1 Lee Kong Chian School of Medicine, Nanyang Technological University, 308232 Singapore, Singapore
2 Collaborative Institute, Interdisciplinary Graduate Programme, Nanyang Technological University, 639798 Singapore, Singapore
3 Department of Respiratory and Critical Care Medicine, Singapore General Hospital, 169608 Singapore, Singapore
4 Duke-NUS Medical School, 169857 Singapore, Singapore
5 Department of Respiratory and Critical Care Medicine, Tan Tock Seng Hospital, 308433 Singapore, Singapore
Abstract
Chronic obstructive pulmonary disease (COPD) is a progressive and debilitating respiratory condition marked by chronic symptoms and frequent exacerbations, contributing to significant morbidity and mortality. The advent of molecular microbiology and next-generation sequencing (NGS) has expanded our understanding of the lung microbiome, and integration of microbiome datasets with other omics reveals important microbial-metabolic-immuno-inflammatory interactions that influence COPD pathogenesis. Recent studies have highlighted dysbiosis of the airway microbiome, with shifts in bacterial, viral, and fungal communities playing a crucial role in disease progression, exacerbations and clinical outcomes. Moreover, microbiome changes are observed in COPD associated overlap syndromes, complicating diagnosis and treatment. This review synthesizes current microbiome research in COPD, focusing on its clinical relevance, including its potential as a diagnostic and prognostic tool. We additionally discuss the challenges of integrating microbiome data into clinical practice, emphasizing the need for personalized, precision medicine approaches to optimize COPD management and improve patient outcomes.
Keywords
- COPD
- microbiome
- metagenomic
- mycobiome
The term ‘microbiome’ refers to the community of all microorganisms, including bacteria, archaea, fungi and viruses, present in a given environment, such as the various ecological niches of the human body [1]. Compared to other major organ systems, lung microbiome research lags significantly due to previously misinformed assumption of lung sterility resulting in its unintended exclusion from the NIH Human Microbiome Project [2, 3, 4, 5]. Nonetheless, research demonstrating the key roles of the lung microbiome in health and disease has emerged steadily over the past decade [6, 7, 8]. This is coupled with the advancements of sequencing technologies to interrogate the various microbial communities. The use of next-generation sequencing (NGS) technologies to assess microbiomes involves amplifying genomic material to profile microbial communities and includes ‘targeted’ amplicon sequencing and ‘untargeted’ shotgun metagenomics [9, 10]. Targeted amplicon sequencing amplifies specific marker genes that serve as universal taxonomic barcodes and include the 16S ribosomal RNA (rRNA) gene for bacterial microbiomes [11], and the internal transcribed spacer (i.e., 18S/ITS) regions for the fungal microbiome (mycobiome) [12, 13]. Shotgun metagenomic sequencing amplifies all microbial genomes present in a given sample [14]. However, it has limited utility in profiling active microbial gene expression for which metatranscriptomic approaches are required [15]. Meanwhile, viromics is an emerging field for the characterization of viromes that entails further development of specific bioinformatic tools and improvement in resourcing viral sequence database annotations [15].
NGS identifies culturable and importantly, unculturable microbes, allowing for a holistic and comprehensive evaluation of lung microbiome composition and diversity [10, 16]. Importantly, NGS requires specialized experimental and bioinformatic capabilities and a lack of gold-standard analytical pipelines has restricted widespread clinical translation [13, 17, 18]. Emerging evidence does support the clinical potential of NGS in clinical settings. In critical care and intensive care unit (ICU) settings, metagenomics have already demonstrated significantly improved sensitivity and clinical outcomes against routine, gold standard clinical tests for the diagnosis of infection in various sample types including blood, sputum, broncho-alveolar lavage (BAL) and cerebrospinal fluid [19, 20, 21, 22, 23, 24, 25, 26]. Most recently, metagenomics demonstrates strong potential for informing antimicrobial treatment and resistance, infection control that can potentially be leveraged for wider public health benefit [26, 27, 28, 29].
A healthy lung microbiome is rich and diverse, characterized by the presence of Prevotella, Streptococcus, Veillonella, Fusobacterium and Haemophilus spp. [8, 30, 31, 32, 33]. The lung microbiome is highly dynamic and largely determined by microbial immigration from the upper respiratory tract and elimination through host immunity and mucociliary clearance [7, 34, 35]. Emerging evidence suggest that the lung microbiome is enriched with oral taxa that confer a protective immunological tone against potential pathogens [36, 37]. Work investigating mycobiome in health have identified Ascomycota and Basidiomycota to be the most commonly detected fungal phyla, with Candida, Saccharomyces, Penicillium, Cladosporium, and Fusarium as the main genera [13, 30, 38, 39, 40, 41, 42]. Lung viromes remain poorly characterized, with small studies proposing a presence of Anelloviridae [43, 44]. In contrast to the healthy state, structural abnormalities in any diseased lung alters the local environment, favouring pathogen growth [34, 45] and microbial dysbiosis that is now described across almost every respiratory disease, including asthma, chronic obstructive pulmonary disease (COPD), interstitial lung disease and bronchiectasis [13, 16, 38, 46, 47, 48, 49].
While other respiratory conditions present with unique challenges and pathophysiological features, COPD remains a leading respiratory cause of global morbidity and mortality accounting for a significant burden of healthcare utilization including hospitalization [50, 51]. The heterogeneity of COPD and its progression through stable, exacerbation states and overlap syndromes further provides a rich framework for studying the role of the respiratory microbiome. As such, this review aims to provide an overview of the lung microbiome in COPD during disease stability and exacerbations and additionally address the emerging clinical entities of COPD with asthma and bronchiectasis overlap.
