1 Genetic and Metabolic Central Laboratory, Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region, 530000 Nanning, Guangxi, China
2 Division of Genetics and Genomics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, USA
Abstract
Background: To evaluate the clinical performance and clinical
implementation of noninvasive prenatal testing (NIPT) using fetal fraction (FF)
enrichment. Methods: Both standard NIPT and NIPT with FF enrichment were
performed concurrently to 277 clinical samples. Results: On average, the
FF of each sample processed by NIPT with FF enrichment was 1.9-fold higher than
without enrichment. 88% (43/49) of samples with low FF in standard NIPT were
recovered by the enrichment method. 11 more copy number variants (CNVs) which
were confirmed by prenatal diagnosis were detected by NIPT with FF enrichment.
NIPT with FF enrichment generated a positive predictive value of 75% for CNVs
(
Keywords
- noninvasive prenatal testing
- fetal fraction enrichment
- body mass index
- copy number variants
The application of maternal plasma-extracted short cell-free DNA (cfDNA) has been widely accepted for screening common fetal aneuploidies [1, 2]. This approach is known as the noninvasive prenatal testing (NIPT), since the discovery of Y chromosome signals in maternal plasma by Lo et al. in 1997 [3]. By 2020, NIPT had been validated and recommended by professional committees, such as the American College of Obstetricians and Gynecologist (ACOG) and the American College of Medical Genetics and Genomics (ACMG) for general pregnancies in order to screen for prenatal trisomy21/trisomy18/trisomy13 (T21/T18/T13) [4, 5].
The accuracy of NIPT increased as sufficient fetal-derived cfDNA was captured during sequencing. The proportion of fetal-derived cfDNA in all circulating cfDNA from the maternal plasma is called the fetal fraction (FF). In maternal plasma, the fetal-derived cfDNA originates from the placenta, and the level of the FF generally ranges from 3 to 30% [6]. Most of the existing methodologies for NIPT do not analyze samples with a low FF, and the cut-off values are defined by the platform used, because analyzing samples below the threshold can result in increased false negative rates [7, 8]. Depending on the platforms used, the reported low FF from the first sample of maternal blood drawn ranges from 1 to 8% [9]. Known factors that can affect the level of the FF include the gestational age of the fetus, maternal body mass index (BMI), maternal age and the size and functional integrity of the placenta [10, 11]. However, these FF-related factors cannot be manipulated during gestational weeks 9 to 12, which is the optimal time window for NIPT. As the sequencing resolution is elevated, the screening targets for NIPT can extend beyond the common trisomies.
Sub-chromosomal imbalanced copy number variants (CNVs) can be associated with structural and developmental abnormalities in the newborns. Collectively, recurrent CNVs associated with genomic disorders can account for 1 to 2% of congenital abnormalities in newborn babies [12, 13]. CNVs-implicated genomic disorders, such as 1q21.1 microdeletion/microduplication as well as Williams-Beuren, Prader-Willi/Angelman and DiGeorge syndromes, are not lethal at birth but require continuous treatment throughout the postal period [14]. DiGeorge syndrome, also known as the 22q11.2 deletion syndrome, has a prevalence of one in 4000 live newborns. This multisystemic disorder manifests as one of the most common CNV-related syndromes [15]. The sensitivity and specificity of CNV detection by NIPT is correlated with the size of the CNV, the FF and the depth of sequencing. Increasing the sequencing depth of each sample can improve CNV detectability, but this approach is economically unwise, and a feasible option is to achieve a higher level of the FF. Several methods for FF enrichment had been reported, mainly through size selection and DNA repair. DNA repair was reported for the recovery of long cfDNA molecules which are more than 250 bp, and the total increase of the FF achieved was less than 5% [16]. Size selection by magnetic nanoparticles showed robust efficiency in FF enrichment, with an average of approximately 10% being achieved [17, 18]. At the same time, the magnetic beads option was able to cope with large scales as well as an automated DNA library construction operation system.
