1 First Clinical Medical College, Nanjing Medical University, 210009 Nanjing, Jiangsu, China
2 Department of Nursing, Jiangsu Cancer Hospital/Jiangsu Institute of Cancer Research/The Affiliated Cancer Hospital of Nanjing Medical University, 210009 Nanjing, Jiangsu, China
Abstract
Background: Most patients with advanced-stage breast cancer present with bone metastasis, which seriously affects their quality of life and prognosis. We aimed to investigate the potential role of and the mechanism of circular Forkhead box protein P1 (FOXP1) (hsa_circ_0008234) in bone metastasis of breast cancer. Methods: The Gene Expression Omnibus database (GEO) database (GSE111504) was used to screen the differentially expressed circular RNAs (circRNAs) in metastatic breast cancer. The related expression of circular FOXP1 (circFOXP1) and miRNA was measured by real-time quantitative reverse transcription polymerase chain reaction (RT-qPCR). Cellular experiments were performed to assess the influence of circFOXP1 in breast cancer cells. After co-culture of circFOXP1 siRNA-transfected MDA-MB-231 cells and bone marrow-derived mesenchymal stem cells (BMSCs), the effect of circFOXP1 on osteogenic genes was detected. Bioinformatic Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analyses were performed using clusterProfilter 4.8.2 and R package version 4.3. Results: circFOXP1 was upregulated in patients with breast cancer, particularly in bone metastasis breast cancer. Silencing of circFOXP1 decreased the abilities of proliferation, migration, and invasion. The increased alkaline phosphatase (ALP) activity and osteogenic gene expression of BMSCs co-cultured with the MDA-MB-231/si-circRNA group was observed. miR-338-3p was a target miRNA of circFOXP1. Bioinformatic enrichment analysis indicated that the targeted mRNAs were involved in in MAPK pathway, regulation of actin cytoskeleton, tight junction, Ras pathway, and PI3K-AKT pathway. Conclusions: circFOXP1 upregulation was related to bone metastasis of breast cancer. Silencing of circFOXP1 in breast cancer cells might repress breast cancer cellular activities and facilitate osteogenetic differentiation of BMSCs in the microenvironment by targeting miR-338-3p. circFOXP1 might be a therapeutic target for patients with bone metastasis of breast cancer.
Keywords
- hsa_circ_0008234
- breast cancer
- bone metastasis
- miR-338-3p
- bioinformatics
The incidence of breast cancer in women is increasing annually and is displaying a tendency to affect younger women [1]. Breast cancer is a malignant tumor prone to distant metastasis, some cases report distant metastasis during the initial diagnosis [2]. Breast cancer metastasis is organ-selective and bone metastasis occurs in approximately 75% of these patients [3]. Bone metastasis of breast cancer can lead to osteolytic bone damage, and often cause a series of bone-related events, such as pathogenic fracture, nerve compression (including spinal cord compression), hypercalcemia, and limb dysfunction, which seriously affects the quality of life of patients and even leads to death [4]. Comprehensive treatment (surgery, radiotherapy, endocrine therapy, chemotherapy, and targeted therapy) is principally used for bone metastasis of breast cancer, and predominantly aims is to improve survival [5]. Therefore, it is particularly important to explore the molecular mechanism underlying the bone metastasis of breast cancer and identify effective therapeutic targets.
Several genes or molecules with a regulatory role are expressed aberrantly
during malignant tumor development, which may serve as novel targets or
biomarkers, and circular RNAs (circRNAs) are one of the research hotspots [6, 7].
circRNAs refer to a group of endogenous non-coding RNAs characterized by
covalently closed-loop structures [8]. The aberrant expression of circRNAs
results in potential diagnostic or prognostic performance in tumors [9, 10]. Some
circRNAs are potential therapeutic targets for cancers and bone metastatic
treatment [11, 12]. For instance, circIKBKB is upregulated in patients with
bone-metastasis of breast cancer and can facilitate bone metastasis by activating
the NF-
This study detected and screened the circRNAs expressed differentially in metastatic breast cancer using the Gene Expression Omnibus database (GEO) GSE111504. The expression of circFOXP1 was validated in breast cancer tissues (BCa/T) and those with bone metastasis (BM-BCa/T). Subsequently, the function of circFOXP1 in breast cancer cells was evaluated. Bioinformatic analysis was performed to predict the potential miRNAs of circFOXP1. The predicted target genes were used to conduct the protein-protein interaction (PPI) network, and to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrich analysis.