COPD is a multifaceted progressive lung disorder characterized by persistent respiratory symptoms and airflow limitation caused by abnormalities in the airway or alveoli [52, 53]. It is the result of lifelong and complex interactions between host, genetics and environmental exposures. While tobacco smoking is a major risk factor, it is now recognized that nearly half of the COPD cases globally occur in non-smokers, particularly in low to middle income countries [54]. The recent Global Initiative for Chronic Obstructive Lung Disease (GOLD) statement categorizes COPD into six different etiotypes (genetically determined COPD, COPD due to abnormal lung development, environmental COPD, COPD due to infections, COPD and asthma and unknown cause), emphasizing the significance of non-smoking related risk factors in disease development and progression [52].
Studies have shown that the airway microbiome, consisting of diverse microbial communities in the respiratory tract, plays a crucial role in maintaining lung homeostasis and influencing COPD outcomes [40, 55]. Exploring the microbiome in stable COPD may enhance our understanding of disease mechanisms, identify biomarkers for patient stratification, and uncover therapeutic targets to improve long-term disease management and clinical outcomes. These concepts are summarized in Fig. 1 (Ref. [56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66]).
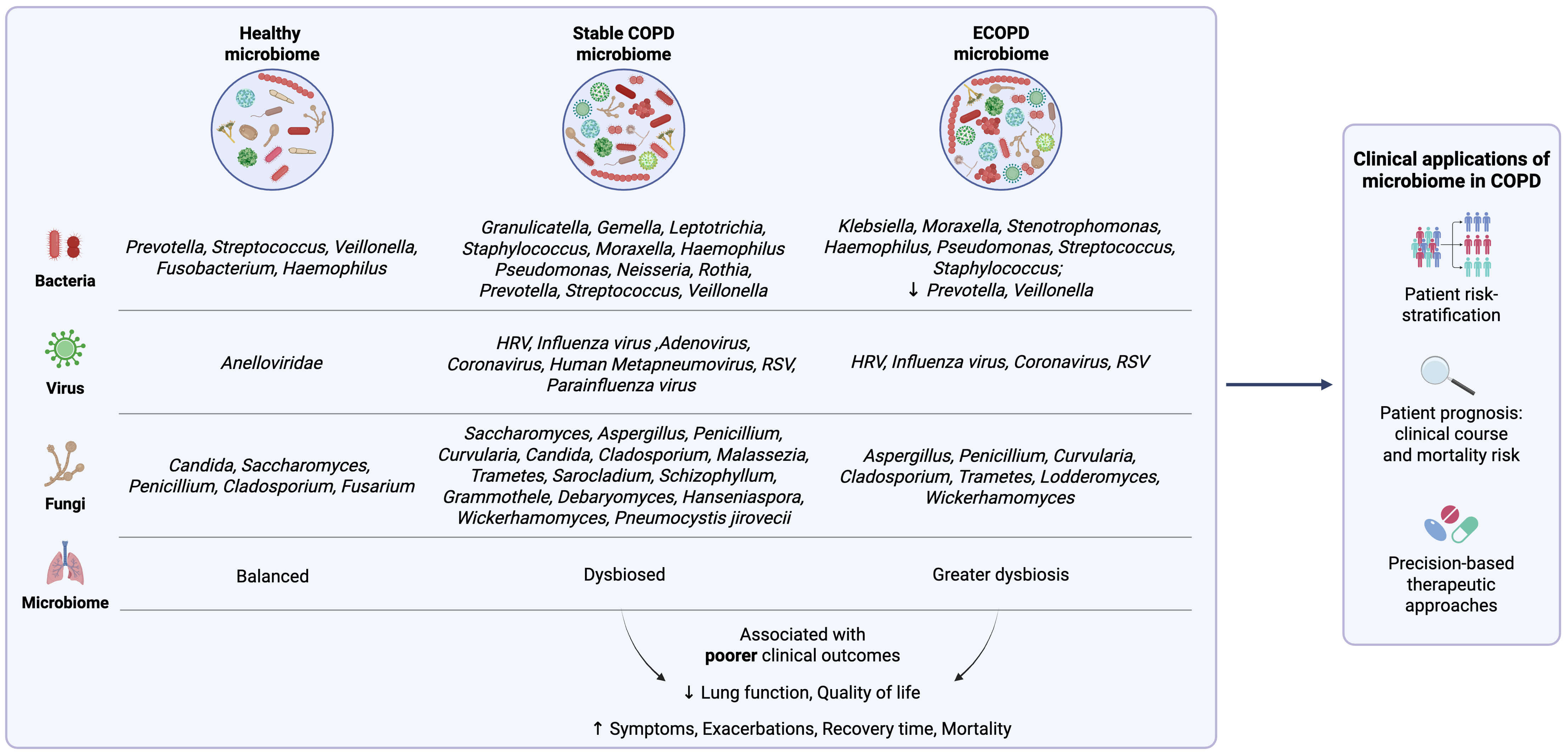 Fig. 1.
Fig. 1.