In this pilot clinical study, we set up a NIPT protocol with FF enrichment using magnetic nanoparticles. Compared to the preliminary method of enriching the FF using magnetic beads, the pipeline we adopted aimed to achieve a more effective enrichment in a fully automated, large-scale DNA extraction and library construction system. We compared the clinical performance of this approach for simultaneous detection of common trisomies and CNVs versus standard NIPT and identified a relationship between the effectiveness of the FF fraction enrichment and BMI.
This retrospective clinical study was approved by the Ethics Committee of the Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region. To evaluate the performance of NIPT with FF enrichment, this study was designed to incorporate plasma samples from three distinct groups: negative, low FF and positive results after standard NIPT (Fig. 1). In the current study, we included 277 plasma samples from singleton pregnant women. From January 2021 to December 2021, our center issued 24,021 standard NIPT reports. All of our standard NIPT patients had consented to participate in this de-identified study. From these patients, we selected plasma samples from those with 200 negative results, 49 with low FF from their first blood samples and 19 with positive results. The positive cases included patients with common trisomies, CNVs and multiple chromosome abnormalities (MCA). Sample selection in these three categories were random.
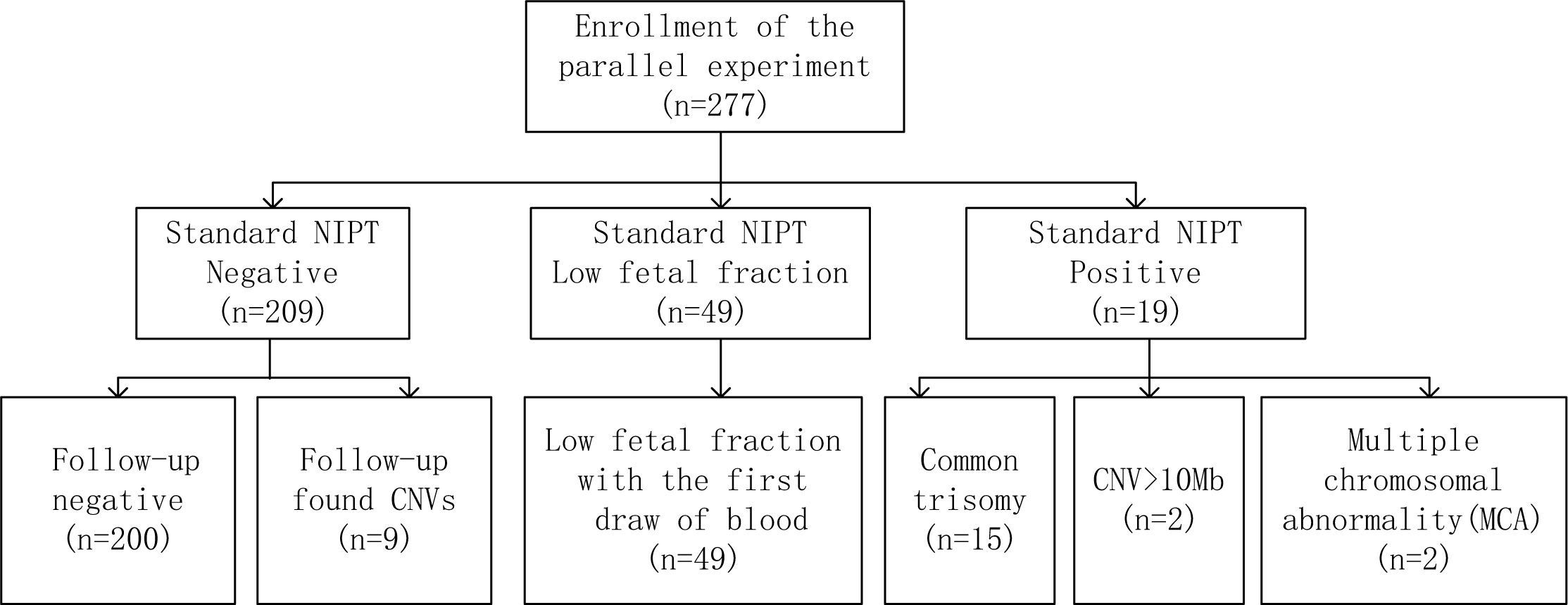 Fig. 1.
