We previously reported that gigantol extracted from Caulis Dendrobii has significant therapeutic benefits for the treatment of galactosemic cataracts through its ability to inhibit aldose reductase (AR) activity. In this study, we identified the binding sites and structurally characterized the interaction between gigantol and AR, to understand the mechanism (s) of the effects of gigantol on cataracts. Gigantol was found to be protective against diabetic cataracts (DC) in rats induced by streptozotocin. Molecular docking predicted the binding sites between AR and gigantol to be residues Trp111, His110, Tyr48 and Trp20. Mutation of each of these residues led to a significant reduction in AR activity. Cold-spray ionization mass spectrometry measurements showed that the binding of gigantol to AR is concentration-dependent and that the maximum stoichiometric ratio of non-covalent bonding is 1:24.4. pH and temperature did not influence the interaction. Taken together, we provide further mechanistic evidence of the beneficial effects of gigantol on DC.
Over time, diabetic patients develop many complications, including cardiovascular disease, retinopathy, neuropathy, and nephropathy. Amongst them, diabetic cataracts (DC) are second most prevalent retinopathy (1-3). With the increased prevalence of overweight/obese patients and increased consumption of refined foods, the incidence of diabetes is steadily growing, which in turn increases the incidence of DC and subsequent blindness. While microsurgery can treat cataract-induced vision impairment, secondary cataracts commonly develop due to surgery-induced proliferation and migration of the remaining lens epithelial cells (4-7). Furthermore, the cost of surgery hampers its affordability. The development of cost-effective treatments to prevent or treat DC are therefore required (2-3).
The pathological mechanism (s) of DC development and progression remain poorly characterized. It is known that DC development can be attributed to a change in lens osmotic pressure, oxidative damage, and non-enzymatic glycosylation of the lens protein (8-10). High, abnormal circulating glucose levels in patients with diabetes can activate the expression of aldose reductase (AR) in lens epithelial cells (2). AR is one of the rate-limiting enzymes in the polyalcohol metabolism pathway. Overexpression of AR enhances polyalcohol metabolism and enables the accumulation of sorbitol, consequently augmenting osmotic pressure and cell membrane permeability, both of which induce cell injury, lens edema, and DC development (8, 11). A wide variety of AR inhibitors (ARIs) can effectively inhibit AR activity, which in turn mitigates polyalcohol-mediated DC progression (12).
Dendrobium plants contain an array of bioactive compounds including alkaloids, polysaccharides, sesquiterpenoids, fluorenones, bibenzyls, and anthrenes (14-18). Gigantol (PubChem CID: 10221179) is a bibenzyl phenol found in Dendrobium with one hydroxy moiety on each aromatic ring. It displays a wide range of bioactivities, including anti-oxidant, anti-mutagen, anti-platelet aggregation, anti-spasm and anti-aging effects (14, 19-21). As a small molecule, gigantol can easily pass through the cell membrane of enterocytes, and no known adverse effects have been reported following its consumption (21). We previously found that gigantol extracted from Dendrobium has significant therapeutic benefits for the treatment of galactosemic cataracts in rats by inhibiting AR activity (22). In this study, we investigated the effects of gigantol on the formation of DC and further explored the interaction between gigantol and AR.
The stems of Dendrobium aurantiacum var. denneanum (kerr) Z.H. Tsi were collected from the Lesan County, Sichuan Province, P.R. China in December 2012, and authenticated by Professors Min Li and Tingmo Zhang of the Chengdu University of Chinese Medicine. A specimen voucher (No. 2012122001) was deposited in Wan’an Dendrobium Industry and Development Co., Ltd. (Sichuan Province, P.R. China). Field studies did not involve endangered or protected species and no specific permission was required for these locations/activities. Gigantol was extracted from Dendrobium, according to the protocol of Zheng et al. (14). The identification of extracted gigantol was verified by thin layer chromatography and its purity determined by HPLC was 98%. Ammonium acetate was purchased from Genebank Gene-Tech Co. Ltd; streptozotocin (STZ), DL-glyceraldehyde from Sigma-Aldrich (St. Louis, MO, USA) and reduced nicotinamide adenine dinucleotide phosphate (NADPH) from Italian Roth Corp. Pirenoxine eye drops were manufactured by Wuhan Tiantianming Pharmacy Co. Ltd. P.R. China. All other reagents were of analytical grade.
Thirty male and female SD rats (5 months old, mean body weight: 150 ± 10 g) were obtained from the Laboratory Animal Center of the Guangzhou University of Chinese Medicine. All animal procedures were approved by the Institutional Animal Care and Use Committee of the Guangzhou University of Chinese Medicine and handled in accordance with institutional guidelines for the care and use of laboratory animals. Animals were acclimatized for 1 week with ad libitum food and water provided. A 12h light/dark cycle was maintained.
After one week of acclimation, rats were divided into four groups of 15 animals. The normal control (Group I; n = 15) received a daily intraperitoneal injection of 10 mL/kg saline and drinking water, whilst experimental rats received a single intraperitoneal injection of freshly prepared STZ (30 mg/kg) in saline (23). After 72 hours, body weight and blood glucose levels were monitored, and cataract formation in each animal was observed. Rats with fasting blood glucose levels over 130 mg/dL and stage 2 lens opacity on day 30 were considered diabetic and were randomly divided into three groups (n = 15): viz. model group, gigantol group and positive control group. The model group (Group II) were treated daily with 3 drops of saline three times a day (TID). The gigantol group (Group III) received 3 drops of 2 mg/mL gigantol three TID. The positive control group (Group IV) received 3 drops of 0.053 mg/mL pirenoxine sodium eye drops three TID. Lens transparency was monitored daily for 90 days using the YZ-5E slit lamp microscope (Suzhou Medical Apparatus and Instruments Factory, China). The initiation and progression of lenticular opacity was graded according to 5 categories (23): stage 0- clear lenses and no vacuoles; stage 1 - vacuoles cover approximately 50% of the surface of the anterior pole, forming a subcapsular cataract; stage 2 - some vacuoles have disappeared and the cortex exhibits a hazy opacity; stage 3 - a hazy cortex remained and dense nuclear opacity is present; and stage 4 - a mature cataract is observed as dense opacity in both the cortex and nucleus. The incidence of cataract appearance was expressed as the percentage of total lenses in each group.
