1 Surgical Oncology, Hangzhou Cancer Hospital, 310002 Hangzhou, Zhejiang, China
Academic Editor: Natascia Tiso
Abstract
Background: The histone lysine methyltransferase Histone-lysine N-methytransferase 2D (KMT2D) is a common mutated gene in a variety of cancers, including papillary thyroid cancer (PTC). However, the mechanism of KMT2D on the progression of PTC remains unclear. Methods: In this study, quantitative real-time polymerase chain reaction (qRT-PCR) and Western blotting were used to evaluate KMT2D expression between human normal cell (Nthy-ori 3-1) and PTC cells (TPC1, IHH-4 and BCPAP). Proliferation, migration and invasion of TPC1, IHH-4 and BCPAP were assessed by Cell Counting Kit-8 (CCK-8), Wound-healing assay and Transwell assay. The mechanism of KMT2D on thyroid papillary cancer was explored with Chromatin immunoprecipitation assay (ChIP), qRT-PCR and Western blotting. Results: The expression of KMT2D in PTC cells was significantly increased. Downregulation of KMT2D significantly decreased the proliferation, migration and invasion of PTC cells, which was correlated with decreased expression levels of H3K4me2, H3K9me2, NCOA6 and THRB. Meanwhile, ChIP assay demonstrated that KMT2D was associated with NCOA6. Conclusions: Study have shown that the downregulation of KMT2D reduces proliferation, migration and invasion of thyroid papillary carcinoma cells through epigenetic modification of NCOA6/THRB signal axis. These results provide a new insight into the role of KMT2D in migration and invasion of PTC.
Keywords
- papillary thyroid cancer
- KMT2D
- epigenetic modification
- migration
- invasion
Thyroid cancer (TC) is the most common malignant disease of endocrine system, and its incidence has been on the rise in recent years. In 2019, there were 233,846 new cases of thyroid cancer and 45,575 deaths from the disease worldwide [1]. China has the highest number of thyroid cancer cases and deaths in the world. Papillary thyroid cancer (PTC) accounts for approximately 90% of all thyroid cancers, and the increased incidence and mortality of PTC leads to an actual increase in all TC cases [2, 3]. The incidence of PTC increases with age, with more women than men [4]. Surgery is usually the first line of treatment for PTC. However, the prevalence of PTC remains a concern, as the recurrence rate of PTC is close to 30% and the cause-specific mortality rate over a 30-year period is 8.6% [5]. Therefore, it is an urgent and necessary task to find diagnostic markers and new therapeutic targets for PTC recurrence and metastasis.
The pathogenesis and progression of PTC has been reported to be associated with genetic and epigenetic alterations, including driver point mutations in BRAF and RAS, the diverse gene fusion (RET/PTC, ETV6-NTRK3) and miR146b, miR21, etc. [6, 7]. The cancer genome atlas project identified 299 genes and 24 pathways that drive tumor progression, including 12 genes involved in histones, histone methylation and demethylation [8]. Our previous studies found that thyroid cancer could induce Histone-lysine N-methytransferase 2D (KMT2D) to inhibit apoptosis by regulating H3K4 site and NCOA1/6 in the protein function and protein network analysis (Supplementary Fig. 1). In TC patients, KMT2D mutation status is associated with shorter disease-specific survival. Thanyawat et al. [9] also found that epigenetic modifications of KMT2D are involved in the malignant progression of TC. is a histone methyltransferase that stimulates tumor growth and triggers metastasis. KMT2D is located on chromosome 12q13.12 and is widely expressed in adult tissues. It is reported that KMT2D plays a role in binding enhancer regions and maintaining global H3K4me1/2 levels. H3K4me2 and H3K9me2 are strongly associated with gene transcription. H3K4me2 is found in a wider range of promoters, enhancers and long-range regulatory elements. H3K9me2 presents chromatin extensively and is enriched in a number of tightly-controlled signaling and cell-type specific pathways. In addition, KMT2D is one of the most commonly mutated genes in many different types of cancer. In breast cancer, the PI3K pathway regulates ER-dependent transcription through the epigenetic regulator KMT2D [10]. Ming et al. [11] indicated that KMT2D regulates H3K4 methylation and participates in the pathology of ovarian cancer. These studies suggest that KMT2D may play an important role in PTC.
Consequently, we focused on the histone methyltransferase KMT2D to explore the specific mechanism of PTC carcinogenesis and invasion. In this study, we analyzed KMT2D expression levels in different thyroid cancer cell lines and evaluated KMT2D-mediated epigenetic modification on the progression of PTC. And, we proposed that inhibitors targeting the KMT2D gene may be used for the treatment of PTC.