Fig. 1.A flowchart to explain the details of patients’ enrolled in this study. NIPT, noninvasive prenatal testing; CNV, copy number variant.
In another nine patients with standard NIPT-negative pregnancies, amniotic fluid
single nucleotide polymorphism (SNP) array testing during their later stages was
performed, and in these, the presence of fetal CNVs was confirmed. These patients
were involved in a parallel study where the effects of NIPT with FF enrichment
(NiftyPro, BGI, Wuhan, Hubei, China) in detecting fetal CNVs were investigated.
Therefore, the total number of samples in the standard NIPT-negative group
totaled 209. The maternal ages and gestational weeks of the three groups were not
different (Table 1). However, the BMI of the low FF group was statistically
higher than the other groups (p
| Category | Standard NIPT results without enrichment | p value | ||
| Negative | Low fetal fraction | Positive | ||
| Group size (N) | 209 | 49 | 19 | N/A |
| Maternal age (years, range) | 32 (18–42) | 32 (22–42) | 33 (16–38) | |
| Gestational week (weeks, range) | 17 (12–32) | 18 (12–22) | 18 (15–22) | |
| Body mass index (BMI, range) | 23 (17–34) | 25* (14–32) | 22 (15–29) | |
*BMI in the group with “Low Fetal Fraction” results were significantly higher than the “Negative” and “Positive” groups. No other statistical significance was found within the groups. N/A, none available.
Pregnancies with positive NIPT results were given a post-test counseling session and advised to proceed with invasive prenatal diagnostic testing. The prenatal invasive diagnostic test was accomplished through amniocentesis, and the samples were analyzed by using SNP arrays (HumanCytoSNP-12, Illumina, San Diago, CA, USA). CNV genomic coordinates were based on the Genome Reference Consortium Human Build 37 (GRCh37). Pregnant women with negative results were also counseled and monitored for their routine pregnancy check-ups. An after-birth follow-up session was provided for each NIPT negative case. The nine standard NIPT negative cases, which were later found to harbor fetal CNVs (Fig. 1), were identified and recovered through this process. The positive cases identified solely by NIPT with enrichment underwent counseling and opted for postnatal SNP array karyotyping using their infants’ peripheral blood.
For each pregnant woman, 5 mL of peripheral blood was collected into a cfDNA storage tube (CoWin Bioscience, Taizhou, Zhejiang, China). For standard NIPT, wet lab procedures were performed according to the NIFTY assay (BGI, Wuhan, Hubei, China) [19]. Procedures for NIPT with enrichment included an extra step of DNA size selection by magnetic beads [18]. This extra size selection was performed after extracting maternal plasma DNA and was followed by DNA-end repair. NIPT with enrichment required five additional cycles of polymerase chain reaction (PCR) compared to the standard NIPT.
For the standard NIPT, the mean raw data for each sample was approximately 15 Mb (35 bp+10 bp index) generated by the BGISEQ500 (BGI, Wuhan, Hubei, China) and each of the mapped unique reads was around 10 Mb. For NIPT with enrichment, 0.15× genome-wide sequence data were generated by the BGISEQ500, which involved extra cycles in PCR amplification. The mean raw data was about 47 Mb (35 bp+10 bp index) for each sample, and the corresponding mapped unique read was around 23 Mb each. K-mer (where K denote the sequencing reads length) coverage was introduced on each chromosome, including the Y chromosome, to eliminate GC bias and determine fetal gender and sex chromosomal abnormalities [19]. A hidden Markov model was proposed as a method for identifying CNVs [20]. When the FF of a sample was lower than 3.5%, the sequencing data from that sample was considered to be inadequate for bioinformatics analysis and denoted as ‘low fetal fraction’.
Statistical significance between two categorical variables was estimated by Fisher’s exact test. Student’s t-test was used to evaluate whether two groups differed from each other. A p value less than 0.05 was considered to be statistically significant. The statistical calculations were performed by using IBM SPSS Statistics (version 22.0, IBM Corp., Armonk, NY, USA).