The sample collection was based on the specimen bank of Nanjing Medical University. The study population included patients with breast cancer who underwent surgery at our hospital from May 2019 to January 2023, including 60 patients with bone metastasis of breast cancer and 110 patients with primary breast cancer. The samples of bone metastasis, primary lesions, and adjacent breast cancer tissues were obtained during the surgery. All the samples were quickly frozen in liquid nitrogen. All the patients and their families were informed of the study content and signed an informed written consent form. The study was approved by the Ethics Committee of the Jiangsu Cancer Hospital (Approval No. 2019017).
The circRNAs expressed differentially in metastatic breast cancer were screened
from the GEO database GSE111504. GSE111504 circRNA expression profile was based
on GPL21825 platform from parental MDA-MB-231 (231-PAR) cells, isogenic brain
metastatic cells, and metastatic cells. The volcano map was conducted based on
the condition of
Human breast cancer cells MCF-7 (HTB-22), SK-BR-3 (HTB-30), T47D (HTB-133), and
MDA-MB-231 (CRM-HTB-26), as well as non-tumorigenic epithelial cell line MCF-10A
(CRL-10317), were acquired from American Type Culture Collection (ATCC; Manassas,
VA, USA) with short tandem repeats (STR) verification. All cancer cell lines were maintained in liquid nitrogen and incubated
in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, Grand Island, NY, USA) with
10% fetal bovine serum (FBS; Gibco, Grand Island, NY, USA) at 37 °C in
a 5% CO
circFOXP1 siRNAs (si-circRNA-1, -2, -3), siRNA negative control (si-NC),
miR-338-3p mimic (miR-338-3p), and mimic NC (miR-NC) were synthesized by
GenePharma (Shanghai, China). Transfection of plasmids into breast cancer cells
was performed with the help of lipofectamine 2000 (Invitrogen, Grand Island, NY,
USA). After 48 h of transfection, a real-time quantitative reverse transcription
polymerase chain reaction (RT-qPCR) was performed to detect the transfection
efficiency. The sequences for circFOXP1 siRNAs were as follows: si-circRNA-1,
5
The co-culture model was performed by applying the medium supernatant of different transfected cells to the BMSCs culture. The co-cultured BMSCs were incubated for 1, 3, 5, 7, and 9 days to detect alkaline phosphatase (ALP) activity.
The clinical samples were cut into sizes measuring approximately 0.5
After co-culture, the cells were collected and homogenized with RIPA lysis buffer to obtain total protein. The protein purity and concentration were measured by standard methods of BCA assays (Beyotime, Shanghai, China). 10 µg per sample were separated by SDS-PAGE for 90 min, then blotted onto nitrocellulose membranes for 1 h, and incubated with primary antibodies (Abcam, Cambridge, UK) overnight at 4 °C. Nitrocellulose membranes were washed four times and treated with appropriate secondary HRP-conjugated goat anti-rabbit or goat anti-mouse antibodies (Abcam, Cambridge, UK) at 37 °C for 1 h. Finally, membranes were scanned and transferred to the Image J program (version 1.53, National Institutes of Health, Bethesda, MD, USA) for gel gray measurement.
The cell proliferative capacities were assessed using cell counting kit-8 (CCK-8, Dojindo, Tokyo, Japan). The transfection-treated breast cancer cells were incubated in 96-well plates (approximately 5000 cells/well). The CCK-8 reagent was added to each well at 0, 24, 48, and 72 h time points. The cells were incubated for further 1 h. The micro-plate Reader (Bio-Rad, Hercules, CA, USA) was used to determine the absorbance at a wavelength of 450 nm (OD value).
ALP activity of co-cultured BMSCs was measured using an Alkaline Phosphatase Activity Assay Kit (Beyotime, Shanghai, China).
DMEM medium without serum was put on the upper layer of the
Transwell culture plate (pore size = 8 µm) coated with or without
Matrigel (Corning, Corning, NY, USA), and DMEM medium containing 10% FBS was added to the
lower layer. The cell suspension (5
The circFOXP1-wild type (WT-circRNA) or circFOXP1-mutant (MUT-circRNA) binding sites with miR-338-3p were inserted into the pGL3 promoter vector (Realgene, Shanghai, China). Breast cancer cells were cultured and seeded in 24-well plates. miR-338-3p mimic, mimic NCs, and luciferase reporter plasmids (WT-circRNA or MUT-circRNA) were co-transfected into the cells when cells grew to 80% confluence. After 48 h of incubation, firefly luciferase activities were measured using Dual-luciferase Reporter Assay System (Promega, Madison, WI, USA).