Summary of the lung microbiome profile in the healthy state, in
stable chronic obstructive pulmonary disease (COPD) and during exacerbations of COPD (ECOPD) with associated clinical outcomes and potential applications. HRV, human
rhinovirus; RSV, respiratory syncytial virus;
In patients with mild-moderate COPD (GOLD stage 1 or 2), a reduced airway microbial diversity is observed with an increased abundance of Granulicatella, Gemella, Leptotrichia, Streptococcus, Prevotella, Staphylococcus, Neisseria, Veillonella and Rothia [60, 61, 67, 68]. Notably, Streptococcus, Prevotella and Veillonella were found to significantly co-occur, and strongly associates with a concurrent change in host transcriptomic profile, including the upregulation of several transcriptional pathways that are important for mucosal immune response towards microbes [67]. Similar pro-inflammatory responses associated with microbiome dysbiosis have been reported, alongside a suppression of genes involved in epithelial repair [69]. Additionally, alterations in the lung bacterial composition driven by enrichment of Streptococcus, Prevotella, Staphylococcus, and Pseudomonas have been associated with key clinical parameters, including increased neutrophil percentage, greater symptom burden and functional impairment measured by six-minute walk distance [61]. While many studies have separately explored the effect of microbiota and metabolomics on COPD outcomes, few have integrated data from both [40]. Madapoosi et al. [60] noted that several lung bacterial genera, Streptococcus, Neisseria and Veillonella, along with metabolites such as glycosphingolipids, glycerophospholipids positively associate with poorer clinical outcomes including decreased lung function and greater symptomology. Conversely, the authors also noted Prevotella bacteria, although were less abundant in severe COPD, was positively associated with the metabolites adenosine monophosphate (AMP), adenosine, spermine and 5’-methylthioadenosine (MTA), which correlate negatively with neutrophil levels and positively with regulation of airway surface liquid, important for maintaining ciliary function [60, 70]. While it remains unclear if the bacteria-metabolite correlations are direct or indirect, as these data suggest the role of both in shaping the lung environment which could potentially impact the COPD pathophysiology. Collectively, these findings suggest that early airway dysbiosis triggers a pro-inflammatory respiratory niche that impairs repair responses and drive clinical decline, underscoring the important role of the microbiome and their associated metabolites, even in stable, non-exacerbating COPD.
As COPD progresses, microbial shifts in airway bacteriome profiles portending
toward enrichment of more commonly recognized pathogens, such as
Haemophilus, Moraxella, Pseudomonas and
Staphylococcus are observed [33, 59, 71, 72]. The outgrowth of
these genera is strongly associated with COPD severity, reduced microbial
diversity and increase airway inflammation [33, 59, 72, 73]. In particular, COPD characterized
by a predominance of the Proteobacteria phylum including
Haemophilus and Moraxella often associates with neutrophilic
inflammations, poorer lung function, increased exacerbations and a higher risk of
mortality compared to those with Firmicutes dominance [74, 75, 76].
Haemophilus-dominated microbiome (
During stable COPD, viruses including human rhinovirus (HRV),
influenza, coronavirus, respiratory syncytial
virus (RSV), adenovirus, human metapneumovirus and
parainfluenza virus are consistently detected in
Stable COPD patients exhibit a distinct mycobiome, with increased abundance of Aspergillus, Candida, Cladosporium, Malassezia, Trametes, Penicillium, Sarocladium and Wickerhamomyces compared to healthy controls [65, 93]. Interestingly, these mycobiome profiles vary across geographical regions, likely influenced by local climate and environmental factors that support fungal growth and survival. For instance, cohorts from Asia (Singapore and Malaysia) show a predominance of Saccharomyces, Curvularia, Aspergillus, Schizophyllum, Penicillium and Grammothele, while European cohorts (United Kingdom) display an enrichment of Debaryomyces, Hanseniaspora, Trametes and Wickerhamomyces [65]. These findings highlight the geographical variability of the COPD-associated mycobiome and the role of environmental factors in shaping its composition. Our group further identifies the home environment as a key source of fungal exposure, particularly Aspergillus fumigatus (A. fumigatus) and its associated allergens. Elevated abundance of A. fumigatus was associated with increased risk of COPD exacerbations [94]. Additionally, a higher sensitization response was observed in COPD patients living in homes with greater abundance of A. fumigatus (Asp f 3) allergens. This in turn precipitates adverse outcomes such as lower lung function and higher exacerbation rates compared to patients with no sensitization response [95, 96, 97, 98]. Environmental parameters, such as temperature and particulate matter (PM2.5 and PM10) levels, were also positively correlated with abundance of A. fumigatus [94]. These findings underscore the complex interplay between environmental pollution, airway mycobiome and adverse clinical outcomes in COPD. Meanwhile, Pneumocystis jirovecii remains the main opportunistic species isolated in COPD patients with co-existing human immunodeficiency virus, where it is associated with increased inflammation and severe airflow obstruction [99, 100]. In stable COPD, two distinct and clinically significant ‘mycobiome clusters’ were established: a Saccharomyces-dominant cluster that associates with increased symptoms and a ‘high-risk’ cluster characterized by Aspergillus, Curvularia and Penicillium associated with increased exacerbations and higher risks of mortality [65]. There is now growing evidence highlighting the emerging and often overlooked role of environmental factors and the airway mycobiome in influencing COPD outcomes including exacerbations, disease progression and prognosis. Further investigations into these interactions could provide the foundation for novel patient stratification strategies, enabling earlier identification of ‘high-risk’ patients necessitating closer clinical monitoring.