TPC1 (iCell-h309), IHH-4 and BCPAP (iCell-h022) cells were purchased from iCell
Bioscience Inc (Shanghai, China). All thyroid papillary carcinoma cell lines were
cultured in Dulbecco’s modified Eagle’s Medium (DMEM, Hyclone, Logan, Utah, USA)
supplemented with 10% fetal bovine serum (FBS, ZhejingTianhang Biotechnology Co.
Ltd, Hangzhou, China). A human normal cell line Nthy-ori 3-1 was required from
iCell Bioscience Inc and cultured in RPMI
medium 1640 with 10% FBS. All cells were maintained in a humidified atmosphere
at 5% CO
TPC1, IHH-4 and BCPAP cells were seeded into 6-well plates, and transfected with two siRNA targeting KMT2D or negative control (NC) using lipofectamine 2000 (Thermo Fisher Scientific, Waltham, MA, USA). The efficacy of RNA interference in thyroid papillary carcinoma cells was assessed by qRT-PCR. Compared with sh-NC group, the mRNA level of KMT2D was significantly decreased in si-KMT2D (1) and si-KMT2D (2) group (Supplementary Fig. 2).
RNA was isolated from Nthy-ori3-1, TPC, IHH-4 and BCPAP cells with Trizol (Sangon Biotech, Shanghai, China). The total RNA was reverse transcribed into cDNA at the following condition: 95 °C, 10 min; 95 °C, 15 s; 60 °C, 60 s; 40 cycles. The mRNA expression was quantified by the SYBR Premix Ex TaqII (Takara, Shiga, Japan) and normalized by Glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The primers were listed in Table 1.
| Gene | Amplicon lengths | Forward primer | Reverse primer |
|---|---|---|---|
| KMT2D | 793 | GACACAACAACACGATGCTCC | ACGTGACCTCTATGCAACCC |
| NCOA6 | 167 | AAAACGTGCCCAATTTGTTACAC | GAGAATCCCTAAATCCCGAAGC |
| THRB | 86 | TGGGACAAACCGAAGCACTG | TGGCTCTTCCTATGTAGGCAG |
| GAPDH | 119 | TGTGGGCATCAATGGATTTGG | ACACCATGTATTCCGGGTCAAT |
The cells were trypsinized, harvested and lysed in Radio-Immunoprecipitation Assay (RIPA) buffer containing phenylmethylsulfonyl fluoride (PMSF) (Beyotime, Shanghai, China) and protease inhibitor (CWBIO, Beijing, China). The concentration of total protein was measured by the BCA Protein Assay Kit (Beijing Solarbio Science & Technology Co., Ltd, Beijing, China). After loading samples, proteins were migrated by Sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes, which were then blocked with non-fat milk for 2 h. The primary antibody was used to incubate membrane overnight at 4 °C. The antibodies were bought from Affinity Biosciences company (KMT2D Antibody: DF9646; H3K4me2 Antibody: AF0589; H3K9me2 Antibody: DF6937; NCOA6 Antibody: DF13176; THRB Antibody: AF0357; H3 Antibody: AF0863; GAPDH Antibody: AF7021). The second antibody was diluted with non-fat milk and incubated membrane for 1.5 h at room temperature. ECL detection kit and ChemiScope 6100 (Clinx Science Instruments Co., Ltd, Shanghai, China) were used to visualize protein bands.
High-sensitivity ChIP Kit (ab185913, abcam, Cambridge, UK) was used in this
study. We assessed the association between KMT2D and NCOA6. The protocol was as
follow: (1) Formaldehyde cross-linking and ultrasonic crushing of cells: the
cells were removed, formaldehyde was added to make the final concentration of
formaldehyde 1%, and the cells were incubated at 37 °C for 10 min. The
cross-linking was terminated, the medium was sucked up, and the cells were washed
twice with cold Polybutylene Succinate (PBS). The cells were collected in a 15 mL centrifuge tube with a
cell scraper. After pre-cooling, the cells were collected by centrifugation, and
the supernatant was poured out. (2) Impurity removal and antibody feeding: after
ultrasonic crushing, centrifuge to remove insoluble substances; A 900μL dilution
Buffer and 20 μL of 50
Cells were seeded in a 96-well plate for 24 h, and 10
Cells were resuspended in serum-free medium, added to a six-well plate (5
Cells were added into the upper chambers of 24-well transwell chambers (Corning, New York, USA) with or without BD Matrigel (invasion assay and migration assay). After 24 h of incubation, cells on the surface of upper chamber were removed with cotton swabs. The invaded/migrated cells on the lower surface were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. Cells were imaged and counted in five random fields.