The average FF from standard NIPT was 9.86%, and after enrichment the average FF was 18.56% (n = 277). The enrichment method, in general, resulted in a 1.9-fold increase in the FF. However, the DNA library for each sample dropped from an average of 24.5 ng/μL when using the standard NIPT method to 12.1 ng/μL with the introduction of FF enrichment. This represents a 2.0-fold decrease in the DNA libraries obtained. All the sample library construction obtained with standard NIPT were normal. However, one sample failed during library construction in NIPT with enrichment due to a low concentration which was less than 2.5 ng/μL.
The BMI of the participants in this study were divided into four groups [21]:
underweight (BMI
| BMI defined groups | Group Size (N) | Mean gestational weeks | Cases of low FF without enrichment | Cases of low FF with enrichment | Mean FF (%) without enrichment | Mean FF (%) with enrichment | Mean FF increasing fold | p value* |
| Underweight ( |
16 | 17.2 | 1 | 0 | 13.5 | 25.1 | 1.9 | |
| Normal (18.6–24.9) | 175 | 17.1 | 22 | 4 | 10.4 | 19.3 | 2.0 | |
| Overweight (25.0–29.9) | 76 | 17.2 | 21 | 2 | 7.6 | 15.4 | 2.2 | |
| Obese ( |
10 | 17.8 | 5 | 0 | 6.1 | 13.5 | 2.4 |
*FF increasing folds in the groups of “Overweight” and “Obese” are significantly higher than the group of “Normal” and “Underweight”. No statistical significance in FF increasing fold was found between the “Normal” and “Underweight” groups, nor between the “Overweight” and “Obese” groups.
277 samples were tested in parallel by NIPT with and without FF enrichment. In
our selected samples, the rate of low FF in NIPT dropped significantly from 17.7
(49/277) to 2.2% (6/277) with the additional enrichment step (p
| Categories | NIPT without FF enrichment (N) | NIPT with FF enrichment (N) | p value | |
| NIPT results: | Negative | 209 | 239 | |
| Low FF | 49 | 6 | ||
| Common Trisomy | 15 | 14 | ||
| Copy number variant (CNV) | 2 | 16 | ||
| Sex Chromosome Abnormality (SCA) | 0 | 1 | ||
| Multiple Chromosome Abnormality (MCA) | 2 | 3 | ||
| Trisomy concordant | 7 | 7 | ||
| Trisomy false positive | 8 | 7* | ||
| Trisomy positive predictive value (PPV, %) | 47 | 50 | N/A | |
| CNV concordant | 1 | 12 | ||
| CNV false positive | 1 | 4 | ||
| CNV PPV (%) | 50 | 75 | N/A | |
| MCA concordant | 0 | 0 | ||
| MCA false positive | 2 | 3* | ||
| SCA concordant | 0 | 0 | ||
| SCA false positive | 0 | 1* | ||
| Failure in DNA library construction | 0 | 1 | ||
The details of all CNVs and samples related to CNVs included in this study can be found in Table 4.
*One sample in each of the three corresponding categories: trisomy false positive, MCA false positive and SCA false positive, were derived from ‘Low FF’ samples without FF enrichment.
The occurrence of calls for seven concordant common trisomies, including five T21 and two T18, remained the same in NIPT with and without FF enrichment (Table 3). Two false positive calls were made by standard NIPT, one T18 and one T13, and they were eliminated by enrichment NIPT. Since there was one extra false positive T13 noted by the enrichment NIPT from the low FF group, the total number of trisomy false positive cases added up to seven in those who had NIPT with FF enrichment. The positive predictive value (PPV) of common trisomies was 47% (7/15) in standard NIPT and 50% (7/14) in NIPT with enrichment. The sample with a result of MCA in standard NIPT yielded the same result in NIPT with enrichment, and the latter also generated one additional false positive MCA from the standard NIPT low FF group.
