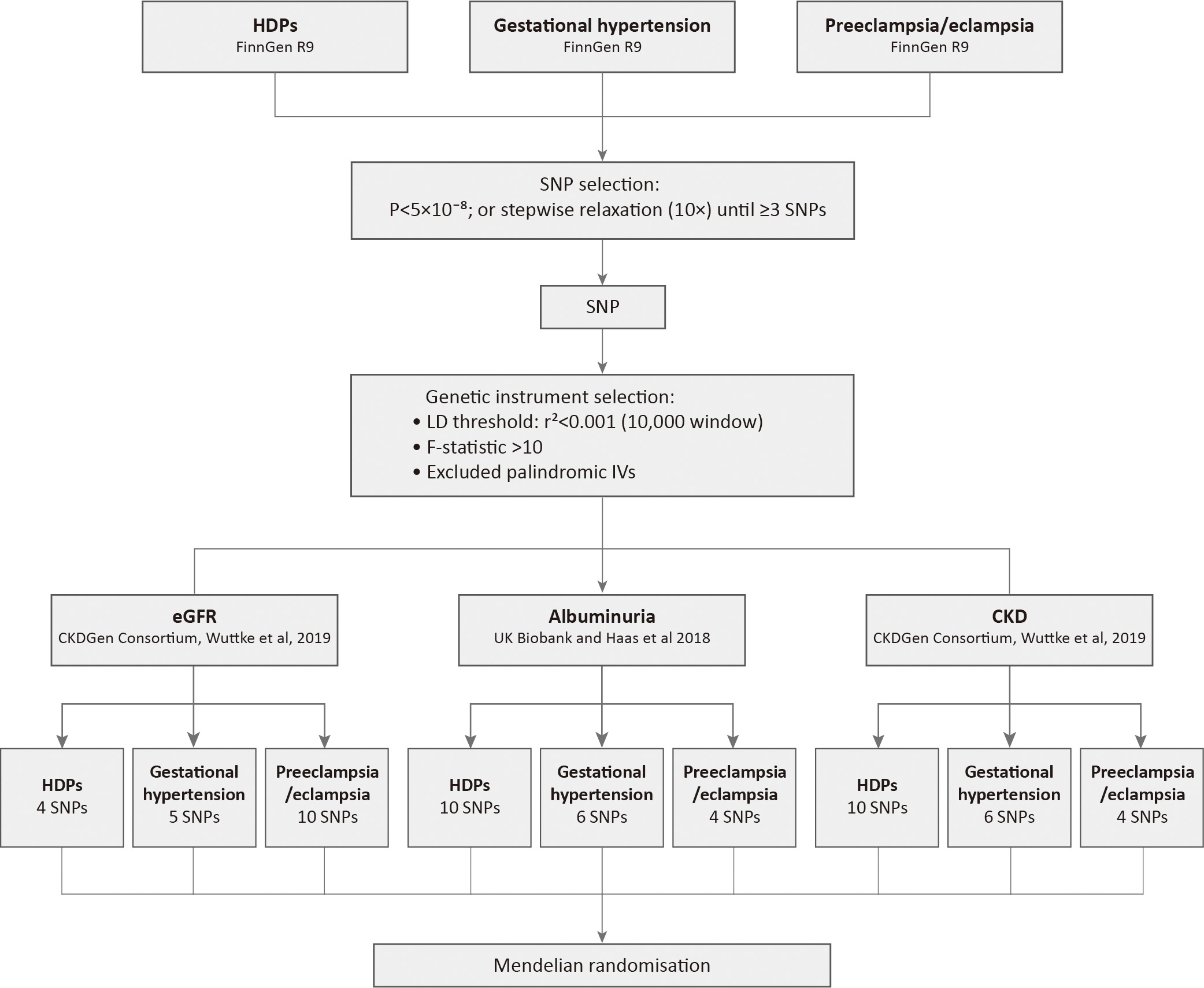 Fig. 1.