Statistical analysis was performed with SPSS 18.0 software (IBM Corp., Chicago,
IL, USA) and GraphPad Prism 8 software (GraphPad Software Inc., San Diego, CA,
USA). Each experiment was repeated three times. Differences between two groups
were made using Independent sample T test or Kruskal-Wallis H Test. All
data were expressed as the means
We compared the expression of KMT2D in thyroid papillary carcinoma cell lines and normal thyroid follicular cell line by qRT-PCR and Western blotting (Fig. 1). The results demonstrated that KMT2D mRNA levels were significantly higher in TPC, IHH-4 and BCPAP cells than in Nthy-ori3-1 cell. Western blotting analysis of KMT2D protein confirmed the RNA expression data, showing remarkably higher expression in thyroid papillary carcinoma cells than in normal thyroid cell.
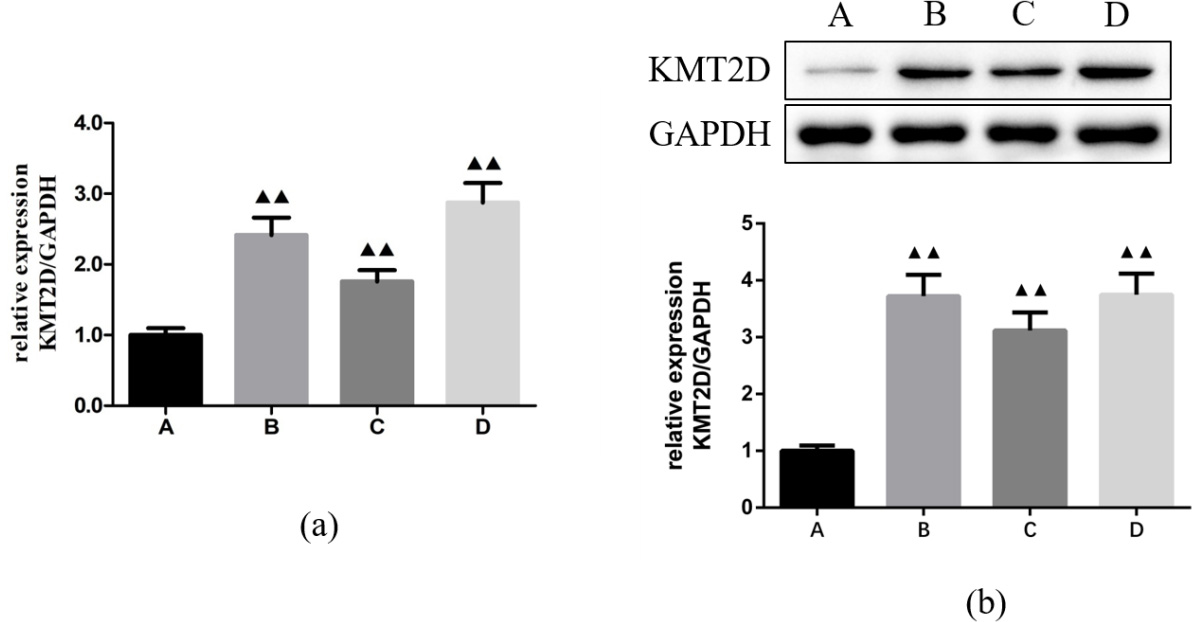 Fig. 1.
Fig. 1.The mRNA and protein expressions of KMT2D in
Nthy-ori3-1, TPC, IHH-4 and BCPAP cells. (a) qRT-PCR analysis.
(b) Western blotting analysis. A: Nthy-ori3-1 cell; B: TPC cell;
C: IHH-4 cell; D: BCPAP cell.
TPC, IHH-4 and BCPAP cells were transfected with KMT2D inhibitors or NC
to evaluate the effect of KMT2D in thyroid papillary carcinoma cells.
CCK-8 assay demonstrated the cell viability in si-KMT2D (1) and
si-KMT2D (2) groups was lower than in si-NC group (Fig. 2a). It
indicated that KMT2D knockdown suppressed the proliferation of TPC,
IHH-4 and BCPAP cells. Then, Wound healing and Transwell assay were used to
investigate the role of KMT2D in the cell invasion and migration. As
shown in Fig. 2b, the migration distances in si-KMT2D (1) and
si-KMT2D (2) groups were less than si-NC group at 12 h after scratch.