The version of NIPT with FF enrichment we used in this study was specifically designed to CNVs of size 3 Mb or above. For chromosomal deletions in the 22q11.2 region, the minimum resolution was set to 1 Mb. In the standard NIPT, out of the two CNV calls (call number 3 and 14, Table 4), only call number 14 was concordant. The PPV for CNV in standard NIPT, without enrichment, was 50% (1/2, Table 3). For NIPT with enrichment, there were significantly more CNV calls and concordant CNVs made. Out of 16 CNV calls, 12 were concordant with the array confirmation results. The PPV for fetal CNVs in NIPT with enrichment was 75% (12/16, Table 3). This study included a total of 13 CNVs confirmed by SNP-array (Table 4). Call number 11 had negative results in NIPT with and without enrichment. The corresponding pregnant woman underwent regular check-ups and fetal sonographic abnormalities were detected at 23 weeks gestation. Invasive prenatal diagnosis confirmed that the fetus had a 20–40% mosaic for 22q11.1q11.21 duplication, with a size 4.2 Mb.
| Sample number | Fetal fraction (%) | NIPT Call Number | Result of standard NIPT without enrichment | NIPT with enrichment | SNP array confirmed CNVs coordinates | NIPT vs. Confirmatory results | Identity | |
| Results | Genomic coordinates of NIPT reported CNVs | |||||||
| 1001 | 31.2 | 1 | Normal | del(4q35.1q35.2,4.1M) | chr4:186952297-191044176 | arr4q35.1q35.2(187017191-190880409)x1 | Concordant | TP |
| 1002 | 13.6 | 2 | Normal | del(13q33.3q34,5.22M) | chr13:108764667-113983625 | arr13q33.3q34(108680001-113900000)x1 | Concordant | TP |
| 1003 | 30.7 | 3 | del(5p15.33p15.2,10.34M) | T16* | N/A | arr16p12.2p11.1(22797852-35069526)x3 | Concordant | TP |
| 1004 | 39.1 | 4 | Normal | dup(9p21.2p24.3,27.45M) | chr9:363154-27814977 | N/A | Discordant | FP |
| 5 | Normal | dup(Xp22.33q12,64.1M) | chrX:3004178-67102979 | N/A | Discordant | FP | ||
| 6 | Normal | del(17p12,1.43M-Maternal)* | N/A | arr17p12(14040338-15551871)x1 | Concordant | TP | ||
| 1005 | 17.0 | 7 | Normal | del(22q11.21,1.72M) | chr22:18778537-20501581 | arr22q11.21(18895703-21928916)x1 | Concordant | TP |
| 1006 | 15.8 | 8 | Normal | del(22q11.21,1.72M) | chr22:18778537-20501581 | arr22q11.21(18889490-21460220)x1 | Concordant | TP |
| 1007 | 15.8 | 9 | Normal | del(7q11.23,1.76M) | chr7:72726282-74482696 | arr7q11.23(72350815-74138121)x1 | Concordant | TP |
| 1008 | 18.8 | 10 | Normal | del(22q11.21,3.17M) | chr22:18778537-21950233 | arr22q11.21(18844632-21460220)x1 | Concordant | TP |
| 1009 | 20.2 | 11 | Normal | Normal | N/A | arr22q11.1q11.21(17227461-21433331)x2~3 | Discordant | FN |
| 1010 | 25.9 | 12 | Normal | del(11p14.1p14.3,6.4M) | chr11:22148844-28728526 | arr11p14.1p14.3(22348844-28989610)x1 | Concordant | TP |
| 13 | Normal | del(21q11.2q21.3,11.7M) | chr21:14653457-27375870 | arr21q11.2q21.3(14795475-27486796)x1 | Concordant | TP | ||
| 1011 | 25.2 | 14 | T11 | dup(11q23.3q25,18.26M) | chr11:116667341-134931182 | arr11q23.3q25(116575754-134944006)x3 | Concordant | TP |
| 15 | Normal | dup(22q11.1q11.21,3.3M) | chr22:18277537-21650233 | arr22q11.1q11.21(17227461-20966125)x3 | Concordant | TP | ||
| 1012 | 19.7 | 16 | Normal | dup(3p26.1p26.3,5.48M) | chr3:563399-6048293 | N/A | Discordant | FP |
| 17 | Normal | del(4p15.31p16.3,17.17M) | chr4:1551181-18725179 | N/A | Discordant | FP | ||
Genomic coordinates of CNVs were based on the Genome Reference Consortium Human Build 37 (GRCh37). TP, true positive; FP, false negative; FN, false negative; SNP, single nucleotide polymorphism.