Rats in Groups I-III were euthanized by CO2 asphyxiation and their eyes enucleated. Lenses were carefully removed from enucleated eyes using the posterior approach, washed with saline, and frozen at -70°C until use. Lens homogenates were prepared according to the methods of Hayman and Kinoshita (24–27), with minor modifications. Briefly, lenses were pooled and then homogenized in 0.1 M phosphate buffer saline (PBS, pH 6.2). After centrifugation at 10,000 rpm for 30 min, AR supernatants were collected. All procedures were performed at 4°C.
The inhibitory effect of gigantol on AR activity was examined as previously described (28, 29). Briefly, AR solution was diluted 100-fold in PBS and gigantol was dissolved in dimethyl sulfoxide (DMSO). The final concentration of DMSO in the experiments was <1%. Enzyme activity was measured during AR purification through monitoring the decrease in NADPH absorbance at 340 nm, for 10 min and at 40s intervals, in a Bio Tek Power Wave XS spectrophotometer (Bio Tek Instruments, Winooski, VT, USA) maintained at 25°C. The 200 μL reaction mixture contained 200 μg/mL enzyme solution (20 μL), 0.104 mM NADPH (50 μL), gigantol (20 μL), 1mM DL-glyceraldehyde (50 μL) and ddH2O (10 μL). Gigantol was added at concentrations ranging from 0.1-10 mM. The inhibitory ratio (%) was calculated according to previously described formula (30): inhibition ratio (%)= (1- (A2-A0)/ (A1 -A0))×100%, where A0 is the decrease in NADPH absorbance every minute without adding AR, substrate, and gigantol; A1 is the decrease in NAPDH absorbance every minute without gigantol; and A2 is the decrease of NAPDH absorbance every minute with gigantol added. The IC50 of gigantol was obtained from inhibition curves. The Km (Michaelis-Menten constant) and the Vm (maximum velocity) were calculated from the double-reciprocal plot. All experiments were performed in triplicate.
The simulation of molecular dynamics (MD) and molecular docking were performed using the Amber 10 Molecular Dynamics Package (31). To prepare the molecular system, we first established the topology and coordinated files, set the force field to parm 99, added water to the gigantol system, applied transferable intermolecular potential 3 points box (TIP3PBOX) water model, and set the distance between the water environment and the molecule (9 angstroms). We designed the molecule as an octahedron to prevent washing of the protein from the water environment and to ensure that the whole system was electrically neutral. Gaussian 03W was adopted and the basic installation was set as HF/6-31G (32). Energy optimization was completed by hydrogenation between gigantol, NADPH, and AR.
Molecular Dynamics Simulation (MDS) in the solvent environment was performed under several important conditions. (1) Energy optimization of the solvent environment: Van der Waals interactions were removed to fix the protein. The maximum frequency of energy reduction optimization (macyc) was x 1000. The MD included: volume immobilization (ntb = 1) and atom position energy was inhibited (ntr = 1). The distance of the cut was 10. The force for stabilizing the peptide chain was 500 kal/mol and the number of peptide chain residues was 1316. (2) Energetic optimization of the system: the interaction with abnormal energy was removed from the whole system. The maximum frequency for energy reduction optimization (macyc) x 2500 times. Following energy optimization, the frequency (ncyc) was changed to x 1000, from the quenching step to the gradient change. The MD volume immobilization (ntb = 1) and atom position energy was not inhibited (ntr = 0) and the distance of the cut was 10 (3). Limited molecular kinetics: the kinetics maintained protein immobility and various layers of the solvent were dissolved. Systemic temperature was gradually increased from 0 to 300 K. After 20 ps, the MD volume immobilization of ntb = 1, step length of 0.002 ps (dt), and arithmetic step length of 10000 (nstlim). The collision frequency at this time was 1 PS, hydrogen bonds were (ntc = 2) and the force was 5-10 kcal/mol (ntr = 1). Molecular dynamic simulation of the solvent environment was performed under the following conditions: one atmospheric pressure at 300K, 100ps, tb=2, the MD volume immobilization (ntb=1), one unit pressure (pes = 1), pressure relief time of 2ps (taup = 2), and arithmetic step width of 2500000 (nstlim). After dynamic simulation for 6 nanoseconds (ns), the molecular movement was tracked by the imaging system. The MDS of the whole system was performed under the following conditions: atmospheric pressure at 300 K, 100 ps, tb = 2, molecular dynamics remaining volume immobilization (ntb = 1), one unit pressure (pes = 1), pressure relief time of 2ps (taup = 2), arithmetic step width of 2500000 (nstlim). Following dynamic simulation, molecular movements were tracked on the imaging system.
MDS was performed using a flexible docking module (FlexX, Sybyl 17.3), and gigantol was treated with the Wash and Energy Minimizing modules of the Molecular Operating Environment drug design software. Gaussian03W was used for energy reduction optimization with the basic installation of B3LYP/6-31+G (d) (33). The crystal structure of AR with the target enzyme structure of 1USO was acquired from the PDB database (34). The 6.5 Å area around the ligand crystal structure of AR was defined as “the active pocket”, and represented the docking location of gigantol.
For AR overexpression studies, full-length cDNA of AR with Hind III or Xho I restriction sites was cloned into the pET28a expression vector (Figure 1). Trp111, His110, Tyr48 or Trp20 residues were mutated to alanine by Quik Change XL site-directed mutagenesis (Stratagene, USA). (28-29). Primers were synthetized by Shanghai Biotechnology Co., Ltd and are listed in Table 1. Recombinant plasmids was sequenced by BGI genomics Co., Ltd, Shenzhen, P.R. China, followed by analysis using the Basic Local Alignment Search Tool (BLAST).