Exacerbations are key events in the clinical course of COPD, characterized by
worsening dyspnea and/or cough with increased sputum production over a period of
Compared to stable COPD,
Beyond direct associations with clinical outcomes, bacteriome composition during
ECOPD differs between bacterial and eosinophilic exacerbation subtypes [73, 109]. Bacterial associated exacerbations are
characterised by decreased
Besides bacteria, viruses are also infectious triggers for ECOPD, accounting for at least half of all exacerbations [89, 121]. Viruses such as HRV, RSV, influenza, and coronavirus, which are present in the stable state of COPD, show a marked increase in prevalence and abundance during ECOPD, including the doubling of HRV abundance, and a 12-times surge of influenza virus [63, 87, 89, 122, 123]. Notably, the incidence of viral-positive ECOPD events were highest in winter months, potentially due to low temperatures favouring many key COPD viral pathogens including influenza virus and RSV, thereby making winter months an important period for closer monitoring of patients [84, 124]. In virus-positive ECOPD patients, elevated blood neutrophil, fibrinogen, IL-6, and C-reactive protein levels were observed, indicating increased neutrophilic inflammation levels within the host [62, 84, 125]. Recent studies further elucidate that viral infection induces higher abundance of airway NETs compared to healthy controls that in turn associated with viral loads, inflammatory and clinical measures, including increased symptomology, exacerbation severity, longer recovery and greater mortality [63, 78, 90, 121, 125].
Vaccinations, a key preventive measure against viral infections, have been
utilized to minimize the incidence of viral infections and their associated
complications [126]. While there is currently no
immunisation for the most prevalent HRV, vaccines targeting other commonly
detected viruses in COPD are available [127, 128].
Influenza vaccines have long been recommended for patients with COPD, yet the
vaccination rates vary globally with suboptimal overall coverage [129, 130]. Evidence indicates that
current-season influenza vaccines moderately reduce the risk of
influenza-associated hospitalizations, COPD exacerbations, and mortality
[129, 131, 132, 133, 134]. Notably, while
current-season vaccination offers modest efficacy, COPD patients who miss the
current season’s vaccine may retain residual protection from prior vaccinations
[133]. These findings reinforce the annual
recommendation of influenza vaccinations while highlighting the need for more
effective influenza vaccines to enhance outcomes for COPD patients. In late 2019,
the unprecedented COVID-19 pandemic (caused by coronavirus) brought about a
serious global health threat, but also significantly accelerated vaccine
innovation, with COVID-19 vaccines being developed and available within two years
of the first reported case [135]. COVID-19 vaccines are
strongly advised for COPD patients to prevent COVID-19 infection, reduce risk of
hospitalisation and mortality [128, 136, 137]. More recently, RSV vaccines have also been approved and
recommended for COPD patients
Although fungi are an uncommon trigger for traditional ECOPD, mycobiome alterations portending towards more pathogenic genera have been observed [65, 143]. The external environment is a common source of fungi, with climatic factors such as air quality and temperature influencing fungal growth and survival [94, 144]. Coupling this with impaired airway structure and defence mechanisms in COPD patients that facilitates conidial-epithelial cell binding, colonisation and persistence of pathogenic fungi may then be promoted [145, 146]. Using a broad panel of fungi allergens, Tiew et al. [98] found that frequent exacerbators of COPD have increased fungal sensitisation to crude fungal allergens, Penicillium, A. fumigatus, Curvularia and recombinant A. fumigatus. Further exploration of fungal sensitisation profiles identified two distinct clusters, one characterised by elevated sensitisation to Aspergillus-related allergens that associated with poorer lung function, worse prognosis and increased exacerbations while the second cluster had higher symptom burden that was linked to Cladosporium sensitisation [147]. Given that crude allergens (and not recombinant allergens) are routinely used in clinical practice to measure patients’ sensitisation, findings from this study thus encourage the use of a more comprehensive allergen panel that includes both crude and recombinant fungal allergens for patient screening and identification of those at high clinical risk. The sputum mycobiome study in COPD further identified increased fungal interactions and Wickerhamomyces in frequent exacerbators [65]. During ECOPD, reduced airway mycobiome diversity with greater proportions of Penicillium, Cladosporium, Trametes and Lodderomyces was associated with an increased 2-year mortality [65]. Importantly, mycobiomes appear unaffected by treatments used during ECOPD including systemic steroids and antibiotics, in contrast to that observed in bacterial microbiomes [65]. Further work assessing inter-kingdom interactions during ECOPD is required to fully delineate their emerging role in ECOPD and disease progression. Comprehensive mycobiome profiling and analysis during ECOPD is likely beneficial in prognostication and identification of patients at greater clinical risk of long-term adverse events.
Coinfection refers to concurrent presence of two or more microbial species, either from the same kingdom or from different kingdoms. In COPD, one of the most important coinfections involves bacteria and viruses. Notably, viruses can precipitate a secondary bacterial infection following the initial viral infection, leading to synergistic worsening of airway dysfunction [57, 148]. Studies have reported that up to 60% of viral infections in COPD are associated with bacterial coinfection, which is linked to microbial shifts including an increase in Proteobacteria and a decrease in Firmicutes and Bacteriodetes [63, 124, 149, 150]. At the genus level, these changes correspond to an upregulation of pathogenic species such as Haemophilus, Moraxella, Pseudomonas and a decline in Streptococcus, Veillonella and Prevotella [87, 150]. Experimental studies using HRV inoculation in COPD patients further confirmed this observations, demonstrating a sustained increase in pathogenic bacterial load 15 days post-inoculation which persisted up to day 42, even after viral loads have lowered, highlighting the prolonged effects of microbiome alterations through viral infections [150]. The mechanism underlying the bacterial overgrowth include viral induction of high sputum neutrophil elastase levels, elevated proinflammatory cytokines and reduced antimicrobial peptides, creating pro-inflammatory environment that impairs airway defence and facilitates pathogen colonisation [121, 149, 151]. Furthermore, HRV disrupts host epithelial cell functions and enhance bacterial adherence to airway epithelium, further promoting bacterial colonisation and persistence [152, 153, 154]. These pathogenic bacteria can then further promote airway inflammation, resulting in disease progression.