The statistical data are expressed as mean
According to GSE111504 circRNA profile data, the volcano map was based on the
condition of
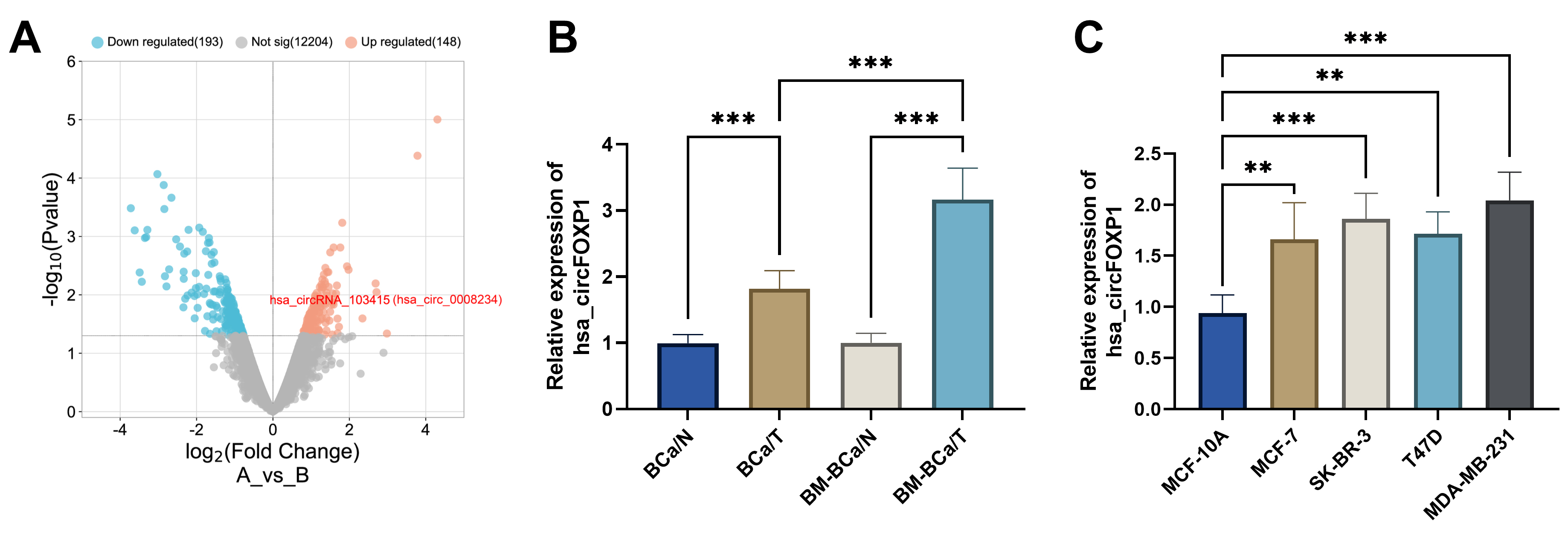 Fig. 1.
Fig. 1.circFOXP1 expression in breast cancer. (A)
Volcano plot of circRNAs from GES111504 and screened upregulated circFOXP1
(hsa_circ_0008234). (B) circFOXP1 expression was detected in breast cancer bone
metastasis tissues, primary breast cancer tissues, and surrounding tissues. (C)
circFOXP1 expression was increased in breast cancer cells in contrast with that
in normal MCF-10A cells. **p
circFOXP1 siRNAs (si-circRNA-1, si-circRNA-2, si-circRNA-3) were transfected
into SK-BR-3 MDA-MB-231 cells. Fig. 2A,B depict that si-circRNA-1 exhibited the
highest inhibitory effect on circFOXP1 expression (p
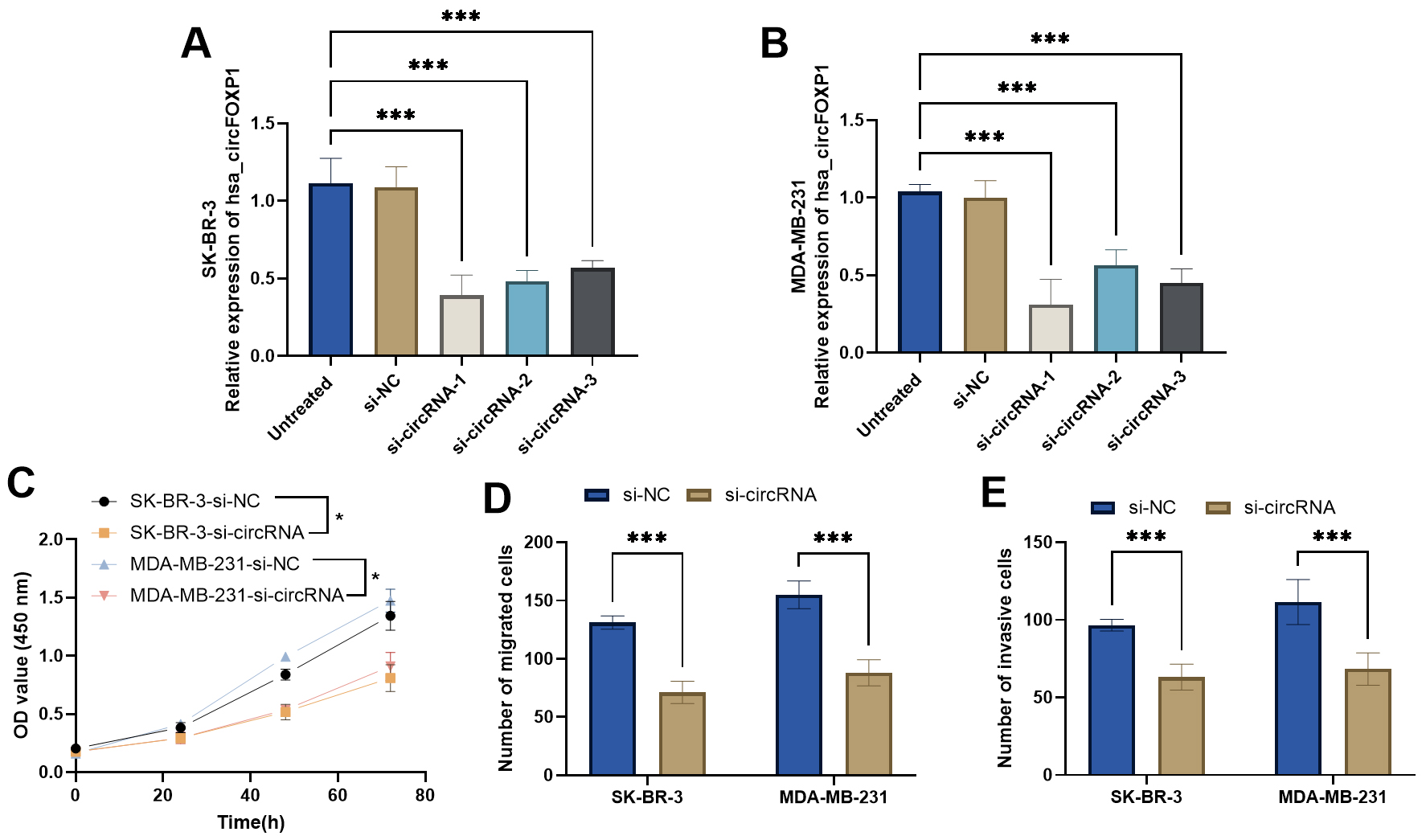 Fig. 2.
Fig. 2.Functional role of