 Figure 1.
Figure 1.
Sequence of WT and mutant pET-28a-AR. (A) Plasmid maps of pET-28a and pET-28a-AR. pET-28a was digested with HindIII and Xhol and ligated with a 951 bp AR and its mutant fragments. (B) Sequences of wt-AR and its mutants.
| Primers | Primer sequence |
|---|---|
| Wild-type AR | 5’ CCCATCCTGGGGTTGGGTACC GCT AAGTCCCCTCCAGGGCAG 3’5’ CTGCCCTGGAGGGGACTTAGCGGTACCCAACCCCAGGATGGG 3’ |
| W20A | HindIIIF:5’ CGACTGTGCCCATGTGGCACAGAATGAGAATGAGGTGG 3’XhoIR:5’ CCACCTCATTCTCATTCTGTGCCACATGGGCACAGTCG 3’ |
| Y48A | HindIIIF: 5’ CGACTGTGCCCATGTGGCACAGAATGAGAATGAGGTGG 3’XhoIR: 5’ CCACCTCATTCTCATTCTGTGCCACATGGGCACAGTCG 3’ |
| H110A | HindIIIF: 5’ CTACCTGGACCTCTACCTTATTGCATGGCCGACTGGCTTTAAG 3’XhoIR: 5’ CTTAAAGCCAGTCGGCCATGCAATAAGGTAGAGGTCCAGGTAG 3’ |
| W111A | HindIIIF: 5’ CTGGACCTCTACCTTATTCAC GCA CCGACTGGCTTTAAGCCT 3’XhoIR: 5’ AGGCTTAAAGCCAGTCGG TGC GTGAATAAGGTAGAGGTCCAG 3’ |
The bacterial expression and purification of AR and its mutants was performed as previously described (24, 27-29, 35). Briefly, pET28a-AR and its corresponding mutant pET28a-AR Δ1-Δ4 expression vectors were transfected into E. coli BL21 (DE3) and co-cultured overnight at 18 °C in LB containing 50 μg/mL kanamycin. Positive clones were screened using 12% sodium dodecyl sulfate and polyacrylamide gel electrophoresis (SDS-PAGE) electrophoresis. For purification, overnight cultures were lysed in 50 mM NaH2PO4, 300 mM NaCl and 20 mM imidazole, pH = 8.0, and ultrasonicated for 20 min at 4°C. After 30 min of centrifugation at 10,000 rpm at 4°C, supernatants were collected and incubated with Ni-NTA beads for 1 h on ice, and subjected to purification using a cartridge (Bio-Rad, Hercules, USA). After washing twice with 8 mL of wash solution (50 mM NaH2PO4, 300 mM NaCl, 20 mM imidazole,pH = 8.0). AR was eluted in 50 mM NaH2PO4, 300 mM NaCl, 250 mM imidazole and pH = 8.0. Protein content and purity was assessed using the Bradford method (36). Purified proteins were stored in 15% glycerol at -80°C.
The activity of recombinant AR was determined as described. The inhibitory effects of gigantol on WT and mutant AR activity was assessed at concentrations of 0.1-10 mM.
Gigantol (15 µM) prepared in dehydrated alcohol was mixed with 25µl of AR (2.8 μM) in water at a series of molarity ratios: 1:3, 1:9, 1:27, 1:81, 1:243, and 1:729. The resulting mixtures were dried under nitrogen, diluted 50-fold using 10 mmol/L ammonium acetate buffer (pH = 6.7), and injected onto CSI-MS (JMS-T100CS, JEOL, Tokyo, Japan) for analysis of the gigantol-AR complex. The following parameters were set in the CSI-MS: CSI source, mass scan range: 100-1000 m/z, needle voltage: -2200 V, camera lens voltage: -10 V, hole 1 voltage: -75 V, hole 2 voltage: -8 V, capillary tube temperature: 10°C, detector voltage: 2500 V, and 1 min negative ion scan. The injection volume was 20 μL.
Data were expressed as mean ± SD, and significance was assessed using a two-tailed Student’s t test. P≤0.05 was considered statistically significant. Analyses were performed using SPSS 16.0.
To examine the effects of gigantol, the change in STZ-induced lens opacity was monitored using slit lamp microscopy. Cataract onset occurred 23 days post-STZ treatment. All lenses in the normal control group were clear (stage 0). However, most lenses (60%) in Group II were at stage 3 or 4 and 0 % were clear after 50 days. Most lenses in Group III and IV rats treated with gigantol and pirenoxine remained at stage 2 after 50 days (Figure 2). After 90 days, 75% of the lenses in Group II developed a mature cataract (stage 4), as opposed to 15% and 20% in Groups III and IV, respectively. Both gigantol and pirenoxine significantly reduced STZ-induced lens opacity in comparison to the model group (p <0.01). Gigantol was more protective than pirenoxine (p <0.01).