Apart from the well-established viral-bacterial coinfection, interactions between airway bacterial and fungi has recently been explored. Liu et al. [155] reported significant negative associations between bacteria and fungi when comparing COPD patients with healthy controls and frequent versus non-frequent exacerbators. Specifically, the authors found that bacteria Prevotella and Veillonella exhibited an inverse relationship with fungi Candida and Aspergillus [155]. These perturbations, characterised by loss of commensals and enriched pathogenic fungi, were notably associated with elevated proinflammatory mediators, such as IL-6 and IL-8. During ECOPD, Huerta et al. [156] also found an increased isolation of pathogenic bacteria in patients with Aspergillus fungal colonisation, with Pseudomonas being the most frequently isolated bacterium. Although both studies did not explore mechanistic details of the coinfection, these findings suggest that bacteria-fungal interactions play a significant role in airway inflammation potentially contributing to disease progression.
Importantly, these studies highlight the significant of inter-kingdom interactions, emphasizing how microbes from different kingdom (bacteria, fungi and virus) can interact to influence disease severity, progression and outcomes. As such, beyond conventional microbiome studies that focus on the effects of individual kingdom, approaches focusing on microbial interactions should also be employed. Such approaches may reveal key microbial relationships that will facilitate our understanding of the underlying disease mechanisms and offer clinical insight that might be overlooked by using conventional study methods [157, 158].
COPD treatments can further influence the lung microbiome profiles.
Bronchodilator therapy, comprising long-acting
Macrolides, as an alternative COPD therapeutic, serves as an add-on treatment
for individuals experiencing frequent exacerbations [52, 169]. Macrolides’ inherent anti-inflammatory and immunomodulatory
properties confer multiple beneficial airway effects including reduced bacterial
load, attenuated mucus secretion and dampened inflammation [170, 171]. Their use has been shown to decrease exacerbations
in chronic airway diseases such as COPD, asthma and bronchiectasis [170, 172, 173, 174].
Studies investigating macrolide therapy report shifts in microbial abundance and
a reduction in
A short course of systemic corticosteroid (5 days) remains the primary treatment for ECOPD and is often combined with antibiotics in cases of bacterial-associa-ted exacerbations or severe exacerbations requiring ventilatory support. Systemic corticosteroid use has been associated with reduced bacteriome diversity, and enrichment of Proteobacteria, while the reverse was observed in antibiotic therapy, with or without corticosteroid during ECOPD [73, 177].
Beyond conventional clinical treatment, other intervention approaches can target respiratory microbiomes. A rapidly evolving field in microbiome-related therapy is probiotics, that has now extended beyond its initial use in gastrointestinal diseases [178]. In COPD, KavianFar et al. [179] utilised a meta-analysis approach to reveal Lactobacillus and Bifidobacterium as two potential probiotic therapeutic candidates since both bacteria not only correlated positively and found in decreasing abundance with increasing disease severity, but also negatively correlated with Neisseria and Haemophilus, known pathogens in COPD. While this finding indicates a promising potential for probiotics in COPD, more research and clinical studies investigating the effects of probiotics on respiratory microbiome will be necessary before application. Meanwhile, phage therapy, which uses the bacterial-specific lytic action of bacteriophages (viruses), offers another promising therapeutic avenue, especially in an era of increasing antimicrobial resistance. Current evidence in COPD stems from a case report detailing provision of phage therapy with concurrent antibiotics to a hospitalised COPD patient in respiratory failure with positive culture of hospital-acquired Carapenem-resistant Acinetobacter baumannii. After a week of treatment, cultures turned negative along with improvements in lung function, indicating a beneficial response to the treatment provided [180]. Despite its potential, the field of phage therapy remains at a nascent stage clinically due to a lack of comprehensive studies and challenges, including phage-resistant bacteria, potential alterations on host microbiome and lack of regulatory oversight [181].
It is now recognized that COPD commonly ‘overlaps’ with other respiratory disease states such as asthma and bronchiectasis, resulting in asthma-COPD overlap (ACO) and bronchiectasis-COPD overlap (BCO) which contribute to the observed heterogeneity of the COPD endophenotype seen in clinical practice [182, 183]. Such overlap syndromes associate with poorer outcomes and add complexity to diagnosis and treatment, underlining a continued need for refined stratification with a focus on precision medicine [182, 184, 185, 186] (Fig. 2, Ref. [52, 182, 187, 188, 189, 190, 191, 192, 193, 194, 195]).
