1 Department of Gynecology, The Affiliated Wuxi People’s Hospital of Nanjing Medical University, 214023 Wuxi, Jiangsu, China
2 Department of Gynecology, Wuxi Medical Center, Nanjing Medical University, 214023 Wuxi, Jiangsu, China
3 Department of Gynecology, Wuxi People's Hospital, 214023 Wuxi, Jiangsu, China
4 Department of Gynecology, Affiliated Hospital of Jiangsu University, 212000 Zhenjiang, Jiangsu, China
Abstract
Endometrial cancer is one of the common cancers in gynecology, which seriously endangers women’s reproductive health. Therefore, it is urgent to search for new diagnostic and prognostic monitoring markers for endometrial cancer. This study aimed to explore the clinical significance and biological role of spindle and kinetochore-associated complex subunit 3 (SKA3) in endometrial cancer.
Weighted gene co-expression network analysis (WGCNA) and identification of differentially expressed genes (DEGs) were conducted to identify the key gene in endometrial cancer. The clinical significance of SKA3 within endometrial cancer was assessed through receiver operating characteristic (ROC) and Kaplan-Meier (KM) curves. Spearman correlation analysis, the STRING database, Cytoscape software, and the molecular complex detection (MCODE) algorithm were employed to investigate genes associated with SKA3. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were carried out for elucidating the functional role and pathways involving SKA3. The cBioPortal online platform was utilized to explore mutations in SKA3. The biological effects of SKA3 were further investigated through Cell Counting Kit-8 (CCK8) and cell scratch experiments.
SKA3 emerges as a pivotal gene in endometrial cancer, exhibiting a statistically significant high expression level. Its area under the curve (AUC) for diagnosing endometrial cancer stands at 0.943. Patients displaying elevated SKA3 expression demonstrated a notably poorer prognosis. In the context of endometrial cancer, 9 genes directly interact with SKA3. The functional pathway of SKA3 in endometrial cancer likely involves the mitotic pathway. The alterations observed in SKA3 in endometrial cancer primarily manifest as “mutations”. Specifically, SKA3 exhibits 26 mutation sites in endometrial cancer, distributed across 7 distinct regions and involving 4 mutation types. Furthermore, SKA3 is implicated in promoting the proliferation and migration of HEC-1A cells.
SKA3, a key gene in endometrial cancer, holds significant diagnostic and prognostic value and may influence the progression in endometrial cancer.
Keywords
- SKA3
- proliferation
- migration
- endometrial cancer
- clinical value
Endometrial cancer, the prevalent gynecological malignancy, is experiencing a global increase in incidence [1]. In 2020, the incidence rate of endometrial cancer exceeded 400,000 globally, ranking it as the sixth most frequent cancer in female [2, 3]. Age, race, socio-economic status, and geographical differences are important determinants of the incidence and mortality of endometrial cancer [2]. Despite traditional surgery remaining the main treatment for endometrial cancer [4], concerns persist regarding the invasive nature of the surgery and the possibility of recurrence [5, 6]. Therefore, exploring new targets is crucial to assist in diagnosis and reduce the postoperative recurrence rate. Spindle and kinetochore-associated complex subunit 3 (SKA3) is a crucial component of the outer kinetochore involved in microtubule binding, crucial for accurate chromosomal segregation and cell division [7, 8, 9]. A recent study has highlighted the significant status of SKA3 in cancer initiation as well as progression [10]. Researches indicates that SKA3 exhibits elevated expression levels in adrenal cortical carcinoma and renal papillary cell carcinoma, correlating with poor prognosis [9, 11]. You et al. [12] found that SKA3 also exhibited high expression in bladder cancer, promoting its proliferation through accelerating G2/M phase transition. Similarly, SKA3 is highly expressed in breast cancer [13, 14, 15], hepatocellular carcinoma [16, 17, 18], gastric cancer [19, 20], lung cancer [21, 22], and cholangiocarcinoma [23], and is often associated with poor prognosis. Nevertheless, it is unclear about the SKA3 expression and functional mechanisms in endometrial cancer. This research aims to utilize various biological methods to investigate the influence of SKA3 on the onset, development, and prognosis in endometrial cancer, thereby assisting in clinical diagnosis and prognosis monitoring of endometrial cancer.
First, we downloaded the original mRNA expression data and endometrial cancer clinical data from TCGA database (https://portal.gdc.cancer.gov/) using R language, excluded missing data, and integrated them into a gene expression matrix. We then used the R package (https://cran.r-project.org/web/packages/WGCNA/index.html) of weighted gene co-expression network analysis (WGCNA) for performing WGCNA on integrated gene expression matrix, constructing a co-expression network based on gene expression matrix and identifying co-expression modules. For further analysis, we calculated the correlation between each color module and endometrial cancer, selecting the two modules with the highest correlation as key modules for subsequent analysis of the genes involved.
We used the edgeR package to perform differential analysis on the key module
genes and selected differential genes by
During the organization of clinical data, we removed patient information with no
survival status and zero follow-up time, then matched it with SKA3 gene
expression data and corresponding clinical data. Next, we utilized pROC R package
for generating receiver operating characteristic (ROC) curves for further
assessing SKA3 gene’s diagnostic efficacy in endometrial cancer. Subsequently, we
applied survival R package for survival analysis and visualized the survival
curve. We then used the Kaplan-Meier (KM) curve to assess its influence upon
patients’ survival time. Finally, we used the
Spearman correlation analysis was utilized for determining these linking of key module genes and SKA3 genes. For retrieving protein-protein interaction (PPI) networks, 10 genes showing the highest positive correlation with 10 genes exhibiting the strongest negative correlation were imported into the STRING database (https://cn.string-db.org/). The results were then inputed into Cytoscape software (v3.9.1, La Jolla, CA, USA), and the molecular complex detection (MCODE) algorithm was used to select the module with the highest score to identify the key proteins that interact with SKA3. We conducted Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis on these key proteins to further explore the possible biological functions of SKA3.
SKA3 was put into Quick Search module through cBioPortal (https://www.cbioportal.org/). The “Cancer Types Summary” module was employed for illustrating mutation type alteration frequency of SKA3 across various cancer types. The “Mutation” module could play a part in screening and displaying the mutation sites of SKA3 in endometrial cancer and the exon regions to which the mutation sites belong.