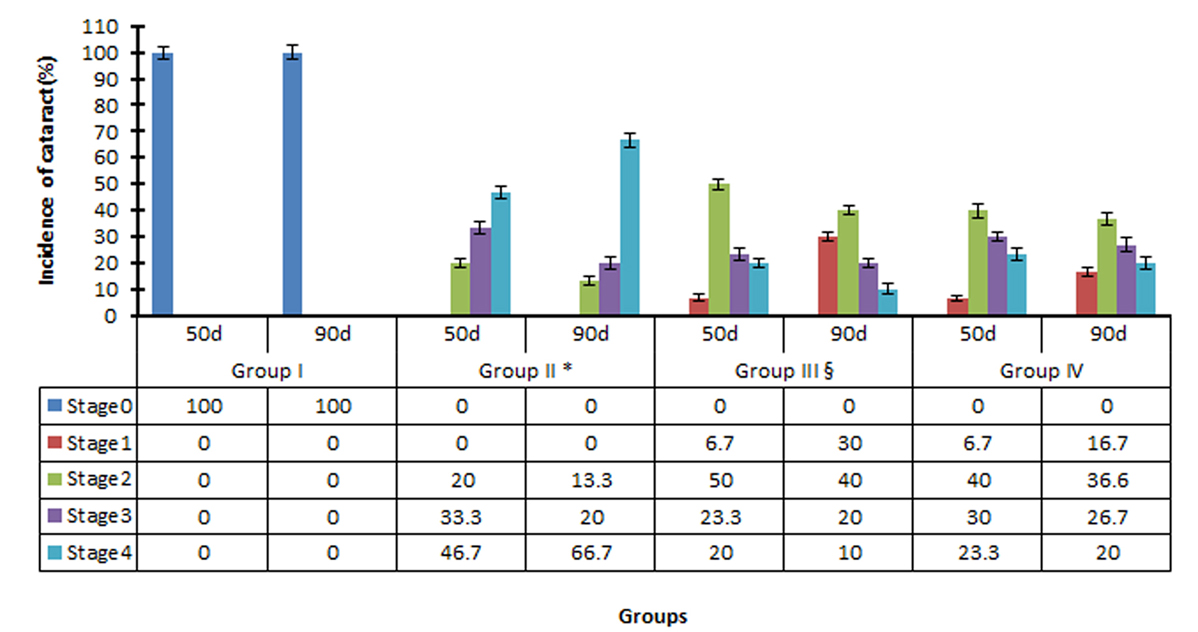 Figure 2.
Figure 2.
Progression of STZ-induced DC in rats treated with saline (Group II), gigantol (Group III) and pirenoxine (Group IV). Rats in group I were normal (control animals). Results are expressed as the mean ± SD (n = 15). *P<0.05 vs. group III and Group IV; §p<0.05 vs. Group IV.
Purified AR showed a single band of molecular weight: 36000 ± 1000 Dalton (Da) on SDS-PAGE.The Km and Vm values obtained from the double-reciprocal plot were 8.09 mM and 11.6 μmol/L/s, respectively (Figure 3). Gigantol inhibited AR activity by 30.7, 53.0, and 68.6%, respectively at concentrations of 0.1, 1, and 10 mM. The IC50 was 0.24 ± 0.01 mM.
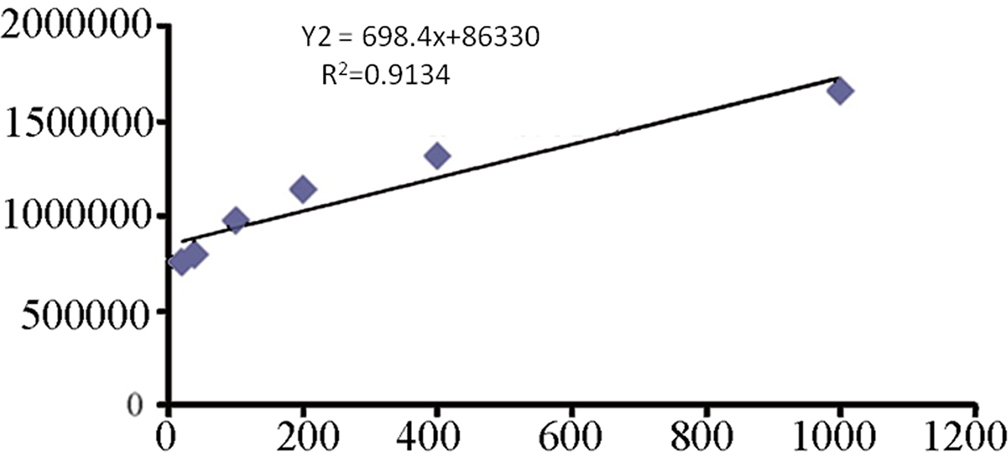 Figure 3.
Figure 3.
Km and Vm values for lens purified AR towards DL-glyceraldehyde. Values were obtained from double-reciprocal plots.
The binding residues for gigantol on AR were identified as Trp111, His110, Tyr 48, and Trp20, which formed “the active pocket” for gigantol. His110 and Tyr48 formed hydrogen bonds with the 1′-hydroxy moiety of gigantol, Trp111 formed a π-π conjugation via Van der Waals interactions with the left benzene ring. Trp20 formed a hydrogen bond with the 5′-methoxy moiety (Figure 4). Hydrophobic forces were present between gigantol and the amino acid residues.
 Figure 4.
Figure 4.
Molecular docking simulation reveal the interaction between gigantol and AR. Gigantol interacts with AR at four residues: Trp111, His110, Tyr48, and Trp20. His110 and Tyr48 formed hydrogen bonds with the 1′-hydroxy moiety of gigantol, Trp111 formed a π-π conjugation via Van der Waals interaction with the left benzene ring. Trp20 formed a hydrogen bond with the 5′-methoxy moiety.
To refine and examine the stability of docking complexes obtained from MD, molecular dynamics simulations were performed using the Amber 10 molecular dynamics package. For each complex, independent simulation runs were performed to generate trajectories. Prior to simulations, internal constraints were relaxed by energy minimization. Docking of gigantol onto AR showed changes in the Root Mean Square Deviation (RMSD) over level 2 within 6 ns and that the balanced phase was in the normal range of <3 at 6 ns (Figure 5A). The changes in RMSD with the NADPH cofactor were slightly above level 0.5 within 6 ns and the model remained stable. The RMSD changes of AR conformation were less than 1.5 within 6 ns and remained constant (Figure 5B). After docking, hydrogen bonds formed with RMSD at less than level 2.5 and were stable (Figure 5C). The RMSD changes of AR-His110 due to gigantol were less than level 5 within 6 ns (Figure 5D). The hydrogen bonds between gigantol and the critical amino acids were stable in the active site of AR in 6 ns simulations. Secondary structural analysis was performed to measure the stability of the simulations. The analyses showed that complex secondary structures persisted throughout the simulations. The results also indicated that the MD model with non-covalent binding between gigantol and AR was stable, reliable, and robust.