circFOXP1 in breast cancer cells. (A,B) circFOXP1 siRNAs decrease circFOXP1
expression in SK-BR-3 and MDA-MB-231 cells. (C) CCK-8 assay measured the
influence of circFOXP1 siRNA in breast cancer cells. (D,E) circFOXP1 interference
decreased breast cancer cell migration (D) and invasion (E) abilities.
*p
To investigate the effects of circFOXP1 on the interaction between breast cancer
cells and BMSCs, the supernatant of MDA-MB-231 cells cultured in si-circRNA group
and si-NC group were collected and co-cultured with BMSCs. The increased ALP
activity of BMSCs co-cultured with MDA-MB-231/si-circRNA group was observed
(p
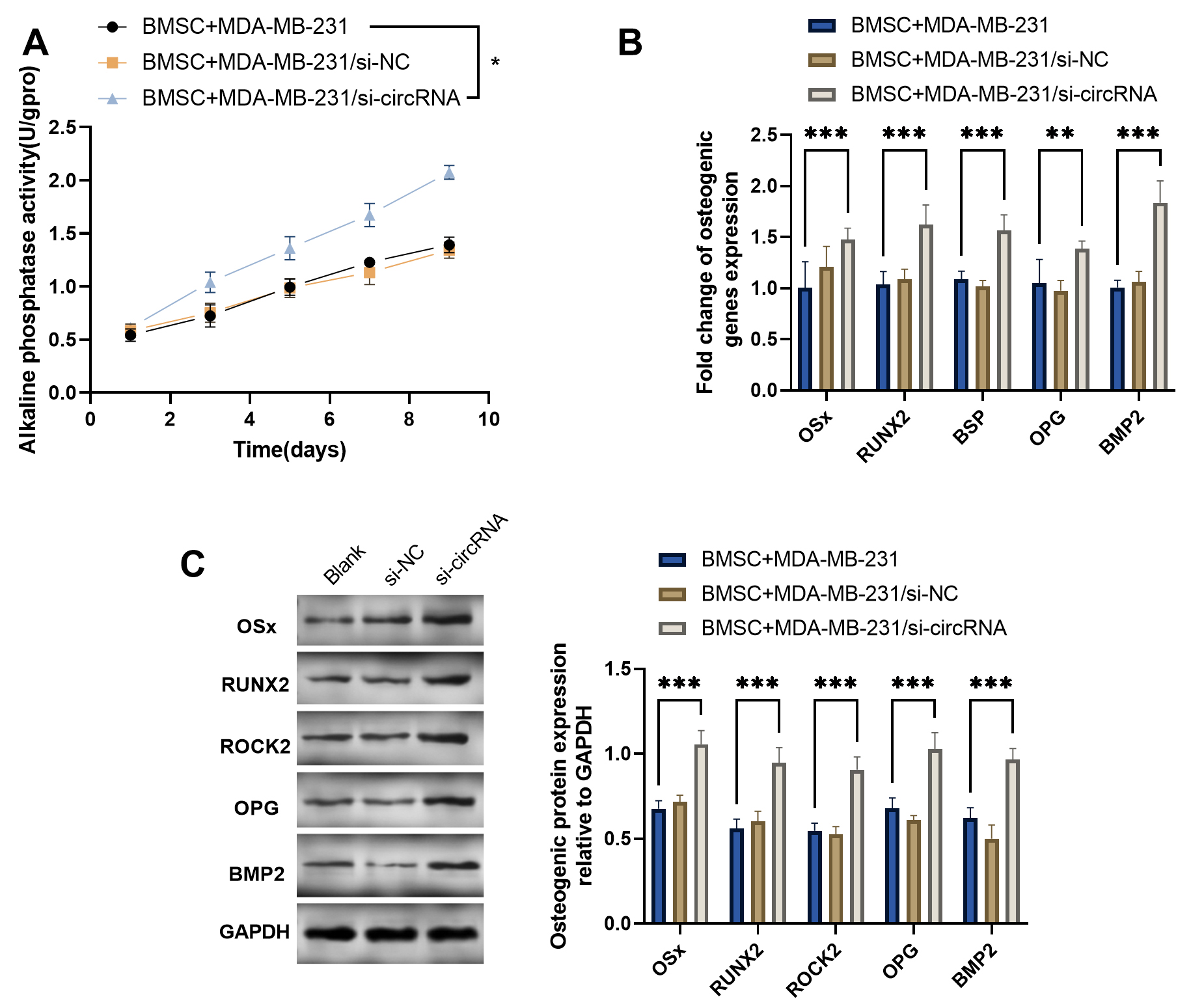 Fig. 3.
Fig. 3.circFOXP1 silencing in MDA-MB-231 improves the BMSCs
osteogenesis. (A) Alkaline phosphatase activity analysis of BMSCs co-cultured
with transfected MDA-MB-231 culture supernatant. (B) The osteogenic genes