Fig. 1.
Study analysis flowchart: from data to instrumental variants. HDPs, hypertensive disorders of pregnancy; CKD, chronic kidney disease; SNP, single nucleotide polymorphism; LD, linkage disequilibrium; eGFR, estimated glomerular filtration rate; IVs, instrumental variables.
| Trait | Study/Source | Participants | Phenotype definition |
| Exposure | |||
| HDPs | FinnGen R9 [13] | Cases: 14,727; Control: 196,143 | Cases: FinnGen code O15_HYPTENSPREG |
| Controls: not classified as a case | |||
| Gestational hypertension | Cases: 8502; Control: 194,266 | Cases: FinnGen code O15_GESTAT_HYPERT | |
| Controls: not classified as a case | |||
| Preeclampsia/eclampsia | Cases: 7212; Control: 194,266 | Cases: FinnGen code O15_PRE_OR_ECLAMPSIA | |
| Controls: not classified as a case | |||
| Outcome | |||
| eGFR | CKDGen Consortium and Wuttke et al., 2019 [14] | 567,460 individuals | eGFR was estimated using the CKD-EPI equation for individuals above 18 years of age and the Schwartz formula for those aged 18 years or younger. |
| Albuminuria | UK Biobank and Haas et al., 2018 [15] | 382,500 individuals | Albuminuria levels were measured during the initial enrollment visit using a Beckman Coulter AU5400 clinical chemistry analyzer with a detection range of 6.7–200 mg/L. |
| CKD | CKDGen Consortium and Wuttke et al., 2019 [14] | Cases: 41,395; Control: 439,303 | Cases: eGFR below 60 mL/min/1.73 m2 |
| Controls: not classified as a case | |||
| Covariate | |||
| BMI | UK Biobank, GIANT consortium and Pulit et al., 2019 [16] | 434,794 female individuals | Body mass index |
| Smoking | deCODE, UK Biobank, and Liu et al., 2019 [18] | Cases: 557,337; Controls: 674,754 | Cases: self-reported smoking behavior |
| Controls: never smoked | |||
| T2D | DIAMANTE Consortium and Mahajan et al., 2022 [17] | Cases: 80,154; Controls: 853,816 | Cases: T2D was defined based on the criteria outlined in the previous study |
| Controls: not classified as a case |
BMI, body mass index; T2D, type 2 diabetes; GIANT, Genetic Investigation of ANthropometric Traits; DIAMANTE, DIAbetes Meta-ANalysis of Trans-Ethnic association studies; CKD-EPI, Chronic Kidney Disease Epidemiology Collaboration.
The MR design estimates the lifetime risk of an outcome associated with genetic
predisposition to an exposure. The kidney outcome data (eGFR, albuminuria, CKD)
were obtained from GWAS involving adult populations, with measurements taken at a
single time point during study enrollment (Table 1). Summary statistics for eGFR
(n = 567,460 individuals) and CKD (n = 480,698 individuals, 41,395 cases and
439,303 controls) were derived from meta-analysis of GWAS of participants of
European ancestry from the CKDGen Consortium [14]. Participant characteristics
and the phenotype definitions have been comprehensively described by Wuttke
et al. [14]. CKD was specifically defined as an eGFR
Three variables—BMI, smoking, and T2D—were selected as potential confounders when evaluating the association between HDPs and subsequent kidney impairment (Table 1). These factors were included due to their well-documented, although not necessarily causal, associations with both the exposures and outcomes of interest [21, 22, 23]. Genetic instruments for each confounder were derived from GWAS with the largest available sample sizes, focusing on individuals of European ancestry. Specifically, summary-level data for BMI were acquired from a meta-analysis including 434,794 female participants, conducted jointly by the UK Biobank and the GIANT (Genetic Investigation of ANthropometric Traits) consortium [16]. Genetic associations for T2D were obtained from a trans-ethnic GWAS meta-analysis by the DIAMANTE (DIAbetes Meta-ANalysis of Trans-Ethnic association studies) Consortium, comprising 80,154 cases and 853,816 controls [17]. Data for smoking initiation, based on 557,337 cases and 674,754 controls, were collected from the deCODE study and the UK Biobank [18].