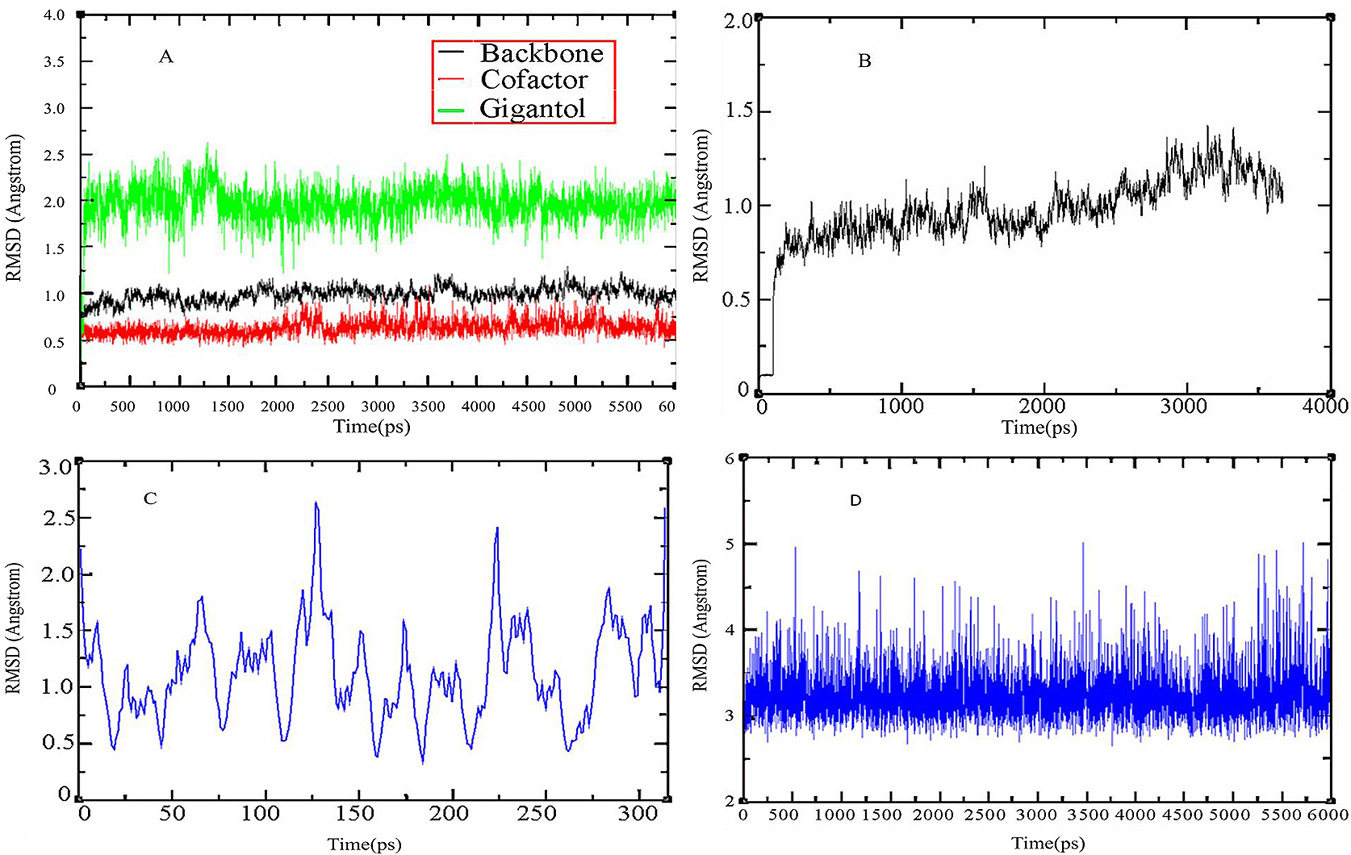 Figure 5.
Figure 5.
Trajectory of the interactions between gigantol and AR. (A) RMSD change of gigantol and NADPH within 6 ns; (B) RMSD change in the AR backbone within 4 ns; (C) RMSD change in the hydrogen bonds of gigantol and amino acid residues; (D) RMSD change of gigantol binding to AR-His110 within 6 ns.
The AR mutants Trp20Ala, Tyr48Ala, His110Ala, and Trp111Ala were generated to test the predicted MD binding sites. WT-AR and its mutants displayed comparable molecular weights of 36,000 Da on SDS PAGE (Figure 6). The concentrations of W20A, Y48A, H110A, and W111A mutants were1.29, 1.77, 1.54, and 2.15 mg/mL, respectively.