OSx, RUNX2, BSP, OPG, and BMP2 in
BMSCs after co-culture. (C) Osteogenic proteins expression of BMSCs after
co-culture. *p
Two circRNA-miRNA interaction databases (circInteractome and circBank) were used
to screen the intersection of miRNAs that may play regulatory roles. These
databases overlapped five potential miRNAs (Fig. 4A). Among the five miRNAs,
miR-338-3p exhibited the highest levels after silencing of circFOXP1 in
MDA-MB-231 cells, which was selected to validate the targeting relationship with
circFOXP1 (p
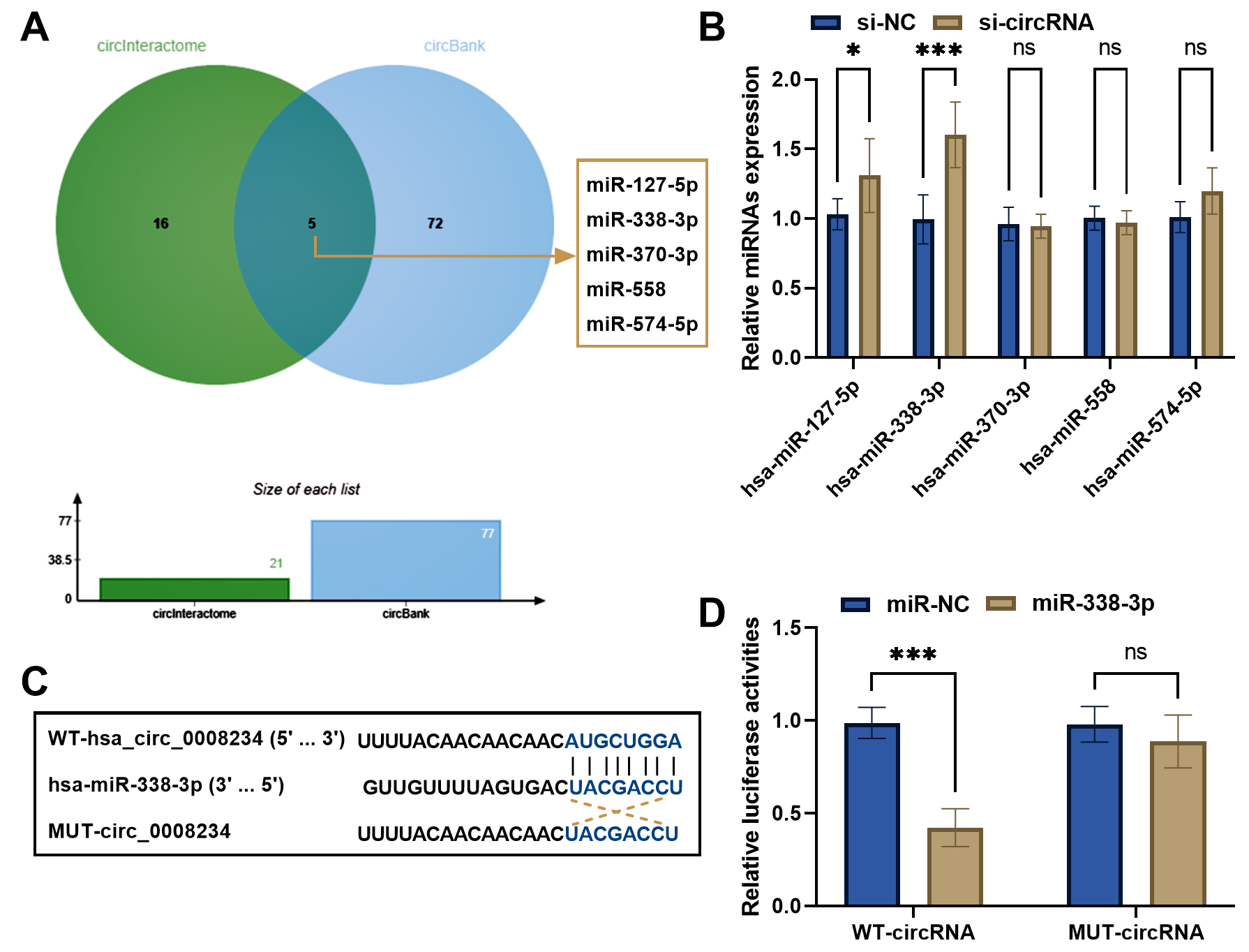 Fig. 4.
Fig. 4.circFOXP1 could sponge miR-338-3p. (A) circInteractome and
circBank databases overlapped the potential predictive miRNAs of circFOXP1. (B)
Relative expression of miRNAs was measured in si-circFXOP1 transfected MDA-MB-231
cells. (C) Binding sites between circFOXP1 and miR-338-3p. (D) Dual-luciferase
reporter assay confirmed the relationship between circFOXP1 and miR-338-3p.
*p
To explore the potential molecular mechanism underlying circFOXP1, TargetScan, miRDB, miRWalk, and starBase algorithms were used to overlap the potential target mRNAs of miR-338-3p (Fig. 5A). The STRING online database predicted PPI using the overlapped 92 mRNAs of miR-338-3p (Fig. 5B) and the top ten target mRNAs (PTEN was the leading mRNA) were visualized by Cytoscape (Fig. 5C). The GO (Fig. 5D) and KEGG (Fig. 5E) pathway enrichment analysis indicated that the targets genes were principally enriched in dendrite development, morphogenesis, and other biological processes, and pathway enriched in MAPK signaling pathway, regulation of actin cytoskeleton, tight junction, Ras signaling pathway, and PI3K-AKT signaling pathway.