As this study utilized summary-level GWAS data, individual-level characteristics and inclusion/exclusion criteria were determined by the original consortia. Details can be found in the referenced publications.
Genetic instruments were selected if associated with each exposure at a
genome-wide significance threshold of p
The selected genetic instruments were grouped for independence by estimating
linkage disequilibrium (LD), applying a cut-off of r2
The strength of the chosen genetic instruments was assessed through the
calculation of F statistics, and genetic instruments with an F statistic below 10
were deemed weak and therefore excluded. The evaluation was based on the
mathematical formula utilized: F = R2
To determine the statistical power of our primary univariable MR analyses, we utilized the online tool mRnd (https://shiny.cnsgenomics.com/mRnd/), which is specifically designed for power calculations in MR studies [24, 25]. This tool estimates statistical power using key parameters such as sample size, case-control ratio, and the proportion of variance in the exposure explained by the genetic instruments.
The inverse variance weighting (IVW) method was employed as the primary approach to infer potential causal relationships between HDPs and kidney damage. IVW is known for its ability to offer robust causal estimates free from directional pleiotropy, making it a commonly used method in MR studies [26]. The combined causal estimate from the IVW method represents a weighted average of the individual SNP effects, which may naturally vary in direction due to the complex genetic architecture of the exposure.
To assess the reliability of the main causal estimates and identify potential
pleiotropic effects, we implemented several sensitivity analyses, including
weighted median MR, MR‑Egger regression, and MR-PRESSO (Pleiotropy RESidual Sum
and Outlier). The weighted median approach produces consistent causal estimates,
provided that at least half of the IVs are valid [27]. MR-Egger regression
enables the detection of directional pleiotropy via its intercept term and yields
pleiotropy-robust effect estimates [28]. MR-PRESSO detects outlier SNPs and
provides effect estimates adjusted from these outliers [29]. Heterogeneity across
IVs was assessed using Cochran’s Q statistic [30]. A significance level of
p
Multivariable MR utilized SNPs from GWAS on BMI, smoking, and T2D, applying IVW to estimate their concurrent effects. This method re-evaluates summary genetic associations by incorporating genetic correlations with both exposure and risk factors into a weighted regression framework, thereby reducing the influence of SNP exposure on their effects on other presumed risk factors through an indirect pathway. All statistical analyses were performed in R (version 4.3.1, R Foundation for Statistical Computing, Vienna, Austria) with the TwoSampleMR, MR-PRESSO, and MVMR (Multivariable Mendelian Randomization) packages.
The complete set of genetic instruments employed for the exposures is detailed in Supplementary Table 1. The F-statistics for these genetic instruments exceeded the commonly selected threshold of 10, indicating robust instrument strength.
Contrary to conventional analyses that suggest a positive association between HDPs and the risk of CKD, the MR analysis did not provide evidence to support such associations between HDPs (OR: 1.01; 95% CI: 0.93–1.09; p = 0.78) and the subtypes (gestational hypertension: OR: 1.02; 95% CI: 0.94–1.11; p = 0.63; preeclampsia/eclampsia: OR: 0.97; 95% CI: 0.81–1.14; p = 0.68; Supplementary Table 2 and Fig. 2) with the risks of CKD.