Similarly, the Transwell assay showed a significantly decrease in the number of
migrated cells in si-KMT2D group (p
 Fig. 2.
Fig. 2.KMT2D regulates migration and invasion in TPC, IHH-4
and BCPAP cells. (a) CCK8 assay. (b) Wound healing. (c) Transwell
assay.
ChIP assay showed that KMT2D is associated with NCOA6 in TPC, IHH-4 and BCPAP cells (Fig. 3a). Compared with anti-IgG group, the input of NCOA6 is significantly increased in anti-KMT2D group. To understand why KMT2D is required for the activities of NCOA6, we examined the levels of H3K4me2 and H3K9me2, which are closely correlated with active enhancers and promoters. Western blotting showed that silencing KMT2D (si-KMT2D) was significantly upregulated H3K4me2 and H3K9me2 at protein levels (Fig. 3c). In addition, western blotting and qRT-PCR analysis were used to explore the association of KMT2D and NCOA6. As shown in Fig. 3b,c, KMT2D knock out markedly decreased the expression of NCOA6 and THRB compared with si-NC group. These results demonstrated that KMT2D regulates the expression of NCOA6 and THRB through demethylation of H3K4me2 and H3K9me2 in thyroid papillary carcinoma cells.
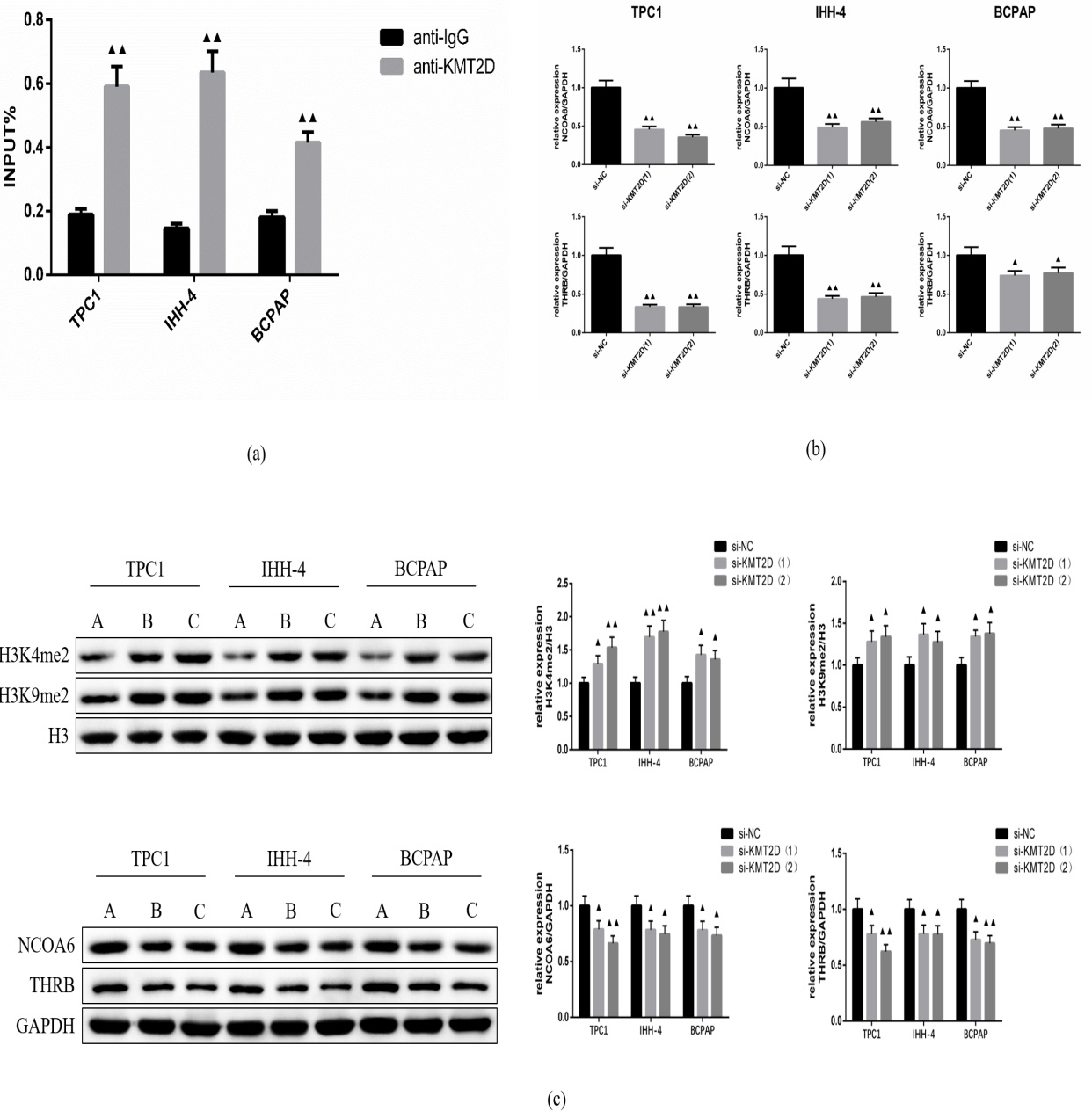 Fig. 3.