All cell lines except for endometrial epithelial cells (hEEC) (Purchased from Shenzhen Haodi Huatuo Biotechnology Co., Ltd., catalog number: HTX2443, Shenzhen, Guangdong, China) were supplied by American Type Culture Collection (ATCC), including 3 endometrial cancer cells: HEC-1A (catalog number: HTB-112), RL95-2 (catalog number: CRL-1671), and AN3CA (catalog number: HTB-111). hEEC and RL95-2 cells were cultured in RPMI-1640 medium (C0893, Beyotime, Shanghai, China) with 10% fetal bovine serum at 5% CO2 and 37 °C. HEC-1A and AN3CA were cultured in Dulbecco’s modified Eagle medium (DMEM) medium (C0891, Beyotime, Shanghai, China) under the same conditions. The cells used in this study have been verified by short tandem repeat (STR) and confirmed to be free of mycoplasma contamination.
Cells were transfected with SKA3-targeting siRNAs or a negative control (NC) at 37 °C in a humidified incubator with 5% CO2 using Lipofectamine® 2000 (Lipo2000) (11668030, Thermo Fisher Scientific, Waltham, MA, USA). The siRNAs were provided (Shanghai Sangon Biotech., Shanghai, China) and confirmed to be SKA3-specific sequences after excluding homology with other genes. Two cell types were transfected with SKA3-specific siRNA-1 and siRNA-2, while a separate group was transfected with a non-specific siRNA labeled with fluorescein amidite (FAM) serving as the NC. The sequence for siRNA-1 is GGAAGAGCCCGUAAUUGUA, and for siRNA-2 is AAUCCAGGCUCAAUGAUAA. HEC-1A cells were transfected when they reached 80% confluence in a 100 mm dish by combining 20 µL of siRNA with 10 µL of Lipo2000 in 10 mL of DMEM medium (11965092, Thermo Fisher Scientific, Waltham, MA, USA) without antibiotics. Subsequently, cell morphology and transfection efficiency were assessed 6 hours post-transfection. Transfections were carried out in triplicate, with each experiment replicated at least 3 times concurrently.
The cells were lysed for subsequent western blot analysis after being harvested
by trypsinization and washed with phosphate buffered saline (PBS). The proteins were obtained by
centrifugation at 24,080
HEC-1A (siRNA-NC) and HEC-1A (siRNA-1) cells of 1000 cells/well were plated into the 96-well plate. To assess the impact of SKA3 on viability, cells were exposed to DMEM supplemented with 10% fetal bovine serum (FBS) for 1 day and 2 days. Following 1 hour incubation with 10 µL Cell Counting Kit-8 (CCK8) solution (C0037, Beyotime, Shanghai, China) at 37 °C, the absorbance was determined by a microplate reader.
First, lines were drawn on the back of the culture dish using a marker. We added
5
Statistical analysis was carried out through the SPSS 22.0 (IBM Corp., Chicago,
IL, USA) and GraphPad Prism 9.0.0 (Dotmatics, Boston, MA, USA). Among them,
quantitative data is presented as mean
We conducted WGCNA analysis based on the gene expression matrix of endometrial cancer to build a co-expression network and detect co-expression modules. Hierarchical clustering confirmed the absence of any outliers. Using the topological matrix constructed from all samples, we established a gene scale-free network with a soft threshold of 5 and a scale independence of 0.9 (Fig. 1A,B). We then applied a dynamic tree cutting algorithm to calculate the feature values of each module, forming a module feature value tree and identifying 9 strongly correlated gene modules (Fig. 1C,D). For further analysis, we calculated the correlation between each color module and endometrial cancer. The MEyellow and MEbrown modules, which showed the highest correlation with endometrial cancer, were selected for further analysis, with SKA3 located in the upregulated module (Fig. 1E).
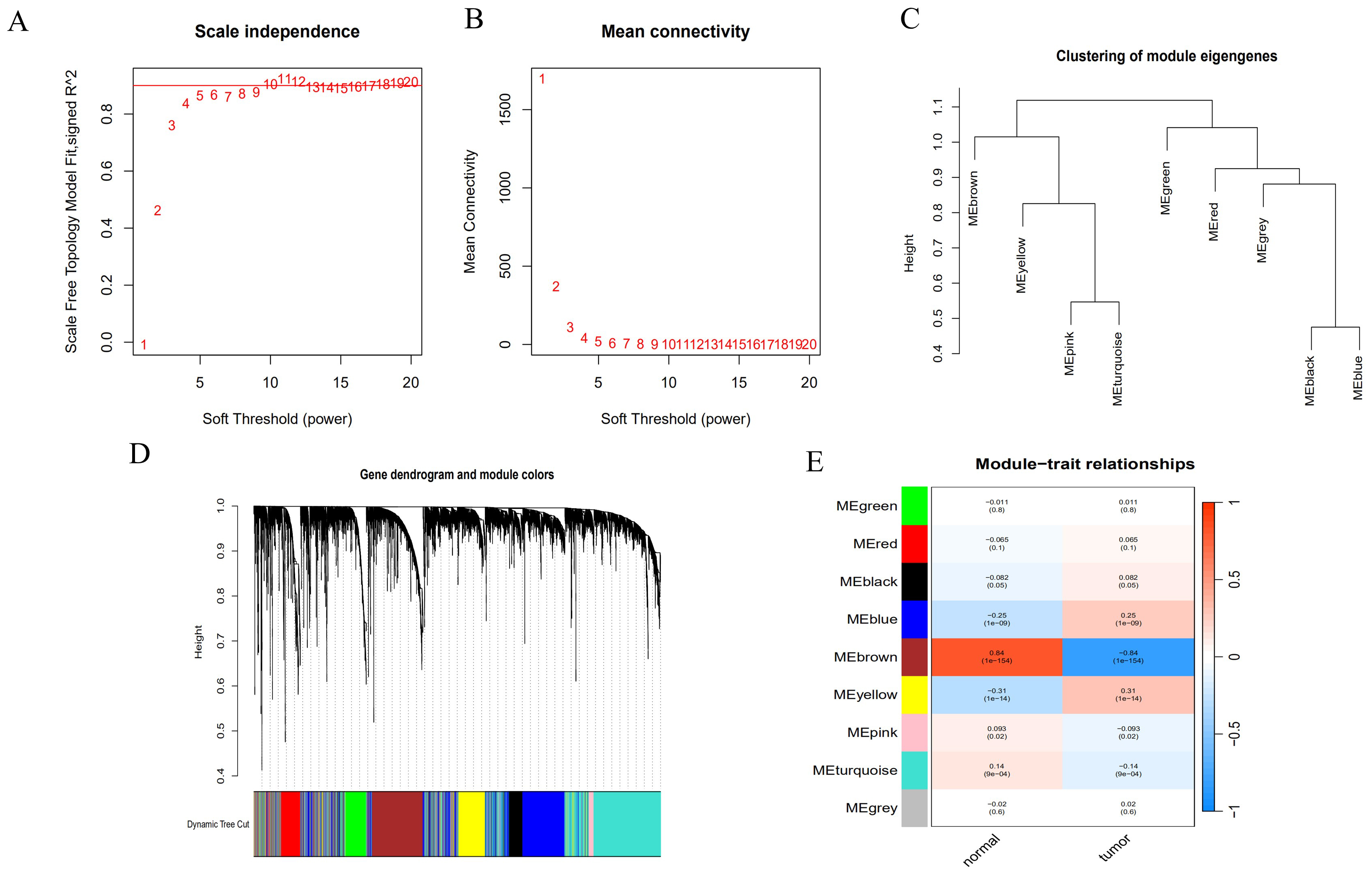 Fig. 1.