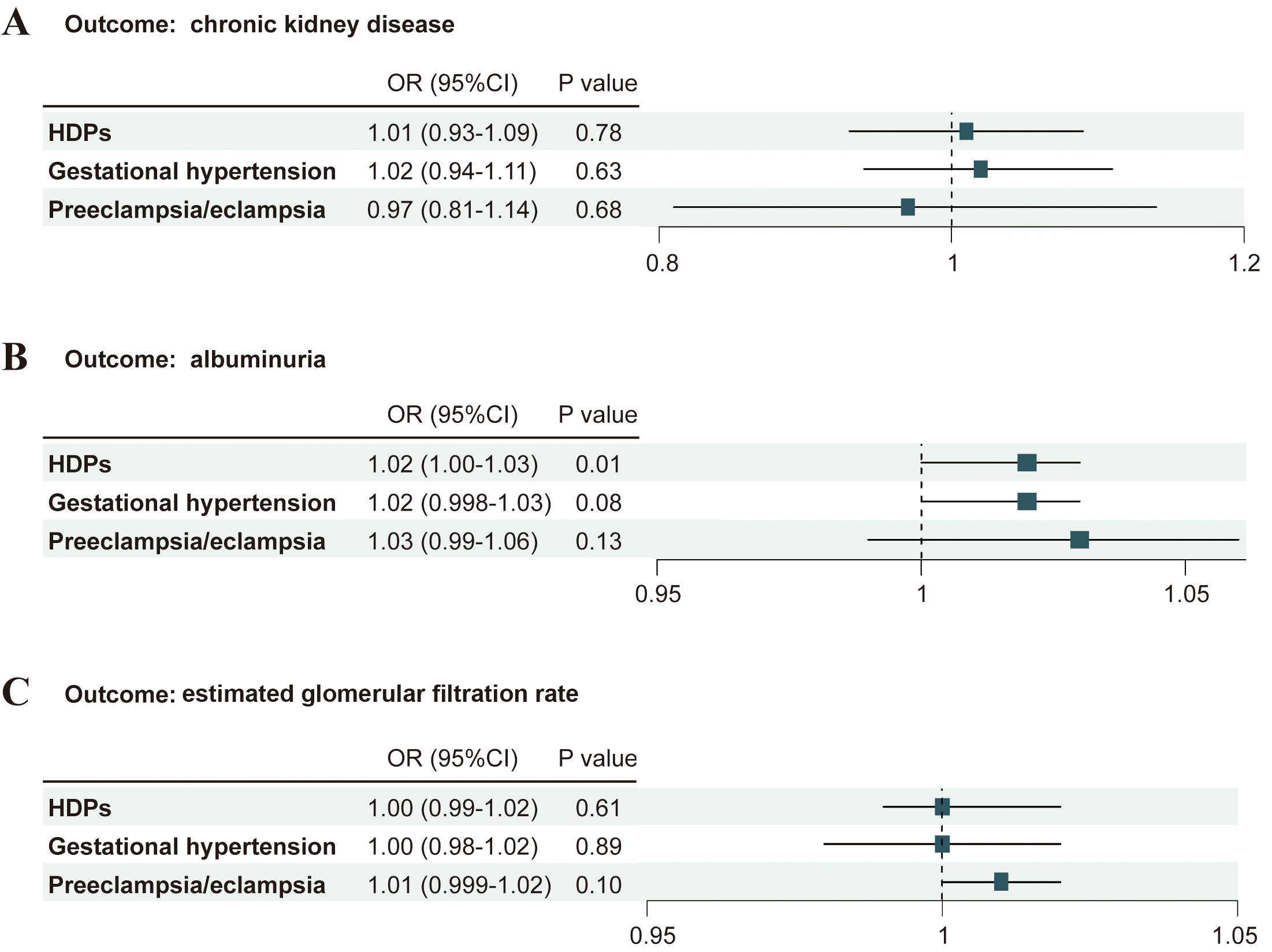 Fig. 2.
Fig. 2.
Associations of genetically predicted HDPs and its subtypes with the risk of kidney function. (A) Associations of HDPs with the risk of CKD. (B) Associations of HDPs with albuminuria. (C) Associations of HDPs with eGFR. OR, odds ratio; CI, confidence interval.