*The version of the NIPT report analysis system we used did not provide genomic
coordinates for trisomy or maternal CNV (
The current clinical guidelines support the use of NIPT in detecting common fetal trisomies in all pregnant women [4, 5]. The use of fetal cfDNA screening for genome-wide fetal CNVs, pathogenic as well as likely pathogenic, has not as yet been recommended by the formal clinical guidelines. This was partly due to concerns about the lack of sufficient and specific experimental parameters (e.g., sensitivity, specificity and PPV) for the application of NIPT in different types of CNVs. However, in some updates and supplements to the formal guidelines, there have been thorough discussions regarding the use of NIPT beyond the screening of common trisomies. Local experiences in the clinical application of expanded NIPT with respect to CNVs have also been shared [22, 23]. Although expanding the target of NIPT will be likely to increase the rate of invasive diagnosis in pregnant women, for now, this is the only screening method of CNVs and SCAs offered that has the potential of detecting CNV-related syndromes that may not show distinct abnormalities on fetal ultrasound imaging [12, 24].
In this study, we tested the performance of NIPT with an additional step of FF enrichment using magnetic nanoparticles. Through parallel testing with clinical samples, we observed a robust increase in the FF by using this new approach, particularly in pregnant women with high BMIs. In addition, significantly fewer cases of low FF were observed in NIPT with enrichment. Samples that failed enrichment NIPT due to a low FF all had an extremely low fetal fraction when the standard NIPT was used. Such conditions may be related to CNVs in the Y chromosome, as we encountered in this study, or they may be associated with other complications during pregnancy [25]. As is recommended in standard NIPT, no-calls in NIPT with enrichment should be considered for prenatal diagnostic procedures. We observed a general decrease in DNA library concentration after adopting FF enrichment. As limited samples were included in this test for NIPT with enrichment, the failure rate of DNA library construction may be lower when pooling from larger experimental cohorts.
Maternal obesity was associated with failures in NIPT due to a low FF, and some guidelines have previously suggested a specific weight limit [26, 27]. Due to the rapid turnover of adipocytes, high concentrations of cfDNA have been linked to obesity [28]. Thus, there is usually an excess of maternal-origin cfDNA present in the peripheral blood plasma of obese pregnant women. The length of fetal-origin cfDNA typically peaks at 143 bp, while maternal cfDNA peaks at 166 bp [29]. The platform we adopted for FF enrichment further selected DNA fragments below 160 bp using magnetic beads. This size threshold was designed to strike a balance between cost, labor and effectiveness. With the implementation of FF enrichment, a process that effectively removes excess maternal and preserves fetal cfDNA, overweight and obese women had an increased likelihood of obtaining more stable results from NIPT [28, 29]. However, the specific weight and BMI limits for NIPT with enrichment remains to be discovered by future studies with larger and different populations.
NIPT with enrichment was reported with enhanced z-scores and sensitivity [18, 30]. Our study reflected similar outcomes, with more distinguished z-scores for the trisomy positive samples (Supplementary Fig. 1) and a significantly increased number of CNVs calls (Table 3). Even though the enrichment method successfully reduced the low FF rate in NIPT, false positive calls for trisomy and MCA still remained. MCA was found to be associated with maternal tumors [31], and false positive calls for trisomy in NIPT were mainly due to feto-placental mosaicism [32]. In addition, maternal background and placental origin interferences are still technical issues that NIPT cannot readily address.