Fig. 1.
Gene modules related to endometrial cancer based on weighted gene co-expression network analysis (WGCNA) analysis. (A,B) The gene scale-free network. (C,D) The module feature value tree. (E) The module-trait relationships.
Using the MEyellow and MEbrown module genes obtained through WGCNA screening, we
applied the edgeR package for differential analysis of key module genes. A
heatmap was utilized to display the differential expression of genes in
endometrial cancer (Fig. 2A). Volcano plots were then utilized to visualize the
high and low expression genes, revealing significant upregulation of the
SKA3 gene (Fig. 2B). To demonstrate the expression of SKA3 in
endometrial cancer, we conducted a Wilcoxon rank sum test and presented the
results with a box plot, showing that SKA3’s expression in endometrial cancer was
extremely elevated contrasted to adjacent tissues (Fig. 2C). For the sake of
ensuring the reliability of our results, we also examined SKA3 expression in
pan-cancer, showing that SKA3 was upregulated in 21 out of 33 types of cancer,
with all p
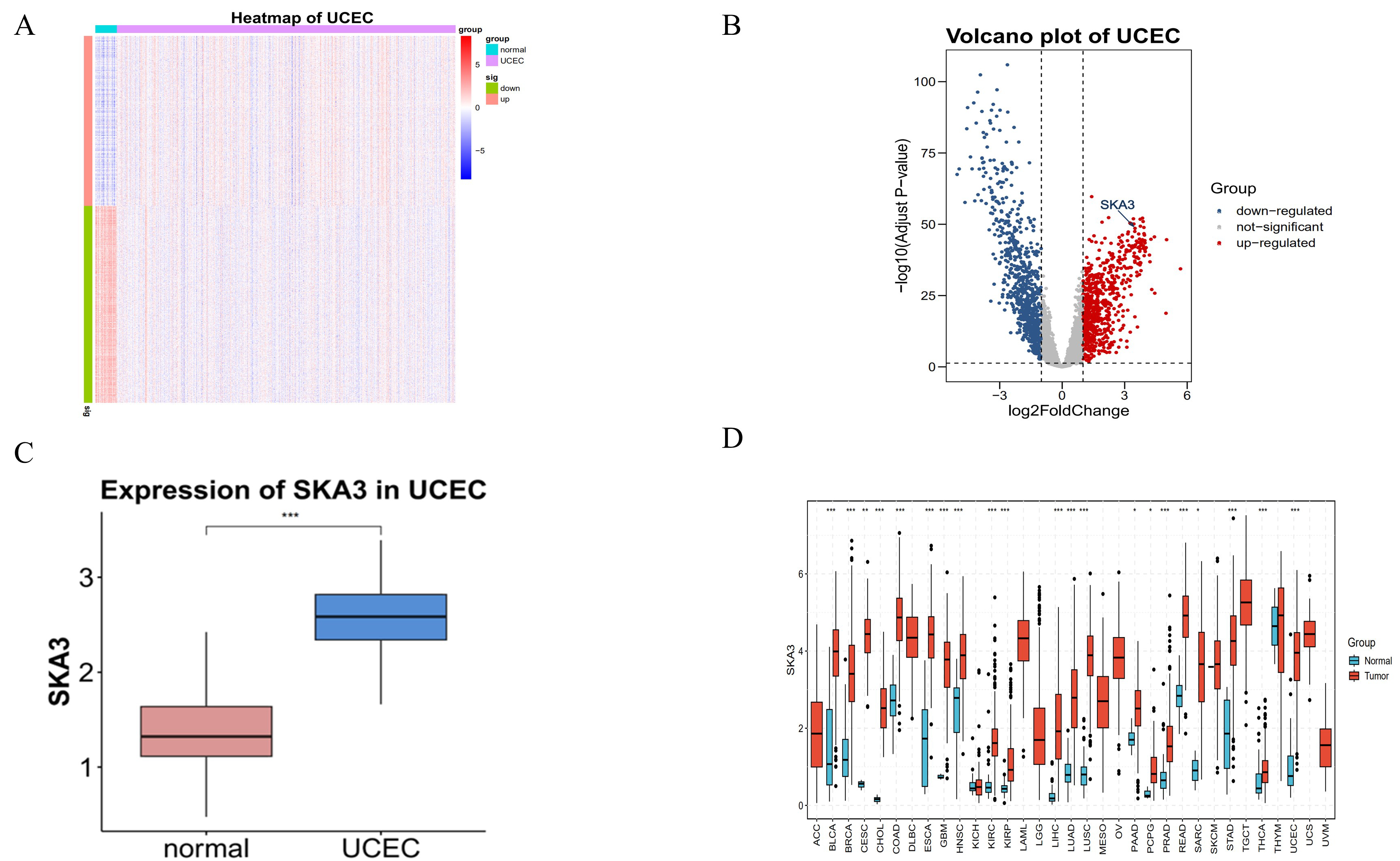 Fig. 2.