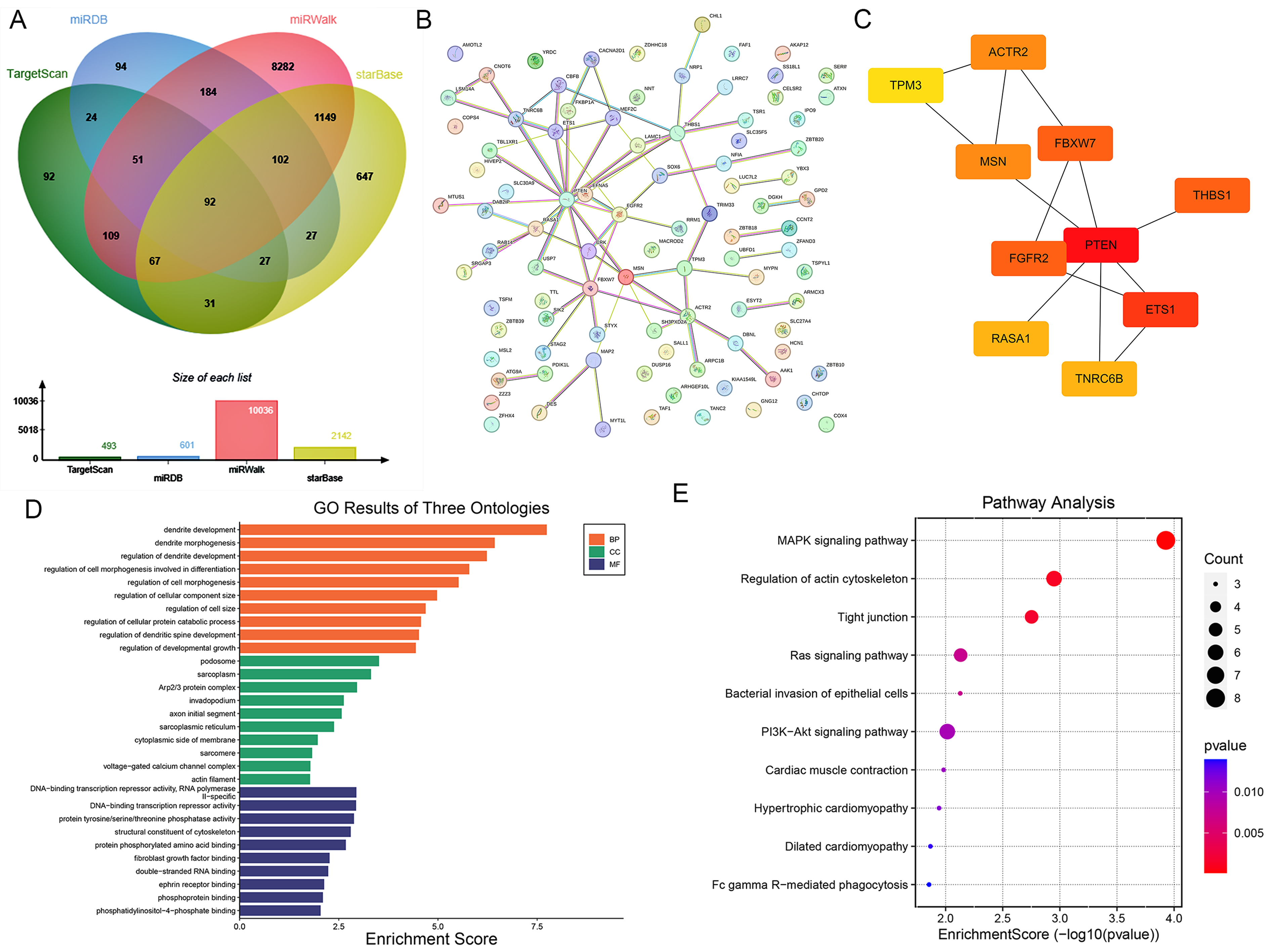 Fig. 5.
Fig. 5.Downstream mRNAs of miR-338-3p and pathway enrichment analysis. (A) Downstream mRNAs of miR-338-3p were overlapped using four online databases. (B) STRING conducted a protein-protein interaction (PPI) network using overlapped 92 mRNAs. (C) The leading ten hub genes were identified using the Cytoscape network. (D) Gene Ontology (GO) analysis was performed using the related genes in the network. (E) Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway enrichment analysis of the genes in the network genes.
Most patients with breast cancer develop bone metastasis during the disease course, and the treatment regimens are unsatisfactory, so it is crucial to seek novel therapeutic targets for treating breast cancer, particularly for patients with bone metastasis of breast cancer. The GSE111504 dataset exhibited upregulated circFOXP1 in metastatic breast cancer cells. Aberrant expression of circFOXP1 participated in the tumorigenesis of several types of cancers [13, 14]. However, the detailed function of circFOXP1 in breast cancer, particularly in bone metastasis remains elusive. Here, the circFOXP1 levels were increased in metastatic breast cancer tissues, especially in bone metastatic breast cancer tissues, as well as breast cancer cells. Interfering circFOXP1 expression could repress cellular activities, including proliferative capacities, migration abilities, and invasion potential. Importantly, silencing of circFOXP1 could increase ALP activities and osteogenic gene expression. Bioinformatic analysis displayed that the network genes were enriched in several signaling pathways related to bone metastasis.