Fig. 3.The mechanism of KMT2D on papillary thyroid carcinoma
cells. (a) ChIP assay. (b) qRT-PCR analysis. (c) Western blotting analysis. A,
si-NC; B, si-KMT2D (1); C, si- KMT2D (1).
KMT2D plays an important role in tumorigenesis, differentiation, metabolism and inhibition. KMT2D mutations are common in various types of cancer, including the brain, lymph nodes, blood, lung, thyroid, intestine, and endometrium [12, 13, 14]. The anti- or pro-tumor effects of KMT2D are related to cell types. In bladder cancer tissues and cell lines, KMT2D is downregulated and its overexpression could significantly alleviate tumor progression [15]. KMT2D also exhibits tumor suppressor effects in melanoma and pancreatic cancer cells [16, 17]. It may depend on the inhibition of downregulated oncogenes, including PTEN, P53 and STAG2 [18]. On the other hand, KMT2D is required for the formation of acute myeloid leukemia and is highly expressed in esophageal squamous cell carcinoma, prostate cancer, pancreatic and gastric cancer [15]. In this study, we found that KMT2D was overexpressed in PTC cell lines by qRT-PCR and Western blotting. It suggests that KMT2D may play an important role in the initiation and development of PTC. KMT2D deficiency has been associated with cell proliferation and apoptosis in gastric, pancreatic, prostate and colon cancer [19]. Consistent with previous findings, KMT2D knockdown inhibited the proliferation, migration and invasion of TPC, IHH-4 and BCPAP cells. It indicated the tumor inhibition of KMT2D on PTC.
Then, we studied the underlying mechanism of KMT2D mutations on PTC. KMT2D mutations lead to defective enhancer regulation and genomic instability during DNA replication and/or transcription [20]. KMT2D is associated with WRAD, NCOA6, PTIP, PA1 and H3K27 demethylase UTX in protein complexes. Cefan et al. [21] reported that Pygo2 is involved in the progression of glioma by upregulating the KMT2D complex and H3K4me3. Whole exome sequencing showed that there were 6 harmful mutations (ARID1A, NCOR2, KMT2D, NCOA6, MECP2 and SUPT6H) in the epigenetic regulation of colorectal cancer in Chinese patients. Moreover, KMT2D and NCOA6 encode subunits of histone H3K4 methyltransferase protein complexes that act as coactivators of p53 [20]. In this study, ChIP assay confirmed the binding of KMT2D and NCOA6 in PTC cells. Meanwhile, the mRNA and protein levels of NCOA6 and THRB were significantly decreased in si-KMT2D thyroid cells, suggesting that KMT2D modulates the expression of NCOA6/THRB signal axis. In addition, western blotting showed that si-KMT2D could upregulate the expression of H3K4me2 and H3K9me2 in TPC, IHH-4 and BCPAP cells. The role of KMT2D in epigenetic regulation was demonstrated.
In summary, we found that KMT2D is markedly upregulated in PTC cells and plays a vital role in the migration and invasion of PTC cells. KMT2D knockdown could regulate the expression of NCOA6, THRB, H3K4me2 and H3K9me2. The results indicated that KMT2D regulated the NCOA6/THRB signaling axis through epigenetic modification, promoting the migration and invasion of PTC.
In conclusion, we have identified that NCOA6/THRB signal axis as a target of KMT2D in PTC cells, which may be regulated by epigenetic modification. It provides new insights into the role of KMT2D in the treatment of PTC.
All data generated or analyzed during this study are included in this published article.
RW and YH mainly performed this study, RW wrote the draft, SX supervised this study, YW analyzed data.
This project was approval by Hangzhou Cancer Hospital ethic commitment. (HZCH-2020-109).
We sincerely thank all who participated in this study.
This study was supported by Hangzhou Science and technology Development plan project (20201231Y031).
The authors declare no conflict of interest.
References
Publisher’s Note: IMR Press stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.