Fig. 2.
Differential expression analysis of Spindle and
kinetochore-associated complex subunit 3 (SKA3) in endometrial cancer. (A) The
heatmap in endometrial cancer. (B) The volcano plots of endometrial cancer. (C)
Box-plots of SKA3 in endometrial cancer. (D) Expression of SKA3 in Pan-Cancer.
*p
To further explore SKA3’s medical value within endometrial cancer, we analyzed the distribution both in high and low SKA3 gene expression characteristics in different diagnostic data. SKA3 expression showed significant differences among endometrial cancer patients based on age, tumor stage, and race (Table 1). Following this, the diagnostic significance of SKA3 was evaluated in individuals diagnosed with endometrial cancer. The results revealed an area under the curve (AUC) of 0.943, with a cutoff value of 1.819, specificity of 0.829, and sensitivity of 0.969 (Fig. 3A). Thus, SKA3 demonstrated strong diagnostic value for endometrial cancer. We also evaluated its prognostic value through the KM curve. The findings revealed that individuals exhibiting high SKA3 expression had a poorer prognosis with a statistically significant difference (Fig. 3B). This suggests the higher SKA3 expression levels correlate with lower survival probability, shorter survival periods, and faster tumor progression.
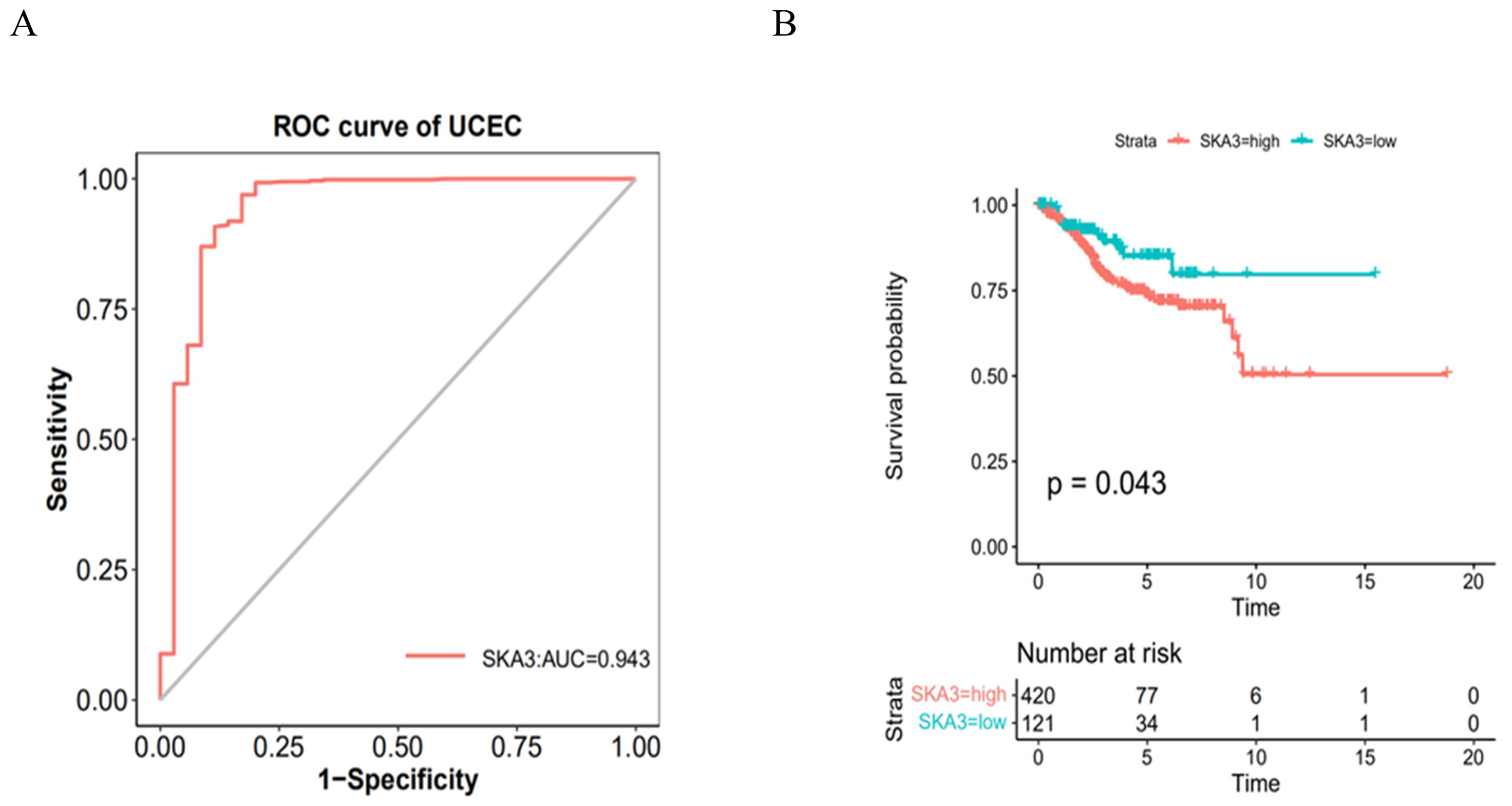 Fig. 3.
Fig. 3.