Based on this pilot study, NIPT with enrichment showed satisfactory performance in detecting common trisomies and CNVs. NIPT with enrichment did not outperform the standard NIPT in terms of the abilities to detect the common trisomies (T21/T18/T13). However, due to the higher FF obtained, the z-scores of the positive cases were higher in the enriched NIPT making them more easily distinguishable. For CNVs, we reported a PPV of 75% for NIPT with enrichment and the detection of more concordant and smaller CNVs: fragments with size of 3 Mb and above in general, and 1 Mb and above for the 22q11.2 deletion. With the FF enrichment, we were able to screen out three cases of 22q11.2 deletion syndrome, one case of 16p11.2-p12.2 duplication syndrome and several other clinically significant pathogenic CNVs. It is noteworthy that the NIPT estimated size for a 2.5 Mb 22q11.2 deletion was 1.72 Mb in the region of chromosome 22q11.21, which suggested there is still a discrepancy between the estimated size of the CNVs using the screening method and the size measured by diagnostic method. This size difference may be due to the bioinformatics analysis of the NIPT platform that we chose. It may also be due to the fact that the NIPT technique was analyzing placental-originated cfDNA [32, 33].
Low percentage of mosaicisms cannot be detected by NIPT even with the inclusion of FF enrichment. A 20–40% mosaicism of a 3.7 Mb duplication at 22q11.1q11.21 (as was seen in sample no. 1009 in Table 4) showed negative results in NIPT with and without enrichment. The PPV for CNVs in our selected samples was 75%, which compared favorably to other studies, which ranged from 19 to 47% [12, 34]. The introduction of CNV screening into NIPT would likely result in additional invasive procedures and genetic counseling. According to this study, CNV screening resulted in a minimum false positive rate of 25%, assuming there were no false negatives in our screened negative cohort. We suggest that this enhanced version of NIPT should only be recommended when comprehensive counseling is available. Clinical validation of this enrichment approach should be further carried out with larger and unselected samples. Our recruitment was limited in number, and therefore, does not represent the general population.
Another study by Liang et al. [35] also performed enrichment of the FF during prenatal screening. However, their study differed from this one as it focused mainly on aneuploidies whereas we were also interested in CNVs. In addition, the technique used for FF enrichment was gel electrophoresis, while we employed magnetic nano-beads, which are more compatible with a large-scale DNA extraction and library construction system. In addition, our interest was focused on the clinical performance of FF enrichment, especially in terms of BMI and its effect on the enrichment obtained.
The data from this small-scale pilot study demonstrated the clinical usefulness of NIPT with FF enrichment. The implementation of FF enrichment requires little additional cost and labor, although it is likely to render the results from NIPT more reliable in obese pregnant women and it is capable of detecting fetal CNVs as small as 3 Mb. In a similar way to standard NIPT, restriction for mosaicism and maternal cancer are retained in NIPT with enrichment. Patients with negative results should always follow routine ultrasound tests during their pregnancies. Comprehensive pre- and post-test genetic counseling should be provided for individual cases. This study will increase the awareness of NIPT with FF enrichment and encourage researchers to conduct future studies with larger and different populations. The combination of an advanced version of NIPT and ultrasound may eventually serve as an effective clinical tool to identify a wider range of fetal sub-chromosomal CNVs.
The supporting data of findings in this study are available from the author, YC, upon reasonable request.
YC, JY, XH, SY, YT and JS carried out the experiments, data collection and data analysis. YL and HW made substantial contributions to the conception and design of the project. YC designed and performed the experiments. YC prepared the manuscript. YS provided substantial contributions to the conception of the manuscript and revised it critically for important intellectual content. All authors contributed to editorial changes in the manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work. All the authors discussed the results and approved the manuscript for submission.
This retrospective clinical study was approved by the Ethics Committee of the Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region, with the approval number of [2019]-2-17. All of our standard NIPT patients gave their written consent for participation in this de-identified research.
We thank Dr. Jingsi Luo from the Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region for administrative support for the experiments and manuscript production. The authors also thank Dr. Dev Sooranna of Imperial College London for editing the language of this manuscript.
This study was supported by The Guangxi Zhuang Autonomous Region Health Commission [project number Z20190827].
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.