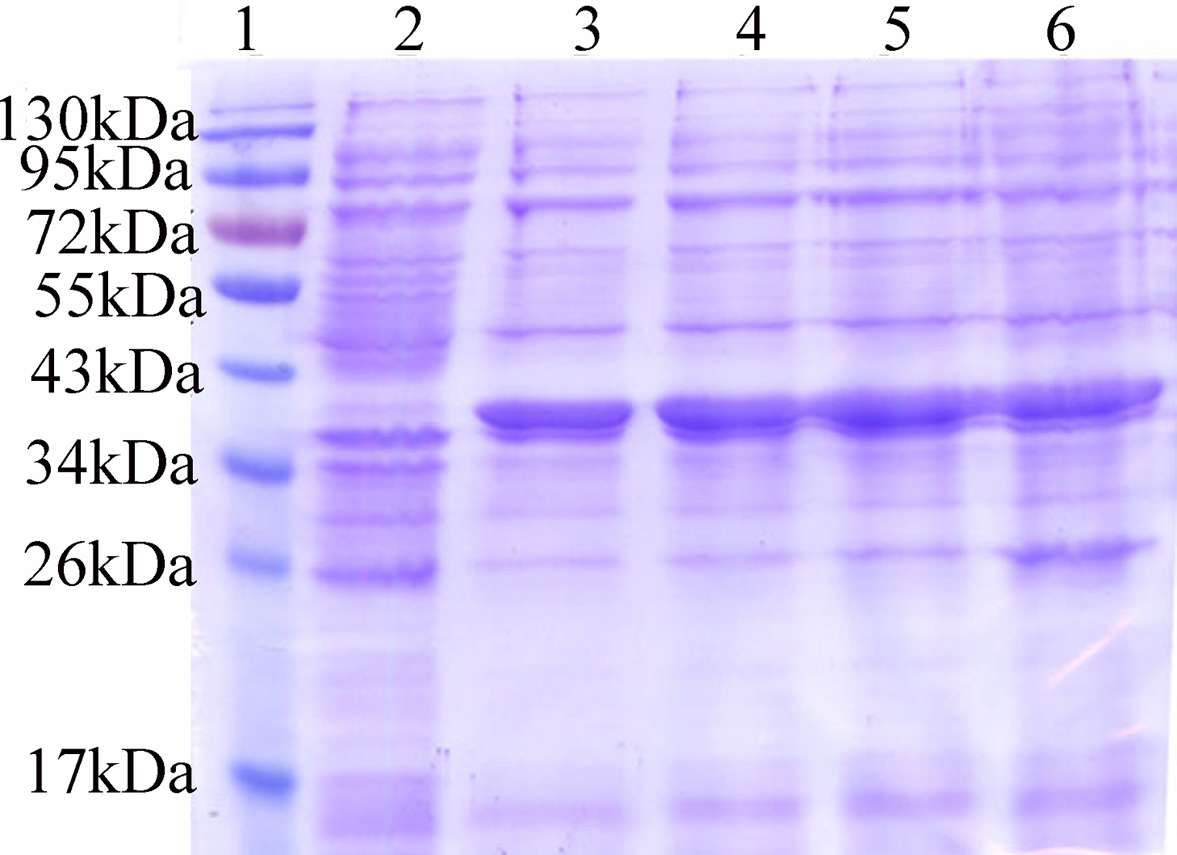 Figure 6.
Figure 6.
Expression of AR and its mutants. BL21 (DE3)-pET28b-AR and mutants were expressed at 18°C following induction by 0.5 mM IPTG. Sample volumes for each well: 8 μg. The MW of AR was ~ 38 kDa after melting with the His marker. (Lane 1:marker; 2:blank control; 3: to 6:W20A, Y48A, H110A and W111A mutants).
The activity of WT-AR and the mutants was assessed according to the rate of NADPH consumption, which was monitored at 340 nm in 10 sec intervals, for 10 mins. After the addition of WT-AR to the reaction system, the absorbance decreased significantly (Figure 7A). The addition of gigantol to the four AR mutants did not lead to a decrease in absorbance (Figure 7B) indicating that they lost the ability to interact with gigantol. Thus, Trp20, Tyr48, His110, and Trp111 are key residues in the interaction of gigantol and AR.
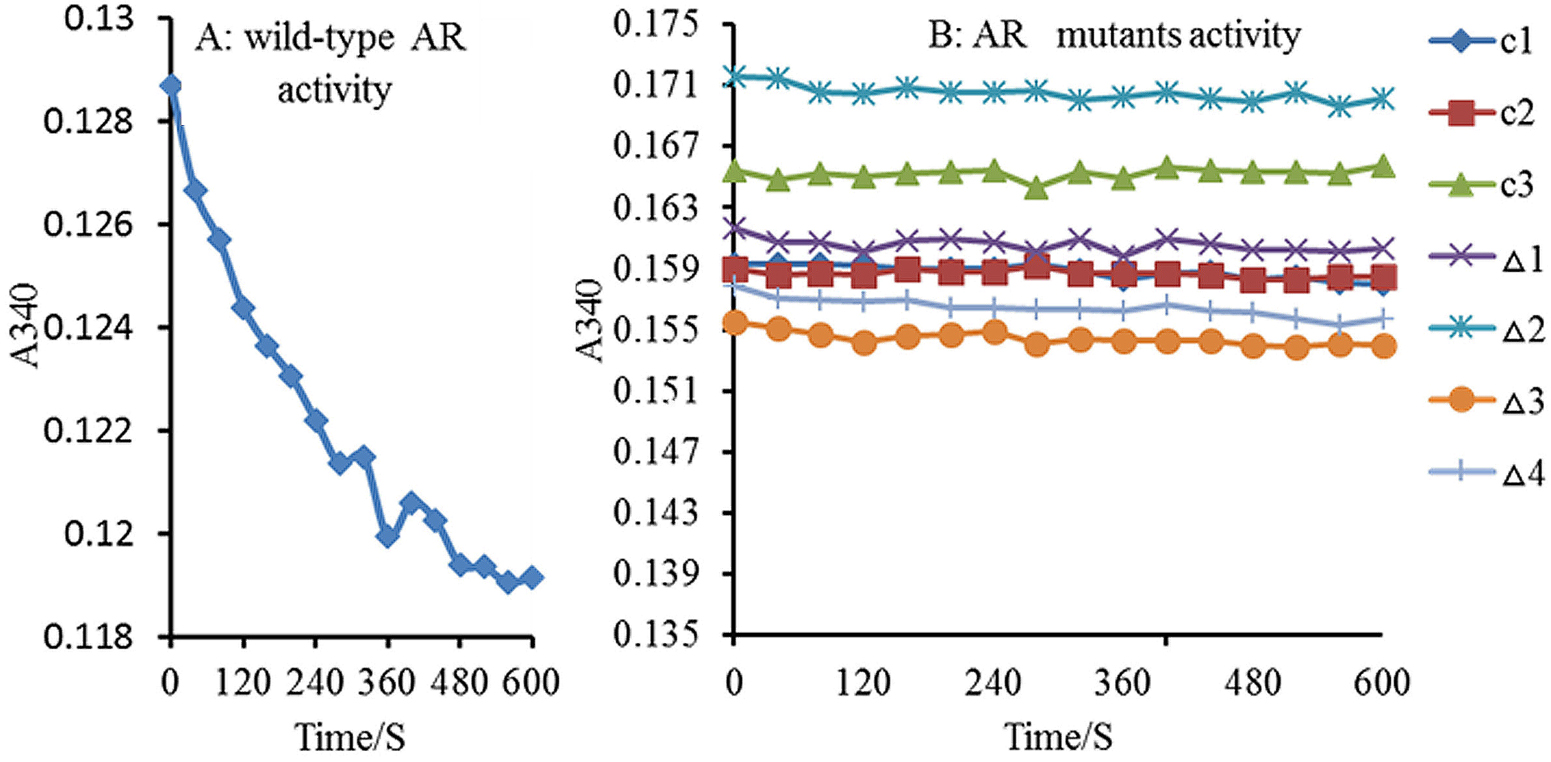 Figure 7.