Evaluation of diagnostic and prognostic value of SKA3 in patients with endometrial cancer. (A) Diagnostic value of SKA3 in patients with endometrial cancer. (B) Survival analysis of SKA3 in patients with endometrial cancer. ROC, receiver operating characteristic; UCEC, uterine corpus endometrial carcinoma; AUC, area under the curve.
| Level | Overall | High | Low | p | ||
| n | 541 | 420 | 121 | |||
| Age (%) | 179 (33.1) | 128 (30.5) | 51 (42.1) | 5.781 | 0.016 | |
| 362 (66.9) | 292 (69.5) | 70 (57.9) | ||||
| Stage (%) | Stage I | 337 (62.3) | 244 (58.1) | 93 (76.9) | 14.355 | 0.002 |
| Stage II | 52 (9.6) | 44 (10.5) | 8 (6.6) | |||
| Stage III | 123 (22.7) | 106 (25.2) | 17 (14.0) | |||
| Stage IV | 29 (5.4) | 26 (6.2) | 3 (2.5) | |||
| Race (%) | Asian | 52 (9.6) | 42 (10.0) | 10 (8.3) | 11.473 | 0.003 |
| Others | 120 (22.2) | 106 (25.2) | 14 (11.6) | |||
| White | 369 (68.2) | 272 (64.8) | 97 (80.2) |
In order to elucidate the SKA3’s potential mechanisms in endometrial cancer, Spearman correlation analysis was carried out to identify SKA3-related genes in the key modules, highlighting 10 genes possessing positive and negative correlations (Fig. 4A, Table 2). Next, we examined the relationship between SKA3and its related genes to obtain a PPI network of SKA3 and 20 related genes. We found that SKA3 directly interacts with 10 genes, indirectly with 3 genes, and has no correlation with 7 genes (Fig. 4B). We then imported the proteins that directly and indirectly interact with SKA3 into Cytoscape, applied the MCODE algorithm, and identified the module with the highest score, which included 9 genes that directly interact with SKA3 (Fig. 4C). To further explore its possible pathways, GO as well as KEGG enrichment analyses were then implemented. The findings suggested that SKA3 was likely implicated in the mitotic pathway in the cell cycle of endometrial cancer (Fig. 4D,E).
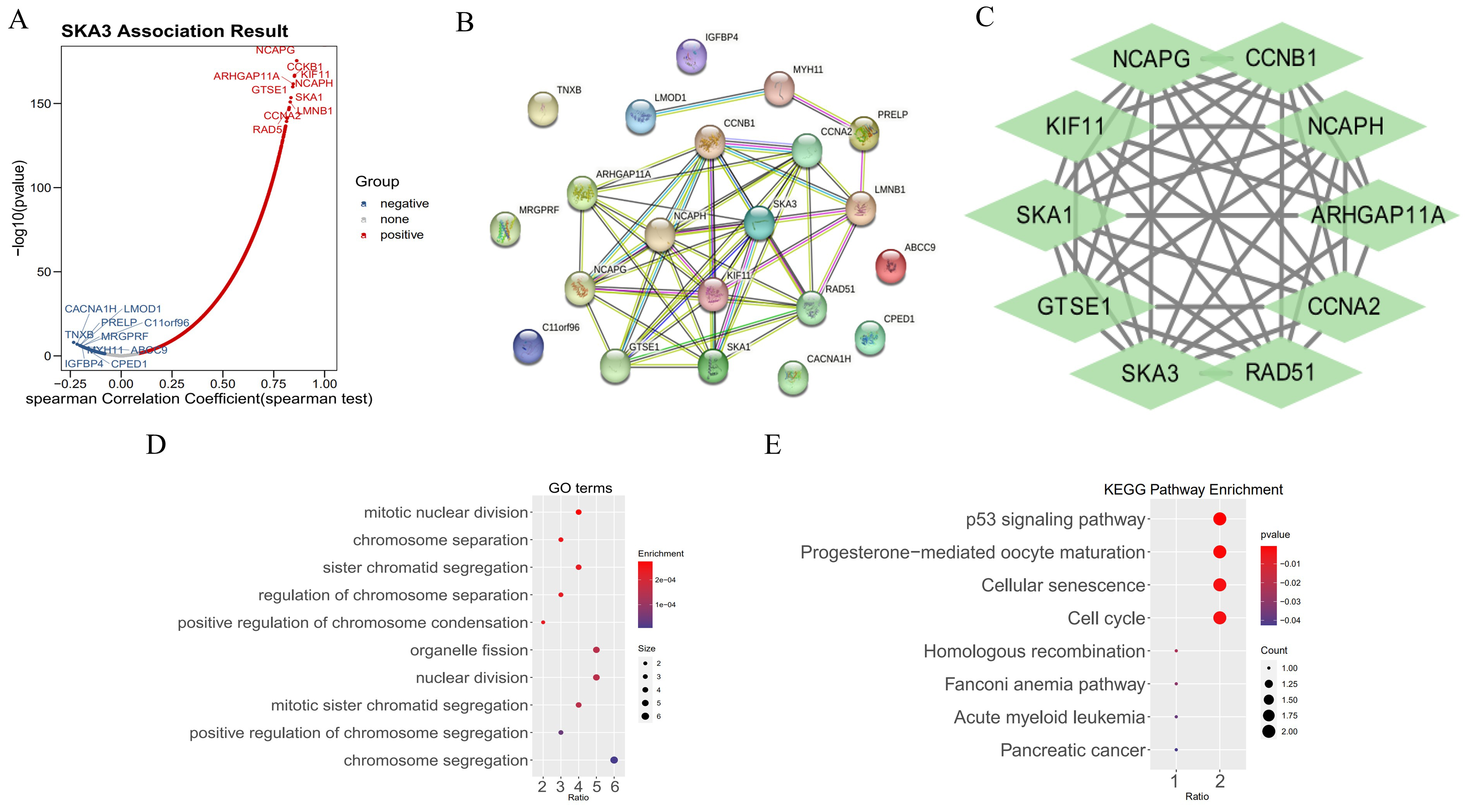 Fig. 4.
Fig. 4.