MR-Egger regression analysis indicated an absence of directional pleiotropy for genetically predicted HDPs (intercept p = 0.92; Supplementary Table 3). Consistent results were observed for both subtypes: gestational hypertension (intercept p = 0.42) and preeclampsia/eclampsia (intercept p = 0.34). Effect estimates derived from the weighted median method were consistent with those obtained from the primary IVW analysis, supporting the robustness of the findings. Some heterogeneity was detected for preeclampsia/eclampsia (Cochran’s Q = 8.88, p = 0.03; Supplementary Table 3). In contrast, neither HDPs overall (Q = 5.17, p = 0.64) nor gestational hypertension (Q = 2.34, p = 0.67) exhibited significant heterogeneity. Furthermore, MR-PRESSO analysis revealed no evidence that outliers substantially influenced the causal estimates (Supplementary Table 4).
Genetically predicted HDPs demonstrated a significant yet weak association with an elevated risk of albuminuria (OR: 1.02; 95% CI: 1.00–1.03; p = 0.01). In contrast, neither gestational hypertension (OR: 1.02; 95% CI: 0.998–1.03; p = 0.08) nor preeclampsia/eclampsia (OR: 1.03; 95% CI: 0.99–1.06; p = 0.13) showed statistically significant associations with albuminuria, as detailed in Supplementary Table 2 and Fig. 2.
As shown in Supplementary Table 3, some heterogeneity was noted for preeclampsia/eclampsia (Q statistic = 10.63; p = 0.01), however, not for HDPs (Q statistic = 7.85; p = 0.35) or gestational hypertension (Q statistic = 5.85; p = 0.21). MR-PRESSO analysis identified no outlier SNPs (Supplementary Table 4), indicating that the heterogeneity likely reflects diffuse mechanisms rather than a few invalid instruments. The MR-Egger test did not detect significant pleiotropy (MR-Egger intercept p = 0.42, 0.28, and 0.11 for HDPs, gestational hypertension, and preeclampsia/eclampsia, respectively). The weighted median analysis indicated a consistent direction of effect between genetically predicted HDPs and albuminuria, aligning with the results obtained from the IVW method.
No significant associations were observed between genetically predicted HDPs and eGFR, as indicated by the following ORs: HDPs overall (OR: 1.00; 95% CI: 0.99–1.02; p = 0.61), gestational hypertension (OR: 1.00; 95% CI: 0.98–1.02; p = 0.89), and preeclampsia/eclampsia (OR: 1.01; 95% CI: 0.999–1.01; p = 0.10) (Supplementary Table 2; Fig. 2). Sensitivity analyses provided no evidence of directional pleiotropy for HDPs (MR-Egger intercept p = 0.48) or its subtypes (gestational hypertension: p = 0.16; preeclampsia/eclampsia: p = 0.55) (Supplementary Tables 3,4). Moreover, the weighted median estimates showed close agreement with the IVW results in both direction and magnitude of effect. Some heterogeneity was detected for HDPs (Cochran’s Q = 12.38; p = 0.01) and gestational hypertension (Q = 12.23; p = 0.01), though not for preeclampsia/eclampsia (Q = 10.30; p = 0.33). The MR-PRESSO analysis identified 4 outlier SNPs in the association between HDPs and eGFR; however, after their removal, the corrected estimates remained consistent with the initial IVW results, confirming that these outliers did not influence the overall null association.
Ensuring the accuracy and validity of the results entails addressing potential
confounding factors by adjusting variables. Prior epidemiological research has
recognized BMI, smoking behavior, and T2D as established confounding variables in
the association between HDPs and subsequent renal impairment [21, 22, 23]. In the
multivariable MR analysis, a consistent and significant association remained
between HDPs and albuminuria (OR: 1.03; 95% CI: 1.01–1.04; p
| Exposure | Outcome | Covariate | nSNP | Standard error | OR (95% CI) | p value | |
| HDPs | Albuminuria | BMI, smoking, and T2D | 6 | 0.03 | 0.01 | 1.03 (1.01–1.04) |
In this MR analysis, we identified a modest yet statistically significant association between the genetic proxy of HDPs and albuminuria, independent of maternal BMI, smoking, and T2D. The results were robust across various sensitivity analyses. Our findings provide novel evidence strengthening the causal association between HDPs and long-term maternal renal dysfunction.