Figure 7.
Activities of WT AR and its mutants against gigantol. A: Wild-type AR activity; B: AR mutant activity; C1: control (water); C2: control (wash buffer); C3: control (blank vector); Δ1 to Δ4 mutant: W20A, Y48A, H110A and W111A mutants. AR activity was represented by the decline in NADPH absorbance per minute at 340 nm, determined every 10s for 10min. Following AR addition to the reaction system, the OD340 significantly decreased: (A) WT-AR activity with almost no change in OD340. Δ1-Δ4 indicate that the AR mutants were inactive.
Negative ion mass-spectrum scans and deconvolution software were employed to analyze the relative molecular mass of the gigantol-AR complex. For these experiments, gigantol and AR were bound at a series of molar ratios (Figure 8). The relative molecular mass of the gigantol-AR complex increased from ratios 3:1 to 9:1, whilst the larger ratios caused no further increase (Table 2), suggesting that the binding of gigantol to AR is concentration-dependent.
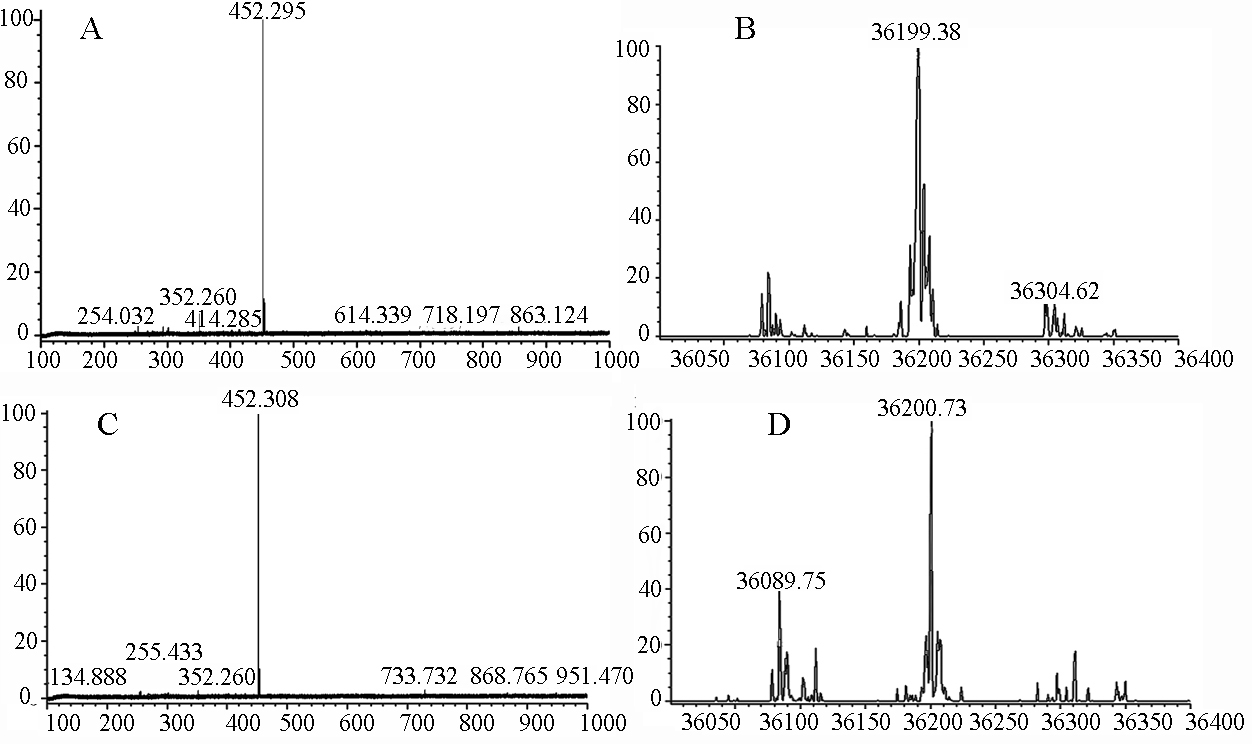 Figure 8.
Figure 8.
CSI-MS spectrum for gigantol and AR. A & B: CSI-MS spectrum and relative molecular mass calculation of AR at 2.8 μmol/L. A: CSI-MS spectrum; B: relative molecular mass calculation. C & D: CSI-MS spectrum and calculated average molecular mass of gigantol/AR at a molar ratio of 27:1. C: CSI-MS spectrum; D: average molecular mass.
| Gigantol (mol)/AR (mol) | Molecular mass of the complex |
|---|---|
| 3:1 | 36198.04±3.25 |
| 9:1 | 36200.06±2.21 |
| 27:1 | 36200.73±2.79 |
| 81:1 | 36200.73±2.59 |
| 243:1 | 36200.06±3.25 |
| 729:1 | 36200.06±2.18 |
| The results are expressed as mean ± SD (n = 5) | |
The maximal chemical constitution stoichiometric ratio (N) calculated using the formula of M (complexes)-M (proteins)/M (reagents) was 1:24.4 for the gigantol-AR complex. The maximum relative MW of the complex was 36200.73 Da (Figure 8D).