Related genes and possible pathways of SKA3 in endometrial cancer. (A) Genes related to SKA3 in the key modules. (B) The PPI network of SKA3 and 20 related genes. (C) Nine genes that directly interact with SKA3. (D) Gene Ontology (GO) enrichment analyses of SKA3 in endometrial cancer. (E) Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses of SKA3 in endometrial cancer.
| Gene | Correlation | p | Group |
| CCNB1 | 0.86295 | 3.62 |
positive |
| NCAPG | 0.862937 | 3.72 |
positive |
| KIF11 | 0.852535 | 1.54 |
positive |
| NCAPH | 0.851683 | 7.28 |
positive |
| ARHGAP11A | 0.845896 | 2.21 |
positive |
| GTSE1 | 0.843549 | 1.29 |
positive |
| SKA1 | 0.834683 | 3.36 |
positive |
| LMNB1 | 0.830989 | 1.23 |
positive |
| RAD51 | 0.826131 | 2.32 |
positive |
| CCNA2 | 0.825942 | 3.09 |
positive |
| CPED1 | –0.18925 | 3.74 |
negative |
| MYH11 | –0.19067 | 3.16 |
negative |
| ABCC9 | –0.19152 | 2.85 |
negative |
| IGFBP4 | –0.1921 | 2.65 |
negative |
| MRGPRF | –0.19335 | 2.28 |
negative |
| C11orf96 | –0.1952 | 1.81 |
negative |
| LMOD1 | –0.20266 | 7.07 |
negative |
| PRELP | –0.20317 | 6.62 |
negative |
| CACNA1H | –0.21647 | 1.12 |
negative |
| TNXB | –0.2337 | 9.51 |
negative |
We investigated the genetic changes of SKA3 across various cancers and found that SKA3 ranks second in terms of mutation frequency in endometrial cancer, with a frequency close to 5%, second only to colorectal cancer. The changes in SKA3 in endometrial cancer are primarily mutations (Fig. 5A). Therefore, we further examined the specific mutation sites in SKA3. Our findings revealed that SKA3 has 26 mutation sites in endometrial cancer, distributed across 7 different exon regions. The mutations included splice mutations, missense mutations, frameshift deletion mutations, and nonsense mutations. Notably, one of these mutation sites can result in 2 different amino acid changes (Fig. 5B) (Table 3).
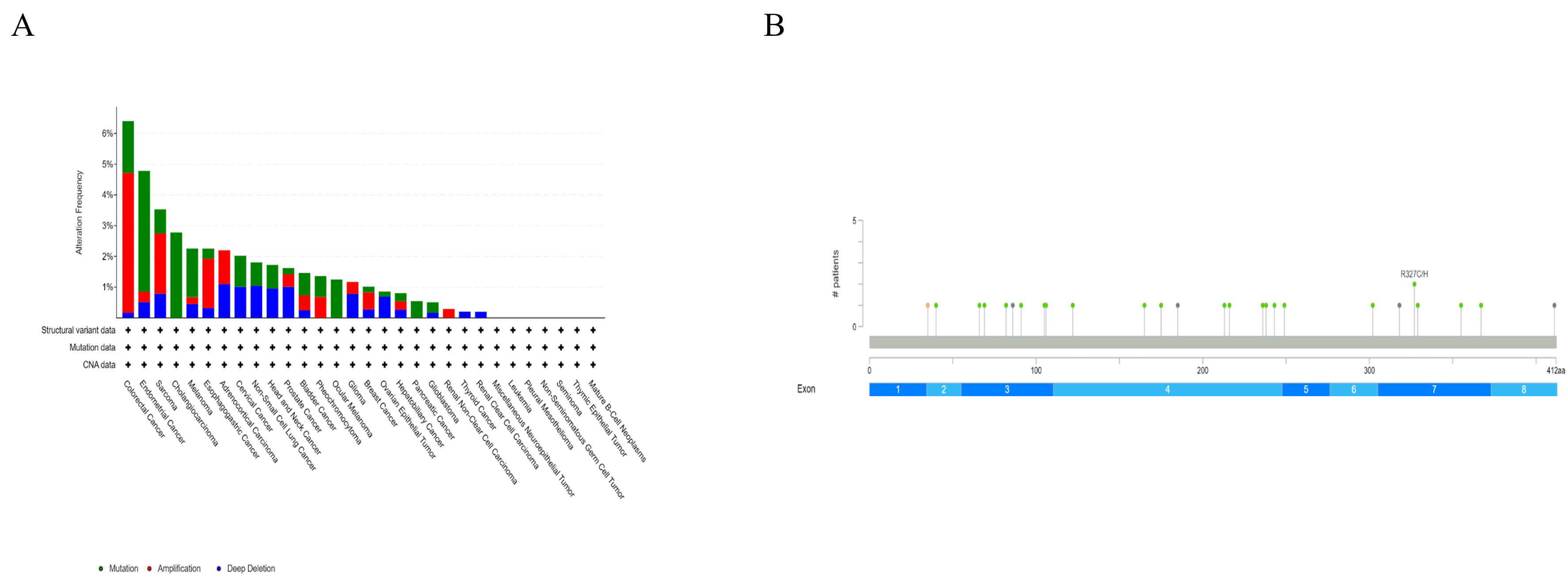 Fig. 5.
Fig. 5.
Mutation of SKA3 in endometrial cancer. (A) Mutation frequency of SKA3 in endometrial cancer. (B) Mutation sites of SKA3 in endometrial cancer.
| Sample ID | Protein Change | Mutation Type | EXON | Nonsynonymous Mutations |
| TCGA-B5-A3FA-01 | X35_splice | Splice | 2 | 9645 |
| TCGA-A5-A0G2-01 | P40S | Missense | 2 | 25696 |
| TCGA-BS-A0UV-01 | R66I | Missense | 3 | 8969 |
| TCGA-B5-A0JY-01 | N69T | Missense | 3 | 9685 |
| TCGA-A5-A1OF-01 | L82I | Missense | 3 | 10487 |
| TCGA-B5-A3FC-01 | N86Ifs*6 | FS del | 3 | 12205 |
| TCGA-B5-A11N-01 | M91I | Missense | 3 | 1909 |
| TCGA-AP-A051-01 | R105C | Missense | 3 | 8311 |
| TCGA-AX-A05Z-01 | V106A | Missense | 3 | 7370 |
| TCGA-D1-A160-01 | E122G | Missense | 4 | 499 |
| TCGA-A5-A0VQ-01 | R165Q | Missense | 4 | 366 |
| TCGA-A5-A0G2-01 | P175H | Missense | 4 | 25696 |
| TCGA-EO-A3KX-01 | E185* | Nonsense | 4 | 3200 |
| TCGA-DF-A2KU-01 | F213S | Missense | 4 | 10042 |
| TCGA-B5-A11E-01 | V216A | Missense | 4 | 9038 |
| TCGA-A5-A0G2-01 | Y236C | Missense | 4 | 25696 |
| TCGA-AP-A1E1-01 | M238V | Missense | 4 | 551 |
| TCGA-BS-A0UV-01 | A243V | Missense | 4 | 8969 |
| TCGA-AP-A059-01 | E249D | Missense | 5 | 10942 |
| TCGA-AX-A063-01 | S302N | Missense | 6 | 1000 |
| TCGA-EO-A22X-01 | S318* | Nonsense | 7 | 9421 |
| TCGA-AX-A1CE-01 | R327H | Missense | 7 | 11440 |
| TCGA-EO-A22U-01 | R327C | Missense | 7 | 13820 |
| TCGA-EY-A1GI-01 | S329L | Missense | 7 | 4487 |
| TCGA-EO-A22R-01 | L355M | Missense | 7 | 12770 |
| TCGA-BS-A0UV-01 | I367S | Missense | 7 | 8969 |
| TCGA-EO-A22U-01 | E411* | Nonsense | 8 | 13820 |
EXON, extron; FS, frame shift. * represents a termination codon mutation.