circFOXP1 was overexpressed in breast cancer tissues and cells, particularly in bone metastatic breast cancer tissues, suggesting the potentially crucial role of circFOXP1 in the tumorigenesis of breast cancer and bone homeostasis. The functional significance of circFOXP1 has been reported in several types of cancers, including the promoting role of circFOXP1 in cutaneous squamous carcinoma by facilitating cell proliferation through miR-127-5p/ADCY7 axis regulation [13]. circFOXP1 has identified as a prognostic biomarker and regulator in gallbladder cancer progression and the Warburg effect by regulating miR-370/PKLR expression [15]. Besides, the inconsistent role of circFOXP1 in different subtypes of lung cancer was reported, which may be attributed to the circFOXP1 expression measurement in different samples [14, 24, 25]. In this study, the functional performance of circFOXP1 was explored. Silencing of circFOXP1 displayed anti-neoplastic properties through inhibiting proliferative, migratory, and invasive potentials, suggesting that circFOXP1 might play a tumor-promoting role in breast cancer tumorigenesis.
The concentration of bone metabolic markers for numerous bone disease diagnoses, the assessment of fracture risk prediction, and drug efficacy, are valuable. The present study tried to measure the ALP activities and osteogenic genes (OSX, RUNX2, BSP, OPG, and BMP2) expression in BMSCs co-cultured with MDA-MB-231/si-circRNA group. OSX, RUNX2, BSP, and BMP2 could reflect osteoblastic differentiation ability and OPG was identified as a refection marker of osteoclastogenesis [26, 27]. Increased ALP activities and osteogenic gene expression in BMSCs co-cultured with MDA-MB-231/si-circRNA group illustrated that circFOXP1 silencing in breast cancer might facilitate the osteogenic capacity of BMSCs in the microenvironment of bone metastasis of breast cancer.
The dysregulation of circRNAs may be a key factor in biological processes such as post-transcriptional regulation and epigenetic modification of malignant rumors by regulating miRNA expression. The current study observed that miR-338-3p was a target of circFOXP1, illustrating that circFOXP1 may participate in tumorigenesis of breast cancer by targeting miR-338-3p. miR-338-3p was a tumor suppressor in breast cancer and could be sponged by circTFF1 [28, 29]. In addition, osteogenic gene RUNX2 might be a target of miR-338-3p, other osteogenic genes did not show targeting relationship with miR-338-3p (Supplementary Fig. 1). These data revealed that circFOXP1/miR-338-3p might affect the osteogenic genes expression by modulating breast cancer bone metastasis, but it might not directly regulate the expression of osteogenic genes through miR-338-3p. Furthermore, the potential mechanism underlying circFOXP1 was explored using bioinformatic enrichment analysis. Among the overlapped targeted mRNAs of miR-338-3p, PTEN was the top hub gene. In addition, the targeted genes were related to biological processes (BP) of dendrite development, podosome cellular component (CC), and DNA-binding transcription repressor activity, RNA polymerase II-specific molecular function (MF). Plasmacytoid dendritic cells were increased with bone metastasis in breast cancer [30]. Pathway enrichment analysis displayed that the targeted mRNAs were involved in various functional pathways, such as MAPK pathway, regulation of actin cytoskeleton, tight junction, Ras pathway, and PI3K-AKT pathway. These pathways were closely related to bone metastasis in tumors [31, 32, 33]. As a complex pathological process, multiple signaling may control the distant metastasis of breast cancer, and the detailed mechanism underlying circFOXP1 in bone metastasis of breast cancer warrants to further investigation.
In conclusion, circFOXP1 upregulation was related to bone metastasis of breast cancer. Silencing of circFOXP1 in breast cancer cells can repress breast cancer cellular activities and facilitate osteogenetic differentiation of BMSCs in the microenvironment by targeting miR-338-3p. circFOXP1 might be a therapeutic target for patients with bone metastasis of breast cancer.
ALP, alkaline phosphatase; BCa/T, breast cancer tissues; BM-BCa/T, bone metastasis breast cancer tissues; BMSCs, bone marrow mesenchymal stem cells; BP, biological processes; CC, cellular component; circRNAs, circular RNAs; GEO, Gene Expression Omnibus database; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; MF, molecular function; miR-NC, mimic NC; RT-qPCR, real-time quantitative reverse transcription polymerase chain reaction; si-NC, siRNA negative control; 231-PAR, parental MDA-MB-231.
All data generated or analyzed during this study are included in this article. Further enquiries can be directed to the corresponding author.
Both authors designed this study. KW and KC conducted the experiment and analyzed the data. KW wrote the manuscript. KC revised the manuscript. Both authors reviewed and approved for publication.
All the enrolled patients and their families were informed of the study content and signed informed written consent. The study was approved by the Ethics Committee of the Jiangsu Cancer Hospital (Approval No. 2019017).
Not applicable.
This research received no external funding.
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.