Epidemiological evidence from multiple observational studies indicates that HDPs, particularly preeclampsia, are associated with increased risk of CKD [10, 31] and a decline in eGFR in later life [7]. A previous meta-analysis revealed that, on average, 7.1 years postpartum, individuals with severe preeclampsia faced an eight-fold higher risk of microalbuminuria [31]. Another study emphasized that, throughout the postpartum years, a history of HDPs correlated with an elevated risk of microalbuminuria [9]. A recent study has also highlighted a significant long-term escalation in kidney disease risk among women with a history of HDPs [32].
However, establishing causal relationships from observational data presents challenges due to potential residual confounding factors and biases. Many pregnant women with preeclampsia may have undiagnosed preexisting kidney dysfunction, as certain diagnostic tests are not routinely conducted. Additionally, biases might arise from residual confounding in the observational setting, especially given the multifaceted nature of preeclampsia, which is influenced by various sociobehavioral factors [33, 34]. This study contributes to existing literature by employing MR, a method that reduces the impact of confounding factors and biases. Findings from this approach revealed a causal relationship between HDPs and long-term maternal albuminuria, a mild manifestation of CKD, although HDPs were not directly associated with CKD or eGFR.
HDPs and CKD may exhibit a bidirectional relationship. Maternal renal disease, even with mild reductions in eGFR or an increase in microalbuminuria, is a recognized risk factor for preeclampsia [35, 36]. HDPs, especially preeclampsia, marked by the dysregulation of angiogenic mediators precipitating systemic endothelial dysfunction, can be exacerbated by CKD, altering the complement and renin-angiotensin-aldosterone systems. Our study showed that HDPs, in turn, have been implicated in the onset of long-term albuminuria. HDPs have the potential to instigate future kidney disorders through mechanisms like acute kidney injury, endothelial impairment, and podocyte loss during pregnancy [37, 38]. Furthermore, we found that HDPs represent a gender-specific risk factor for future kidney dysfunction, underscoring the importance of incorporating obstetric history into the routine CKD assessments. Heightened awareness among healthcare providers regarding the link between HDPs and CKD can contribute significantly to the prevention of long-term kidney complications in women with a history of HDPs.
Notably, functional annotation of the genetic variants associated with HDPs revealed several genes involved in biological pathways crucial to both hypertensive pregnancy and renal physiology. Among these, PLCE1 (phospholipase C epsilon 1) (rs10882398) is particularly significant as mutations in this gene are a well-established cause of early-onset nephrotic syndrome. This directly links it to podocyte function and glomerular integrity, suggesting a compelling mechanism for albuminuria due to disruption of the renal filtration barrier [39, 40]. Similarly, TCF7L2 (transcription factor 7 like 2) (rs6585195), a major diabetes risk gene, indicates a potential shared genetic predisposition between HDPs and diabetic nephropathy, possibly mediated through metabolic disturbances influencing renal health [41]. Additional variants highlight more pathways: PREX1 (phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1) (rs2208589), involved in Angiotensin II signaling and endothelial function [42], offers a plausible link to the systemic endothelial dysfunction characteristic of preeclampsia. Likewise, MTHFR (methylenetetrahydrofolate reductase) (rs13306561), critical in folate metabolism and associated with hyperhomocysteinemia [43], may contribute to long-term vascular and renal injury via endothelial dysfunction and thrombosis. Furthermore, THADA (THADA armadillo repeat containing) (represented by multiple SNPs), associated with apoptosis regulation, diabetes, and polycystic ovary syndrome [44], suggests potential roles in both placental development and podocyte survival. Together, these genetic insights strengthen the biological plausibility of a causal pathway from HDPs to albuminuria, implicating podocyte injury, endothelial damage, and metabolic dysregulation as key mediators.