For further investigating the SKA3’s function in endometrial cancer cells, we
measured its expression in HEC-1A, RL95-2, and AN3CA, as well as its expression
in endometrial epithelial cell lines (hEEC). These findings revealed that
relative to hEEC, SKA3 was expressed in the highest levels in HEC-1A with a
statistically significant difference (Fig. 6A). Thus, the HEC-1A was chosen for
subsequent experiments. We constructed 2 HEC-1A cell lines with silenced SKA3
using siRNA, namely HEC-1A (siRNA-1) and HEC-1A (siRNA-2). The results indicated
that the silencing efficiency of HEC-1A (siRNA-1) cell lines was higher with
p
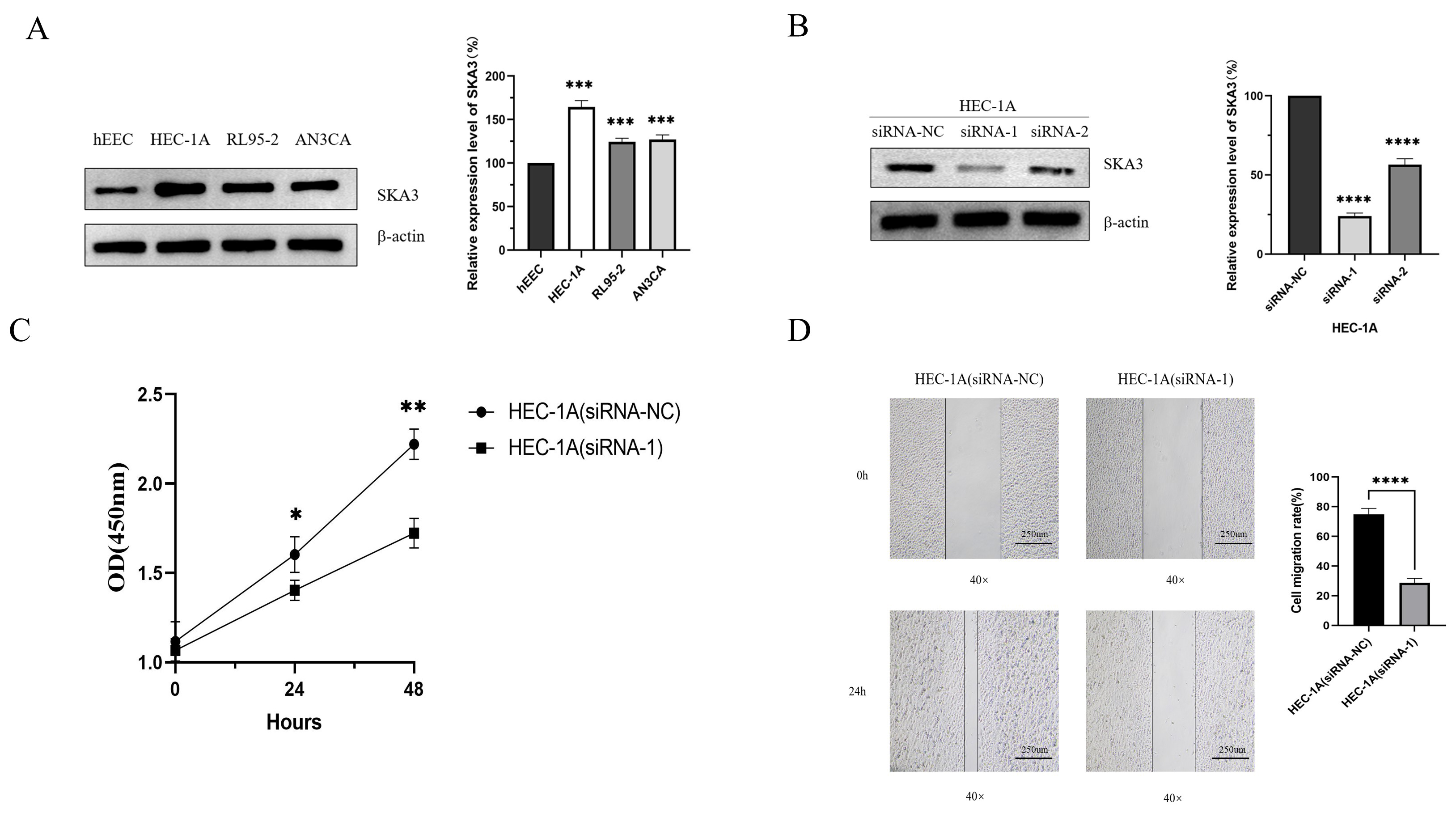 Fig. 6.
Fig. 6.