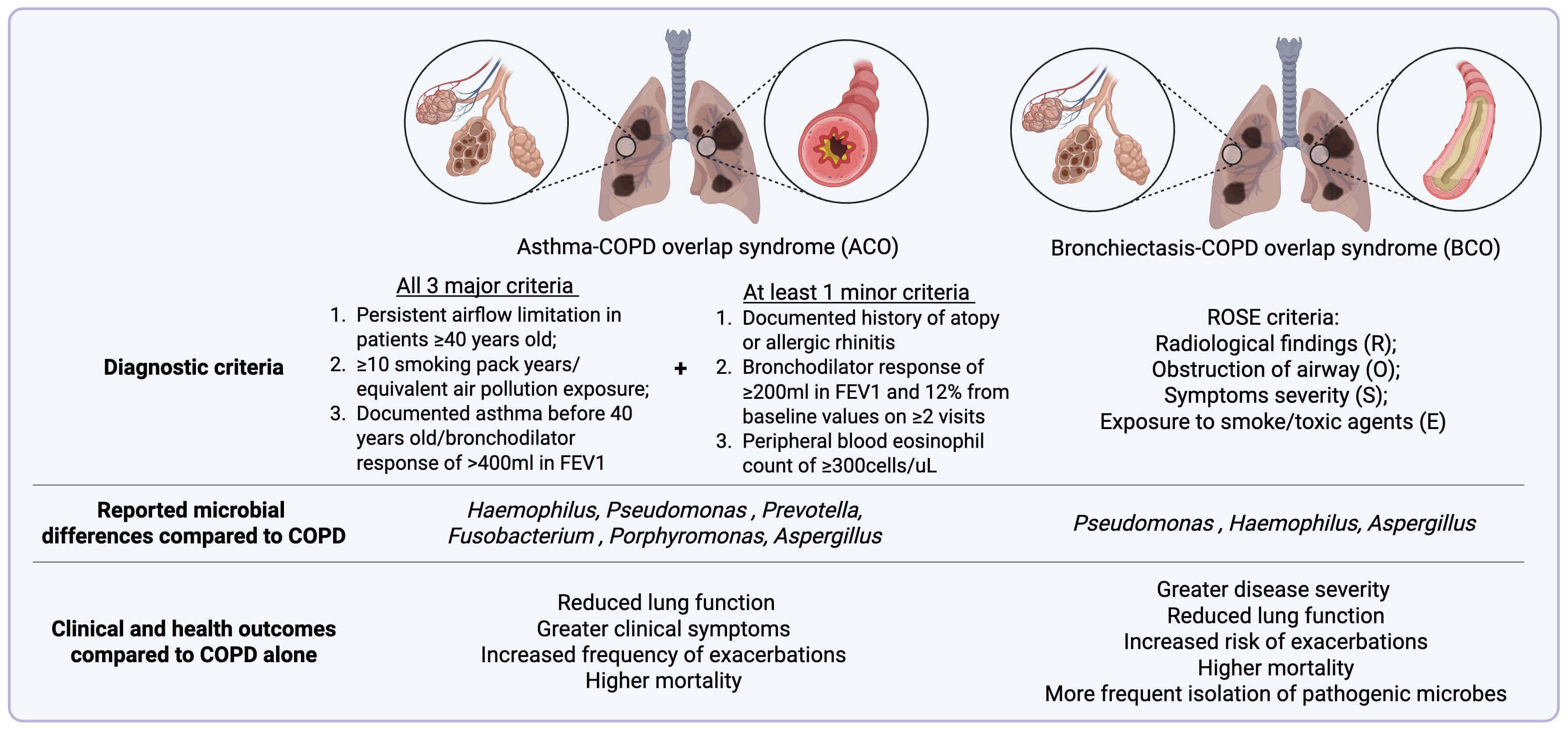 Fig. 2.
Fig. 2.
Summary of the diagnostic criteria and key microbial differences with associated clinical and health outcomes in asthma-COPD overlap (ACO) and bronchiectasis-COPD overlap (BCO). FEV1, forced expiratory volume in one second; ROSE, radiology, obstruction, symptoms and exposure. Created in BioRender (https://BioRender.com). Information on diagnostic criteria and clinical outcomes synthesized from references [52, 182, 187, 188, 189, 190, 191, 192, 193, 194, 195].
Clinically, ACO is difficult to diagnose due to the lack of a clear definition,
compounded by the heterogeneity of both conditions and their overlapping clinical
features [194, 196]. A roundtable
discussion by global experts was held, resulting in a consensus definition based
on three major criteria: (1) post bronchodilator forced expiratory volume in one
second over forced vital capacity ratio (FEV1/FVC)
Given the heterogeneity and unclear pathophysiology of ACO, several studies have explored the use of respiratory microbiome to better characterize this condition. Most of these studies focused primarily on bacterial, and to a lesser extent, fungal alterations. Research on viral involvement in ACO remains limited. Relative to patients with COPD alone, patients with ACO have significantly lower abundance of Fusobacterium and Porphyromonas. Notably, a decrease in the abundance of Porphyromonas was associated with elevated levels of Fractional Exhaled Nitric Oxide (FeNO), indicating a greater degree of airway inflammation [200]. Additionally, greater abundance of Haemophilus and pseudomonas found in ACO patients has been linked to higher mucus plug scores [201]. Further alterations in the airway microbiome were reported during ACO exacerbations, with a decrease in bacteriome richness, increase in evenness and abundance of Prevotella [196]. Although bacteriome changes in ACO are increasingly being studied, research exploring mycobiome changes remains scarce. Nonetheless, a study found that prevalence of A. fumigatus sensitisation was higher in ACO patients compared to patients with asthma alone, although this difference did not reach statistical significance. This finding suggests a potential role of A. fumigatus in ACO which could have implications for screening and management [202]. While more studies into ACO and its biomarkers are necessary to improve our current understanding of ACO, the combinations of ICS and LABA remains the recommended treatment options for ACO patients as LABA prescription alone is reported to increase the risk of hospitalisation and mortality [198, 203].