The binding of gigantol to AR was not influenced by pH or temperature. Temperatures of 5, 10, 15, and 25°C were used to examine the formation of the gigantol-AR complex at the same molar ratio (27:1) at pH 6.7. The relative molecular mass of the complex in these conditions was 36200.73 Da. When a pH of 6, 6.7, 8, and 9 were assessed at a molar ratio of 27:1 and at 37 °C, the relative MW remained constant.
DC is a major complication that can develop in diabetic patients. Whilst the precise mechanism (s) of its development have not been elucidated, the hyperglycemia-activated polyol pathway has been recognized to contribute to both initiation and progression. AR is an enzyme of the polyol pathway that transforms glucose to D-glucitol, a compound that cannot readily cross cell membranes. Thus, chronic activation of the polyol pathway leads to the accumulation of D-glucitol in the lens, leading to an increased osmotic pressure, and the consequent development of DCs (37, 38). To protect patients with diabetes against polyol pathway-induced cataract development, ARI related drugs are the first line therapy. In theory, ARIs can dock into the AR active site via polar and non-polar interactions (39), leading to non-competitive or mixed competitive inhibition (40). The binding mode of ARI to AR differs from that of glucose or NADPH.
The benefits of ARI to patients with diabetes have been well documented (41, 42). A series of synthetic ARIs, including tolrestat and sorbinil, have been developed, but these have drawbacks including poor permeability and safety issues. Pharmaceutical companies have therefore focused their efforts on the discovery of new, potent, and safe ARIs from natural sources (43-46).
Caulis Dendrobii has been historically used to protect and improve vision in Chinese herbal literature (47). However, the biological mechanism (s) responsible for its beneficial effects have not been elucidated (48). In this study, gigantol, a polyphenolic compound found in Caulis Dendrobii was shown to benefit DC through its ability to bind AR. The interaction between gigantol and AR was also characterized. Mutation of Trp20Ala, Tyr48Ala, His110Ala, and Trp111Ala resulted in a loss of AR binding to gigantol. These data support the hypothesis that these residues are critical for the formation of the gigantol-AR complex, in agreement with MD experiments that identified Tyr48, His110, and Trp111 to be located in the AR active site (24-27, 49). Given the identification of four binding sites between gigantol and AR, we speculate that the interaction is stronger than that of AR-ARI. This stronger binding will enhance the therapeutic efficacy of gigantol in the protection/prevention of DC.
Molecular dynamic simulations have been used to formulate new drugs and characterize interactions between drugs and target molecules. Using this approach, we established the linkages between gigantol and AR. RMSD changes of AR-His110 with gigantol were less than level 5 within 6 ns. We thus concluded that both gigantol-AR complexes remained energetically and structurally stable after acquiring the equilibrium state. These results confirmed that the MD model of the gigantol-AR complex was stable and reliable.
Mass spectrometry, particularly ESI-MS, can characterize the interaction between small molecules and biomacromolecules. However, rigid ionization conditions in ESI-MS limit its application to the characterization of non-covalent interactions between small molecules and proteins. Our study employed low-temperature atomization in CSI-MS to maintain the naturally active conformation of gigantol and AR for characterization of their non-covalent interactions. Using this technology, we identified the maximum interaction ratio of the gigantol-AR complex to be 24.4. We further noted that the MW of the gigantol-AR complex was dependent on the concentration of gigantol when the saturation level was attained. The gigantol and AR interaction was independent of pH and temperature.
In summary, this study adds to previous evidence supporting the protective effects of gigantol on DC in rats. As a potent ARI inhibitor, gigantol occupies the AR active site and stably binds through hydrophobic interactions that mediate its anti-cataract activity. The AR-gigantol interaction is not influenced by pH or temperature, but is influenced by concentration. Whilst the ARIs primarily belong to three major classes, namely acetic acid derivatives, cyclic imides and phenolic derivatives (50), exploring gigantol as a new anti-cataractogenic agent for prevention and/or treatment of DC is warranted in future clinical trials. We anticipate that the active sites of AR identified in this study will provide new insight for the design of novel ARIs with improved efficacy and fewer side effects.
Yong Yang and Qiaohong Yang contributed equally to the project. This work was supported by the grants from the Guangdong High-level university construction project (A1-AFD018171Z11023, A1-AFD018171Z1310), the Natural Science Foundation of Guangdong (2017A030313814), and the National Key R&D Program of China (2017YFC1701100).
Abbreviations: diabetic cataracts (DC), aldose reductase (AR), aldose reductase inhibitors (ARI), AR,Ala substitution at Trp20 (W20A), AR,Ala substitution at Tyr48 (Y48A), AR,Ala substitution at His110 (H110A), AR,Ala substitution at Trp111 (W111A), phosphate buffer saline (PBS), dimethyl sulfoxide (DMSO), molecular docking (MD), protein data bank (PDB), 50% inhibition of enzyme activity (IC50), transferable intermolecular potential 3 points (TIP3P), wild-type (WT), root mean square deviation (RMSD), cold spray ionization mass spetrometry (CSI-MS), Streptozotocin (STZ), Kanamycin (Kan), Dalton (Da), Nanosecond (ns), reduced nicotinamide adenine dinucleotide phosphate (NADPH), three times a day (TID), complementary DNA (cDNA), polymerase chain reaction (PCR), basic local alignment search tool (BLAST), sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), Michaelis-Menten constant (Km), maximum velocity (Vax)