SKA3 promotes proliferation and migration of
endometrial cancer. (A) Expression of SKA3 in endometrial cancer cell lines and
endometrial epithelial cell lines. (B) Silencing efficiency of SKA3 in HEC-1A
(siRNA-1) and HEC-1A (siRNA-2) cells. (C) Effect of silencing SKA3 on the
proliferation of endometrial cancer HEC-1A cells. (D) Effect of silencing SKA3 on
the migration of endometrial cancer HEC-1A cells. The specific length of the
scale bar is 250 um. *, p
The pathogenesis and progression of tumors are closely linked to changes in organelles during the cell cycle, particularly those like the spindle and kinetochore that are directly involved in chromosome separation [24, 25]. SKA3 is a microtubule-binding subunit of exosomes, occupying an important position in ensuring accurate chromosome separation and cell cycle division [26, 27]. Research has shown that SKA3 holds a significant state in developing and progressing multiple types of tumors, like bladder cancer [12], gastric cancer [19, 28], prostate cancer [8, 29], and lung cancer [26, 30, 31]. Recently, SKA3 has been identified to hold a more prominent state, especially in gynecological tumors. In a study by Hu et al. [27], it was discovered that SKA3 enhances cervical cancer cells proliferation and migration by activating the PI3K/Akt signaling pathway, potentially offering potential new targets in cervical cancer therapy. Zhong et al. [13] discovered that elevated SKA3 mRNA levels are observed in cancer tissues, and this high expression is linked to a poor prognosis in early-stage female patients. Furthermore, SKA3 overexpression could potentially serve as an independent risk factor in the female group with early breast cancer, indicating its potential in acting as a biomarker. Ruan et al. [15] found that SKA3 can regulate the degradation of polo-like kinase-1 (PLK-1), controlling the growth as well as proliferation of breast cancers. Sheng and Wang [32] found that extracellular vesicle circ-SKA3 mediates cellular communication between ovarian cancer cells, suggesting a promising target for therapeutic interventions in cervical cancer. However, SKA3’s expression and clinical value have no current reports. This study represents a pioneering investigation into its role in endometrial cancer, aiming to fill the research gap and improve the understanding of SKA3 in gynecological tumors.
This study utilized WGCNA and differential analysis to identify SKA3 as a key gene in endometrial cancer from the TCGA database, establishing a foundation for further research. This study revealed the SKA3 exhibits high expression levels in endometrial cancer. We also detected SKA3 expression in 3 endometrial cancer cell types and found elevated levels of SKA3 in all 3, consistent with the results of the TCGA database. For exploring its widespread expression within various cancers, we examined its expression in 33 common cancers and found the SKA3 exhibits high expression levels in 21 types, suggesting that SKA3 may act as an oncogene in many cancers. This aligns with findings by Feng et al. [10], who reported that SKA3 is more upregulated in tumor samples than in normal samples in many cancers and plays an oncogenic role. To further investigate the clinical value of SKA3 in endometrial cancer, we evaluated its diagnostic and prognostic significance using ROC curves and KM survival analysis. The diagnostic value of SKA3 for endometrial cancer was found to be 0.943, indicating a high diagnostic potential. Additionally, patients with high SKA3 expression experienced a shorter prognosis and survival duration compared to the patients with low SKA3 expression, with the difference being statistically significant. Therefore, SKA3 has substantial diagnostic and prognostic value for endometrial cancer. In gynecological tumors, elevated SKA3 expression is also observed in cervical and breast cancer, promoting their proliferation and migration [13, 14, 15, 27]. Patients with breast cancer and high SKA3 expression have a poor prognosis [13]. Next, we explored the possible mechanisms by which SKA3 affects endometrial cancer. We initially screened the 10 genes most positively and negatively correlated with SKA3 in endometrial cancer. We found that SKA3 directly interacts with 10 genes, indirectly with 3 genes, and has no correlation with 7 genes. To further identify the interacting genes of SKA3, we used the MCODE algorithm to obtain the highest-scoring module, which included SKA3 and 9 directly interacting genes. We discovered that SKA3 may be associated with mitosis-related pathways in the cell cycle of endometrial cancer. Next, we explored the genetic changes of SKA3 in endometrial cancer. Among common cancers, SKA3 ranks second in overall mutation frequency in endometrial cancer, close to 5%, second only to colorectal cancer. The changes in SKA3 in endometrial cancer are primarily due to gene mutations. Furthermore, we analyzed its mutation sites and identified 26 mutation sites distributed across 7 different exon regions. These mutations included splice site mutations, missense mutations, frameshift deletion mutations, and nonsense mutations. One of the mutation sites can result in two different amino acids, indicating the complexity of SKA3 genetic changes in endometrial cancer. Finally, we explored the biological functions of SKA3 at the cellular level. Compared to endometrial epithelial cells (hEEC), SKA3 expression was increased in all 3 types of endometrial cancer cells. In all of them, HEC-1A showed the highest expression level. Therefore, we used HEC-1A cells for further exploration. We constructed 2 HEC-1A cell lines with silenced SKA3 expression using siRNA interference technology, and the results demonstrated that the HEC-1A (siRNA-1) cell line had higher silencing efficiency. Subsequently, we investigated its biological effects in proliferation and migration within HEC-1A lines. Silencing SKA3 expression resulted in a deceleration in all these cells, suggesting that SKA3 was likely to be an oncogene in endometrial cancer and influence its development.
However, there are still some limitations to this study. This study focuses on statistical analysis of samples in the TCGA database, lacking validation of clinical samples. In addition, this study did not delve into the pathway of SKA3 in endometrial cancer. In the future, we will collect a large number of clinical samples for validation, making the conclusions of this study more scientific and credible. And we will continue to explore the pathway of SKA3 in endometrial cancer to fill the gap in the mechanism of SKA3 in endometrial cancer.
In conclusion, SKA3 is a key gene in endometrial cancer, with significant diagnostic and prognostic value, and may also impact the development of endometrial cancer.
The data supporting the conclusions of this research are available upon request from the corresponding author.
XD and JZ were responsible for the conception of the study; XD, YueZ, JQ and YuZ were responsible for bioinformatics analysis and basic experiment; XD and YuZ wrote the original draft; XD reviewed and revised the final manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
We would like to express our gratitude to all those who helped us during the writing of this manuscript. Thanks to all the peer reviewers for their opinions and suggestions.
This research was supported by the Maternal and Child Scientific Research Project of Wuxi City (Grant No. FYKY202301).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.