Similarly, the overlapping clinical features and lack of unifying definition of BCO has led to a wide variability in the reported prevalence, ranging from 4% to 70% across different studies [204, 205, 206]. To address this, the radiology, obstruction, symptoms and exposure (ROSE) criteria, incorporating radiological findings (R), airflow obstruction (O), symptom severity (S), and exposure to smoking or toxic air history (E), offers a more objective diagnostic framework [193]. Using this criterion, Polverino et al. [191] reported a high rate of COPD overdiagnosis in a large cohort of patients with primary bronchiectasis. The study also reported that patients who met the ROSE criteria had worse clinical outcomes, including an increased risk of exacerbation and hospitalisation compared to bronchiectasis-only patients [191]. Their findings aligned with previous studies depicting poorer lung function, increased exacerbations, greater disease severity, higher mortality and more frequent isolation of pathogenic microbes in BCO patients relative to patients with either COPD or bronchiectasis alone [52, 190, 205]. Huang et al. [182] reported an important, albeit ‘partial overlap’ between the microbiome-proteome profiles of stable COPD and BCO. In the BCO group, there was an increase in Proteobacteria and a decreased Firmicutes and Bacteroidetes, alongside the upregulation of proteins involved in the ‘neutrophil degranulation’ pathway, suggestive of potential pathophysiological differences between the entities [182]. Similarly, a separate study reports a substantial increase in the abundance of the Proteobacteria, specifically Pseudomonas and Haemophilus in BCO with the latter persist across both stable and exacerbation states when compared to COPD without bronchiectasis [109, 205, 207].
Beyond alterations in bacteriome, the role of the mycobiome in BCO remains less explored. Using culture and PCR techniques combined with sputum galactomannan testing, Everaerts et al. [208] reported higher sputum galactomannan levels and an increased abundance of A. fumigatus in BCO patients compared to COPD patients without bronchiectasis. A separate study evaluating the association of BCO, COPD and Aspergillus sensitisation found that BCO patients experience more frequent episodes of allergic bronchopulmonary aspergillosis (ABPA), and BCO-ABPA is clinically more severe compared to ABPA in COPD or bronchiectasis without overlap [93, 209]. While current research in BCO highlights the clinical relevance of specific microbes, further investigations into BCO-specific mechanisms, particularly microbiome-host interactions is needed to develop more effective diagnostic, management and therapeutic strategies for this patient group.
In addition to asthma and bronchiectasis overlap, COPD patients with other comorbidities, including cardiovascular disease and previous pulmonary tuberculosis (ex-TB) also experience poor outcomes, such as more frequent exacerbations and higher mortality [64, 210, 211, 212]. Notably, cytokine profiling on a large, multicentered cohort of Asian Chinese reveal a highly intricate cytokine network in a novel cluster of stable COPD patients with ex-TB compared to patients with other comorbidities [209]. Specifically, cytokines involved in tissue inflammation and airway remodeling such as TNF and platelet-derived growth factor (PDGF), were found to be more prominent in the ex-TB group. These findings suggest that even after clinical resolution, TB-recovered patients may experience a “memory effect” of the disease, leading to irreversible structural lung damage and pulmonary obstruction, which increases the risk of COPD progression and worse outcomes [64, 213, 214, 215]. Taken together, COPD patients with co-existing conditions are highly complex and further studies are required to explore the role of microbiome in these settings including its interaction with host and systemic inflammation.
Understanding microbiomes must move beyond single-kingdom assessment and evaluating inter-kingdom and even inter-organ interactions will be essential in providing novel and translatable clinical insight for COPD endophenotyping, therapeutic intervention and patient prognostication [158, 216]. Inter-kingdom assessment by microbial integration and co-occurrence analysis of bacterial, fungal and viral microbiomes demonstrates significant clinical utility for risk profiling exacerbations in bronchiectasis. Our group have identified significant changes in microbial interactions (“the interactome”) across bacteria, viruses and fungi in frequent exacerbators with bronchiectasis, changes undetectable when studying individual microbial kingdoms [158]. Future work in COPD should explore “interactomes” to refine risk-profiling COPD exacerbators.
Emerging evidence highlight a role for inter-organ axis, including the gut-lung axis, where microbial crosstalk occurs between these two anatomically and physiologically distinct organ sites. The gut microbiome and its metabolites influence COPD disease progression through various immune and neuroendocrine signaling pathways [217, 218, 219, 220]. Key metabolites including lipopolysaccharides, short chain fatty acids (SCFAs), trimethylamine-N-oxide, phytohemagglutinin and peptidoglycan are all implicated, although their clinical utility in COPD remains unexplored. Nevertheless, beneficial microbes in probiotics, which support a healthy gut microbiome and the production of these metabolites, have demonstrated anti-inflammatory properties, and symptomatic improvement in COPD [221, 222, 223, 224]. Similarly, SCFAs derived from the gut microbial fermentation of dietary fiber supplementation have demonstrable anti-inflammatory effects and symptom alleviation in COPD [225]. Further, therapeutic alteration of the gut microbiome via fecal microbiota transplantation (FMT) exhibits promising anti-inflammatory effects, reduces cell apoptosis and even emphysema in mouse models [226, 227, 228]. Taken together, therapeutic modulation of the gut microbiome through oral dietary supplements and FMT offers a promising avenue for exploration in the future management of COPD [229].