Although our MR study addressed many limitations commonly found in observational designs, the results should be interpreted with caution due to constraints specific to our study and the broader MR framework. First, the use of GWAS data that included various forms of CKD may have introduced heterogeneity in the outcome, potentially affecting the robustness of our findings. Second, the lack of detailed information on preeclampsia severity and gestational age at onset prevented meaningful stratification of outcomes, which is essential for clinical translation. Future investigations into the relationship between HDPs and renal function should therefore incorporate more refined clinical phenotypes. Third, reliance on registry-based ICD codes for defining exposure may have resulted in underascertainment of subclinical or mild HDP cases and constrained our ability to evaluate dose-response relationships. Fourth, the number of strong genetic instruments varied across HDP subtypes, with fewer instruments available for gestational hypertension compared to preeclampsia. This likely reduced statistical power to detect significant associations for gestational hypertension, as suggested by the borderline association observed with albuminuria. As GWAS sample sizes for these phenotypes increase, future studies will be better equipped to examine the causal effects of specific HDP subtypes. Additionally, statistical power remained limited for certain analyses, such as between preeclampsia/eclampsia and eGFR, which may have hindered the detection of a true causal effect despite a large outcome sample size (Supplementary Table 7). Finally, the use of mixed-sex outcome data, along with the absence of sex-stratified summary statistics, likely diluted effect estimates due to the inclusion of males who are not at risk for female-specific exposures. Eliminating this sex-related bias was not feasible.
To our knowledge, this study represents the first application of MR to examine the potential causal effect of HDPs on long-term renal outcomes. Incorporating a wide range of sensitivity analyses to validate the robustness of the results. This study found that the genetic proxy for HDPs was linked to maternal long-term albuminuria, and this association appeared to be independent of BMI, smoking, and T2D. Although the observed genetic association is modest in magnitude, it represents the lifelong, population-level effect of HDPs on albuminuria risk. Our findings provide novel evidence strengthening the causality between HDPs and long-term maternal kidney dysfunction.
The GWAS summary statistics analyzed in this study are publicly available for download in the FinnGen (https://www.finngen.fi/en), CKDGen (https://ckdgen.imbi.uni-freiburg.de/), GIANT (https://giant-consortium.web.broadinstitute.org/GIANT_consortium), deCODE (https://www.decode.com/), and DIAMANTE (https://diagram-consortium.org/) consortia, as detailed in Table 1.
XL and YX designed the study and supervised its implementation. HZ, PW, SS, QX, XZ, SC and ZW performed the data curation. HZ and YZ conducted the analysis. SS and HZ drafted the original version of the manuscript. HZ, YZ, QX, XZ and SC provided data interpretation, critical review, and commentary to the revised versions of the manuscript. All authors contributed to critical revision of the manuscript for important intellectual content. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
We extend our deep appreciation to the participants and researchers involved in the FinnGen study for their indispensable contributions to our research. We would like to acknowledge the UK Biobank (https://www.ukbiobank.ac.uk/) for providing the summary statistics data for our analysis. We also acknowledge and appreciate the generosity of the GIANT, DIAMANTE, CKDGen, and deCODE consortia for sharing their invaluable resources.
This work was supported by Shenzhen Medical Research Fund (B2404004, A2303073), National Key Research and Development Program (2021YFC2701600), Shenzhen Science and Technology Program (RCYX20231211090407012, JCYJ20240813115007009, JCYJ20240813115015019, JCYJ20240813115014018), National Natural Science Foundation of China (82371698, 82502041), Medical Scientific Research Foundation of Guangdong Province (B2025763).
The authors declare no conflict of interest.
Supplementary material associated with this article can be found, in the online version, at https://doi.org/10.31083/CEOG44838.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.