The lung-brain axis similarly represents an evolving paradigm although work
involving the role and influence of microbiome remains in its infancy [230]. Indeed, respiratory infections can increase the permeability of
blood-brain barrier to immune cells and induce neuroinflammation leading to
neuronal cell death of the brain, thereby illustrating immune mediated cross-talk
between lung and brain [231, 232].
Respiratory infections have also been associated with increased risk of ischemic
stroke [233], where bacterial lipopolysaccharides (LPS)
triggers systemic inflammation and blood clotting [234, 235]. Notably, COPD and air pollution elevates stroke risk whereby
systemic inflammation and oxidative stress results in vascular dysfunction
[236, 237, 238]. More
recently, the lung microbiome has been shown to regulate brain autoimmunity in
rats where neomycin-mediated modulation for the enrichment of LPS-producing
microbes suppressed neuroinflammation associated with the development of
autoimmune disease including multiple sclerosis [239].
Exposure to air pollutant Ozone (O3) has also been shown to be implicated in
neuropathology of Alzheimer’s Disease (AD) whereby phagocytic clearance of
amyloid-
The emerging role of microbial metabolites in organ-organ crosstalk should therefore also prompt the integration of microbiome with metabolomics alongside clinical characteristics to further delineation of endophenotypes [60, 241]. Such integrative analysis should also extend to incorporate multi-omics approaches encompassing proteomics and lipidomics in a systems biology approach could also identify novel pathobionts, further delineating the microbiome across the spectrum of pathogen, pathobiont and commensals as have been demonstrated in other chronic respiratory disease namely bronchiectasis [242, 243].
The role of the environment in COPD disease pathophysiology represents an additional and emerging frontier, with evidence demonstrating close relationships between environmental parameters and the airway microbiome in relation to negative clinical consequence [94, 98, 244]. Geographic and seasonal variation in microbiomes are documented across various organ systems including the lungs, reflective of the environmental influence on microbiome composition [93, 113, 115, 125, 245, 246, 247]. Given the constant exchange of air between lungs and the environment, the effects of the exposome, including climate and air microbiome, on the COPD lung microbiome and disease progression warrants further investigation as we enter the era of climate change [248].
The advent of NGS has revolutionized our understanding of the lung microbiome in COPD, demonstrating the remarkable microbial heterogeneity and trajectory across various disease states—from mild-moderate and stable phases to acute exacerbations—and highlighting its potential clinical value for early disease detection, patient stratification for personalised treatment approaches, and prognostic assessment of disease severity, exacerbations and mortality. Investigating these microbial ‘switches’ could further uncover underlying mechanisms driving COPD progression and pave the way for targeted therapeutic interventions to halt disease progression. Furthermore, the microbiome holds potential for endotyping overlap syndromes, thereby delineating heterogeneity and improve diagnosis.
Crucially, the translational implementation of NGS-approaches to bedside application in COPD will need to overcome several challenges. Areas requiring focus include standardization of sample collection and processing, timeliness of result availability, complexity and interpretability of bioinformatic requirements and infrastructural and sequencing costs [249, 250]. The constantly evolving field of bioinformatics to analyze sequencing data may further impede the ‘true’ implementation of standardized workflows for NGS. Accepted workflows in clinical settings typically require robust validation before routine implementation, and the dynamic nature of informatics needs to be carefully considered.
All supporting data are included in the article.
WLYK: Conceptualization, writing—original draft preparation, reviewing and editing, figure curation; KXT: Visualization, writing—original draft preparation, reviewing and editing; PYT: Visualization, writing—reviewing and editing; SHC: Conceptualization of paper, writing—reviewing and editing, obtained study funding and writing of the final manuscript. All authors contributed to important editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Not applicable.
The authors wish to thank The Academic Respiratory Initiative for Pulmonary Health (TARIPH) and the Lee Kong Chian School of Medicine Centre for Microbiome Medicine for collaboration support.
This research is supported by the National Research Foundation Singapore under its Open Fund-Large Collaborative Grant (MOH-001636) and administered by the Singapore Ministry of Health’s National Medical Research Council; the Singapore Ministry of Health’s National Medical Research Council under its Clinician-Scientist Individual Research Grant (MOH-001356) (S.H.C); Clinician Scientist Award (MOH-000710) (S.H.C); Transition Award (MOH-001275-00) (P.Y.T); Open Fund Individual Research Grant (MOH-000955) (S.H.C) and the Singapore Ministry of Education under its AcRF Tier 1 Grant (RT1/22) (S.H.C).
Fig. 1 and Fig. 2 were created using BioRender (https://BioRender.com). The authors have no financial or personal relationship with BioRender (https://BioRender.com), and the use of this tool does not imply any endorsement. PYT has served on advisory boards for Sanofi and AstraZeneca outside the submitted work. SHC has served on advisory boards for CSL Behring, Pneumagen Ltd., Zaccha Pte Ltd., Boehringer-Ingelheim and Sanofi, on DSMBs for Inovio Pharmaceuticals and Imam Abdulrahman Bin Faisal University and has received personal fees from Astra-Zeneca and Chiesi Farmaceutici, all unrelated to this work. All other authors have no conflict of interest to disclose.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.


